Correlations between the NMR Lipoprotein Profile, APOE Genotype, and Cholesterol Efflux Capacity of Fasting Plasma from Cognitively Healthy Elderly Adults
Abstract
1. Introduction
2. Results
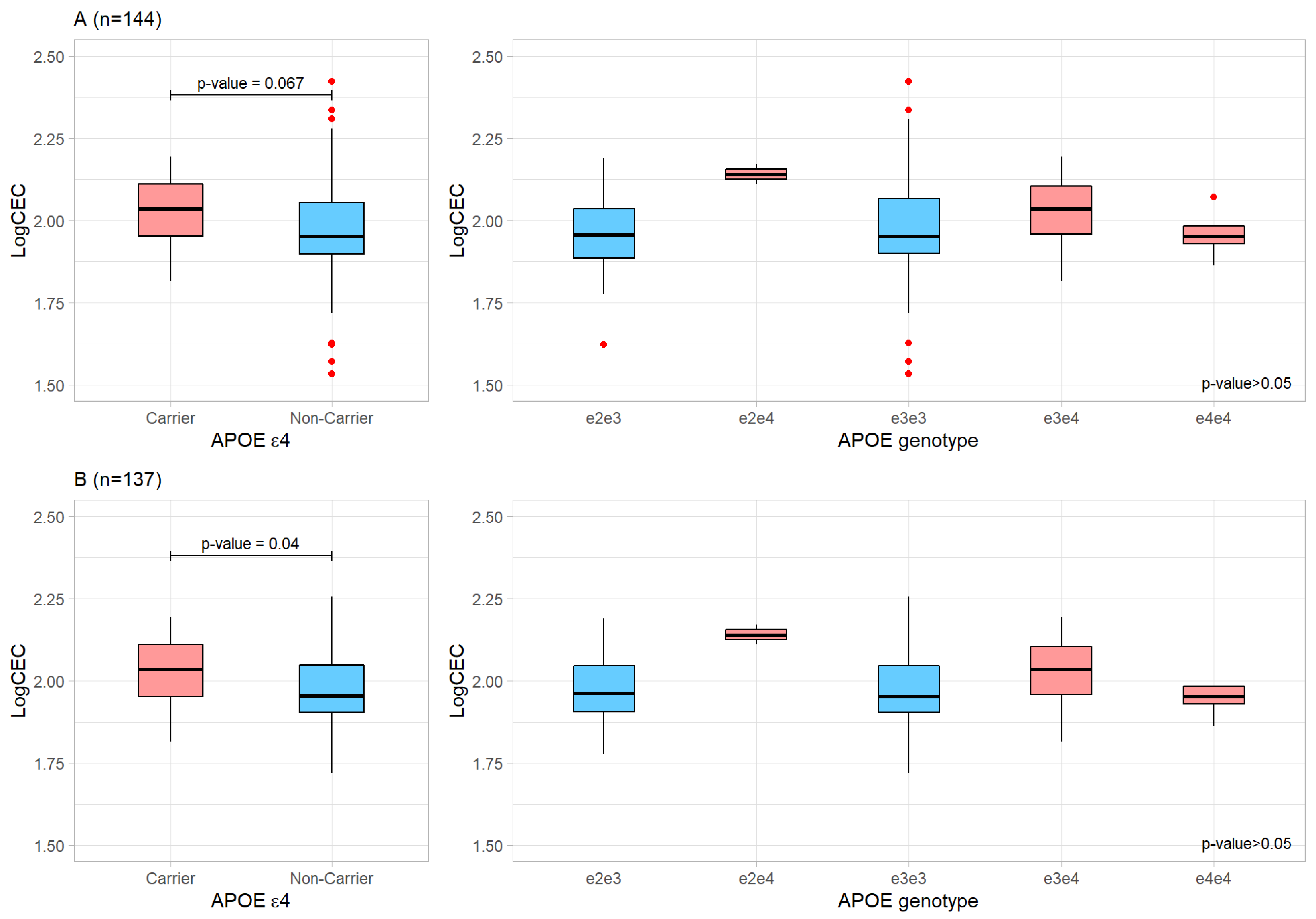
2.1. Effect of the APOE Genotype on the Plasma Cholesterol Efflux Capacity
2.2. Factors Associated with Plasma CEC
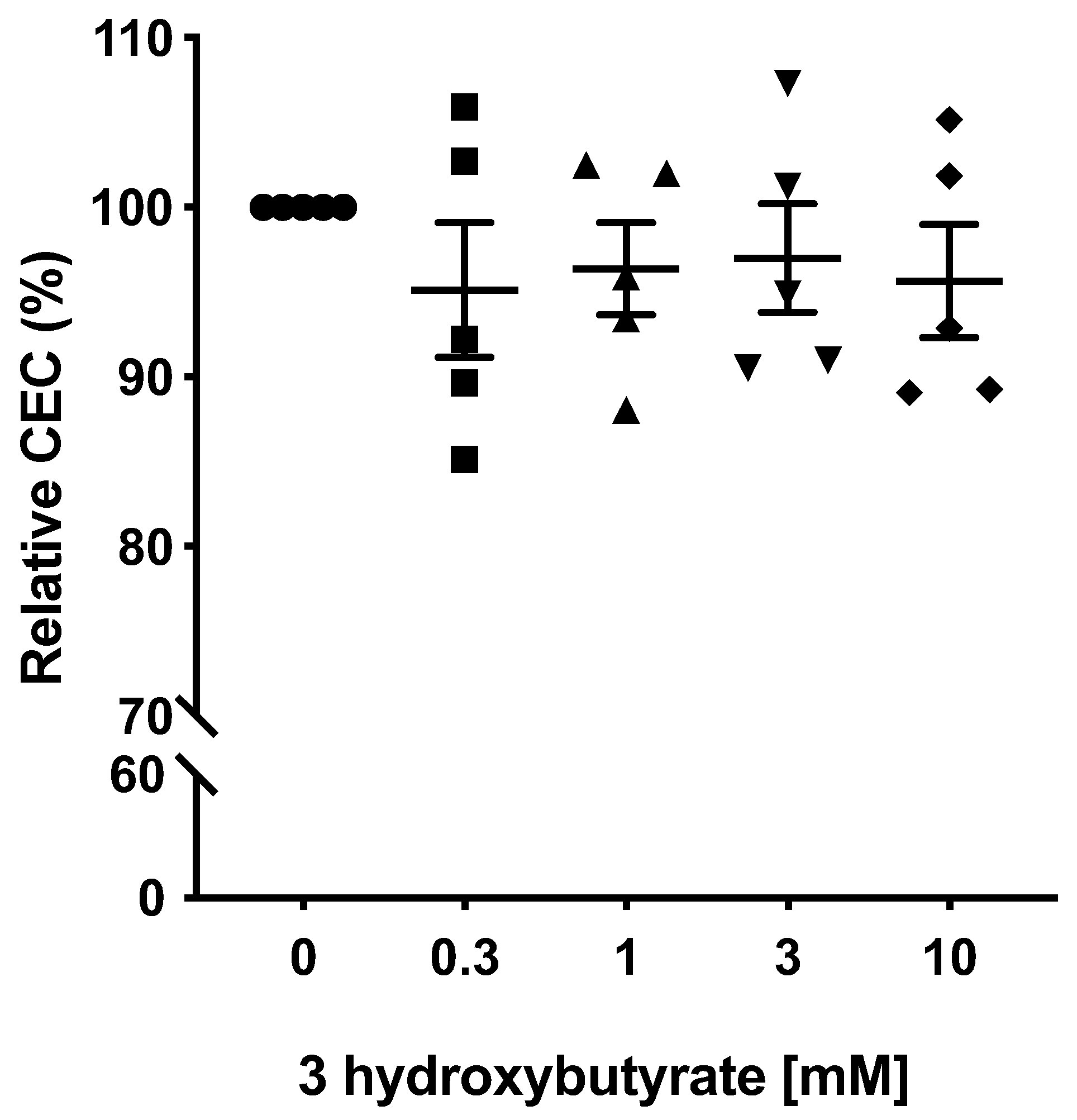
2.3. 3-Hydroxybutyrate Has No Direct Effect on Plasma Cholesterol Efflux Capacity
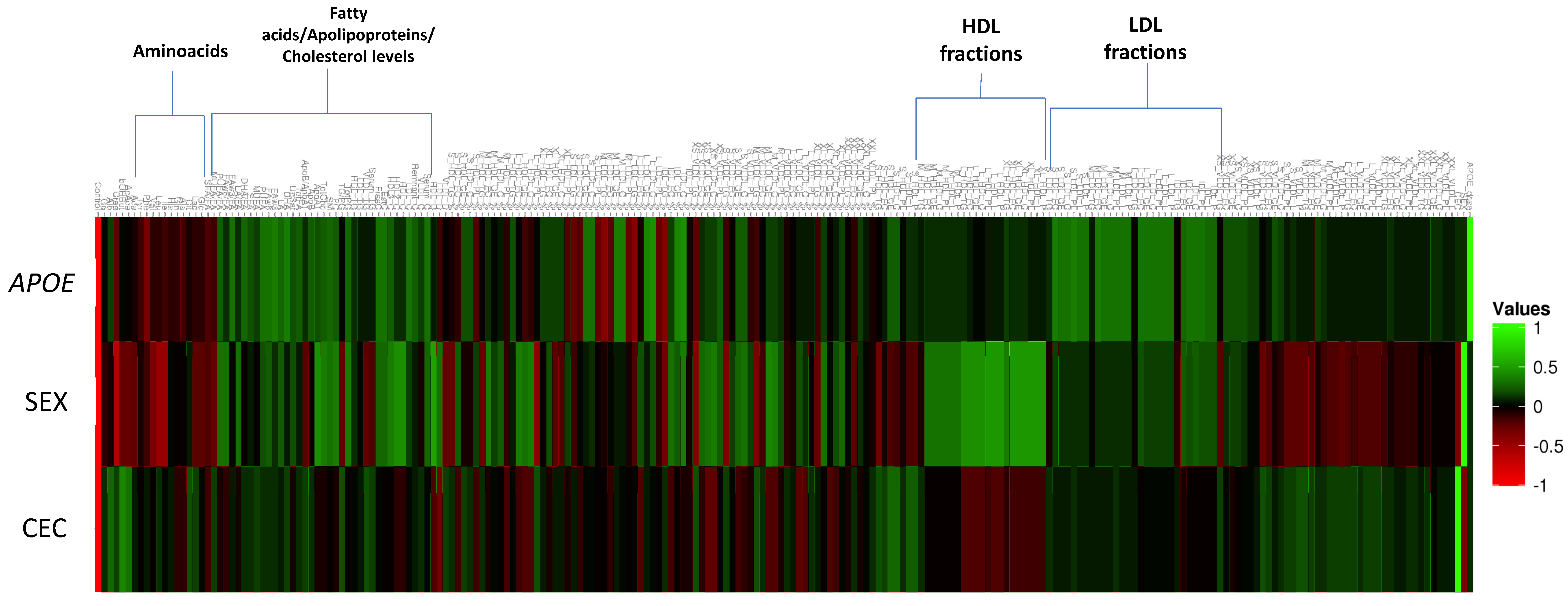
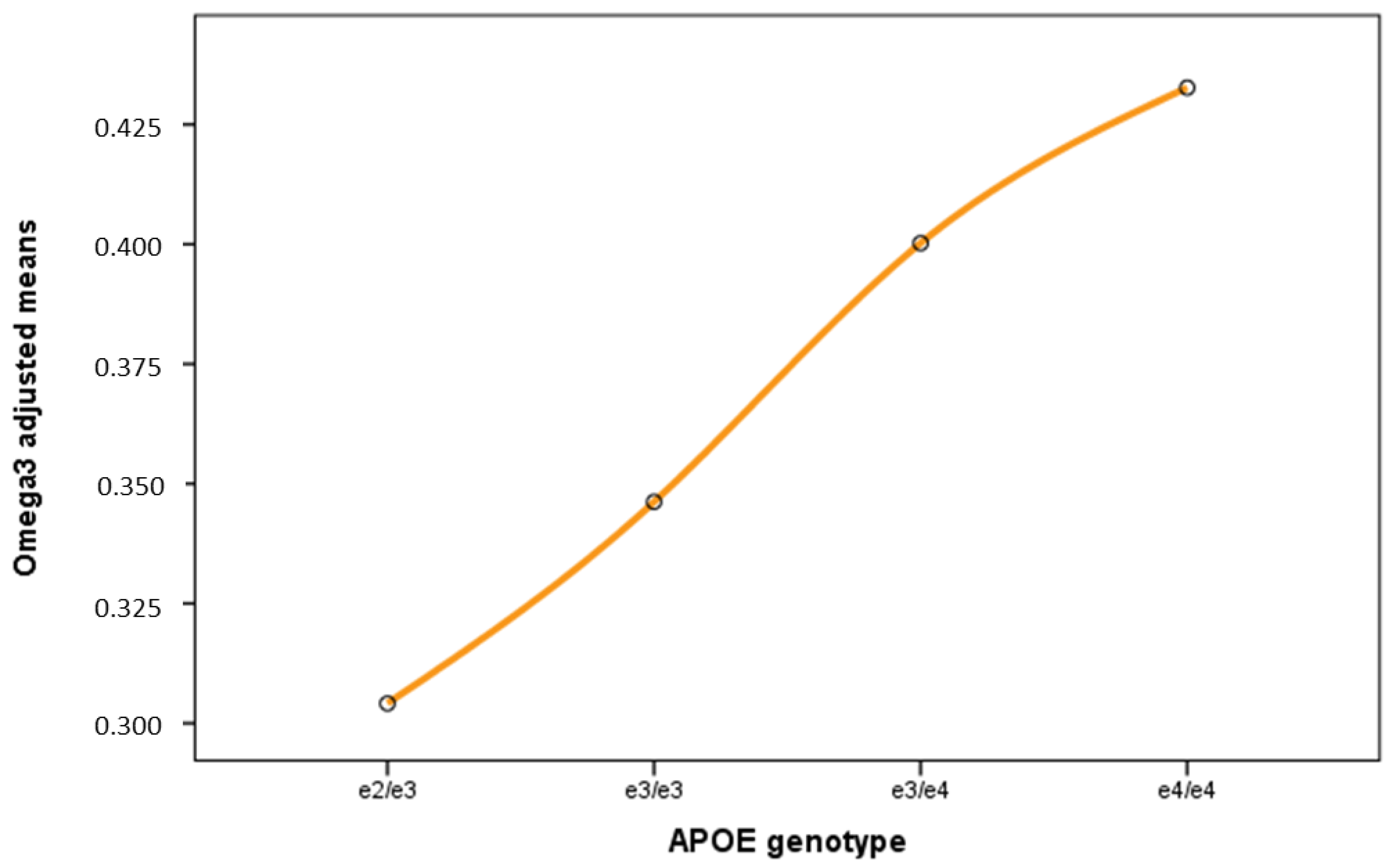
2.4. Effect of APOE on the Human Plasma NMR Profile
3. Discussion
4. Methods
4.1. Patients and Samples
4.2. Determination of the Cholesterol Efflux Capacity
4.3. High-Throughput Proton Nuclear Magnetic Resonance Metabolomics Profiling
4.4. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mira, E.; Manes, S. Immunomodulatory and Anti-Inflammatory Activities of Statins. Endocr. Metab Immune Disord Drug Targets 2009, 9, 237–247. [Google Scholar] [CrossRef] [PubMed]
- Talbot, C.P.J.; Plat, J.; Ritsch, A.; Mensink, R.P. Progress in Lipid Research. In Determinants of Cholesterol Efflux Capacity in Humans; Elsevier Ltd.: Amsterdam, The Netherlands, 2018; Volume 69, pp. 21–32. Available online: https://pubmed.ncbi.nlm.nih.gov/29269048/ (accessed on 27 August 2020).
- Anastasius, M.; Luquain-Costaz, C.; Kockx, M.; Jessup, W.; Kritharides, L. A Critical Appraisal of the Measurement of Serum ‘Cholesterol Efflux Capacity’ and Its Use as Surrogate Marker of Risk of Cardiovascular Disease. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2018, 1863, 1257–1273. [Google Scholar] [CrossRef] [PubMed]
- Sankaranarayanan, S.; De La Llera-Moya, M.; Drazul-Schrader, D.; Phillips, M.C.; Kellner-Weibel, G.; Rothblat, G.H. Serum Albumin Acts as a Shuttle to Enhance Cholesterol effl ux from Cells. J. Lipid Res. 2013, 54, 671–676. [Google Scholar] [CrossRef] [PubMed]
- Sankaranarayanan, S.; Kellner-Weibel, G.; de la Llera-Moya, M.; Phillips, M.C.; Asztalos, B.F.; Bittman, R.; Rothblat, G.H. A sensitive assay for ABCA1-mediated cholesterol efflux using BODIPY-cholesterol. J. Lipid Res. 2011, 52, 2332–2340. [Google Scholar] [CrossRef]
- Roe, A.; Hillman, J.; Butts, S.; Smith, M.; Rader, D.; Playford, M.; Mehta, N.N.; Dokras, A. Decreased Cholesterol Efflux Capacity and Atherogenic Lipid Profile in Young Women with PCOS. J. Clin. Endocrinol. Metab. 2014, 99, E841–E847. [Google Scholar] [CrossRef]
- Rohatgi, A.; Khera, A.; Berry, J.D.; Givens, E.G.; Ayers, C.R.; Wedin, K.E.; Neeland, I.J.; Yuhanna, I.S.; Rader, D.R.; de Lemos, J.A.; et al. HDL Cholesterol Efflux Capacity and Incident Cardiovascular Events. N. Engl. J. Med. 2014, 371, 2383–2393. [Google Scholar] [CrossRef]
- Gall, J.; Frisdal, E.; Bittar, R.; Le Goff, W.; Bruckert, E.; Lesnik, P.; Guerin, M.; Giral, P. Association of Cholesterol Efflux Capacity with Clinical Features of Metabolic Syndrome: Relevance to Atherosclerosis. J. Am. Heart Assoc. 2016, 5, e004808. [Google Scholar] [CrossRef]
- Yassine, H.N.; Feng, Q.; Chiang, J.; Petrosspour, L.M.; Fonteh, A.N.; Chui, H.C.; Harrington, M.G. ABCA1-Mediated Cholesterol Efflux Capacity to Cerebrospinal Fluid Is Reduced in Patients with Mild Cognitive Impairment and Alzheimer’s Disease. J. Am. Heart Assoc. 2016, 5, e002886. [Google Scholar] [CrossRef]
- Jones, L.; Lambert, J.C.; Wang, L.S.; Choi, S.H.; Harold, D.; Vedernikov, A.; Escott-Price, V.; Stone, T.; Richards, A.; Bellenguez, C. Convergent Genetic and Expression Data Implicate Immunity in Alzheimer’s Disease. Alzheimer Dement. 2015, 11, 658–671. [Google Scholar]
- Moreno-Grau, S.; Rodríguez-Gómez, O.; Sanabria, A.; Pérez-Cordón, A.; SAnchez-Ruiz, D.; Abdelnour, C.; Valero, S.; Hernandez, I.; Rosende-Roca, M.; Mauleon, A.; et al. Exploring APOE Genotype Effects on Alzheimer’s Disease Risk and Amyloid Amyloid Beta Burden in Individuals with Subjective Cognitive Decline: The FundacioACE Healthy Brain Initiative (FACEHBI) study baseline results. Alzheimer Dement. 2018, 14, 634–643. [Google Scholar] [CrossRef]
- Tudorache, I.F.; Trusca, V.G.; Gafencu, A.V. Apolipoprotein E: A Multifunctional Protein with Implications in Various Pathologies as a Result of Its Structural Features. Computat. Struct. Biotechnol. J. 2017, 15, 359–365. [Google Scholar] [CrossRef] [PubMed]
- Farrer, L.A.; Cupples, L.A.; Haines, J.L.; Hyman, B.; Kukull, W.A.; Mayeux, R.; Myers, R.H.; Pericak-Vance, M.A.; Risch, N.; van Duijn, C.M. Effects of age, sex, and ethnicity on the association between apolipoprotein E genotype and Alzheimer disease: A meta-analysis. J. Am. Med. Assoc. 1997, 278, 1349–1356. [Google Scholar] [CrossRef]
- Zhu, Y.; Bellosta, S.; Langer, C.; Bernini, F.; Pitas, R.E.; Mahley, R.W.; Assmann, G.; von Eckardstein, A. Low-Dose Expression of a Human Apolipoprotein E Transgene in Macrophages Restores Cholesterol Efflux Capacity of Apolipoprotein E-Deficient Mouse Plasma. Proc. Natl. Acad. Sci. USA 1998, 95, 7585–7590. [Google Scholar] [CrossRef] [PubMed]
- Hara, M.; Matsushima, T.; Satoh, H.; Iso-o, N.; Noto, H.; Togo, M.; Kimura, S.; Hashimoto, Y.; Tsukamoto, K. Isoform-Dependent Cholesterol Efflux from Macrophages by Apolipoprotein E is Modulated by Cell Surface Proteoglycans. Arterioscler. Thromb. Vasc. Biol. 2003, 23, 269–274. [Google Scholar] [CrossRef] [PubMed]
- Altenburg, M.; Johnson, L.; Wilder, J.; Maeda, N. Apolipoprotein E4 in Macrophages Enhances Atherogenesis in a Low Density Lipoprotein Receptor-Dependent Manner. J. Biol. Chem. 2007, 282, 7817–7824. [Google Scholar] [CrossRef]
- Heinsinger, N.M.; Gachechiladze, M.A.; Rebeck, G.W. Apolipoprotein E Genotype Affects Size of ApoE Complexes in Cerebrospinal Fluid. J. Neuropathol. Exp. Neurol. 2016, 75, 918–924. [Google Scholar] [CrossRef]
- Rawat, V.; Wang, S.; Sima, J.; Bar, R.; Liraz, O.; Gundimeda, U.; Parekh, T.; Chan, J.; Johansson, J.; Tang, C.; et al. ApoE4 Alters ABCA1 Membrane Trafficking in Astrocytes. J. Neurosci. 2019, 39, 9611–9622. [Google Scholar] [CrossRef]
- Camponova, P.; Le Page, A.; Berrougui, H.; Lamoureux, J.; Pawelec, G.; Witkowski, M.J.; Fulop, T.; Khalil, A. Alteration of high-density lipoprotein functionality in Alzheimer’s disease patients. Can. J. Physiol. Pharmacol. 2017, 95, 894–903. [Google Scholar] [CrossRef]
- Low-Kam, C.; Rhainds, D.; Lo, K.S.; Barhdadi, A.; Boulé, M.; Alem, S.; Pedneault-Gagnon, V.; Rhéaume, E.; Dubé, M.; Busseuil, D.; et al. Variants at the APOE/C1/C2/C4 Locus Modulate Cholesterol Efflux Capacity Independently of High-Density Lipoprotein Cholesterol. J. Am. Heart Assoc. 2018, 7, e009545. [Google Scholar] [CrossRef]
- Dedkova, E.N.; Blatter, L.A.; Aon, M.A.; Hopkins, J.; Pavlov, E. Role of β-Hydroxybutyrate, Its Polymer Poly-β-hydroxybutyrate and Inorganic Polyphosphate in Mammalian Health and Disease. 2014. Available online: www.frontiersin.org (accessed on 27 August 2020).
- Castaño, D.; Rattanasopa, C.; Monteiro-Cardoso, V.F.; Corlianò, M.; Liu, Y.; Zhong, S.; Rusu, M.; Liehn, E.A.; Singaraja, R.R. Lipid Efflux Mechanisms, Relation to Disease and Potential Therapeutic Aspects. Adv. Drug Deliv. Rev. 2020, 159, 54–93. [Google Scholar] [CrossRef]
- Pamir, N.; Hutchins, P.M.; Ronsein, G.E.; Wei, H.; Tang, C.; Das, R.; Vaisar, T.; Plow, E.; Schuster, V.; Reardon, C.A.; et al. Plasminogen Promotes Cholesterol Efflux by the ABCA1 Pathway. JCI Insight 2017, 2, e92176. [Google Scholar] [CrossRef]
- Kockx, M.; Traini, M.; Kritharides, L. Cell-Specific Production, Secretion, and Function of Apolipoprotein E. J. Mol. Med. 2018, 96, 361–371. [Google Scholar] [CrossRef] [PubMed]
- Schilcher, I.; Kern, S.; Hrzenjak, A.; Eichmann, T.O.; Stojakovic, T.; Scharnagl, H.; Duta-Mare, M.; Kratky, D.; Marsche, G.; Frank, S. Impact of Endothelial Lipase on Cholesterol Efflux Capacity of Serum and High-Density Lipoprotein. Sci. Rep. 2017, 7, 12485. [Google Scholar] [CrossRef] [PubMed]
- Kypreos, K.E.; Zannis, V.I. Pathway of Biogenesis of Apolipoprotein E-Containing HDL In Vivo with the Participation of ABCA1 and LCAT. Biochem. J. 2007, 403, 359–367. [Google Scholar] [CrossRef]
- Laffel, L. Ketone Bodies: A Review of Physiology, Pathophysiology and Application of Monitoring to Diabetes. Diabetes Metab. Res. Rev. 1999, 15, 412–426. [Google Scholar] [CrossRef]
- Nazih, H.; Nazih-Sanderson, F.; Krempf, M.; Huvelin, J.M.; Mercier, S.; Bard, J.M. Butyrate Stimulates apoA-IV-Containing Lipoprotein Secretion in Differentiated Caco-2 Cells: Role in Cholesterol Efflux. J. Cell Biochem. 2001, 83, 230–238. [Google Scholar] [CrossRef]
- Du, Y.; Li, X.; Su, C.; Xi, M.; Zhang, X.; Jiang, Z.; Wang, L.; Hong, B. Butyrate Protects against High-Fat Diet-Induced Atherosclerosis via Up-Regulating ABCA1 Expression in Apolipoprotein E-Deficiency Mice. Br. J. Pharmacol. 2020, 177, 1754–1772. [Google Scholar] [CrossRef]
- Bai, H.-B.; Yang, P.; Zhang, H.-B.; Liu, Y.-L.; Fang, S.-X.; Xu, X.-Y. Short-Chain Fatty Acid Butyrate Acid Attenuates Atherosclerotic Plaque Formation in Apolipoprotein E-Knockout Mice and the Underlying Mechanism. Sheng Li Xue Bao 2021, 73, 42–50. [Google Scholar]
- Nomura, S.; Umeda, T.; Tomiyama, T.; Mori, H. The E693Δ (Osaka) Mutation in Amyloid Precursor Protein Potentiates Cholesterol-Mediated Intracellular Amyloid β Toxicity via Its Impaired Cholesterol Efflux. J. Neurosci. Res. 2013, 91, 1541–1550. [Google Scholar] [CrossRef]
- Rebeck, G.W. Cholesterol Efflux as a Critical Component of Alzheimer’s Disease Pathogenesis. J. Mol. Neurosci. 2004, 23, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Simons, M.; Keller, P.; De Strooper, B.; Beyreuther, K.; Dotti, C.G.; Simons, K. Cholesterol Depletion Inhibits the Generation of β-Amyloid in Hippocampal Neurons. Proc. Natl. Acad. Sci. USA 1998, 95, 6460–6464. [Google Scholar] [CrossRef] [PubMed]
- Fassbender, K.; Simons, M.; Bergmann, C.; Stroick, M.; Lütjohann, D.; Keller, P.; Runz, H.; Kühl, S.; Bertsch, T.; von Bergmann, K.; et al. Simvastatin Strongly Reduces Levels of Alzheimer’s Disease β-Amyloid Peptides Aβ42 and Aβ40 In Vitro and In Vivo. Proc. Natl. Acad. Sci. USA 2001, 98, 5856–5861. [Google Scholar] [CrossRef]
- Puglielli, L.; Konopka, G.; Pack-Chung, E.; Ingano, L.A.M.; Berezovska, O.; Hyman, B.T.; Chang, T.-Y.; Tanzi, R.E.; Kovacs, D.M. Acyl-Coenzyme A: Cholesterol Acyltransferase Modulates the Generation of the Amyloid β-Peptide. Nat. Cell Biol. 2001, 3, 905–912. [Google Scholar] [CrossRef] [PubMed]
- Refolo, L.M.; Pappolla, M.A.; Malester, B.; LaFrancois, J.; Bryant-Thomas, T.; Wang, R.; Tint, G.S.; Sambamurti, K.; Duff, K.; Pappolla, M.A. Hypercholesterolemia Accelerates the Alzheimer’s Amyloid Pathology in a Transgenic Mouse Model. Neurobiol. Dis. 2000, 7, 321–331. [Google Scholar] [CrossRef]
- Petanceska, S.S.; DeRosa, S.; Olm, V.; Diaz, N.; Sharma, A.; Thomas-Bryant, T.; Duff, K.; Pappolla, M.; Refolo, L.M. Statin Therapy for Alzheimer’s Disease. Will It Work? J. Mol. Neurosci. 2002, 19, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Dafnis, I.; Raftopoulou, C.; Mountaki, C.; Megalou, E.; Zannis, V.I.; Chroni, A. ApoE Isoforms and Carboxyl-Terminal-Truncated apoE4 Forms Affect Neuronal BACE1 Levels and Aβ Production Independently of Their Cholesterol Efflux Capacity. Biochem. J. 2018, 475, 1839–1859. [Google Scholar] [CrossRef]
- Henderson, S.T. Ketone Bodies as a Therapeutic for Alzheimer’s Disease. Neurotherapeutics 2008, 5, 470–480. [Google Scholar] [CrossRef]
- Rodriguez-Gomez, O.; Sanabria, A.; Perez-Cordon, A.; Sanchez-Ruiz, D.; Abdelnour, C.; Valero, S.; Hernandez, I.; Rosende-Roca, M.; Mauleon, A.; Vargas, L.; et al. FACEHBI: A Prospective Study of Risk Factors, Biomarkers and Cognition in a Cohort of Individuals with Subjective Cognitive Decline. Study Rationale and Research Protocols. J. Prev. Alzheimer Dis. 2017, 4, 100–108. [Google Scholar] [CrossRef]
- Boada, M.; Tárraga, L.; Hernández, I.; Valero, S.; Alegret, M.; Ruiz, A.; Lopez, O.L.; Becker, J.T.; Fundació ACE Alzheimer Research Center and Memory Clinic. Design of a comprehensive Alzheimer’s disease clinic and research center in Spain to meet critical patient and family needs. Alzheimers Dement. 2014, 10, 409–415. [Google Scholar] [CrossRef]
- Alegret, M.; Espinosa, A.; Valero, S.; Vinyes-Junqué, G.; Ruiz, A.; Hernández, I.; Rosende-Roca, M.; Mauleón, A.; Becker, J.T.; Tárraga, L.; et al. Cut-off Scores of a Brief Neuropsychological Battery (NBACE) for Spanish Individual Adults Older than 44 Years Old. PLoS ONE 2013, 8, e76436. [Google Scholar] [CrossRef]
- Soininen, P.; Kangas, A.J.; Würtz, P.; Tukiainen, T.; Tynkkynen, T.; Laatikainen, R.; Jarvelin, M.-R.; Kähönen, M.; Lehtimäki, T.; Viikari, J.; et al. High-Throughput Serum NMR Metabonomics for Cost-Effective Holistic Studies on Systemic Metabolism. Analyst 2009, 134, 1781–1785. [Google Scholar] [CrossRef]
- Kettunen, J.; Demirkan, A.; Würtz, P.; Draisma, H.H.M.; Haller, T.; Rawal, R.; Vaarhorst, A.; Kangas, A.J.; Lyytikäinen, L.; Pirinen, M.; et al. Genome-Wide Study for Circulating Metabolites Identifies 62 loci and Reveals Novel Systemic Effects of LPA. Nat. Commun. 2016, 7, 11122. [Google Scholar] [CrossRef] [PubMed]
- Lacour, A.; Espinosa, A.; Louwersheimer, E.; Heilmann-Heimbach, S.; Hernández, I.; Wolfsgruber, S.; Fernández, V.; Wagner, H.; Rosende-Roca, M.; Mauleón, A.; et al. Genome-Wide Significant Risk Factors for Alzheimer’s Disease: Role in Progression to Dementia due to Alzheimer’s Disease among Subjects with Mild Cognitive Impairment. Mol. Psychiatry 2016, 22, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Breiman, L. Technical Note: Some Properties of Splitting Criteria. Mach. Learn. 1996, 24, 41–47. [Google Scholar] [CrossRef]
| Variable | Effect in CEC (beta) | p-Value (CEC Effect) | APOE Impact (beta) | p-Value (APOE Effect) |
|---|---|---|---|---|
| 3-Hydroxybutyrate | 0.365 | 0.0000073 * | 0.025 | 0.765 |
| Sex | −0.326 | 0.000068 * | 0.096 | 0.254 |
| Acetoacetate | 0.317 | 0.00012 * | −0.013 | 0.88 |
| Mean diameter for LDL particles | −0.282 | 0.001 | −0.147 | 0.08 |
| Albumin | 0.279 | 0.001 | 0.147 | 0.08 |
| Total cholesterol in small HDL particles | 0.254 | 0.002 | 0.221 | 0.008 |
| Cholesterol esters in small HDL particles | 0.245 | 0.003 | 0.23 | 0.006 |
| Concentration of small HDL particles | 0.229 | 0.006 | 0.109 | 0.195 |
| Total lipids in small HDL particles | 0.229 | 0.006 | 0.117 | 0.165 |
| Familial antecedents of Parkinson’s disease | −0.22 | 0.009 | −0.18 | 0.034 |
| Total-cholesterol-to-total-lipid ratio in very small VLDL particles | −0.216 | 0.009 | 0.156 | 0.064 |
| NMR Parameter | APOE (beta) | p-Value APOE | CEC (beta) | p-Value CEC |
|---|---|---|---|---|
| Total-cholesterol-to-total-lipid ratio in large LDL particles | 0.406 | 0.00000054 | 0.002 | 0.984 |
| Total-cholesterol-to-total-lipid ratio in IDL particles | 0.405 | 0.00000058 | −0.073 | 0.386 |
| Triglyceride-to-total-lipid ratio in large LDL particles | −0.398 | 0.00000094 | 0.024 | 0.778 |
| Triglyceride-to-total-lipid ratio in medium LDL particles | −0.386 | 0.000002 | −0.008 | 0.925 |
| Triglyceride-to-total-lipid ratio in IDL particles | −0.378 | 0.000004 | 0.094 | 0.262 |
| Cholesterol-ester-to-total-lipid ratio in large LDL particles | 0.376 | 0.000004 | 0.074 | 0.377 |
| Cholesterol-ester-to-total-lipid ratio in IDL particles | 0.375 | 0.000004 | −0.003 | 0.968 |
| FAw3: Omega-3 fatty acids | 0.359 | 0.000011 | 0.105 | 0.212 |
| Total-cholesterol-to-total-lipid ratio in medium LDL particles | 0.357 | 0.000013 | 0.024 | 0.774 |
| Free cholesterol in small LDL particles | 0.346 | 0.000025 | 0.054 | 0.524 |
| Cholesterol-ester-to-total-lipid ratio in medium LDL particles | 0.337 | 0.000041 | 0.053 | 0.529 |
| DHA: 22:6, docosahexaenoic acid | 0.334 | 0.000048 | 0.123 | 0.142 |
| Phospholipids in small LDL particles | 0.334 | 0.00005 | 0.098 | 0.246 |
| Free cholesterol in medium LDL particles | 0.332 | 0.000055 | 0.055 | 0.516 |
| EstC: Esterified cholesterol | 0.32 | 0.000105 | 0 | 0.999 |
| Total cholesterol in small LDL particles | 0.319 | 0.000111 | 0.049 | 0.564 |
| FAw3/FA: Ratio of omega-3 fatty acids to total fatty acids | 0.318 | 0.000115 | 0.075 | 0.373 |
| Total lipids in small LDL particles | 0.317 | 0.000121 | 0.066 | 0.434 |
| PUFAs: Polyunsaturated fatty acids | 0.316 | 0.000129 | 0.09 | 0.284 |
| Total cholesterol in medium LDL particles | 0.316 | 0.000131 | 0.051 | 0.548 |
| Serum total cholesterol | 0.313 | 0.000148 | 0.001 | 0.994 |
| Concentration of small LDL particles | 0.312 | 0.000156 | 0.07 | 0.408 |
| Total cholesterol in LDL particles | 0.312 | 0.000157 | 0.033 | 0.698 |
| Cholesterol esters in medium LDL particles | 0.311 | 0.000166 | 0.049 | 0.557 |
| Cholesterol esters in small LDL particles | 0.311 | 0.000169 | 0.047 | 0.574 |
| Phospholipids in medium LDL particles | 0.31 | 0.000173 | 0.085 | 0.315 |
| Total-cholesterol-to-total-lipid ratio in small LDL particles | 0.308 | 0.000196 | 0 | 1 |
| Phospholipids in large LDL particles | 0.307 | 0.000203 | 0.026 | 0.757 |
| Cholesterol esters in large LDL particles | 0.306 | 0.000216 | 0.025 | 0.766 |
| Total lipids in medium LDL particles | 0.306 | 0.000218 | 0.06 | 0.48 |
| Total cholesterol in large LDL particles | 0.305 | 0.000219 | 0.016 | 0.852 |
| Variable | beta (Effect) | p-Value (beta) |
|---|---|---|
| (Regression constant) | 0.697 | |
| Hypertension | −0.184 | 0.016 |
| Dyslipidemia | 0.137 | 0.091 |
| Total-cholesterol-to-total-lipid ratio in IDL particles (IDL-C-%) | 5.373 | 0.000016 * |
| Triglyceride-to-total-lipid ratio in large LDL particles (L-LDL-TG-%) | 0.627 | 0.08 |
| Cholesterol-ester-to-total-lipid ratio in large LDL particles (L-LDL-CE-%) | −4.202 | 0.02 |
| Cholesterol-ester-to-total-lipid ratio in IDL particles (IDL-CE-%) | −2.395 | 0.003 |
| Omega-3 fatty acids (FAw3) | 1.718 | 0.002 |
| Free cholesterol in small LDL particles (S-LDL-FC) | 2.076 | 0.033 |
| Cholesterol-ester-to-total-lipid ratio in medium LDL particles (M-LDL-CE-%) | 8.641 | 0.001 |
| Esterified cholesterol (EstC) | −4.59 | 0.000017 * |
| Ratio of omega-3 fatty acids to total fatty acids (FAw3/FA) | −0.944 | 0.024 |
| Concentration of small LDL particles (S-LDL-P) | −5.025 | 0.091 |
| Cholesterol esters in medium LDL particles (M-LDL-CE) | 3.588 | 0.033 |
| Phospholipids in medium LDL particles (M-LDL-PL) | 3.228 | 0.012 |
| Total-cholesterol-to-total-lipid ratio in small LDL particles (S-LDL-C-%) | −7.003 | 0.00018 * |
| Demographics | Valid Sample | Mean|n | Std|% |
|---|---|---|---|
| Age (years) | 144 | 65.6 | 8.9 |
| Sex (female) | 144 | 82 | 56.9 |
| Height (cm) | 140 | 165.7 | 9.2 |
| Weight (kg) | 141 | 73.4 | 13.2 |
| BMI (kg/m2) | 140 | 26.6 | 3.97 |
| Familiar antecedent dementia (% subjects) | 137 | 90 | 65.7 |
| Familial antecedent Parkinson’s disease (% subjects) | 138 | 13 | 9.4 |
| Dyslipidemia (% subjects) | 137 | 64 | 46.7 |
| Hypertension (% subjects) | 138 | 46 | 33.7 |
| APOE (E4 carriers) (% subjects) | 143 | 31 | 21.7 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
de Rojas, I.; del Barrio, L.; Hernández, I.; Montrreal, L.; García-González, P.; Marquié, M.; Valero, S.; Cano, A.; Orellana, A.; Boada, M.; et al. Correlations between the NMR Lipoprotein Profile, APOE Genotype, and Cholesterol Efflux Capacity of Fasting Plasma from Cognitively Healthy Elderly Adults. Int. J. Mol. Sci. 2023, 24, 2186. https://doi.org/10.3390/ijms24032186
de Rojas I, del Barrio L, Hernández I, Montrreal L, García-González P, Marquié M, Valero S, Cano A, Orellana A, Boada M, et al. Correlations between the NMR Lipoprotein Profile, APOE Genotype, and Cholesterol Efflux Capacity of Fasting Plasma from Cognitively Healthy Elderly Adults. International Journal of Molecular Sciences. 2023; 24(3):2186. https://doi.org/10.3390/ijms24032186
Chicago/Turabian Stylede Rojas, Itziar, Laura del Barrio, Isabel Hernández, Laura Montrreal, Pablo García-González, Marta Marquié, Sergi Valero, Amanda Cano, Adelina Orellana, Mercè Boada, and et al. 2023. "Correlations between the NMR Lipoprotein Profile, APOE Genotype, and Cholesterol Efflux Capacity of Fasting Plasma from Cognitively Healthy Elderly Adults" International Journal of Molecular Sciences 24, no. 3: 2186. https://doi.org/10.3390/ijms24032186
APA Stylede Rojas, I., del Barrio, L., Hernández, I., Montrreal, L., García-González, P., Marquié, M., Valero, S., Cano, A., Orellana, A., Boada, M., Mañes, S., & Ruiz, A. (2023). Correlations between the NMR Lipoprotein Profile, APOE Genotype, and Cholesterol Efflux Capacity of Fasting Plasma from Cognitively Healthy Elderly Adults. International Journal of Molecular Sciences, 24(3), 2186. https://doi.org/10.3390/ijms24032186







