Classification of COVID-19 Patients into Clinically Relevant Subsets by a Novel Machine Learning Pipeline Using Transcriptomic Features
Abstract
1. Introduction
2. Results
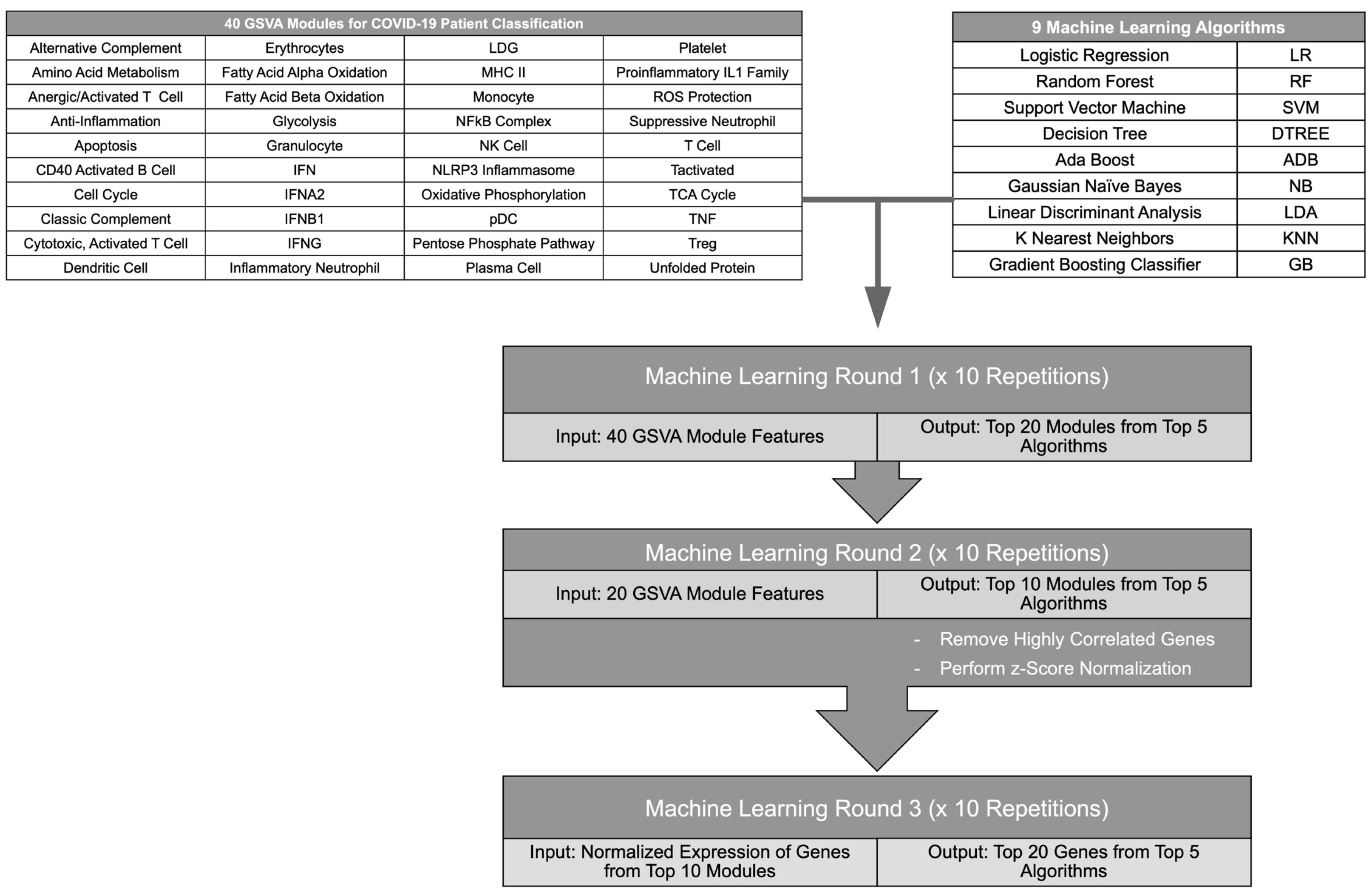
2.1. A Gene Expression-Based Iterative Machine Learning Approach to Classification of COVID-19 Patients
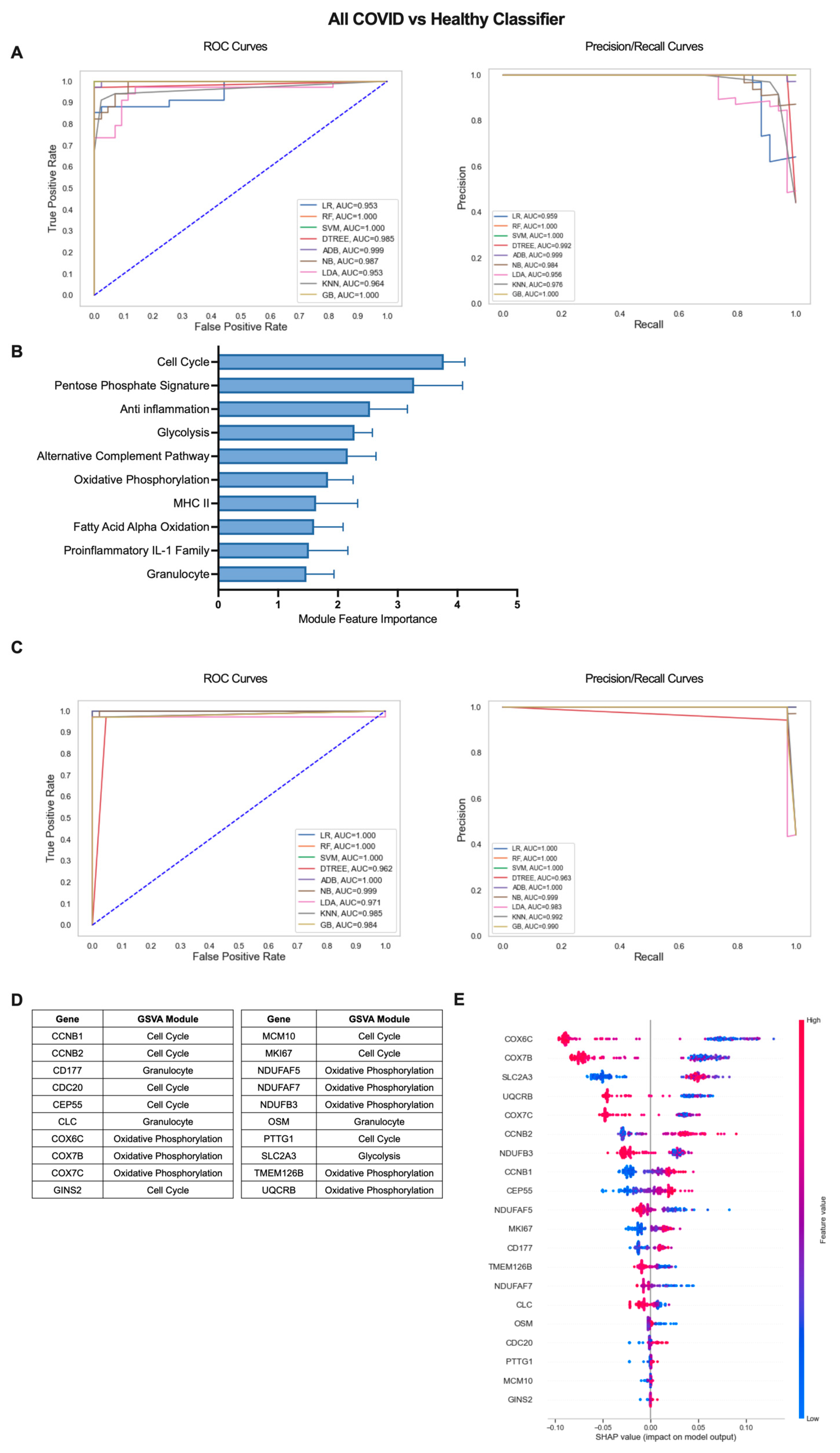
2.2. Machine Learning Models Reveal Genes Critical for Classification of COVID-19 Patients from Healthy Individuals
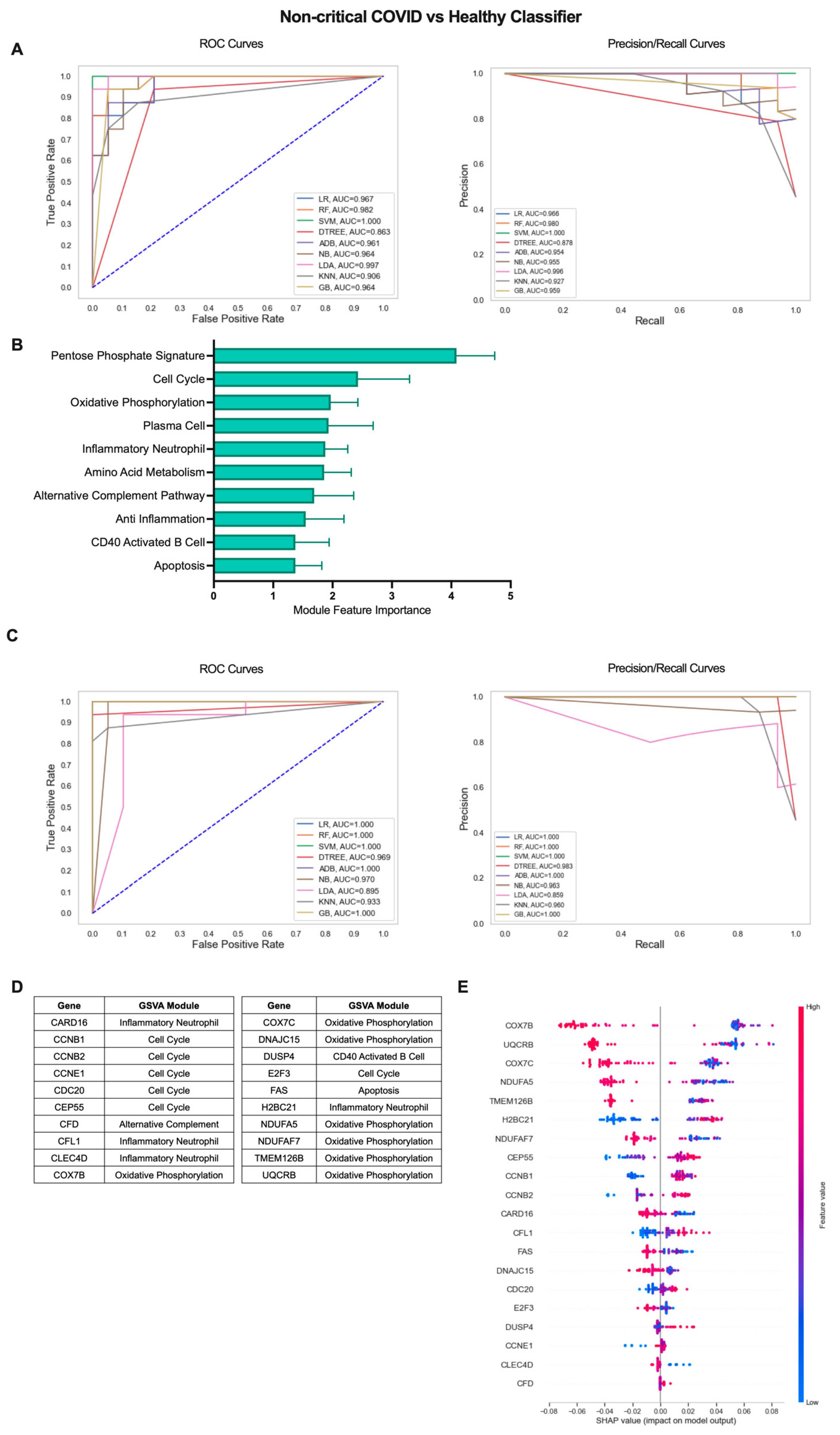
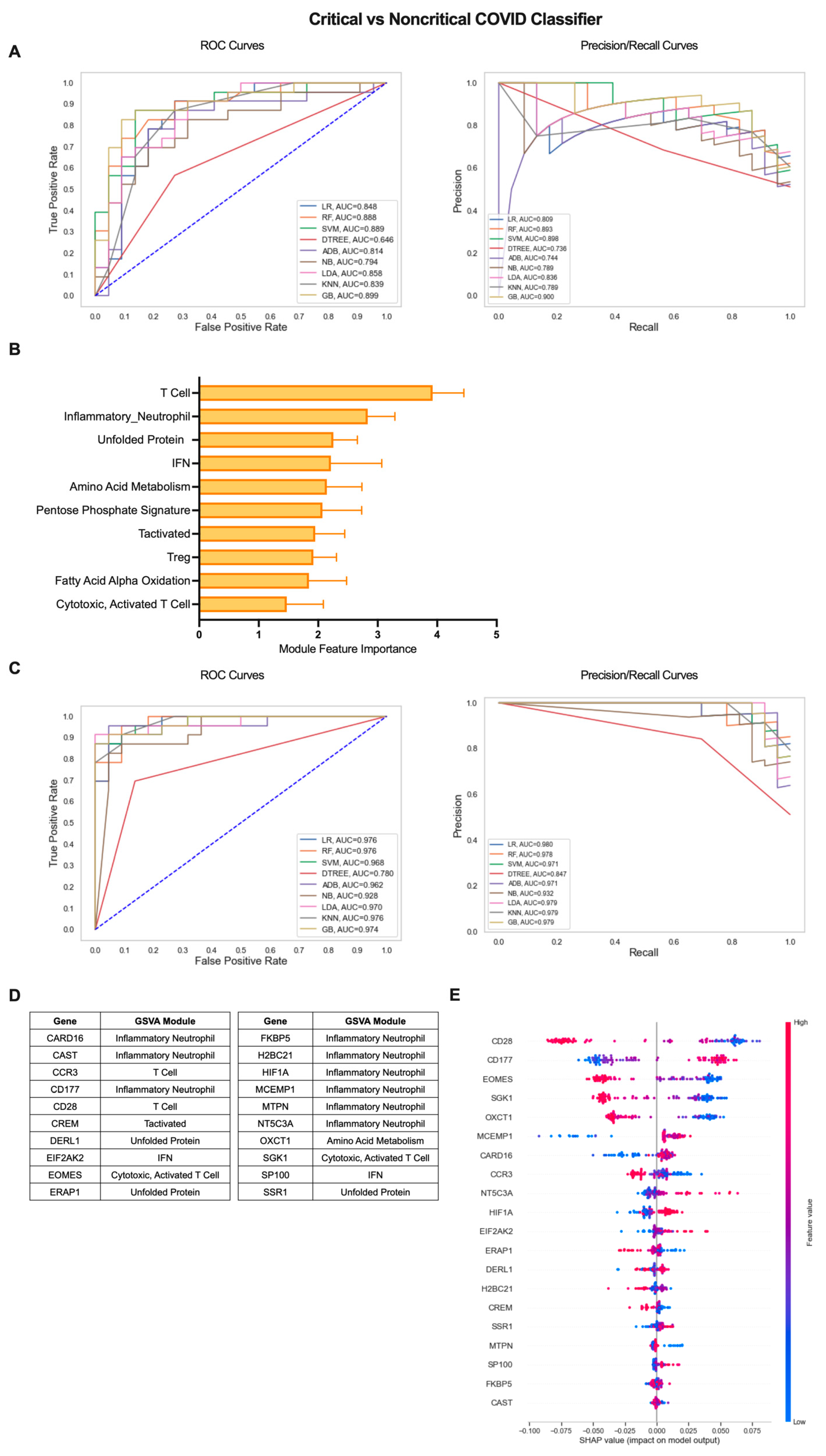
2.3. Iterative Machine Learning Effectively Predicts Disease Severity of COVID-19 Patients Based on Gene Expression
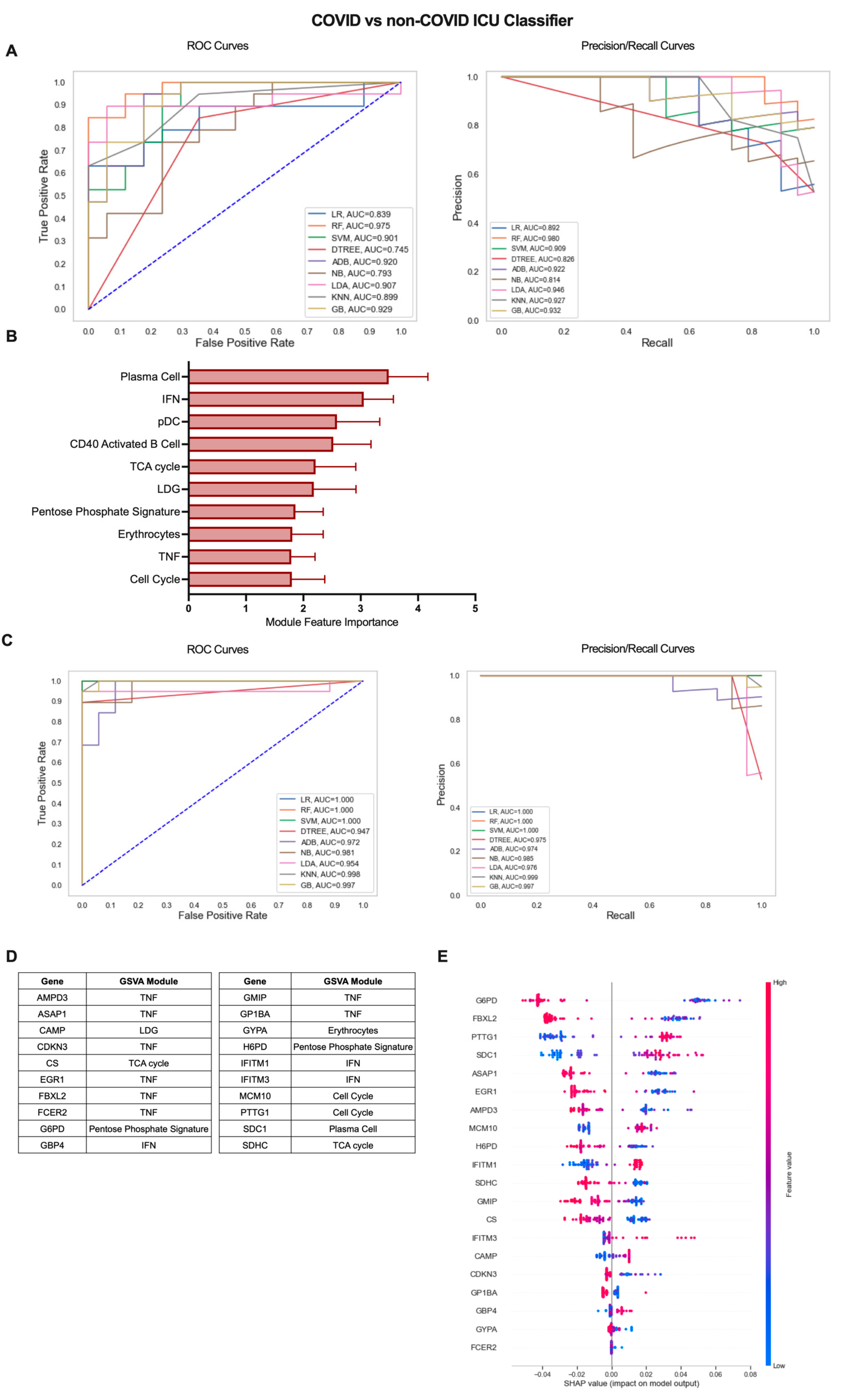
2.4. A Gene Expression-Based Machine Learning Approach Identifies Genes Distinguishing COVID-19 ICU Patients from Other Patients Admitted to the ICU
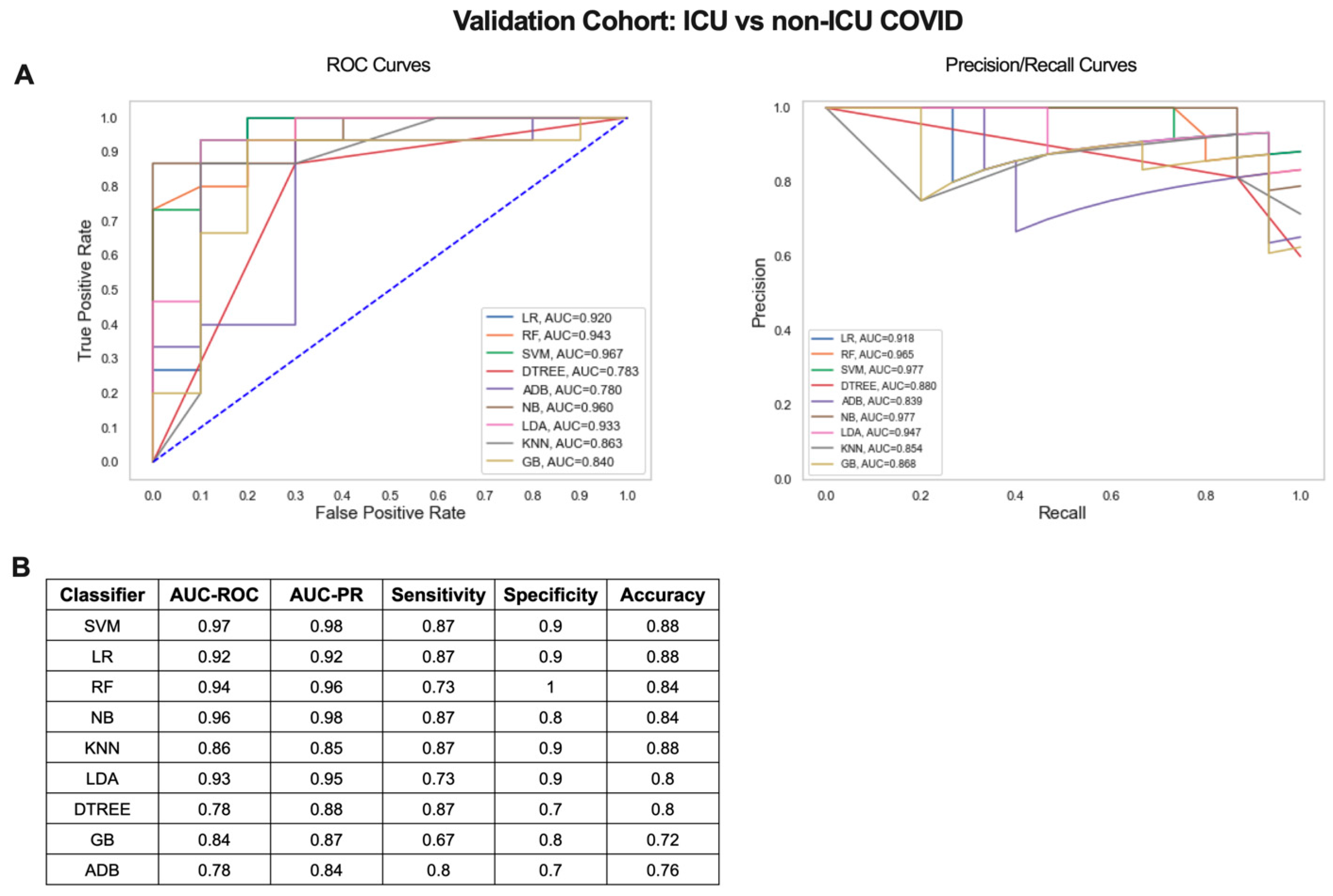
2.5. Validation of the Iterative ML Pipeline in an Independent COVID-19 Patient Dataset
3. Discussion
4. Materials and Methods
4.1. Study Datasets
4.2. Gene Set Variation Analysis (GSVA)
4.3. Iterative Machine Learning (ML) Pipeline
4.4. ML Classification Algorithms
4.5. Feature Importance Calculation
4.6. Gene Feature Correlation Analysis
4.7. z-Score Gene Expression Normalization
4.8. SHAP Value Calculation
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Guan, W.-J.; Ni, Z.-Y.; Hu, Y.; Liang, W.-H.; Ou, C.-Q.; He, J.-X.; Liu, L.; Shan, H.; Lei, C.-L.; Hui, D.S.C.; et al. Clinical Characteristics of Coronavirus Disease 2019 in China. N. Engl. J. Med. 2020, 382, 1708–1720. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical Characteristics of 138 Hospitalized Patients with 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA J. Am. Med. Assoc. 2020, 323, 1061–1069. [Google Scholar] [CrossRef] [PubMed]
- WHO. WHO Coronavirus WHO Coronavirus. Available online: https://covid19.who.int/ (accessed on 1 January 2023).
- Wiersinga, W.J.; Rhodes, A.; Cheng, A.C.; Peacock, S.J.; Prescott, H.C. Pathophysiology, Transmission, Diagnosis, and Treatment of Coronavirus Disease 2019 (COVID-19): A Review. JAMA 2020, 324, 782–793. [Google Scholar] [CrossRef] [PubMed]
- Nalbandian, A.; Sehgal, K.; Gupta, A.; Madhavan, M.V.; McGroder, C.; Stevens, J.S.; Cook, J.R.; Nordvig, A.S.; Shalev, D.; Sehrawat, T.S.; et al. Post-Acute COVID-19 Syndrome. Nat. Med. 2021, 27, 601–615. [Google Scholar] [CrossRef]
- Williamson, E.J.; Walker, A.J.; Bhaskaran, K.; Bacon, S.; Bates, C.; Morton, C.E.; Curtis, H.J.; Mehrkar, A.; Evans, D.; Inglesby, P.; et al. Factors Associated with COVID-19-Related Death Using OpenSAFELY. Nature 2020, 584, 430–436. [Google Scholar] [CrossRef]
- Takahashi, T.; Ellingson, M.K.; Wong, P.; Israelow, B.; Lucas, C.; Klein, J.; Silva, J.; Mao, T.; Oh, J.E.; Tokuyama, M.; et al. Sex Differences in Immune Responses That Underlie COVID-19 Disease Outcomes. Nature 2020, 588, 315–320. [Google Scholar] [CrossRef]
- Mokhtari, T.; Hassani, F.; Ghaffari, N.; Ebrahimi, B.; Yarahmadi, A.; Hassanzadeh, G. COVID-19 and Multiorgan Failure: A Narrative Review on Potential Mechanisms. J. Mol. Histol. 2020, 51, 613–628. [Google Scholar] [CrossRef]
- Michalski, J.E.; Kurche, J.S.; Schwartz, D.A. From ARDS to Pulmonary Fibrosis: The next Phase of the COVID-19 Pandemic? Transl. Res. 2022, 241, 13–24. [Google Scholar] [CrossRef]
- Chen, Y.-T.; Shao, S.-C.; Hsu, C.-K.; Wu, I.-W.; Hung, M.-J.; Chen, Y.-C. Incidence of Acute Kidney Injury in COVID-19 Infection: A Systematic Review and Meta-Analysis. Crit. Care 2020, 24, 346. [Google Scholar] [CrossRef]
- Merad, M.; Blish, C.A.; Sallusto, F.; Iwasaki, A. The Immunology and Immunopathology of COVID-19. Science 2022, 375, 1122–1127. [Google Scholar] [CrossRef]
- Ilieva, M.; Tschaikowski, M.; Vandin, A.; Uchida, S. The Current Status of Gene Expression Profilings in COVID-19 Patients. Clin. Transl. Discov. 2022, 2, e104. [Google Scholar] [CrossRef]
- Daamen, A.R.; Bachali, P.; Owen, K.A.; Kingsmore, K.M.; Hubbard, E.L.; Labonte, A.C.; Robl, R.; Shrotri, S.; Grammer, A.C.; Lipsky, P.E. Comprehensive Transcriptomic Analysis of COVID-19 Blood, Lung, and Airway. Sci. Rep. 2021, 11, 7052. [Google Scholar] [CrossRef]
- Daamen, A.R.; Bachali, P.; Bonham, C.A.; Somerville, L.; Sturek, J.M.; Grammer, A.C.; Kadl, A.; Lipsky, P.E. COVID-19 Patients Exhibit Unique Transcriptional Signatures Indicative of Disease Severity. Front. Immunol. 2022, 13, 989556. [Google Scholar] [CrossRef]
- Mathew, D.; Giles, J.R.; Baxter, A.E.; Oldridge, D.A.; Greenplate, A.R.; Wu, J.E.; Alanio, C.; Kuri-Cervantes, L.; Pampena, M.B.; D’Andrea, K.; et al. Deep Immune Profiling of COVID-19 Patients Reveals Distinct Immunotypes with Therapeutic Implications. Science 2020, 369, eabc8511. [Google Scholar] [CrossRef]
- Lucas, C.; Wong, P.; Klein, J.; Castro, T.B.R.; Silva, J.; Sundaram, M.; Ellingson, M.K.; Mao, T.; Oh, J.E.; Israelow, B.; et al. Longitudinal Analyses Reveal Immunological Misfiring in Severe COVID-19. Nature 2020, 584, 463–469. [Google Scholar] [CrossRef]
- Wilk, A.J.; Rustagi, A.; Zhao, N.Q.; Roque, J.; Martínez-Colón, G.J.; McKechnie, J.L.; Ivison, G.T.; Ranganath, T.; Vergara, R.; Hollis, T.; et al. A Single-Cell Atlas of the Peripheral Immune Response in Patients with Severe COVID-19. Nat. Med. 2020, 26, 1070–1076. [Google Scholar] [CrossRef]
- Wilk, A.J.; Lee, M.J.; Wei, B.; Parks, B.; Pi, R.; Martínez-Colón, G.J.; Ranganath, T.; Zhao, N.Q.; Taylor, S.; Becker, W.; et al. Multi-Omic Profiling Reveals Widespread Dysregulation of Innate Immunity and Hematopoiesis in COVID-19. J. Exp. Med. 2021, 218, e20210582. [Google Scholar] [CrossRef]
- McClain, M.T.; Constantine, F.J.; Henao, R.; Liu, Y.; Tsalik, E.L.; Burke, T.W.; Steinbrink, J.M.; Petzold, E.; Nicholson, B.P.; Rolfe, R.; et al. Dysregulated Transcriptional Responses to SARS-CoV-2 in the Periphery. Nat. Commun. 2021, 12, 1079. [Google Scholar] [CrossRef]
- Stephenson, E.; Reynolds, G.; Botting, R.A.; Calero-Nieto, F.J.; Morgan, M.D.; Tuong, Z.K.; Bach, K.; Sungnak, W.; Worlock, K.B.; Yoshida, M.; et al. Single-Cell Multi-Omics Analysis of the Immune Response in COVID-19. Nat. Med. 2021, 27, 904–916. [Google Scholar] [CrossRef]
- Overmyer, K.A.; Shishkova, E.; Miller, I.J.; Balnis, J.; Bernstein, M.N.; Peters-Clarke, T.M.; Meyer, J.G.; Quan, Q.; Muehlbauer, L.K.; Trujillo, E.A.; et al. Large-Scale Multi-Omic Analysis of COVID-19 Severity. Cell Syst. 2021, 12, 23–40.e7. [Google Scholar] [CrossRef]
- Carapito, R.; Li, R.; Helms, J.; Carapito, C.; Gujja, S.; Rolli, V.; Guimaraes, R.; Malagon-Lopez, J.; Spinnhirny, P.; Lederle, A.; et al. Identification of Driver Genes for Critical Forms of COVID-19 in a Deeply Phenotyped Young Patient Cohort. Sci. Transl. Med. 2022, 14, eabj7521. [Google Scholar] [CrossRef]
- Shah, P.; Kendall, F.; Khozin, S.; Goosen, R.; Hu, J.; Laramie, J.; Ringel, M.; Schork, N. Artificial Intelligence and Machine Learning in Clinical Development: A Translational Perspective. NPJ Digit. Med. 2019, 2, 69. [Google Scholar] [CrossRef]
- Meraihi, Y.; Gabis, A.B.; Mirjalili, S.; Ramdane-Cherif, A.; Alsaadi, F.E. Machine Learning-Based Research for COVID-19 Detection, Diagnosis, and Prediction: A Survey. SN Comput. Sci. 2022, 3, 286. [Google Scholar] [CrossRef] [PubMed]
- Quiroz-Juárez, M.A.; Torres-Gómez, A.; Hoyo-Ulloa, I.; León-Montiel, R.d.J.; U’Ren, A.B. Identification of High-Risk COVID-19 Patients Using Machine Learning. PLoS ONE 2021, 16, e0257234. [Google Scholar] [CrossRef] [PubMed]
- Guhan, B.; Almutairi, L.; Sowmiya, S.; Snekhalatha, U.; Rajalakshmi, T.; Aslam, S.M. Automated System for Classification of COVID-19 Infection from Lung CT Images Based on Machine Learning and Deep Learning Techniques. Sci. Rep. 2022, 12, 17417. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, D.; Kay, F.; Tan, J.; Yan, Y.; Ng, Y.S.; Iyengar, P.; Peshock, R.; Jiang, S. Deep Learning–Based COVID-19 Pneumonia Classification Using Chest CT Images: Model Generalizability. Front. Artif. Intell. 2021, 4, 694875. [Google Scholar] [CrossRef]
- Zargari Khuzani, A.; Heidari, M.; Shariati, S.A. COVID-Classifier: An Automated Machine Learning Model to Assist in the Diagnosis of COVID-19 Infection in Chest X-Ray Images. Sci. Rep. 2021, 11, 9887. [Google Scholar] [CrossRef]
- Penrice-Randal, R.; Dong, X.; Shapanis, A.G.; Gardner, A.; Harding, N.; Legebeke, J.; Lord, J.; Vallejo, A.F.; Poole, S.; Brendish, N.J.; et al. Blood Gene Expression Predicts Intensive Care Unit Admission in Hospitalised Patients with COVID-19. Front. Immunol. 2022, 13, 988685. [Google Scholar] [CrossRef]
- Zarei Ghobadi, M.; Emamzadeh, R.; Teymoori-Rad, M.; Afsaneh, E. Exploration of Blood-Derived Coding and Non-Coding RNA Diagnostic Immunological Panels for COVID-19 through a Co-Expressed-Based Machine Learning Procedure. Front. Immunol. 2022, 13, 1001070. [Google Scholar] [CrossRef]
- Song, X.; Zhu, J.; Tan, X.; Yu, W.; Wang, Q.; Shen, D.; Chen, W. XGBoost-Based Feature Learning Method for Mining COVID-19 Novel Diagnostic Markers. Front. Public Health 2022, 10, 926069. [Google Scholar] [CrossRef]
- Li, H.; Huang, F.; Liao, H.; Li, Z.; Feng, K.; Huang, T.; Cai, Y.-D. Identification of COVID-19-Specific Immune Markers Using a Machine Learning Method. Front. Mol. Biosci. 2022, 9, 952626. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhou, X.; Ding, S.; Chen, L.; Feng, K.; Li, H.; Huang, T.; Cai, Y.-D. Identification of Transcriptome Biomarkers for Severe COVID-19 with Machine Learning Methods. Biomolecules 2022, 12, 1735. [Google Scholar] [CrossRef] [PubMed]
- Lohmann, S.; Herold, A.; Bergauer, T.; Belousov, A.; Betzl, G.; Demario, M.; Dietrich, M.; Luistro, L.; Poignée-Heger, M.; Schostack, K.; et al. Gene Expression Analysis in Biomarker Research and Early Drug Development Using Function Tested Reverse Transcription Quantitative Real-Time PCR Assays. Methods 2013, 59, 10–19. [Google Scholar] [CrossRef] [PubMed]
- El-Deiry, W.S.; Goldberg, R.M.; Lenz, H.-J.; Shields, A.F.; Gibney, G.T.; Tan, A.R.; Brown, J.; Eisenberg, B.; Heath, E.I.; Phuphanich, S.; et al. The Current State of Molecular Testing in the Treatment of Patients with Solid Tumors, 2019. CA. Cancer J. Clin. 2019, 69, 305–343. [Google Scholar] [CrossRef]
- Marshall, K.W.; Mohr, S.; Khettabi, F.E.; Nossova, N.; Chao, S.; Bao, W.; Ma, J.; Li, X.-J.; Liew, C.-C. A Blood-Based Biomarker Panel for Stratifying Current Risk for Colorectal Cancer. Int. J. Cancer 2010, 126, 1177–1186. [Google Scholar] [CrossRef] [PubMed]
- Uddin, S.; Khan, A.; Hossain, M.E.; Moni, M.A. Comparing Different Supervised Machine Learning Algorithms for Disease Prediction. BMC Med. Inform. Decis. Mak. 2019, 19, 281. [Google Scholar] [CrossRef]
- Mazlan, A.U.; Sahabudin, N.A.; Remli, M.A.; Ismail, N.S.; Mohamad, M.S.; Nies, H.W.; Abd Warif, N.B. A Review on Recent Progress in Machine Learning and Deep Learning Methods for Cancer Classification on Gene Expression Data. Processes 2021, 9, 1466. [Google Scholar] [CrossRef]
- Kingsmore, K.M.; Puglisi, C.E.; Grammer, A.C.; Lipsky, P.E. An Introduction to Machine Learning and Analysis of Its Use in Rheumatic Diseases. Nat. Rev. Rheumatol. 2021, 17, 710–730. [Google Scholar] [CrossRef]
- Saeys, Y.; Inza, I.; Larranaga, P. A Review of Feature Selection Techniques in Bioinformatics. Bioinformatics 2007, 23, 2507–2517. [Google Scholar] [CrossRef]
- Ang, J.C.; Mirzal, A.; Haron, H.; Hamed, H.N.A. Supervised, Unsupervised, and Semi-Supervised Feature Selection: A Review on Gene Selection. IEEE/ACM Trans. Comput. Biol. Bioinforma. 2016, 13, 971–989. [Google Scholar] [CrossRef]
- Samy, A.; Maher, M.A.; Abdelsalam, N.A.; Badr, E. SARS-CoV-2 Potential Drugs, Drug Targets, and Biomarkers: A Viral-Host Interaction Network-Based Analysis. Sci. Rep. 2022, 12, 11934. [Google Scholar] [CrossRef]
- Cavalcante, L.T.D.F.; da Fonseca, G.C.; Amado Leon, L.A.; Salvio, A.L.; Brustolini, O.J.; Gerber, A.L.; Guimarães, A.P.D.C.; Marques, C.A.B.; Fernandes, R.A.; Ramos Filho, C.H.F.; et al. Buffy Coat Transcriptomic Analysis Reveals Alterations in Host Cell Protein Synthesis and Cell Cycle in Severe COVID-19 Patients. Int. J. Mol. Sci. 2022, 23, 13588. [Google Scholar] [CrossRef] [PubMed]
- Prado, C.A.D.S.; Fonseca, D.L.M.; Singh, Y.; Filgueiras, I.S.; Baiocchi, G.C.; Plaça, D.R.; Marques, A.H.; Dantas-Komatsu, R.C.S.; Usuda, J.N.; Freire, P.P.; et al. Integrative Systems Immunology Uncovers Molecular Networks of the Cell Cycle That Stratify COVID-19 Severity. J. Med. Virol. 2023, 95, e28450. [Google Scholar] [CrossRef] [PubMed]
- Duan, C.; Ma, R.; Zeng, X.; Chen, B.; Hou, D.; Liu, R.; Li, X.; Liu, L.; Li, T.; Huang, H. SARS-CoV-2 Achieves Immune Escape by Destroying Mitochondrial Quality: Comprehensive Analysis of the Cellular Landscapes of Lung and Blood Specimens From Patients With COVID-19. Front. Immunol. 2022, 13, 946731. [Google Scholar] [CrossRef] [PubMed]
- Chernyak, B.V.; Popova, E.N.; Prikhodko, A.S.; Grebenchikov, O.A.; Zinovkina, L.A.; Zinovkin, R.A. COVID-19 and Oxidative Stress. Biochemistry 2020, 85, 1543–1553. [Google Scholar] [CrossRef] [PubMed]
- Guarnieri, J.W.; Dybas, J.M.; Fazelinia, H.; Kim, M.S.; Frere, J.; Zhang, Y.; Albrecht, Y.S.; Murdock, D.G.; Angelin, A.; Singh, L.N.; et al. Targeted Down Regulation of Core Mitochondrial Genes During SARS-CoV-2 Infection. bioRxiv 2022. preprint. [Google Scholar] [CrossRef]
- McKenna, E.; Wubben, R.; Isaza-Correa, J.M.; Melo, A.M.; Mhaonaigh, A.U.; Conlon, N.; O’Donnell, J.S.; Ní Cheallaigh, C.; Hurley, T.; Stevenson, N.J.; et al. Neutrophils in COVID-19: Not Innocent Bystanders. Front. Immunol. 2022, 13, 2548. [Google Scholar] [CrossRef]
- Aschenbrenner, A.C.; Mouktaroudi, M.; Kraemer, B.; Antonakos, N.; Oestreich, M.; Gkizeli, K.; Nuesch-Germano, M.; Saridaki, M.; Bonaguro, L.; Reusch, N.; et al. Disease Severity-Specific Neutrophil Signatures in Blood Transcriptomes Stratify COVID-19 Patients. medRxiv 2020, 13, 1–25. [Google Scholar] [CrossRef]
- Schulte-Schrepping, J.; Reusch, N.; Paclik, D.; Baßler, K.; Schlickeiser, S.; Zhang, B.; Krämer, B.; Krammer, T.; Brumhard, S.; Bonaguro, L.; et al. Severe COVID-19 Is Marked by a Dysregulated Myeloid Cell Compartment. Cell 2020, 182, 1419–1440.e23. [Google Scholar] [CrossRef]
- Lee, J.S.; Park, S.; Jeong, H.W.; Ahn, J.Y.; Choi, S.J.; Lee, H.; Choi, B.; Nam, S.K.; Sa, M.; Kwon, J.S.; et al. Immunophenotyping of Covid-19 and Influenza Highlights the Role of Type i Interferons in Development of Severe Covid-19. Sci. Immunol. 2020, 5, eabd1554. [Google Scholar] [CrossRef]
- Dong, Z.; Yan, Q.; Cao, W.; Liu, Z.; Wang, X. Identification of Key Molecules in COVID-19 Patients Significantly Correlated with Clinical Outcomes by Analyzing Transcriptomic Data. Front. Immunol. 2022, 13, 930866. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.; To, K.K.-W.; Wong, Y.-C.; Liu, L.; Zhou, B.; Li, X.; Huang, H.; Mo, Y.; Luk, T.-Y.; Lau, T.T.-K.; et al. Acute SARS-CoV-2 Infection Impairs Dendritic Cell and T Cell Responses. Immunity 2020, 53, 864–877.e5. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; John Wherry, E. T Cell Responses in Patients with COVID-19. Nat. Rev. Immunol. 2020, 20, 529–536. [Google Scholar] [CrossRef] [PubMed]
- Cross, A.R.; de Andrea, C.E.; Villalba-Esparza, M.; Landecho, M.F.; Cerundolo, L.; Weeratunga, P.; Etherington, R.E.; Denney, L.; Ogg, G.; Ho, L.-P.; et al. Spatial Transcriptomic Characterization of COVID-19 Pneumonitis Identifies Immune Circuits Related to Tissue Injury. JCI Insight 2022, 8, e157837. [Google Scholar] [CrossRef]
- Izadi, Z.; Brenner, E.J.; Mahil, S.K.; Dand, N.; Yiu, Z.Z.N.; Yates, M.; Ungaro, R.C.; Zhang, X.; Agrawal, M.; Colombel, J.-F.; et al. Association Between Tumor Necrosis Factor Inhibitors and the Risk of Hospitalization or Death Among Patients with Immune-Mediated Inflammatory Disease and COVID-19. JAMA Netw. Open 2021, 4, e2129639. [Google Scholar] [CrossRef]
- Hänzelmann, S.; Castelo, R.; Guinney, J. GSVA: Gene Set Variation Analysis for Microarray and RNA-Seq Data. BMC Bioinform. 2013, 14, 7. [Google Scholar] [CrossRef] [PubMed]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-Learn: Machine Learning in {P}ython. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Blagus, R.; Lusa, L. SMOTE for High-Dimensional Class-Imbalanced Data. BMC Bioinform. 2013, 14, 106. [Google Scholar] [CrossRef]
- Hunter, J.D. Matplotlib: A 2D Graphics Environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Lundberg, S.M.; Lee, S.-I. A Unified Approach to Interpreting Model Predictions. In Proceedings of the Advances in Neural Information Processing Systems; Guyon, I., Luxburg, U., Von Bengio, S., Wallach, H., Fergus, R., Vishwanathan, S., Garnett, R., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2017; Volume 30. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Daamen, A.R.; Bachali, P.; Grammer, A.C.; Lipsky, P.E. Classification of COVID-19 Patients into Clinically Relevant Subsets by a Novel Machine Learning Pipeline Using Transcriptomic Features. Int. J. Mol. Sci. 2023, 24, 4905. https://doi.org/10.3390/ijms24054905
Daamen AR, Bachali P, Grammer AC, Lipsky PE. Classification of COVID-19 Patients into Clinically Relevant Subsets by a Novel Machine Learning Pipeline Using Transcriptomic Features. International Journal of Molecular Sciences. 2023; 24(5):4905. https://doi.org/10.3390/ijms24054905
Chicago/Turabian StyleDaamen, Andrea R., Prathyusha Bachali, Amrie C. Grammer, and Peter E. Lipsky. 2023. "Classification of COVID-19 Patients into Clinically Relevant Subsets by a Novel Machine Learning Pipeline Using Transcriptomic Features" International Journal of Molecular Sciences 24, no. 5: 4905. https://doi.org/10.3390/ijms24054905
APA StyleDaamen, A. R., Bachali, P., Grammer, A. C., & Lipsky, P. E. (2023). Classification of COVID-19 Patients into Clinically Relevant Subsets by a Novel Machine Learning Pipeline Using Transcriptomic Features. International Journal of Molecular Sciences, 24(5), 4905. https://doi.org/10.3390/ijms24054905





