In Vitro Cross-Linking MS Reveals SMG1–UPF2–SMG7 Assembly as Molecular Partners within the NMD Surveillance
Abstract
1. Introduction
2. Results and Discussions
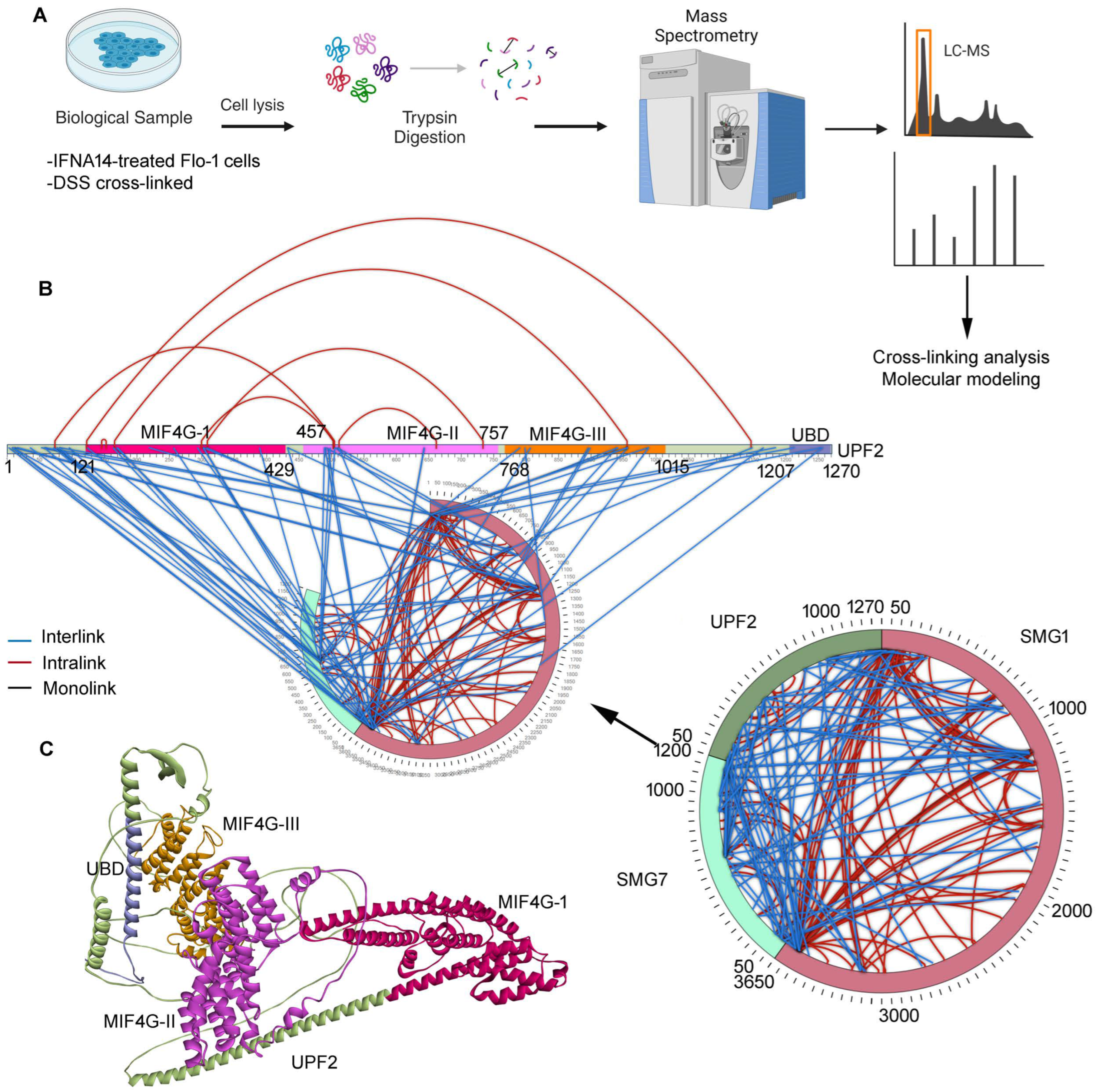
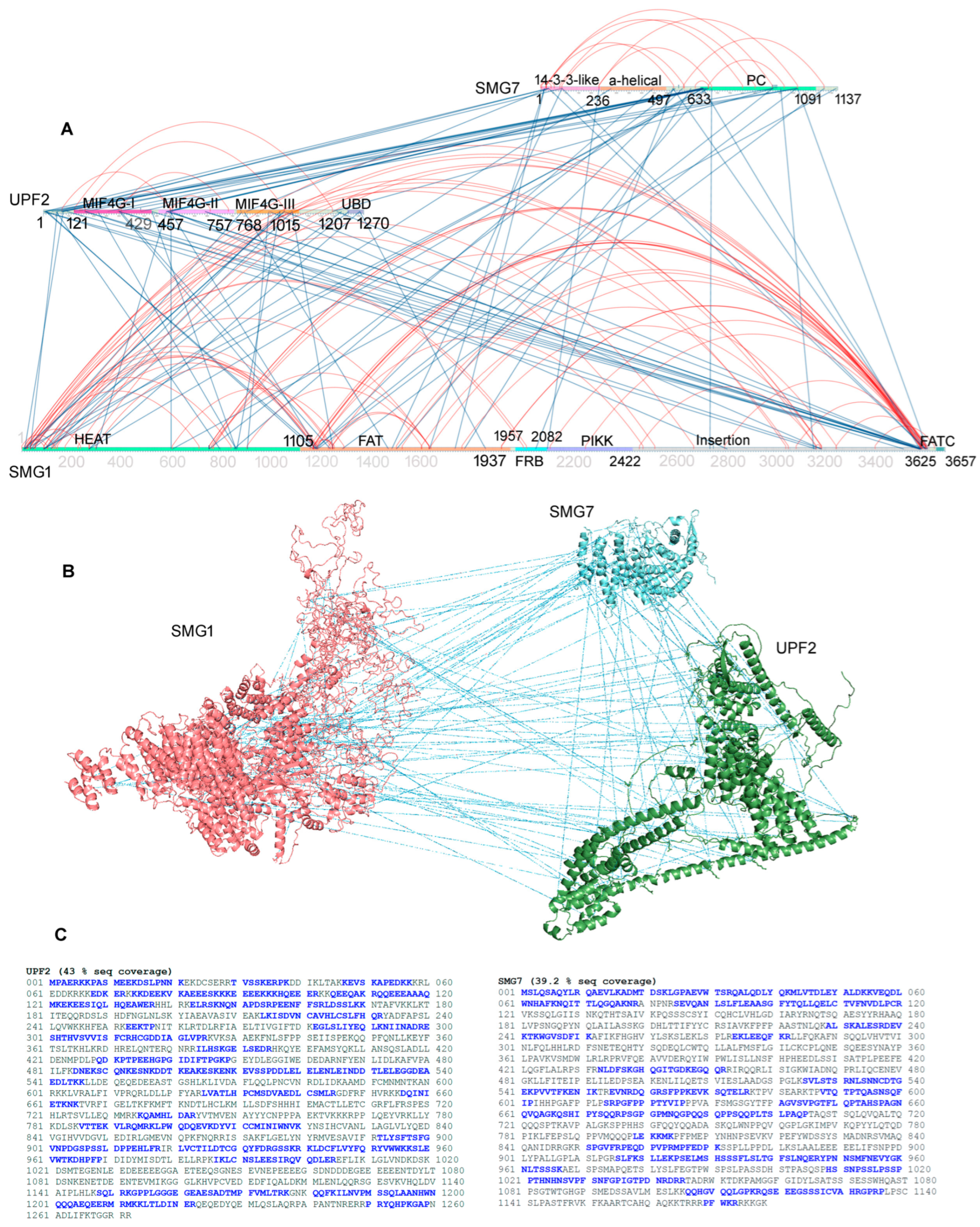
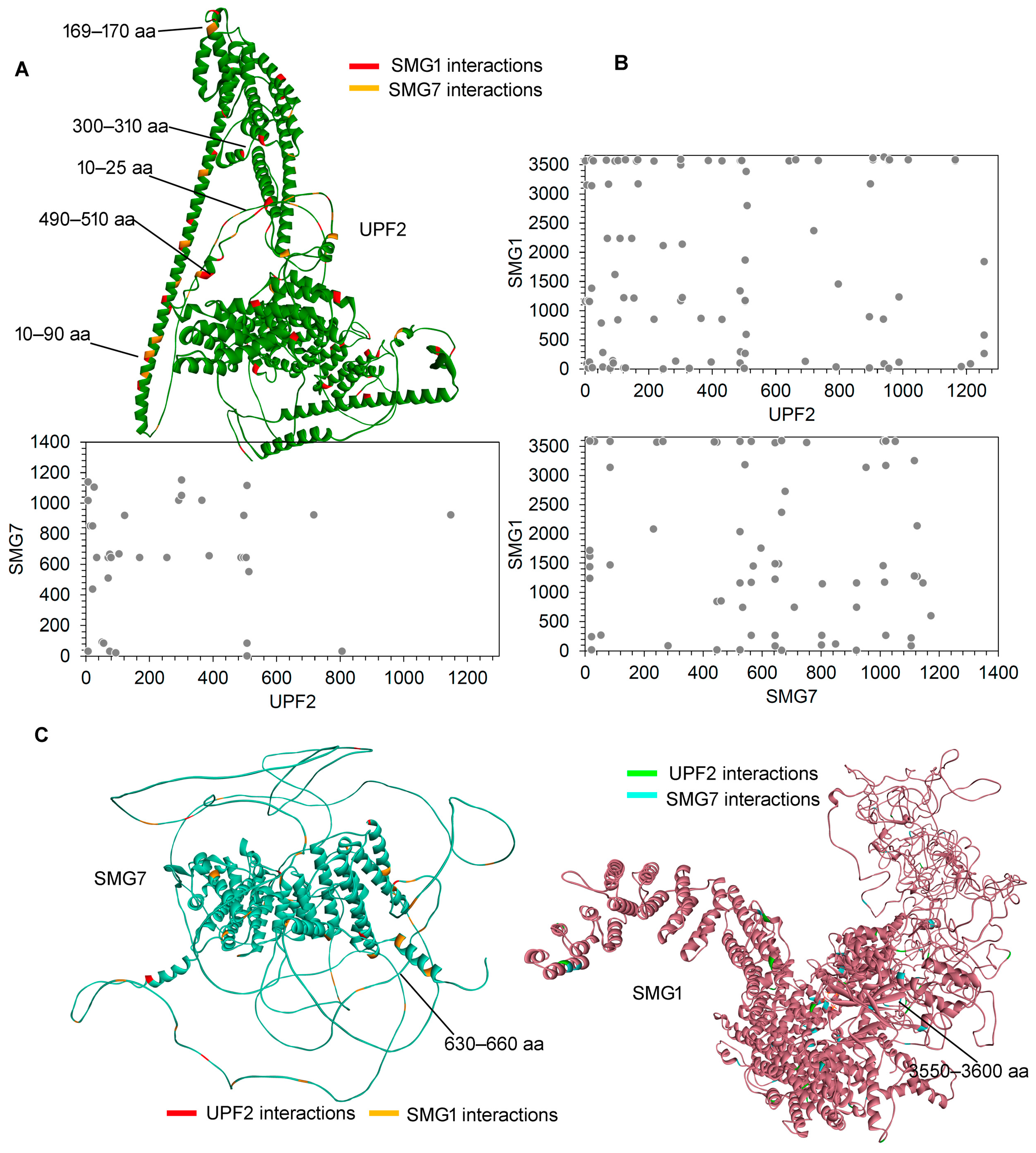
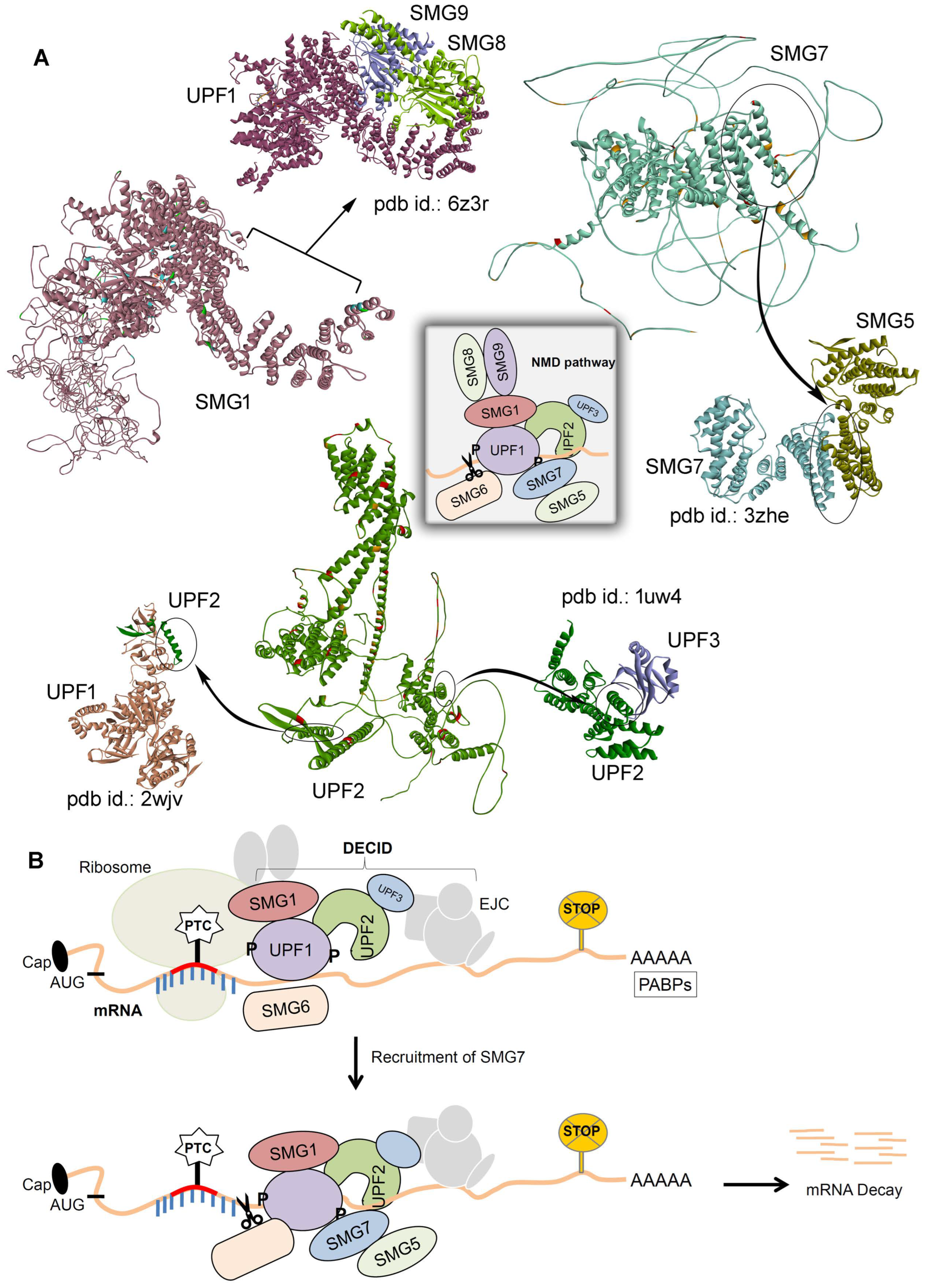
2.1. Inter- and Intramolecular Interactions between UPF2, SMG1, and SMG7
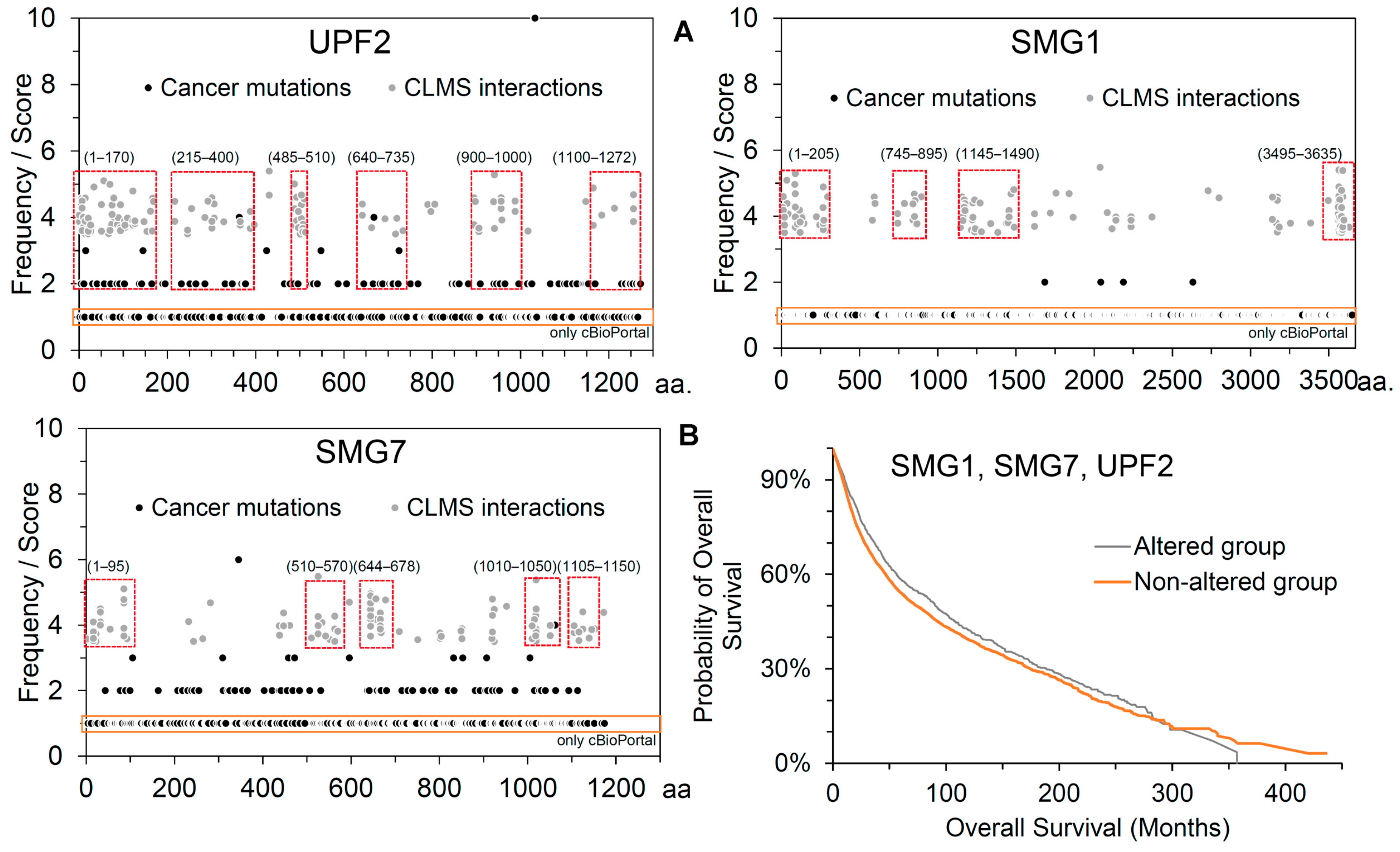
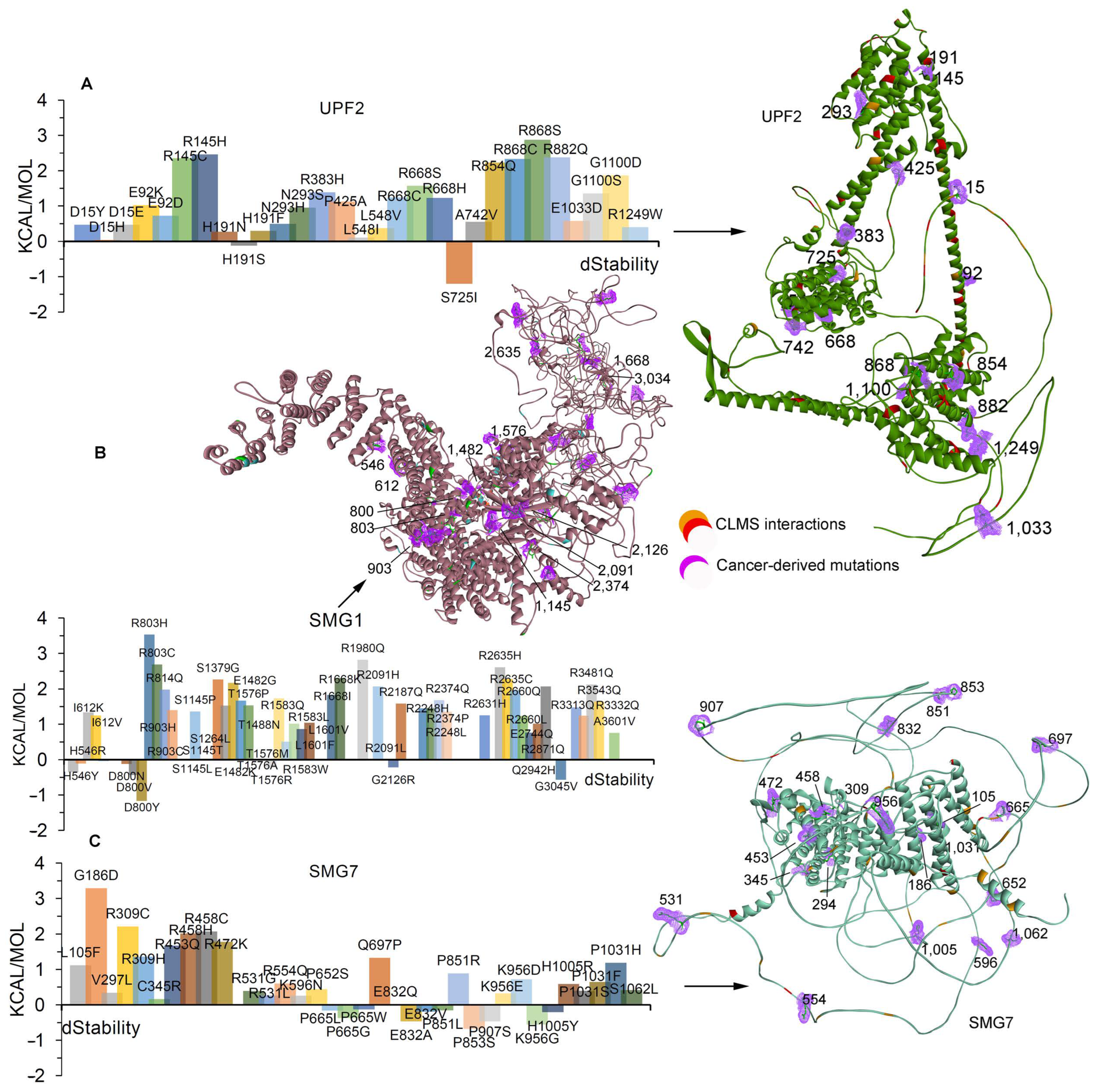
2.2. Effects of Cancer-Derived Mutations on the Identified Protein Cross-Links
3. Materials and Methods
3.1. Identification of Protein–Protein Networks within the NMD Machinery
3.2. Homology Modelling and Mutational Landscape of Cross-Linked Genes
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nickless, A.; Bailis, J.M.; You, Z. Control of gene expression through the nonsense-mediated RNA decay pathway. Cell Biosci. 2017, 7, 26. [Google Scholar] [CrossRef]
- Popp, M.W.; Maquat, L.E. Nonsense-mediated mRNA decay and cancer. Curr. Opin. Genet. Dev. 2018, 48, 44–50. [Google Scholar] [CrossRef]
- Gatfield, D. Nonsense-mediated mRNA decay in Drosophila:at the intersection of the yeast and mammalian pathways. EMBO J. 2003, 22, 3960–3970. [Google Scholar] [CrossRef]
- Bühler, M.; Steiner, S.; Mohn, F.; Paillusson, A.; Mühlemann, O. EJC-independent degradation of nonsense immunoglobulin-μ mRNA depends on 3′ UTR length. Nat. Struct. Mol. Biol. 2006, 13, 462–464. [Google Scholar] [CrossRef] [PubMed]
- Bobadilla, J.L.; Macek, M.; Fine, J.P.; Farrell, P.M. Cystic fibrosis: A worldwide analysis ofCFTR mutations?correlation with incidence data and application to screening. Hum. Mutat. 2002, 19, 575–606. [Google Scholar] [CrossRef] [PubMed]
- Lykke-Andersen, J.; Shu, M.-D.; Steitz, J.A. Human upf proteins target an mRNA for nonsense-mediated decay when bound downstream of a termination Codon. Cell 2000, 103, 1121–1131. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Z.; Saito, K.; Pisarev, A.V.; Wada, M.; Pisareva, V.P.; Pestova, T.V.; Gajda, M.; Round, A.; Kong, C.; Lim, M.; et al. Structural insights into eRF3 and stop codon recognition by eRF1. Genes Dev. 2009, 23, 1106–1118. [Google Scholar] [CrossRef] [PubMed]
- Martin, L.; Grigoryan, A.; Wang, D.; Wang, J.; Breda, L.; Rivella, S.; Cardozo, T.; Gardner, L.B. Identification and characterization of small molecules that inhibit nonsense-mediated RNA decay and suppress nonsense p53 mutations. Cancer Res. 2014, 74, 3104–3113. [Google Scholar] [CrossRef] [PubMed]
- Boehm, V.; Gehring, N.H. Exon junction complexes: Supervising the gene expression assembly line. Trends Genet. 2016, 32, 724–735. [Google Scholar] [CrossRef] [PubMed]
- Kashima, I.; Yamashita, A.; Izumi, N.; Kataoka, N.; Morishita, R.; Hoshino, S.; Ohno, M.; Dreyfuss, G.; Ohno, S. Binding of a novel SMG-1–Upf1–eRF1–eRF3 complex (SURF) to the exon junction complex triggers Upf1 phosphorylation and nonsense-mediated mRNA decay. Genes Dev. 2006, 20, 355–367. [Google Scholar] [CrossRef] [PubMed]
- Schoenberg, D.R.; Maquat, L.E. Regulation of cytoplasmic mRNA decay. Nat. Rev. Genet. 2012, 13, 246–259. [Google Scholar] [CrossRef] [PubMed]
- Chan, W.-K.; Bhalla, A.D.; Le Hir, H.; Nguyen, L.S.; Huang, L.; Gécz, J.; Wilkinson, M.F. A UPF3-mediated regulatory switch that maintains RNA surveillance. Nat. Struct. Mol. Biol. 2009, 16, 747–753. [Google Scholar] [CrossRef]
- Padariya, M.; Fahraeus, R.; Hupp, T.; Kalathiya, U. Molecular determinants and specificity of mRNA with alternatively-spliced UPF1 isoforms, influenced by an insertion in the ‘regulatory loop’. Int. J. Mol. Sci. 2021, 22, 12744. [Google Scholar] [CrossRef] [PubMed]
- Kalathiya, U.; Padariya, M.; Pawlicka, K.; Verma, C.S.; Houston, D.; Hupp, T.R.; Alfaro, J.A. Insights into the effects of cancer associated mutations at the UPF2 and ATP-binding sites of NMD master regulator: UPF1. Int. J. Mol. Sci. 2019, 20, 5644. [Google Scholar] [CrossRef]
- Wu, C.; Roy, B.; He, F.; Yan, K.; Jacobson, A. Poly(A)-binding protein regulates the efficiency of translation termination. Cell Rep. 2020, 33, 108399. [Google Scholar] [CrossRef] [PubMed]
- Lueck, J.D.; Yoon, J.S.; Perales-Puchalt, A.; Mackey, A.L.; Infield, D.T.; Behlke, M.A.; Pope, M.R.; Weiner, D.B.; Skach, W.R.; McCray, P.B., Jr.; et al. Engineered transfer RNAs for suppression of premature termination codons. Nat. Commun. 2019, 10, 822. [Google Scholar] [CrossRef]
- Bordeira-Carriço, R.; Pêgo, A.P.; Santos, M.; Oliveira, C. Cancer syndromes and therapy by stop-codon readthrough. Trends Mol. Med. 2012, 18, 667–678. [Google Scholar] [CrossRef]
- Nomakuchi, T.T.; Rigo, F.; Aznarez, I.; Krainer, A.R. Antisense oligonucleotide–directed inhibition of nonsense-mediated mRNA decay. Nat. Biotechnol. 2016, 34, 164–166. [Google Scholar] [CrossRef]
- Prokhorova, I.; Altman, R.B.; Djumagulov, M.; Shrestha, J.P.; Urzhumtsev, A.; Ferguson, A.; Chang, C.-W.T.; Yusupov, M.; Blanchard, S.C.; Yusupova, G. Aminoglycoside interactions and impacts on the eukaryotic ribosome. Proc. Natl. Acad. Sci. USA 2017, 114, E10899–E10908. [Google Scholar] [CrossRef]
- Kalathiya, U.; Padariya, M.; Faktor, J.; Coyaud, E.; Alfaro, J.A.; Fahraeus, R.; Hupp, T.R.; Goodlett, D.R. Interfaces with structure dynamics of the workhorses from cells revealed through cross-linking mass spectrometry (CLMS). Biomolecules 2021, 11, 382. [Google Scholar] [CrossRef]
- Singh, A.; Padariya, M.; Faktor, J.; Kote, S.; Mikac, S.; Dziadosz, A.; Lam, T.W.; Brydon, J.; Wear, M.A.; Ball, K.L.; et al. Identification of novel interferon responsive protein partners of human leukocyte antigen A (HLA-A) using cross-linking mass spectrometry (CLMS) approach. Sci. Rep. 2022, 12, 19422. [Google Scholar] [CrossRef]
- Guerrero, C.; Tagwerker, C.; Kaiser, P.; Huang, L. An integrated mass spectrometry-based proteomic approach. Mol. Cell. Proteom. 2006, 5, 366–378. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [PubMed]
- Pawlicka, K.; Kalathiya, U.; Alfaro, J. Nonsense-mediated mRNA decay: Pathologies and the potential for novel therapeutics. Cancers 2020, 12, 765. [Google Scholar] [CrossRef] [PubMed]
- McCann, J.J.; Fleenor, D.E.; Chen, J.; Lai, C.-H.; Bass, T.E.; Kastan, M.B. Participation of ATM, SMG1, and DDX5 in a DNA damage-induced alternative splicing pathway. Radiat. Res. 2023, 199, 406–421. [Google Scholar] [CrossRef] [PubMed]
- Leeksma, A.C.; Derks, I.A.M.; Garrick, B.; Jongejan, A.; Colombo, M.; Bloedjes, T.; Trowe, T.; Leisten, J.C.; Howarth, M.; Malek, M.; et al. SMG1, a nonsense-mediated mRNA decay (NMD) regulator, as a candidate therapeutic target in multiple myeloma. Mol. Oncol. 2023, 17, 284–297. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Chen, P.; Qi, D.; Chen, L. Glaucocalyxin A inhibits the malignancies of gastric cancer cells by downregulating MDM2 and RNF6 via MiR-3658 and the SMG1-UPF mRNA decay pathway. Front. Oncol. 2022, 12, 871169. [Google Scholar] [CrossRef] [PubMed]
- Nasif, S.; Colombo, M.; Uldry, A.-C.; Schröder, M.S.; de Brot, S.; Mühlemann, O. Inhibition of nonsense-mediated mRNA decay reduces the tumorigenicity of human fibrosarcoma cells. NAR Cancer 2023, 5, zcad048. [Google Scholar] [CrossRef]
- Steiner, A.J.; Zheng, Y.; Tang, Y. Characterization of a rhabdomyosarcoma reveals a critical role for SMG7 in cancer cell viability and tumor growth. Sci. Rep. 2023, 13, 10152. [Google Scholar] [CrossRef] [PubMed]
- Cowen, L.E.; Luo, H.; Tang, Y. Characterization of SMG7 14-3-3-like domain reveals phosphoserine binding-independent regulation of p53 and UPF1. Sci. Rep. 2019, 9, 13097. [Google Scholar] [CrossRef]
- Luo, H.; Cowen, L.; Yu, G.; Jiang, W.; Tang, Y. SMG7 is a critical regulator of p53 stability and function in DNA damage stress response. Cell Discov. 2016, 2, 15042. [Google Scholar] [CrossRef]
- Berman, H.M. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Langer, L.M.; Gat, Y.; Bonneau, F.; Conti, E. Structure of substrate-bound SMG1-8-9 kinase complex reveals molecular basis for phosphorylation specificity. Elife 2020, 9, e57127. [Google Scholar] [CrossRef] [PubMed]
- Jonas, S.; Weichenrieder, O.; Izaurralde, E. An unusual arrangement of two 14-3-3-like domains in the SMG5–SMG7 heterodimer is required for efficient nonsense-mediated mRNA decay. Genes Dev. 2013, 27, 211–225. [Google Scholar] [CrossRef] [PubMed]
- Clerici, M.; Mourão, A.; Gutsche, I.; Gehring, N.H.; Hentze, M.W.; Kulozik, A.; Kadlec, J.; Sattler, M.; Cusack, S. Unusual bipartite mode of interaction between the nonsense-mediated decay factors, UPF1 and UPF2. EMBO J. 2009, 28, 2293–2306. [Google Scholar] [CrossRef] [PubMed]
- Kadlec, J.; Izaurralde, E.; Cusack, S. The structural basis for the interaction between nonsense-mediated mRNA decay factors UPF2 and UPF3. Nat. Struct. Mol. Biol. 2004, 11, 330–337. [Google Scholar] [CrossRef] [PubMed]
- Behm-Ansmant, I.; Izaurralde, E. Quality control of gene expression: A stepwise assembly pathway for the surveillance complex that triggers nonsense-mediated mRNA decay. Genes Dev. 2006, 20, 391–398. [Google Scholar] [CrossRef] [PubMed]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio Cancer Genomics Portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef] [PubMed]
- Tate, J.G.; Bamford, S.; Jubb, H.C.; Sondka, Z.; Beare, D.M.; Bindal, N.; Boutselakis, H.; Cole, C.G.; Creatore, C.; Dawson, E.; et al. COSMIC: The catalogue of somatic mutations in cancer. Nucleic Acids Res. 2019, 47, D941–D947. [Google Scholar] [CrossRef]
- Lima, D.B.; Melchior, J.T.; Morris, J.; Barbosa, V.C.; Chamot-Rooke, J.; Fioramonte, M.; Souza, T.A.C.B.; Fischer, J.S.G.; Gozzo, F.C.; Carvalho, P.C.; et al. Characterization of homodimer interfaces with cross-linking mass spectrometry and isotopically labeled proteins. Nat. Protoc. 2018, 13, 431–458. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Padariya, M.; Vojtesek, B.; Hupp, T.; Kalathiya, U. In Vitro Cross-Linking MS Reveals SMG1–UPF2–SMG7 Assembly as Molecular Partners within the NMD Surveillance. Int. J. Mol. Sci. 2024, 25, 3182. https://doi.org/10.3390/ijms25063182
Padariya M, Vojtesek B, Hupp T, Kalathiya U. In Vitro Cross-Linking MS Reveals SMG1–UPF2–SMG7 Assembly as Molecular Partners within the NMD Surveillance. International Journal of Molecular Sciences. 2024; 25(6):3182. https://doi.org/10.3390/ijms25063182
Chicago/Turabian StylePadariya, Monikaben, Borivoj Vojtesek, Ted Hupp, and Umesh Kalathiya. 2024. "In Vitro Cross-Linking MS Reveals SMG1–UPF2–SMG7 Assembly as Molecular Partners within the NMD Surveillance" International Journal of Molecular Sciences 25, no. 6: 3182. https://doi.org/10.3390/ijms25063182
APA StylePadariya, M., Vojtesek, B., Hupp, T., & Kalathiya, U. (2024). In Vitro Cross-Linking MS Reveals SMG1–UPF2–SMG7 Assembly as Molecular Partners within the NMD Surveillance. International Journal of Molecular Sciences, 25(6), 3182. https://doi.org/10.3390/ijms25063182







