NR3C1/GLMN-Mediated FKBP12.6 Ubiquitination Disrupts Calcium Homeostasis and Impairs Mitochondrial Quality Control in Stress-Induced Myocardial Damage
Abstract
1. Introduction
2. Results
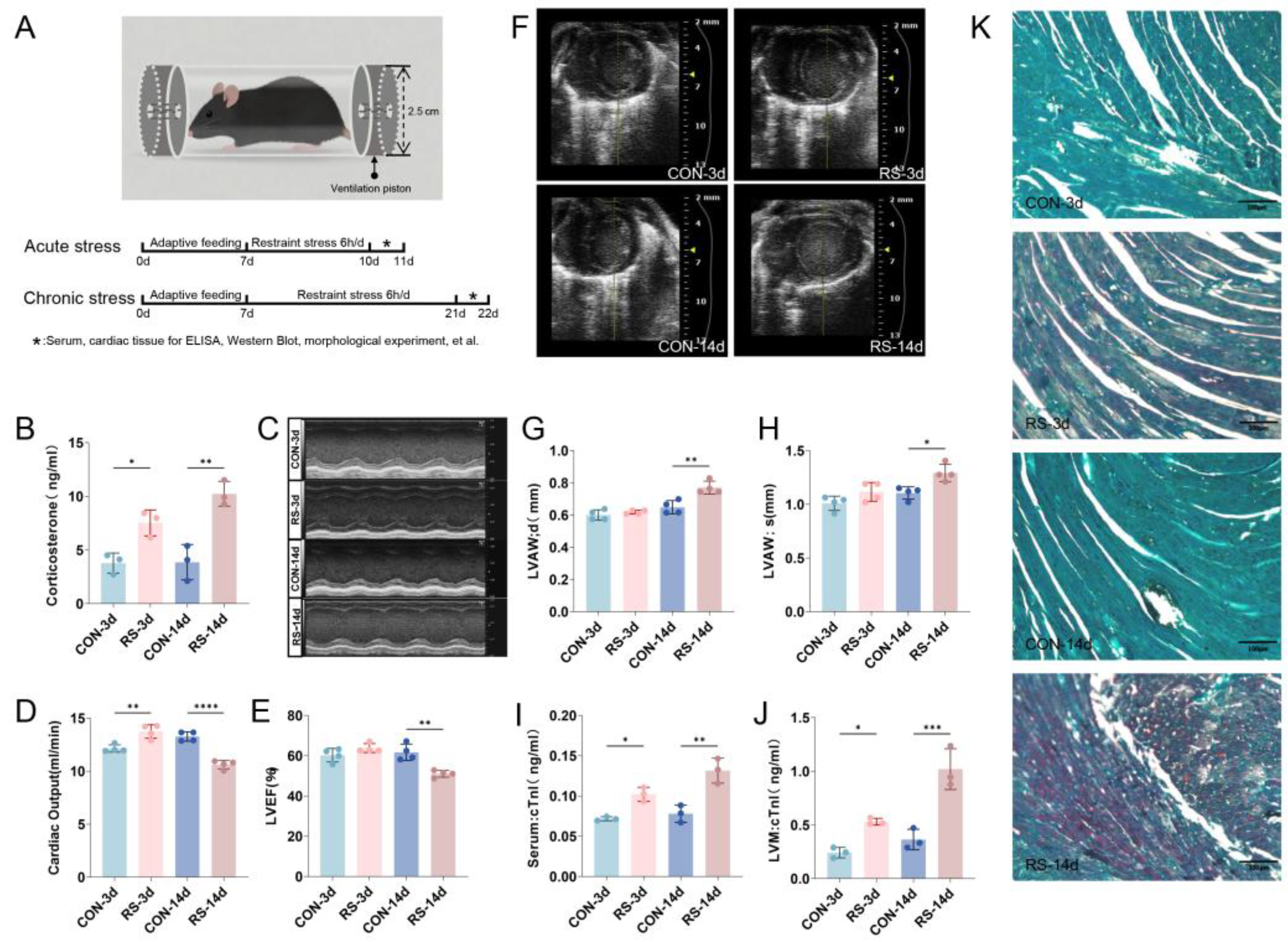
2.1. Stress-Induced Cardiac Dysfunction and Myocardial Cell Damage
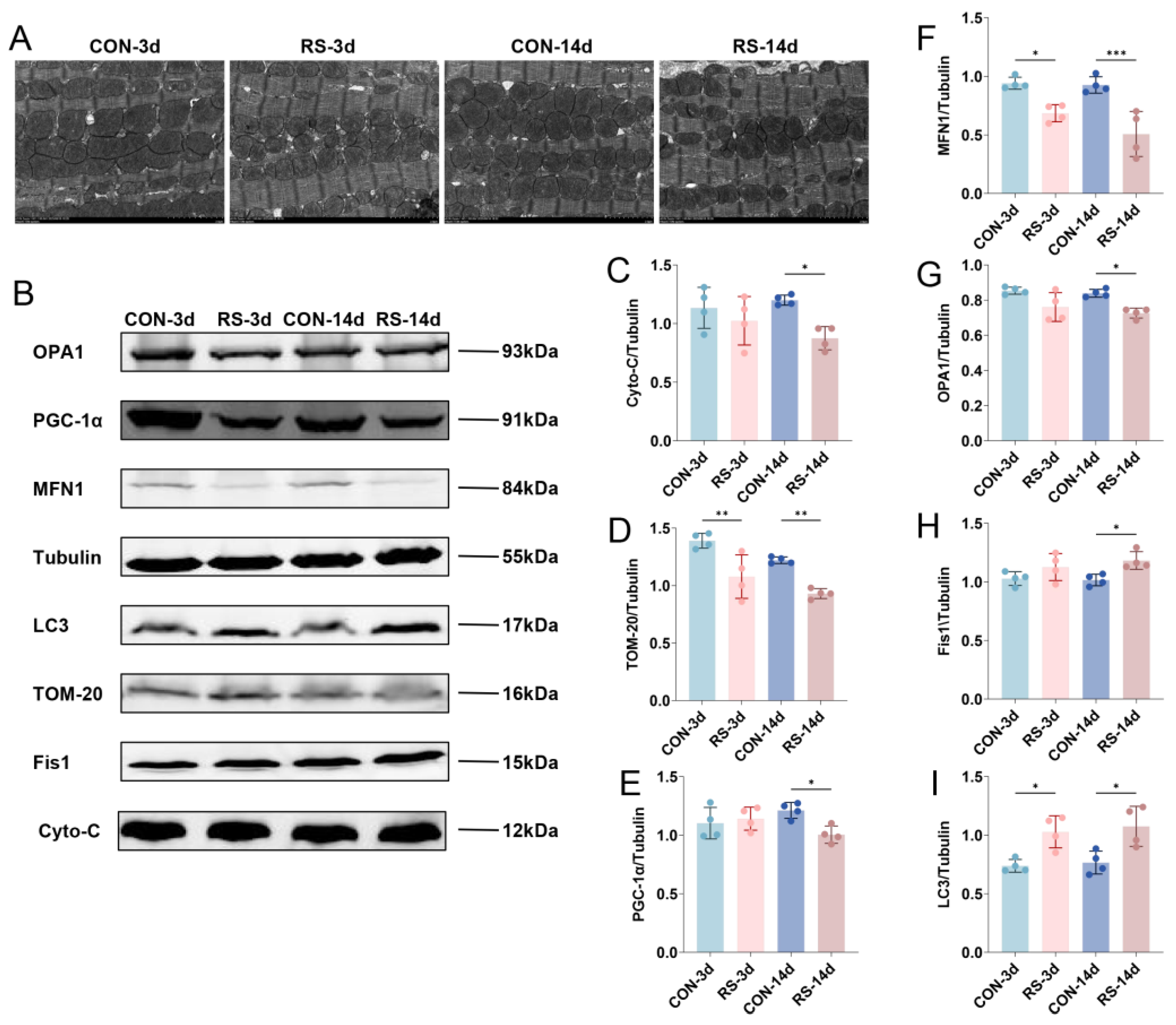
2.2. Stress-Induced Mitochondrial Quality Control Disorder in Cardiomyocytes
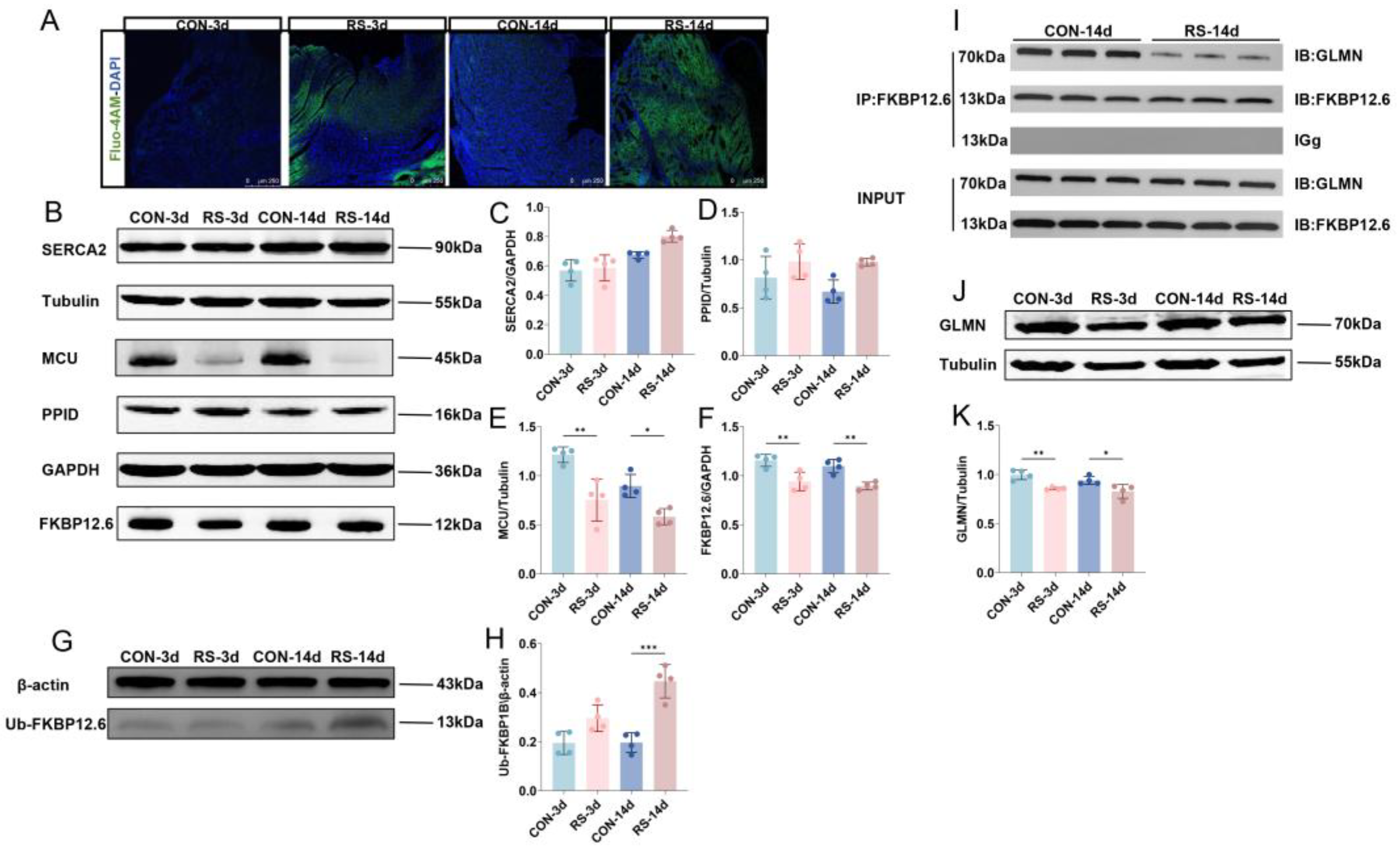
2.3. GLMN-Mediated FKBP12.6 Ubiquitination Induced Intracellular Ca2+ Abnormalities
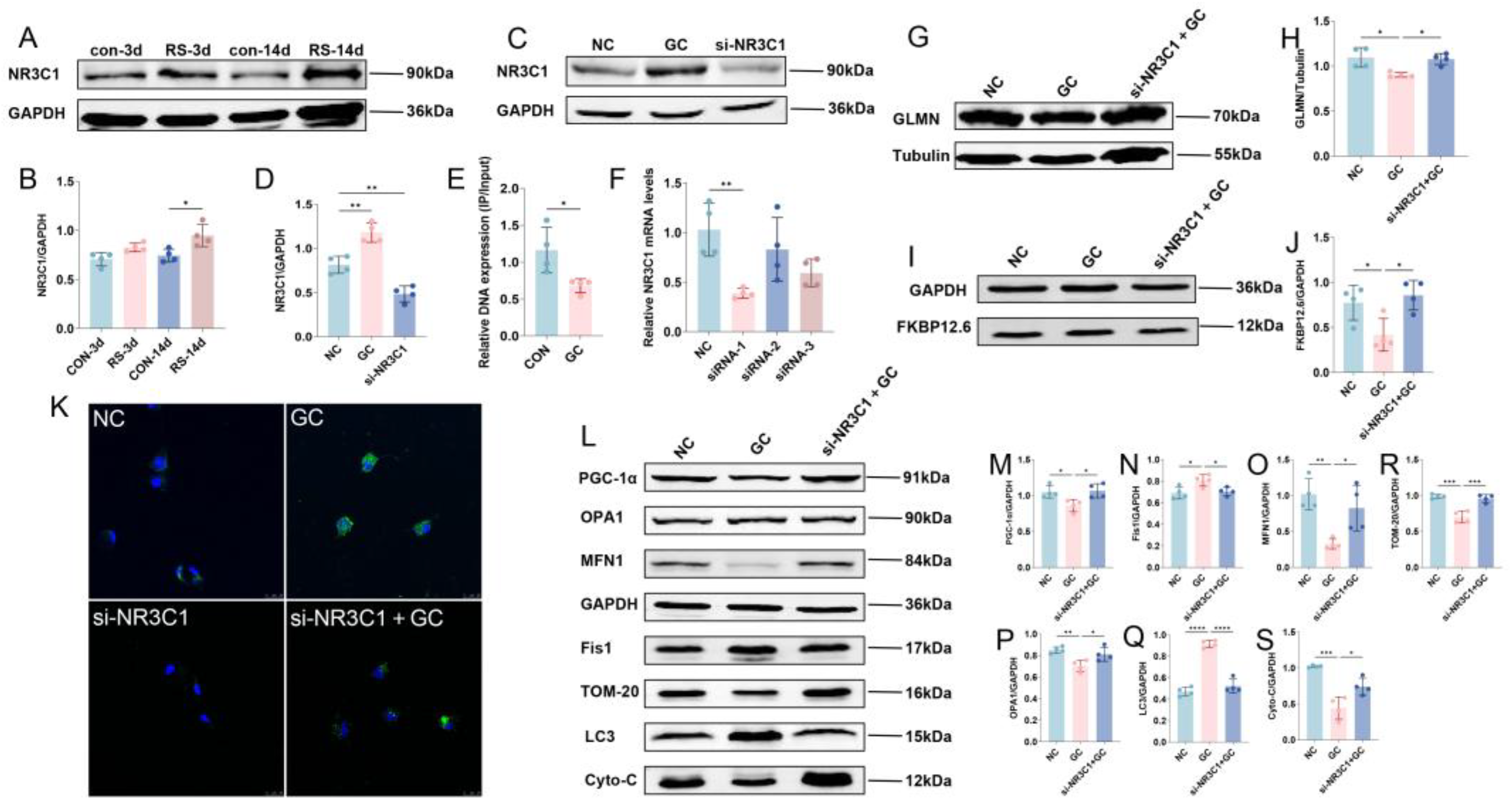
2.4. Stress Inhibits GLMN Transcription-Mediated Ubiquitination of FKBP12.6 Through NR3C1 Inhibition
3. Discussion
4. Materials and Methods
4.1. Stress Model Establishment
4.2. Echocardiographic Analysis of Cardiac Function
4.3. ELISA Detection of cTnI and Corticosterone
4.4. Chromotrope-2R Brilliant Green Staining
4.5. Changes in Mitochondria Morphology
4.6. Western BLOTTING
4.7. Ca2+ Probe
4.8. Co-IP
4.9. ChIP
4.10. RNA Extraction and PCR
4.11. Ubiquitination Assay
4.12. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Neylon, A.; Canniffe, C.; Anand, S.; Kreatsoulas, C.; Blake, G.J.; Sugrue, D.; McGorrian, C. A global perspective on psychosocial risk factors for cardiovascular disease. Prog. Cardiovasc. Dis. 2013, 55, 574–581. [Google Scholar] [CrossRef] [PubMed]
- Osborne, M.T.; Shin, L.M.; Mehta, N.N.; Pitman, R.K.; Fayad, Z.A.; Tawakol, A. Disentangling the Links Between Psychosocial Stress and Cardiovascular Disease. Circ. Cardiovasc. Imaging 2020, 13, e010931. [Google Scholar] [CrossRef]
- Dong, F.; Yin, L.; Sisakian, H.; Hakobyan, T.; Jeong, L.S.; Joshi, H.; Hoff, E.; Chandler, S.; Srivastava, G.; Jabir, A.R.; et al. Takotsubo syndrome is a coronary microvascular disease: Experimental evidence. Eur. Heart J. 2023, 44, 2244–2253. [Google Scholar] [CrossRef] [PubMed]
- Salazar-Ramírez, F.; Ramos-Mondragón, R.; García-Rivas, G. Mitochondrial and Sarcoplasmic Reticulum Interconnection in Cardiac Arrhythmia. Front. Cell Dev. Biol. 2020, 8, 623381. [Google Scholar] [CrossRef]
- Avila, G.; de la Rosa, J.A.; Monsalvo-Villegas, A.; Montiel-Jaen, M.G. Ca2+ Channels Mediate Bidirectional Signaling between Sarcolemma and Sarcoplasmic Reticulum in Muscle Cells. Cells 2019, 9, 55. [Google Scholar] [CrossRef]
- Adamcová, M.; Stĕrba, M.; Simůnek, T.; Potácová, A.; Popelová, O.; Gersl, V. Myocardial regulatory proteins and heart failure. Eur. J. Heart Fail. 2006, 8, 333–342. [Google Scholar] [CrossRef]
- Abriel, H.; Rougier, J.S.; Jalife, J. Ion channel macromolecular complexes in cardiomyocytes: Roles in sudden cardiac death. Circ. Res. 2015, 116, 1971–1988. [Google Scholar] [CrossRef]
- Wu, Q.R.; Zheng, D.L.; Liu, P.M.; Yang, H.; Li, L.A.; Kuang, S.J.; Lai, Y.Y.; Rao, F.; Xue, Y.M.; Lin, J.J.; et al. High glucose induces Drp1-mediated mitochondrial fission via the Orai1 calcium channel to participate in diabetic cardiomyocyte hypertrophy. Cell Death Dis. 2021, 12, 216. [Google Scholar] [CrossRef]
- Hu, Y.; Li, Y.; Li, M.; Zhao, T.; Zhang, W.; Wang, Y.; He, Y.; Zhao, H.; Li, H.; Wang, T.; et al. Calcium supplementation attenuates fluoride-induced bone injury via PINK1/Parkin-mediated mitophagy and mitochondrial apoptosis in mice. J. Hazard. Mater. 2024, 465, 133411. [Google Scholar] [CrossRef]
- Walkon, L.L.; Strubbe-Rivera, J.O.; Bazil, J.N. Calcium Overload and Mitochondrial Metabolism. Biomolecules 2022, 12, 1891. [Google Scholar] [CrossRef]
- Ai, X.; Curran, J.W.; Shannon, T.R.; Bers, D.M.; Pogwizd, S.M. Ca2+/calmodulin-dependent protein kinase modulates cardiac ryanodine receptor phosphorylation and sarcoplasmic reticulum Ca2+ leak in heart failure. Circ. Res. 2005, 97, 1314–1322. [Google Scholar] [CrossRef]
- Miotto, M.C.; Reiken, S.; Wronska, A.; Yuan, Q.; Dridi, H.; Liu, Y.; Weninger, G.; Tchagou, C.; Marks, A.R. Structural basis for ryanodine receptor type 2 leak in heart failure and arrhythmogenic disorders. Nat. Commun. 2024, 15, 8080. [Google Scholar] [CrossRef]
- Dridi, H.; Kushnir, A.; Zalk, R.; Yuan, Q.; Melville, Z.; Marks, A.R. Intracellular calcium leak in heart failure and atrial fibrillation: A unifying mechanism and therapeutic target. Nat. Rev. Cardiol. 2020, 17, 732–747. [Google Scholar] [CrossRef] [PubMed]
- Santulli, G.; Xie, W.; Reiken, S.R.; Marks, A.R. Mitochondrial calcium overload is a key determinant in heart failure. Proc. Natl. Acad. Sci. USA 2015, 112, 11389–11394. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.; Santulli, G.; Reiken, S.R.; Yuan, Q.; Osborne, B.W.; Chen, B.X.; Marks, A.R. Mitochondrial oxidative stress promotes atrial fibrillation. Sci. Rep. 2015, 5, 11427. [Google Scholar] [CrossRef]
- Popovic, D.; Vucic, D.; Dikic, I. Ubiquitination in disease pathogenesis and treatment. Nat. Med. 2014, 20, 1242–1253. [Google Scholar] [CrossRef]
- Varshavsky, A. The Ubiquitin System, Autophagy, and Regulated Protein Degradation. Annu. Rev. Biochem. 2017, 86, 123–128. [Google Scholar] [CrossRef]
- de Kloet, E.R.; Oitzl, M.S.; Joëls, M. Stress and cognition: Are corticosteroids good or bad guys? Trends Neurosci. 1999, 22, 422–426. [Google Scholar] [CrossRef]
- Zhang, S.; Li, Y.; Zhu, W.; Zhang, L.; Lei, L.; Tian, X.; Chen, K.; Shi, W.; Cong, B. Endoplasmic reticulum stress induced by turbulence of mitochondrial fusion and fission was involved in stressed cardiomyocyte injury. J. Cell Mol. Med. 2023, 27, 3313–3325. [Google Scholar] [CrossRef]
- Pernas, L.; Scorrano, L. Mito-Morphosis: Mitochondrial Fusion, Fission, and Cristae Remodeling as Key Mediators of Cellular Function. Annu. Rev. Physiol. 2016, 78, 505–531. [Google Scholar] [CrossRef]
- Archer, S.L. Mitochondrial dynamics--mitochondrial fission and fusion in human diseases. N. Engl. J. Med. 2013, 369, 2236–2251. [Google Scholar] [CrossRef]
- Liu, B.H.; Xu, C.Z.; Liu, Y.; Lu, Z.L.; Fu, T.L.; Li, G.R.; Deng, Y.; Luo, G.Q.; Ding, S.; Li, N.; et al. Mitochondrial quality control in human health and disease. Mil. Med. Res. 2024, 11, 32. [Google Scholar] [CrossRef] [PubMed]
- Chang, X.; Liu, R.; Li, R.; Peng, Y.; Zhu, P.; Zhou, H. Molecular Mechanisms of Mitochondrial Quality Control in Ischemic Cardiomyopathy. Int. J. Biol. Sci. 2023, 19, 426–448. [Google Scholar] [CrossRef]
- Boyman, L.; Karbowski, M.; Lederer, W.J. Regulation of Mitochondrial ATP Production: Ca2+ Signaling and Quality Control. Trends Mol. Med. 2020, 26, 21–39. [Google Scholar] [CrossRef]
- Patra, S.; Mahapatra, K.K.; Praharaj, P.P.; Panigrahi, D.P.; Bhol, C.S.; Mishra, S.R.; Behera, B.P.; Singh, A.; Jena, M.; Bhutia, S.K. Intricate role of mitochondrial calcium signalling in mitochondrial quality control for regulation of cancer cell fate. Mitochondrion 2021, 57, 230–240. [Google Scholar] [CrossRef]
- Ranjbarvaziri, S.; Kooiker, K.B.; Ellenberger, M.; Fajardo, G.; Zhao, M.; Vander Roest, A.S.; Woldeyes, R.A.; Koyano, T.T.; Fong, R.; Ma, N.; et al. Altered Cardiac Energetics and Mitochondrial Dysfunction in Hypertrophic Cardiomyopathy. Circulation 2021, 144, 1714–1731. [Google Scholar] [CrossRef]
- Li, L.; Niemann, B.; Knapp, F.; Werner, S.; Mühlfeld, C.; Schneider, J.P.; Jurida, L.M.; Molenda, N.; Schmitz, M.L.; Yin, X.; et al. Comparison of the stage-dependent mitochondrial changes in response to pressure overload between the diseased right and left ventricle in the rat. Basic. Res. Cardiol. 2024, 119, 587–611. [Google Scholar] [CrossRef]
- Vassalle, M.; Lin, C.I. Calcium overload and cardiac function. J. Biomed. Sci. 2004, 11, 542–565. [Google Scholar] [CrossRef]
- Duong, Q.V.; Hoffman, A.; Zhong, K.; Dessinger, M.J.; Zhang, Y.; Bazil, J.N. Calcium overload decreases net free radical emission in cardiac mitochondria. Mitochondrion 2020, 51, 126–139. [Google Scholar] [CrossRef]
- Fan, M.; Zhang, J.; Tsai, C.W.; Orlando, B.J.; Rodriguez, M.; Xu, Y.; Liao, M.; Tsai, M.F.; Feng, L. Structure and mechanism of the mitochondrial Ca2+ uniporter holocomplex. Nature 2020, 582, 129–133. [Google Scholar] [CrossRef]
- Duda, D.M.; Olszewski, J.L.; Tron, A.E.; Hammel, M.; Lambert, L.J.; Waddell, M.B.; Mittag, T.; DeCaprio, J.A.; Schulman, B.A. Structure of a glomulin-RBX1-CUL1 complex: Inhibition of a RING E3 ligase through masking of its E2-binding surface. Mol. Cell 2012, 47, 371–382. [Google Scholar] [CrossRef]
- Puvar, K.; Luo, Z.Q.; Das, C. Uncovering the Structural Basis of a New Twist in Protein Ubiquitination. Trends Biochem. Sci. 2019, 44, 467–477. [Google Scholar] [CrossRef]
- Kwong, J.Q.; Lu, X.; Correll, R.N.; Schwanekamp, J.A.; Vagnozzi, R.J.; Sargent, M.A.; York, A.J.; Zhang, J.; Bers, D.M.; Molkentin, J.D. The Mitochondrial Calcium Uniporter Selectively Matches Metabolic Output to Acute Contractile Stress in the Heart. Cell Rep. 2015, 12, 15–22. [Google Scholar] [CrossRef]
- Wu, Y.; Rasmussen, T.P.; Koval, O.M.; Joiner, M.L.; Hall, D.D.; Chen, B.; Luczak, E.D.; Wang, Q.; Rokita, A.G.; Wehrens, X.H.; et al. The mitochondrial uniporter controls fight or flight heart rate increases. Nat. Commun. 2015, 6, 6081, Correction in Nat. Commun. 2015, 6, 7241. [Google Scholar] [CrossRef]
- Dong, Z.; Shanmughapriya, S.; Tomar, D.; Siddiqui, N.; Lynch, S.; Nemani, N.; Breves, S.L.; Zhang, X.; Tripathi, A.; Palaniappan, P.; et al. Mitochondrial Ca2+ Uniporter Is a Mitochondrial Luminal Redox Sensor that Augments MCU Channel Activity. Mol. Cell 2017, 65, 1014–1028.e1017. [Google Scholar] [CrossRef]
- Herman, J.P.; McKlveen, J.M.; Ghosal, S.; Kopp, B.; Wulsin, A.; Makinson, R.; Scheimann, J.; Myers, B. Regulation of the Hypothalamic-Pituitary-Adrenocortical Stress Response. Compr. Physiol. 2016, 6, 603–621. [Google Scholar] [CrossRef]
- Poller, W.C.; Downey, J.; Mooslechner, A.A.; Khan, N.; Li, L.; Chan, C.T.; McAlpine, C.S.; Xu, C.; Kahles, F.; He, S.; et al. Brain motor and fear circuits regulate leukocytes during acute stress. Nature 2022, 607, 578–584. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.Y.; Zhu, X.; Tang, H.D.; Huang, J.Y.; Zhu, M.Y.; Cheng, P.K.; Wang, H.; Wang, X.Y.; Wang, H.; Mao, Y.; et al. Brain regulation of gastric dysfunction induced by stress. Nat. Metab. 2023, 5, 1494–1505. [Google Scholar] [CrossRef]
- Rhana, P.; Matsumoto, C.; Fong, Z.; Costa, A.D.; Del Villar, S.G.; Dixon, R.E.; Santana, L.F. Fueling the heartbeat: Dynamic regulation of intracellular ATP during excitation-contraction coupling in ventricular myocytes. Proc. Natl. Acad. Sci. USA 2024, 121, e2318535121. [Google Scholar] [CrossRef] [PubMed]
- Izu, L.T.; Kohl, P.; Boyden, P.A.; Miura, M.; Banyasz, T.; Chiamvimonvat, N.; Trayanova, N.; Bers, D.M.; Chen-Izu, Y. Mechano-electric and mechano-chemo-transduction in cardiomyocytes. J. Physiol. 2020, 598, 1285–1305. [Google Scholar] [CrossRef]
- Zhou, B.; Tian, R. Mitochondrial dysfunction in pathophysiology of heart failure. J. Clin. Investig. 2018, 128, 3716–3726. [Google Scholar] [CrossRef]
- Jencks, W.P. On the mechanism of ATP-driven Ca2+ transport by the calcium ATPase of sarcoplasmic reticulum. Ann. N. Y. Acad. Sci. 1992, 671, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.T.; Valdivia, C.R.; Gurrola, G.B.; Hernández, J.J.; Valdivia, H.H. Arrhythmogenic mechanisms in ryanodine receptor channelopathies. Sci. China Life Sci. 2015, 58, 54–58. [Google Scholar] [CrossRef]
- Do, T.Q.; Knollmann, B.C. Inhibitors of Intracellular RyR2 Calcium Release Channels as Therapeutic Agents in Arrhythmogenic Heart Diseases. Annu. Rev. Pharmacol. Toxicol. 2025, 65, 443–463. [Google Scholar] [CrossRef]
- Tian, X.; Li, Y.; Lei, L.; Feng, X.; Xin, H.; Chen, H.; Zhang, G.; Zuo, M.; Shi, W.; Cong, B. The TF/Nrf2/GSTP1 pathway is involved in stress-induced hepatocellular injury through ferroptosis. J. Cell Mol. Med. 2024, 28, e18494. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cong, J.; Liu, L.; Shi, R.; He, M.; An, Y.; Feng, X.; Yin, X.; Li, Y.; Cong, B.; Shi, W. NR3C1/GLMN-Mediated FKBP12.6 Ubiquitination Disrupts Calcium Homeostasis and Impairs Mitochondrial Quality Control in Stress-Induced Myocardial Damage. Int. J. Mol. Sci. 2025, 26, 8245. https://doi.org/10.3390/ijms26178245
Cong J, Liu L, Shi R, He M, An Y, Feng X, Yin X, Li Y, Cong B, Shi W. NR3C1/GLMN-Mediated FKBP12.6 Ubiquitination Disrupts Calcium Homeostasis and Impairs Mitochondrial Quality Control in Stress-Induced Myocardial Damage. International Journal of Molecular Sciences. 2025; 26(17):8245. https://doi.org/10.3390/ijms26178245
Chicago/Turabian StyleCong, Jingze, Lihui Liu, Rui Shi, Mengting He, Yuchuan An, Xiaowei Feng, Xiaoyu Yin, Yingmin Li, Bin Cong, and Weibo Shi. 2025. "NR3C1/GLMN-Mediated FKBP12.6 Ubiquitination Disrupts Calcium Homeostasis and Impairs Mitochondrial Quality Control in Stress-Induced Myocardial Damage" International Journal of Molecular Sciences 26, no. 17: 8245. https://doi.org/10.3390/ijms26178245
APA StyleCong, J., Liu, L., Shi, R., He, M., An, Y., Feng, X., Yin, X., Li, Y., Cong, B., & Shi, W. (2025). NR3C1/GLMN-Mediated FKBP12.6 Ubiquitination Disrupts Calcium Homeostasis and Impairs Mitochondrial Quality Control in Stress-Induced Myocardial Damage. International Journal of Molecular Sciences, 26(17), 8245. https://doi.org/10.3390/ijms26178245





