CLEC16A—An Emerging Master Regulator of Autoimmunity and Neurodegeneration
Abstract
1. Introduction
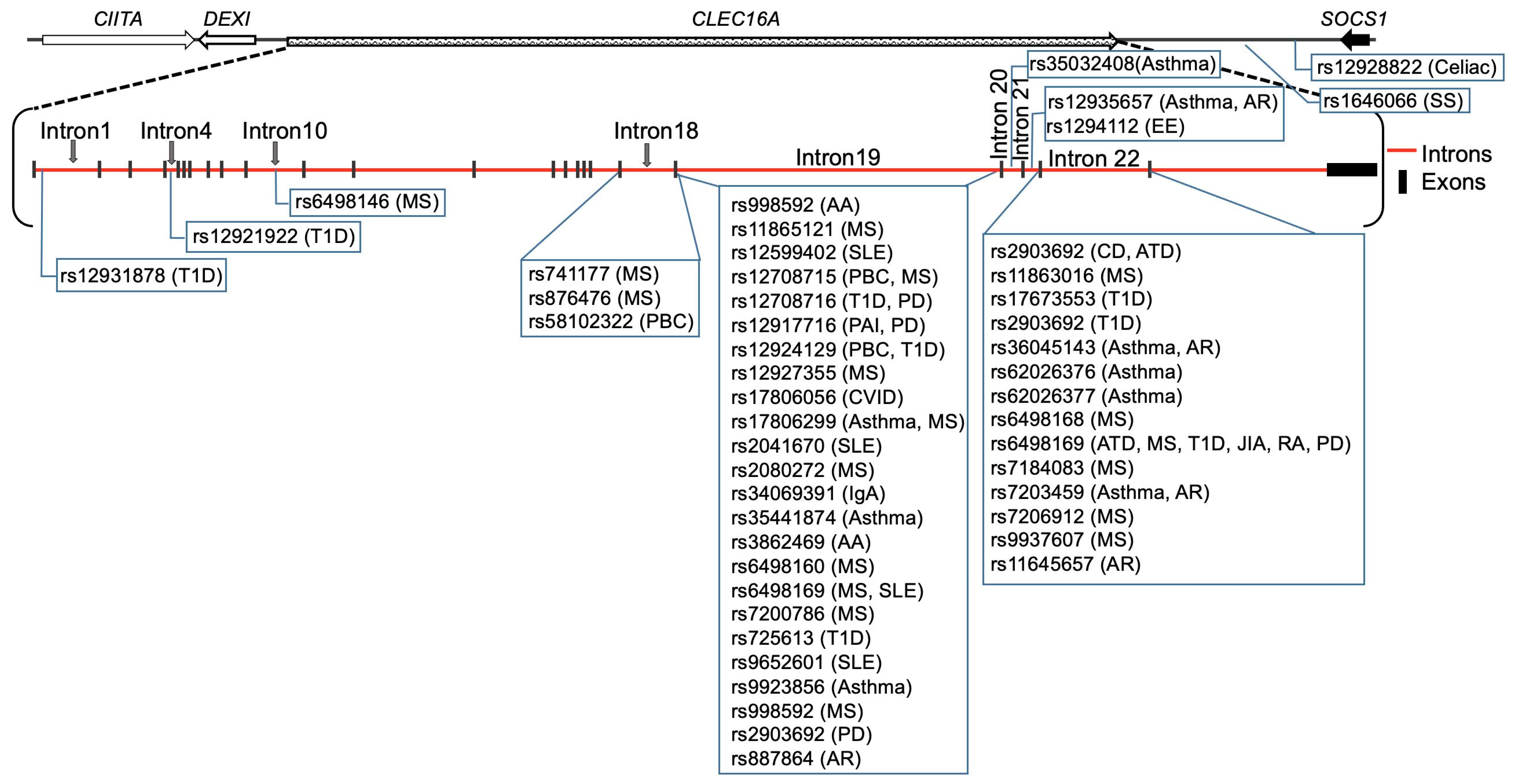
2. CLEC16A (16p13)—An Autoimmune Candidate Gene
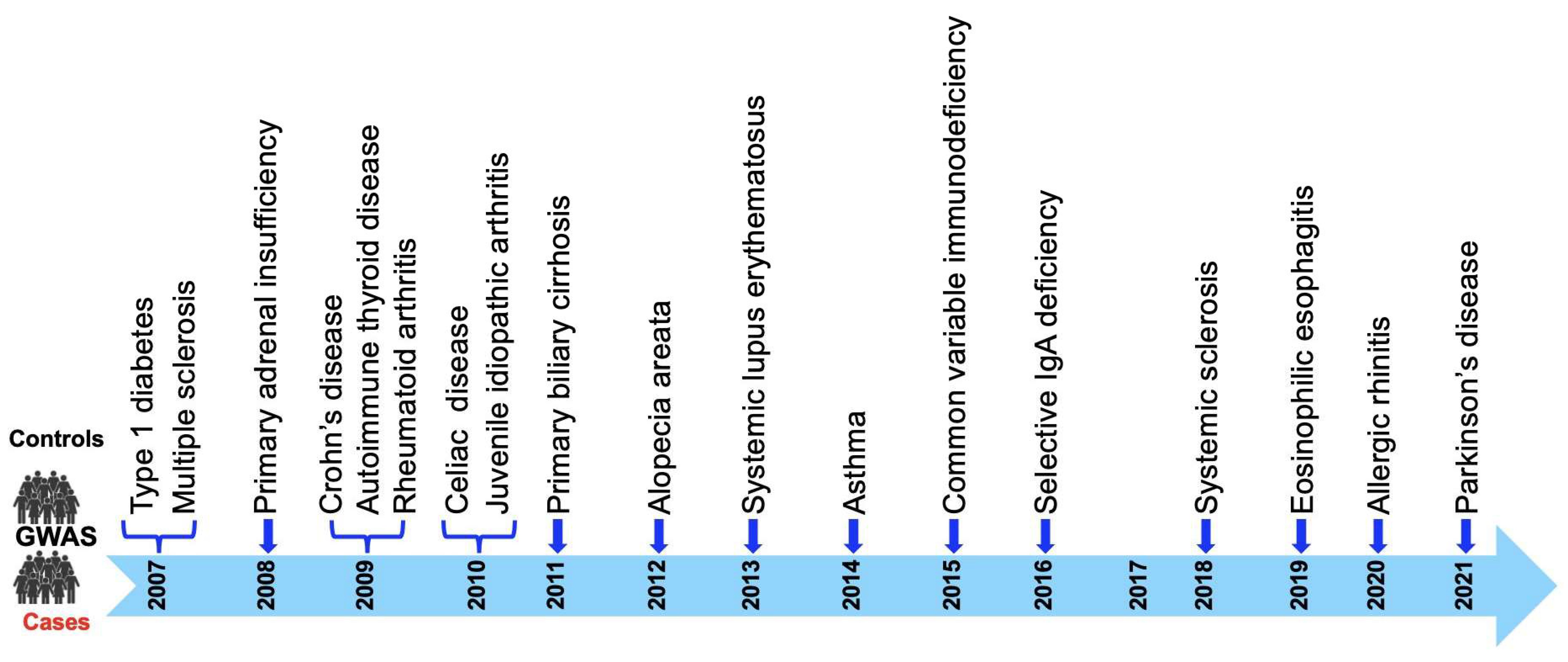
3. CLEC16A in Health and Diseases through GWAS
4. Biological Role of CLEC16A
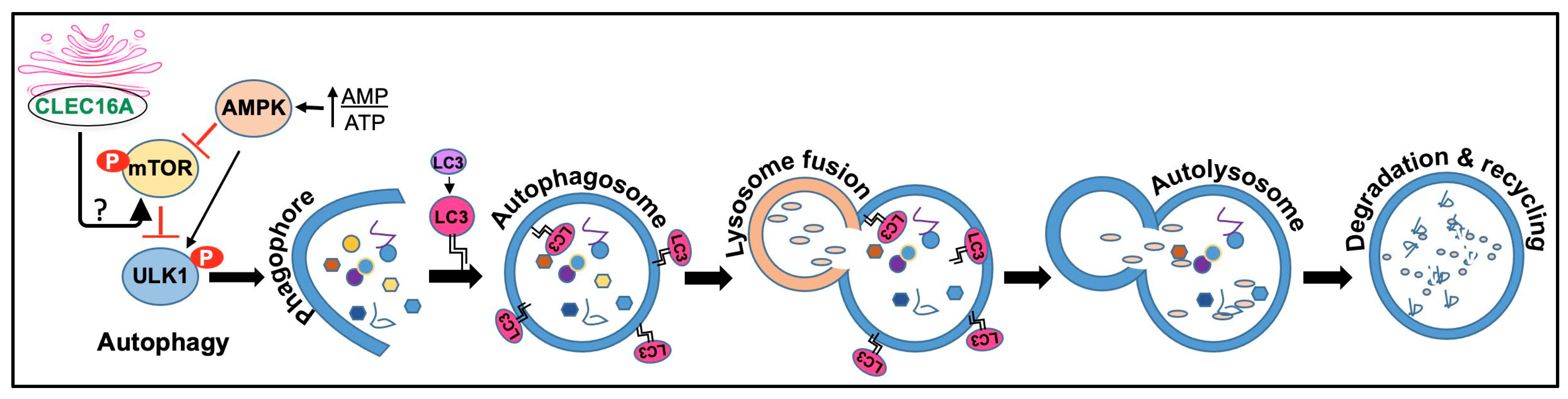
4.1. Autophagy Regulation
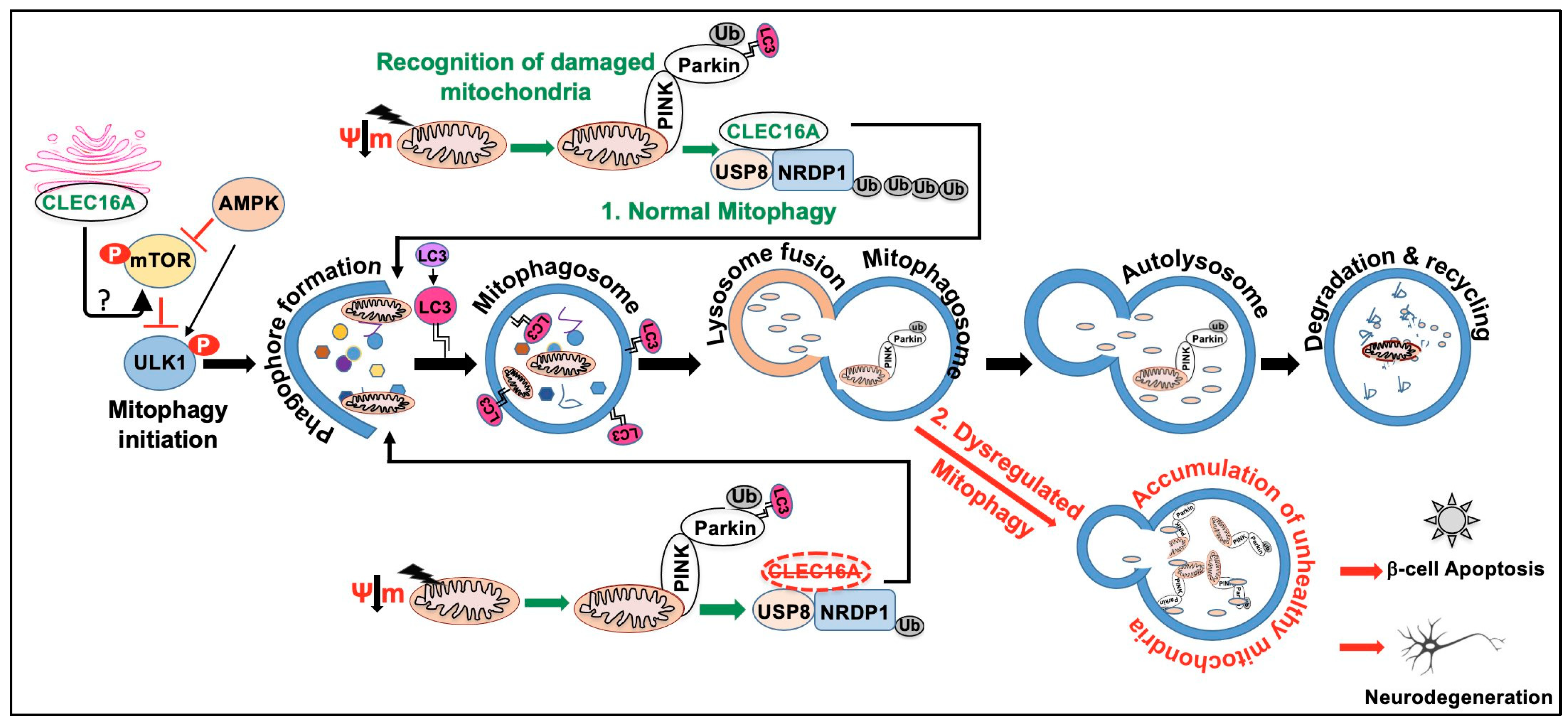
4.2. Regulator of Mitophagy
5. CLEC16A in Neurodegeneration
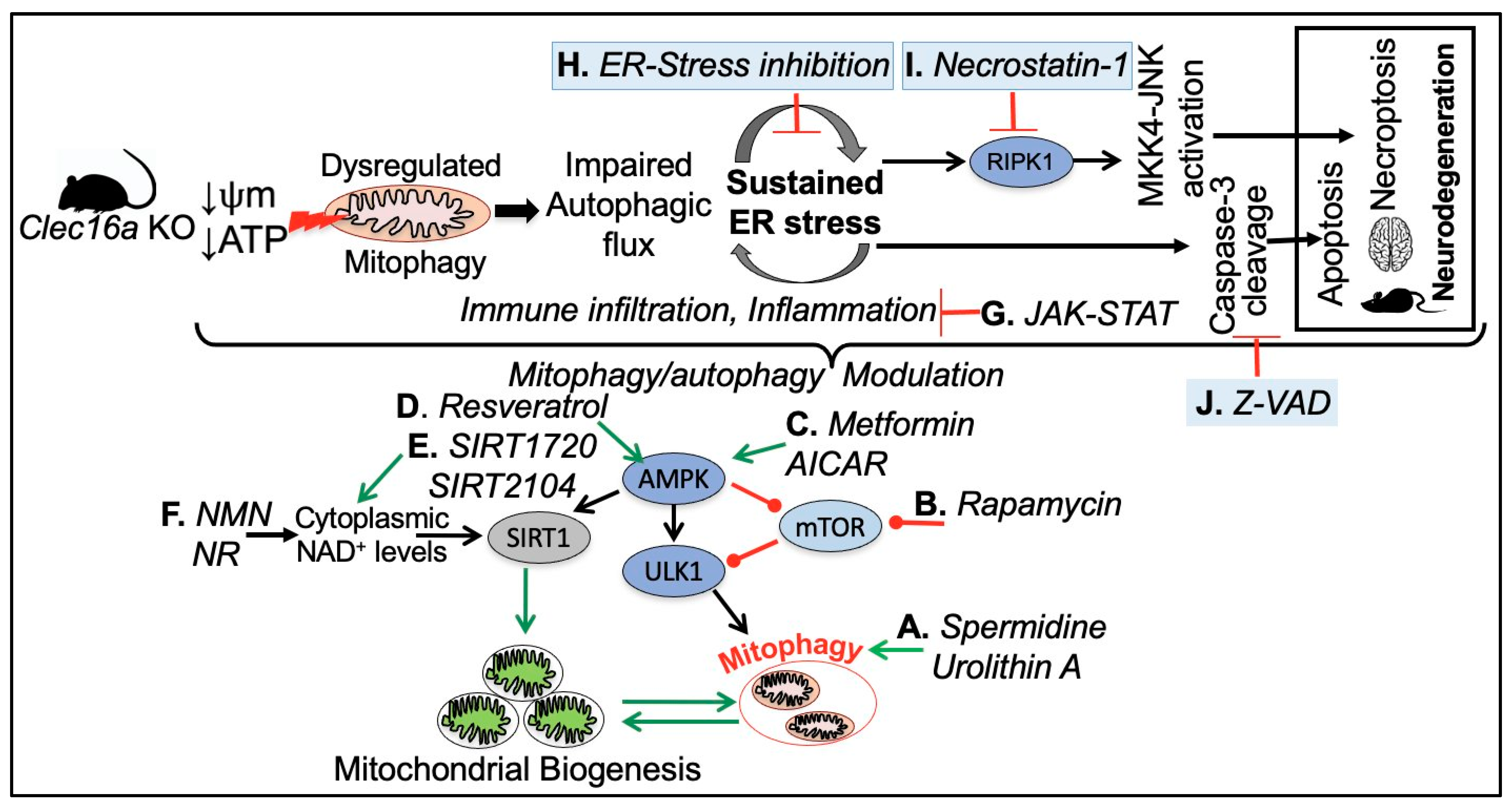
6. Therapeutic Targeting of CLEC16A
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hakonarson, H.; Grant, S.F.; Bradfield, J.P.; Marchand, L.; Kim, C.E.; Glessner, J.T.; Grabs, R.; Casalunovo, T.; Taback, S.P.; Frackelton, E.C.; et al. A genome-wide association study identifies KIAA0350 as a type 1 diabetes gene. Nature 2007, 448, 591–594. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Naylor, S.A.; DiAntonio, A. Drosophila Golgi membrane protein Ema promotes autophagosomal growth and function. Proc. Natl. Acad. Sci. USA 2012, 109, E1072–E1081. [Google Scholar] [CrossRef]
- Kim, S.; Wairkar, Y.P.; Daniels, R.W.; DiAntonio, A. The novel endosomal membrane protein Ema interacts with the class C Vps-HOPS complex to promote endosomal maturation. J. Cell Biol. 2010, 188, 717–734. [Google Scholar] [CrossRef] [PubMed]
- Schuster, C.; Gerold, K.D.; Schober, K.; Probst, L.; Boerner, K.; Kim, M.J.; Ruckdeschel, A.; Serwold, T.; Kissler, S. The Autoimmunity-Associated Gene CLEC16A Modulates Thymic Epithelial Cell Autophagy and Alters T Cell Selection. Immunity 2015, 42, 942–952. [Google Scholar] [CrossRef]
- Tam, R.C.; Li, M.W.; Gao, Y.P.; Pang, Y.T.; Yan, S.; Ge, W.; Lau, C.S.; Chan, V.S. Human CLEC16A regulates autophagy through modulating mTOR activity. Exp. Cell Res. 2017, 352, 304–312. [Google Scholar] [CrossRef]
- Soleimanpour, S.A.; Gupta, A.; Bakay, M.; Ferrari, A.M.; Groff, D.N.; Fadista, J.; Spruce, L.A.; Kushner, J.A.; Groop, L.; Seeholzer, S.H.; et al. The diabetes susceptibility gene Clec16a regulates mitophagy. Cell 2014, 157, 1577–1590. [Google Scholar] [CrossRef]
- Betz, R.C.; Petukhova, L.; Ripke, S.; Huang, H.; Menelaou, A.; Redler, S.; Becker, T.; Heilmann, S.; Yamany, T.; Duvic, M.; et al. Genome-wide meta-analysis in alopecia areata resolves HLA associations and reveals two new susceptibility loci. Nat. Commun. 2015, 6, 5966. [Google Scholar] [CrossRef]
- Rijvers, L.; Melief, M.J.; van Langelaar, J.; van der Vuurst de Vries, R.M.; Wierenga-Wolf, A.F.; Koetzier, S.C.; Priatel, J.J.; Jorritsma, T.; van Ham, S.M.; Hintzen, R.Q.; et al. The Role of Autoimmunity-Related Gene CLEC16A in the B Cell Receptor-Mediated HLA Class II Pathway. J. Immunol. 2020, 205, 945–956. [Google Scholar] [CrossRef] [PubMed]
- van Luijn, M.M.; Kreft, K.L.; Jongsma, M.L.; Mes, S.W.; Wierenga-Wolf, A.F.; van Meurs, M.; Melief, M.J.; der Kant, R.; Janssen, L.; Janssen, H.; et al. Multiple sclerosis-associated CLEC16A controls HLA class II expression via late endosome biogenesis. Brain 2015, 138, 1531–1547. [Google Scholar] [CrossRef]
- Pandey, R.; Bakay, M.; Hain, H.S.; Strenkowski, B.; Elsaqa, B.Z.B.; Roizen, J.D.; Kushner, J.A.; Orange, J.S.; Hakonarson, H. CLEC16A regulates splenocyte and NK cell function in part through MEK signaling. PLoS ONE 2018, 13, e0203952. [Google Scholar] [CrossRef]
- Pandey, R.; Bakay, M.; Hain, H.S.; Strenkowski, B.; Yermakova, A.; Kushner, J.A.; Orange, J.S.; Hakonarson, H. The Autoimmune Disorder Susceptibility Gene CLEC16A Restrains NK Cell Function in YTS NK Cell Line and Clec16a Knockout Mice. Front. Immunol. 2019, 10, 68. [Google Scholar] [CrossRef] [PubMed]
- Pandey, R.; Bakay, M.; Strenkowski, B.P.; Hain, H.S.; Hakonarson, H. JAK/STAT inhibitor therapy partially rescues the lipodystrophic autoimmune phenotype in Clec16a KO mice. Sci. Rep. 2021, 11, 7372. [Google Scholar] [CrossRef]
- Hain, H.S.; Pandey, R.; Bakay, M.; Strenkowski, B.P.; Harrington, D.; Romer, M.; Motley, W.W.; Li, J.; Lancaster, E.; Roth, L.; et al. Inducible knockout of Clec16a in mice results in sensory neurodegeneration. Sci. Rep. 2021, 11, 9319. [Google Scholar] [CrossRef]
- Redmann, V.; Lamb, C.A.; Hwang, S.; Orchard, R.C.; Kim, S.; Razi, M.; Milam, A.; Park, S.; Yokoyama, C.C.; Kambal, A.; et al. Clec16a is Critical for Autolysosome Function and Purkinje Cell Survival. Sci. Rep. 2016, 6, 23326. [Google Scholar] [CrossRef] [PubMed]
- Tomlinson, M.J.t.; Pitsillides, A.; Pickin, R.; Mika, M.; Keene, K.L.; Hou, X.; Mychaleckyj, J.; Chen, W.M.; Concannon, P.; Onengut-Gumuscu, S. Fine mapping and functional studies of risk variants for type 1 diabetes at chromosome 16p13.13. Diabetes 2014, 63, 4360–4368. [Google Scholar] [CrossRef]
- Davison, L.J.; Wallace, C.; Cooper, J.D.; Cope, N.F.; Wilson, N.K.; Smyth, D.J.; Howson, J.M.; Saleh, N.; Al-Jeffery, A.; Angus, K.L.; et al. Long-range DNA looping and gene expression analyses identify DEXI as an autoimmune disease candidate gene. Hum. Mol. Genet. 2012, 21, 322–333. [Google Scholar] [CrossRef]
- Nieves-Bonilla, J.M.; Kiaf, B.; Schuster, C.; Kissler, S. The type 1 diabetes candidate gene Dexi does not affect disease risk in the nonobese diabetic mouse model. Genes Immun. 2020, 21, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Mero, I.L.; Ban, M.; Lorentzen, A.R.; Smestad, C.; Celius, E.G.; Saether, H.; Saeedi, H.; Viken, M.K.; Skinningsrud, B.; Undlien, D.E.; et al. Exploring the CLEC16A gene reveals a MS-associated variant with correlation to the relative expression of CLEC16A isoforms in thymus. Genes Immun. 2011, 12, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Zuvich, R.L.; Bush, W.S.; McCauley, J.L.; Beecham, A.H.; De Jager, P.L.; Ivinson, A.J.; Compston, A.; Hafler, D.A.; Hauser, S.L.; Sawcer, S.J.; et al. Interrogating the complex role of chromosome 16p13.13 in multiple sclerosis susceptibility: Independent genetic signals in the CIITA-CLEC16A-SOCS1 gene complex. Hum. Mol. Genet. 2011, 20, 3517–3524. [Google Scholar] [CrossRef]
- Cooney, R.N. Suppressors of cytokine signaling (SOCS): Inhibitors of the JAK/STAT pathway. Shock 2002, 17, 83–90. [Google Scholar] [CrossRef]
- Leikfoss, I.S.; Mero, I.L.; Dahle, M.K.; Lie, B.A.; Harbo, H.F.; Spurkland, A.; Berge, T. Multiple sclerosis-associated single-nucleotide polymorphisms in CLEC16A correlate with reduced SOCS1 and DEXI expression in the thymus. Genes Immun. 2013, 14, 62–66. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Suh, J.M.; Hwang, E.S.; Kim, D.W.; Chung, H.K.; Song, J.H.; Hwang, J.H.; Park, K.C.; Ro, H.K.; Jo, E.K.; et al. Thyrotropin-mediated repression of class II trans-activator expression in thyroid cells: Involvement of STAT3 and suppressor of cytokine signaling. J. Immunol. 2003, 171, 616–627. [Google Scholar] [CrossRef]
- Yoshimura, A.; Aki, D.; Ito, M. SOCS, SPRED, and NR4a: Negative regulators of cytokine signaling and transcription in immune tolerance. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2021, 97, 277–291. [Google Scholar] [CrossRef] [PubMed]
- Sobah, M.L.; Liongue, C.; Ward, A.C. SOCS Proteins in Immunity, Inflammatory Diseases, and Immune-Related Cancer. Front. Med. (Lausanne) 2021, 8, 727987. [Google Scholar] [CrossRef] [PubMed]
- Pahlevan Kakhki, M.; Rakhshi, N.; Heidary, M.; Behmanesh, M.; Nikravesh, A. Expression of suppressor of cytokine signaling 1 (SOCS1) gene dramatically increases in relapsing-remitting multiple sclerosis. J. Neurol. Sci. 2015, 350, 40–45. [Google Scholar] [CrossRef]
- Berge, T.; Leikfoss, I.S.; Harbo, H.F. From Identification to Characterization of the Multiple Sclerosis Susceptibility Gene CLEC16A. Int. J. Mol. Sci. 2013, 14, 4476–4497. [Google Scholar] [CrossRef] [PubMed]
- Gingerich, M.A.; Liu, X.; Chai, B.; Pearson, G.L.; Vincent, M.P.; Stromer, T.; Zhu, J.; Sidarala, V.; Renberg, A.; Sahu, D.; et al. An intrinsically disordered protein region encoded by the human disease gene CLEC16A regulates mitophagy. Autophagy 2023, 19, 525–543. [Google Scholar] [CrossRef]
- Babu, M.M. The contribution of intrinsically disordered regions to protein function, cellular complexity, and human disease. Biochem. Soc. Trans. 2016, 44, 1185–1200. [Google Scholar] [CrossRef] [PubMed]
- Wright, P.E.; Dyson, H.J. Intrinsically disordered proteins in cellular signalling and regulation. Nat. Rev. Mol. Cell Biol. 2015, 16, 18–29. [Google Scholar] [CrossRef]
- The International HapMap Consortium. The International HapMap Project. Nature 2003, 426, 789–796. [Google Scholar] [CrossRef]
- The International HapMap Consortium. A haplotype map of the human genome. Nature 2005, 437, 1299–1320. [Google Scholar] [CrossRef] [PubMed]
- The Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3000 shared controls. Nature 2007, 447, 661–678. [Google Scholar] [CrossRef]
- Cooper, J.D.; Smyth, D.J.; Smiles, A.M.; Plagnol, V.; Walker, N.M.; Allen, J.E.; Downes, K.; Barrett, J.C.; Healy, B.C.; Mychaleckyj, J.C.; et al. Meta-analysis of genome-wide association study data identifies additional type 1 diabetes risk loci. Nat. Genet. 2008, 40, 1399–1401. [Google Scholar] [CrossRef] [PubMed]
- Todd, J.A.; Walker, N.M.; Cooper, J.D.; Smyth, D.J.; Downes, K.; Plagnol, V.; Bailey, R.; Nejentsev, S.; Field, S.F.; Payne, F.; et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat. Genet. 2007, 39, 857–864. [Google Scholar] [CrossRef]
- Zoledziewska, M.; Costa, G.; Pitzalis, M.; Cocco, E.; Melis, C.; Moi, L.; Zavattari, P.; Murru, R.; Lampis, R.; Morelli, L.; et al. Variation within the CLEC16A gene shows consistent disease association with both multiple sclerosis and type 1 diabetes in Sardinia. Genes Immun. 2009, 10, 15–17. [Google Scholar] [CrossRef] [PubMed]
- Martinez, A.; Perdigones, N.; Cenit, M.C.; Espino, L.; Varade, J.; Lamas, J.R.; Santiago, J.L.; Fernandez-Arquero, M.; de la Calle, H.; Arroyo, R.; et al. Chromosomal region 16p13: Further evidence of increased predisposition to immune diseases. Ann. Rheum. Dis. 2010, 69, 309–311. [Google Scholar] [CrossRef]
- Reddy, M.V.; Wang, H.; Liu, S.; Bode, B.; Reed, J.C.; Steed, R.D.; Anderson, S.W.; Steed, L.; Hopkins, D.; She, J.X. Association between type 1 diabetes and GWAS SNPs in the southeast US Caucasian population. Genes Immun. 2011, 12, 208–212. [Google Scholar] [CrossRef]
- Sang, Y.; Zong, W.; Yan, J.; Liu, M. The Correlation between the CLEC16A Gene and Genetic Susceptibility to Type 1 Diabetes in Chinese Children. Int. J. Endocrinol. 2012, 2012, 245384. [Google Scholar] [CrossRef]
- Wu, X.; Zhu, X.; Wang, X.; Ma, J.; Zhu, S.; Li, J.; Liu, Y. Intron polymorphism in the KIAA0350 gene is reproducibly associated with susceptibility to type 1 diabetes (T1D) in the Han Chinese population. Clin. Endocrinol. 2009, 71, 46–49. [Google Scholar] [CrossRef]
- Yamashita, H.; Awata, T.; Kawasaki, E.; Ikegami, H.; Tanaka, S.; Maruyama, T.; Shimada, A.; Nakanishi, K.; Takahashi, K.; Kobayashi, T.; et al. Analysis of the HLA and non-HLA susceptibility loci in Japanese type 1 diabetes. Diabetes/Metab. Res. Rev. 2011, 27, 844–848. [Google Scholar] [CrossRef]
- Howson, J.M.; Rosinger, S.; Smyth, D.J.; Boehm, B.O.; Todd, J.A. Genetic analysis of adult-onset autoimmune diabetes. Diabetes 2011, 60, 2645–2653. [Google Scholar] [CrossRef]
- Awata, T.; Kawasaki, E.; Tanaka, S.; Ikegami, H.; Maruyama, T.; Shimada, A.; Nakanishi, K.; Kobayashi, T.; Iizuka, H.; Uga, M.; et al. Association of type 1 diabetes with two Loci on 12q13 and 16p13 and the influence coexisting thyroid autoimmunity in Japanese. J. Clin. Endocrinol. Metab. 2009, 94, 231–235. [Google Scholar] [CrossRef] [PubMed]
- Barrett, J.C.; Clayton, D.G.; Concannon, P.; Akolkar, B.; Cooper, J.D.; Erlich, H.A.; Julier, C.; Morahan, G.; Nerup, J.; Nierras, C.; et al. Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat. Genet. 2009, 41, 703–707. [Google Scholar] [CrossRef] [PubMed]
- Plagnol, V.; Howson, J.M.; Smyth, D.J.; Walker, N.; Hafler, J.P.; Wallace, C.; Stevens, H.; Jackson, L.; Simmonds, M.J.; Type 1 Diabetes Genetics, C.; et al. Genome-wide association analysis of autoantibody positivity in type 1 diabetes cases. PLoS Genet. 2011, 7, e1002216. [Google Scholar] [CrossRef]
- Onengut-Gumuscu, S.; Chen, W.M.; Burren, O.; Cooper, N.J.; Quinlan, A.R.; Mychaleckyj, J.C.; Farber, E.; Bonnie, J.K.; Szpak, M.; Schofield, E.; et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat. Genet. 2015, 47, 381–386. [Google Scholar] [CrossRef]
- Hafler, D.A.; Compston, A.; Sawcer, S.; Lander, E.S.; Daly, M.J.; De Jager, P.L.; de Bakker, P.I.; Gabriel, S.B.; Mirel, D.B.; Ivinson, A.J.; et al. Risk alleles for multiple sclerosis identified by a genomewide study. N. Engl. J. Med. 2007, 357, 851–862. [Google Scholar] [CrossRef] [PubMed]
- Rubio, J.P.; Stankovich, J.; Field, J.; Tubridy, N.; Marriott, M.; Chapman, C.; Bahlo, M.; Perera, D.; Johnson, L.J.; Tait, B.D.; et al. Replication of KIAA0350, IL2RA, RPL5 and CD58 as multiple sclerosis susceptibility genes in Australians. Genes Immun. 2008, 9, 624–630. [Google Scholar] [CrossRef]
- D’Netto, M.J.; Ward, H.; Morrison, K.M.; Ramagopalan, S.V.; Dyment, D.A.; DeLuca, G.C.; Handunnetthi, L.; Sadovnick, A.D.; Ebers, G.C. Risk alleles for multiple sclerosis in multiplex families. Neurology 2009, 72, 1984–1988. [Google Scholar] [CrossRef]
- De Jager, P.L.; Jia, X.; Wang, J.; de Bakker, P.I.; Ottoboni, L.; Aggarwal, N.T.; Piccio, L.; Raychaudhuri, S.; Tran, D.; Aubin, C.; et al. Meta-analysis of genome scans and replication identify CD6, IRF8 and TNFRSF1A as new multiple sclerosis susceptibility loci. Nat. Genet. 2009, 41, 776–782. [Google Scholar] [CrossRef]
- International Multiple Sclerosis Genetics, C. The expanding genetic overlap between multiple sclerosis and type I diabetes. Genes Immun. 2009, 10, 11–14. [Google Scholar] [CrossRef]
- Perera, D.; Stankovich, J.; Butzkueven, H.; Taylor, B.V.; Foote, S.J.; Kilpatrick, T.J.; Rubio, J.P. Fine mapping of multiple sclerosis susceptibility genes provides evidence of allelic heterogeneity at the IL2RA locus. J. Neuroimmunol. 2009, 211, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Johnson, B.A.; Wang, J.; Taylor, E.M.; Caillier, S.J.; Herbert, J.; Khan, O.A.; Cross, A.H.; De Jager, P.L.; Gourraud, P.A.; Cree, B.C.; et al. Multiple sclerosis susceptibility alleles in African Americans. Genes Immun. 2010, 11, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Nischwitz, S.; Cepok, S.; Kroner, A.; Wolf, C.; Knop, M.; Muller-Sarnowski, F.; Pfister, H.; Rieckmann, P.; Hemmer, B.; Ising, M.; et al. More CLEC16A gene variants associated with multiple sclerosis. Acta Neurol. Scand 2011, 123, 400–406. [Google Scholar] [CrossRef] [PubMed]
- Pandit, L.; Ban, M.; Sawcer, S.; Singhal, B.; Nair, S.; Radhakrishnan, K.; Shetty, R.; Misri, Z.; Hegde, S.; Bhat, I.G. Evaluation of the established non-MHC multiple sclerosis loci in an Indian population. Mult. Scler. 2011, 17, 139–143. [Google Scholar] [CrossRef]
- Sawcer, S.; Hellenthal, G.; Pirinen, M.; Spencer, C.C.; Patsopoulos, N.A.; Moutsianas, L.; Dilthey, A.; Su, Z.; Freeman, C.; Hunt, S.E.; et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 2011, 476, 214–219. [Google Scholar] [CrossRef]
- International Multiple Sclerosis Genetics, C.; Beecham, A.H.; Patsopoulos, N.A.; Xifara, D.K.; Davis, M.F.; Kemppinen, A.; Cotsapas, C.; Shah, T.S.; Spencer, C.; Booth, D.; et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat. Genet. 2013, 45, 1353–1360. [Google Scholar] [CrossRef]
- Andlauer, T.F.; Buck, D.; Antony, G.; Bayas, A.; Bechmann, L.; Berthele, A.; Chan, A.; Gasperi, C.; Gold, R.; Graetz, C.; et al. Novel multiple sclerosis susceptibility loci implicated in epigenetic regulation. Sci. Adv. 2016, 2, e1501678. [Google Scholar] [CrossRef]
- Steri, M.; Orru, V.; Idda, M.L.; Pitzalis, M.; Pala, M.; Zara, I.; Sidore, C.; Faa, V.; Floris, M.; Deiana, M.; et al. Overexpression of the Cytokine BAFF and Autoimmunity Risk. N. Engl. J. Med. 2017, 376, 1615–1626. [Google Scholar] [CrossRef]
- Eriksson, D.; Bianchi, M.; Landegren, N.; Dalin, F.; Skov, J.; Hultin-Rosenberg, L.; Mathioudaki, A.; Nordin, J.; Hallgren, A.; Andersson, G.; et al. Common genetic variation in the autoimmune regulator (AIRE) locus is associated with autoimmune Addison’s disease in Sweden. Sci. Rep. 2018, 8, 8395. [Google Scholar] [CrossRef]
- Skinningsrud, B.; Husebye, E.S.; Pearce, S.H.; McDonald, D.O.; Brandal, K.; Wolff, A.B.; Lovas, K.; Egeland, T.; Undlien, D.E. Polymorphisms in CLEC16A and CIITA at 16p13 are associated with primary adrenal insufficiency. J. Clin. Endocrinol. Metab. 2008, 93, 3310–3317. [Google Scholar] [CrossRef]
- Langefeld, C.D.; Ainsworth, H.C.; Cunninghame Graham, D.S.; Kelly, J.A.; Comeau, M.E.; Marion, M.C.; Howard, T.D.; Ramos, P.S.; Croker, J.A.; Morris, D.L.; et al. Transancestral mapping and genetic load in systemic lupus erythematosus. Nat. Commun. 2017, 8, 16021. [Google Scholar] [CrossRef] [PubMed]
- Morris, D.L.; Sheng, Y.; Zhang, Y.; Wang, Y.F.; Zhu, Z.; Tombleson, P.; Chen, L.; Cunninghame Graham, D.S.; Bentham, J.; Roberts, A.L.; et al. Genome-wide association meta-analysis in Chinese and European individuals identifies ten new loci associated with systemic lupus erythematosus. Nat. Genet. 2016, 48, 940–946. [Google Scholar] [CrossRef] [PubMed]
- Bentham, J.; Morris, D.L.; Graham, D.S.C.; Pinder, C.L.; Tombleson, P.; Behrens, T.W.; Martin, J.; Fairfax, B.P.; Knight, J.C.; Chen, L.; et al. Genetic association analyses implicate aberrant regulation of innate and adaptive immunity genes in the pathogenesis of systemic lupus erythematosus. Nat. Genet. 2015, 47, 1457–1464. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Tang, H.; Zhang, Y.; Tang, X.; Zhang, J.; Sun, L.; Yang, J.; Cui, Y.; Zhang, L.; Hirankarn, N.; et al. Meta-analysis followed by replication identifies loci in or near CDKN1B, TET3, CD80, DRAM1, and ARID5B as associated with systemic lupus erythematosus in Asians. Am. J. Hum. Genet. 2013, 92, 41–51. [Google Scholar] [CrossRef]
- Marquez, A.; Varade, J.; Robledo, G.; Martinez, A.; Mendoza, J.L.; Taxonera, C.; Fernandez-Arquero, M.; Diaz-Rubio, M.; Gomez-Garcia, M.; Lopez-Nevot, M.A.; et al. Specific association of a CLEC16A/KIAA0350 polymorphism with NOD2/CARD15(-) Crohn’s disease patients. Eur. J. Hum. Genet. 2009, 17, 1304–1308. [Google Scholar] [CrossRef]
- Bronson, P.G.; Chang, D.; Bhangale, T.; Seldin, M.F.; Ortmann, W.; Ferreira, R.C.; Urcelay, E.; Pereira, L.F.; Martin, J.; Plebani, A.; et al. Common variants at PVT1, ATG13-AMBRA1, AHI1 and CLEC16A are associated with selective IgA deficiency. Nat. Genet. 2016, 48, 1425–1429. [Google Scholar] [CrossRef]
- Jagielska, D.; Redler, S.; Brockschmidt, F.F.; Herold, C.; Pasternack, S.M.; Garcia Bartels, N.; Hanneken, S.; Eigelshoven, S.; Refke, M.; Barth, S.; et al. Follow-up study of the first genome-wide association scan in alopecia areata: IL13 and KIAA0350 as susceptibility loci supported with genome-wide significance. J. Investig. Dermatol. 2012, 132, 2192–2197. [Google Scholar] [CrossRef]
- Skinningsrud, B.; Lie, B.A.; Husebye, E.S.; Kvien, T.K.; Forre, O.; Flato, B.; Stormyr, A.; Joner, G.; Njolstad, P.R.; Egeland, T.; et al. A CLEC16A variant confers risk for juvenile idiopathic arthritis and anti-cyclic citrullinated peptide antibody negative rheumatoid arthritis. Ann. Rheum. Dis. 2010, 69, 1471–1474. [Google Scholar] [CrossRef]
- Cordell, H.J.; Han, Y.; Mells, G.F.; Li, Y.; Hirschfield, G.M.; Greene, C.S.; Xie, G.; Juran, B.D.; Zhu, D.; Qian, D.C.; et al. International genome-wide meta-analysis identifies new primary biliary cirrhosis risk loci and targetable pathogenic pathways. Nat. Commun. 2015, 6, 8019. [Google Scholar] [CrossRef]
- Liu, J.Z.; Almarri, M.A.; Gaffney, D.J.; Mells, G.F.; Jostins, L.; Cordell, H.J.; Ducker, S.J.; Day, D.B.; Heneghan, M.A.; Neuberger, J.M.; et al. Dense fine-mapping study identifies new susceptibility loci for primary biliary cirrhosis. Nat. Genet. 2012, 44, 1137–1141. [Google Scholar] [CrossRef]
- Hirschfield, G.M.; Xie, G.; Lu, E.; Sun, Y.; Juran, B.D.; Chellappa, V.; Coltescu, C.; Mason, A.L.; Milkiewicz, P.; Myers, R.P.; et al. Association of primary biliary cirrhosis with variants in the CLEC16A, SOCS1, SPIB and SIAE immunomodulatory genes. Genes Immun. 2012, 13, 328–335. [Google Scholar] [CrossRef] [PubMed]
- Mells, G.F.; Floyd, J.A.; Morley, K.I.; Cordell, H.J.; Franklin, C.S.; Shin, S.Y.; Heneghan, M.A.; Neuberger, J.M.; Donaldson, P.T.; Day, D.B.; et al. Genome-wide association study identifies 12 new susceptibility loci for primary biliary cirrhosis. Nat. Genet. 2011, 43, 329–332. [Google Scholar] [CrossRef]
- Valette, K.; Li, Z.; Bon-Baret, V.; Chignon, A.; Berube, J.C.; Eslami, A.; Lamothe, J.; Gaudreault, N.; Joubert, P.; Obeidat, M.; et al. Prioritization of candidate causal genes for asthma in susceptibility loci derived from UK Biobank. Commun. Biol. 2021, 4, 700. [Google Scholar] [CrossRef]
- Shrine, N.; Portelli, M.A.; John, C.; Soler Artigas, M.; Bennett, N.; Hall, R.; Lewis, J.; Henry, A.P.; Billington, C.K.; Ahmad, A.; et al. Moderate-to-severe asthma in individuals of European ancestry: A genome-wide association study. Lancet Respir. Med. 2019, 7, 20–34. [Google Scholar] [CrossRef]
- Pividori, M.; Schoettler, N.; Nicolae, D.L.; Ober, C.; Im, H.K. Shared and distinct genetic risk factors for childhood-onset and adult-onset asthma: Genome-wide and transcriptome-wide studies. Lancet Respir. Med. 2019, 7, 509–522. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Lee, P.H.; Chaffin, M.D.; Chung, W.; Loh, P.R.; Lu, Q.; Christiani, D.C.; Liang, L. A genome-wide cross-trait analysis from UK Biobank highlights the shared genetic architecture of asthma and allergic diseases. Nat. Genet. 2018, 50, 857–864. [Google Scholar] [CrossRef] [PubMed]
- Demenais, F.; Margaritte-Jeannin, P.; Barnes, K.C.; Cookson, W.O.C.; Altmuller, J.; Ang, W.; Barr, R.G.; Beaty, T.H.; Becker, A.B.; Beilby, J.; et al. Multiancestry association study identifies new asthma risk loci that colocalize with immune-cell enhancer marks. Nat. Genet. 2018, 50, 42–53. [Google Scholar] [CrossRef]
- Almoguera, B.; Vazquez, L.; Mentch, F.; Connolly, J.; Pacheco, J.A.; Sundaresan, A.S.; Peissig, P.L.; Linneman, J.G.; McCarty, C.A.; Crosslin, D.; et al. Identification of Four Novel Loci in Asthma in European American and African American Populations. Am. J. Respir. Crit. Care Med. 2017, 195, 456–463. [Google Scholar] [CrossRef]
- Pickrell, J.K.; Berisa, T.; Liu, J.Z.; Segurel, L.; Tung, J.Y.; Hinds, D.A. Detection and interpretation of shared genetic influences on 42 human traits. Nat. Genet. 2016, 48, 709–717. [Google Scholar] [CrossRef]
- Niu, Y.; Wang, H.; Li, Z.; Shamsi, B.H.; Liu, M.; Liu, J.; Wang, Q.; Liu, Y. CLEC16A variants conferred a decreased risk to allergic rhinitis in the Chinese population. Front. Genet. 2022, 13, 1053761. [Google Scholar] [CrossRef]
- Gao, Y.; Li, J.; Zhang, Y.; Zhang, L. Replication study of susceptibility variants associated with allergic rhinitis and allergy in Han Chinese. Allergy Asthma Clin. Immunol. 2020, 16, 13. [Google Scholar] [CrossRef]
- Muhali, F.S.; Cai, T.T.; Zhu, J.L.; Qin, Q.; Xu, J.; He, S.T.; Shi, X.H.; Jiang, W.J.; Xiao, L.; Li, D.F.; et al. Polymorphisms of CLEC16A Region and Autoimmune Thyroid Diseases. G3 (Bethesda) 2014, 4, 973–977. [Google Scholar] [CrossRef]
- Li, J.; Jorgensen, S.F.; Maggadottir, S.M.; Bakay, M.; Warnatz, K.; Glessner, J.; Pandey, R.; Salzer, U.; Schmidt, R.E.; Perez, E.; et al. Association of CLEC16A with human common variable immunodeficiency disorder and role in murine B cells. Nat. Commun. 2015, 6, 6804. [Google Scholar] [CrossRef]
- Kottyan, L.C.; Maddox, A.; Braxton, J.R.; Stucke, E.M.; Mukkada, V.; Putnam, P.E.; Abonia, J.P.; Chehade, M.; Wood, R.A.; Pesek, R.D.; et al. Genetic variants at the 16p13 locus confer risk for eosinophilic esophagitis. Genes Immun. 2019, 20, 281–292. [Google Scholar] [CrossRef]
- Dubois, P.C.; Trynka, G.; Franke, L.; Hunt, K.A.; Romanos, J.; Curtotti, A.; Zhernakova, A.; Heap, G.A.; Adany, R.; Aromaa, A.; et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 2010, 42, 295–302. [Google Scholar] [CrossRef]
- Gorlova, O.Y.; Li, Y.; Gorlov, I.; Ying, J.; Chen, W.V.; Assassi, S.; Reveille, J.D.; Arnett, F.C.; Zhou, X.; Bossini-Castillo, L.; et al. Gene-level association analysis of systemic sclerosis: A comparison of African-Americans and White populations. PLoS ONE 2018, 13, e0189498. [Google Scholar] [CrossRef]
- Strafella, C.; Caputo, V.; Termine, A.; Assogna, F.; Pellicano, C.; Pontieri, F.E.; Macchiusi, L.; Minozzi, G.; Gambardella, S.; Centonze, D.; et al. Immune System and Neuroinflammation in Idiopathic Parkinson’s Disease: Association Analysis of Genetic Variants and miRNAs Interactions. Front. Genet. 2021, 12, 651971. [Google Scholar] [CrossRef] [PubMed]
- Fan, H.H.; Cui, L.; Jiang, X.X.; Song, Y.D.; Liu, S.S.; Wu, K.Y.; Dong, H.J.; Mao, M.; Ovlyakulov, B.; Wu, H.M.; et al. Autoimmune Disease Associated CLEC16A Variants Convey Risk of Parkinson’s Disease in Han Chinese. Front. Genet. 2022, 13, 856493. [Google Scholar] [CrossRef] [PubMed]
- Polymeropoulos, M.H.; Lavedan, C.; Leroy, E.; Ide, S.E.; Dehejia, A.; Dutra, A.; Pike, B.; Root, H.; Rubenstein, J.; Boyer, R.; et al. Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 1997, 276, 2045–2047. [Google Scholar] [CrossRef] [PubMed]
- Kitada, T.; Asakawa, S.; Hattori, N.; Matsumine, H.; Yamamura, Y.; Minoshima, S.; Yokochi, M.; Mizuno, Y.; Shimizu, N. Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 1998, 392, 605–608. [Google Scholar] [CrossRef]
- Bonifati, V.; Rizzu, P.; van Baren, M.J.; Schaap, O.; Breedveld, G.J.; Krieger, E.; Dekker, M.C.; Squitieri, F.; Ibanez, P.; Joosse, M.; et al. Mutations in the DJ-1 gene associated with autosomal recessive early-onset parkinsonism. Science 2003, 299, 256–259. [Google Scholar] [CrossRef]
- Valente, E.M.; Abou-Sleiman, P.M.; Caputo, V.; Muqit, M.M.; Harvey, K.; Gispert, S.; Ali, Z.; Del Turco, D.; Bentivoglio, A.R.; Healy, D.G.; et al. Hereditary early-onset Parkinson’s disease caused by mutations in PINK1. Science 2004, 304, 1158–1160. [Google Scholar] [CrossRef]
- Zimprich, A.; Muller-Myhsok, B.; Farrer, M.; Leitner, P.; Sharma, M.; Hulihan, M.; Lockhart, P.; Strongosky, A.; Kachergus, J.; Calne, D.B.; et al. The PARK8 locus in autosomal dominant parkinsonism: Confirmation of linkage and further delineation of the disease-containing interval. Am. J. Hum. Genet. 2004, 74, 11–19. [Google Scholar] [CrossRef]
- Paisan-Ruiz, C.; Jain, S.; Evans, E.W.; Gilks, W.P.; Simon, J.; van der Brug, M.; Lopez de Munain, A.; Aparicio, S.; Gil, A.M.; Khan, N.; et al. Cloning of the gene containing mutations that cause PARK8-linked Parkinson’s disease. Neuron 2004, 44, 595–600. [Google Scholar] [CrossRef]
- Schreglmann, S.R.; Houlden, H. VPS13C-Another Hint at Mitochondrial Dysfunction in Familial Parkinson’s Disease. Mov. Disord. 2016, 31, 1340. [Google Scholar] [CrossRef] [PubMed]
- Monfrini, E.; Spagnolo, F.; Canesi, M.; Seresini, A.; Rini, A.; Passarella, B.; Percetti, M.; Seia, M.; Goldwurm, S.; Cereda, V.; et al. VPS13C-associated Parkinson’s disease: Two novel cases and review of the literature. Park. Relat. Disord. 2022, 94, 37–39. [Google Scholar] [CrossRef]
- Darvish, H.; Bravo, P.; Tafakhori, A.; Azcona, L.J.; Ranji-Burachaloo, S.; Johari, A.H.; Paisan-Ruiz, C. Identification of a large homozygous VPS13C deletion in a patient with early-onset Parkinsonism. Mov. Disord. 2018, 33, 1968–1970. [Google Scholar] [CrossRef] [PubMed]
- Aharon-Peretz, J.; Rosenbaum, H.; Gershoni-Baruch, R. Mutations in the glucocerebrosidase gene and Parkinson’s disease in Ashkenazi Jews. N. Engl. J. Med. 2004, 351, 1972–1977. [Google Scholar] [CrossRef]
- Witoelar, A.; Jansen, I.E.; Wang, Y.; Desikan, R.S.; Gibbs, J.R.; Blauwendraat, C.; Thompson, W.K.; Hernandez, D.G.; Djurovic, S.; Schork, A.J.; et al. Genome-wide Pleiotropy Between Parkinson Disease and Autoimmune Diseases. JAMA Neurol. 2017, 74, 780–792. [Google Scholar] [CrossRef]
- Sliter, D.A.; Martinez, J.; Hao, L.; Chen, X.; Sun, N.; Fischer, T.D.; Burman, J.L.; Li, Y.; Zhang, Z.; Narendra, D.P.; et al. Parkin and PINK1 mitigate STING-induced inflammation. Nature 2018, 561, 258–262. [Google Scholar] [CrossRef] [PubMed]
- van Horssen, J.; van Schaik, P.; Witte, M. Inflammation and mitochondrial dysfunction: A vicious circle in neurodegenerative disorders? Neurosci. Lett. 2019, 710, 132931. [Google Scholar] [CrossRef]
- Li, M.; Wan, J.; Xu, Z.; Tang, B. The association between Parkinson’s disease and autoimmune diseases: A systematic review and meta-analysis. Front. Immunol. 2023, 14, 1103053. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, M.A.; Matheson, M.C.; Tang, C.S.; Granell, R.; Ang, W.; Hui, J.; Kiefer, A.K.; Duffy, D.L.; Baltic, S.; Danoy, P.; et al. Genome-wide association analysis identifies 11 risk variants associated with the asthma with hay fever phenotype. J. Allergy Clin. Immunol. 2013, 13, 1564–1571. [Google Scholar] [CrossRef]
- Ferreira, M.A.R.; Mathur, R.; Vonk, J.M.; Szwajda, A.; Brumpton, B.; Granell, R.; Brew, B.K.; Ullemar, V.; Lu, Y.; Jiang, Y.; et al. Genetic Architectures of Childhood- and Adult-Onset Asthma Are Partly Distinct. Am. J. Hum. Genet. 2019, 104, 665–684. [Google Scholar] [CrossRef] [PubMed]
- Hoppenbrouwers, I.A.; Aulchenko, Y.S.; Janssens, A.C.; Ramagopalan, S.V.; Broer, L.; Kayser, M.; Ebers, G.C.; Oostra, B.A.; van Duijn, C.M.; Hintzen, R.Q. Replication of CD58 and CLEC16A as genome-wide significant risk genes for multiple sclerosis. J. Hum. Genet. 2009, 54, 676–680. [Google Scholar] [CrossRef]
- Condello, M.; Pellegrini, E.; Caraglia, M.; Meschini, S. Targeting Autophagy to Overcome Human Diseases. Int. J. Mol. Sci. 2019, 20, 725. [Google Scholar] [CrossRef]
- Grosjean, I.; Romeo, B.; Domdom, M.A.; Belaid, A.; D’Andrea, G.; Guillot, N.; Gherardi, R.K.; Gal, J.; Milano, G.; Marquette, C.H.; et al. Autophagopathies: From autophagy gene polymorphisms to precision medicine for human diseases. Autophagy 2022, 18, 2519–2536. [Google Scholar] [CrossRef]
- Chong, Z.Z. mTOR: A Novel Therapeutic Target for Diseases of Multiple Systems. Curr. Drug Targets 2015, 16, 1107–1132. [Google Scholar] [CrossRef]
- Ballesteros-Alvarez, J.; Andersen, J.K. mTORC2: The other mTOR in autophagy regulation. Aging Cell 2021, 20, e13431. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.C.; Guan, K.L. mTOR: A pharmacologic target for autophagy regulation. J. Clin. Investig. 2015, 125, 25–32. [Google Scholar] [CrossRef]
- Jung, C.H.; Ro, S.H.; Cao, J.; Otto, N.M.; Kim, D.H. mTOR regulation of autophagy. FEBS Lett. 2010, 584, 1287–1295. [Google Scholar] [CrossRef] [PubMed]
- Leikfoss, I.S.; Keshari, P.K.; Gustavsen, M.W.; Bjolgerud, A.; Brorson, I.S.; Celius, E.G.; Spurkland, A.; Bos, S.D.; Harbo, H.F.; Berge, T. Multiple Sclerosis Risk Allele in CLEC16A Acts as an Expression Quantitative Trait Locus for CLEC16A and SOCS1 in CD4+ T Cells. PLoS ONE 2015, 10, e0132957. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, A.M.; Leikfoss, I.S.; Abrahamsen, G.; Sundvold, V.; Isom, M.M.; Keshari, P.K.; Rognes, T.; Landsverk, O.J.B.; Bos, S.D.; Harbo, H.F.; et al. Exploring the role of the multiple sclerosis susceptibility gene CLEC16A in T cells. Scand J. Immunol. 2021, 94, e13050. [Google Scholar] [CrossRef] [PubMed]
- Hardman, J.A.; Nicu, C.; Tai, C.; Harries, M.; Mironov, A.; Purba, T.S.; Paus, R. Does dysfunctional autophagy contribute to immune privilege collapse and alopecia areata pathogenesis? J. Derm. Sci. 2020, 100, 75–78. [Google Scholar] [CrossRef]
- Sundberg, J.P.; Boggess, D.; Montagutelli, X.; Hogan, M.E.; King, L.E., Jr. C3H/HeJ mouse model for alopecia areata. J. Investig. Derm. 1995, 104, 16S–17S. [Google Scholar] [CrossRef]
- Bjorkoy, G.; Lamark, T.; Brech, A.; Outzen, H.; Perander, M.; Overvatn, A.; Stenmark, H.; Johansen, T. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J. Cell Biol. 2005, 171, 603–614. [Google Scholar] [CrossRef]
- Gund, R.; Christiano, A.M. Impaired autophagy promotes hair loss in the C3H/HeJ mouse model of alopecia areata. Autophagy 2023, 19, 296–305. [Google Scholar] [CrossRef]
- Zia, A.; Farkhondeh, T.; Pourbagher-Shahri, A.M.; Samarghandian, S. The Roles of Mitochondrial Dysfunction and Reactive Oxygen Species in Aging and Senescence. Curr. Mol. Med. 2022, 22, 37–49. [Google Scholar] [CrossRef]
- Ko, T.K.; Tan, D.J.Y. Is Disrupted Mitophagy a Central Player to Parkinson’s Disease Pathology? Cureus 2023, 15, e35458. [Google Scholar] [CrossRef]
- Miller, S.; Muqit, M.M.K. Therapeutic approaches to enhance PINK1/Parkin mediated mitophagy for the treatment of Parkinson’s disease. Neurosci. Lett. 2019, 705, 7–13. [Google Scholar] [CrossRef]
- Vives-Bauza, C.; Przedborski, S. Mitophagy: The latest problem for Parkinson’s disease. Trends Mol. Med. 2011, 17, 158–165. [Google Scholar] [CrossRef]
- Pearson, G.; Chai, B.; Vozheiko, T.; Liu, X.; Kandarpa, M.; Piper, R.C.; Soleimanpour, S.A. Clec16a, Nrdp1, and USP8 Form a Ubiquitin-Dependent Tripartite Complex That Regulates beta-Cell Mitophagy. Diabetes 2018, 67, 265–277. [Google Scholar] [CrossRef]
- Smits, D.J.; Dekker, J.; Schot, R.; Tabarki, B.; Alhashem, A.; Demmers, J.A.A.; Dekkers, D.H.W.; Romito, A.; van der Spek, P.J.; van Ham, T.J.; et al. CLEC16A interacts with retromer and TRIM27, and its loss impairs endosomal trafficking and neurodevelopment. Hum. Genet. 2023, 142, 379–397. [Google Scholar] [CrossRef] [PubMed]
- Su, A.I.; Cooke, M.P.; Ching, K.A.; Hakak, Y.; Walker, J.R.; Wiltshire, T.; Orth, A.P.; Vega, R.G.; Sapinoso, L.M.; Moqrich, A.; et al. Large-scale analysis of the human and mouse transcriptomes. Proc. Natl. Acad. Sci. USA 2002, 99, 4465–4470. [Google Scholar] [CrossRef]
- Wu, C.; Orozco, C.; Boyer, J.; Leglise, M.; Goodale, J.; Batalov, S.; Hodge, C.L.; Haase, J.; Janes, J.; Huss, J.W., 3rd; et al. BioGPS: An extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol. 2009, 10, R130. [Google Scholar] [CrossRef]
- Askenase, M.H. Yo GABA GABA! A neurotransmitter interrupts DC-NK crosstalk. J. Leukoc. Biol. 2021, 110, 611–612. [Google Scholar] [CrossRef]
- Thomas, R.; Yang, X. NK-DC Crosstalk in Immunity to Microbial Infection. J. Immunol. Res. 2016, 2016, 6374379. [Google Scholar] [CrossRef]
- Marcenaro, E.; Carlomagno, S.; Pesce, S.; Moretta, A.; Sivori, S. NK/DC crosstalk in anti-viral response. Adv. Exp. Med. Biol. 2012, 946, 295–308. [Google Scholar] [CrossRef]
- Zouk, H.; D’Hennezel, E.; Du, X.; Ounissi-Benkalha, H.; Piccirillo, C.A.; Polychronakos, C. Functional evaluation of the role of C-type lectin domain family 16A at the chromosome 16p13 locus. Clin. Exp. Immunol. 2014, 175, 485–497. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Li, J.; Chen, C.; Yan, Y.; Jiang, S.; Wu, X.; Shao, B.; Xu, J.; Kang, L.; Huang, Y.; et al. Involvement of CLEC16A in activation of astrocytes after LPS treated. Neurochem. Res. 2012, 37, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Villarroya-Beltri, C.; Guerra, S.; Sanchez-Madrid, F. ISGylation—A key to lock the cell gates for preventing the spread of threats. J. Cell Sci. 2017, 130, 2961–2969. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Zhang, D.E. Interferon-stimulated gene 15 and the protein ISGylation system. J. Interferon Cytokine Res. 2011, 31, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.G.; Kaul, M.; Zhang, D.X. Interferon-stimulated gene 15 as a general marker for acute and chronic neuronal injuries. Sheng Li Xue Bao 2012, 64, 577–583. [Google Scholar]
- Desai, S.; Juncker, M.; Kim, C. Regulation of mitophagy by the ubiquitin pathway in neurodegenerative diseases. Exp. Biol. Med. (Maywood) 2018, 243, 554–562. [Google Scholar] [CrossRef]
- Albert, M.; Becares, M.; Falqui, M.; Fernandez-Lozano, C.; Guerra, S. ISG15, a Small Molecule with Huge Implications: Regulation of Mitochondrial Homeostasis. Viruses 2018, 10, 629. [Google Scholar] [CrossRef]
- Im, E.; Yoo, L.; Hyun, M.; Shin, W.H.; Chung, K.C. Covalent ISG15 conjugation positively regulates the ubiquitin E3 ligase activity of parkin. Open Biol. 2016, 6, 160193. [Google Scholar] [CrossRef]
- D’Cunha, J.; Knight, E., Jr.; Haas, A.L.; Truitt, R.L.; Borden, E.C. Immunoregulatory properties of ISG15, an interferon-induced cytokine. Proc. Natl. Acad. Sci. USA 1996, 93, 211–215. [Google Scholar] [CrossRef]
- Hofer, S.J.; Liang, Y.; Zimmermann, A.; Schroeder, S.; Dengjel, J.; Kroemer, G.; Eisenberg, T.; Sigrist, S.J.; Madeo, F. Spermidine-induced hypusination preserves mitochondrial and cognitive function during aging. Autophagy 2021, 17, 2037–2039. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Zhang, M.; Dai, Y.; Sun, Y.; Aman, Y.; Xu, Y.; Yu, P.; Zheng, Y.; Yang, J.; Zhu, X. Spermidine inhibits neurodegeneration and delays aging via the PINK1-PDR1-dependent mitophagy pathway in C. elegans. Aging (Albany NY) 2020, 12, 16852–16866. [Google Scholar] [CrossRef]
- Qi, Y.; Qiu, Q.; Gu, X.; Tian, Y.; Zhang, Y. ATM mediates spermidine-induced mitophagy via PINK1 and Parkin regulation in human fibroblasts. Sci. Rep. 2016, 6, 24700. [Google Scholar] [CrossRef]
- Ryu, D.; Mouchiroud, L.; Andreux, P.A.; Katsyuba, E.; Moullan, N.; Nicolet-Dit-Felix, A.A.; Williams, E.G.; Jha, P.; Lo Sasso, G.; Huzard, D.; et al. Urolithin A induces mitophagy and prolongs lifespan in C. elegans and increases muscle function in rodents. Nat. Med. 2016, 22, 879–888. [Google Scholar] [CrossRef]
- Cho, S.I.; Jo, E.R.; Song, H. Urolithin A attenuates auditory cell senescence by activating mitophagy. Sci. Rep. 2022, 12, 7704. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Song, Y.; Chen, L.; Chen, P.; Yuan, M.; Meng, Y.; Wang, Q.; Zheng, G.; Qiu, Z. Urolithin A Attenuates Hyperuricemic Nephropathy in Fructose-Fed Mice by Impairing STING-NLRP3 Axis-Mediated Inflammatory Response via Restoration of Parkin-Dependent Mitophagy. Front. Pharm. 2022, 13, 907209. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.; Chen, Y.; Zhuo, J.; Zhang, L.; Liu, J.; Wang, B.; Sun, D.; Yu, S.; Lou, H. Urolithin A promotes mitophagy and suppresses NLRP3 inflammasome activation in lipopolysaccharide-induced BV2 microglial cells and MPTP-induced Parkinson’s disease model. Neuropharmacology 2022, 207, 108963. [Google Scholar] [CrossRef]
- Wang, H.; Fu, J.; Xu, X.; Yang, Z.; Zhang, T. Rapamycin Activates Mitophagy and Alleviates Cognitive and Synaptic Plasticity Deficits in a Mouse Model of Alzheimer’s Disease. J. Gerontol. A Biol. Sci. Med. Sci. 2021, 76, 1707–1713. [Google Scholar] [CrossRef]
- Ramalingam, M.; Huh, Y.J.; Lee, Y.I. The Impairments of alpha-Synuclein and Mechanistic Target of Rapamycin in Rotenone-Induced SH-SY5Y Cells and Mice Model of Parkinson’s Disease. Front. Neurosci. 2019, 13, 1028. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Zhang, T.; Wang, J.; Zhang, Z.; Zhai, Y.; Yang, G.Y.; Sun, X. Rapamycin attenuates mitochondrial dysfunction via activation of mitophagy in experimental ischemic stroke. Biochem. Biophys. Res. Commun. 2014, 444, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Bove, J.; Martinez-Vicente, M.; Vila, M. Fighting neurodegeneration with rapamycin: Mechanistic insights. Nat. Rev. Neurosci. 2011, 12, 437–452. [Google Scholar] [CrossRef]
- Tsang, C.K.; Qi, H.; Liu, L.F.; Zheng, X.F. Targeting mammalian target of rapamycin (mTOR) for health and diseases. Drug Discov. Today 2007, 12, 112–124. [Google Scholar] [CrossRef]
- Rowinsky, E.K. Targeting the molecular target of rapamycin (mTOR). Curr. Opin. Oncol. 2004, 16, 564–575. [Google Scholar] [CrossRef]
- Shah, S.Z.A.; Zhao, D.; Hussain, T.; Yang, L. Role of the AMPK pathway in promoting autophagic flux via modulating mitochondrial dynamics in neurodegenerative diseases: Insight into prion diseases. Ageing Res. Rev. 2017, 40, 51–63. [Google Scholar] [CrossRef]
- Hardie, D.G. AMPK: A target for drugs and natural products with effects on both diabetes and cancer. Diabetes 2013, 62, 2164–2172. [Google Scholar] [CrossRef] [PubMed]
- Bharath, L.P.; Agrawal, M.; McCambridge, G.; Nicholas, D.A.; Hasturk, H.; Liu, J.; Jiang, K.; Liu, R.; Guo, Z.; Deeney, J.; et al. Metformin Enhances Autophagy and Normalizes Mitochondrial Function to Alleviate Aging-Associated Inflammation. Cell Metab. 2020, 32, 44–55 e46. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.Y.; Moon, S.J.; Kim, E.K.; Seo, H.B.; Yang, E.J.; Son, H.J.; Kim, J.K.; Min, J.K.; Park, S.H.; Cho, M.L. Metformin Suppresses Systemic Autoimmunity in Roquin(san/san) Mice through Inhibiting B Cell Differentiation into Plasma Cells via Regulation of AMPK/mTOR/STAT3. J. Immunol. 2017, 198, 2661–2670. [Google Scholar] [CrossRef]
- Song, Y.M.; Lee, W.K.; Lee, Y.H.; Kang, E.S.; Cha, B.S.; Lee, B.W. Metformin Restores Parkin-Mediated Mitophagy, Suppressed by Cytosolic p53. Int. J. Mol. Sci. 2016, 17, 122. [Google Scholar] [CrossRef] [PubMed]
- Paintlia, A.S.; Mohan, S.; Singh, I. Combinatorial Effect of Metformin and Lovastatin Impedes T-cell Autoimmunity and Neurodegeneration in Experimental Autoimmune Encephalomyelitis. J. Clin. Cell Immunol. 2013, 4, 3. [Google Scholar] [CrossRef] [PubMed]
- Pineda-Ramirez, N.; Alquisiras-Burgos, I.; Ortiz-Plata, A.; Ruiz-Tachiquin, M.E.; Espinoza-Rojo, M.; Aguilera, P. Resveratrol Activates Neuronal Autophagy Through AMPK in the Ischemic Brain. Mol. Neurobiol. 2020, 57, 1055–1069. [Google Scholar] [CrossRef]
- Sebori, R.; Kuno, A.; Hosoda, R.; Hayashi, T.; Horio, Y. Resveratrol Decreases Oxidative Stress by Restoring Mitophagy and Improves the Pathophysiology of Dystrophin-Deficient mdx Mice. Oxid. Med. Cell Longev. 2018, 2018, 9179270. [Google Scholar] [CrossRef] [PubMed]
- Wan, W.; Hua, F.; Fang, P.; Li, C.; Deng, F.; Chen, S.; Ying, J.; Wang, X. Regulation of Mitophagy by Sirtuin Family Proteins: A Vital Role in Aging and Age-Related Diseases. Front. Aging Neurosci. 2022, 14, 845330. [Google Scholar] [CrossRef]
- Chu, X.; Raju, R.P. Regulation of NAD(+) metabolism in aging and disease. Metabolism 2022, 126, 154923. [Google Scholar] [CrossRef]
- Kulkarni, S.S.; Canto, C. Mitochondrial Post-translational Modifications and Metabolic Control: Sirtuins and Beyond. Curr. Diabetes Rev. 2017, 13, 338–351. [Google Scholar] [CrossRef] [PubMed]
- Procaccio, V.; Bris, C.; Chao de la Barca, J.M.; Oca, F.; Chevrollier, A.; Amati-Bonneau, P.; Bonneau, D.; Reynier, P. Perspectives of drug-based neuroprotection targeting mitochondria. Rev. Neurol. 2014, 170, 390–400. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Yuan, J. Targeting RIPK1 kinase for modulating inflammation in human diseases. Front. Immunol. 2023, 14, 1159743. [Google Scholar] [CrossRef]
- Kaur, S.; Sharma, N.; Kumar, V.; Sharma, D.; Devi, B.; Kapil, L.; Singh, C.; Singh, A. The Role of Mitophagy in Various Neurological Diseases as a Therapeutic Approach. Cell Mol. Neurobiol. 2022. [Google Scholar] [CrossRef] [PubMed]
- Slee, E.A.; Zhu, H.; Chow, S.C.; MacFarlane, M.; Nicholson, D.W.; Cohen, G.M. Benzyloxycarbonyl-Val-Ala-Asp (OMe) fluoromethylketone (Z-VAD.FMK) inhibits apoptosis by blocking the processing of CPP32. Biochem. J. 1996, 315 Pt 1, 21–24. [Google Scholar] [CrossRef] [PubMed]


| Disease | SNP | Localization | Cases/Controls/Trios | Subject Cohort | Reference | PubMed Link (Accessed on 27 April 2023) |
|---|---|---|---|---|---|---|
| Primary Adrenal Insufficiency (PAI) | rs12917716 | Intron 19 | 542/1220 | Norway, UK | [60] | https://www.ncbi.nlm.nih.gov/pubmed/18593762 |
| rs12917716 | Intron 19 | 479/2394 | Sweden | [59] | https://pubmed.ncbi.nlm.nih.gov/29849176 | |
| Allergic Rhinitis (AR) | rs7203459 | Intron 22 | 762/760 | Han Chinese | [81] | https://pubmed.ncbi.nlm.nih.gov/32082391 |
| rs2286973 | Exon 19 | 995/1004 | Chinese population | [80] | https://pubmed.ncbi.nlm.nih.gov/36588789 | |
| rs887864 | Intron 19 | |||||
| rs12935657 | Intron 21 | |||||
| rs11645657 | Intron 22 | |||||
| rs36045143 | Intron 22 | |||||
| Alopecia Areata (AA) | rs998592 | Intron 19 | 1702/1723 | European ancestry | [67] | https://www.ncbi.nlm.nih.gov/pubmed/22534877 |
| rs3862469 | Intron 19 | 2332/5233 | European ancestry | [7] | www.ncbi.nlm.nih.gov/pubmed/25608926 | |
| Asthma | rs62026376 | Intron 22 | 6685/14,091 | European ancestry | [103] | www.ncbi.nlm.nih.gov/pubmed/24388013 |
| rs7203459 | Intron 22 | 28,399/128,843 | European ancestry | [79] | www.ncbi.nlm.nih.gov/pubmed/27182965 | |
| rs9923856 | Intron 19 | 1750/9245 | African and European Americans | [78] | www.ncbi.nlm.nih.gov/pubmed/27611488 | |
| rs62026377 | Intron 22 | |||||
| rs17806299 | Intron 19 | 19,954/107,715 | European ancestry | [77] | www.ncbi.nlm.nih.gov/pubmed/29273806 | |
| rs36045143 | Intron 22 | 39,770/76,768 | European ancestry | [76] | www.ncbi.nlm.nih.gov/pubmed/29785011 | |
| rs12935657 | Intron 21 | 40,544/300,671 | European ancestry | [104] | www.ncbi.nlm.nih.gov/pubmed/30929738 | |
| rs35441874 | Intron 19 | |||||
| rs35032408 | Intron 20 | 21,564/318,237 | British ancestry | [75] | www.ncbi.nlm.nih.gov/pubmed/31036433 | |
| rs7203459 | Intron 22 | 5135/25,675 | European ancestry | [74] | www.ncbi.nlm.nih.gov/pubmed/30552067 | |
| rs35441874 | Intron 19 | 46,802/347,481 | European ancestry | [76] | www.ncbi.nlm.nih.gov/pubmed/31619474 | |
| rs35441874 | Intron 19 | 56,167/352,255 | European ancestry | [73] | https://pubmed.ncbi.nlm.nih.gov/34103634 | |
| Autoimmune Thyroid Diseases (ATD) | rs2903692 | Intron 22 | 330621 | Japanese | [42] | https://pubmed.ncbi.nlm.nih.gov/18940880 |
| rs6498169 | Intron 22 | 667/301 | Chinese Han | [82] | https://www.ncbi.nlm.nih.gov/pubmed/24646814 | |
| Celiac Disease | rs12928822 | intergenic | 4,533/10,700 | European populations | [85] | https://pubmed.ncbi.nlm.nih.gov/20190752 |
| Common Variable Immunodeficiency (CVID) | rs17806056 | Intron 19 | 778/10,999 | Sweden, Norway, USA, UK, Germany | [83] | https://www.ncbi.nlm.nih.gov/pubmed/25891430 |
| Crohn’s Disease (CD) | rs2903692 | Intron 22 | 1264/890 | Spain | [65] | https://www.ncbi.nlm.nih.gov/pubmed/19337309 |
| Eosinophilic Esophagitis (EE) | rs12924112 | Intron 21 | 1214/3734 | European ancestry | [84] | https://pubmed.ncbi.nlm.nih.gov/29904099 |
| Juvenile Idiopathic Arthritis (JIA) | rs6498169 | Intron 22 | 1318/2149 | Norway | [68] | https://www.ncbi.nlm.nih.gov/pubmed/19734133 |
| Multiple Sclerosis (MS) | rs6498169 | Intron 22 | 2322/5418/1540 | European ancestry | [46] | www.ncbi.nlm.nih.gov/pubmed/17660530 |
| rs6498169 | Intron 22 | 1146/1309 | Australia | [47] | https://www.ncbi.nlm.nih.gov/pubmed/18650830 | |
| rs11865121 | Intron 19 | 2624/7220 | European ancestry | [49] | www.ncbi.nlm.nih.gov/pubmed/19525953 | |
| rs725613 | Intron 19 | 1498/1706 | Sardinia | [35] | https://www.ncbi.nlm.nih.gov/pubmed/18946483 | |
| rs12708716 | Intron 19 | 5737/10,296/ 2369 | Australia, Belgium, Norway, Sweden, UK, USA | [50] | https://www.ncbi.nlm.nih.gov/pubmed/18987646 | |
| rs6498169 | Intron 19 | 1146/1309 | Europe | [51] | https://www.ncbi.nlm.nih.gov/pubmed/19375175 | |
| rs6498146 | Intron 10 | |||||
| rs741177 | Intron 18 | |||||
| rs876476 | Intron 18 | |||||
| rs11863016 | Intron 22 | |||||
| rs9937607 | Intron 22 | |||||
| rs6498169 | Intron 22 | 211/182 (+521 multiplex controls) | UK | [48] | https://www.ncbi.nlm.nih.gov/pubmed/19506219 | |
| rs6498169 | Intron 22 | 1853/2128 | Holland, Canada | [105] | https://www.ncbi.nlm.nih.gov/pubmed/19834503 | |
| rs6498169 | Intron 22 | 435/550 | Spain | [36] | https://www.ncbi.nlm.nih.gov/pubmed/19221398 | |
| rs12708716 | Intron 19 | 918/656 | African Americans | [52] | https://www.ncbi.nlm.nih.gov/pubmed/19865102 | |
| rs6498169 | Intron 22 | |||||
| rs2080272 | Intron 19 | |||||
| rs7200786 | Intron 19 | 9772/16,849 | European ancestry | [55] | www.ncbi.nlm.nih.gov/pubmed/21833088 | |
| rs2041670 | Intron 19 | 603/825 | Europe | [53] | https://www.ncbi.nlm.nih.gov/pubmed/20849399 | |
| rs998592 | Intron 19 | 197/197 | India | [54] | https://www.ncbi.nlm.nih.gov/pubmed/20952449 | |
| rs12708716 | Intron 19 | |||||
| rs12708716 | Intron 19 | 3102/5047/1113 | Norway, UK | [18] | https://www.ncbi.nlm.nih.gov/pubmed/21179112 | |
| rs7206912 | Intron 22 | |||||
| rs6498169 | Intron 22 | |||||
| rs7184083 | Intron 22 | 1343/1379 | UK, US | [19] | https://www.ncbi.nlm.nih.gov/pubmed/21653641 | |
| rs12927355 | Intron 19 | 14,498/24,091 | European ancestry | [56] | www.ncbi.nlm.nih.gov/pubmed/24076602 | |
| rs4780346 | Intragenic region | |||||
| rs6498168 | Intron 22 | 4888/10,395 | German ancestry | [57] | www.ncbi.nlm.nih.gov/pubmed/27386562 | |
| rs6498160 | Intron 19 | 2273/2148 | Sardinian | [58] | www.ncbi.nlm.nih.gov/pubmed/28445677 | |
| Parkinson’s Disease (PD) | rs7200786 | Intron 19 | 342/503 | European population | [87] | https://pubmed.ncbi.nlm.nih.gov/34149802 |
| rs6498169 | Intron 22 | 515/504 | Han Chinese | [88] | https://pubmed.ncbi.nlm.nih.gov/35432448 | |
| rs12708716 | Intron 19 | |||||
| rs12917716 | Intron 19 | |||||
| rs7200786 | Intron 19 | |||||
| rs2903692 | Intron 19 | |||||
| Primary Biliary Cirrhosis (PBC) | rs12924729 | Intron 19 | 2460/7677 | UK | [72] | www.ncbi.nlm.nih.gov/pubmed/21399635 |
| rs12708715 | Intron 19 | 2861/8514 | British and Irish ancestry | [70] | www.ncbi.nlm.nih.gov/pubmed/22961000 | |
| rs12708715 | Intron 19 | http://www.ncbi.nlm.nih.gov/pubmed/22961000 | ||||
| rs58102322 | Intron 18 | 1450/2967 | Europe | [71] | https://www.ncbi.nlm.nih.gov/pubmed/22257840 | |
| rs12924129 | Intron 19 | |||||
| rs12924729 | Intron 19 | 2764/10,475 | European ancestry | [69] | www.ncbi.nlm.nih.gov/pubmed/26394269 | |
| Rheumatoid Arthritis (RA) | rs6498169 | Intron 22 | 600/550 | Spain | [36] | https://www.ncbi.nlm.nih.gov/pubmed/19221398 |
| rs6498169 | Intron 22 | 1318/2149 | Norway | [68] | https://www.ncbi.nlm.nih.gov/pubmed/19734133 | |
| Selective IgA Deficiency | rs34069391 | Intron 19 | 1635/4852 | European ancestry | [66] | www.ncbi.nlm.nih.gov/pubmed/27723758 |
| Systemic Lupus Erythematosus (SLE) | rs12599402 | Intron 19 | 1656/3394 | Han Chinese | [64] | www.ncbi.nlm.nih.gov/pubmed/23273568 |
| rs7200786 | Intron 19 | 5201/9066 | European ancestry | [63] | www.ncbi.nlm.nih.gov/pubmed/26502338 | |
| rs9652601 | Intron 19 | |||||
| rs9652601 | Intron 19 | 5695/10,352 | Chinese and European ancestry | [62] | www.ncbi.nlm.nih.gov/pubmed/27399966 | |
| rs9652601 | Intron 19 | 6748/11,516 | European ancestry | [61] | www.ncbi.nlm.nih.gov/pubmed/28714469 | |
| rs2041670 | Intron 19 | |||||
| rs8054198 | UTR | |||||
| Systemic Sclerosis | rs1646066 | intergenic | 291/260 | African Americans | [86] | https://pubmed.ncbi.nlm.nih.gov/29293537 |
| Type 1 Diabetes (T1D) | rs12708716 | Intron 19 | 2000/3000 | European ancestry | [34] | www.ncbi.nlm.nih.gov/pubmed/17554260 |
| rs12708716 | Intron 19 | 1963/2938 | European ancestry | [32] | www.ncbi.nlm.nih.gov/pubmed/17554300 | |
| rs725613 | Intron 19 | 1896/1146/873 | European ancestry | [1] | www.ncbi.nlm.nih.gov/pubmed/17632545 | |
| rs2903692 | Intron 22 | |||||
| rs17673553 | Intron 22 | |||||
| rs12708716 | Intron 19 | 3561/4646 | European ancestry | [33] | www.ncbi.nlm.nih.gov/pubmed/18978792 | |
| rs12708716 | Intron 19 | 7514/9045 | European ancestry | [43] | www.ncbi.nlm.nih.gov/pubmed/19430480 | |
| rs2903692 | Intron 22 | 735/621 | Japan | [42] | https://www.ncbi.nlm.nih.gov/pubmed/18940880 | |
| rs725613 | Intron 19 | 1037/1706 | Italy (Sardinia) | [35] | https://www.ncbi.nlm.nih.gov/pubmed/18946483 | |
| rs725613 | Intron 19 | 205/422 | China (Han) | [39] | https://www.ncbi.nlm.nih.gov/pubmed/19178520 | |
| rs6498169 | Intron 22 | 316/550 | Spain | [36] | https://pubmed.ncbi.nlm.nih.gov/19221398 | |
| rs12708716 | Intron 19 | 1212/2513 | Germany | [41] | https://www.ncbi.nlm.nih.gov/pubmed/21873553 | |
| rs2903692 | Intron 22 | 1743/790 | Japan | [40] | https://www.ncbi.nlm.nih.gov/pubmed/22069271 | |
| rs12708716 | Intron 19 | 8506/10,596 | European ancestry | [44] | www.ncbi.nlm.nih.gov/pubmed/21829393 | |
| rs12921922 | Intron 4 | 131/121 | China | [38] | https://www.ncbi.nlm.nih.gov/pubmed/22778732 | |
| rs12931878 | Intron 1 | |||||
| rs12927355 | Intron 19 | 6683/12,173/69 and 2601 affected sibling pair families | European ancestry | [45] | www.ncbi.nlm.nih.gov/pubmed/25751624 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pandey, R.; Bakay, M.; Hakonarson, H. CLEC16A—An Emerging Master Regulator of Autoimmunity and Neurodegeneration. Int. J. Mol. Sci. 2023, 24, 8224. https://doi.org/10.3390/ijms24098224
Pandey R, Bakay M, Hakonarson H. CLEC16A—An Emerging Master Regulator of Autoimmunity and Neurodegeneration. International Journal of Molecular Sciences. 2023; 24(9):8224. https://doi.org/10.3390/ijms24098224
Chicago/Turabian StylePandey, Rahul, Marina Bakay, and Hakon Hakonarson. 2023. "CLEC16A—An Emerging Master Regulator of Autoimmunity and Neurodegeneration" International Journal of Molecular Sciences 24, no. 9: 8224. https://doi.org/10.3390/ijms24098224
APA StylePandey, R., Bakay, M., & Hakonarson, H. (2023). CLEC16A—An Emerging Master Regulator of Autoimmunity and Neurodegeneration. International Journal of Molecular Sciences, 24(9), 8224. https://doi.org/10.3390/ijms24098224





