Methanogenic Archaea in the Pediatric Inflammatory Bowel Disease in Relation to Disease Type and Activity
Abstract
1. Introduction
2. Results
2.1. Characteristics of the Subjects
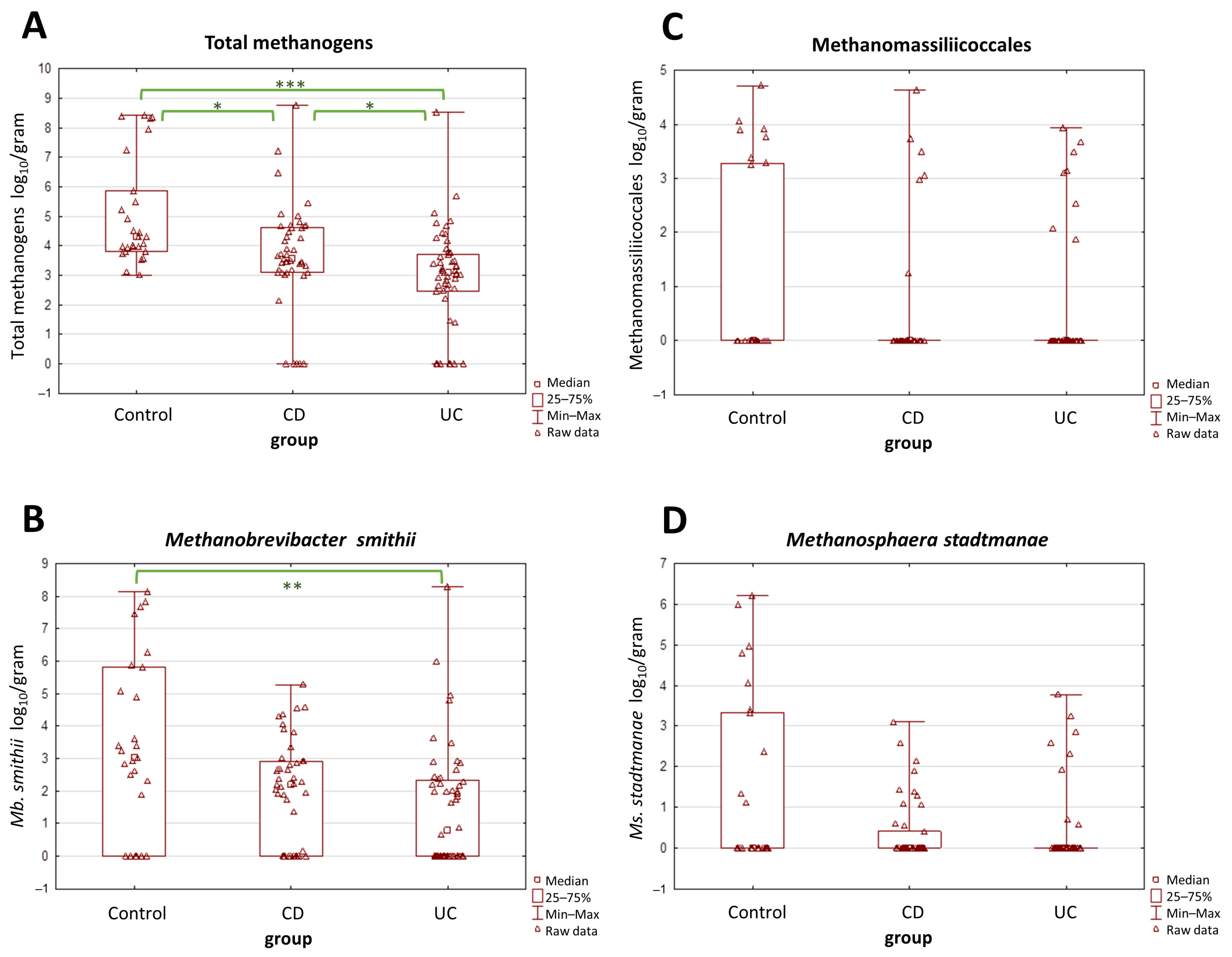
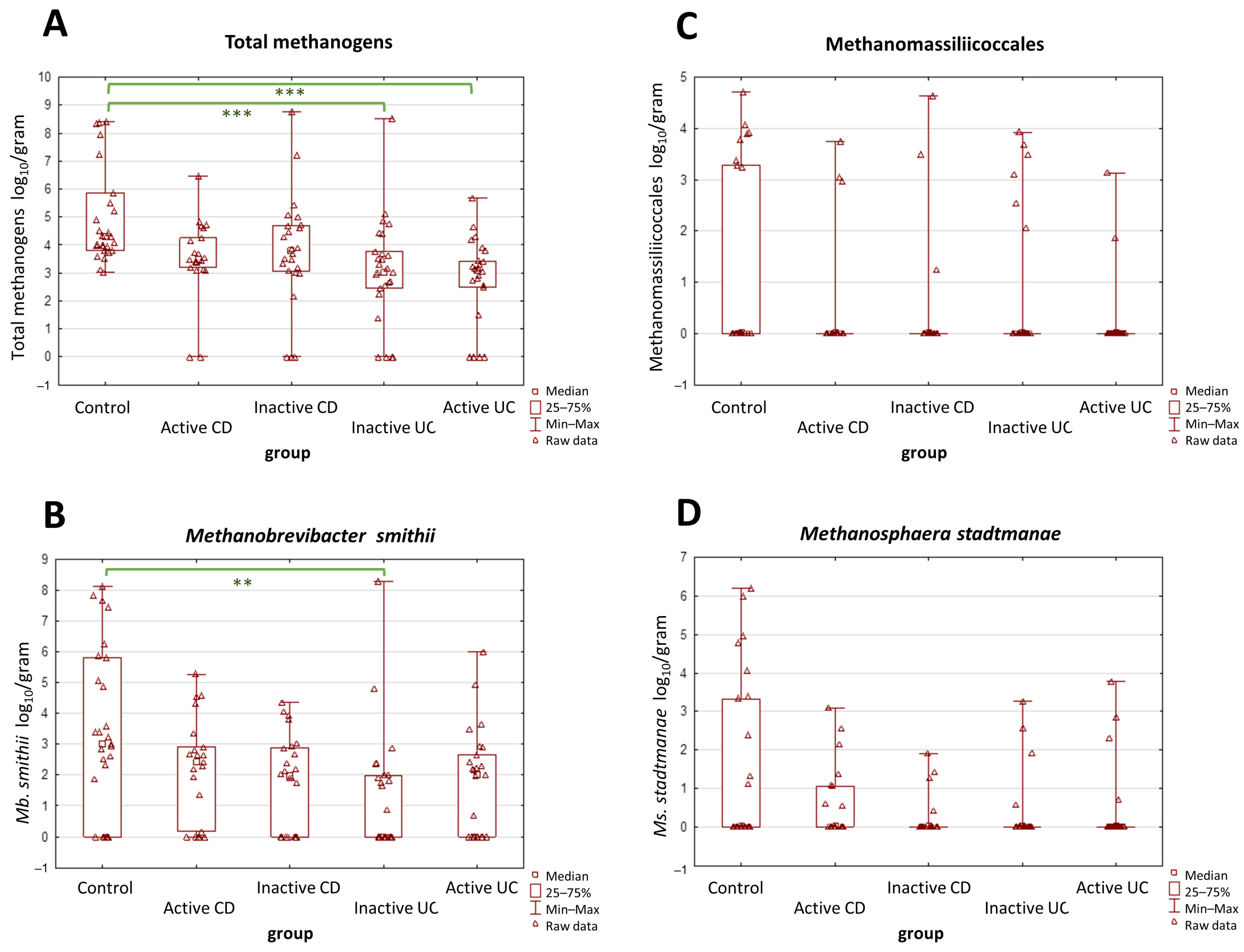
2.2. The Prevalence and Abundance of Gut Methanogens
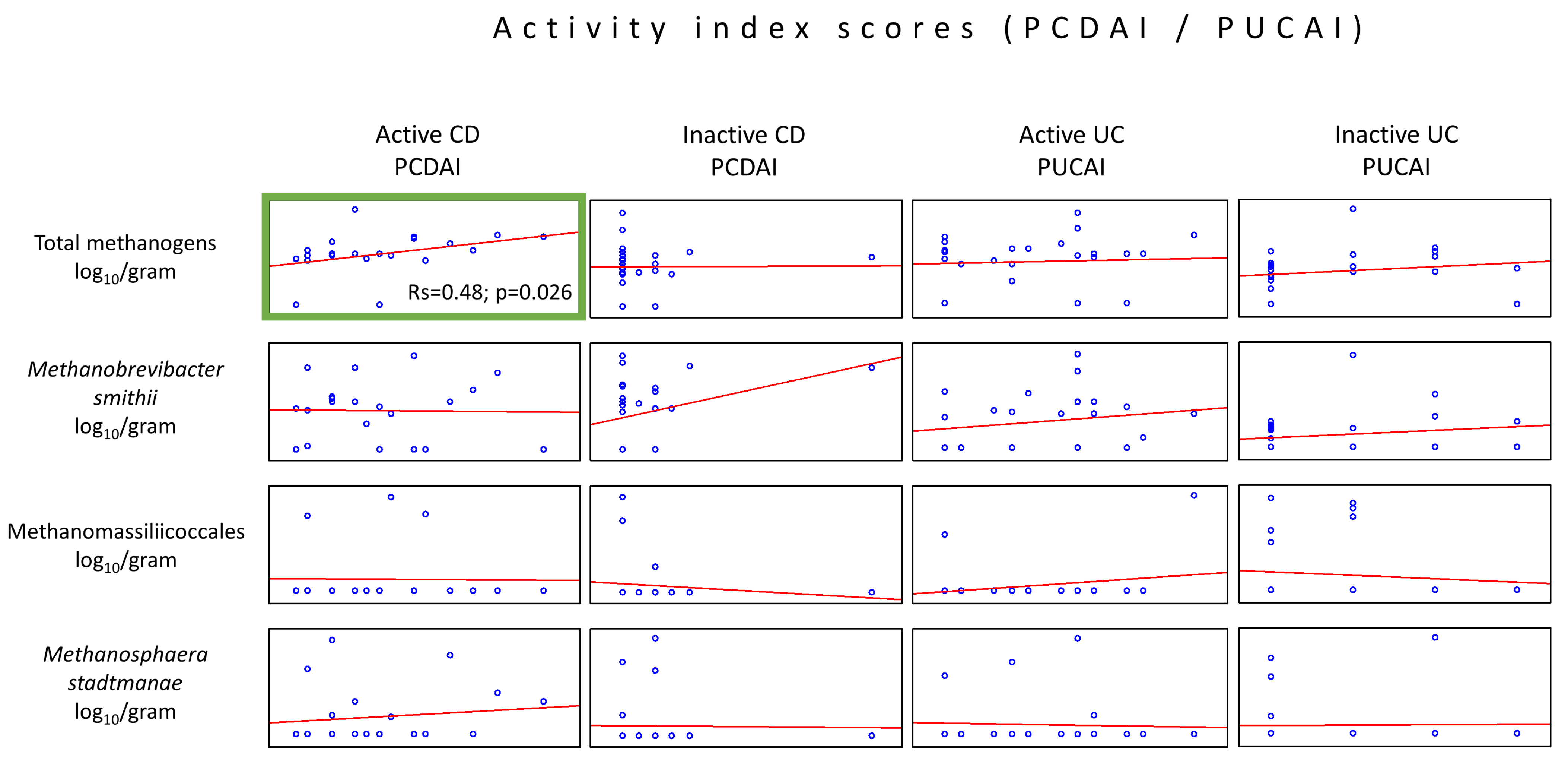
2.3. Relationship between IBD Activity Indices and the Number of Methanogenic Archaea
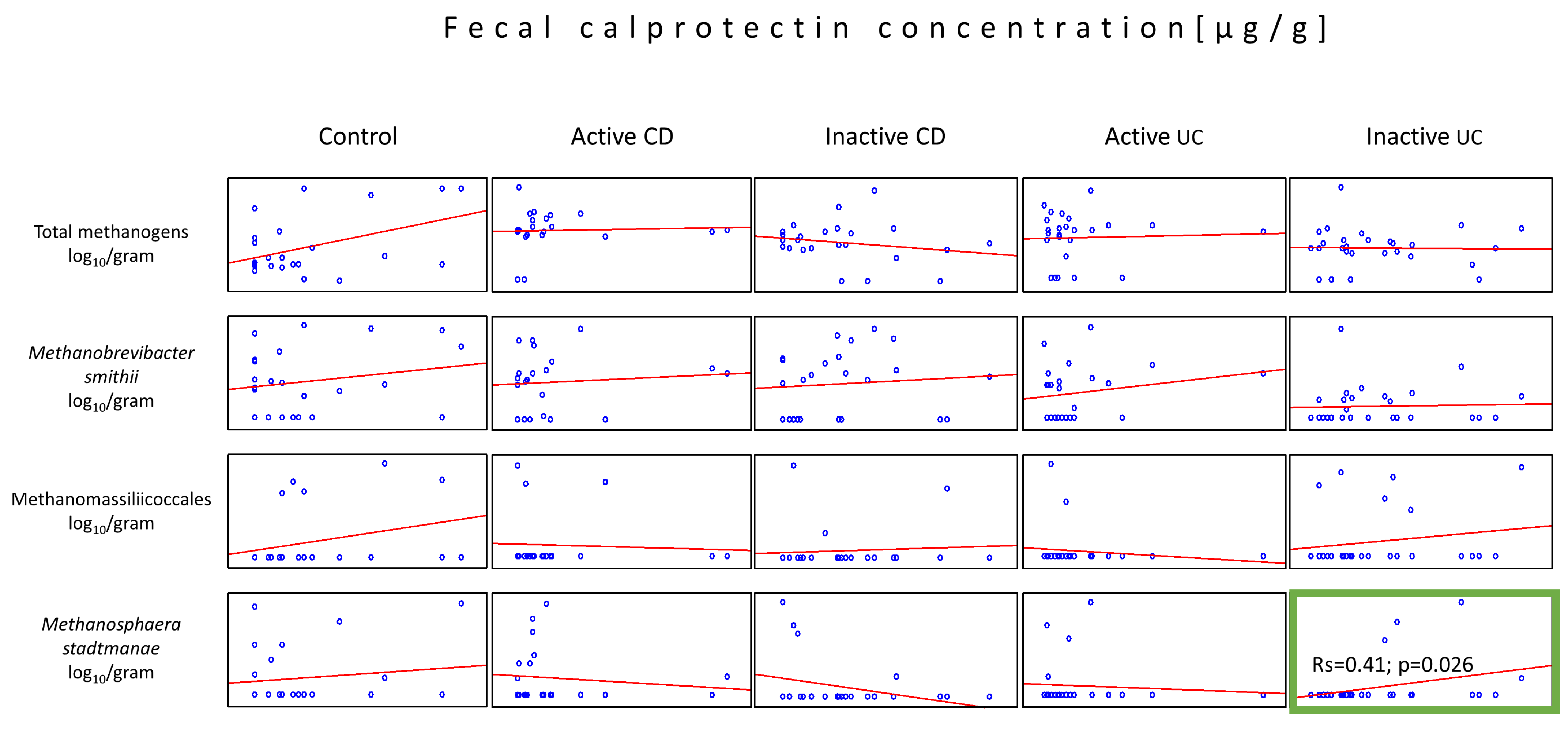
2.4. Relationship between FCP and the Number of Methanogenic Archaea
2.5. Relationship between Age and the Number of Methanogenic Archaea
3. Discussion
Limitation and Strength of the Study
4. Materials and Methods
4.1. Subjects
4.2. FCP Measurement
4.3. DNA Isolation
4.4. Quantitative Real-Time PCR
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Matsuoka, K.; Kobayashi, T.; Ueno, F.; Matsui, T.; Hirai, F.; Inoue, N.; Kato, J.; Kobayashi, K.; Kobayashi, K.; Koganei, K.; et al. Evidence-Based Clinical Practice Guidelines for Inflammatory Bowel Disease. J. Gastroenterol. 2018, 53, 305–353. [Google Scholar] [CrossRef] [PubMed]
- Ng, S.C.; Shi, H.Y.; Hamidi, N.; Underwood, F.E.; Tang, W.; Benchimol, E.I.; Panaccione, R.; Ghosh, S.; Wu, J.C.Y.; Chan, F.K.L.; et al. Worldwide Incidence and Prevalence of Inflammatory Bowel Disease in the 21st Century: A Systematic Review of Population-Based Studies. Lancet 2017, 390, 2769–2778. [Google Scholar] [CrossRef] [PubMed]
- Putowski, M.; Padala, O.; Krupa, A.; Konopelko, M.; Piasek, E.; Mazurek, M. Inflammatory Bowel Disease in Children. J. Educ. Health Sport 2019, 9, 406–411. [Google Scholar] [CrossRef]
- Witanowska, A.; Rydzewska, G. Epidemiologia i Przebieg Kliniczny Choroby Leśniowskiego-Crohna. In Choroba Leśniowskiego-Crohna—100 lat Diagnostyki i Terapii; Rydzewska, G., Małecka-Panas, E., Eds.; Termedia Wydawnictwa Medyczne: Poznań, Poland, 2008; pp. 23–36. ISBN 978-83-89825-53-7. [Google Scholar]
- Socha, P.; Dądalski, M.; Kierkuś, J.; Szymańska, S.; Ryżko, J. Genetyczne Uwarunkowania Nieswoistych Chorób Zapalnych Jelit. Stand. Med. 2010, 7, 100–105. [Google Scholar]
- Cho, J.H. Inflammatory Bowel Disease: Genetic and Epidemiologic Considerations. World J. Gastroenterol. 2008, 14, 338. [Google Scholar] [CrossRef] [PubMed]
- Guan, Q. A Comprehensive Review and Update on the Pathogenesis of Inflammatory Bowel Disease. J. Immunol. Res. 2019, 2019, 7247238. [Google Scholar] [CrossRef] [PubMed]
- Mottawea, W.; Chiang, C.K.; Mühlbauer, M.; Starr, A.E.; Butcher, J.; Abujamel, T.; Deeke, S.A.; Brandel, A.; Zhou, H.; Shokralla, S.; et al. Altered Intestinal Microbiota–Host Mitochondria Crosstalk in New Onset Crohn’s Disease. Nat. Commun. 2016, 7, 13419. [Google Scholar] [CrossRef]
- Scanlan, P.D.; Shanahan, F.; Marchesi, J.R. Human Methanogen Diversity and Incidence in Healthy and Diseased Colonic Groups Using McrA Gene Analysis. BMC Microbiol. 2008, 8, 79. [Google Scholar] [CrossRef]
- Valentine, D.L. Adaptations to Energy Stress Dictate the Ecology and Evolution of the Archaea. Nat. Rev. Microbiol. 2007, 5, 316–323. [Google Scholar] [CrossRef]
- Nottingham, P.M.; Hungate, R.E. Isolation of Methanogenic Bacteria from Feces of Man. J. Bacteriol. 1968, 96, 2178. [Google Scholar] [CrossRef]
- Gaci, N.; Borrel, G.; Tottey, W.; O’Toole, P.W.; Brugère, J.F. Archaea and the Human Gut: New Beginning of an Old Story. World J. Gastroenterol. 2014, 20, 16062–16078. [Google Scholar] [CrossRef] [PubMed]
- Moissl-Eichinger, C.; Pausan, M.; Taffner, J.; Berg, G.; Bang, C.; Schmitz, R.A. Archaea Are Interactive Components of Complex Microbiomes. Trends Microbiol. 2018, 26, 70–85. [Google Scholar] [CrossRef] [PubMed]
- Mohammadzadeh, R.; Mahnert, A.; Duller, S.; Moissl-Eichinger, C. Archaeal Key-Residents within the Human Microbiome: Characteristics, Interactions and Involvement in Health and Disease. Curr. Opin. Microbiol. 2022, 67, 102146. [Google Scholar] [CrossRef] [PubMed]
- Sereme, Y.; Mezouar, S.; Grine, G.; Mege, J.L.; Drancourt, M.; Corbeau, P.; Vitte, J. Methanogenic Archaea: Emerging Partners in the Field of Allergic Diseases. Clin. Rev. Allergy Immunol. 2019, 57, 456–466. [Google Scholar] [CrossRef]
- Lecours, P.B.; Marsolais, D.; Cormier, Y.; Berberi, M.; Haché, C.; Bourdages, R.; Duchaine, C. Increased Prevalence of Methanosphaera stadtmanae in Inflammatory Bowel Diseases. PLoS ONE 2014, 9, e87734. [Google Scholar] [CrossRef]
- Vierbuchen, T.; Bang, C.; Rosigkeit, H.; Schmitz, R.A.; Heine, H. The Human-Associated Archaeon Methanosphaera stadtmanae Is Recognized through Its RNA and Induces TLR8-Dependent NLRP3 Inflammasome Activation. Front. Immunol. 2017, 8, 313829. [Google Scholar] [CrossRef]
- Chehoud, C.; Albenberg, L.G.; Judge, C.; Hoffmann, C.; Grunberg, S.; Bittinger, K.; Baldassano, R.N.; Lewis, J.D.; Bushman, F.D.; Wu, G.D. A Fungal Signature in the Gut Microbiota of Pediatric Patients with Inflammatory Bowel Disease. Inflamm. Bowel Dis. 2015, 21, 1948. [Google Scholar] [CrossRef]
- Krawczyk, A.; Salamon, D.; Kowalska-Duplaga, K.; Bogiel, T.; Gosiewski, T. Association of Fungi and Archaea of the Gut Microbiota with Crohn’s Disease in Pediatric Patients—Pilot Study. Pathogens 2021, 10, 1119. [Google Scholar] [CrossRef]
- Ruel, J.; Ruane, D.; Mehandru, S.; Gower-Rousseau, C.; Colombel, J.F. IBD across the Age Spectrum: Is It the Same Disease? Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 88–98. [Google Scholar] [CrossRef]
- Cisek, A.A.; Bak, I.; Cukrowska, B. Improved Quantitative Real-Time PCR Protocol for Detection and Quantification of Methanogenic Archaea in Stool Samples. Microorganisms 2023, 11, 660. [Google Scholar] [CrossRef]
- OpenEpi—2 × 2 Table Statistics. Available online: https://www.openepi.com/TwobyTwo/TwobyTwo.htm (accessed on 25 December 2023).
- Ghavami, S.B.; Rostami, E.; Sephay, A.A.; Shahrokh, S.; Balaii, H.; Aghdaei, H.A.; Zali, M.R. Alterations of the Human Gut Methanobrevibacter smithii as a Biomarker for Inflammatory Bowel Diseases. Microb. Pathog. 2018, 117, 285–289. [Google Scholar] [CrossRef] [PubMed]
- Houshyar, Y.; Massimino, L.; Lamparelli, L.A.; Danese, S.; Ungaro, F. Going Beyond Bacteria: Uncovering the Role of Archaeome and Mycobiome in Inflammatory Bowel Disease. Front. Physiol. 2021, 12, 783295. [Google Scholar] [CrossRef] [PubMed]
- Massimino, L.; Lamparelli, L.A.; Houshyar, Y.; D’Alessio, S.; Peyrin-Biroulet, L.; Vetrano, S.; Danese, S.; Ungaro, F. The Inflammatory Bowel Disease Transcriptome and Metatranscriptome Meta-Analysis (IBD TaMMA) Framework. Nat. Comput. Sci. 2021, 1, 511–515. [Google Scholar] [CrossRef]
- Zhang, X.; Deeke, S.A.; Ning, Z.; Starr, A.E.; Butcher, J.; Li, J.; Mayne, J.; Cheng, K.; Liao, B.; Li, L.; et al. Metaproteomics Reveals Associations between Microbiome and Intestinal Extracellular Vesicle Proteins in Pediatric Inflammatory Bowel Disease. Nat. Commun. 2018, 9, 2873. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Wu, M.; Xiao, H.; Liu, H.; Yang, G. Mesalamine-Mediated Amelioration of Experimental Colitis in Piglets Involves Gut Microbiota Modulation and Intestinal Immune Cell Infiltration. Front. Immunol. 2022, 13, 883682. [Google Scholar] [CrossRef]
- Bang, C.; Weidenbach, K.; Gutsmann, T.; Heine, H.; Schmitz, R.A. The Intestinal Archaea Methanosphaera stadtmanae and Methanobrevibacter smithii Activate Human Dendritic Cells. PLoS ONE 2014, 9, 99411. [Google Scholar] [CrossRef]
- D’Amico, F.; Nancey, S.; Danese, S.; Peyrin-Biroulet, L. A Practical Guide for Faecal Calprotectin Measurement: Myths and Realities. J. Crohn’s Colitis 2021, 15, 152–161. [Google Scholar] [CrossRef]
- Pimentel, M.; Mayer, A.G.; Park, S.; Chow, E.J.; Hasan, A.; Kong, Y. Methane Production during Lactulose Breath Test Is Associated with Gastrointestinal Disease Presentation. Dig. Dis. Sci. 2003, 48, 86–92. [Google Scholar] [CrossRef]
- Rana, S.V.; Sharma, S.; Malik, A.; Kaur, J.; Prasad, K.K.; Sinha, S.K.; Singh, K. Small Intestinal Bacterial Overgrowth and Orocecal Transit Time in Patients of Inflammatory Bowel Disease. Dig. Dis. Sci. 2013, 58, 2594–2598. [Google Scholar] [CrossRef]
- Shah, A.; Morrison, M.; Burger, D.; Martin, N.; Rich, J.; Jones, M.; Koloski, N.; Walker, M.M.; Talley, N.J.; Holtmann, G.J.; et al. Systematic Review with Meta-Analysis: The Prevalence of Small Intestinal Bacterial Overgrowth in Inflammatory Bowel Disease Summary Background: Current Data on Small Intestinal Bacterial Overgrowth (SIBO) in Patients. Aliment. Pharmacol. Ther. 2019, 49, 624–635. [Google Scholar] [CrossRef]
- Gandhi, A.; Shah, A.; Jones, M.P.; Koloski, N.; Talley, N.J.; Morrison, M.; Holtmann, G. Methane Positive Small Intestinal Bacterial Overgrowth in Inflammatory Bowel Disease and Irritable Bowel Syndrome: A Systematic Review and Meta-Analysis. Gut Microbes 2021, 13, 1933313. [Google Scholar] [CrossRef]
- Weaver, G.A.; Krause, J.A.; Miller, T.L.; Wolin, M.J. Incidence of Methanogenic Bacteria in a Sigmoidoscopy Population: An Association of Methanogenic Bacteria and Diverticulosis. Gut 1986, 27, 698–704. [Google Scholar] [CrossRef]
- Peled, Y.; Gilat, T.; Liberman, E.; Bujanover, Y. The Development of Methane Production in Childhood and Adolescence. J. Pediatr. Gastroenterol. Nutr. 1985, 4, 575–579. [Google Scholar] [CrossRef]
- Mihajlovski, A.; Doré, J.; Levenez, F.; Alric, M.; Brugère, J.F. Molecular Evaluation of the Human Gut Methanogenic Archaeal Microbiota Reveals an Age-Associated Increase of the Diversity. Environ. Microbiol. Rep. 2010, 2, 272–280. [Google Scholar] [CrossRef]
- Dridi, B.; Henry, M.; Richet, H.; Raoult, D.; Drancourt, M. Age-Related Prevalence of Methanomassiliicoccus luminyensis in the Human Gut Microbiome. APMIS 2012, 120, 773–777. [Google Scholar] [CrossRef]
- Vanderhaeghen, S.; Lacroix, C.; Schwab, C. Methanogen Communities in Stools of Humans of Different Age and Health Status and Co-Occurrence with Bacteria. FEMS Microbiol. Lett. 2015, 362, fnv092. [Google Scholar] [CrossRef]
- Busquets, D.; Oliver, L.; Amoedo, J.; Ramió-Pujol, S.; Malagón, M.; Serrano, M.; Bahí, A.; Capdevila, M.; Lluansí, A.; Torrealba, L.; et al. RAID Prediction: Pilot Study of Fecal Microbial Signature With Capacity to Predict Response to Anti-TNF Treatment. Inflamm. Bowel Dis. 2021, 27, S63–S66. [Google Scholar] [CrossRef]
- Kucharzik, T.; Ellul, P.; Greuter, T.; Rahier, J.F.; Verstockt, B.; Abreu, C.; Albuquerque, A.; Allocca, M.; Esteve, M.; Farraye, F.A.; et al. ECCO Guidelines on the Prevention, Diagnosis, and Management of Infections in Inflammatory Bowel Disease. J. Crohn’s Colitis 2021, 15, 879–913. [Google Scholar] [CrossRef]
- Cisek, A.A.; Bąk, I.; Stefańska, I.; Binek, M. Selection and Optimization of High-Yielding DNA Isolation Protocol for Quantitative Analyses of Methanogenic Archaea. Microorganisms 2022, 10, 523. [Google Scholar] [CrossRef]
- Salonen, A.; Nikkilä, J.; Jalanka-Tuovinen, J.; Immonen, O.; Rajilić-Stojanović, M.; Kekkonen, R.A.; Palva, A.; de Vos, W.M. Comparative Analysis of Fecal DNA Extraction Methods with Phylogenetic Microarray: Effective Recovery of Bacterial and Archaeal DNA Using Mechanical Cell Lysis. J. Microbiol. Methods 2010, 81, 127–134. [Google Scholar] [CrossRef] [PubMed]
- The Ribosomal RNA Database. Available online: https://rrndb.umms.med.umich.edu/ (accessed on 24 October 2023).
- Science Primer. Available online: http://scienceprimer.com/copy-number-calculator-for-realtime-pcr (accessed on 1 October 2023).
- Ufnar, J.A.; Wang, S.Y.; Christiansen, J.M.; Yampara-Iquise, H.; Carson, C.A.; Ellender, R.D. Detection of the NifH Gene of Methanobrevibacter smithii: A Potential Tool to Identify Sewage Pollution in Recreational Waters. J. Appl. Microbiol. 2006, 101, 44–52. [Google Scholar] [CrossRef]
- Cozannet, M.; Borrel, G.; Roussel, E.; Moalic, Y.; Allioux, M.; Sanvoisin, A.; Toffin, L.; Alain, K. New Insights into the Ecology and Physiology of Methanomassiliicoccales from Terrestrial and Aquatic Environments. Microorganisms 2021, 9, 30. [Google Scholar] [CrossRef]
- Prion, S.; Haerling, K.A. Making Sense of Methods and Measurement: Spearman-Rho Ranked-Order Correlation Coefficient. Clin. Simul. Nurs. 2014, 10, 535–536. [Google Scholar] [CrossRef]

| Group | Mean Age [Years] ± SD (Mdn) | Female to Male Ratio | Mean PCDAI ± SD (Mdn) | Mean PUCAI ± SD (Mdn) | Mean FCP [µg/g] ± SD (Mdn) |
|---|---|---|---|---|---|
| CD (n = 45) | 14.2 ± 3.1 (15.0) * | 25:20 | 10.0 ± 13.5 (5.0) | n/a | 781.6 ± 1800.7 (119.0) * |
| Active CD (n = 21) | 13.9 ± 3.4 (15.0) * | 15:6 | 17.5 ± 14.7 (15.0) # | n/a | 1601.8 ± 2409.4 (699.0) *# |
| Inactive CD (n = 24) | 14.6 ± 2.8 (15.0) * | 10:14 | 3.4 ± 7.8 (0.0) | n/a | 64.0 ± 61.0 (59.5) |
| UC (n = 52) | 13.0 ± 4.7 (14.0) * | 27:25 | n/a | 15.2 ± 20.3 (5.0) | 396.9 ± 810.3 (100.5) * |
| Active UC (n = 25) | 12.2 ± 5.0 (13.0) | 12:13 | n/a | 27.6 ± 23.3 (25.0) # | 758.5 ± 1062.9 (403.0) *# |
| Inactive UC (n = 27) | 13.8 ± 4.3 (15.0) * | 15:12 | n/a | 3.7 ± 4.9 (0.0) | 62.0 ± 54.9 (38.0) |
| Controls (n = 27) | 10.0 ± 4.0 (10.0) | 14:13 | n/a | n/a | 19.3 ± 24.1 (10.0) |
| Group | No. of Positive/Total No. of Tested Samples and Percentage Values [%] | |||
|---|---|---|---|---|
| Total Methanogens | Mb. smithii | Ms. stadtmanae | Methanomasiilicoccales | |
| CD (n = 45) | 40/45 [88.9%] | 31/45 [68.9%] | 12/45 [26.7%] | 6/45 [13.3%] |
| Active CD (n = 21) | 19/21 [90.5%] | 16/21 [76.2%] | 8/21 [38.1%] | 3/21 [14.3%] |
| Inactive CD (n = 24) | 21/24 [87.5%] | 15/24 [62.5%] | 4//24 [16.7%] | 3/24 [12.5%] |
| UC (n = 52) | 43/52 [82.7%] * | 27/52 [51.9%] | 8/52 [15.4%] * | 8/52 [15.4%] |
| Active UC (n = 25) | 20/25 [80.0%] * | 15/25 [60.0%] | 4/25 [16.0%] | 2/25 [8.0%] |
| Inactive UC (n = 27) | 23/27 [85.2%] | 12/27 [44.4%] | 4/27 [14.8%] | 6/27 [22.2%] |
| Controls (n = 27) | 27/27 [100%] | 20/27 [74.1%] | 10/27 [37.0%] | 8/27 [29.6%] |
| Group | Odds Ratios [95% Confidence Intervals] Compared to Control Group | |||
|---|---|---|---|---|
| Total Methanogens * | Mb. smithii | Ms. stadtmanae | Methanomasiilicoccales | |
| CD (n = 45) | 0 | 0.78 [0.27, 2.25] | 0.62 [0.22, 1.72] | 0.37 [0.11, 1.20] |
| Active CD (n = 21) | 0 | 1.12 [0.30, 4.20] | 1.05 [0.32, 3.40] | 0.40 [0.09, 1.73] |
| Inactive CD (n = 24) | 0 | 0.58 [0.18, 1.92] | 0.34 [0.09, 1.28] | 0.34 [0.08, 1.47] |
| UC (n = 52) | 0 | 0.38 [0.14, 1.05] | 0.31 [0.10, 0.91] | 0.43 [0.14, 1.32] |
| Active UC (n = 25) | 0 | 0.53 [0.16, 1.70] | 0.32 [0.09, 1.22] | 0.21 [0.04, 1.09] |
| Inactive UC (n = 27) | 0 | 0.28 [0.09, 0.88] | 0.30 [0.08, 1.11] | 0.68 [0.20, 2.31] |
| Microorganism | Target Gene | Forward/Reverse Primer 5′–3′ Sequence * | Amplicon Length [bp] | Reference |
|---|---|---|---|---|
| Total methanogenic archaea | mcrA | CTTGAARMTCACTTCGGTGGWTC/CGTTCATBGCGTAGTTVGGRTAGT | Approx. 270 | [21] |
| Methanobrevibacter smithii | nifH | AACAGAAAACCCAGTGAAGAGGATA/ACGTAAAGGCACTGAAAAACCTCC | 222 | Modified [45] |
| Methanomassiliicoccales | 16S rDNA | GGGGTAGGGGTAAAATCCTGTAATCC/AACAACTTCTCTCCGGCACTGG | 194 | Modified [46] |
| Methanosphaera stadtmanae | mtaB | GTAGTTCCTAACATCAAAGTAGCTCC/TCCTCTAAGACCGTTTTCTTCTTCTCTCA | 300 | Modified [16] |
| Real-Time PCR Step | Total Methanogens | Mb. smithii | Methanomassiliicoccales | Ms. stadtmanae |
|---|---|---|---|---|
| Initial Denaturation | 95 °C—5 min | |||
| Denaturation | 94 °C—20 s | 94 °C—20 s | 94 °C—20 s | 94 °C—20 s |
| Annealing | 60 °C—20 s | 66 °C—20 s | 70 °C—20 s | 68 °C—20 s |
| Elongation | 72 °C—20 s | 72 °C—20 s | 72 °C—20 s | 72 °C—25 s |
| Signal acquisition * | 81 °C—20 s + Acq | 82 °C—20 s + Acq | 87 °C—20 s + Acq | 81 °C—20 s + Acq |
| Melt analysis * | 95 °C—5 s, then 60 °C—1 min, and 95 °C—continuous Acq with ramp rate 0.11 °C/s | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cisek, A.A.; Szymańska, E.; Wierzbicka-Rucińska, A.; Aleksandrzak-Piekarczyk, T.; Cukrowska, B. Methanogenic Archaea in the Pediatric Inflammatory Bowel Disease in Relation to Disease Type and Activity. Int. J. Mol. Sci. 2024, 25, 673. https://doi.org/10.3390/ijms25010673
Cisek AA, Szymańska E, Wierzbicka-Rucińska A, Aleksandrzak-Piekarczyk T, Cukrowska B. Methanogenic Archaea in the Pediatric Inflammatory Bowel Disease in Relation to Disease Type and Activity. International Journal of Molecular Sciences. 2024; 25(1):673. https://doi.org/10.3390/ijms25010673
Chicago/Turabian StyleCisek, Agata Anna, Edyta Szymańska, Aldona Wierzbicka-Rucińska, Tamara Aleksandrzak-Piekarczyk, and Bożena Cukrowska. 2024. "Methanogenic Archaea in the Pediatric Inflammatory Bowel Disease in Relation to Disease Type and Activity" International Journal of Molecular Sciences 25, no. 1: 673. https://doi.org/10.3390/ijms25010673
APA StyleCisek, A. A., Szymańska, E., Wierzbicka-Rucińska, A., Aleksandrzak-Piekarczyk, T., & Cukrowska, B. (2024). Methanogenic Archaea in the Pediatric Inflammatory Bowel Disease in Relation to Disease Type and Activity. International Journal of Molecular Sciences, 25(1), 673. https://doi.org/10.3390/ijms25010673






