Convergent Evolution Dynamics of SARS-CoV-2 and HIV Surface Envelope Glycoproteins Driven by Host Cell Surface Receptors and Lipid Rafts: Lessons for the Future
Abstract
1. Introduction
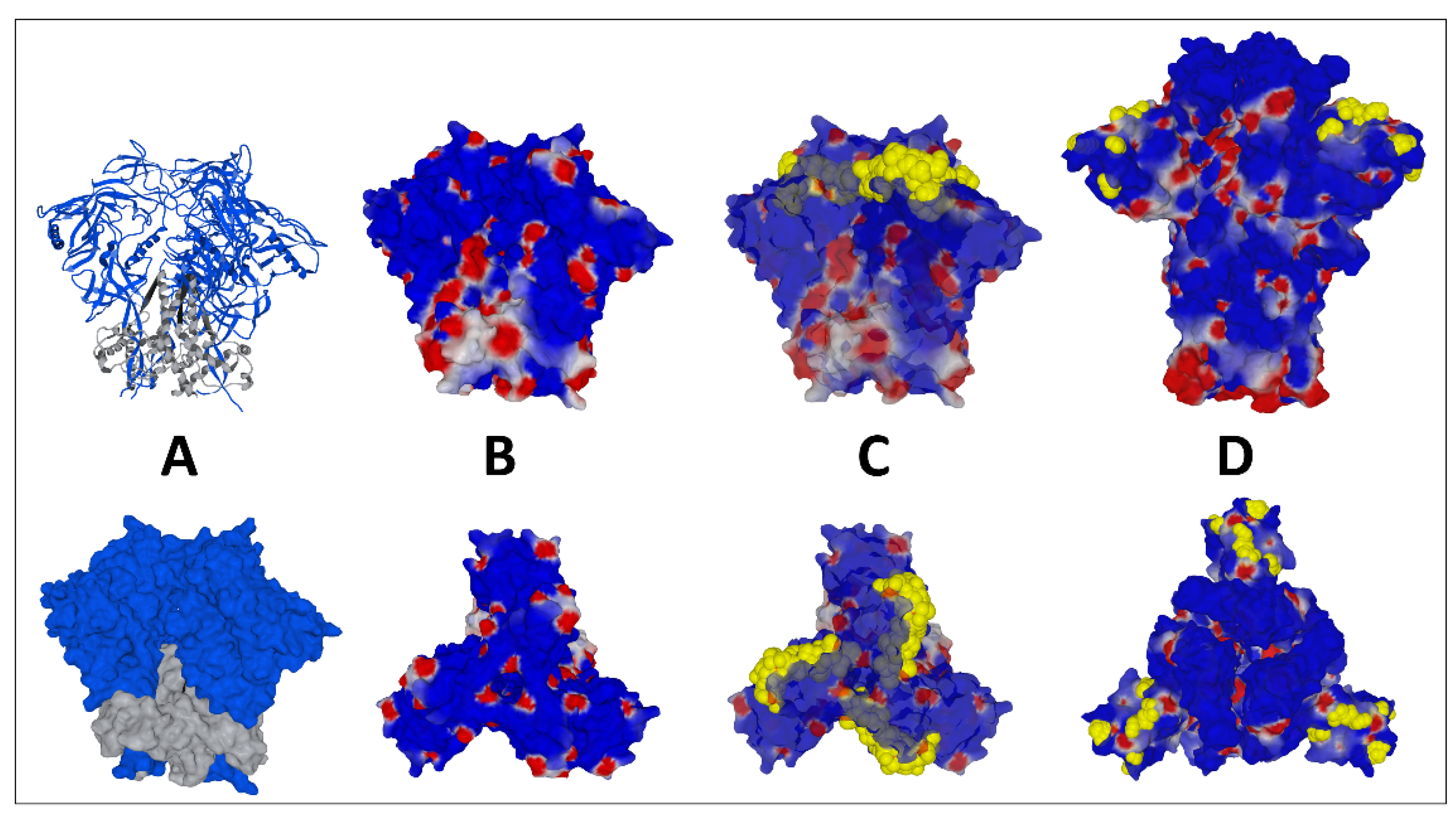
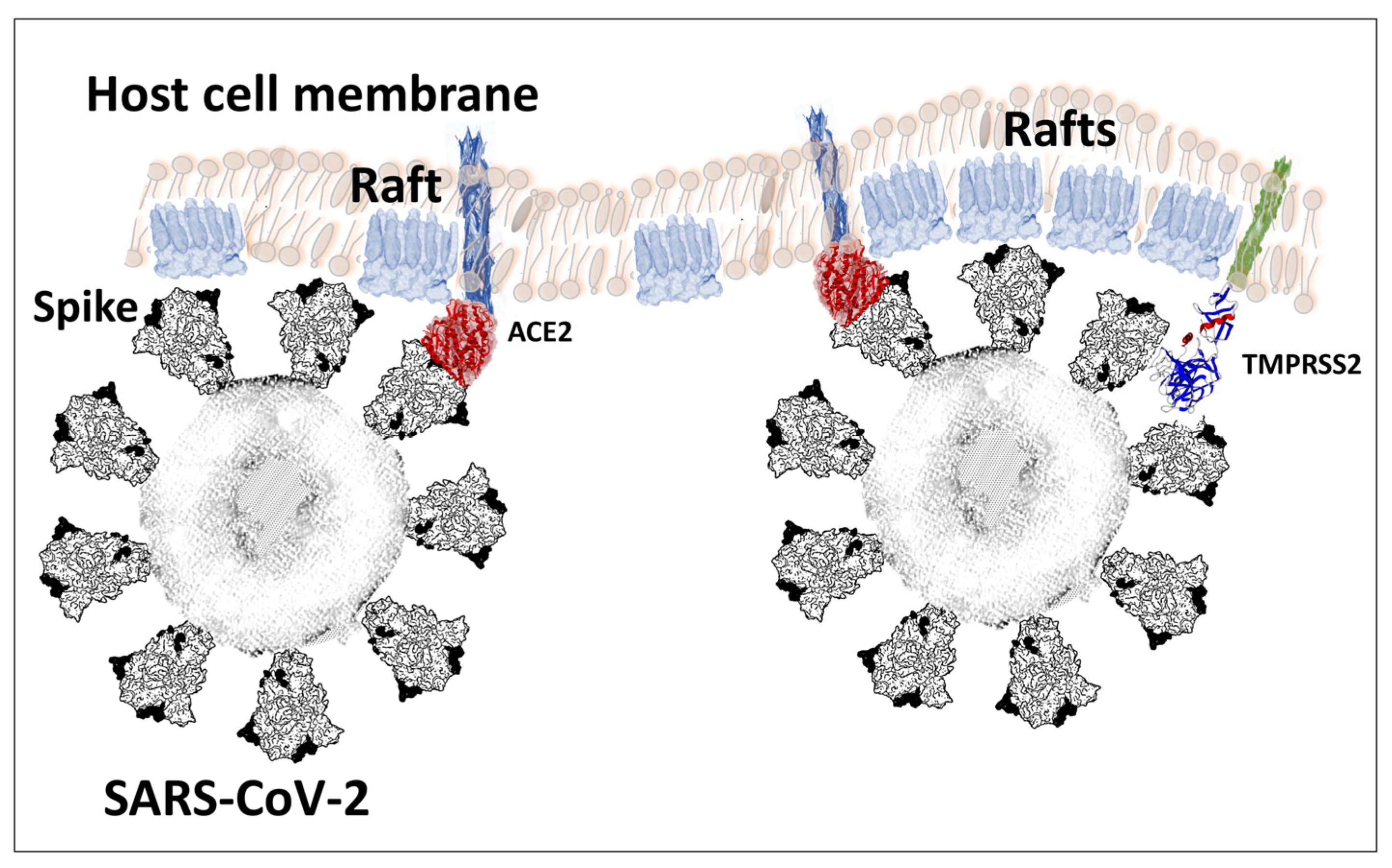
2. Structural and Functional Analysis of the SARS-CoV-2 Spike Protein
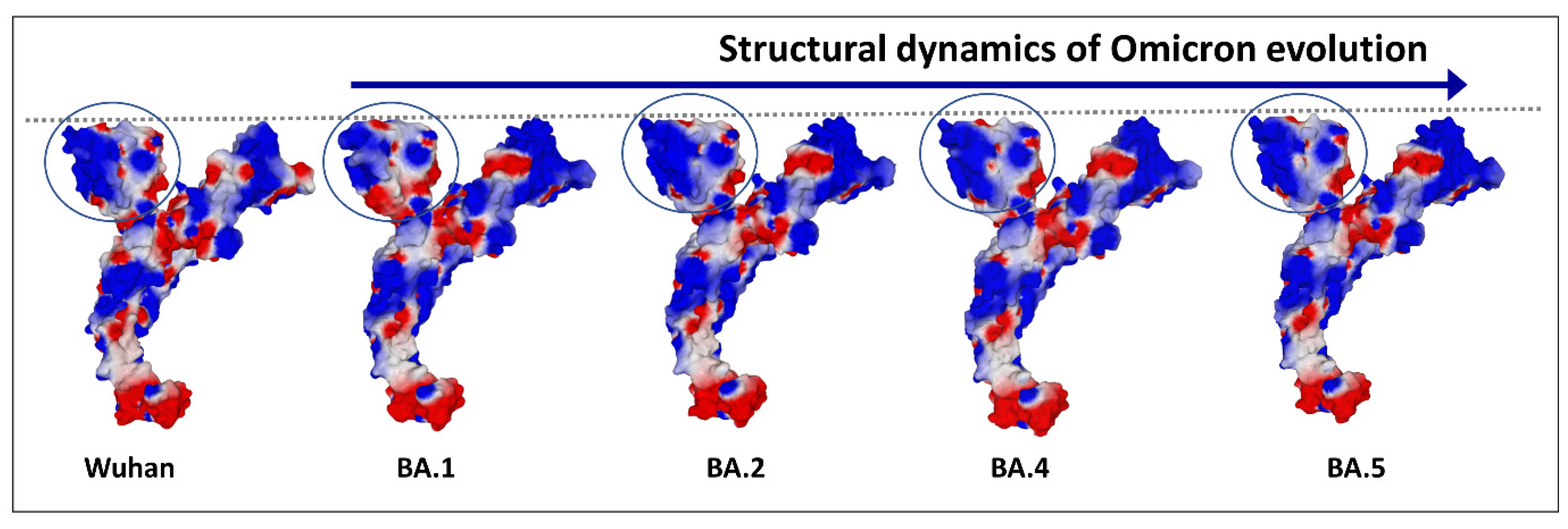
3. Structural Dynamics of SARS-CoV-2 Spike Protein Evolution
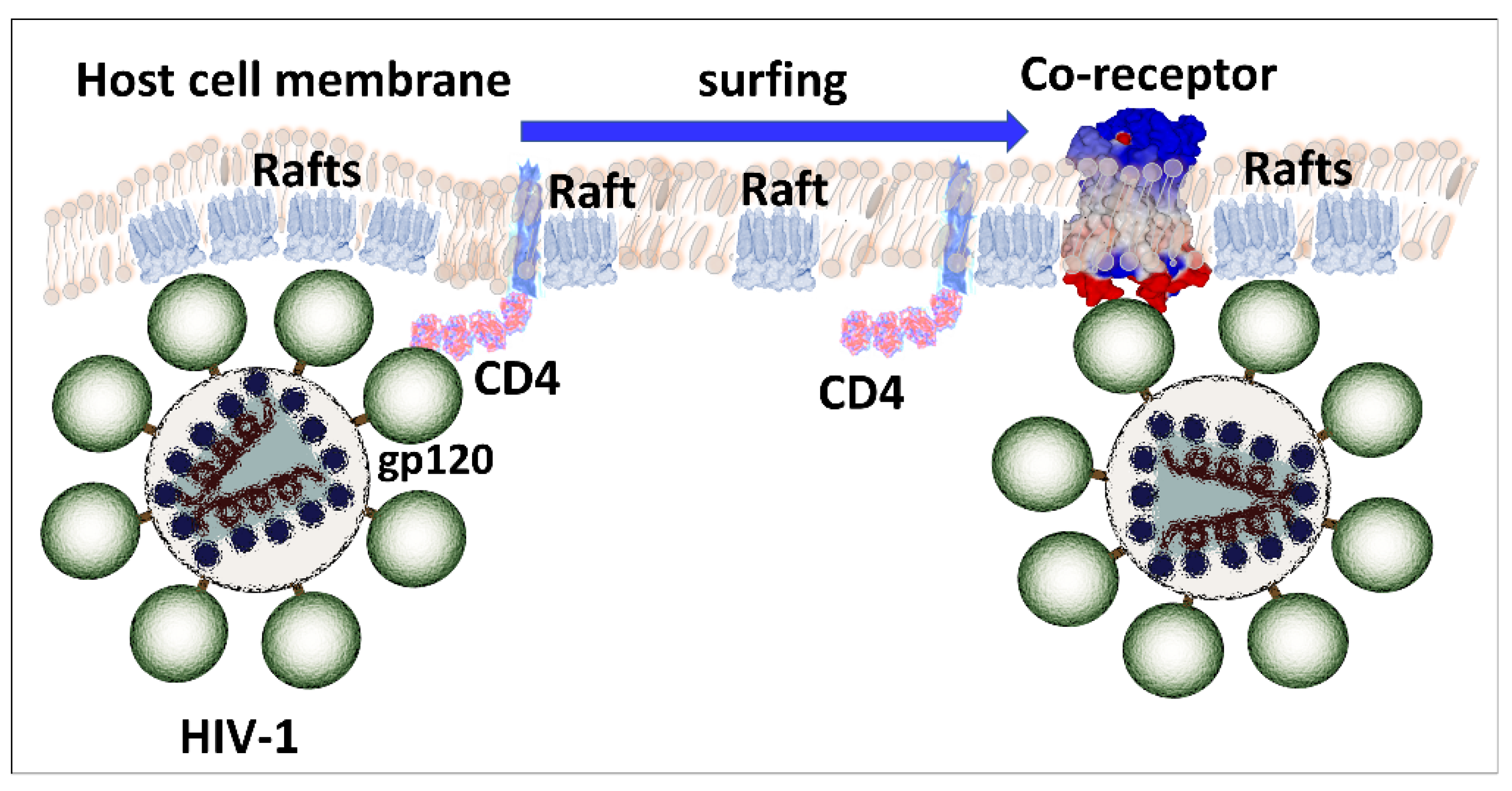
4. Structural and Functional Analysis of HIV-1 gp120 Surface Envelope Glycoprotein
5. Structural Dynamics of HIV-1 gp120 Surface Envelope Glycoprotein Evolution
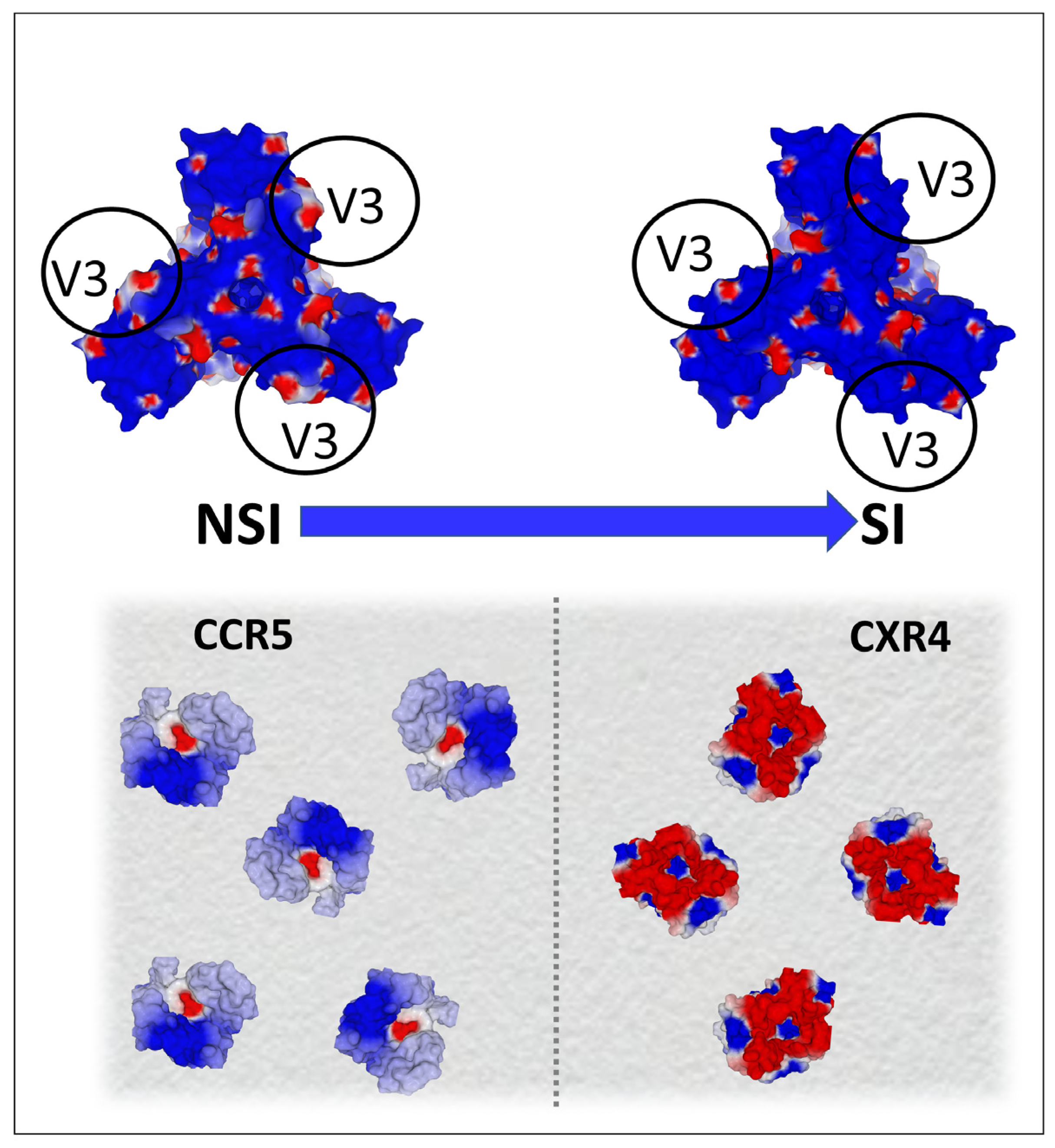
6. Considerations on the Electrostatic Surface Potential
7. Impact on Immune Responses
8. Key Role of Lipid Rafts
9. Common Mechanisms of Pathology
10. Therapeutic Applications
Author Contributions
Funding
Conflicts of Interest
References
- Chermann, J.C.; Donker, G.; Yahi, N.; Salaun, D.; Guettari, N.; Gayet, O.; Hirsh, I. Discrepancies in AIDS virus data. Nature 1991, 351, 277–278. [Google Scholar] [CrossRef]
- Venaud, S.; Yahi, N.; Fehrentz, J.L.; Guettari, N.; Nisato, D.; Hirsch, I.; Chermann, J.C. Inhibition of HIV by an anti-HIV protease synthetic peptide blocks an early step of viral replication. Res. Virol. 1992, 143, 311–319. [Google Scholar] [CrossRef] [PubMed]
- Fantini, J.; Yahi, N.; Chermann, J.C. Human immunodeficiency virus can infect the apical and basolateral surfaces of human colonic epithelial cells. Proc. Natl. Acad. Sci. USA 1991, 88, 9297–9301. [Google Scholar] [CrossRef]
- Yahi, N.; Baghdiguian, S.; Moreau, H.; Fantini, J. Galactosyl ceramide (or a closely related molecule) is the receptor for human immunodeficiency virus type 1 on human colon epithelial HT29 cells. J. Virol. 1992, 66, 4848–4854. [Google Scholar] [CrossRef]
- Harouse, J.M.; Bhat, S.; Spitalnik, S.L.; Laughlin, M.; Stefano, K.; Silberberg, D.H.; Gonzalez-Scarano, F. Inhibition of entry of HIV-1 in neural cell lines by antibodies against galactosyl ceramide. Science 1991, 253, 320–323. [Google Scholar] [CrossRef] [PubMed]
- Fantini, J.; Yahi, N.; Baghdiguian, S.; Chermann, J.C. Human colon epithelial cells productively infected with human immunodeficiency virus show impaired differentiation and altered secretion. J. Virol. 1992, 66, 580–585. [Google Scholar] [CrossRef]
- Dayanithi, G.; Yahi, N.; Baghdiguian, S.; Fantini, J. Intracellular calcium release induced by human immunodeficiency virus type 1 (HIV-1) surface envelope glycoprotein in human intestinal epithelial cells: A putative mechanism for HIV-1 enteropathy. Cell Calcium 1995, 18, 9–18. [Google Scholar] [CrossRef]
- Delézay, O.; Yahi, N.; Tamalet, C.; Baghdiguian, S.; Boudier, J.A.; Fantini, J. Direct effect of type 1 human immunodeficiency virus (HIV-1) on intestinal epithelial cell differentiation: Relationship to HIV-1 enteropathy. Virology 1997, 238, 231–242. [Google Scholar] [CrossRef] [PubMed]
- Fantini, J.; Maresca, M.; Hammache, D.; Yahi, N.; Delézay, O. Glycosphingolipid (GSL) microdomains as attachment platforms for host pathogens and their toxins on intestinal epithelial cells: Activation of signal transduction pathways and perturbations of intestinal absorption and secretion. Glycoconj. J. 2000, 17, 173–179. [Google Scholar] [CrossRef]
- Maresca, M.; Mahfoud, R.; Garmy, N.; Kotler, D.P.; Fantini, J.; Clayton, F. The virotoxin model of HIV-1 enteropathy: Involvement of GPR15/Bob and galactosylceramide in the cytopathic effects induced by HIV-1 gp120 in the HT-29-D4 intestinal cell line. J. Biomed. Sci. 2003, 10, 156–166. [Google Scholar] [CrossRef]
- Hammache, D.; Yahi, N.; Maresca, M.; Piéroni, G.; Fantini, J. Human erythrocyte glycosphingolipids as alternative cofactors for human immunodeficiency virus type 1 (HIV-1) entry: Evidence for CD4-induced interactions between HIV-1 gp120 and reconstituted membrane microdomains of glycosphingolipids (Gb3 and GM3). J. Virol. 1999, 73, 5244–5248. [Google Scholar] [CrossRef]
- Puri, A.; Hug, P.; Jernigan, K.; Barchi, J.; Kim, H.Y.; Hamilton, J.; Wiels, J.; Murray, G.J.; Brady, R.O.; Blumenthal, R. The neutral glycosphingolipid globotriaosylceramide promotes fusion mediated by a CD4-dependent CXCR4-utilizing HIV type 1 envelope glycoprotein. Proc. Natl. Acad. Sci. USA 1998, 95, 14435–14440. [Google Scholar] [CrossRef]
- Hammache, D.; Piéroni, G.; Yahi, N.; Delézay, O.; Koch, N.; Lafont, H.; Tamalet, C.; Fantini, J. Specific interaction of HIV-1 and HIV-2 surface envelope glycoproteins with monolayers of galactosylceramide and ganglioside GM3. J. Biol. Chem. 1998, 273, 7967–7971. [Google Scholar] [CrossRef]
- Hug, P.; Lin, H.M.; Korte, T.; Xiao, X.; Dimitrov, D.S.; Wang, J.M.; Puri, A.; Blumenthal, R. Glycosphingolipids promote entry of a broad range of human immunodeficiency virus type 1 isolates into cell lines expressing CD4, CXCR4, and/or CCR5. J. Virol. 2000, 74, 6377–6385. [Google Scholar] [CrossRef]
- Rawat, S.S.; Johnson, B.T.; Puri, A. Sphingolipids: Modulators of HIV-1 infection and pathogenesis. Biosci. Rep. 2005, 25, 329–343. [Google Scholar] [CrossRef]
- Simons, K.; Ikonen, E. Functional rafts in cell membranes. Nature 1997, 387, 569–572. [Google Scholar] [CrossRef]
- Fantini, J.; Garmy, N.; Mahfoud, R.; Yahi, N. Lipid rafts: Structure, function and role in HIV, Alzheimer’s and prion diseases. Expert Rev. Mol. Med. 2002, 4, 1–22. [Google Scholar] [CrossRef]
- Fantini, J.; Hammache, D.; Piéroni, G.; Yahi, N. Role of glycosphingolipid microdomains in CD4-dependent HIV-1 fusion. Glycoconj. J. 2000, 17, 199–204. [Google Scholar] [CrossRef]
- Hammache, D.; Yahi, N.; Piéroni, G.; Ariasi, F.; Tamalet, C.; Fantini, J. Sequential interaction of CD4 and HIV-1 gp120 with a reconstituted membrane patch of ganglioside GM3: Implications for the role of glycolipids as potential HIV-1 fusion cofactors. Biochem. Biophys. Res. Commun. 1998, 246, 117–122. [Google Scholar] [CrossRef]
- Deng, H.; Liu, R.; Ellmeier, W.; Choe, S.; Unutmaz, D.; Burkhart, M.; Di Marzio, P.; Marmon, S.; Sutton, R.E.; Hill, C.M.; et al. Identification of a major co-receptor for primary isolates of HIV-1. Nature 1996, 381, 661–666. [Google Scholar] [CrossRef]
- Feng, Y.; Broder, C.C.; Kennedy, P.E.; Berger, E.A. HIV-1 entry cofactor: Functional cDNA cloning of a seven-transmembrane, G protein-coupled receptor. Science 1996, 272, 872–877. [Google Scholar] [CrossRef]
- Puri, A.; Hug, P.; Muñoz-Barroso, I.; Blumenthal, R. Human erythrocyte glycolipids promote HIV-1 envelope glycoprotein-mediated fusion of CD4+ cells. Biochem. Biophys. Res. Commun. 1998, 242, 219–225. [Google Scholar] [CrossRef]
- Puri, A.; Rawat, S.S.; Lin, H.M.; Finnegan, C.M.; Mikovits, J.; Ruscetti, F.W.; Blumenthal, R. An inhibitor of glycosphingolipid metabolism blocks HIV-1 infection of primary T-cells. AIDS 2004, 18, 849–858. [Google Scholar] [CrossRef]
- Ablan, S.; Rawat, S.S.; Viard, M.; Wang, J.M.; Puri, A.; Blumenthal, R. The role of cholesterol and sphingolipids in chemokine receptor function and HIV-1 envelope glycoprotein-mediated fusion. Virol. J. 2006, 3, 104. [Google Scholar] [CrossRef][Green Version]
- Mahfoud, R.; Garmy, N.; Maresca, M.; Yahi, N.; Puigserver, A.; Fantini, J. Identification of a common sphingolipid-binding domain in Alzheimer, prion, and HIV-1 proteins. J. Biol. Chem. 2002, 277, 11292–11296. [Google Scholar] [CrossRef]
- Yahi, N.; Fantini, J. Deciphering the glycolipid code of Alzheimer’s and Parkinson’s amyloid proteins allowed the creation of a universal ganglioside-binding peptide. PloS ONE 2014, 9, e104751. [Google Scholar] [CrossRef]
- Fantini, J.; Chahinian, H.; Yahi, N. Leveraging coronavirus binding to gangliosides for innovative vaccine and therapeutic strategies against COVID-19. Biochem. Biophys. Res. Commun. 2021, 538, 132–136. [Google Scholar] [CrossRef]
- Andreani, J.; Le Bideau, M.; Duflot, I.; Jardot, P.; Rolland, C.; Boxberger, M.; Wurtz, N.; Rolain, J.M.; Colson, P.; La Scola, B.; et al. In vitro testing of combined hydroxychloroquine and azithromycin on SARS-CoV-2 shows synergistic effect. Microb. Pathog. 2020, 145, 104228. [Google Scholar] [CrossRef]
- Fantini, J.; Di Scala, C.; Chahinian, H.; Yahi, N. Structural and molecular modelling studies reveal a new mechanism of action of chloroquine and hydroxychloroquine against SARS-CoV-2 infection. Int. J. Antimicrob. Agents 2020, 55, 105960. [Google Scholar] [CrossRef]
- Fantini, J.; Chahinian, H.; Yahi, N. Synergistic antiviral effect of hydroxychloroquine and azithromycin in combination against SARS-CoV-2: What molecular dynamics studies of virus-host interactions reveal. Int. J. Antimicrob. Agents 2020, 56, 106020. [Google Scholar] [CrossRef]
- Fournier, P.E.; Colson, P.; Levasseur, A.; Devaux, C.A.; Gautret, P.; Bedotto, M.; Delerce, J.; Brechard, L.; Pinault, L.; Lagier, J.C.; et al. Emergence and outcomes of the SARS-CoV-2 ‘Marseille-4’ variant. Int. J. Infect. Dis. IJID Off. Publ. Int. Soc. Infect. Dis. 2021, 106, 228–236. [Google Scholar] [CrossRef]
- Vilar, S.; Isom, D.G. One Year of SARS-CoV-2: How Much Has the Virus Changed? Biology 2021, 10, 91. [Google Scholar] [CrossRef]
- Smith, E.C.; Sexton, N.R.; Denison, M.R. Thinking outside the triangle: Replication fidelity of the largest RNA viruses. Annu. Rev. Virol. 2014, 1, 111–132. [Google Scholar] [CrossRef]
- Fantini, J.; Yahi, N.; Azzaz, F.; Chahinian, H. Structural dynamics of SARS-CoV-2 variants: A health monitoring strategy for anticipating Covid-19 outbreaks. J. Infect. 2021, 83, 197–206. [Google Scholar] [CrossRef]
- Al-Khatib, H.A.; Smatti, M.K.; Ali, F.H.; Zedan, H.T.; Thomas, S.; Ahmed, M.N.; El-Kahlout, R.A.; Al Bader, M.A.; Elgakhlab, D.; Coyle, P.V.; et al. Comparative analysis of within-host diversity among vaccinated COVID-19 patients infected with different SARS-CoV-2 variants. iScience 2022, 25, 105438. [Google Scholar] [CrossRef]
- Bull, J.J.; Meyers, L.A.; Lachmann, M. Quasispecies made simple. PLoS Comput. Biol. 2005, 1, e61. [Google Scholar] [CrossRef]
- Domingo, E.; Menéndez-Arias, L.; Holland, J.J. RNA virus fitness. Rev. Med. Virol. 1997, 7, 87–96. [Google Scholar] [CrossRef]
- Yu, F.; Wen, Y.; Wang, J.; Gong, Y.; Feng, K.; Ye, R.; Jiang, Y.; Zhao, Q.; Pan, P.; Wu, H.; et al. The Transmission and Evolution of HIV-1 Quasispecies within One Couple: A Follow-up Study based on Next-Generation Sequencing. Sci. Rep. 2018, 8, 1404. [Google Scholar] [CrossRef]
- Guérin, P.; Yahi, N.; Azzaz, F.; Chahinian, H.; Sabatier, J.M.; Fantini, J. Structural Dynamics of the SARS-CoV-2 Spike Protein: A 2-Year Retrospective Analysis of SARS-CoV-2 Variants (from Alpha to Omicron) Reveals an Early Divergence between Conserved and Variable Epitopes. Molecules 2022, 27, 3851. [Google Scholar] [CrossRef]
- Welsby, P.D. A personal historical perspective of HIV. Postgrad. Med. J. 2020, 96, 408–411. [Google Scholar] [CrossRef]
- Al-Harthi, L.; Campbell, E.; Schneider, J.A.; Bennett, D.A. What HIV in the Brain Can Teach Us About SARS-CoV-2 Neurological Complications? AIDS Res. Hum. Retrovir. 2021, 37, 255–265. [Google Scholar] [CrossRef]
- Illanes-Álvarez, F.; Márquez-Ruiz, D.; Márquez-Coello, M.; Cuesta-Sancho, S.; Girón-González, J.A. Similarities and differences between HIV and SARS-CoV-2. Int. J. Med. Sci. 2021, 18, 846–851. [Google Scholar] [CrossRef]
- Fischer, W.; Giorgi, E.E.; Chakraborty, S.; Nguyen, K.; Bhattacharya, T.; Theiler, J.; Goloboff, P.A.; Yoon, H.; Abfalterer, W.; Foley, B.T.; et al. HIV-1 and SARS-CoV-2: Patterns in the evolution of two pandemic pathogens. Cell Host Microbe 2021, 29, 1093–1110. [Google Scholar] [CrossRef]
- Negi, G.; Sharma, A.; Dey, M.; Dhanawat, G.; Parveen, N. Membrane attachment and fusion of HIV-1, influenza A, and SARS-CoV-2: Resolving the mechanisms with biophysical methods. Biophys. Rev. 2022, 14, 1109–1140. [Google Scholar] [CrossRef]
- Demoliou, C.; Papaneophytou, C.; Nicolaidou, V. SARS-CoV-2 and HIV-1: So Different yet so Alike. Immune Response at the Cellular and Molecular Level. Int. J. Med. Sci. 2022, 19, 1787–1795. [Google Scholar] [CrossRef]
- Abernathy, M.E.; Dam, K.-M.A.; Esswein, S.R.; Jette, C.A.; Bjorkman, P.J. How Antibodies Recognize Pathogenic Viruses: Structural Correlates of Antibody Neutralization of HIV-1, SARS-CoV-2, and Zika. Viruses 2021, 13, 2106. [Google Scholar] [CrossRef]
- Saleemi, M.A.; Ahmad, B.; Benchoula, K.; Vohra, M.S.; Mea, H.J.; Chong, P.P.; Palanisamy, N.K.; Wong, E.H. Emergence and molecular mechanisms of SARS-CoV-2 and HIV to target host cells and potential therapeutics. Infect. Genet. Evol. 2020, 85, 104583. [Google Scholar] [CrossRef]
- Jackson, C.B.; Farzan, M.; Chen, B.; Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 2022, 23, 3–20. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Pöhlmann, S. A Multibasic Cleavage Site in the Spike Protein of SARS-CoV-2 Is Essential for Infection of Human Lung Cells. Mol. Cell 2020, 78, 779–784.e775. [Google Scholar] [CrossRef]
- Sasaki, M.; Uemura, K.; Sato, A.; Toba, S.; Sanaki, T.; Maenaka, K.; Hall, W.W.; Orba, Y.; Sawa, H. SARS-CoV-2 variants with mutations at the S1/S2 cleavage site are generated in vitro during propagation in TMPRSS2-deficient cells. PLoS Pathog. 2021, 17, e1009233. [Google Scholar] [CrossRef]
- Kawase, M.; Shirato, K.; Matsuyama, S.; Taguchi, F. Protease-Mediated Entry via the Endosome of Human Coronavirus 229E. J. Virol. 2009, 83, 712–721. [Google Scholar] [CrossRef]
- Cai, Y.; Zhang, J.; Xiao, T.; Peng, H.; Sterling, S.M.; Walsh, R.M., Jr.; Rawson, S.; Rits-Volloch, S.; Chen, B. Distinct conformational states of SARS-CoV-2 spike protein. Science 2020, 369, 1586–1592. [Google Scholar] [CrossRef]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef]
- Sun, X.L. The role of cell surface sialic acids for SARS-CoV-2 infection. Glycobiology 2021, 31, 1245–1253. [Google Scholar] [CrossRef]
- Palacios-Rápalo, S.N.; De Jesús-González, L.A.; Cordero-Rivera, C.D.; Farfan-Morales, C.N.; Osuna-Ramos, J.F.; Martínez-Mier, G.; Quistián-Galván, J.; Muñoz-Pérez, A.; Bernal-Dolores, V.; Del Ángel, R.M.; et al. Cholesterol-Rich Lipid Rafts as Platforms for SARS-CoV-2 Entry. Front. Immunol. 2021, 12, 796855. [Google Scholar] [CrossRef]
- Seyran, M.; Takayama, K.; Uversky, V.N.; Lundstrom, K.; Palù, G.; Sherchan, S.P.; Attrish, D.; Rezaei, N.; Aljabali, A.A.A.; Ghosh, S.; et al. The structural basis of accelerated host cell entry by SARS-CoV-2. FEBS J. 2021, 288, 5010–5020. [Google Scholar] [CrossRef]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 2020, 181, 271–280.e278. [Google Scholar] [CrossRef]
- Pak, A.J.; Yu, A.; Ke, Z.; Briggs, J.A.G.; Voth, G.A. Cooperative multivalent receptor binding promotes exposure of the SARS-CoV-2 fusion machinery core. Nat. Commun. 2022, 13, 1002. [Google Scholar] [CrossRef]
- Colson, P.; Lavagna, C.; Delerce, J.; Groshenry, G.; Yahi, N.; Fantini, J.; La Scola, B.; Althaus, T. First Detection of the SARS-CoV-2 Omicron BA.5/22B in Monaco. Microorganisms 2022, 10, 1952. [Google Scholar] [CrossRef]
- Yahi, N.; Chahinian, H.; Fantini, J. Infection-enhancing anti-SARS-CoV-2 antibodies recognize both the original Wuhan/D614G strain and Delta variants. A potential risk for mass vaccination? J. Infect. 2021, 83, 607–635. [Google Scholar] [CrossRef]
- Azzaz, F.; Yahi, N.; Di Scala, C.; Chahinian, H.; Fantini, J. Ganglioside binding domains in proteins: Physiological and pathological mechanisms. Adv. Protein Chem. Struct. Biol. 2022, 128, 289–324. [Google Scholar] [CrossRef]
- Gowrisankar, A.; Priyanka, T.M.C.; Banerjee, S. Omicron: A mysterious variant of concern. Eur. Phys. J. Plus 2022, 137, 100. [Google Scholar] [CrossRef]
- Mahase, E. Omicron: South Africa says fourth wave peak has passed as it lifts curfew. BMJ Br. Med. J. 2022, 376, o7. [Google Scholar] [CrossRef]
- Haschka, T.; Vergu, E.; Roche, B.; Poletto, C.; Opatowski, L. Retrospective analysis of SARS-CoV-2 omicron invasion over delta in French regions in 2021-22: A status-based multi-variant model. BMC Infect. Dis. 2022, 22, 815. [Google Scholar] [CrossRef]
- Bal, A.; Simon, B.; Destras, G.; Chalvignac, R.; Semanas, Q.; Oblette, A.; Quéromès, G.; Fanget, R.; Regue, H.; Morfin, F.; et al. Detection and prevalence of SARS-CoV-2 co-infections during the Omicron variant circulation in France. Nat. Commun. 2022, 13, 6316. [Google Scholar] [CrossRef]
- Grabowski, F.; Kochańczyk, M.; Lipniacki, T. The Spread of SARS-CoV-2 Variant Omicron with a Doubling Time of 2.0-3.3 Days Can Be Explained by Immune Evasion. Viruses 2022, 14, 294. [Google Scholar] [CrossRef]
- Zhang, L.; Li, Q.; Liang, Z.; Li, T.; Liu, S.; Cui, Q.; Nie, J.; Wu, Q.; Qu, X.; Huang, W. The significant immune escape of pseudotyped SARS-CoV-2 variant Omicron. Emerg. Microbes Infect. 2022, 11, 1–5. [Google Scholar] [CrossRef]
- Willett, B.J.; Grove, J.; MacLean, O.A.; Wilkie, C.; De Lorenzo, G.; Furnon, W.; Cantoni, D.; Scott, S.; Logan, N.; Ashraf, S.; et al. SARS-CoV-2 Omicron is an immune escape variant with an altered cell entry pathway. Nat. Microbiol. 2022, 7, 1161–1179. [Google Scholar] [CrossRef]
- Zhao, H.; Lu, L.; Peng, Z.; Chen, L.-L.; Meng, X.; Zhang, C.; Ip, J.D.; Chan, W.-M.; Chu, A.W.-H.; Chan, K.-H.; et al. SARS-CoV-2 Omicron variant shows less efficient replication and fusion activity when compared with Delta variant in TMPRSS2-expressed cells. Emerg. Microbes Infect. 2022, 11, 277–283. [Google Scholar] [CrossRef]
- Willett, B.J.; Grove, J.; MacLean, O.; Wilkie, C.; Logan, N.; De Lorenzo, G.; Furnon, W.; Scott, S.; Manali, M.; Szemiel, A. The hyper-transmissible SARS-CoV-2 Omicron variant exhibits significant antigenic change, vaccine escape and a switch in cell entry mechanism. MedRxiv 2022. [Google Scholar] [CrossRef]
- Fantini, J.; Yahi, N.; Colson, P.; Chahinian, H.; La Scola, B.; Raoult, D. The puzzling mutational landscape of the SARS-2-variant Omicron. J. Med. Virol. 2022, 94, 2019–2025. [Google Scholar] [CrossRef]
- Jaafar, R.; Boschi, C.; Aherfi, S.; Bancod, A.; Le Bideau, M.; Edouard, S.; Colson, P.; Chahinian, H.; Raoult, D.; Yahi, N.; et al. High Individual Heterogeneity of Neutralizing Activities against the Original Strain and Nine Different Variants of SARS-CoV-2. Viruses 2021, 13, 2177. [Google Scholar] [CrossRef] [PubMed]
- Akaishi, T.; Fujiwara, K.; Ishii, T. Insertion/deletion hotspots in the Nsp2, Nsp3, S1, and ORF8 genes of SARS-related coronaviruses. BMC Ecol. Evol. 2022, 22, 123. [Google Scholar] [CrossRef] [PubMed]
- Tamalet, C.; Izopet, J.; Koch, N.; Fantini, J.; Yahi, N. Stable rearrangements of the beta3-beta4 hairpin loop of HIV-1 reverse transcriptase in plasma viruses from patients receiving combination therapy. AIDS 1998, 12, F161–F166. [Google Scholar] [CrossRef] [PubMed]
- Tamalet, C.; Yahi, N.; Tourrès, C.; Colson, P.; Quinson, A.M.; Poizot-Martin, I.; Dhiver, C.; Fantini, J. Multidrug resistance genotypes (insertions in the beta3-beta4 finger subdomain and MDR mutations) of HIV-1 reverse transcriptase from extensively treated patients: Incidence and association with other resistance mutations. Virology 2000, 270, 310–316. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Li, Z.; Li, W.; Lu, M.; Bess, J., Jr.; Chao, C.W.; Gorman, J.; Terry, D.S.; Zhang, B.; Zhou, T.; Blanchard, S.C.; et al. Subnanometer structures of HIV-1 envelope trimers on aldrithiol-2-inactivated virus particles. Nat. Struct. Mol. Biol. 2020, 27, 726–734. [Google Scholar] [CrossRef] [PubMed]
- Willey, R.L.; Bonifacino, J.S.; Potts, B.J.; Martin, M.A.; Klausner, R.D. Biosynthesis, cleavage, and degradation of the human immunodeficiency virus 1 envelope glycoprotein gp160. Proc. Natl. Acad. Sci. USA 1988, 85, 9580–9584. [Google Scholar] [CrossRef]
- Chatterjee, S.; Basak, S.; Khan, N.C. Morphogenesis of human immunodeficiency virus type 1. Pathobiol. J. Immunopathol. Mol. Cell. Biol. 1992, 60, 181–186. [Google Scholar] [CrossRef]
- Turner, B.G.; Summers, M.F. Structural biology of HIV. J. Mol. Biol. 1999, 285, 1–32. [Google Scholar] [CrossRef]
- Fantini, J.; Chahinian, H.; Yahi, N. A Vaccine Strategy Based on the Identification of an Annular Ganglioside Binding Motif in Monkeypox Virus Protein E8L. Viruses 2022, 14, 2531. [Google Scholar] [CrossRef]
- Popik, W.; Alce, T.M.; Au, W.C. Human immunodeficiency virus type 1 uses lipid raft-colocalized CD4 and chemokine receptors for productive entry into CD4(+) T cells. J. Virol. 2002, 76, 4709–4722. [Google Scholar] [CrossRef]
- Sorice, M.; Garofalo, T.; Misasi, R.; Longo, A.; Mattei, V.; Sale, P.; Dolo, V.; Gradini, R.; Pavan, A. Evidence for cell surface association between CXCR4 and ganglioside GM3 after gp120 binding in SupT1 lymphoblastoid cells. FEBS Lett. 2001, 506, 55–60. [Google Scholar] [CrossRef]
- Nguyen, D.H.; Giri, B.; Collins, G.; Taub, D.D. Dynamic reorganization of chemokine receptors, cholesterol, lipid rafts, and adhesion molecules to sites of CD4 engagement. Exp. Cell Res. 2005, 304, 559–569. [Google Scholar] [CrossRef]
- Carter, G.C.; Bernstone, L.; Sangani, D.; Bee, J.W.; Harder, T.; James, W. HIV entry in macrophages is dependent on intact lipid rafts. Virology 2009, 386, 192–202. [Google Scholar] [CrossRef] [PubMed]
- Campbell, S.M.; Crowe, S.M.; Mak, J. Lipid rafts and HIV-1: From viral entry to assembly of progeny virions. J. Clin. Virol. 2001, 22, 217–227. [Google Scholar] [CrossRef]
- Mañes, S.; del Real, G.; Lacalle, R.A.; Lucas, P.; Gómez-Moutón, C.; Sánchez-Palomino, S.; Delgado, R.; Alcamí, J.; Mira, E.; Martínez, A.C. Membrane raft microdomains mediate lateral assemblies required for HIV-1 infection. EMBO Rep. 2000, 1, 190–196. [Google Scholar] [CrossRef] [PubMed]
- Kozak, S.L.; Heard, J.M.; Kabat, D. Segregation of CD4 and CXCR4 into distinct lipid microdomains in T lymphocytes suggests a mechanism for membrane destabilization by human immunodeficiency virus. J. Virol. 2002, 76, 1802–1815. [Google Scholar] [CrossRef]
- Fragoso, R.; Ren, D.; Zhang, X.; Su, M.W.; Burakoff, S.J.; Jin, Y.J. Lipid raft distribution of CD4 depends on its palmitoylation and association with Lck, and evidence for CD4-induced lipid raft aggregation as an additional mechanism to enhance CD3 signaling. J. Immunol. 2003, 170, 913–921. [Google Scholar] [CrossRef] [PubMed]
- Sorice, M.; Garofalo, T.; Misasi, R.; Longo, A.; Mikulak, J.; Dolo, V.; Pontieri, G.M.; Pavan, A. Association between GM3 and CD4-Ick complex in human peripheral blood lymphocytes. Glycoconj. J. 2000, 17, 247–252. [Google Scholar] [CrossRef]
- Sattentau, Q.J.; Moore, J.P. Conformational changes induced in the human immunodeficiency virus envelope glycoprotein by soluble CD4 binding. J. Exp. Med. 1991, 174, 407–415. [Google Scholar] [CrossRef]
- Pinter, A.; Honnen, W.J.; Tilley, S.A. Conformational changes affecting the V3 and CD4-binding domains of human immunodeficiency virus type 1 gp120 associated with env processing and with binding of ligands to these sites. J. Virol. 1993, 67, 5692–5697. [Google Scholar] [CrossRef] [PubMed]
- Cormier, E.G.; Dragic, T. The crown and stem of the V3 loop play distinct roles in human immunodeficiency virus type 1 envelope glycoprotein interactions with the CCR5 coreceptor. J. Virol. 2002, 76, 8953–8957. [Google Scholar] [CrossRef] [PubMed]
- Gallo, S.A.; Finnegan, C.M.; Viard, M.; Raviv, Y.; Dimitrov, A.; Rawat, S.S.; Puri, A.; Durell, S.; Blumenthal, R. The HIV Env-mediated fusion reaction. Biochim. Et Biophys. Acta (BBA)—Biomembr. 2003, 1614, 36–50. [Google Scholar] [CrossRef]
- Richman, D.D.; Bozzette, S.A. The Impact of the Syncytium-Inducing Phenotype of Human Immunodeficiency Virus on Disease Progression. J. Infect. Dis. 1994, 169, 968–974. [Google Scholar] [CrossRef] [PubMed]
- Rosen, O.; Sharon, M.; Quadt-Akabayov, S.R.; Anglister, J. Molecular switch for alternative conformations of the HIV-1 V3 region: Implications for phenotype conversion. Proc. Natl. Acad. Sci. USA 2006, 103, 13950–13955. [Google Scholar] [CrossRef]
- Kuiken, C.L.; De Jong, J.; Baan, E.; Keulen, W.; Tersmette, M.; Goudsmit, J. Evolution of the V3 envelope domain in proviral sequences and isolates of human immunodeficiency virus type 1 during transition of the viral biological phenotype. J. Virol. 1992, 66, 4622–4627. [Google Scholar] [CrossRef]
- Xiao, L.; Owen, S.M.; Goldman, I.; Lal, A.A.; deJong, J.J.; Goudsmit, J.; Lal, R.B. CCR5 Coreceptor Usage of Non-Syncytium-Inducing Primary HIV-1 Is Independent of Phylogenetically Distinct Global HIV-1 Isolates: Delineation of Consensus Motif in the V3 Domain That Predicts CCR-5 Usage. Virology 1998, 240, 83–92. [Google Scholar] [CrossRef]
- Björndal, A.; Deng, H.; Jansson, M.; Fiore, J.R.; Colognesi, C.; Karlsson, A.; Albert, J.; Scarlatti, G.; Littman, D.R.; Fenyö, E.M. Coreceptor usage of primary human immunodeficiency virus type 1 isolates varies according to biological phenotype. J. Virol. 1997, 71, 7478–7487. [Google Scholar] [CrossRef]
- Balasubramanian, C.; Chillemi, G.; Abbate, I.; Capobianchi, M.R.; Rozera, G.; Desideri, A. Importance of V3 loop flexibility and net charge in the context of co-receptor recognition. A molecular dynamics study on HIV gp120. J. Biomol. Struct. Dyn. 2012, 29, 879–891. [Google Scholar] [CrossRef]
- Repits, J.; Sterjovski, J.; Badia-Martinez, D.; Mild, M.; Gray, L.; Churchill, M.J.; Purcell, D.F.; Karlsson, A.; Albert, J.; Fenyö, E.M. Primary HIV-1 R5 isolates from end-stage disease display enhanced viral fitness in parallel with increased gp120 net charge. Virology 2008, 379, 125–134. [Google Scholar] [CrossRef] [PubMed]
- Kalinina, O.V.; Pfeifer, N.; Lengauer, T. Modelling binding between CCR5 and CXCR4 receptors and their ligands suggests the surface electrostatic potential of the co-receptor to be a key player in the HIV-1 tropism. Retrovirology 2013, 10, 130. [Google Scholar] [CrossRef] [PubMed]
- López de Victoria, A.; Kieslich, C.A.; Rizos, A.K.; Krambovitis, E.; Morikis, D. Clustering of HIV-1 subtypes based on gp120 V3 loop electrostatic properties. BMC Biophys. 2012, 5, 3. [Google Scholar] [CrossRef]
- Shioda, T.; Levy, J.A.; Cheng-Mayer, C. Small amino acid changes in the V3 hypervariable region of gp120 can affect the T-cell-line and macrophage tropism of human immunodeficiency virus type 1. Proc. Natl. Acad. Sci. USA 1992, 89, 9434–9438. [Google Scholar] [CrossRef]
- Clayton, F.; Kotler, D.P.; Kuwada, S.K.; Morgan, T.; Stepan, C.; Kuang, J.; Le, J.; Fantini, J. Gp120-induced Bob/GPR15 activation: A possible cause of human immunodeficiency virus enteropathy. Am. J. Pathol. 2001, 159, 1933–1939. [Google Scholar] [CrossRef]
- Brenneman, D.E.; Westbrook, G.L.; Fitzgerald, S.P.; Ennist, D.L.; Elkins, K.L.; Ruff, M.R.; Pert, C.B. Neuronal cell killing by the envelope protein of HIV and its prevention by vasoactive intestinal peptide. Nature 1988, 335, 639–642. [Google Scholar] [CrossRef]
- Yokoyama, M.; Naganawa, S.; Yoshimura, K.; Matsushita, S.; Sato, H. Structural dynamics of HIV-1 envelope Gp120 outer domain with V3 loop. PloS ONE 2012, 7, e37530. [Google Scholar] [CrossRef] [PubMed]
- Javaherian, K.; Langlois, A.J.; LaRosa, G.J.; Profy, A.T.; Bolognesi, D.P.; Herlihy, W.C.; Putney, S.D.; Matthews, T.J. Broadly neutralizing antibodies elicited by the hypervariable neutralizing determinant of HIV-1. Science 1990, 250, 1590–1593. [Google Scholar] [CrossRef] [PubMed]
- Lyke, K.E.; Atmar, R.L.; Islas, C.D.; Posavad, C.M.; Szydlo, D.; Paul Chourdhury, R.; Deming, M.E.; Eaton, A.; Jackson, L.A.; Branche, A.R.; et al. Rapid decline in vaccine-boosted neutralizing antibodies against SARS-CoV-2 Omicron variant. Cell Rep. Med. 2022, 3, 100679. [Google Scholar] [CrossRef] [PubMed]
- Frutos, R.; Yahi, N.; Gavotte, L.; Fantini, J.; Devaux, C.A. Role of spike compensatory mutations in the interspecies transmission of SARS-CoV-2. One Health 2022, 15, 100429. [Google Scholar] [CrossRef]
- Fantini, J.; Devaux, C.A.; Yahi, N.; Frutos, R. The novel hamster-adapted SARS-CoV-2 Delta variant may be selectively advantaged in humans. J. Infect. 2022, 84, e53–e54. [Google Scholar] [CrossRef] [PubMed]
- Hammache, D.; Piéroni, G.; Maresca, M.; Ivaldi, S.; Yahi, N.; Fantini, J. Reconstitution of sphingolipid-cholesterol plasma membrane microdomains for studies of virus-glycolipid interactions. Methods Enzymol. 2000, 312, 495–506. [Google Scholar] [PubMed]
- Di Scala, C.; Fantini, J. Hybrid in silico/in vitro approaches for the identification of functional cholesterol-binding domains in membrane proteins. In Cholesterol Homeostasis; Humana Press: New York, NY, USA, 2017; pp. 7–19. [Google Scholar]
- Fantini, J.; Yahi, N.; Garmy, N. Cholesterol accelerates the binding of Alzheimer’s β-amyloid peptide to ganglioside GM1 through a universal hydrogen-bond-dependent sterol tuning of glycolipid conformation. Front. Physiol. 2013, 4, 120. [Google Scholar] [CrossRef] [PubMed]
- Träger, J.; Widder, K.; Kerth, A.; Harauz, G.; Hinderberger, D. Effect of Cholesterol and Myelin Basic Protein (MBP) Content on Lipid Monolayers Mimicking the Cytoplasmic Membrane of Myelin. Cells 2020, 9, 529. [Google Scholar] [CrossRef]
- Barnett, B.P.; Wahlin, K.; Krawczyk, M.; Spencer, D.; Welsbie, D.; Afshari, N.; Chao, D. Potential of ocular transmission of SARS-CoV-2: A review. Vision 2020, 4, 40. [Google Scholar] [CrossRef]
- Mulay, A.; Konda, B.; Garcia, G.; Yao, C.; Beil, S.; Villalba, J.M.; Koziol, C.; Sen, C.; Purkayastha, A.; Kolls, J.K.; et al. SARS-CoV-2 infection of primary human lung epithelium for COVID-19 modeling and drug discovery. Cell Rep. 2021, 35, 109055. [Google Scholar] [CrossRef]
- Furuta, Y.; Eriksson, K.; Svennerholm, B.; Fredman, P.; Horal, P.; Jeansson, S.; Vahlne, A.; Holmgren, J.; Czerkinsky, C. Infection of vaginal and colonic epithelial cells by the human immunodeficiency virus type 1 is neutralized by antibodies raised against conserved epitopes in the envelope glycoprotein gp120. Proc. Natl. Acad. Sci. USA 1994, 91, 12559–12563. [Google Scholar] [CrossRef]
- Fantini, J.; Yahi, N.; Tourres, C.; Delézay, O.; Tamalet, C. HIV-1 transmission across the vaginal epithelium. AIDS 1997, 11, 1663–1664. [Google Scholar]
- Amerongen, H.M.; Weltzin, R.; Farnet, C.M.; Michetti, P.; Haseltine, W.A.; Neutra, M.R. Transepithelial transport of HIV-1 by intestinal M cells: A mechanism for transmission of AIDS. J. Acquir. Immune Defic. Syndr. 1991, 4, 760–765. [Google Scholar]
- Lawrence, P.; Portran, D.; Terrasse, R.; Palle, S.; Olivier, T.; Fantini, J.; Bourlet, T.; Pozzetto, B.; Delezay, O. Selective transmigration of monocyte-associated HIV-1 across a human cervical monolayer and its modulation by seminal plasma. AIDS 2012, 26, 785–796. [Google Scholar] [CrossRef]
- Kotler, D.P.; Gaetz, H.P.; Lange, M.; Klein, E.B.; Holt, P.R. Enteropathy associated with the acquired immunodeficiency syndrome. Ann. Intern. Med. 1984, 101, 421–428. [Google Scholar] [CrossRef]
- Kotler, D.P. HIV infection and the gastrointestinal tract. AIDS 2005, 19, 107–117. [Google Scholar] [CrossRef]
- Pandrea, I.; Brooks, K.; Desai, R.P.; Tare, M.; Brenchley, J.M.; Apetrei, C. I’ve looked at gut from both sides now: Gastrointestinal tract involvement in the pathogenesis of SARS-CoV-2 and HIV/SIV infections. Front. Immunol. 2022, 13, 899559. [Google Scholar] [CrossRef] [PubMed]
- Fischer, P.A.; Enzensberger, W. Neurological complications in AIDS. J. Neurol. 1987, 234, 269–279. [Google Scholar] [CrossRef] [PubMed]
- Perry, S.; Marotta, R.F. AIDS dementia: A review of the literature. Alzheimer Dis. Assoc. Disord. 1987, 1, 221–235. [Google Scholar] [CrossRef]
- Osman, I.O.; Garrec, C.; de Souza, G.A.P.; Zarubica, A.; Belhaouari, D.B.; Baudoin, J.P.; Lepidi, H.; Mege, J.L.; Malissen, B.; Scola, B.; et al. Control of CDH1/E-Cadherin Gene Expression and Release of a Soluble Form of E-Cadherin in SARS-CoV-2 Infected Caco-2 Intestinal Cells: Physiopathological Consequences for the Intestinal Forms of COVID-19. Front. Cell. Infect. Microbiol. 2022, 12, 798767. [Google Scholar] [CrossRef] [PubMed]
- Engin, A.B.; Engin, E.D.; Engin, A. Dual function of sialic acid in gastrointestinal SARS-CoV-2 infection. Environ. Toxicol. Pharmacol. 2020, 79, 103436. [Google Scholar] [CrossRef]
- Pistollato, F.; Petrillo, M.; Clerbaux, L.A.; Leoni, G.; Ponti, J.; Bogni, A.; Brogna, C.; Cristoni, S.; Sanges, R.; Mendoza-de Gyves, E.; et al. Effects of spike protein and toxin-like peptides found in COVID-19 patients on human 3D neuronal/glial model undergoing differentiation: Possible implications for SARS-CoV-2 impact on brain development. Reprod. Toxicol. 2022, 111, 34–48. [Google Scholar] [CrossRef]
- Kim, E.S.; Jeon, M.-T.; Kim, K.-S.; Lee, S.; Kim, S.; Kim, D.-G. Spike Proteins of SARS-CoV-2 Induce Pathological Changes in Molecular Delivery and Metabolic Function in the Brain Endothelial Cells. Viruses 2021, 13, 2021. [Google Scholar] [CrossRef]
- Buzhdygan, T.P.; DeOre, B.J.; Baldwin-Leclair, A.; Bullock, T.A.; McGary, H.M.; Khan, J.A.; Razmpour, R.; Hale, J.F.; Galie, P.A.; Potula, R.; et al. The SARS-CoV-2 spike protein alters barrier function in 2D static and 3D microfluidic in-vitro models of the human blood–brain barrier. Neurobiol. Dis. 2020, 146, 105131. [Google Scholar] [CrossRef]
- El-Arif, G.; Farhat, A.; Khazaal, S.; Annweiler, C.; Kovacic, H.; Wu, Y.; Cao, Z.; Fajloun, Z.; Khattar, Z.A.; Sabatier, J.M. The Renin-Angiotensin System: A Key Role in SARS-CoV-2-Induced COVID-19. Molecules 2021, 26, 6945. [Google Scholar] [CrossRef] [PubMed]
- Kumar, N.; Zuo, Y.; Yalavarthi, S.; Hunker, K.L.; Knight, J.S.; Kanthi, Y.; Obi, A.T.; Ganesh, S.K. SARS-CoV-2 Spike Protein S1-Mediated Endothelial Injury and Pro-Inflammatory State Is Amplified by Dihydrotestosterone and Prevented by Mineralocorticoid Antagonism. Viruses 2021, 13, 2209. [Google Scholar] [CrossRef]
- DeOre, B.J.; Tran, K.A.; Andrews, A.M.; Ramirez, S.H.; Galie, P.A. SARS-CoV-2 spike protein disrupts blood–brain barrier integrity via RhoA activation. J. Neuroimmune Pharmacol. 2021, 16, 722–728. [Google Scholar] [CrossRef]
- Rhea, E.M.; Logsdon, A.F.; Hansen, K.M.; Williams, L.M.; Reed, M.J.; Baumann, K.K.; Holden, S.J.; Raber, J.; Banks, W.A.; Erickson, M.A. The S1 protein of SARS-CoV-2 crosses the blood–brain barrier in mice. Nat. Neurosci. 2021, 24, 368–378. [Google Scholar] [CrossRef] [PubMed]
- Kanmogne, G.D.; Primeaux, C.; Grammas, P. HIV-1 gp120 proteins alter tight junction protein expression and brain endothelial cell permeability: Implications for the pathogenesis of HIV-associated dementia. J. Neuropathol. Exp. Neurol. 2005, 64, 498–505. [Google Scholar] [CrossRef] [PubMed]
- Kanmogne, G.D.; Kennedy, R.C.; Grammas, P. HIV-1 gp120 Proteins and gp160 Peptides Are Toxic to Brain Endothelial Cells and Neurons: Possible Pathway for HIV Entry into the Brain and HIV-Associated Dementia. J. Neuropathol. Exp. Neurol. 2002, 61, 992–1000. [Google Scholar] [CrossRef]
- Bansal, A.K.; Mactutus, C.F.; Nath, A.; Maragos, W.; Hauser, K.F.; Booze, R.M. Neurotoxicity of HIV-1 proteins gp120 and Tat in the rat striatum. Brain Res. 2000, 879, 42–49. [Google Scholar] [CrossRef]
- Alshammary, A.F.; Al-Sulaiman, A.M. The journey of SARS-CoV-2 in human hosts: A review of immune responses, immunosuppression, and their consequences. Virulence 2021, 12, 1771–1794. [Google Scholar] [CrossRef] [PubMed]
- Reizine, F.; Lesouhaitier, M.; Gregoire, M.; Pinceaux, K.; Gacouin, A.; Maamar, A.; Painvin, B.; Camus, C.; Le Tulzo, Y.; Tattevin, P.; et al. SARS-CoV-2-Induced ARDS Associates with MDSC Expansion, Lymphocyte Dysfunction, and Arginine Shortage. J. Clin. Immunol. 2021, 41, 515–525. [Google Scholar] [CrossRef]
- De Martini, R.M.; Parker, J.W. Immunologic alterations in human immunodeficiency virus infection: A review. J. Clin. Lab. Anal. 1989, 3, 56–70. [Google Scholar] [CrossRef]
- Zeberg, H.; Pääbo, S. The major genetic risk factor for severe COVID-19 is inherited from Neanderthals. Nature 2020, 587, 610–612. [Google Scholar] [CrossRef]
- Zeberg, H. The major genetic risk factor for severe COVID-19 is associated with protection against HIV. Proc. Natl. Acad. Sci. USA 2022, 119, e2116435119. [Google Scholar] [CrossRef] [PubMed]
- Simons, K.; Ehehalt, R. Cholesterol, lipid rafts, and disease. J. Clin. Investig. 2002, 110, 597–603. [Google Scholar] [CrossRef]
- Ripa, I.; Andreu, S.; López-Guerrero, J.A.; Bello-Morales, R. Membrane rafts: Portals for viral entry. Front. Microbiol. 2021, 12, 631274. [Google Scholar] [CrossRef]
- Schneider-Schaulies, J.; Schneider-Schaulies, S. Sphingolipids in viral infection. Biol. Chem. 2015, 396, 585–595. [Google Scholar] [CrossRef] [PubMed]
- Chazal, N.; Gerlier, D. Virus entry, assembly, budding, and membrane rafts. Microbiol. Mol. Biol. Rev. 2003, 67, 226–237. [Google Scholar] [CrossRef] [PubMed]
- Mañes, S.; del Real, G.; Martínez, A.C. Pathogens: Raft hijackers. Nat. Reviews. Immunol. 2003, 3, 557–568. [Google Scholar] [CrossRef]
- Hanada, K. Sphingolipids in infectious diseases. Jpn. J. Infect. Dis. 2005, 58, 131–148. [Google Scholar]
- Yager, E.J.; Konan, K.V. Sphingolipids as Potential Therapeutic Targets against Enveloped Human RNA Viruses. Viruses 2019, 11, 912. [Google Scholar] [CrossRef]
- Schneider-Schaulies, S.; Schumacher, F.; Wigger, D.; Schöl, M.; Waghmare, T.; Schlegel, J.; Seibel, J.; Kleuser, B. Sphingolipids: Effectors and Achilles Heals in Viral Infections? Cells 2021, 10, 2175. [Google Scholar] [CrossRef]
- Fantini, J.; Tamalet, C.; Hammache, D.; Tourrès, C.; Duclos, N.; Yahi, N. HIV-1-induced perturbations of glycosphingolipid metabolism are cell-specific and can be detected at early stages of HIV-1 infection. J. Acquir. Immune Defic. Syndr. Hum. Retrovirology Off. Publ. Int. Retrovirology Assoc. 1998, 19, 221–229. [Google Scholar] [CrossRef]
- Rawat, S.S.; Viard, M.; Gallo, S.A.; Rein, A.; Blumenthal, R.; Puri, A. Modulation of entry of enveloped viruses by cholesterol and sphingolipids. Mol. Membr. Biol. 2003, 20, 243–254. [Google Scholar] [CrossRef]
- Di Scala, C.; Armstrong, N.; Chahinian, H.; Chabrière, E.; Fantini, J.; Yahi, N. AmyP53, a Therapeutic Peptide Candidate for the Treatment of Alzheimer’s and Parkinson’s Disease: Safety, Stability, Pharmacokinetics Parameters and Nose-to Brain Delivery. Int. J. Mol. Sci. 2022, 23, 13383. [Google Scholar] [CrossRef] [PubMed]
- Di Scala, C.; Yahi, N.; Boutemeur, S.; Flores, A.; Rodriguez, L.; Chahinian, H.; Fantini, J. Common molecular mechanism of amyloid pore formation by Alzheimer’s β-amyloid peptide and α-synuclein. Sci. Rep. 2016, 6, 28781. [Google Scholar] [CrossRef] [PubMed]
- Di Scala, C.; Yahi, N.; Flores, A.; Boutemeur, S.; Kourdougli, N.; Chahinian, H.; Fantini, J. Broad neutralization of calcium-permeable amyloid pore channels with a chimeric Alzheimer/Parkinson peptide targeting brain gangliosides. Biochim. Et Biophys. Acta 2016, 1862, 213–222. [Google Scholar] [CrossRef]
- Fantini, J.; Chahinian, H.; Yahi, N. Progress toward Alzheimer’s disease treatment: Leveraging the Achilles’ heel of Aβ oligomers? Protein Sci. A Publ. Protein Soc. 2020, 29, 1748–1759. [Google Scholar] [CrossRef]
- Yahi, N.; Di Scala, C.; Chahinian, H.; Fantini, J. Innovative treatment targeting gangliosides aimed at blocking the formation of neurotoxic α-synuclein oligomers in Parkinson’s disease. Glycoconj. J. 2022, 39, 1–11. [Google Scholar] [CrossRef]
- Mohamed, A.; Robinson, H.; Erramouspe, P.J.; Hill, M.M. Advances and challenges in understanding the role of the lipid raft proteome in human health. Expert Rev. Proteom. 2018, 15, 1053–1063. [Google Scholar] [CrossRef] [PubMed]
- Alipoor, S.D.; Mirsaeidi, M. SARS-CoV-2 cell entry beyond the ACE2 receptor. Mol. Biol. Rep. 2022, 49, 10715–10727. [Google Scholar] [CrossRef]
- Grass, G.D.; Bratoeva, M.; Toole, B.P. Regulation of invadopodia formation and activity by CD147. J. Cell Sci. 2012, 125, 777–788. [Google Scholar] [CrossRef]
- Wang, H.-B.; Zhang, H.; Zhang, J.-P.; Li, Y.; Zhao, B.; Feng, G.-K.; Du, Y.; Xiong, D.; Zhong, Q.; Liu, W.-L.; et al. Neuropilin 1 is an entry factor that promotes EBV infection of nasopharyngeal epithelial cells. Nat. Commun. 2015, 6, 6240. [Google Scholar] [CrossRef] [PubMed]
- Ohnuma, K.; Takahashi, N.; Yamochi, T.; Hosono, O.; Dang, N.H.; Morimoto, C. Role of CD26/dipeptidyl peptidase IV in human T cell activation and function. Front. Biosci.-Landmark 2008, 13, 2299–2310. [Google Scholar] [CrossRef] [PubMed]
- Geijtenbeek, T.B.; Kwon, D.S.; Torensma, R.; van Vliet, S.J.; van Duijnhoven, G.C.; Middel, J.; Cornelissen, I.L.; Nottet, H.S.; KewalRamani, V.N.; Littman, D.R.; et al. DC-SIGN, a dendritic cell-specific HIV-1-binding protein that enhances trans-infection of T cells. Cell 2000, 100, 587–597. [Google Scholar] [CrossRef]
- Tilghman, R.W.; Hoover, R.L. E-selectin and ICAM-1 are incorporated into detergent-insoluble membrane domains following clustering in endothelial cells. FEBS Lett. 2002, 525, 83–87. [Google Scholar] [CrossRef]
- Cambi, A.; de Lange, F.; van Maarseveen, N.M.; Nijhuis, M.; Joosten, B.; van Dijk, E.M.; de Bakker, B.R.I.; Fransen, J.A.; Bovee-Geurts, P.H.; van Leeuwen, F.N. Microdomains of the C-type lectin DC-SIGN are portals for virus entry into dendritic cells. J. Cell Biol. 2004, 164, 145–155. [Google Scholar] [CrossRef]
- Serafin, M.B.; Bottega, A.; Foletto, V.S.; da Rosa, T.F.; Hörner, A.; Hörner, R. Drug repositioning is an alternative for the treatment of coronavirus COVID-19. Int. J. Antimicrob. Agents 2020, 55, 105969. [Google Scholar] [CrossRef] [PubMed]
- Gautret, P.; Lagier, J.-C.; Parola, P.; Meddeb, L.; Mailhe, M.; Doudier, B.; Courjon, J.; Giordanengo, V.; Vieira, V.E.; Dupont, H.T. Hydroxychloroquine and azithromycin as a treatment of COVID-19: Results of an open-label non-randomized clinical trial. Int. J. Antimicrob. Agents 2020, 56, 105949. [Google Scholar] [CrossRef]
- Lagier, J.-C.; Million, M.; Gautret, P.; Colson, P.; Cortaredona, S.; Giraud-Gatineau, A.; Honoré, S.; Gaubert, J.-Y.; Fournier, P.-E.; Tissot-Dupont, H. Outcomes of 3,737 COVID-19 patients treated with hydroxychloroquine/azithromycin and other regimens in Marseille, France: A retrospective analysis. Travel Med. Infect. Dis. 2020, 36, 101791. [Google Scholar] [CrossRef]
- Adalja, A.; Inglesby, T. Broad-Spectrum Antiviral Agents: A Crucial Pandemic Tool. Expert Rev. Anti-Infect. Ther. 2019, 17, 467–470. [Google Scholar] [CrossRef] [PubMed]
- Khoshnood, S.; Shirani, M.; Dalir, A.; Moradi, M.; Haddadi, M.H.; Sadeghifard, N.; Birjandi, F.S.; Yashmi, I.; Heidary, M. Antiviral effects of azithromycin: A narrative review. Biomed. Pharmacother. 2022, 147, 112682. [Google Scholar] [CrossRef]
- Fantini, J.; Hammache, D.; Delézay, O.; Yahi, N.; André-Barrès, C.; Rico-Lattes, I.; Lattes, A. Synthetic soluble analogs of galactosylceramide (GalCer) bind to the V3 domain of HIV-1 gp120 and inhibit HIV-1-induced fusion and entry. J. Biol. Chem. 1997, 272, 7245–7252. [Google Scholar] [CrossRef]
- Mahfoud, R.; Mylvaganam, M.; Lingwood, C.A.; Fantini, J. A novel soluble analog of the HIV-1 fusion cofactor, globotriaosylceramide (Gb(3)), eliminates the cholesterol requirement for high affinity gp120/Gb(3) interaction. J. Lipid Res. 2002, 43, 1670–1679. [Google Scholar] [CrossRef] [PubMed]
- Lund, N.; Branch, D.R.; Mylvaganam, M.; Chark, D.; Ma, X.Z.; Sakac, D.; Binnington, B.; Fantini, J.; Puri, A.; Blumenthal, R.; et al. A novel soluble mimic of the glycolipid, globotriaosyl ceramide inhibits HIV infection. AIDS 2006, 20, 333–343. [Google Scholar] [CrossRef]
- Garg, H.; Francella, N.; Tony, K.A.; Augustine, L.A.; Barchi, J.J., Jr.; Fantini, J.; Puri, A.; Mootoo, D.R.; Blumenthal, R. Glycoside analogs of beta-galactosylceramide, a novel class of small molecule antiviral agents that inhibit HIV-1 entry. Antivir. Res. 2008, 80, 54–61. [Google Scholar] [CrossRef]
- Weber, K.T.; Hammache, D.; Fantini, J.; Ganem, B. Synthesis of glycolipid analogues that disrupt binding of HIV-1 gp120 to galactosylceramide. Bioorganic Med. Chem. Lett. 2000, 10, 1011–1014. [Google Scholar] [CrossRef] [PubMed]
- Fantini, J. Interaction of proteins with lipid rafts through glycolipid-binding domains: Biochemical background and potential therapeutic applications. Curr. Med. Chem. 2007, 14, 2911–2917. [Google Scholar] [CrossRef] [PubMed]
- Fantini, J. [49] Synthetic soluble analogs of glycolipids for studies of virus-glycolipid interactions. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 2000; Volume 311, pp. 626–638. [Google Scholar]
- Richichi, B.; Pastori, C.; Gherardi, S.; Venuti, A.; Cerreto, A.; Sanvito, F.; Toma, L.; Lopalco, L.; Nativi, C. GM-3 Lactone Mimetic Interacts with CD4 and HIV-1 Env Proteins, Hampering HIV-1 Infection without Inducing a Histopathological Alteration. ACS Infect. Dis. 2016, 2, 564–571. [Google Scholar] [CrossRef]
- Matsubara, T.; Sumi, M.; Kubota, H.; Taki, T.; Okahata, Y.; Sato, T. Inhibition of Influenza Virus Infections by Sialylgalactose-Binding Peptides Selected from a Phage Library. J. Med. Chem. 2009, 52, 4247–4256. [Google Scholar] [CrossRef]
- Delézay, O.; Hammache, D.; Fantini, J.; Yahi, N. SPC3, a V3 loop-derived synthetic peptide inhibitor of HIV-1 infection, binds to cell surface glycosphingolipids. Biochemistry 1996, 35, 15663–15671. [Google Scholar] [CrossRef] [PubMed]
- Nehete, P.N.; Vela, E.M.; Hossain, M.M.; Sarkar, A.K.; Yahi, N.; Fantini, J.; Sastry, K.J. A post-CD4-binding step involving interaction of the V3 region of viral gp120 with host cell surface glycosphingolipids is common to entry and infection by diverse HIV-1 strains. Antivir. Res. 2002, 56, 233–251. [Google Scholar] [CrossRef] [PubMed]
- Warren, C.J.; Yu, S.; Peters, D.K.; Barbachano-Guerrero, A.; Yang, Q.; Burris, B.L.; Worwa, G.; Huang, I.C.; Wilkerson, G.K.; Goldberg, T.L.; et al. Primate hemorrhagic fever-causing arteriviruses are poised for spillover to humans. Cell 2022, 185, 3980–3991.e3918. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Zhang, Q.; Tang, J.; Feng, W.-H. Lipid rafts both in cellular membrane and viral envelope are critical for PRRSV efficient infection. Virology 2015, 484, 170–180. [Google Scholar] [CrossRef] [PubMed]




| Characteristic | HIV-1 | SARS-CoV-2 |
|---|---|---|
| Genome | RNA 10 kb | RNA 30 kb |
| Entry into host cells and propagation | Lipid raft mediated fusion or endocytosis, cell-cell fusion | Lipid raft mediated fusion or endocytosis, cell-cell fusion |
| Main receptor | CD4 | ACE2 |
| Alternative receptor | DC-sign, galactosylceramide | CD147, neuropilin, CD26 |
| Receptor binding protein | Gp120 | Spike protein S1 |
| Fusion domain | Gp41 | Spike protein S2 |
| Raft cofactor | GM3, Gb3, cholesterol | GM1, cholesterol |
| Raft binding domain | V3 loop | Spike protein NTD |
| Coreceptor | CCR5, CXCR4, GPR15 | TMPRSS2 |
| Mucosal site of infection | Vaginal or rectal mucosa | Respiratory mucosa |
| Target cells | Lymphocytes Macrophages Intestinal cells Neural cells | Respiratory epithelial cells Intestinal epithelial cells Lymphocytes Macrophages Neural cells Ocular cells |
| Virotoxin | Gp120 | Spike protein |
| Characteristic | HIV-1 | SARS-CoV-2 |
|---|---|---|
| Animal reservoir | Non-human primates | Bats, minks, hamsters, domestic animals, zoo animals (non-human primates), wild animals |
| Low fidelity enzyme | Reverse transcriptase (RT) | RNA polymerase (RdRp) |
| Genetic variability | Quasi-species, subtypes | Quasi-species, variants, recombinants |
| Mutational pattern | Mutations, insertions, deletions, recombinants | Mutations, insertions, deletions, recombinants |
| Selection pressure | Antiretroviral drugs, immunity | Immunity, vaccines |
| Pathology | lymphopenia Gastrointestinal dysfunction Microbiome alteration Neurological disorders Thrombosis Proinflammatory cytokines synthesis | lymphopenia Gastrointestinal dysfunction Microbiome alteration Neurological disorders Thrombosis Proinflammatory cytokines synthesis |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fantini, J.; Chahinian, H.; Yahi, N. Convergent Evolution Dynamics of SARS-CoV-2 and HIV Surface Envelope Glycoproteins Driven by Host Cell Surface Receptors and Lipid Rafts: Lessons for the Future. Int. J. Mol. Sci. 2023, 24, 1923. https://doi.org/10.3390/ijms24031923
Fantini J, Chahinian H, Yahi N. Convergent Evolution Dynamics of SARS-CoV-2 and HIV Surface Envelope Glycoproteins Driven by Host Cell Surface Receptors and Lipid Rafts: Lessons for the Future. International Journal of Molecular Sciences. 2023; 24(3):1923. https://doi.org/10.3390/ijms24031923
Chicago/Turabian StyleFantini, Jacques, Henri Chahinian, and Nouara Yahi. 2023. "Convergent Evolution Dynamics of SARS-CoV-2 and HIV Surface Envelope Glycoproteins Driven by Host Cell Surface Receptors and Lipid Rafts: Lessons for the Future" International Journal of Molecular Sciences 24, no. 3: 1923. https://doi.org/10.3390/ijms24031923
APA StyleFantini, J., Chahinian, H., & Yahi, N. (2023). Convergent Evolution Dynamics of SARS-CoV-2 and HIV Surface Envelope Glycoproteins Driven by Host Cell Surface Receptors and Lipid Rafts: Lessons for the Future. International Journal of Molecular Sciences, 24(3), 1923. https://doi.org/10.3390/ijms24031923







