Disparate Effects of Two Clerodane Diterpenes of Giant Goldenrod (Solidago gigantea Ait.) on Bacillus spizizenii
Abstract
1. Introduction
2. Results and Discussion
2.1. Antibacterial Effects of Different Diterpenes Isolated from Giant Goldenrod Root
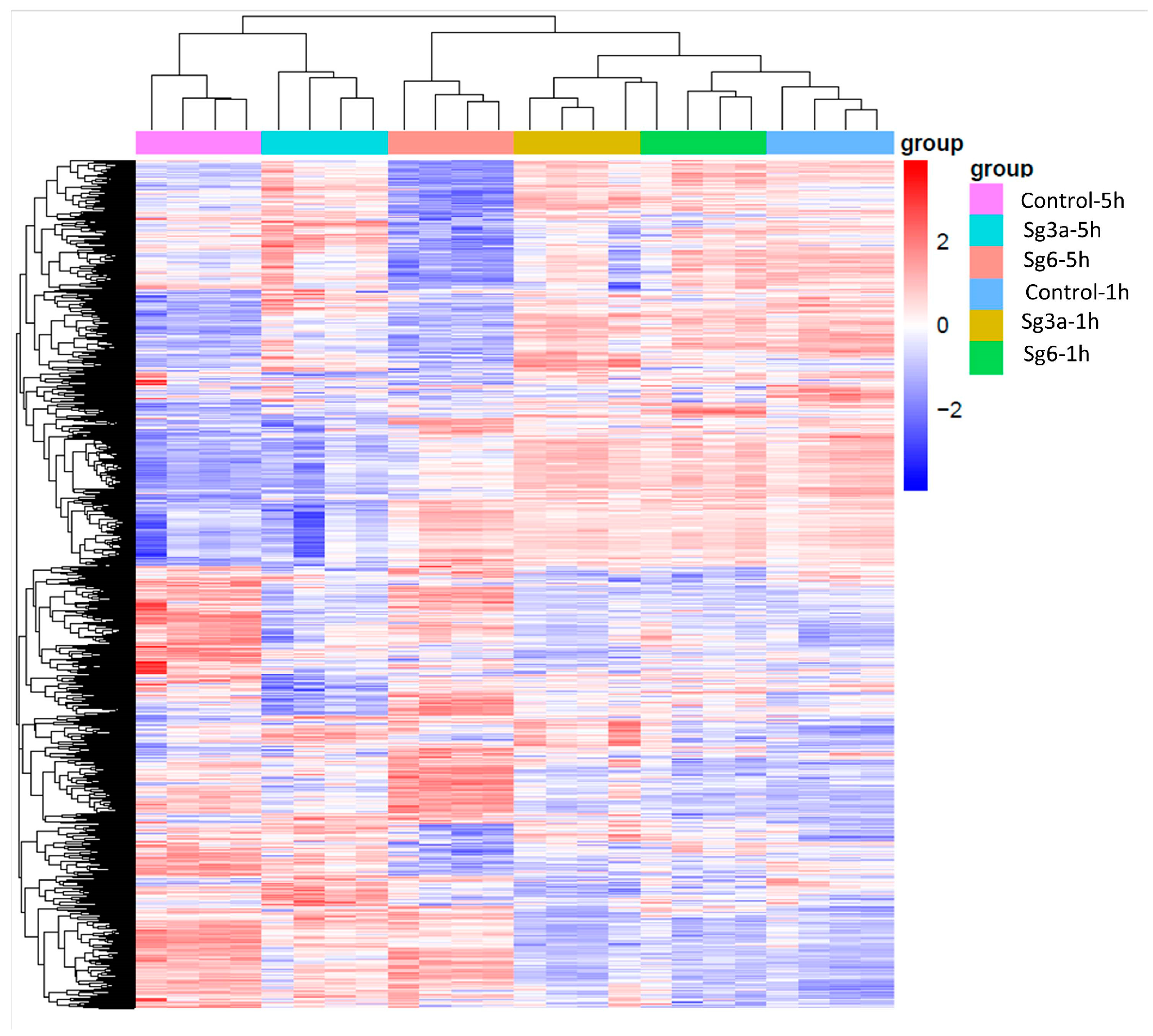
2.2. Genome Wide Transcriptome Analysis Induced by Sg3a and Sg6 Diterpenes
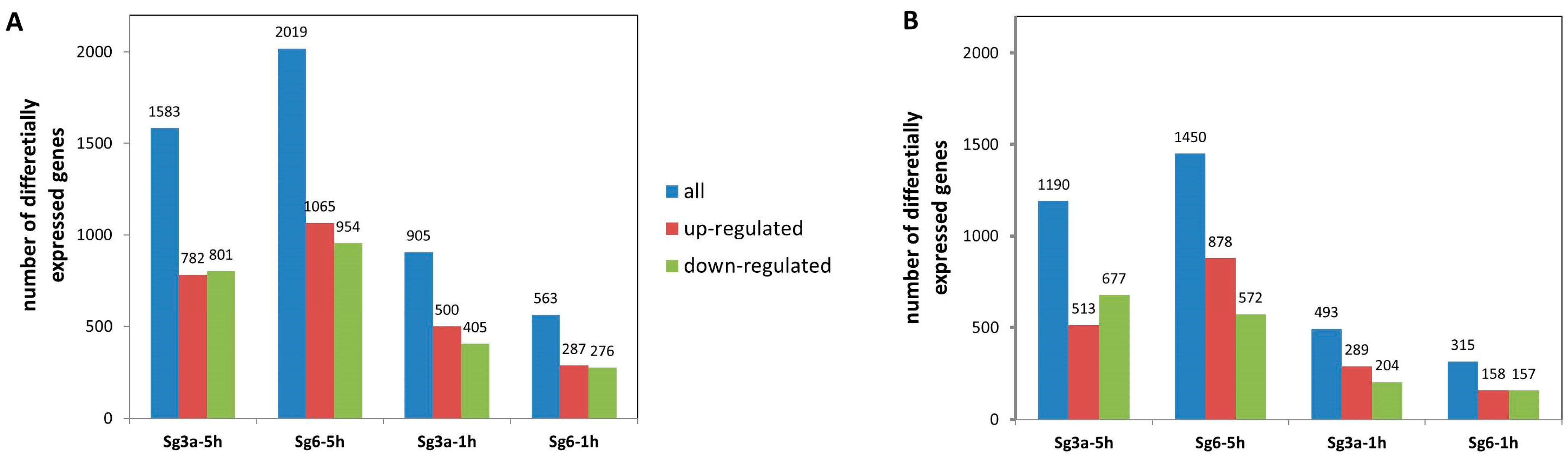
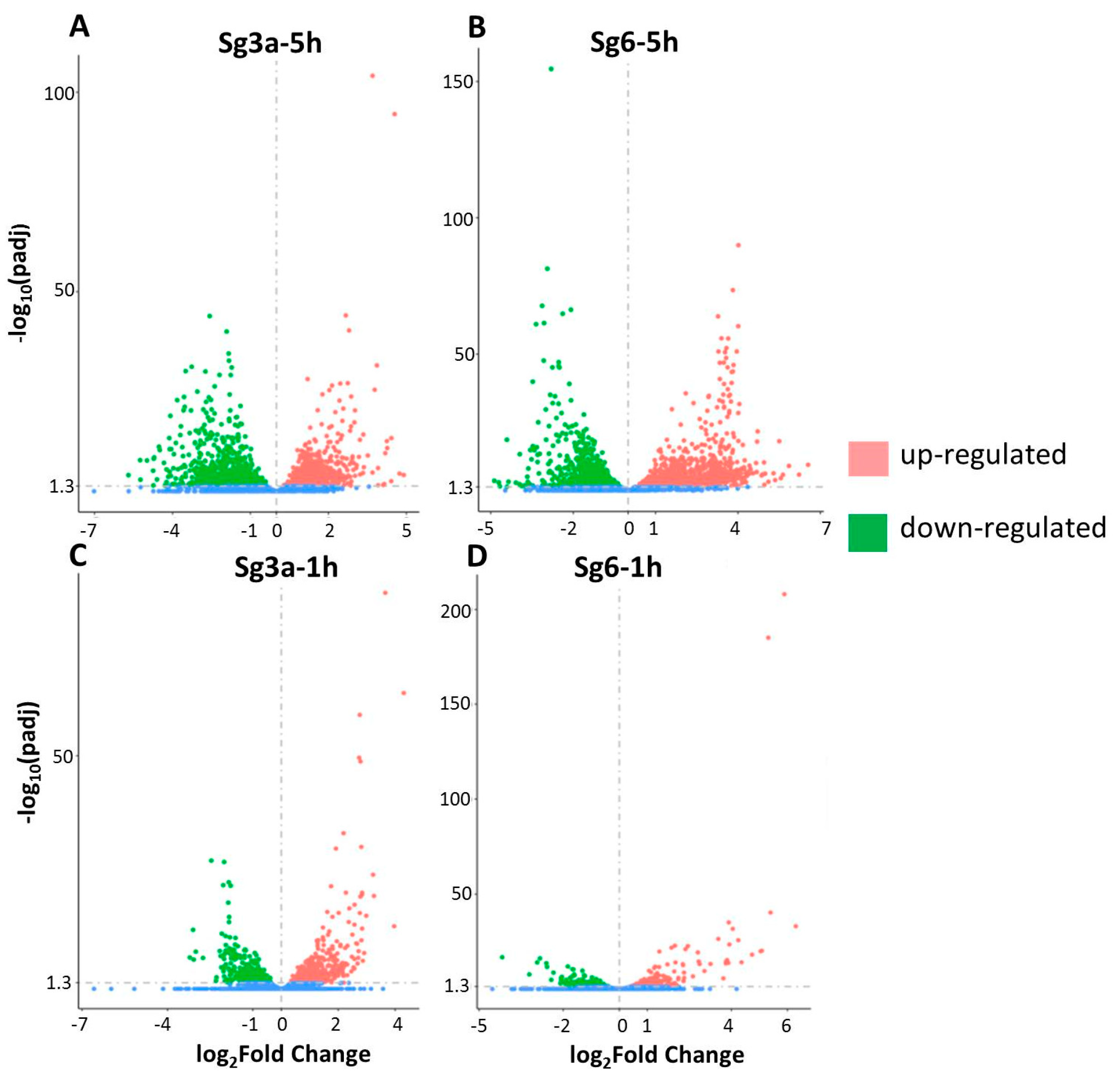
2.3. Differentially Expressed Genes in B. spizizenii after Sg3a- and Sg6-Treatments
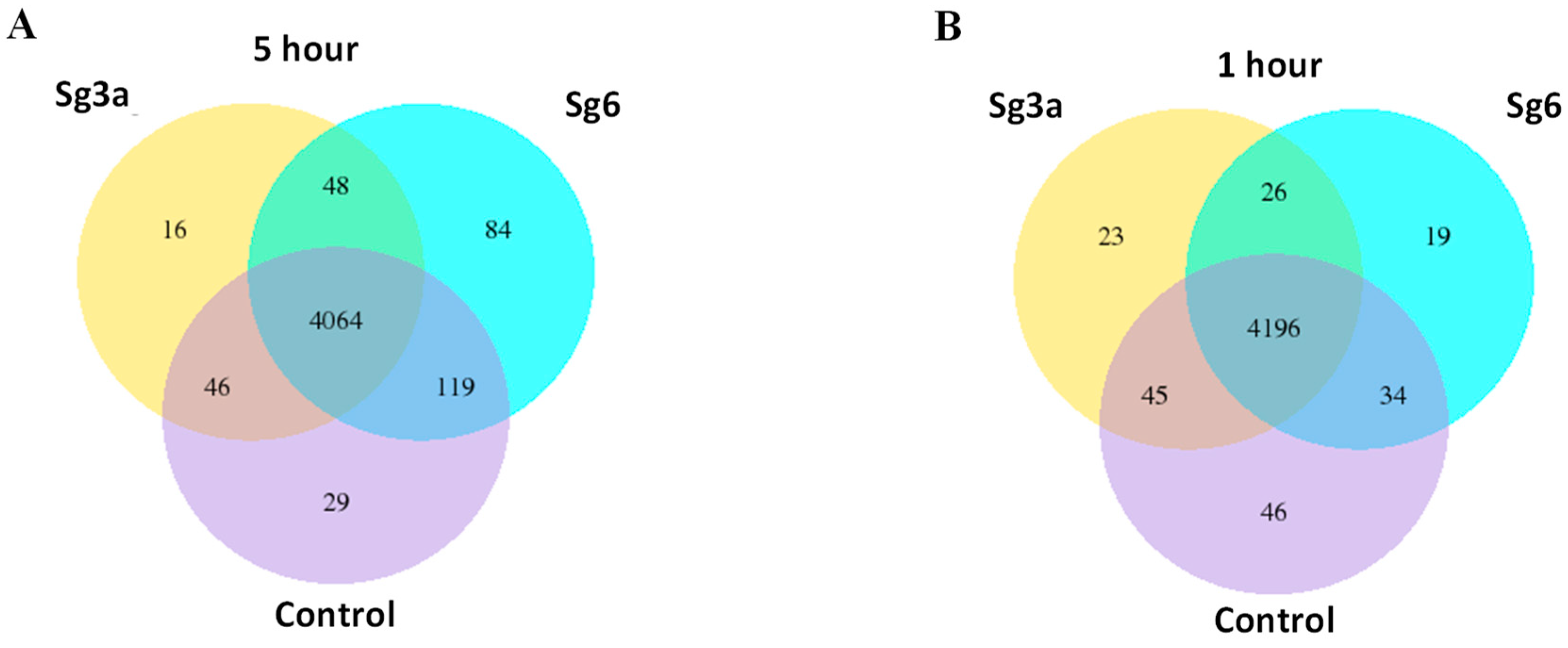
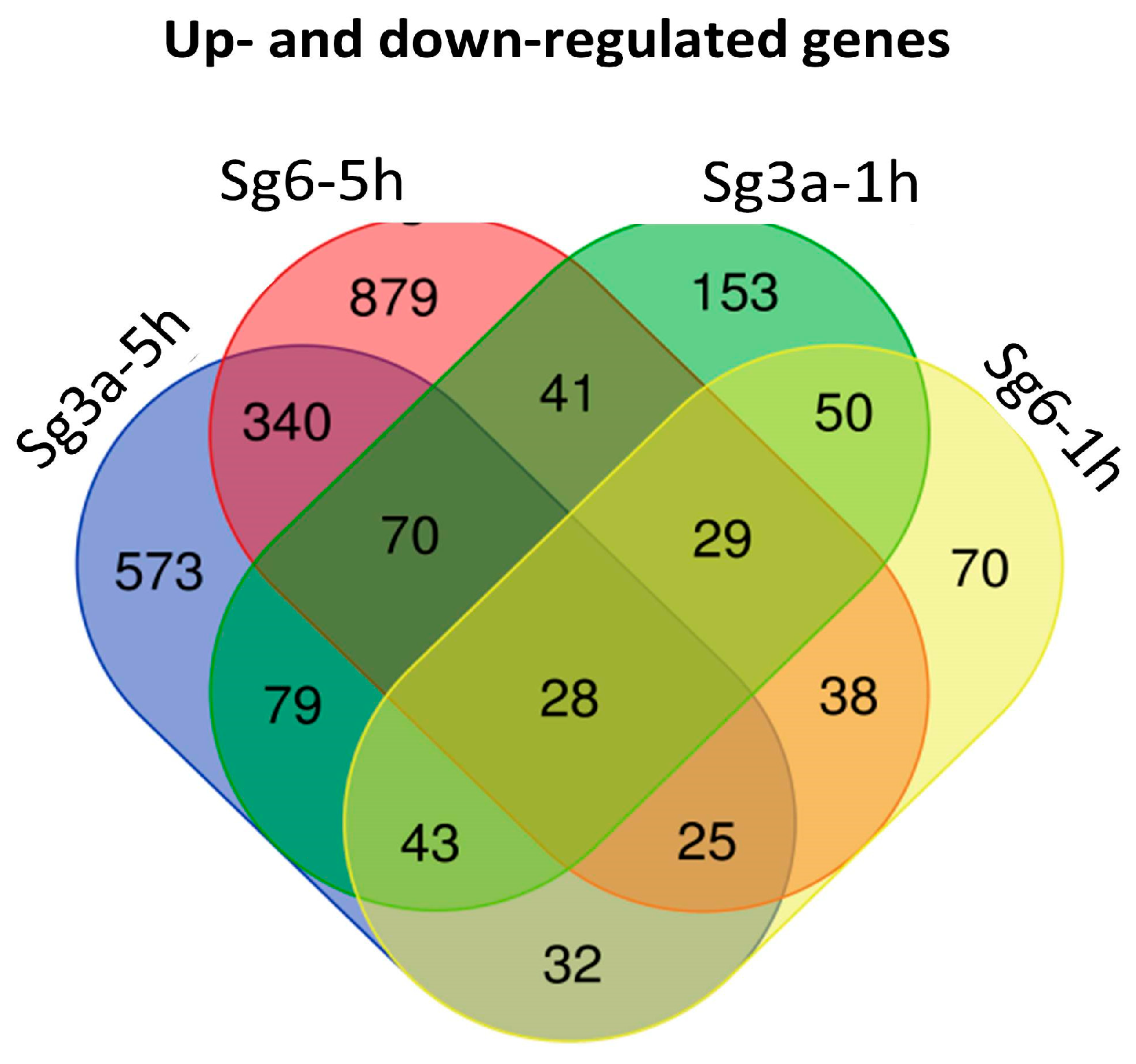
2.4. Overlapping B. spizizenii Genes That Changed Their Activity after Different Types of Treatments
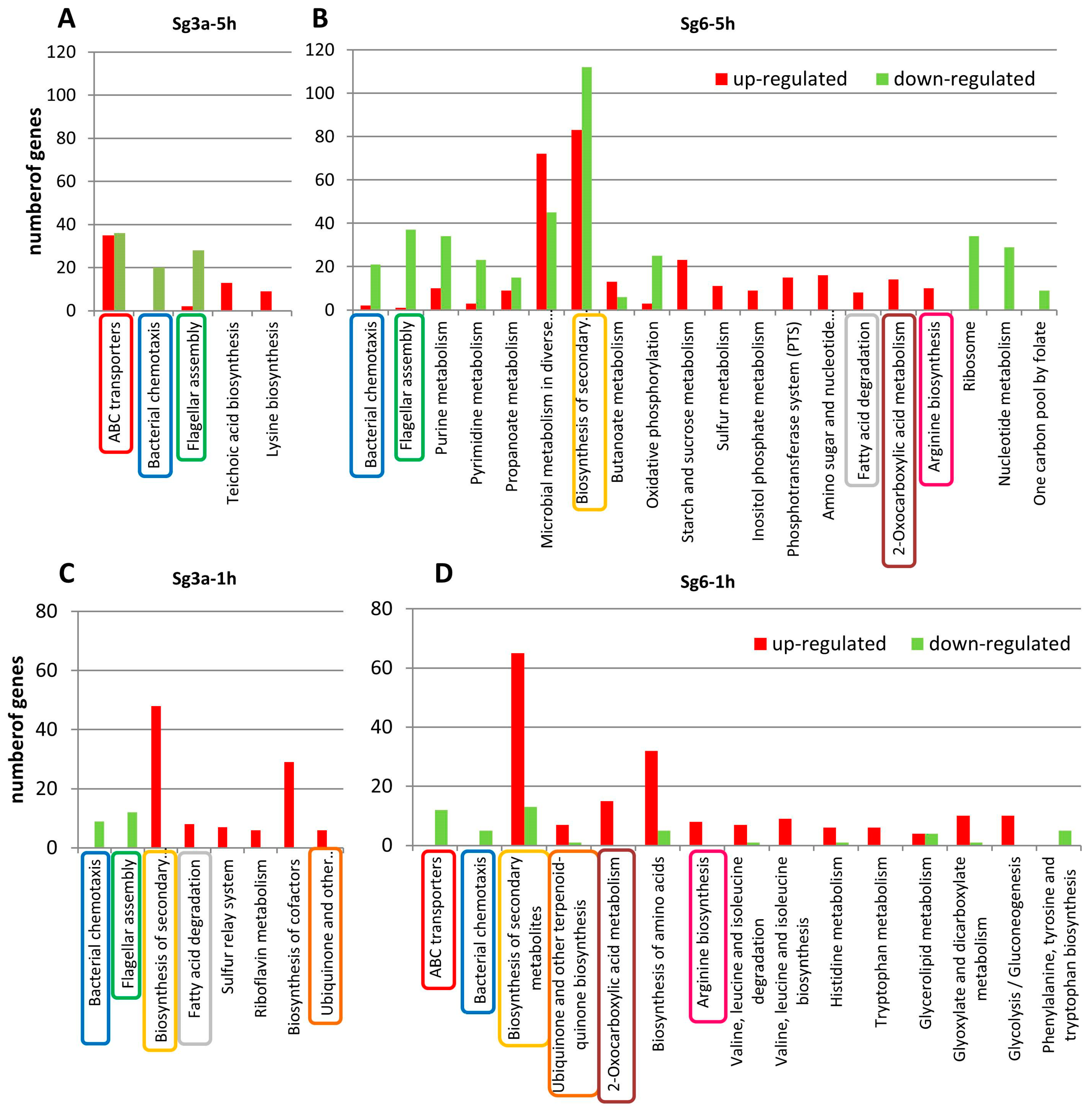
2.5. Enrichment Analyses Identified Significantly Up- or Down-Regulated Genes and Pathways Affected by Diterpene Treatments
2.6. Common Processes with Altered Gene Activities after Diterpene Treatments
2.7. Treatment Specific Up-Regulated Processes Enriched after Diterpene Treatments
2.8. Treatment Specific Down-Regulated Processes Enriched after Diterpene Treatments
2.9. Comparison of Effect of Diterpene Treatments on Transcriptome with Changes Induced under Different Environmental and Nutritional Conditions
3. Materials and Methods
3.1. Bacterial Strains and Microdilution Assays
3.2. Treatment of B. spizizenii with Diterpenes Sg3a and Sg6 for RNA Purification
3.3. RNA Purification from B. spizizenii
3.4. RNA Sequencing, Data Processing and Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sundin, G.W.; Wang, N. Antibiotic resistance in plant-pathogenic bacteria. Annu. Rev. Phytopathol. 2018, 56, 161–180. [Google Scholar] [CrossRef]
- Thomford, N.; Senthebane, D.; Rowe, A.; Munro, D.; Seele, P.; Maroyi, A.; Dzobo, K. Natural products for drug discovery in the 21st century: Innovations for novel drug discovery. Int. J. Mol. Sci. 2018, 19, 1578. [Google Scholar] [CrossRef]
- Choudhury, D.; Dobhal, P.; Srivastava, S.; Saha, S.; Kundu, S.S. Role of botanical plant extracts to control plant pathogens. Indian J. Agric. Res. 2018, 52, 341–346. [Google Scholar] [CrossRef]
- Schlaepfer, D.R.; Edwards, P.J.; Billeter, R. Why only tetraploid Solidago gigantea (Asteraceae) became invasive: A common garden comparison of ploidy levels. Oecologia 2010, 163, 661–673. [Google Scholar] [CrossRef]
- Szymura, M.; Szymura, T.H. Interactions between alien goldenrods (Solidago and Euthamia species) and comparison with native species in Central Europe. Flora-Morphol. Distrib. Funct. Ecol. Plants 2016, 218, 51–61. [Google Scholar] [CrossRef]
- Jakobs, G.; Weber, E.; Edwards, P.J. Introduced plants of the invasive Solidago gigantea (Asteraceae) are larger and grow denser than conspecifics in the native range. Divers. Distrib. 2004, 10, 11–19. [Google Scholar] [CrossRef]
- Reznicek, G.; Jurenitsch, J.; Michl, G.; Haslinger, E. The first structurally confirmed saponin from Solidago gigantea: Structure elucidation by modern NMR techniques. Tetrahedron Lett. 1989, 30, 4097–4100. [Google Scholar] [CrossRef]
- Johnson, R.H.; Hull-Sanders, H.M.; Meyer, G.A. Comparison of foliar terpenes between native and invasive Solidago gigantea. Biochem. Syst. Ecol. 2007, 35, 821–830. [Google Scholar] [CrossRef]
- Radusiene, J.; Marska, M.; Ivanauskas, L.; Jakstas, V.; Karpaviciene, B. Assessment of phenolic compound accumulation in two widespread goldenrods. Ind. Crops Prod. 2015, 63, 158–166. [Google Scholar] [CrossRef]
- Benelli, G.; Pavela, R.; Cianfaglione, K.; Nagy, D.U.; Canale, A.; Maggi, F. Evaluation of two invasive plant invaders in Europe (Solidago canadensis and Solidago gigantea) as possible sources of botanical insecticides. J. Pest Sci. 2019, 92, 805–821. [Google Scholar] [CrossRef]
- Zekič, J.; Vovk, I.; Glavnik, V. Extraction and analyses of flavonoids and phenolic acids from Canadian goldenrod and giant goldenrod. Forests 2020, 12, 40. [Google Scholar] [CrossRef]
- Móricz, Á.M.; Krüzselyi, D.; Ott, P.G.; Garádi, Z.; Béni, S.; Morlock, G.E.; Bakonyi, J. Bioactive clerodane diterpenes of giant goldenrod (Solidago gigantea Ait.) root extract. J. Chromatogr. A 2021, 1635, 461727. [Google Scholar] [CrossRef]
- Baglyas, M.; Ott, P.G.; Garádi, Z.; Glavnik, V.; Béni, S.; Vovk, I.; Móricz, Á.M. High-performance thin-layer chromatography—antibacterial assay first reveals bioactive clerodane diterpenes in giant goldenrod (Solidago gigantea Ait.). J. Chromatogr. A 2022, 1677, 463308. [Google Scholar] [CrossRef]
- Kalemba, D.; Marschall, H.; Bradesi, P. Constituents of the essential oil of Solidago gigantea Ait. Flavour Fragr. J. 2001, 16, 19–26. [Google Scholar] [CrossRef]
- Webster, D.; Taschereau, P.; Belland, R.J.; Sand, C.; Rennie, R.P. Antifungal activity of medicinal plant extracts; preliminary screening studies. J. Ethnopharmacol. 2008, 115, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Kołodziej, B.; Kowalski, R.; Kedzia, B. Antibacterial and antimutagenic activity of extracts aboveground parts of three Solidago species: Solidago virgaurea L., Solidago canadensis L. and Solidago gigantea Ait. J. Med. Plants Res. 2011, 5, 6770–6779. [Google Scholar] [CrossRef]
- Anžlovar, S.; Koce, J.D. Antibacterial and antifungal activity of aqueous and organic extracts from indigenous and invasive species of goldenrod (Solidago spp.) grown in Slovenia. Phyton 2014, 54, 135–147. [Google Scholar]
- Krüzselyi, D.; Bakonyi, J.; Ott, P.G.; Darcsi, A.; Csontos, P.; Morlock, G.E.; Móricz, Á.M. Goldenrod root compounds active against crop pathogenic fungi. J. Agric. Food Chem. 2021, 69, 12686–12694. [Google Scholar] [CrossRef]
- Ninkuu, V.; Zhang, L.; Yan, J.; Fu, Z.; Yang, T.; Zeng, H. Biochemistry of terpenes and recent advances in plant protection. Int. J. Mol. Sci. 2021, 22, 5710. [Google Scholar] [CrossRef]
- Li, R.; Morris-Natschke, S.L.; Lee, K.-H. Clerodane diterpenes: Sources, structures, and biological activities. Nat. Prod. Rep. 2016, 33, 1166–1226. [Google Scholar] [CrossRef] [PubMed]
- Urzúa, A.; Torres, R.; Mendoza, L.; Monache, F.D. Antibacterial new clerodane diterpenes from the surface of Haplopappus foliosus. Planta Med. 2003, 69, 675–677. [Google Scholar] [CrossRef]
- Marthanda Murthy, M.; Subramanyam, M.; Hima Bindu, M.; Annapurna, J. Antimicrobial activity of clerodane diterpenoids from Polyalthia longifolia seeds. Fitoterapia 2005, 76, 336–339. [Google Scholar] [CrossRef]
- Mahizan, N.A.; Yang, S.-K.; Moo, C.-L.; Song, A.A.-L.; Chong, C.-M.; Chong, C.-W.; Abushelaibi, A.; Lim, S.-H.E.; Lai, K.-S. Terpene derivatives as a potential agent against antimicrobial resistance (AMR) pathogens. Molecules 2019, 24, 2631. [Google Scholar] [CrossRef] [PubMed]
- Wiart, C.; Kathirvalu, G.; Raju, C.S.; Nissapatorn, V.; Rahmatullah, M.; Paul, A.K.; Rajagopal, M.; Sathiya Seelan, J.S.; Rusdi, N.A.; Lanting, S.; et al. Antibacterial and antifungal terpenes from the medicinal angiosperms of Asia and the Pacific: Haystacks and gold needles. Molecules 2023, 28, 3873. [Google Scholar] [CrossRef]
- Guimarães, A.C.; Meireles, L.M.; Lemos, M.F.; Guimarães, M.C.C.; Endringer, D.C.; Fronza, M.; Scherer, R. Antibacterial activity of terpenes and terpenoids present in essential oils. Molecules 2019, 24, 2471. [Google Scholar] [CrossRef]
- Su, T.; Qiu, Y.; Hua, X.; Ye, B.; Luo, H.; Liu, D.; Qu, P.; Qiu, Z. Novel opportunity to reverse antibiotic resistance: To explore traditional Chinese medicine with potential activity against antibiotics-resistance bacteria. Front. Microbiol. 2020, 11, 610070. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Yang, L.; Zou, Y.; Luo, S.; Wang, X.; Liang, Y.; Du, Y.; Feng, R.; Wei, Q. Antibacterial activity and mechanism of three isomeric terpineols of Cinnamomum longepaniculatum leaf oil. Folia Microbiol. 2021, 66, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Haraguchi, H.; Oike, S.; Muroi, H.; Kubo, I. Mode of Antibacterial action of totarol, a diterpene from Podocarpus nagi. Planta Med. 1996, 62, 122–125. [Google Scholar] [CrossRef] [PubMed]
- Torres-Bustos, J.; Farías, L.; Urzúa, A.; Mendoza, L.; Wilkens, M. The diterpenoid ent -16-kauren-19-oic acid acts as an uncoupler of the bacterial respiratory chain. Planta Med. 2009, 75, 823–828. [Google Scholar] [CrossRef]
- Zengin, H.; Baysal, A. Antibacterial and antioxidant activity of essential oil terpenes against pathogenic and spoilage-forming bacteria and cell structure-activity relationships evaluated by SEM microscopy. Molecules 2014, 19, 17773–17798. [Google Scholar] [CrossRef]
- Yang, S.-K.; Yusoff, K.; Ajat, M.; Thomas, W.; Abushelaibi, A.; Akseer, R.; Lim, S.-H.E.; Lai, K.-S. Disruption of KPC-producing Klebsiella pneumoniae membrane via induction of oxidative stress by cinnamon bark (Cinnamomum verum J. Presl) essential oil. PLoS ONE 2019, 14, e0214326. [Google Scholar] [CrossRef] [PubMed]
- Wink, M.; Ashour, M.L.; El-Readi, M.Z. Secondary metabolites from plants inhibiting ABC transporters and reversing resistance of cancer cells and microbes to cytotoxic and antimicrobial agents. Front. Microbiol. 2012, 3, 130. [Google Scholar] [CrossRef] [PubMed]
- Merghni, A.; Marzouki, H.; Hentati, H.; Aouni, M.; Mastouri, M. Antibacterial and antibiofilm activities of Laurus nobilis L. essential oil against Staphylococcus aureus strains associated with oral infections. Curr. Res. Transl. Med. 2016, 64, 29–34. [Google Scholar] [CrossRef] [PubMed]
- Barbieri, R.; Coppo, E.; Marchese, A.; Daglia, M.; Sobarzo-Sánchez, E.; Nabavi, S.F.; Nabavi, S.M. Phytochemicals for human disease: An update on plant-derived compounds antibacterial activity. Microbiol. Res. 2017, 196, 44–68. [Google Scholar] [CrossRef] [PubMed]
- Rooney, A.P.; Price, N.P.J.; Ehrhardt, C.; Swezey, J.L.; Bannan, J.D. Phylogeny and molecular taxonomy of the Bacillus subtilis species complex and description of Bacillus subtilis subsp. inaquosorum subsp. nov. Int. J. Syst. Evol. Microbiol. 2009, 59, 2429–2436. [Google Scholar] [CrossRef]
- Dhaouadi, S.; Mougou, A.H.; Rhouma, A. The plant pathogen Rhodococcus fascians. History, disease symptomatology, host range, pathogenesis and plant–pathogen interaction. Ann. Appl. Biol. 2020, 177, 4–15. [Google Scholar] [CrossRef]
- Gundlach, J.; Herzberg, C.; Kaever, V.; Gunka, K.; Hoffmann, T.; Weiß, M.; Gibhardt, J.; Thürmer, A.; Hertel, D.; Daniel, R.; et al. Control of potassium homeostasis is an essential function of the second messenger cyclic di-AMP in Bacillus subtilis. Sci. Signal. 2017, 10, eaal3011. [Google Scholar] [CrossRef]
- Ito, M.; Guffanti, A.A.; Wang, W.; Krulwich, T.A. Effects of nonpolar mutations in each of the seven Bacillus subtilis mrp genes suggest complex interactions among the gene products in support of Na(+) and alkali but not cholate resistance. J. Bacteriol. 2000, 182, 5663–5670. [Google Scholar] [CrossRef]
- Wilks, J.C.; Kitko, R.D.; Cleeton, S.H.; Lee, G.E.; Ugwu, C.S.; Jones, B.D.; BonDurant, S.S.; Slonczewski, J.L. Acid and base stress and transcriptomic responses in Bacillus subtilis. Appl. Environ. Microbiol. 2009, 75, 981–990. [Google Scholar] [CrossRef]
- Kitko, R.D.; Cleeton, R.L.; Armentrout, E.I.; Lee, G.E.; Noguchi, K.; Berkmen, M.B.; Jones, B.D.; Slonczewski, J.L. Cytoplasmic acidification and the benzoate transcriptome in Bacillus subtilis. PLoS ONE 2009, 4, e8255. [Google Scholar] [CrossRef]
- Domínguez-Escobar, J.; Wolf, D.; Fritz, G.; Höfler, C.; Wedlich-Söldner, R.; Mascher, T. Subcellular localization, interactions and dynamics of the phage-shock protein-like Lia response in Bacillus subtilis. Mol. Microbiol. 2014, 92, 716–732. [Google Scholar] [CrossRef] [PubMed]
- Marciniak, B.C.; Trip, H.; Fusetti, F.; Kuipers, O.P. Regulation of ykrL (htpX) by Rok and YkrK, a novel type of regulator in Bacillus subtilis. J. Bacteriol. 2012, 194, 2837–2845. [Google Scholar] [CrossRef] [PubMed]
- Basta, D.W.; Angeles-Albores, D.; Spero, M.A.; Ciemniecki, J.A.; Newman, D.K. Heat-shock proteases promote survival of Pseudomonas aeruginosa during growth arrest. Proc. Natl. Acad. Sci. USA 2020, 117, 4358–4367. [Google Scholar] [CrossRef] [PubMed]
- Gustafsson, M.C.U.; Roitel, O.; Marshall, K.R.; Noble, M.A.; Chapman, S.K.; Pessegueiro, A.; Fulco, A.J.; Cheesman, M.R.; von Wachenfeldt, C.; Munro, A.W. Expression, purification, and characterization of Bacillus subtilis cytochromes P450 CYP102A2 and CYP102A3: Flavocytochrome homologues of P450 BM3 from Bacillus megaterium. Biochemistry 2004, 43, 5474–5487. [Google Scholar] [CrossRef] [PubMed]
- Omardien, S.; Drijfhout, J.W.; van Veen, H.; Schachtschabel, S.; Riool, M.; Hamoen, L.W.; Brul, S.; Zaat, S.A.J. Synthetic antimicrobial peptides delocalize membrane bound proteins thereby inducing a cell envelope stress response. Biochim. Biophys. Acta-Biomembr. 2018, 1860, 2416–2427. [Google Scholar] [CrossRef] [PubMed]
- Yeh, J.I.; Chinte, U.; Du, S. Structure of glycerol-3-phosphate dehydrogenase, an essential monotopic membrane enzyme involved in respiration and metabolism. Proc. Natl. Acad. Sci. USA 2008, 105, 3280–3285. [Google Scholar] [CrossRef]
- Yang, C.-K.; Kashyap, D.R.; Kowalczyk, D.A.; Rudner, D.Z.; Wang, X.; Gupta, D.; Dziarski, R. Respiratory chain components are required for peptidoglycan recognition protein-induced thiol depletion and killing in Bacillus subtilis and Escherichia coli. Sci. Rep. 2021, 11, 64. [Google Scholar] [CrossRef]
- Thierbach, S.; Sartor, P.; Yücel, O.; Fetzner, S. Efficient modification of the Pseudomonas aeruginosa toxin 2-heptyl-1-hydroxyquinolin-4-one by three Bacillus glycosyltransferases with broad substrate ranges. J. Biotechnol. 2020, 308, 74–81. [Google Scholar] [CrossRef]
- Chen, L.; Keramati, L.; Helmann, J.D. Coordinate regulation of Bacillus subtilis peroxide stress genes by hydrogen peroxide and metal ions. Proc. Natl. Acad. Sci. USA 1995, 92, 8190–8194. [Google Scholar] [CrossRef]
- Que, Q.; Helmann, J.D. Manganese homeostasis in Bacillus subtilis is regulated by MntR, a bifunctional regulator related to the diphtheria toxin repressor family of proteins. Mol. Microbiol. 2000, 35, 1454–1468. [Google Scholar] [CrossRef] [PubMed]
- Atalla, A.; Schumann, W. The pst operon of Bacillus subtilis is specifically induced by alkali stress. J. Bacteriol. 2003, 185, 5019–5022. [Google Scholar] [CrossRef]
- Bender, R.A. Regulation of the histidine utilization (hut) system in bacteria. Microbiol. Mol. Biol. Rev. 2012, 76, 565–584. [Google Scholar] [CrossRef]
- Koskiniemi, S.; Lamoureux, J.G.; Nikolakakis, K.C.; t’Kint de Roodenbeke, C.; Kaplan, M.D.; Low, D.A.; Hayes, C.S. Rhs proteins from diverse bacteria mediate intercellular competition. Proc. Natl. Acad. Sci. USA 2013, 110, 7032–7037. [Google Scholar] [CrossRef] [PubMed]
- Rismondo, J.; Schulz, L.M. Not just transporters: Alternative functions of ABC transporters in Bacillus subtilis and Listeria monocytogenes. Microorganisms 2021, 9, 163. [Google Scholar] [CrossRef] [PubMed]
- Benda, M.; Schulz, L.M.; Stülke, J.; Rismondo, J. Influence of the ABC transporter YtrBCDEF of Bacillus subtilis on competence, biofilm formation and cell wall thickness. Front. Microbiol. 2021, 12, 587035. [Google Scholar] [CrossRef] [PubMed]
- Hughes, A.M.; Darby, J.F.; Dodson, E.J.; Wilson, S.J.; Turkenburg, J.P.; Thomas, G.H.; Wilkinson, A.J. Peptide transport in Bacillus subtilis—Structure and specificity in the extracellular solute binding proteins OppA and DppE. Microbiology 2022, 168. [Google Scholar] [CrossRef] [PubMed]
- Maurer, L.M.; Yohannes, E.; Bondurant, S.S.; Radmacher, M.; Slonczewski, J.L. PH Regulates genes for flagellar motility, catabolism, and oxidative stress in Escherichia coli K-12. J. Bacteriol. 2005, 187, 304–319. [Google Scholar] [CrossRef] [PubMed]
- He, S.; Cui, Y.; Dong, R.; Chang, J.; Cai, H.; Liu, H.; Shi, X. Global transcriptomic analysis of ethanol tolerance response in Salmonella enteritidis. Curr. Res. Food Sci. 2022, 5, 798–806. [Google Scholar] [CrossRef]
- Higgins, D.; Dworkin, J. Recent progress in Bacillus subtilis sporulation. FEMS Microbiol. Rev. 2012, 36, 131–148. [Google Scholar] [CrossRef]
- Brown, S.; Santa Maria, J.P.; Walker, S. Wall teichoic acids of Gram-positive bacteria. Annu. Rev. Microbiol. 2013, 67, 313–336. [Google Scholar] [CrossRef]
- Adak, S.; Aulak, K.S.; Stuehr, D.J. Direct evidence for nitric oxide production by a nitric-oxide synthase-like protein from Bacillus subtilis. J. Biol. Chem. 2002, 277, 16167–16171. [Google Scholar] [CrossRef]
- Chen, J.; Liu, L.; Wang, W.; Gao, H. Nitric oxide, nitric oxide formers and their physiological impacts in bacteria. Int. J. Mol. Sci. 2022, 23, 10778. [Google Scholar] [CrossRef]
- Shah, P.; Swiatlo, E. A multifaceted role for polyamines in bacterial pathogens. Mol. Microbiol. 2008, 68, 4–16. [Google Scholar] [CrossRef] [PubMed]
- Burrell, M.; Hanfrey, C.C.; Murray, E.J.; Stanley-Wall, N.R.; Michael, A.J. Evolution and multiplicity of arginine decarboxylases in polyamine biosynthesis and essential role in Bacillus subtilis biofilm formation. J. Biol. Chem. 2010, 285, 39224–39238. [Google Scholar] [CrossRef] [PubMed]
- Nicolas, P.; Mäder, U.; Dervyn, E.; Rochat, T.; Leduc, A.; Pigeonneau, N.; Bidnenko, E.; Marchadier, E.; Hoebeke, M.; Aymerich, S.; et al. Condition-dependent transcriptome reveals high-level regulatory architecture in Bacillus subtilis. Science 2012, 335, 1103–1106. [Google Scholar] [CrossRef]
- Huang, Y.; Smith, W.; Harwood, C.; Wipat, A.; Bacardit, J. Computational strategies for the identification of a transcriptional biomarker panel to sense cellular growth states in Bacillus subtilis. Sensors 2021, 21, 2436. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. FeatureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. ClusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Khameneh, B.; Iranshahy, M.; Soheili, V.; Fazly Bazzaz, B.S. Review on plant antimicrobials: A mechanistic viewpoint. Antimicrob. Resist. Infect. Control 2019, 8, 118. [Google Scholar] [CrossRef]
- Dey, S.; Rathod, S.; Gumphalwad, K.; Yadav, N.; Choudhari, P.; Rajakumara, E.; Dhavale, R.; Mahuli, D. Exploring α, β-unsaturated carbonyl compounds against bacterial efflux pumps via computational approach. J. Biomol. Struct. Dyn. 2023, 1–14. [Google Scholar] [CrossRef]
| Sg1 a | Sg2 | Sg3a | Sg3b | Sg4 | Sg5 | Sg6 | Gent b | |
|---|---|---|---|---|---|---|---|---|
| B. subtilis | ||||||||
| mean | 23.74 | 44.15 | 4.39 | - e | 7.89 | 56.67 | 2.07 | 0.77 |
| SD c | 1.42 | 17.63 | 0.44 | - | 0.62 | 22.76 | 0.07 | 0.14 |
| RSD% d | 5.97 | 39.92 | 10.01 | - | 7.88 | 40.16 | 3.21 | 18.50 |
| B. spizizenii | ||||||||
| mean | 16.36 | 36.00 | 4.49 | - | 8.48 | 32.03 | 1.65 | 0.79 |
| SD | 0.61 | 0.80 | 0.13 | - | 0.21 | 3.67 | 0.14 | 0.15 |
| RSD% | 3.75 | 2.23 | 2.90 | - | 2.51 | 11.46 | 8.43 | 18.66 |
| R. fascians | ||||||||
| mean | 8.32 | 15.83 | 1.59 | 7.09 | 4.01 | 1.93 | 1.43 | 3.58 |
| SD | 0.27 | 1.22 | 0.07 | 0.14 | 0.04 | 0.25 | 0.22 | 0.20 |
| RSD% | 3.25 | 7.72 | 4.65 | 1.98 | 0.98 | 12.87 | 15.43 | 5.52 |
| Sg1 a | Sg2 | Sg3a | Sg3b | Sg4 | Sg5 | Sg6 | Gent b | ||
|---|---|---|---|---|---|---|---|---|---|
| B. subtilis | MIC | 41.66 | 83.33 | 5.2 | - c | 10.41 | - | 2.6 | 1.04 |
| MBC | 41.66 | 83.33 | 5.2 | - | 41.66 | - | 2.6 | 1.66 | |
| B. spizizenii | MIC | 41.66 | 83.33 | 8.3 | - | 10.4 | - | 2.08 | 1.66 |
| MBC | 41.66 | 83.33 | 8.3 | - | 20.83 | - | 2.08 | 1.66 | |
| R. fascians | MIC | 10.4 | 20.8 | 4.1 | 16.7 | 5.2 | 4.1 | 2.6 | 8.3 |
| MBC | 41.7 | 41.7 | 8.3 | - | 10.4 | - | 8.3 | 16.7 |
| Gene Id a | Sg3a-5h | Sg6-5h | Sg3a-1h | Sg6-1h | Similarity, Function |
|---|---|---|---|---|---|
| GYO_0650 | 1.6 b | 1.2 | 2.2 | 1.9 | B. subtilis (BSU_04320) ydaO/kimA, high-affinity K+/H+ symporter |
| GYO_1109 | 2.9 | 1.5 | 4.3 | 1.4 | B. subtilis (BSU_08400) yfiU, similar to multidrug-efflux transporter |
| GYO_1188 | 3.2 | 2.5 | 3.0 | 2.8 | B. subtilis (BSU_08990) yhbI similar to transcriptional regulator |
| GYO_1189 | 2.5 | 2.2 | 2.8 | 4.0 | B. subtilis (BSU_09000) yhbJ, putative efflux system component |
| GYO_1190 | 2.8 | 4.0 | 2.8 | 3.9 | B. subtilis (BSU_09010) yhcA, c-di-AMP exporter |
| GYO_1191 | 2.5 | 3.6 | 2.8 | 3.9 | B. subtilis (BSU_09020) yhcB similar to oxidoreductase |
| GYO_1192 | 2.7 | 3.6 | 2.6 | 3.5 | B. subtilis (BSU_09030) yhcC unknown, hypothetical membrane protein |
| GYO_1218 | 1.2 | 1.1 | 1.3 | 2.1 | B. subtilis (BSU_09300) glpD, glycerol-3-phosphate dehydrogenase |
| GYO_1674 | 2.3 | 1.8 | 2.8 | 1.2 | B. subtilis (BSU_13490) htpX, stress-responsive membrane protease |
| GYO_2344 | 2.4 | 2.5 | 2.6 | 2.9 | B. subtilis (BSU_19420) yojK similar to macrolide glycosyltransferase |
| GYO_2953 | 1.1 | 1.0 | 2.1 | 1.7 | B. subtilis (BSU_27160) yrhJ, cytochrome P450, fatty acid metabolism |
| GYO_3335 | 1.1 | 2.1 | 1.0 | 1.7 | B. subtilis (BSU_30810) ytxM, ubiquinone and menaquinone biosynthesis |
| GYO_3455 | 1.2 | 1.4 | 2.3 | 1.1 | B. subtilis (BSU_31650) mrpF, Na+/H+ antiporter subunit, sodium export |
| GYO_3490 | 2.3 | 2.0 | 1.0 | 1.0 | B. subtilis (BSU_31990) dhbC, isochorismate synthase |
| GYO_3618 | 1.6 | 2.3 | 2.9 | 2.8 | B. subtilis (BSU_33120) liaH, Two-component system, accessory subunit of the TatAY-TatCY protein secretion complex, resistance against oxidative stress and cell wall antibiotics, protein secretion |
| GYO_3619 | 1.3 | 1.6 | 2.9 | 2.3 | B. subtilis (BSU_33130) liaI, Two-component system, membrane anchor for LiaH |
| GYO_0666 | −1.7 | −1.2 | −1.3 | −1.3 | B. subtilis (BSU_04470) dctP, C4-dicarboxylate transport protein, Two-component system, uptake of succinate, fumarate, malate and oxaloacetate via proton symport |
| GYO_3331 | −2.0 | −1.2 | −1.8 | −1.8 | B. subtilis (BSU_30770) mntA, manganese ABC transporter (binding protein, lipoprotein), manganese uptake |
| Treatments | Common Up-Regulated Genes | Common Down-Regulated Genes | Oppositely Regulated Genes |
|---|---|---|---|
| Sg3a-5h vs. Sg6-5h | 150 | 194 | 121 |
| Sg3a-1h vs. Sg6-1h | 67 | 82 | 1 |
| Sg3a-5h vs. Sg3a-1h | 83 | 29 | 108 |
| Sg6-5h vs. Sg6-1h | 56 | 32 | 32 |
| GO Term a | 5 h | 1 h | |||
|---|---|---|---|---|---|
| Sg3a | Sg6 | Sg3a | Sg6 | ||
| GO:0005215 | transporter activity | up/down b 58/51 | down 18 | up/down 24/46 | up/down 14/30 |
| GO:0016491 | oxidoreductase activity | up/down 3/7 | up 66 | up/down 60/4 | |
| GO:0016798 | hydrolase activity, acting on glycosyl bonds | down 16 | up 17 | ||
| GO:0015144 | carbohydrate transmembrane transporter activity | up 16 | |||
| GO:0020037 | heme binding | down 16 | |||
| GO:0003723 | RNA binding | down 39 | |||
| GO:0003735 | structural constituent of ribosome | down 27 | |||
| GO:0140101 | catalytic activity, acting on a tRNA | down 20 | |||
| GO:0035639 | purine ribonucleoside triphosphate binding (including GTP binding) | down 94 | |||
| GO:0046943 | carboxylic acid transmembrane transporter activity | down 8 | down 6 | ||
| GO:0140110 | transcription regulator activity | down 14 | |||
| GO Term a | 5 h | 1 h | |||
|---|---|---|---|---|---|
| Sg3a | Sg6 | Sg3a | Sg6 | ||
| GO:0006810 | transport | up/down b 79/90 | down 57 | down 37 | |
| GO:0055114 | oxidation–reduction process | up 55 | up 47 | ||
| GO:0032502 | developmental process | up 23 | |||
| GO:0043934 | sporulation | up 15 | |||
| GO:0006935 | chemotaxis | down 9 | down 10 | down 5 | |
| GO:0007165 | signal transduction | down 29 | |||
| GO:0005975 | carbohydrate metabolic process | down 36 | |||
| GO:1901135 | carbohydrate derivative metabolic process | down 35 | |||
| GO:1901564 | organonitrogen compound metabolic process | down 127 | |||
| GO:0019538 | protein metabolic process | down 69 | |||
| GO:0006412 | translation | down 46 | |||
| GO:0006754 | ATP biosynthetic process | down 8 | |||
| GO:0008610 | lipid biosynthetic process | down 16 | |||
| GO:0046942 | carboxylic acid transport | down 8 | down 7 | ||
| GO:0006355 | regulation of transcription, DNA-templated | down 20 | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bozsó, Z.; Lapat, V.; Ott, P.G.; Móricz, Á.M. Disparate Effects of Two Clerodane Diterpenes of Giant Goldenrod (Solidago gigantea Ait.) on Bacillus spizizenii. Int. J. Mol. Sci. 2024, 25, 1531. https://doi.org/10.3390/ijms25031531
Bozsó Z, Lapat V, Ott PG, Móricz ÁM. Disparate Effects of Two Clerodane Diterpenes of Giant Goldenrod (Solidago gigantea Ait.) on Bacillus spizizenii. International Journal of Molecular Sciences. 2024; 25(3):1531. https://doi.org/10.3390/ijms25031531
Chicago/Turabian StyleBozsó, Zoltán, Virág Lapat, Péter G. Ott, and Ágnes M. Móricz. 2024. "Disparate Effects of Two Clerodane Diterpenes of Giant Goldenrod (Solidago gigantea Ait.) on Bacillus spizizenii" International Journal of Molecular Sciences 25, no. 3: 1531. https://doi.org/10.3390/ijms25031531
APA StyleBozsó, Z., Lapat, V., Ott, P. G., & Móricz, Á. M. (2024). Disparate Effects of Two Clerodane Diterpenes of Giant Goldenrod (Solidago gigantea Ait.) on Bacillus spizizenii. International Journal of Molecular Sciences, 25(3), 1531. https://doi.org/10.3390/ijms25031531






