Profiling of Breast Cancer Stem Cell Types/States Shows the Role of CD44hi/CD24lo-ALDH1hi as an Independent Prognostic Factor After Neoadjuvant Chemotherapy
Abstract
1. Introduction
2. Results
2.1. Patient Characteristics
2.2. CD44hi/CD24lo CSCs Phenotype Correlates with EMT, Proliferation, and PD-L1 Expression
2.3. Ep-CAMhi Phenotype Correlates with EMT, Proliferation, and Immune Evasion
2.4. CD24 Overexpression Correlates with Markers of Immune Evasion and Higher Proliferation
2.5. ALDH1hi Phenotype Correlates with Higher Proliferation
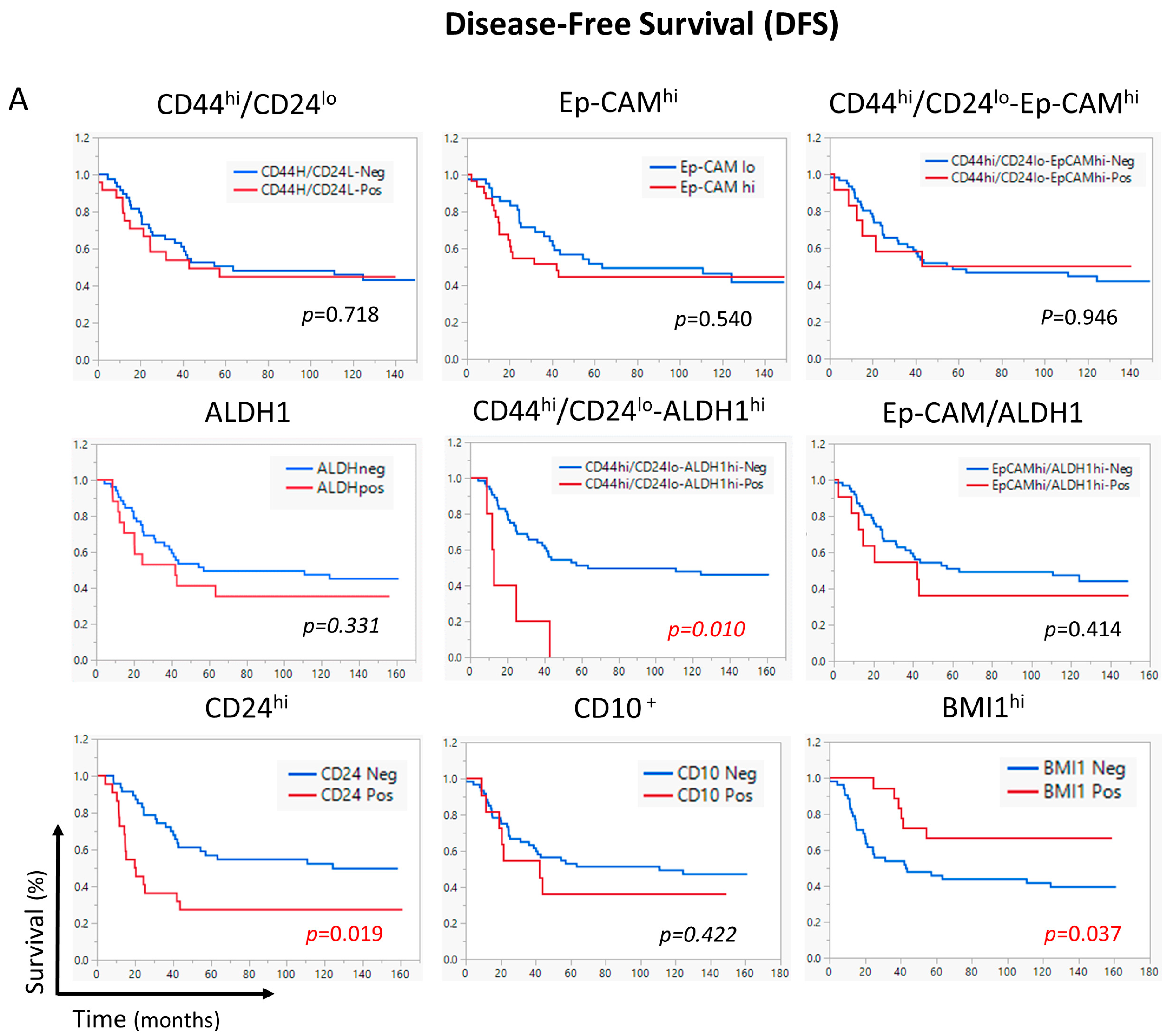
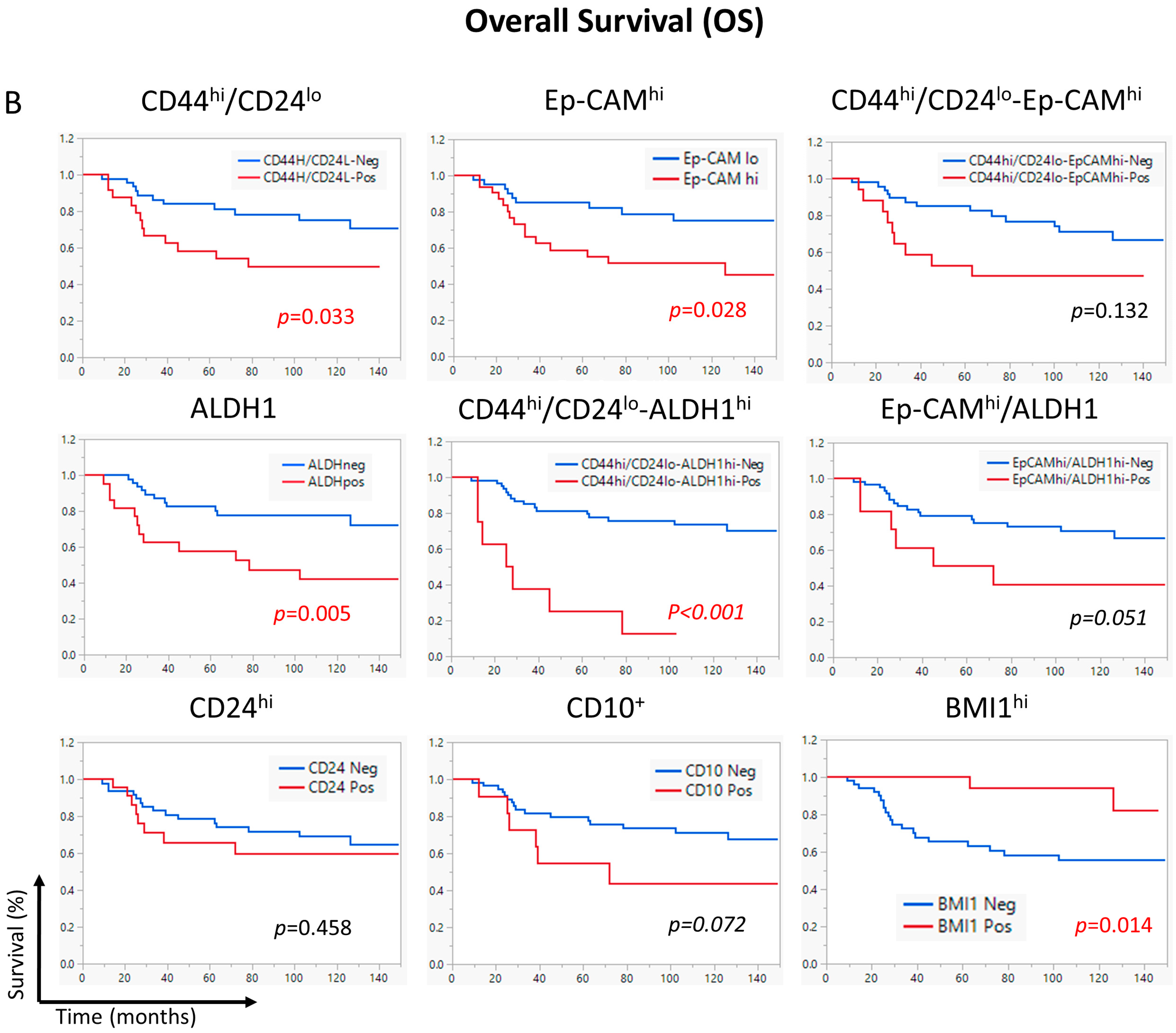
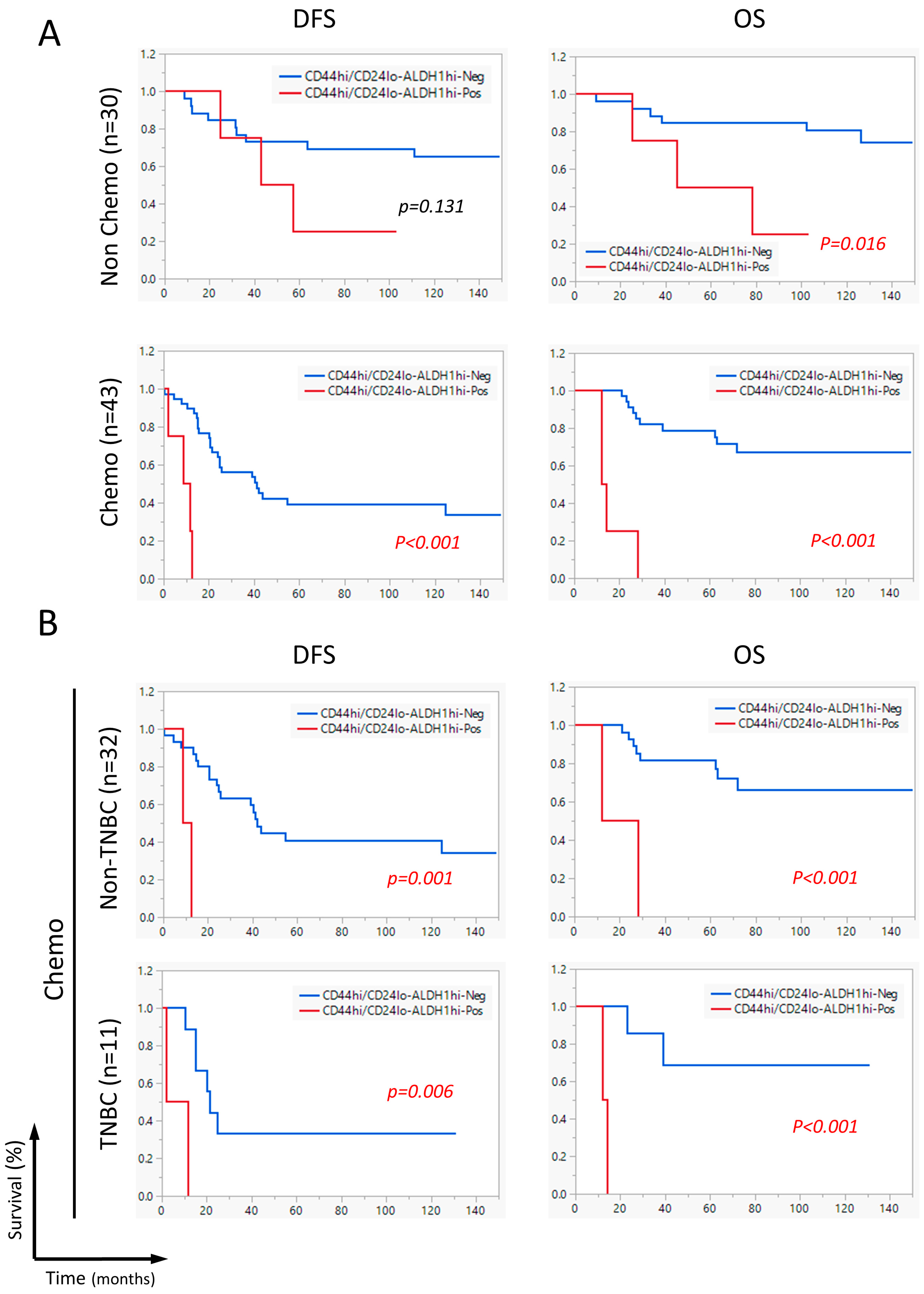
2.6. The CD44hi/CD24lo-ALDHhi Combination Correlated with Shorter DFS and OS in Neoadjuvant Chemotherapy-Receiving Patients
2.7. CD44hi/CD24lo-ALDH1hi Is an Independent Prognostic Factor for Worse Survival
3. Discussion
4. Materials and Methods
4.1. Patients
4.2. Immunohistochemistry
4.3. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| PD-L1 | Programmed death ligand-1 |
| CSCs | Cancer Stem Cells |
| EMT | Epithelial-to-Mesenchymal Transition |
References
- McDermott, S.P.; Wicha, M.S. Targeting breast cancer stem cells. Mol. Oncol. 2010, 4, 404–419. [Google Scholar] [CrossRef]
- Hiraga, T.; Ito, S.; Nakamura, H. EpCAM expression in breast cancer cells is associated with enhanced bone metastasis formation. Int. J. Cancer 2016, 138, 1698–1708. [Google Scholar] [CrossRef]
- Al-Hajj, M.; Wicha, M.S.; Benito-Hernandez, A.; Morrison, S.J.; Clarke, M.F. Prospective identification of tumorigenic breast cancer cells. Proc. Natl. Acad. Sci. USA 2003, 100, 3983–3988. [Google Scholar] [CrossRef]
- Ghebeh, H.; Sleiman, G.M.; Manogaran, P.S.; Al-Mazrou, A.; Barhoush, E.; Al-Mohanna, F.H.; Tulbah, A.; Al-Faqeeh, K.; Adra, C.N. Profiling of normal and malignant breast tissue show CD44high/CD24low phenotype as a predominant stem/progenitor marker when used in combination with Ep-CAM/CD49f markers. BMC Cancer 2013, 13, 289. [Google Scholar] [CrossRef] [PubMed]
- Fillmore, C.M.; Kuperwasser, C. Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast Cancer Res. 2008, 10, R25. [Google Scholar] [CrossRef] [PubMed]
- Maguer-Satta, V.; Besancon, R.; Bachelard-Cascales, E. Concise review: Neutral endopeptidase (CD10): A multifaceted environment actor in stem cells, physiological mechanisms, and cancer. Stem Cells 2011, 29, 389–396. [Google Scholar] [CrossRef] [PubMed]
- Deng, X.; Apple, S.; Zhao, H.; Song, J.; Lee, M.; Luo, W.; Wu, X.; Chung, D.; Pietras, R.J.; Chang, H.R. CD24 Expression and differential resistance to chemotherapy in triple-negative breast cancer. Oncotarget 2017, 8, 38294–38308. [Google Scholar] [CrossRef]
- Srinivasan, M.; Bharali, D.J.; Sudha, T.; Khedr, M.; Guest, I.; Sell, S.; Glinsky, G.V.; Mousa, S.A. Downregulation of Bmi1 in breast cancer stem cells suppresses tumor growth and proliferation. Oncotarget 2017, 8, 38731–38742. [Google Scholar] [CrossRef]
- Liu, S.; Cong, Y.; Wang, D.; Sun, Y.; Deng, L.; Liu, Y.; Martin-Trevino, R.; Shang, L.; McDermott, S.P.; Landis, M.D.; et al. Breast cancer stem cells transition between epithelial and mesenchymal states reflective of their normal counterparts. Stem Cell Rep. 2014, 2, 78–91. [Google Scholar] [CrossRef]
- Warrier, N.M.; Kelkar, N.; Johnson, C.T.; Govindarajan, T.; Prabhu, V.; Kumar, P. Understanding cancer stem cells and plasticity: Towards better therapeutics. Eur. J. Cell Biol. 2023, 102, 151321. [Google Scholar] [CrossRef]
- Butterfield, L.H.; Howard, L.; Kaufman, M.D.F.; Francesco, M.; Marincola, M.D.F. Cancer Immunotherapy Principles and Practice, 2nd ed.; Springer Publishing Company: New York, NY, USA, 2021. [Google Scholar]
- Guha, A.; Goswami, K.K.; Sultana, J.; Ganguly, N.; Choudhury, P.R.; Chakravarti, M.; Bhuniya, A.; Sarkar, A.; Bera, S.; Dhar, S.; et al. Cancer stem cell-immune cell crosstalk in breast tumor microenvironment: A determinant of therapeutic facet. Front. Immunol. 2023, 14, 1245421. [Google Scholar] [CrossRef]
- Almozyan, S.; Colak, D.; Mansour, F.; Alaiya, A.; Al-Harazi, O.; Qattan, A.; Al-Mohanna, F.; Al-Alwan, M.; Ghebeh, H. PD-L1 promotes OCT4 and Nanog expression in breast cancer stem cells by sustaining PI3K/AKT pathway activation. Int. J. Cancer 2017, 141, 1402–1412. [Google Scholar] [CrossRef]
- Mansour, F.A.; Al-Mazrou, A.; Al-Mohanna, F.; Al-Alwan, M.; Ghebeh, H. PD-L1 is overexpressed on breast cancer stem cells through notch3/mTOR axis. Oncoimmunology 2020, 9, 1729299. [Google Scholar] [CrossRef]
- Ponomarev, A.; Gilazieva, Z.; Solovyeva, V.; Allegrucci, C.; Rizvanov, A. Intrinsic and Extrinsic Factors Impacting Cancer Stemness and Tumor Progression. Cancers 2022, 14, 970. [Google Scholar] [CrossRef] [PubMed]
- Polónia, A.; Pinto, R.; Cameselle-Teijeiro, J.F.; Schmitt, F.C.; Paredes, J. Prognostic value of stromal tumour infiltrating lymphocytes and programmed cell death-ligand 1 expression in breast cancer. J. Clin. Pathol. 2017, 70, 860–867. [Google Scholar] [CrossRef]
- Flores, M.L.; Franco, E.H.; Cousido, L.F.S.; Minguito-Carazo, C.; Guadarrama, O.S.; González, L.L.; Pascual, M.E.V.; de la Torre, A.J.M.; Palomo, A.G.; González, A.L. Relationship between Aldehyde Dehydrogenase, PD-L1 and Tumor-Infiltrating Lymphocytes with Pathologic Response and Survival in Breast Cancer. Cancers 2022, 14, 4418. [Google Scholar] [CrossRef]
- Guler, G.; Balci, S.; Costinean, S.; Ussakli, C.H.; Irkkan, C.; Suren, D.; Sari, E.; Altundag, K.; Ozisik, Y.; Jones, S.; et al. Stem cell-related markers in primary breast cancers and associated metastatic lesions. Mod. Pathol. 2012, 25, 949–955. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Song, J.; Jiang, Y.; Yu, C.; Ma, Z. Predictive value of CD44 and CD24 for prognosis and chemotherapy response in invasive breast ductal carcinoma. Int. J. Clin. Exp. Pathol. 2015, 8, 11287–11295. [Google Scholar]
- Lee, H.E.; Kim, J.H.; Kim, Y.J.; Choi, S.Y.; Kim, S.W.; Kang, E.; Chung, I.Y.; Kim, I.A.; Kim, E.J.; Choi, Y.; et al. An increase in cancer stem cell population after primary systemic therapy is a poor prognostic factor in breast cancer. Br. J. Cancer 2011, 104, 1730–1738. [Google Scholar] [CrossRef]
- Ali, H.R.; Dawson, S.J.; Blows, F.M.; Provenzano, E.; Pharoah, P.D.; Caldas, C. Cancer stem cell markers in breast cancer: Pathological, clinical and prognostic significance. Breast Cancer Res. 2011, 13, R118. [Google Scholar] [CrossRef] [PubMed]
- Ricardo, S.; Vieira, A.F.; Gerhard, R.; Leitao, D.; Pinto, R.; Cameselle-Teijeiro, J.F.; Milanezi, F.; Schmitt, F.; Paredes, J. Breast cancer stem cell markers CD44, CD24 and ALDH1: Expression distribution within intrinsic molecular subtype. J. Clin. Pathol. 2011, 64, 937–946. [Google Scholar] [CrossRef]
- Mani, S.A.; Guo, W.; Liao, M.J.; Eaton, E.N.; Ayyanan, A.; Zhou, A.Y.; Brooks, M.; Reinhard, F.; Zhang, C.C.; Shipitsin, M.; et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008, 133, 704–715. [Google Scholar] [CrossRef]
- Blick, T.; Hugo, H.; Widodo, E.; Waltham, M.; Pinto, C.; Mani, S.A.; Weinberg, R.A.; Neve, R.M.; Lenburg, M.E.; Thompson, E.W. Epithelial mesenchymal transition traits in human breast cancer cell lines parallel the CD44(hi/)CD24 (lo/-) stem cell phenotype in human breast cancer. J. Mammary Gland. Biol. Neoplasia 2010, 15, 235–252. [Google Scholar] [CrossRef]
- May, C.D.; Sphyris, N.; Evans, K.W.; Werden, S.J.; Guo, W.; Mani, S.A. Epithelial-mesenchymal transition and cancer stem cells: A dangerously dynamic duo in breast cancer progression. Breast Cancer Res. 2011, 13, 202. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.S.; Tang, H.T.; Xiao, Q.H.; He, M.; Zhao, L.; Fu, Y.Z.; Wu, H.Z.; Yu, Z.J.; Jiang, Q.; Yan, Y.Y.; et al. The Hedgehog signaling pathway is associated with poor prognosis in breast cancer patients with the CD44/CD24 phenotype. Mol. Med. Rep. 2016, 14, 5261–5270. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, Q.Q.; Wang, Q.; Wang, Y.P.; Chen, J. Prognostic significance of CD24 and CD44 in breast cancer: A meta-analysis. Int. J. Biol. Marker 2017, 32, E75–E82. [Google Scholar] [CrossRef] [PubMed]
- Kristiansen, G.; Machado, E.; Bretz, N.; Rupp, C.; Winzer, K.J.; Konig, A.K.; Moldenhauer, G.; Marme, F.; Costa, J.; Altevogt, P. Molecular and clinical dissection of CD24 antibody specificity by a comprehensive comparative analysis. Lab. Investig. 2010, 90, 1102–1116. [Google Scholar] [CrossRef]
- Abd El-Maqsoud, N.M.; Abd El-Rehim, D.M. Clinicopathologic implications of EpCAM and Sox2 expression in breast cancer. Clin. Breast Cancer 2014, 14, e1–e9. [Google Scholar] [CrossRef]
- Agboola, A.J.; Paish, E.C.; Rakha, E.A.; Powe, D.G.; Macmillan, R.D.; Ellis, I.O.; Green, A.R. EpCAM expression is an indicator of recurrence in basal-like breast cancer. Breast Cancer Res. Tr. 2012, 133, 575–582. [Google Scholar] [CrossRef]
- Soysal, S.D.; Muenst, S.; Barbie, T.; Fleming, T.; Gao, F.; Spizzo, G.; Oertli, D.; Viehl, C.T.; Obermann, E.C.; Gillanders, W.E. EpCAM expression varies significantly and is differentially associated with prognosis in the luminal B HER2+, basal-like, and HER2 intrinsic subtypes of breast cancer. Br. J. Surg. 2013, 100, 11–12. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.N.; Liang, K.H.; Lai, J.K.; Lan, C.H.; Liao, M.Y.; Hung, S.H.; Chuang, Y.T.; Chen, K.C.; Tsuei, W.W.F.; Wu, H.C. EpCAM Signaling Promotes Tumor Progression and Protein Stability of PD-L1 through the EGFR Pathway. Cancer Res. 2020, 80, 5035–5050. [Google Scholar] [CrossRef]
- Zheng, X.; Fan, X.; Fu, B.; Zheng, M.; Zhang, A.; Zhong, K.; Yan, J.; Sun, R.; Tian, Z.; Wei, H. EpCAM Inhibition Sensitizes Chemoresistant Leukemia to Immune Surveillance. Cancer Res. 2017, 77, 482–493. [Google Scholar] [CrossRef]
- Spizzo, G.; Went, P.; Dirnhofer, S.; Obrist, P.; Simon, R.; Spichtin, H.; Maurer, R.; Metzger, U.; von Castelberg, B.; Bart, R.; et al. High Ep-CAM expression is associated with poor prognosis in node-positive breast cancer. Breast Cancer Res. Treat. 2004, 86, 207–213. [Google Scholar] [CrossRef] [PubMed]
- Gastl, G.; Spizzo, G.; Obrist, P.; Dünser, M.; Mikuz, G. Ep-CAM overexpression in breast cancer as a predictor of survival. Lancet 2000, 356, 1981–1982. [Google Scholar] [CrossRef]
- Barkal, A.A.; Brewer, R.E.; Markovic, M.; Kowarsky, M.; Barkal, S.A.; Zaro, B.W.; Krishnan, V.; Hatakeyama, J.; Dorigo, O.; Barkal, L.J.; et al. CD24 signalling through macrophage Siglec-10 is a target for cancer immunotherapy. Nature 2019, 572, 392–396. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Lu, X.M.; Tao, K.X.; Shi, L.; Li, W.; Wang, G.B.; Wu, K. Siglec-10 is associated with survival and natural killer cell dysfunction in hepatocellular carcinoma. J. Surg. Res. 2015, 194, 107–113. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Chen, J.; Ge, C.; Zhao, F.; Chen, T.; Tian, H.; Li, J.; Li, H. CD24 isoform a promotes cell proliferation, migration and invasion and is downregulated by EGR1 in hepatocellular carcinoma. OncoTargets Ther. 2019, 12, 1705–1716. [Google Scholar] [CrossRef]
- Smith, S.C.; Oxford, G.; Wu, Z.; Nitz, M.D.; Conaway, M.; Frierson, H.F.; Hampton, G.; Theodorescu, D. The metastasis-associated gene CD24 is regulated by Ral GTPase and is a mediator of cell proliferation and survival in human cancer. Cancer Res. 2006, 66, 1917–1922. [Google Scholar] [CrossRef]
- Lee, K.M.; Ju, J.H.; Jang, K.; Yang, W.; Yi, J.Y.; Noh, D.Y.; Shin, I. CD24 regulates cell proliferation and transforming growth factor beta-induced epithelial to mesenchymal transition through modulation of integrin beta1 stability. Cell Signal 2012, 24, 2132–2142. [Google Scholar] [CrossRef]
- Kaipparettu, B.A.; Malik, S.; Konduri, S.D.; Liu, W.; Rokavec, M.; van der Kuip, H.; Hoppe, R.; Hammerich-Hille, S.; Fritz, P.; Schroth, W.; et al. Estrogen-mediated downregulation of CD24 in breast cancer cells. Int. J. Cancer 2008, 123, 66–72, Erratum in Int. J. Cancer 2019, 145, E21. [Google Scholar] [CrossRef]
- Khoury, T.; Ademuyiwa, F.O.; Chandrasekhar, R.; Jabbour, M.; Deleo, A.; Ferrone, S.; Wang, Y.; Wang, X. Aldehyde dehydrogenase 1A1 expression in breast cancer is associated with stage, triple negativity, and outcome to neoadjuvant chemotherapy. Mod. Pathol. 2012, 25, 388–397, Erratum in Mod. Pathol. 2012, 25, 917. [Google Scholar] [CrossRef]
- Li, J.; Zhang, B.; Yang, Y.F.; Jin, J.; Liu, Y.H. Aldehyde dehydrogenase 1 as a predictor of the neoadjuvant chemotherapy response in breast cancer: A meta-analysis. Medicine 2018, 97, e12056. [Google Scholar] [CrossRef]
- Seo, A.N.; Lee, H.J.; Kim, E.J.; Kim, H.J.; Jang, M.H.; Lee, H.E.; Kim, Y.J.; Kim, J.H.; Park, S.Y. Tumour-infiltrating CD8+lymphocytes as an independent predictive factor for pathological complete response to primary systemic therapy in breast cancer. Br. J. Cancer 2013, 109, 2705–2713. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Lv, D.L.; Duan, J.J.; Xu, S.L.; Zhang, J.F.; Yang, X.J.; Zhang, X.; Cui, Y.H.; Bian, X.W.; Yu, S.C. ALDH1A1 expression correlates with clinicopathologic features and poor prognosis of breast cancer patients: A systematic review and meta-analysis. BMC Cancer 2014, 14, 444. [Google Scholar] [CrossRef] [PubMed]
- Peng, D.; Tanikawa, T.; Li, W.; Zhao, L.; Vatan, L.; Szeliga, W.; Wan, S.; Wei, S.; Wang, Y.; Liu, Y.; et al. Myeloid-Derived Suppressor Cells Endow Stem-like Qualities to Breast Cancer Cells through IL6/STAT3 and NO/NOTCH Cross-talk Signaling. Cancer Res. 2016, 76, 3156–3165. [Google Scholar] [CrossRef] [PubMed]
- Louhichi, T.; Saad, H.; Ben Dhiab, M.; Ziadi, S.; Trimeche, M. Stromal CD10 expression in breast cancer correlates with tumor invasion and cancer stem cell phenotype. BMC Cancer 2018, 18, 49. [Google Scholar] [CrossRef]
- Lee, Y.C.; Chang, W.W.; Chen, Y.Y.; Tsai, Y.H.; Chou, Y.H.; Tseng, H.C.; Chen, H.L.; Wu, C.C.; Chang-Chien, J.; Lee, H.T.; et al. Hsp90α Mediates BMI1 Expression in Breast Cancer Stem/Progenitor Cells through Facilitating Nuclear Translocation of c-Myc and EZH2. Int. J. Mol. Sci. 2017, 18, 1986. [Google Scholar] [CrossRef]
- Choi, Y.J.; Choi, Y.L.; Cho, E.Y.; Shin, Y.K.; Sung, K.W.; Hwang, Y.K.; Lee, S.J.; Kong, G.; Lee, J.E.; Kim, J.S.; et al. Expression of Bmi-1 protein in tumor tissues is associated with favorable prognosis in breast cancer patients. Breast Cancer Res. Tr. 2009, 113, 83–93. [Google Scholar] [CrossRef]
- Wang, Y.Y.; Zhe, H.; Ding, Z.; Gao, P.; Zhang, N.M.; Li, G.F. Cancer Stem Cell Marker Bmi-1 Expression is Associated with Basal-like Phenotype and Poor Survival in Breast Cancer. World J. Surg. 2012, 36, 1189–1194. [Google Scholar] [CrossRef]
- Althobiti, M.; Muftah, A.A.; Aleskandarany, M.A.; Joseph, C.; Toss, M.S.; Green, A.; Rakha, E. The prognostic significance of BMI1 expression in invasive breast cancer is dependent on its molecular subtypes. Breast Cancer Res. Tr. 2020, 182, 581–589. [Google Scholar] [CrossRef]
- Alsuliman, A.; Colak, D.; Al-Harazi, O.; Fitwi, H.; Tulbah, A.; Al-Tweigeri, T.; Al-Alwan, M.; Ghebeh, H. Bidirectional crosstalk between PD-L1 expression and epithelial to mesenchymal transition: Significance in claudin-low breast cancer cells. Mol. Cancer 2015, 14, 149. [Google Scholar] [CrossRef]
- Ghebeh, H.; Tulbah, A.; Mohammed, S.; Elkum, N.; Bin Amer, S.M.; Al-Tweigeri, T.; Dermime, S. Expression of B7-H1 in breast cancer patients is strongly associated with high proliferative Ki-67-expressing tumor cells. Int. J. Cancer 2007, 121, 751–758. [Google Scholar] [CrossRef] [PubMed]
- Ghebeh, H.; Mansour, F.A.; Colak, D.; Alfuraydi, A.A.; Al-Thubiti, A.A.; Monies, D.; Al-Alwan, M.; Al-Tweigeri, T.; Tulbah, A. Higher PD-L1 Immunohistochemical Detection Signal in Frozen Compared to Matched Paraffin-Embedded Formalin-Fixed Tissues. Antibodies 2021, 10, 24. [Google Scholar] [CrossRef] [PubMed]
- Elfoly, M.; Mirza, J.Y.; Alaiya, A.; Al-Hazzani, A.A.; Tulbah, A.; Al-Alwan, M.; Ghebeh, H. PD-L1 intrinsically promotes the proliferation of breast cancer cells through the SKP2-p27/p21 axis. Cancer Cell Int. 2024, 24, 161. [Google Scholar] [CrossRef] [PubMed]
- Ghebeh, H.; Barhoush, E.; Tulbah, A.; Elkum, N.; Al-Tweigeri, T.; Dermime, S. FOXP3 Tregs and B7-HI+/PD-I+T lymphocytes co-infiltrate the tumor tissues of high-risk breast cancer patients:: Implication for immunotherapy. BMC Cancer 2008, 8, 57. [Google Scholar] [CrossRef]


| (A) | ||||||||||
| CD44hi/CD24lo | Ep-CAMhi | CD44hi/CD24lo-Ep-CAMhi | ||||||||
| <10% | ≥10% | ♣ p | H-Score < 200 | H-Score ≥ 200 | ♣ p | − | + | ♣ p | ||
| Immune Markers | PD-L1 | |||||||||
| <5% | 45 (76) * | 14 (24) | 0.001 | 38 (64) | 21 (36) | 0.019 | 53 (90) | 6 (10) | 0.008 | |
| ≥5% | 4 (29) | 10 (71) | 4 (29) | 10 (71) | 8 (57) | 6 (43) | ||||
| TIL | ||||||||||
| Low (Score 1 and 2) | 40 (70) | 17 (30) | 0.369 | 37 (65) | 20 (35) | 0.024 | 50 (88) | 7 (12) | 0.120 | |
| High (Score 3) | 9 (56) | 7 (44) | 5 (31) | 11 (69) | 11 (69) | 5 (31) | ||||
| § FOXP3 + TIL | ||||||||||
| <10% | 27 (73) | 10 (27) | 0.784 | 26 (70) | 11 (30) | 0.012 | 33 (89) | 4 (11) | 0.483 | |
| ≥10% | 19 (68) | 9 (32) | 11 (39) | 17 (61) | 23 (82) | 5 (18) | ||||
| § PD-1 + TIL | ||||||||||
| <10% | 24 (75) | 8 (25) | 0.314 | 23 (72) | 9 (28) | 0.052 | 30 (94) | 2 (6) | 0.097 | |
| ≥10% | 24 (63) | 14 (37) | 18 (47) | 20 (53) | 30 (79) | 8 (21) | ||||
| Proliferation Markers | Ki-67 | |||||||||
| <20% | 29 (81) | 7 (19) | 0.024 | 26 (72) | 10 (28) | 0.018 | 34 (94) | 2 (6) | 0.024 | |
| ≥20% | 20 (54) | 17 (46) | 16 (43) | 21 (57) | 27 (73) | 10 (27) | ||||
| § SKP2 | ||||||||||
| <10% | 35 (76) | 11 (24) | 0.062 | 29 (63) | 17 (37) | 0.314 | 40 (87) | 6 (13) | 0.322 | |
| ≥10% | 13 (52) | 12 (48) | 12 (48) | 13 (52) | 19 (76) | 6 (24) | ||||
| § p21 | ||||||||||
| <10% | 28 (65) | 15 (35) | 0.615 | 22 (51) | 21 (49) | 0.221 | 34 (79) | 9 (21) | 0.341 | |
| ≥10% | 20 (71) | 8 (29) | 19 (68) | 9 (32) | 25 (89) | 3 (11) | ||||
| § p27 | ||||||||||
| <50% | 33 (65) | 18 (35) | 0.574 | 25 (49) | 26 (51) | 0.031 | 41 (80) | 10 (20) | 0.488 | |
| ≥50% | 15 (75) | 5 (25) | 16 (80) | 4 (20) | 18 (90) | 2 (10) | ||||
| § Loss of both p21/p27 | ||||||||||
| NO | 17 (55) | 14 (45) | 0.072 | 28 (70) | 12 (30) | 0.029 | 23 (74) | 8 (26) | 0.112 | |
| YES | 31 (78) | 9 (23) | 13 (42) | 18 (58) | 36 (90) | 4 (10) | ||||
| (B) | ||||||||||
| CD44hi CD24lo | Ep-CAMhi | CD44hi CD24lo/Ep-CAMhi | ||||||||
| <10% | ≥10% | ♣ p | H-Score < 200 | H-Score ≥ 200 | ♣ p | − | + | ♣ p | ||
| Clinicopathological Markers | Age | |||||||||
| <40 years | 14 (58) * | 10 (42) | 0.297 | 13 (54) | 11 (46) | 0.802 | 17 (71) | 7 (29) | 0.051 | |
| ≥40 years | 35 (71) | 14 (29) | 29 (59) | 20 (41) | 44 (90) | 5 (10) | ||||
| Tumor Size | ||||||||||
| <4 cm | 30 (77) | 9 (23) | 0.081 | 27 (69) | 12 (31) | 0.036 | 34 (87) | 5 (13) | 0.529 | |
| ≥4 cm | 19 (56) | 15 (44) | 15 (44) | 19 (56) | 27 (79) | 7 (21) | ||||
| Invasion | ||||||||||
| Absent | 21 (70) | 9 (30) | 0.801 | 16 (53) | 14 (47) | 0.633 | 25 (83) | 5 (17) | 1.000 | |
| Present | 28 (65) | 15 (35) | 26 (60) | 17 (40) | 36 (84) | 7 (16) | ||||
| Histological Grade | ||||||||||
| 1 and 2 | 30 (81) | 7 (19) | 0.013 | 25 (68) | 12 (32) | 0.100 | 34 (92) | 3 (8) | 0.064 | |
| 3 | 19 (53) | 17 (47) | 17 (47) | 19 (53) | 27 (75) | 9 (25) | ||||
| § Lymph Node Metastasis | ||||||||||
| Absent | 18 (69) | 8 (31) | 1.000 | 14 (54) | 12 (46) | 0.810 | 20 (77) | 6 (23) | 0.331 | |
| Present | 31 (67) | 15 (33) | 27 (59) | 19 (41) | 40 (87) | 6 (13) | ||||
| Neoadjuvant Chemotherapy | ||||||||||
| Absent | 21 (70) | 9 (30) | 0.801 | 19 (63) | 11 (37) | 0.476 | 27 (90) | 3 (10) | 0.337 | |
| Present | 28 (65) | 15 (35) | 23 (54) | 20 (47) | 34 (79) | 9 (21) | ||||
| BC Subtype Markers | ER Status | |||||||||
| Negative | 12 (44) | 15 (56) | 0.002 | 10 (37) | 17 (63) | 0.008 | 18 (67) | 9 (33) | 0.007 | |
| Positive | 37 (80) | 9 (20) | 32 (70) | 14 (30) | 43 (93) | 3 (7) | ||||
| PR Status | ||||||||||
| Negative | 23 (58) | 17 (42) | 0.080 | 17 (42) | 23 (58) | 0.005 | 29 (72) | 11 (28) | 0.009 | |
| Positive | 26 (79) | 7 (21) | 25 (76) | 8 (24) | 32 (97) | 1 (3) | ||||
| HER2/neu Status | ||||||||||
| Negative | 33 (67) | 16 (33) | 1.000 | 29 (59) | 20 (41) | 0.802 | 40 (82) | 9 (18) | 0.739 | |
| Positive | 16 (67) | 8 (33) | 13 (54) | 11 (46) | 21 (88) | 3 (12) | ||||
| TNBC Status | ||||||||||
| Negative | 42 (72) | 16 (28) | 0.071 | 38 (66) | 20 (34) | 0.009 | 53 (91) | 5 (9) | 0.002 | |
| Positive | 7 (47) | 8 (53) | 4 (27) | 11 (73) | 8 (53) | 7 (47) | ||||
| EMT Markers | Vimentin | |||||||||
| Negative | 42 (82) | 9 (18) | <0.001 | 33 (65) | 18 (35) | 0.074 | 47 (92) | 4 (8) | 0.005 | |
| Positive | 7 (32) | 15 (68) | 9 (41) | 13 (59) | 14 (64) | 8 (36) | ||||
| Loss of E-Cadherin | ||||||||||
| Negative | 39 (76) | 12 (24) | 0.015 | 32 (63) | 19 (37) | 0.203 | 47 (92) | 4 (8) | 0.005 | |
| Positive | 10 (45) | 12 (55) | 10 (45) | 12 (55) | 14 (64) | 8 (36) | ||||
| Vimentin/Loss of E-Cadherin | ||||||||||
| Negative | 47 (76) | 15 (24) | <0.001 | 39 (63) | 23 (37) | 0.045 | 56 (90) | 6 (10) | 0.002 | |
| Positive | 2 (18) | 9 (82) | 3 (27) | 8 (73) | 5 (45) | 6 (55) | ||||
| (A) | ||||||||||
| ALDH1hi | CD44hi/CD24lo-ALDH1hi | Ep-CAMhi/ALDH1hi | ||||||||
| H-Score < 35 | H-Score ≥ 35 | ♣ p | − | + | ♣ p | − | + | ♣ p | ||
| Immune Markers | PD-L1 | |||||||||
| <5% | 41 (69) * | 18 (31) | 1.000 | 54 (92) | 5 (8) | 0.174 | 51 (86) | 8 (14) | 0.432 | |
| ≥5% | 10 (71) | 4 (29) | 11 (79) | 3 (21) | 11 (79) | 3 (21) | ||||
| TIL | ||||||||||
| Low (Score 1 and 2) | 44 (77) | 13 (23) | 0.015 | 53 (93) | 4 (7) | 0.064 | 52 (91) | 5 (9) | 0.011 | |
| High (Score 3) | 7 (44) | 9 (56) | 12 (75) | 4 (25) | 10 (62) | 6 (38) | ||||
| § FOXP3 + TIL | ||||||||||
| <10% | 29 (78) | 8 (22) | 0.170 | 34 (92) | 3 (8) | 1.000 | 33 (89) | 4 (11) | 0.483 | |
| ≥10% | 17 (61) | 11 (39) | 25 (89) | 3 (11) | 23 (82) | 5 (18) | ||||
| § PD-1 + TIL | ||||||||||
| <10% | 24 (75) | 8 (25) | 0.603 | 30 (94) | 2 (6) | 0.681 | 29 (91) | 3 (9) | 0.494 | |
| ≥10% | 26 (68) | 12 (32) | 34 (89) | 4 (11) | 32 (84) | 6 (16) | ||||
| Proliferation Markers | Ki-67 | |||||||||
| <20% | 29 (81) | 7 (19) | 0.074 | 35 (97) | 1 (3) | 0.056 | 34 (94) | 2 (6) | 0.046 | |
| ≥20% | 22 (59) | 15 (41) | 30 (81) | 7 (19) | 28 (80) | 9 (20) | ||||
| § SKP2 | ||||||||||
| <10% | 34 (74) | 12 (26) | 0.423 | 42 (91) | 4 (9) | 0.440 | 41 (89) | 5 (11) | 0.307 | |
| ≥10% | 16 (64) | 9 (36) | 21 (84) | 4 (16) | 20 (80) | 5 (20) | ||||
| § p21 | ||||||||||
| <10% | 29 (67) | 14 (33) | 0.599 | 36 (84) | 7 (16) | 0.136 | 34 (79) | 9 (21) | 0.077 | |
| ≥10% | 21 (75) | 7 (25) | 27 (96) | 1 (4) | 27 (96) | 1 (4) | ||||
| § p27 | ||||||||||
| <50% | 32 (63) | 19 (37) | 0.041 | 43 (84) | 8 (16) | 0.095 | 41 (80) | 10 (20) | 0.053 | |
| ≥50% | 18 (90) | 2 (10) | 20 (100) | 0 (0) | 20(100) | 0 (0) | ||||
| Loss of both p21/p27 | ||||||||||
| NO | 19 (61) | 12 (39) | 0.191 | 24 (77) | 7 (23) | 0.018 | 22 (71) | 9 (29) | 0.002 | |
| YES | 31 (78) | 9 (22) | 39 (98) | 1 (2) | 39 (98) | 1 (3) | ||||
| (B) | ||||||||||
| ALDH1hi | CD44hi/CD24lo-ALDH1hi | ALDH1hi/Ep-CAMhi | ||||||||
| H-Score < 35 | H-Score ≥ 35 | ♣ p | − | + | ♣ p | − | + | ♣ p | ||
| Clinicopathological Markers | Age | |||||||||
| <40 years | 19 (79) * | 5 (21) | 0.284 | 21 (88) | 3 (12) | 1.000 | 20 (83) | 4 (17) | 1.000 | |
| ≥40 years | 32 (65) | 17 (35) | 44 (90) | 5 (10) | 42 (86) | 7 (14) | ||||
| Tumor Size | ||||||||||
| <4 cm | 34 (87) | 5 (13) | <0.001 | 38 (97) | 1 (3) | 0.022 | 37 (95) | 2 (5) | 0.019 | |
| ≥4 cm | 17 (50) | 17 (50) | 27 (79) | 7 (21) | 25 (74) | 9 (26) | ||||
| Invasion | ||||||||||
| Absent | 22 (73) | 8 (27) | 0.616 | 28 (93) | 2 (7) | 0.458 | 26 (87) | 4 (13) | 1.000 | |
| Present | 29 (67) | 14 (33) | 37 (86) | 6 (14) | 36 (84) | 7 (16) | ||||
| Histological Grade | ||||||||||
| 1 and 2 | 31 (84) | 6 (16) | 0.011 | 36 (97) | 1 (3) | 0.028 | 34 (92) | 3 (8) | 0.112 | |
| 3 | 20 (56) | 16 (44) | 29 (81) | 7 (19) | 28 (78) | 8 (22) | ||||
| § Lymph Node Metastasis | ||||||||||
| Absent | 19 (73) | 7 (27) | 0.794 | 25 (96) | 1 (4) | 0.410 | 24 (92) | 2 (8) | 0.307 | |
| Present | 32 (70) | 14 (30) | 40 (87) | 6 (13) | 37 (80) | 9 (20) | ||||
| Neoadjuvant Chemotherapy | ||||||||||
| Absent | 17 (57) | 13 (43) | 0.068 | 26 (87) | 4 (13) | 0.710 | 25 (83) | 5 (17) | 0.752 | |
| Present | 34 (79) | 9 (21) | 39 (91) | 4 (9) | 37 (86) | 6 (14) | ||||
| BC Subtype Markers | ER Status | |||||||||
| Negative | 16 (59) | 11 (41) | 0.187 | 21 (78) | 6 (22) | 0.045 | 19 (70) | 8 (30) | 0.015 | |
| Positive | 35 (76) | 11 (24) | 44 (96) | 2 (4) | 43 (93) | 3 (7) | ||||
| PR Status | ||||||||||
| Negative | 26 (65) | 14 (35) | 0.443 | 33 (83) | 7 (17) | 0.065 | 31 (78) | 9 (23) | 0.097 | |
| Positive | 25 (76) | 8 (24) | 32 (97) | 1 (3) | 31 (94) | 2 (6) | ||||
| HER2/neu Status | ||||||||||
| Negative | 37 (76) | 12 (24) | 0.176 | 44 (90) | 5 (10) | 1.000 | 43 (88) | 6 (12) | 0.487 | |
| Positive | 14 (58) | 10 (42) | 21 (88) | 3 (12) | 19 (79) | 5 (21) | ||||
| TNBC Status | ||||||||||
| Negative | 41 (71) | 17 (29) | 0.760 | 53 (91) | 5 (9) | 0.348 | 51 (88) | 7 (12) | 0.221 | |
| Positive | 10 (67) | 5 (33) | 12 (80) | 3 (20) | 11 (73) | 4 (27) | ||||
| EMT Markers | Vimentin | |||||||||
| Negative | 35 (69) | 16 (31) | 0.788 | 48 (94) | 3 (6) | 0.049 | 44 (86) | 7 (14) | 0.724 | |
| Positive | 16 (73) | 6 (27) | 17 (77) | 5 (23) | 18 (82) | 4 (18) | ||||
| Loss of E-Cadherin | ||||||||||
| Negative | 38 (75) | 13 (25) | 0.266 | 47 (92) | 4 (8) | 0.232 | 46 (90) | 5 (10) | 0.077 | |
| Positive | 13 (59) | 9 (41) | 18 (82) | 4 (18) | 16 (72) | 6 (27) | ||||
| Vimentin/Loss of E-Cadherin | ||||||||||
| Negative | 44 (71) | 18 (29) | 0.724 | 57 (92) | 5 (8) | 0.095 | 55 (89) | 7 (11) | 0.055 | |
| Positive | 7 (64) | 4 (36) | 8 (73) | 3 (27) | 7 (64) | 4 (36) | ||||
| Relapse | RFS | Death | OS | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Univariate | Multivariate | Univariate | Multivariate | Multivariate | |||||||||||||||
| − | + | HR | 95% CI | * p | HR | 95%CI | ♣ p | − | + | HR | 95% CI | * p | HR | 95% CI | * p | ||||
| CD44hi CD24lo | |||||||||||||||||||
| <10% | 22 (45) * | 27 (55) | 1 | 38 (78) | 11 (22) | 1 | 1 | ||||||||||||
| ≥10% | 11 (46) | 13 (54) | 1.1 | 0.6–2.2 | 0.718 | 12 (50) | 12 (50) | 2.4 | 1.0–5.4 | 0.039 | 2.4 | 1.1–5.6 | 0.039 | ||||||
| Ep-CAMhi | |||||||||||||||||||
| <H-Score 200 | 19 (45) | 23 (55) | 1 | 33 (79) | 9 (21) | 1 | 1 | 1 | |||||||||||
| ≥H-Score 200 | 14 (45) | 17 (55) | 1.2 | 0.6–2.3 | 0.541 | 17 (55) | 14 (45) | 2.5 | 1.1–5.7 | 0.034 | 1.9 | 0.8–4.6 | 0.131 | 2.0 | 0.8–4.7 | 0.113 | |||
| CD44hi CD24lo-Ep-CAMhi | |||||||||||||||||||
| <H-Score 200 | 27 (44) | 34 (56) | 1 | 44 (72) | 17 (28) | 1 | |||||||||||||
| ≥H-Score 200 | 6 (50) | 6 (50) | 1.0 | 0.4–2.3 | 0.946 | 6 (50) | 6 (50) | 2.0 | 0.8–5.1 | 0.140 | |||||||||
| ALDH1hi | |||||||||||||||||||
| <H-Score 35 | 24 (47) | 27 (53) | 1 | 40 (78) | 11 (22) | 1 | 1 | ||||||||||||
| ≥H-Score 35 | 9 (41) | 13 (59) | 1.2 | 0.6–2.3 | 0.630 | 10 (45) | 12 (55) | 3.0 | 1.3–6.9 | 0.008 | 3.0 | 1.3–7.0 | 0.009 | ||||||
| CD44hi/CD24lo-ALDH1hi | |||||||||||||||||||
| Negative | 32 (49) | 33 (51) | 1 | 1 | 49 (75) | 16 (25) | 1 | 1 | |||||||||||
| Positive | 1 (12) | 7 (88) | 2.8 | 1.2–6.4 | 0.014 | 2.8 | 1.2–6.4 | 0.014 | 1 (13) | 7 (87) | 6.4 | 2.6–15.8 | <0.001 | 5.4 | 2.1–13.6 | <0.001 | |||
| Ep-CAMhi/ALDH1hi | |||||||||||||||||||
| Negative | 29 (47) | 33 (53) | 1 | 45 (73) | 17 (27) | 1 | |||||||||||||
| Positive | 4 (36) | 7 (64) | 1.4 | 0.6–3.2 | 0.417 | 5 (45) | 6 (55) | 2.5 | 1.0–6.2 | 0.059 | |||||||||
| CD24hi | |||||||||||||||||||
| <H-Score 140 | 25 (52) | 23 (48) | 1 | 1 | 33 (69) | 15 (31) | 1 | ||||||||||||
| ≥H-Score 140 | 8 (32) | 17 (68) | 2.1 | 1.1–3.9 | 0.022 | 2.2 | 1.1–4.3 | 0.023 | 17 (68) | 8 (32) | 1.4 | 0.6–3.3 | 0.460 | ||||||
| CD10 | |||||||||||||||||||
| <10% | 28 (47) | 32 (53) | 1 | 44 (73) | 16 (27) | 1 | |||||||||||||
| ≥10% | 4 (36) | 7 (64) | 1.3 | 0.6–3.0 | 0.491 | 5 (45) | 6 (55) | 2.3 | 0.9–5.9 | 0.081 | |||||||||
| BMI1 | |||||||||||||||||||
| Negative | 20 (38) | 32 (62) | 1 | 32 (62) | 20 (38) | 1 | |||||||||||||
| Positive | 12 (63) | 7 (37) | 0.4 | 0.2–1.0 | 0.043 | 17 (89) | 2 (11) | 0.2 | 0.05–0.8 | 0.027 | |||||||||
| CD44/CD24 | Ep-CAM | CD44/CD24- Ep-CAM | ALDH1 | CD44/CD24-ALDH1 | Ep-CAM/ ALDH1 | CD24 | CD10 | |
|---|---|---|---|---|---|---|---|---|
| EMT | √√√ * | √ | √√√ | √ | √ | |||
| Higher Proliferation | √ | √√√√ | √ | √ | √ | √√ | √√√ | |
| Immune Evasion | √ | √√√ | √ | √√√ | ||||
| Higher Histological Grade | √ | √ | √ | √ | ||||
| ER negative | √ | √√√ | √ | √ | √ | √ |
| Antibody | Antigen Retrieval | Primary Antibody | Secondary Antibody | |||
|---|---|---|---|---|---|---|
| Solution | Conditions * | Dilution | Clone/Cat# | |||
| CD44/CD24 | Tris-EDTA pH9 | 121 °C/7 min | CD24, 1:100 CD44, 1:500 | ML5/ HPA005785 | Biotinylated goat anti-mouse AP and Envision goat anti-rabbit HRP | |
| ALDH1 | CC1 # | 121 °C/6 min | 1:200 | 44/ALDH | Envision goat anti-mouse | |
| Ep-CAM | Citrate | 121 °C/8 min | 1:500 | VU-1D9 | Envision goat anti-mouse | |
| BMI1 | CC1 | 121 °C/6 min | 1:500 | F6 | Biotinylated goat anti-mouse (1/1000) (JL) ⊥ | |
| CD10 # | CC1 | (clone SP67), Ventana Benchmark Ultra system | ||||
| PD-L1 # | (clone SP263), Ventana Benchmark Ultra system, previously described [54] | |||||
| FOXP3 and PD-1 | (Clones 236A/E7 and J116, respectively), previously described [56] | |||||
| Vimentin #/E-Cadherin | (Clones V9 and EP700Y, respectively), previously described [52] | |||||
| SKP2, p21, and p27 | (Clones D3G5, 12D1, and F-8 respectively), previously described [55] | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ghebeh, H.; Mirza, J.Y.; Al-Tweigeri, T.; Al-Alwan, M.; Tulbah, A. Profiling of Breast Cancer Stem Cell Types/States Shows the Role of CD44hi/CD24lo-ALDH1hi as an Independent Prognostic Factor After Neoadjuvant Chemotherapy. Int. J. Mol. Sci. 2025, 26, 8219. https://doi.org/10.3390/ijms26178219
Ghebeh H, Mirza JY, Al-Tweigeri T, Al-Alwan M, Tulbah A. Profiling of Breast Cancer Stem Cell Types/States Shows the Role of CD44hi/CD24lo-ALDH1hi as an Independent Prognostic Factor After Neoadjuvant Chemotherapy. International Journal of Molecular Sciences. 2025; 26(17):8219. https://doi.org/10.3390/ijms26178219
Chicago/Turabian StyleGhebeh, Hazem, Jumanah Y. Mirza, Taher Al-Tweigeri, Monther Al-Alwan, and Asma Tulbah. 2025. "Profiling of Breast Cancer Stem Cell Types/States Shows the Role of CD44hi/CD24lo-ALDH1hi as an Independent Prognostic Factor After Neoadjuvant Chemotherapy" International Journal of Molecular Sciences 26, no. 17: 8219. https://doi.org/10.3390/ijms26178219
APA StyleGhebeh, H., Mirza, J. Y., Al-Tweigeri, T., Al-Alwan, M., & Tulbah, A. (2025). Profiling of Breast Cancer Stem Cell Types/States Shows the Role of CD44hi/CD24lo-ALDH1hi as an Independent Prognostic Factor After Neoadjuvant Chemotherapy. International Journal of Molecular Sciences, 26(17), 8219. https://doi.org/10.3390/ijms26178219







