Accurate Detection of SARS-CoV-2 by Next-Generation Sequencing in Low Viral Load Specimens
Abstract
1. Introduction
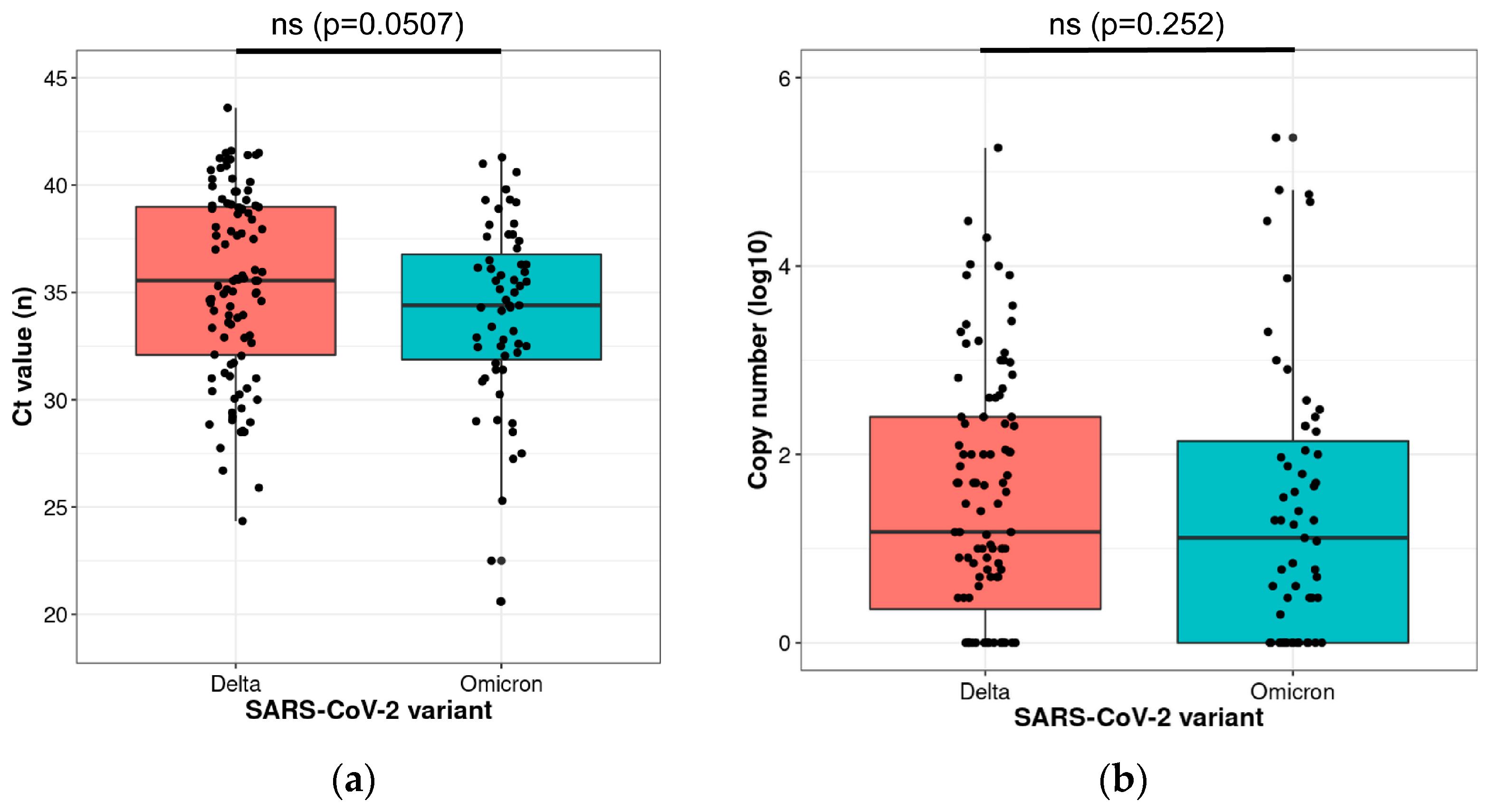
2. Results
3. Discussion
4. Materials and Methods
4.1. Patients and Samples
4.2. SARS-CoV-2 Detection and Genome Sequencing
4.3. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Munnink, B.B.O.; Nieuwenhuijse, D.F.; Stein, M.; O’Toole, A.; Haverkate, M.; Mollers, M.; Kamga, S.K.; Schapendonk, C.; Pronk, M.; Lexmond, P.; et al. Rapid SARS-CoV-2 whole-genome sequencing and analysis for informed public health decision-making in the Netherlands. Nat. Med. 2020, 26, 1405–1410. [Google Scholar] [CrossRef]
- Gaymard, A.; Bosetti, P.; Feri, A.; Destras, G.; Enouf, V.; Andronico, A.; Burrel, S.; Behillil, S.; Sauvage, C.; Bal, A.; et al. Early assessment of diffusion and possible expansion of SARS-CoV-2 Lineage 20I/501Y.V1 (B.1.1.7, variant of concern 202012/01) in France, January to March 2021. Eurosurveillance 2021, 26, 2100133. [Google Scholar] [CrossRef]
- Sabino, E.C.; Buss, L.F.; Carvalho, M.P.S.; Prete, C.A., Jr.; Crispim, M.A.E.; Fraiji, N.A.; Pereira, R.H.M.; Parag, K.V.; da Silva Peixoto, P.; Kraemer, M.U.G.; et al. Resurgence of COVID-19 in Manaus, Brazil, despite high seroprevalence. Lancet 2021, 397, 452–455. [Google Scholar] [CrossRef]
- Thompson, C.N.; Hughes, S.; Ngai, S.; Baumgartner, J.; Wang, J.C.; McGibbon, E.; Devinney, K.; Luoma, E.; Bertolino, D.; Hwang, C.; et al. Rapid Emergence and Epidemiologic Characteristics of the SARS-CoV-2 B.1.526 Variant—New York City, New York, January 1-April 5, 2021. MMWR Morb. Mortal. Wkly. Rep. 2021, 70, 712–716. [Google Scholar] [CrossRef]
- Zhang, W.; Davis, B.D.; Chen, S.S.; Martinez, J.M.S.; Plummer, J.T.; Vail, E. Emergence of a Novel SARS-CoV-2 Variant in Southern California. JAMA 2021, 325, 1324–1326. [Google Scholar] [CrossRef]
- Callaway, E. Why does the Omicron sub-variant spread faster than the original? Nature 2022, 602, 556–557. [Google Scholar] [CrossRef]
- Sentis, C.; Billaud, G.; Bal, A.; Frobert, E.; Bouscambert, M.; Destras, G.; Josset, L.; Lina, B.; Morfin, F.; Gaymard, A.; et al. SARS-CoV-2 Omicron Variant, Lineage BA.1, Is Associated with Lower Viral Load in Nasopharyngeal Samples Compared to Delta Variant. Viruses 2022, 14, 919. [Google Scholar] [CrossRef]
- Chen, Z.; Azman, A.S.; Chen, X.; Zou, J.; Tian, Y.; Sun, R.; Xu, X.; Wu, Y.; Lu, W.; Ge, S.; et al. Global landscape of SARS-CoV-2 genomic surveillance and data sharing. Nat. Genet. 2022, 54, 499–507. [Google Scholar] [CrossRef]
- Khare, S.; Gurry, C.; Freitas, L.; Schultz, M.B.; Bach, G.; Diallo, A.; Akite, N.; Ho, J.; Lee, R.T.; Yeo, W.; et al. GISAID’s Role in Pandemic Response. China CDC Wkly. 2021, 3, 1049–1051. [Google Scholar] [CrossRef]
- Puhach, O.; Adea, K.; Hulo, N.; Sattonnet, P.; Genecand, C.; Iten, A.; Jacquerioz, F.; Kaiser, L.; Vetter, P.; Eckerle, I.; et al. Infectious viral load in unvaccinated and vaccinated individuals infected with ancestral, Delta or Omicron SARS-CoV-2. Nat. Med. 2022, 28, 1491–1500. [Google Scholar] [CrossRef]
- Jain, A.; Rophina, M.; Mahajan, S.; Krishnan, B.B.; Sharma, M.; Mandal, S.; Fernandez, T.; Sultanji, S.; Jolly, B.; Mathew, S.; et al. Analysis of the potential impact of genomic variants in global SARS-CoV-2 genomes on molecular diagnostic assays. Int. J. Infect. Dis. 2021, 102, 460–462. [Google Scholar] [CrossRef]
- Frampton, D.; Rampling, T.; Cross, A.; Bailey, H.; Heaney, J.; Byott, M.; Scott, R.; Sconza, R.; Price, J.; Margaritis, M.; et al. Genomic characteristics and clinical effect of the emergent SARS-CoV-2 B.1.1.7 lineage in London, UK: A whole-genome sequencing and hospital-based cohort study. Lancet Infect. Dis. 2021, 21, 1246–1256. [Google Scholar]
- Mascola, J.R.; Graham, B.S.; Fauci, A.S. SARS-CoV-2 Viral Variants-Tackling a Moving Target. JAMA 2021, 325, 1261–1262. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control; World Health Organization Regional Office for Europe. Methods for the Detection and Identification of SARS-CoV-2 Variants: Second Update, August; ECDC and WHO European Region: Stockholm, Sweden; Copenhagen, Denmark, 2022.
- Fontanet, A.; Autran, B.; Lina, B.; Kieny, M.P.; Karim, S.S.A.; Sridhar, D. SARS-CoV-2 variants and ending the COVID-19 pandemic. Lancet 2021, 397, 952–954. [Google Scholar] [CrossRef]
- Khiabani, K.; Amirzade-Iranaq, M.H. Are saliva and deep throat sputum as reliable as common respiratory specimens for SARS-CoV-2 detection? A systematic review and meta-analysis. Am. J. Infect. Control 2021, 49, 1165–1176. [Google Scholar] [CrossRef]
- Lambrou, A.S.; Shirk, P.; Steele, M.K.; Paul, P.; Paden, C.R.; Cadwell, B.; Reese, H.E.; Aoki, Y.; Hassell, N.; Zheng, X.Y.; et al. Genomic Surveillance for SARS-CoV-2 Variants: Predominance of the Delta (B.1.617.2) and Omicron (B.1.1.529) Variants—United States, June 2021–January 2022. MMWR Morb. Mortal. Wkly. Rep. 2022, 71, 206–211. [Google Scholar] [CrossRef]
- Saxena, S.K.; Kumar, S.; Ansari, S.; Paweska, J.T.; Maurya, V.K.; Tripathi, A.K.; Abdel-Moneim, A.S. Characterization of the novel SARS-CoV-2 Omicron (B.1.1.529) variant of concern and its global perspective. J. Med. Virol. 2022, 94, 1738–1744. [Google Scholar] [CrossRef]
- Jung, C.; Kmiec, D.; Koepke, L.; Zech, F.; Jacob, T.; Sparrer, K.M.J.; Kirchhoff, F. Omicron: What Makes the Latest SARS-CoV-2 Variant of Concern So Concerning? J. Virol. 2022, 96, e0207721. [Google Scholar] [CrossRef]
- Laitman, A.M.; Lieberman, J.A.; Hoffman, N.G.; Roychoudhury, P.; Mathias, P.C.; Greninger, A.L. The SARS-CoV-2 Omicron Variant Does Not Have Higher Nasal Viral Loads Compared to the Delta Variant in Symptomatic and Asymptomatic Individuals. J. Clin. Microbiol. 2022, 60, e0013922. [Google Scholar] [CrossRef]
- Qiu, J. Covert coronavirus infections could be seeding new outbreaks. Nature 2020. [Google Scholar] [CrossRef]
- Braunstein, G.D.; Schwartz, L.; Hymel, P.; Fielding, J. False Positive Results With SARS-CoV-2 RT-PCR Tests and How to Evaluate a RT-PCR-Positive Test for the Possibility of a False Positive Result. J. Occup. Environ. Med. 2021, 63, e159–e162. [Google Scholar] [CrossRef]
- Pecoraro, V.; Negro, A.; Pirotti, T.; Trenti, T. Estimate false-negative RT-PCR rates for SARS-CoV-2. A systematic review and meta-analysis. Eur. J. Clin. Investig. 2022, 52, e13706. [Google Scholar] [CrossRef]
- Kucirka, L.M.; Lauer, S.A.; Laeyendecker, O.; Boon, D.; Lessler, J. Variation in False-Negative Rate of Reverse Transcriptase Polymerase Chain Reaction-Based SARS-CoV-2 Tests by Time Since Exposure. Ann. Intern. Med. 2020, 173, 262–267. [Google Scholar] [CrossRef]
- Wikramaratna, P.S.; Paton, R.S.; Ghafari, M.; Lourenco, J. Estimating the false-negative test probability of SARS-CoV-2 by RT-PCR. Eurosurveillance 2020, 25, 2000568. [Google Scholar] [CrossRef]
- Penarrubia, L.; Ruiz, M.; Porco, R.; Rao, S.N.; Juanola-Falgarona, M.; Manissero, D.; Lopez-Fontanals, M.; Pareja, J. Multiple assays in a real-time RT-PCR SARS-CoV-2 panel can mitigate the risk of loss of sensitivity by new genomic variants during the COVID-19 outbreak. Int. J. Infect. Dis. 2020, 97, 225–229. [Google Scholar] [CrossRef]
- Bal, A.; Destras, G.; Gaymard, A.; Stefic, K.; Marlet, J.; Eymieux, S.; Regue, H.; Semanas, Q.; d’Aubarede, C.; Billaud, G.; et al. Two-step strategy for the identification of SARS-CoV-2 variant of concern 202012/01 and other variants with spike deletion H69-V70, France, August to December 2020. Eurosurveillance 2021, 26, 2100008. [Google Scholar] [CrossRef]
- Mulder, M.; van der Vegt, D.; Oude Munnink, B.B.; GeurtsvanKessel, C.H.; van de Bovenkamp, J.; Sikkema, R.S.; Jacobs, E.M.G.; Koopmans, M.P.G.; Wegdam-Blans, M.C.A. Reinfection of SARS-CoV-2 in an immunocompromised patient: A case report. Clin. Infect. Dis. 2020, 73, e2841–e2842. [Google Scholar] [CrossRef]
- Rambaut, A.; Holmes, E.C.; O’Toole, A.; Hill, V.; McCrone, J.T.; Ruis, C.; du Plessis, L.; Pybus, O.G. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 2020, 5, 1403–1407. [Google Scholar] [CrossRef]
- Lam, C.; Gray, K.; Gall, M.; Sadsad, R.; Arnott, A.; Johnson-Mackinnon, J.; Fong, W.; Basile, K.; Kok, J.; Dwyer, D.E.; et al. SARS-CoV-2 Genome Sequencing Methods Differ in Their Abilities To Detect Variants from Low-Viral-Load Samples. J. Clin. Microbiol. 2021, 59, e0104621. [Google Scholar] [CrossRef]
- Sun, Y.; Ding, C.; Chen, Q.; Xie, J.; Yu, J.; Shi, Y.; Jiang, C.; Zhang, Z.; He, H.; Ge, Y.; et al. Digital PCR assay for the effective detection of COVID-19 patients with SARS-CoV-2 low viral load. J. Virol. Methods 2021, 295, 114185. [Google Scholar] [CrossRef]
- Lu, R.; Wang, J.; Li, M.; He, J.; Wang, Y.; Dong, J.; Cai, W. Retrospective quantitative detection of SARS-CoV-2 by digital PCR showing high accuracy for low viral load specimens. J. Infect. Dev. Ctries. 2022, 16, 10–15. [Google Scholar] [CrossRef]
- Fomenko, A.; Weibel, S.; Moezi, H.; Menger, K.; Schmucker, C.; Metzendorf, M.I.; Motschall, E.; Falcone, V.; Huzly, D.; Panning, M.; et al. Assessing severe acute respiratory syndrome coronavirus 2 infectivity by reverse-transcription polymerase chain reaction: A systematic review.w and meta-analysis. Rev. Med. Virol. 2022, 32, e2342. [Google Scholar] [CrossRef]
- Tegally, H.; San, J.E.; Cotten, M.; Tegomoh, B.; Mboowa, G.; Martin, D.P.; Baxter, C.; Moir, M.; Lambisia, A.; Diallo, A.; et al. The evolving SARS-CoV-2 epidemic in Africa: Insights from rapidly expanding genomic surveillance. medRxiv 2022, 378, eabq5358. [Google Scholar] [CrossRef]
- Hourdel, V.; Kwasiborski, A.; Baliere, C.; Matheus, S.; Batejat, C.F.; Manuguerra, J.C.; Vanhomwegen, J.; Caro, V. Rapid Genomic Characterization of SARS-CoV-2 by Direct Amplicon-Based Sequencing Through Comparison of MinION and Illumina iSeq100(TM) System. Front. Microbiol. 2020, 11, 571328. [Google Scholar] [CrossRef]
- Bull, R.A.; Adikari, T.N.; Ferguson, J.M.; Hammond, J.M.; Stevanovski, I.; Beukers, A.G.; Naing, Z.; Yeang, M.; Verich, A.; Gamaarachchi, H.; et al. Analytical validity of nanopore sequencing for rapid SARS-CoV-2 genome analysis. Nat. Commun. 2020, 11, 6272. [Google Scholar] [CrossRef]
- WHO. Genomic Sequencing of SARS-CoV-2: A Guide to Implementation for Maximum Impact on Public Health; World Health Organization: Geneva, Switzerland, 2021. [Google Scholar]
- Bhoyar, R.C.; Senthivel, V.; Jolly, B.; Imran, M.; Jain, A.; Divakar, M.K.; Scaria, V.; Sivasubbu, S. An optimized, amplicon-based approach for sequencing of SARS-CoV-2 from patient samples using COVIDSeq assay on Illumina MiSeq sequencing platforms. STAR Protoc. 2021, 2, 100755. [Google Scholar] [CrossRef]
- Pembaur, A.; Sallard, E.; Weil, P.P.; Ortelt, J.; Ahmad-Nejad, P.; Postberg, J. Simplified Point-of-Care Full SARS-CoV-2 Genome Sequencing Using Nanopore Technology. Microorganisms 2021, 9, 2598. [Google Scholar] [CrossRef]
- Lambisia, A.W.; Mohammed, K.S.; Makori, T.O.; Ndwiga, L.; Mburu, M.W.; Morobe, J.M.; Moraa, E.O.; Musyoki, J.; Murunga, N.; Mwangi, J.N.; et al. Optimization of the SARS-CoV-2 ARTIC Network V4 Primers and Whole Genome Sequencing Protocol. Front. Med. 2022, 9, 836728. [Google Scholar] [CrossRef]
- Hofman, P.; Boutros, J.; Benchetrit, D.; Benzaquen, J.; Leroy, S.; Tanga, V.; Bordone, O.; Allégra, M.; Lespinet, V.; Fayada, J.; et al. A rapid near-patient RT-PCR test for suspected COVID-19: A study of the diagnostic accuracy. Ann. Transl. Med. 2021, 9, 921. [Google Scholar] [CrossRef]
- Tanga, V.; Leroy, S.; Fayada, J.; Hamila, M.; Allegra, M.; Messaoudi, Z.; Bonnetaud, C.; Lespinet, V.; Bordone, O.; Washetine, K.; et al. Establishment of a Collection of Blood-Derived Products from COVID-19 Patients for Translational Research: Experience of the LPCE Biobank (Nice, France). Biopreservation Biobanking 2020, 18, 517–524. [Google Scholar] [CrossRef]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef]
- Hofman, P.; Bordone, O.; Chamorey, E.; Benzaquen, J.; Schiappa, R.; Lespinet-Fabre, V.; Lanteri, E.; Brest, P.; Mograbi, B.; Maniel, C.; et al. Setting-Up a Rapid SARS-CoV-2 Genome Assessment by Next-Generation Sequencing in an Academic Hospital Center (LPCE, Louis Pasteur Hospital, Nice, France). Front. Med. 2021, 8, 730577. [Google Scholar] [CrossRef]
- Alcazer, V. StatAid: An R package with a graphical user interface for data analysis. J. Open Source Softw. 2020, 5, 2630. [Google Scholar] [CrossRef]

| Genexus Integrated Sequencer | Illumina NextSeq RUO COVID-Seq | Midnight ONT | |
|---|---|---|---|
| Workflow TAT | 1 working day | 3 working days | 1 working day |
| Total run time | 24 h | 40 h | 11 h |
| Hands-on time | 5 min | 3 h | 1 h 10 min |
| Touchpoints | 1 | >80 | >55 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ilié, M.; Benzaquen, J.; Hofman, V.; Long-Mira, E.; Lassalle, S.; Boutros, J.; Bontoux, C.; Lespinet-Fabre, V.; Bordone, O.; Tanga, V.; et al. Accurate Detection of SARS-CoV-2 by Next-Generation Sequencing in Low Viral Load Specimens. Int. J. Mol. Sci. 2023, 24, 3478. https://doi.org/10.3390/ijms24043478
Ilié M, Benzaquen J, Hofman V, Long-Mira E, Lassalle S, Boutros J, Bontoux C, Lespinet-Fabre V, Bordone O, Tanga V, et al. Accurate Detection of SARS-CoV-2 by Next-Generation Sequencing in Low Viral Load Specimens. International Journal of Molecular Sciences. 2023; 24(4):3478. https://doi.org/10.3390/ijms24043478
Chicago/Turabian StyleIlié, Marius, Jonathan Benzaquen, Véronique Hofman, Elodie Long-Mira, Sandra Lassalle, Jacques Boutros, Christophe Bontoux, Virginie Lespinet-Fabre, Olivier Bordone, Virginie Tanga, and et al. 2023. "Accurate Detection of SARS-CoV-2 by Next-Generation Sequencing in Low Viral Load Specimens" International Journal of Molecular Sciences 24, no. 4: 3478. https://doi.org/10.3390/ijms24043478
APA StyleIlié, M., Benzaquen, J., Hofman, V., Long-Mira, E., Lassalle, S., Boutros, J., Bontoux, C., Lespinet-Fabre, V., Bordone, O., Tanga, V., Allegra, M., Salah, M., Fayada, J., Leroy, S., Vassallo, M., Touitou, I., Courjon, J., Contenti, J., Carles, M., ... Hofman, P. (2023). Accurate Detection of SARS-CoV-2 by Next-Generation Sequencing in Low Viral Load Specimens. International Journal of Molecular Sciences, 24(4), 3478. https://doi.org/10.3390/ijms24043478








