Umbilical Cord Mesenchymal Stem Cell-Derived Extracellular Vesicles Enhance Chondrocyte Function by Reducing Oxidative Stress in Chondrocytes
Abstract
1. Introduction
2. Results
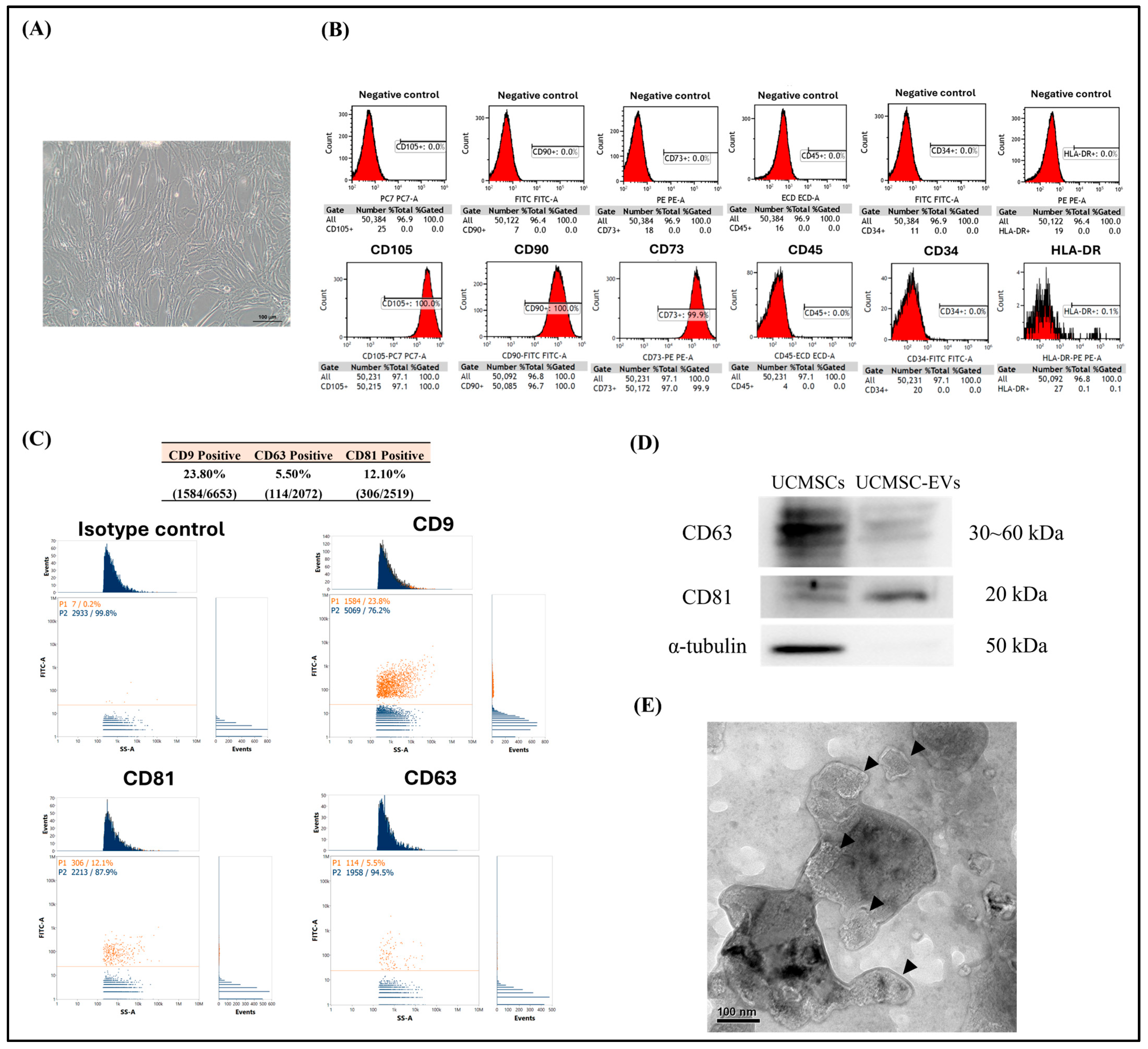
2.1. Characterization of UCMSCs and UCMSC-EVs
2.2. Visualization of UCMSC-EV Internalization in Chondrocytes
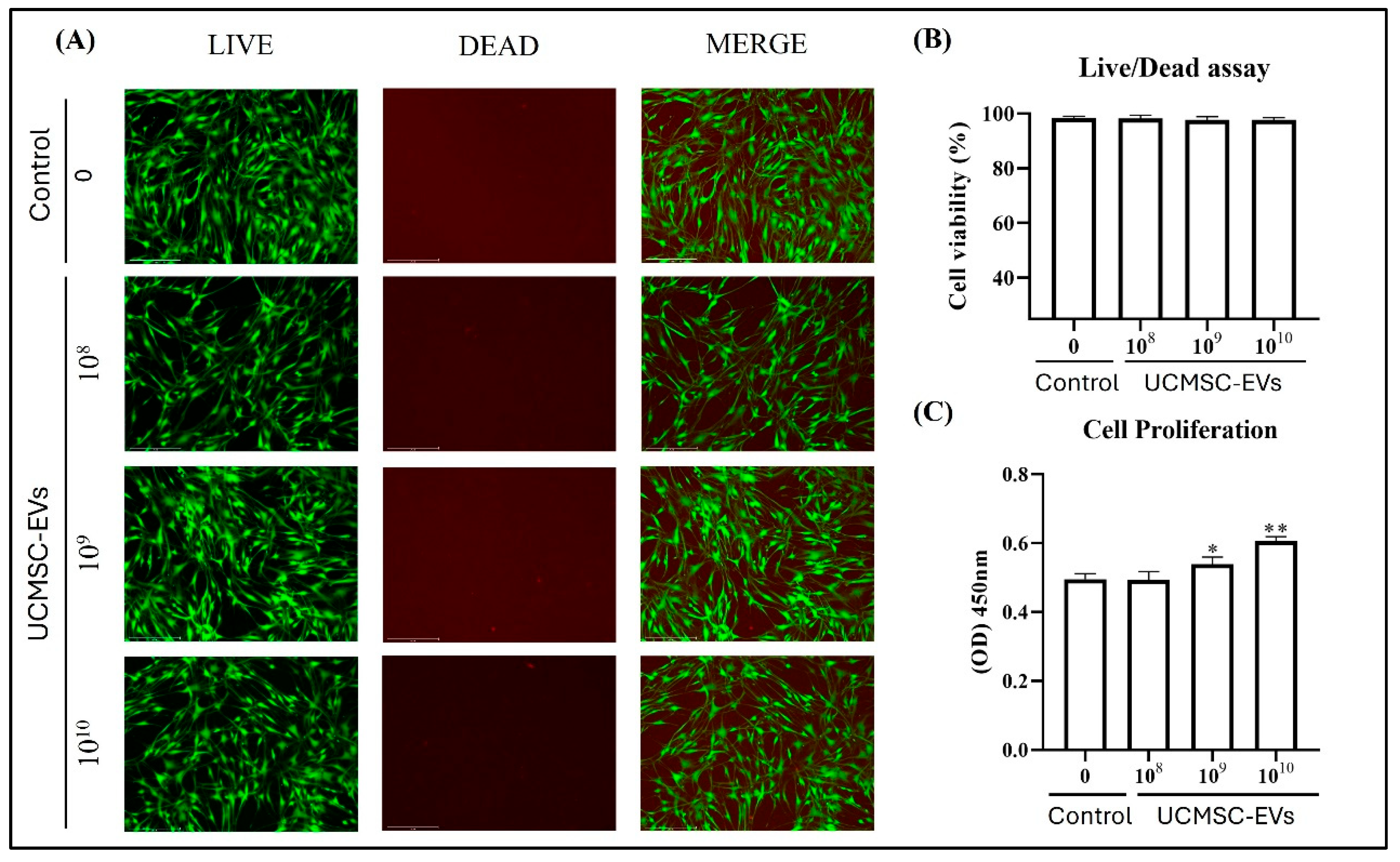
2.3. Effect of UCMSC-EVs on Survival and Proliferation of Chondrocytes
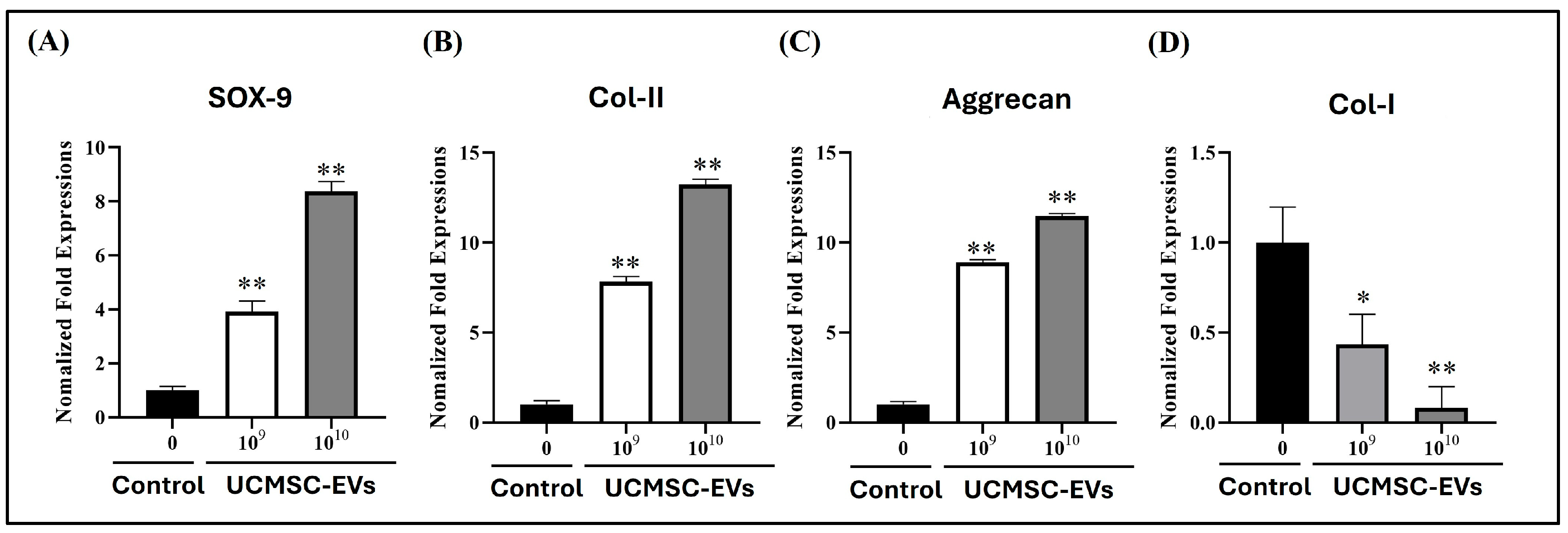
2.4. Effect of UCMSC-EVs on mRNA Expression of Chondrogenic Genes (SOX-9, Col-II, and Aggrecan) and Fribrocartilgous Gene (Col-I) of Chondrocytes
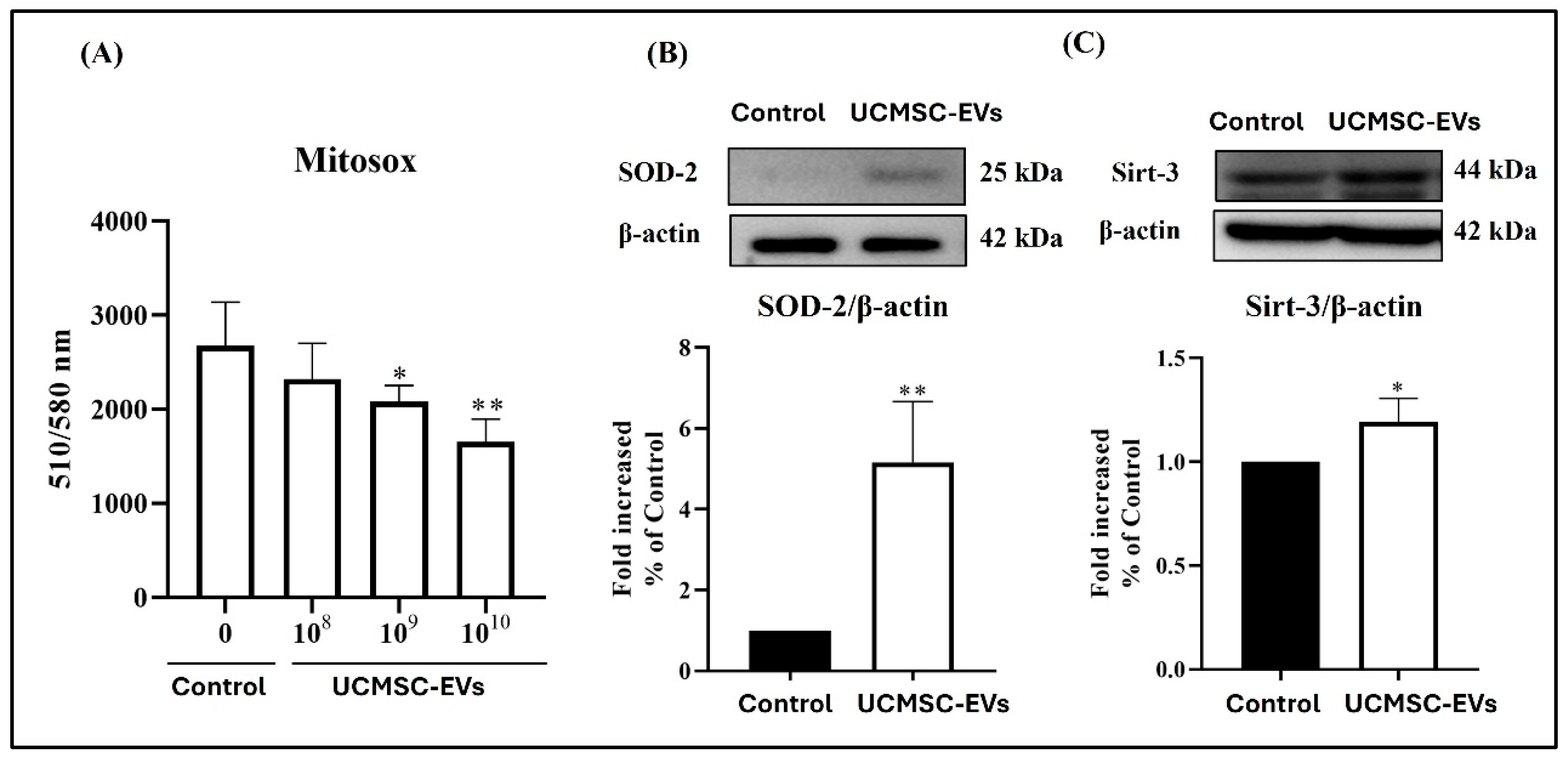
2.5. Effect of UCMSC-EVs on Mitochondrial Superoxide Production and the Protein Levels of SOD-2 and Sirt-3 of Chondrocytes
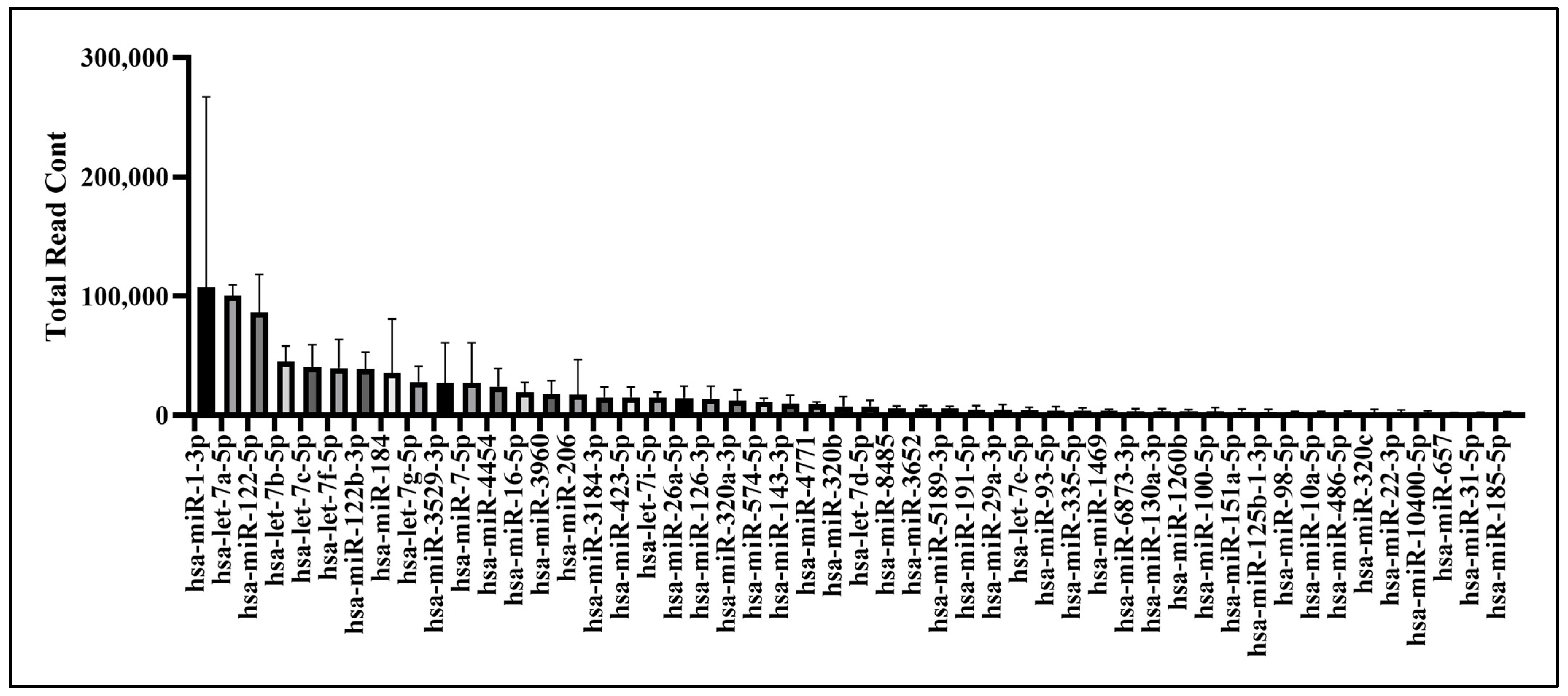
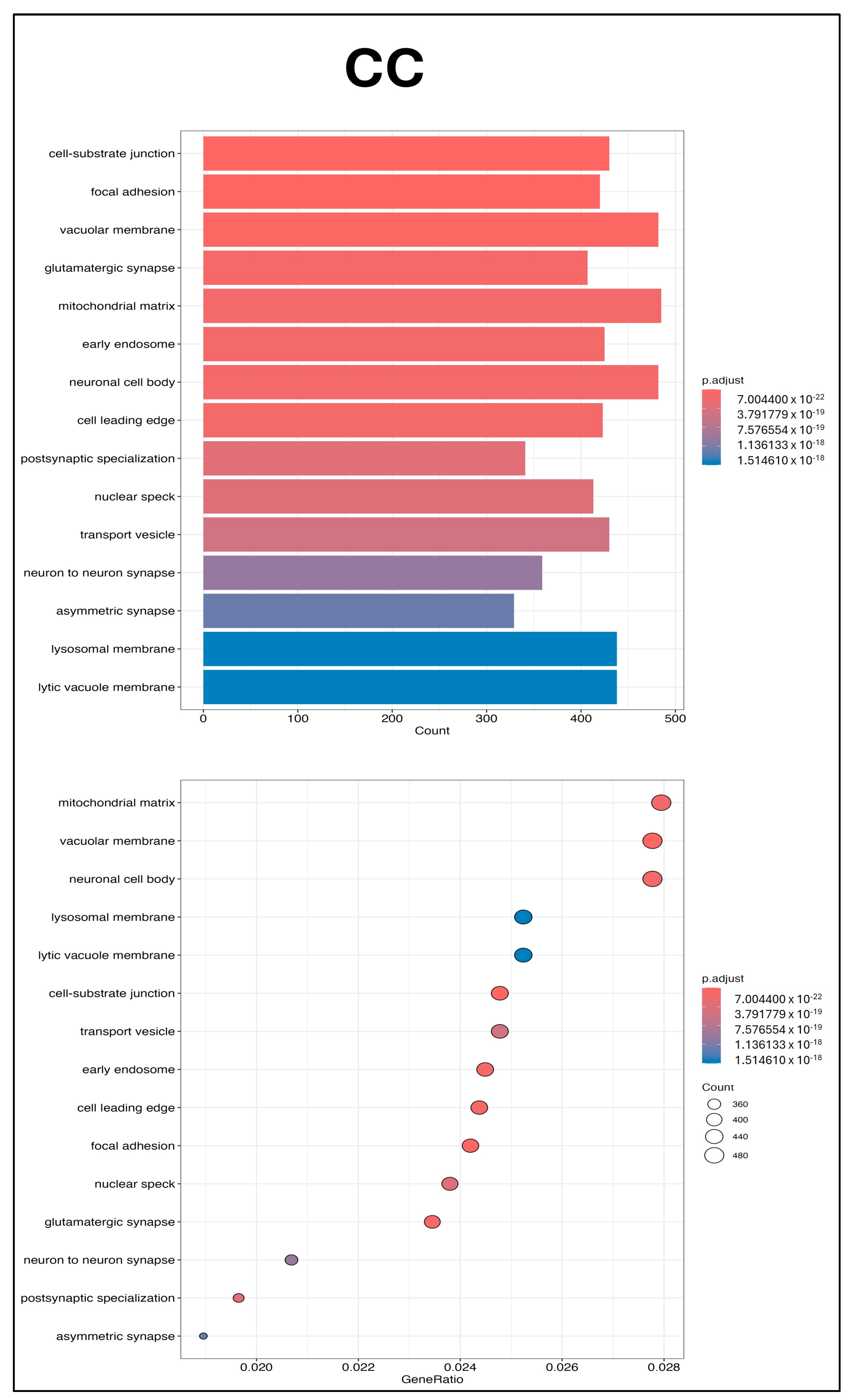
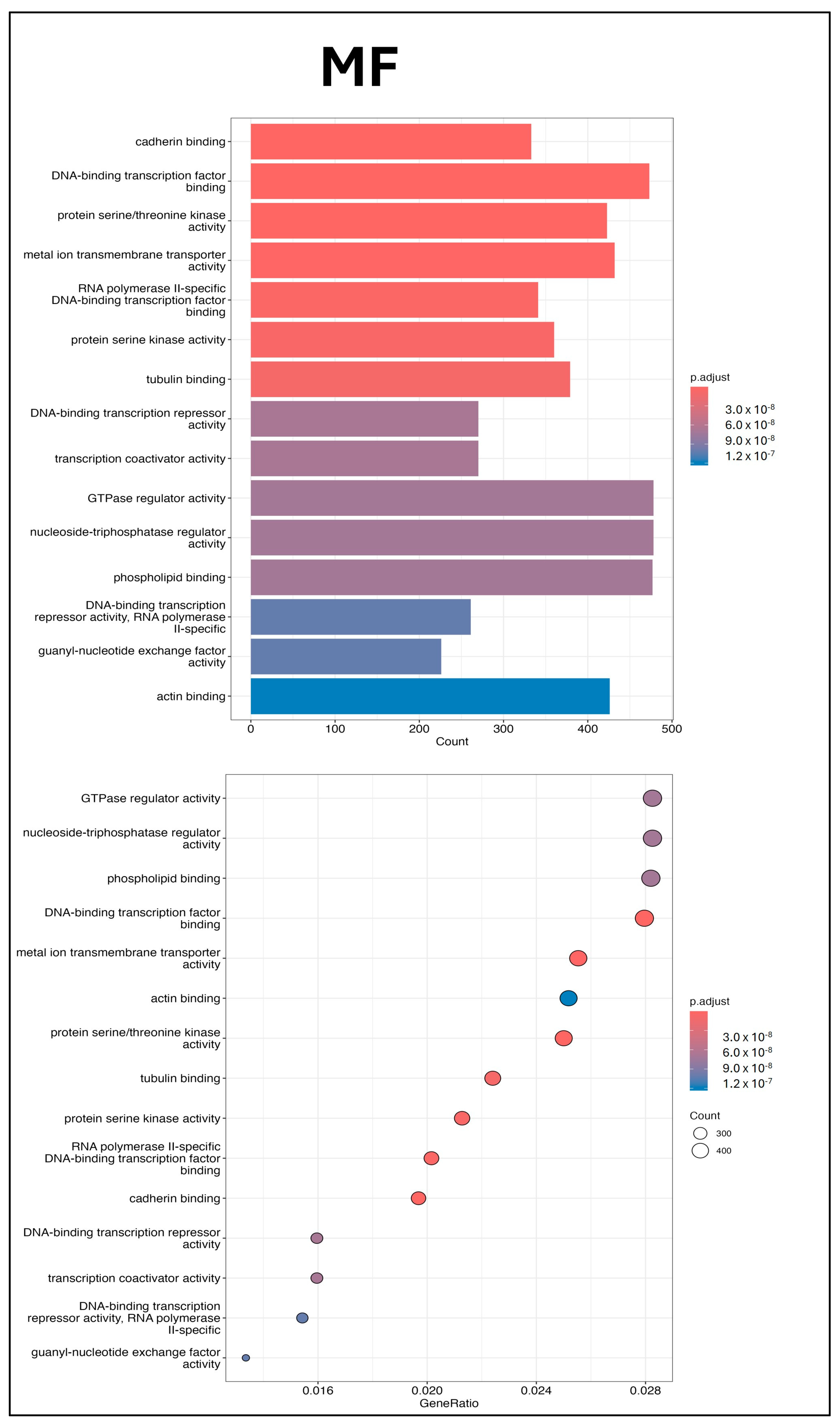
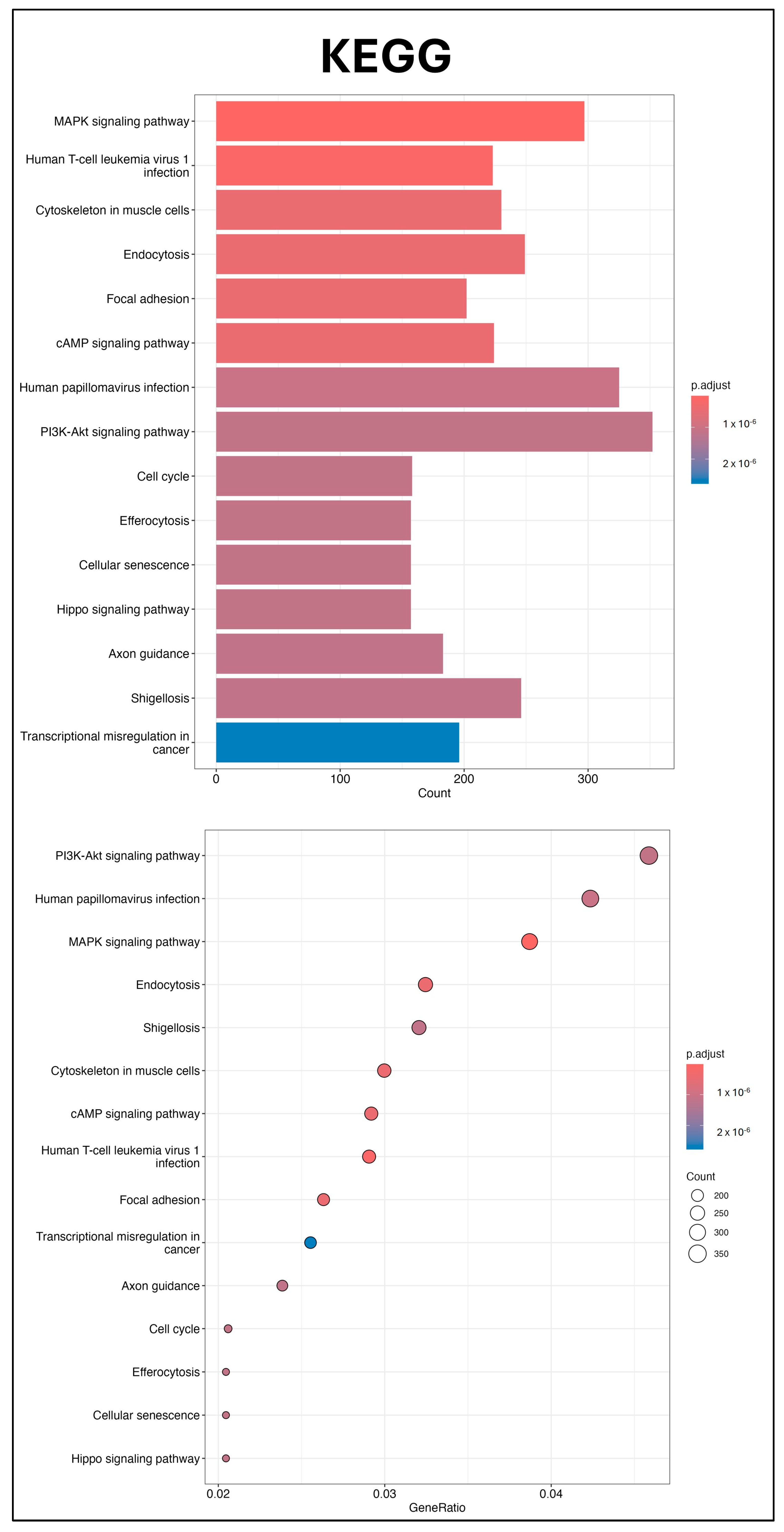
2.6. Bioinformatics Analysis of UCMSC-EV miRNAs
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Human Chondrocyte Culture
4.3. Human UCMSC Culture and Isolation of UCMSC-EVs
4.4. UCMSC-EV Protein Composition and Size Analysis Using Nano Flow Cytometry (nFCM)
4.5. UCMSC-EV Protein Composition Analysis Using Western Blotting
4.6. Morphological Characterization of UCMSC-EVs via TEM
4.7. UCMSC-EV Treatment of Chondrocytes and Study Groups
4.8. Visualization of UCMSC-EV Internalization in Chondrocytes
4.9. Cell Survival of Chondrocytes After UCMSC-EV Treatment
4.10. Evaluation of Chondrocyte Proliferation Following UCMSC-EV Treatment
4.11. Detection and Quantification of Mitochondrial Superoxide
4.12. Extraction of RNA and Analysis of Gene Expression Using Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
4.13. Antibodies
4.14. Western Blot Analysis
4.15. Small RNA Sequencing and Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway and Gene Ontology (GO) Enrichment Analyses of UCMSC-EVs
miRNA Extraction
4.16. miRNA Library Construction for Next-Generation Sequencing
4.17. Bioinformatics Analysis
4.18. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Caldwell, K.L.; Wang, J. Cell-based articular cartilage repair: The link between development and regeneration. Osteoarthr. Cartil. 2015, 23, 351–362. [Google Scholar] [CrossRef]
- Makris, E.A.; Gomoll, A.H.; Malizos, K.N.; Hu, J.C.; Athanasiou, K.A. Repair and tissue engineering techniques for articular cartilage. Nat. Rev. Rheumatol. 2015, 11, 21–34. [Google Scholar] [CrossRef]
- Focsa, M.A.; Florescu, S.; Gogulescu, A. Emerging Strategies in Cartilage Repair and Joint Preservation. Medicina 2024, 61, 24. [Google Scholar] [CrossRef]
- Charlier, E.; Deroyer, C.; Ciregia, F.; Malaise, O.; Neuville, S.; Plener, Z.; Malaise, M.; de Seny, D. Chondrocyte dedifferentiation and osteoarthritis (OA). Biochem. Pharmacol. 2019, 165, 49–65. [Google Scholar] [CrossRef]
- Kloppenburg, M.; Berenbaum, F. Osteoarthritis year in review 2019: Epidemiology and therapy. Osteoarthr. Cartil. 2020, 28, 242–248. [Google Scholar] [CrossRef] [PubMed]
- Davies, R.L.; Kuiper, N.J. Regenerative Medicine: A Review of the Evolution of Autologous Chondrocyte Implantation (ACI) Therapy. Bioengineering 2019, 6, 22. [Google Scholar] [CrossRef] [PubMed]
- Messner, K.; Maletius, W. The long-term prognosis for severe damage to weight-bearing cartilage in the knee: A 14-year clinical and radiographic follow-up in 28 young athletes. Acta Orthop. 1996, 67, 165–168. [Google Scholar] [CrossRef] [PubMed]
- Gou, G.H.; Tseng, F.J.; Wang, S.H.; Chen, P.J.; Shyu, J.F.; Weng, C.F.; Pan, R.Y. Autologous Chondrocyte Implantation Versus Microfracture in the Knee: A Meta-analysis and Systematic Review. Arthroscopy 2020, 36, 289–303. [Google Scholar] [CrossRef] [PubMed]
- Ogura, T.; Mosier, B.A.; Bryant, T.; Minas, T. A 20-Year Follow-up After First-Generation Autologous Chondrocyte Implantation. Am. J. Sports Med. 2017, 45, 2751–2761. [Google Scholar] [CrossRef]
- Roelofs, A.J.; McClure, J.J.; Hay, E.A.; De Bari, C. Stem and progenitor cells in the synovial joint as targets for regenerative therapy. Nat. Rev. Rheumatol. 2025, 21, 211–220. [Google Scholar] [CrossRef]
- Schulze-Tanzil, G. Activation and dedifferentiation of chondrocytes: Implications in cartilage injury and repair. Ann. Anat. 2009, 191, 325–338. [Google Scholar] [CrossRef]
- Fares, M.Y.; Daher, M.; Boufadel, P.; Haikal, E.; Haj Shehade, T.; Koa, J.; Khan, A.Z.; Abboud, J.A. The use of autologous chondrocyte transplantation for the treatment of osteoarthritis: A systematic review of clinical trials. Cell Tissue Bank. 2024, 26, 5. [Google Scholar] [CrossRef]
- Welsh, B.L.; Sikder, P. Advancements in Cartilage Tissue Engineering: A Focused Review. J. Biomed. Mater. Res. Part B Appl. Biomater. 2025, 113, e35520. [Google Scholar] [CrossRef]
- Ashraf, S.; Cha, B.H.; Kim, J.S.; Ahn, J.; Han, I.; Park, H.; Lee, S.H. Regulation of senescence associated signaling mechanisms in chondrocytes for cartilage tissue regeneration. Osteoarthr. Cartil. 2016, 24, 196–205. [Google Scholar] [CrossRef]
- Di Micco, R.; Krizhanovsky, V.; Baker, D.; d’Adda di Fagagna, F. Cellular senescence in ageing: From mechanisms to therapeutic opportunities. Nat. Rev. Mol. Cell Biol. 2021, 22, 75–95. [Google Scholar] [CrossRef]
- Picca, A.; Mankowski, R.T.; Burman, J.L.; Donisi, L.; Kim, J.S.; Marzetti, E.; Leeuwenburgh, C. Mitochondrial quality control mechanisms as molecular targets in cardiac ageing. Nat. Rev. Cardiol. 2018, 15, 543–554. [Google Scholar] [CrossRef]
- Hong, E.H.; Lee, S.J.; Kim, J.S.; Lee, K.H.; Um, H.D.; Kim, J.H.; Kim, S.J.; Kim, J.I.; Hwang, S.G. Ionizing radiation induces cellular senescence of articular chondrocytes via negative regulation of SIRT1 by p38 kinase. J. Biol. Chem. 2010, 285, 1283–1295. [Google Scholar] [CrossRef] [PubMed]
- Morita, K.; Miyamoto, T.; Fujita, N.; Kubota, Y.; Ito, K.; Takubo, K.; Miyamoto, K.; Ninomiya, K.; Suzuki, T.; Iwasaki, R.; et al. Reactive oxygen species induce chondrocyte hypertrophy in endochondral ossification. J. Exp. Med. 2007, 204, 1613–1623. [Google Scholar] [CrossRef] [PubMed]
- Vago, J.; Takacs, R.; Kovacs, P.; Hajdu, T.; van der Veen, D.R.; Matta, C. Combining biomechanical stimulation and chronobiology: A novel approach for augmented chondrogenesis? Front. Bioeng. Biotechnol. 2023, 11, 1232465. [Google Scholar] [CrossRef] [PubMed]
- da Silva Meirelles, L.; Chagastelles, P.C.; Nardi, N.B. Mesenchymal stem cells reside in virtually all post-natal organs and tissues. J. Cell Sci. 2006, 119, 2204–2213. [Google Scholar] [CrossRef]
- Jiang, R.; Han, Z.; Zhuo, G.; Qu, X.; Li, X.; Wang, X.; Shao, Y.; Yang, S.; Han, Z.C. Transplantation of placenta-derived mesenchymal stem cells in type 2 diabetes: A pilot study. Front. Med. 2011, 5, 94–100. [Google Scholar] [CrossRef]
- Attaelmanan, G.A.; Khalil, H.B. Assessment of Umbilical Cord Mesenchymal Stem Cell Cultivation Using Fetal Bovine Serum or Platelet Lysate. Cureus 2025, 17, e78044. [Google Scholar] [CrossRef]
- Carvalho, M.M.; Teixeira, F.G.; Reis, R.L.; Sousa, N.; Salgado, A.J. Mesenchymal stem cells in the umbilical cord: Phenotypic characterization, secretome and applications in central nervous system regenerative medicine. Curr. Stem Cell Res. Ther. 2011, 6, 221–228. [Google Scholar] [CrossRef]
- Ghaneialvar, H.; Soltani, L.; Rahmani, H.R.; Lotfi, A.S.; Soleimani, M. Characterization and Classification of Mesenchymal Stem Cells in Several Species Using Surface Markers for Cell Therapy Purposes. Indian J. Clin. Biochem. 2018, 33, 46–52. [Google Scholar] [CrossRef] [PubMed]
- Liew, L.C.; Katsuda, T.; Gailhouste, L.; Nakagama, H.; Ochiya, T. Mesenchymal stem cell-derived extracellular vesicles: A glimmer of hope in treating Alzheimer’s disease. Int. Immunol. 2017, 29, 11–19. [Google Scholar] [CrossRef]
- Jiang, S.; Tian, G.; Yang, Z.; Gao, X.; Wang, F.; Li, J.; Tian, Z.; Huang, B.; Wei, F.; Sang, X.; et al. Enhancement of acellular cartilage matrix scaffold by Wharton’s jelly mesenchymal stem cell-derived exosomes to promote osteochondral regeneration. Bioact. Mater. 2021, 6, 2711–2728. [Google Scholar] [CrossRef]
- Altamirano, D.E.; Noller, K.; Mihaly, E.; Grayson, W.L. Recent advances toward understanding the role of transplanted stem cells in tissue-engineered regeneration of musculoskeletal tissues. F1000Research 2020, 9, 118. [Google Scholar] [CrossRef]
- Daneshmandi, L.; Shah, S.; Jafari, T.; Bhattacharjee, M.; Momah, D.; Saveh-Shemshaki, N.; Lo, K.W.; Laurencin, C.T. Emergence of the Stem Cell Secretome in Regenerative Engineering. Trends Biotechnol. 2020, 38, 1373–1384. [Google Scholar] [CrossRef]
- Velot, E.; Madry, H.; Venkatesan, J.K.; Bianchi, A.; Cucchiarini, M. Is Extracellular Vesicle-Based Therapy the Next Answer for Cartilage Regeneration? Front. Bioeng. Biotechnol. 2021, 9, 645039. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.H.; Vuong, C.K.; Ngo, N.H.; Yamashita, T.; Ye, X.; Futamura, Y.; Fukushige, M.; Obata-Yasuoka, M.; Hamada, H.; Osaka, M.; et al. Extracellular vesicles derived from Wharton’s Jelly mesenchymal stem cells inhibit the tumor environment via the miR-125b/HIF1alpha signaling pathway. Sci. Rep. 2022, 12, 13550. [Google Scholar] [CrossRef]
- Li, P.; Lv, S.; Jiang, W.; Si, L.; Liao, B.; Zhao, G.; Xu, Z.; Wang, L.; Zhang, J.; Wu, H.; et al. Exosomes derived from umbilical cord mesenchymal stem cells protect cartilage and regulate the polarization of macrophages in osteoarthritis. Ann. Transl. Med. 2022, 10, 976. [Google Scholar] [CrossRef]
- Figueroa-Valdes, A.I.; Luz-Crawford, P.; Herrera-Luna, Y.; Georges-Calderon, N.; Garcia, C.; Tobar, H.E.; Araya, M.J.; Matas, J.; Donoso-Meneses, D.; de la Fuente, C.; et al. Clinical-grade extracellular vesicles derived from umbilical cord mesenchymal stromal cells: Preclinical development and first-in-human intra-articular validation as therapeutics for knee osteoarthritis. J. Nanobiotechnol. 2025, 23, 13. [Google Scholar] [CrossRef]
- Yang, H.; Zhou, Y.; Ying, B.; Dong, X.; Qian, Q.; Gao, S. Effects of human umbilical cord mesenchymal stem cell-derived exosomes in the rat osteoarthritis models. Stem Cells Transl. Med. 2024, 13, 803–811. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.; Chen, M.; Cui, X.; Liu, Y.; Liu, Y.; Deng, S.; Yuan, T.; Fan, Y.; Wang, Q.; Zhang, X. Cell-Free Osteoarthritis Treatment with Sustained-Release of Chondrocyte-Targeting Exosomes from Umbilical Cord-Derived Mesenchymal Stem Cells to Rejuvenate Aging Chondrocytes. ACS Nano 2023, 17, 13358–13376. [Google Scholar] [CrossRef]
- Zhou, H.; Shen, X.; Yan, C.; Xiong, W.; Ma, Z.; Tan, Z.; Wang, J.; Li, Y.; Liu, J.; Duan, A.; et al. Extracellular vesicles derived from human umbilical cord mesenchymal stem cells alleviate osteoarthritis of the knee in mice model by interacting with METTL3 to reduce m6A of NLRP3 in macrophage. Stem Cell Res. Ther. 2022, 13, 322. [Google Scholar] [CrossRef]
- Roberts, S.; Menage, J.; Sandell, L.J.; Evans, E.H.; Richardson, J.B. Immunohistochemical study of collagen types I and II and procollagen IIA in human cartilage repair tissue following autologous chondrocyte implantation. Knee 2009, 16, 398–404. [Google Scholar] [CrossRef]
- He, C.; Zheng, S.; Luo, Y.; Wang, B. Exosome Theranostics: Biology and Translational Medicine. Theranostics 2018, 8, 237–255. [Google Scholar] [CrossRef]
- Lotvall, J.; Hill, A.F.; Hochberg, F.; Buzas, E.I.; Di Vizio, D.; Gardiner, C.; Gho, Y.S.; Kurochkin, I.V.; Mathivanan, S.; Quesenberry, P.; et al. Minimal experimental requirements for definition of extracellular vesicles and their functions: A position statement from the International Society for Extracellular Vesicles. J. Extracell. Vesicles 2014, 3, 26913. [Google Scholar] [CrossRef]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef] [PubMed]
- Welsh, J.A.; Goberdhan, D.C.I.; O’Driscoll, L.; Buzas, E.I.; Blenkiron, C.; Bussolati, B.; Cai, H.; Di Vizio, D.; Driedonks, T.A.P.; Erdbrugger, U.; et al. Minimal information for studies of extracellular vesicles (MISEV2023): From basic to advanced approaches. J. Extracell. Vesicles 2024, 13, e12404. [Google Scholar] [CrossRef] [PubMed]
- Mende, W.; Gotzl, R.; Kubo, Y.; Pufe, T.; Ruhl, T.; Beier, J.P. The Role of Adipose Stem Cells in Bone Regeneration and Bone Tissue Engineering. Cells 2021, 10, 975. [Google Scholar] [CrossRef]
- Gyorgy, B.; Szabo, T.G.; Pasztoi, M.; Pal, Z.; Misjak, P.; Aradi, B.; Laszlo, V.; Pallinger, E.; Pap, E.; Kittel, A.; et al. Membrane vesicles, current state-of-the-art: Emerging role of extracellular vesicles. Cell Mol. Life Sci. 2011, 68, 2667–2688. [Google Scholar] [CrossRef]
- Camussi, G.; Deregibus, M.C.; Bruno, S.; Cantaluppi, V.; Biancone, L. Exosomes/microvesicles as a mechanism of cell-to-cell communication. Kidney Int. 2010, 78, 838–848. [Google Scholar] [CrossRef]
- Losche, W.; Scholz, T.; Temmler, U.; Oberle, V.; Claus, R.A. Platelet-derived microvesicles transfer tissue factor to monocytes but not to neutrophils. Platelets 2004, 15, 109–115. [Google Scholar] [CrossRef]
- van Niel, G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Lee, A.J.; Gangi, L.R.; Zandkarimi, F.; Stockwell, B.R.; Hung, C.T. Red blood cell exposure increases chondrocyte susceptibility to oxidative stress following hemarthrosis. Osteoarthr. Cartil. 2023, 31, 1365–1376. [Google Scholar] [CrossRef]
- Zhuang, C.; Ni, S.; Yang, Z.C.; Liu, R.P. Oxidative Stress Induces Chondrocyte Apoptosis through Caspase-Dependent and Caspase-Independent Mitochondrial Pathways and the Antioxidant Mechanism of Angelica Sinensis Polysaccharide. Oxid. Med. Cell Longev. 2020, 2020, 3240820. [Google Scholar] [CrossRef]
- Guan, M.; Yu, Q.; Zhou, G.; Wang, Y.; Yu, J.; Yang, W.; Li, Z. Mechanisms of chondrocyte cell death in osteoarthritis: Implications for disease progression and treatment. J. Orthop. Surg. Res. 2024, 19, 550. [Google Scholar] [CrossRef] [PubMed]
- Yagi, M.; Endo, K.; Komori, K.; Sekiya, I. Comparison of the effects of oxidative and inflammatory stresses on rat chondrocyte senescence. Sci. Rep. 2023, 13, 7697. [Google Scholar] [CrossRef] [PubMed]
- Lowe, J.; Almarza, A.J. A review of in-vitro fibrocartilage tissue engineered therapies with a focus on the temporomandibular joint. Arch. Oral. Biol. 2017, 83, 193–201. [Google Scholar] [CrossRef] [PubMed]
- Cheung, C.; Tu, S.; Feng, Y.; Wan, C.; Ai, H.; Chen, Z. Mitochondrial quality control dysfunction in osteoarthritis: Mechanisms, therapeutic strategies & future prospects. Arch. Gerontol. Geriatr. 2024, 125, 105522. [Google Scholar] [CrossRef]
- Heywood, H.K.; Lee, D.A. Monolayer expansion induces an oxidative metabolism and ROS in chondrocytes. Biochem. Biophys. Res. Commun. 2008, 373, 224–229. [Google Scholar] [CrossRef] [PubMed]
- Heywood, H.K.; Lee, D.A. Low oxygen reduces the modulation to an oxidative phenotype in monolayer-expanded chondrocytes. J. Cell Physiol. 2010, 222, 248–253. [Google Scholar] [CrossRef]
- Bause, A.S.; Haigis, M.C. SIRT3 regulation of mitochondrial oxidative stress. Exp. Gerontol. 2013, 48, 634–639. [Google Scholar] [CrossRef]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef] [PubMed]
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef]
- Ferro, E.; Goitre, L.; Retta, S.F.; Trabalzini, L. The Interplay between ROS and Ras GTPases: Physiological and Pathological Implications. J. Signal Transduct. 2012, 2012, 365769. [Google Scholar] [CrossRef]
- Liu, J.; Song, X.; Yan, Y.; Liu, B. Role of GTPase-Dependent Mitochondrial Dynamins in Heart Diseases. Front. Cardiovasc. Med. 2021, 8, 720085. [Google Scholar] [CrossRef]
- Werner, E.; Werb, Z. Integrins engage mitochondrial function for signal transduction by a mechanism dependent on Rho GTPases. J. Cell Biol. 2002, 158, 357–368. [Google Scholar] [CrossRef] [PubMed]
- Shiau, J.P.; Chuang, Y.T.; Tang, J.Y.; Yang, K.H.; Chang, F.R.; Hou, M.F.; Yen, C.Y.; Chang, H.W. The Impact of Oxidative Stress and AKT Pathway on Cancer Cell Functions and Its Application to Natural Products. Antioxidants 2022, 11, 1845. [Google Scholar] [CrossRef]
- Signorile, A.; Ferretta, A.; Ruggieri, M.; Paolicelli, D.; Lattanzio, P.; Trojano, M.; De Rasmo, D. Mitochondria, Oxidative Stress, cAMP Signalling and Apoptosis: A Crossroads in Lymphocytes of Multiple Sclerosis, a Possible Role of Nutraceutics. Antioxidants 2020, 10, 21. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.K.; Chang, L.H.; Hung, S.H.; Wu, S.C.; Lee, H.Y.; Lin, Y.S.; Chen, C.H.; Fu, Y.C.; Wang, G.J.; Ho, M.L. Parathyroid hormone 1-34 inhibits terminal differentiation of human articular chondrocytes and osteoarthritis progression in rats. Arthritis Rheum. 2009, 60, 3049–3060. [Google Scholar] [CrossRef]
- Purushothaman, A. Exosomes from Cell Culture-Conditioned Medium: Isolation by Ultracentrifugation and Characterization. Methods Mol. Biol. 2019, 1952, 233–244. [Google Scholar] [CrossRef] [PubMed]
- Relic, B.; Guicheux, J.; Mezin, F.; Lubberts, E.; Togninalli, D.; Garcia, I.; van den Berg, W.B.; Guerne, P.A. Il-4 and IL-13, but not IL-10, protect human synoviocytes from apoptosis. J. Immunol. 2001, 166, 2775–2782. [Google Scholar] [CrossRef]
- Ma, W.J.; Ruys, A.J.; Mason, R.S.; Martin, P.J.; Bendavid, A.; Liu, Z.; Ionescu, M.; Zreiqat, H. DLC coatings: Effects of physical and chemical properties on biological response. Biomaterials 2007, 28, 1620–1628. [Google Scholar] [CrossRef]
- Magne, D.; Bluteau, G.; Faucheux, C.; Palmer, G.; Vignes-Colombeix, C.; Pilet, P.; Rouillon, T.; Caverzasio, J.; Weiss, P.; Daculsi, G.; et al. Phosphate is a specific signal for ATDC5 chondrocyte maturation and apoptosis-associated mineralization: Possible implication of apoptosis in the regulation of endochondral ossification. J. Bone Min. Res. 2003, 18, 1430–1442. [Google Scholar] [CrossRef]
- Wu, S.C.; Huang, P.Y.; Chen, C.H.; Teong, B.; Chen, J.W.; Wu, C.W.; Chang, J.K.; Ho, M.L. Hyaluronan microenvironment enhances cartilage regeneration of human adipose-derived stem cells in a chondral defect model. Int. J. Biol. Macromol. 2018, 119, 726–740. [Google Scholar] [CrossRef]
- Wu, S.C.; Chen, C.H.; Wang, J.Y.; Lin, Y.S.; Chang, J.K.; Ho, M.L. Hyaluronan size alters chondrogenesis of adipose-derived stem cells via the CD44/ERK/SOX-9 pathway. Acta Biomater. 2018, 66, 224–237. [Google Scholar] [CrossRef]
- Wu, S.C.; Hsiao, H.F.; Ho, M.L.; Hung, Y.L.; Chang, J.K.; Wang, G.J.; Wang, C.Z. Suppression of discoidin domain receptor 1 expression enhances the chondrogenesis of adipose-derived stem cells. Am. J. Physiol. Cell Physiol. 2015, 308, C685–C696. [Google Scholar] [CrossRef]
- Wu, S.C.; Chen, C.H.; Chang, J.K.; Fu, Y.C.; Wang, C.K.; Eswaramoorthy, R.; Lin, Y.S.; Wang, Y.H.; Lin, S.Y.; Wang, G.J.; et al. Hyaluronan initiates chondrogenesis mainly via CD44 in human adipose-derived stem cells. J. Appl. Physiol. 2013, 114, 1610–1618. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.C.; Chang, J.K.; Wang, C.K.; Wang, G.J.; Ho, M.L. Enhancement of chondrogenesis of human adipose derived stem cells in a hyaluronan-enriched microenvironment. Biomaterials 2010, 31, 631–640. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.Z.; Chen, S.M.; Chen, C.H.; Wang, C.K.; Wang, G.J.; Chang, J.K.; Ho, M.L. The effect of the local delivery of alendronate on human adipose-derived stem cell-based bone regeneration. Biomaterials 2010, 31, 8674–8683. [Google Scholar] [CrossRef] [PubMed]
- Chang, L.H.; Wu, S.C.; Chen, C.H.; Wang, G.J.; Chang, J.K.; Ho, M.L. Parathyroid hormone 1-34 reduces dexamethasone-induced terminal differentiation in human articular chondrocytes. Toxicology 2016, 368–369, 116–128. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Liu, X.; Yan, Z.; Wu, C.; Yang, Y.; Li, X.; Zhang, G. FastProNGS: Fast preprocessing of next-generation sequencing reads. BMC Bioinform. 2019, 20, 345. [Google Scholar] [CrossRef]
- Blankenberg, D.; Gordon, A.; Von Kuster, G.; Coraor, N.; Taylor, J.; Nekrutenko, A.; Galaxy, T. Manipulation of FASTQ data with Galaxy. Bioinformatics 2010, 26, 1783–1785. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. clusterProfiler: An R package for comparing biological themes among gene clusters. Omics 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, C.-W.; Huang, Y.-H.; Shao, P.-L.; Chang, L.-H.; Lu, C.-C.; Chen, C.-H.; Fu, Y.-C.; Ho, M.-L.; Chang, J.-K.; Wu, S.-C. Umbilical Cord Mesenchymal Stem Cell-Derived Extracellular Vesicles Enhance Chondrocyte Function by Reducing Oxidative Stress in Chondrocytes. Int. J. Mol. Sci. 2025, 26, 7683. https://doi.org/10.3390/ijms26167683
Wu C-W, Huang Y-H, Shao P-L, Chang L-H, Lu C-C, Chen C-H, Fu Y-C, Ho M-L, Chang J-K, Wu S-C. Umbilical Cord Mesenchymal Stem Cell-Derived Extracellular Vesicles Enhance Chondrocyte Function by Reducing Oxidative Stress in Chondrocytes. International Journal of Molecular Sciences. 2025; 26(16):7683. https://doi.org/10.3390/ijms26167683
Chicago/Turabian StyleWu, Che-Wei, Yao-Hui Huang, Pei-Lin Shao, Ling-Hua Chang, Cheng-Chang Lu, Chung-Hwan Chen, Yin-Chih Fu, Mei-Ling Ho, Je-Ken Chang, and Shun-Cheng Wu. 2025. "Umbilical Cord Mesenchymal Stem Cell-Derived Extracellular Vesicles Enhance Chondrocyte Function by Reducing Oxidative Stress in Chondrocytes" International Journal of Molecular Sciences 26, no. 16: 7683. https://doi.org/10.3390/ijms26167683
APA StyleWu, C.-W., Huang, Y.-H., Shao, P.-L., Chang, L.-H., Lu, C.-C., Chen, C.-H., Fu, Y.-C., Ho, M.-L., Chang, J.-K., & Wu, S.-C. (2025). Umbilical Cord Mesenchymal Stem Cell-Derived Extracellular Vesicles Enhance Chondrocyte Function by Reducing Oxidative Stress in Chondrocytes. International Journal of Molecular Sciences, 26(16), 7683. https://doi.org/10.3390/ijms26167683






