MiRNA-21a, miRNA-145, and miRNA-221 Expression and Their Correlations with WNT Proteins in Patients with Obstructive and Non-Obstructive Coronary Artery Disease
Abstract
:1. Introduction
2. Results
2.1. Basic Clinical Characteristics
2.2. Concentration of WNT Proteins in Plasma
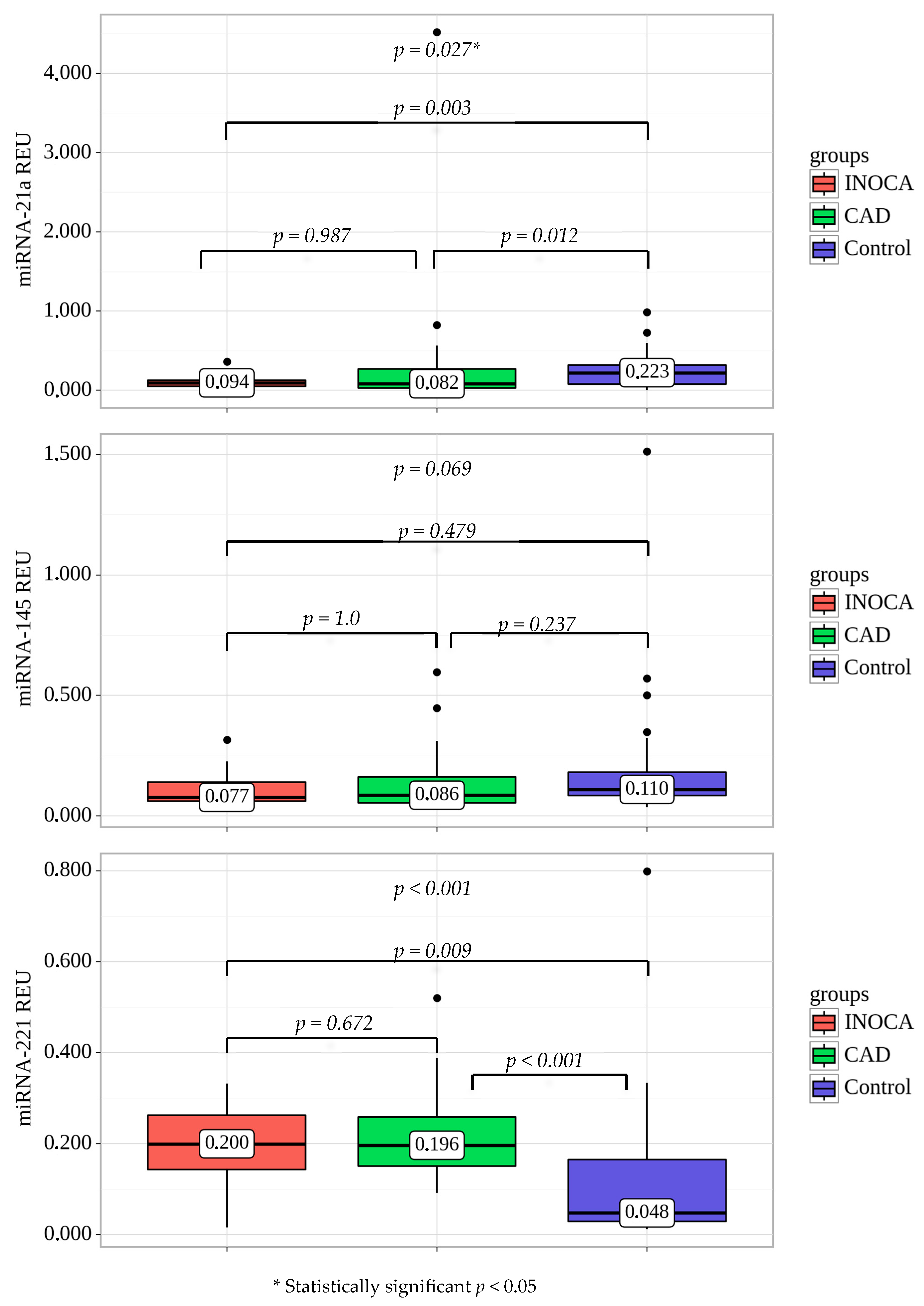
2.3. MiRNA Expression in Plasma of Patients with CAD
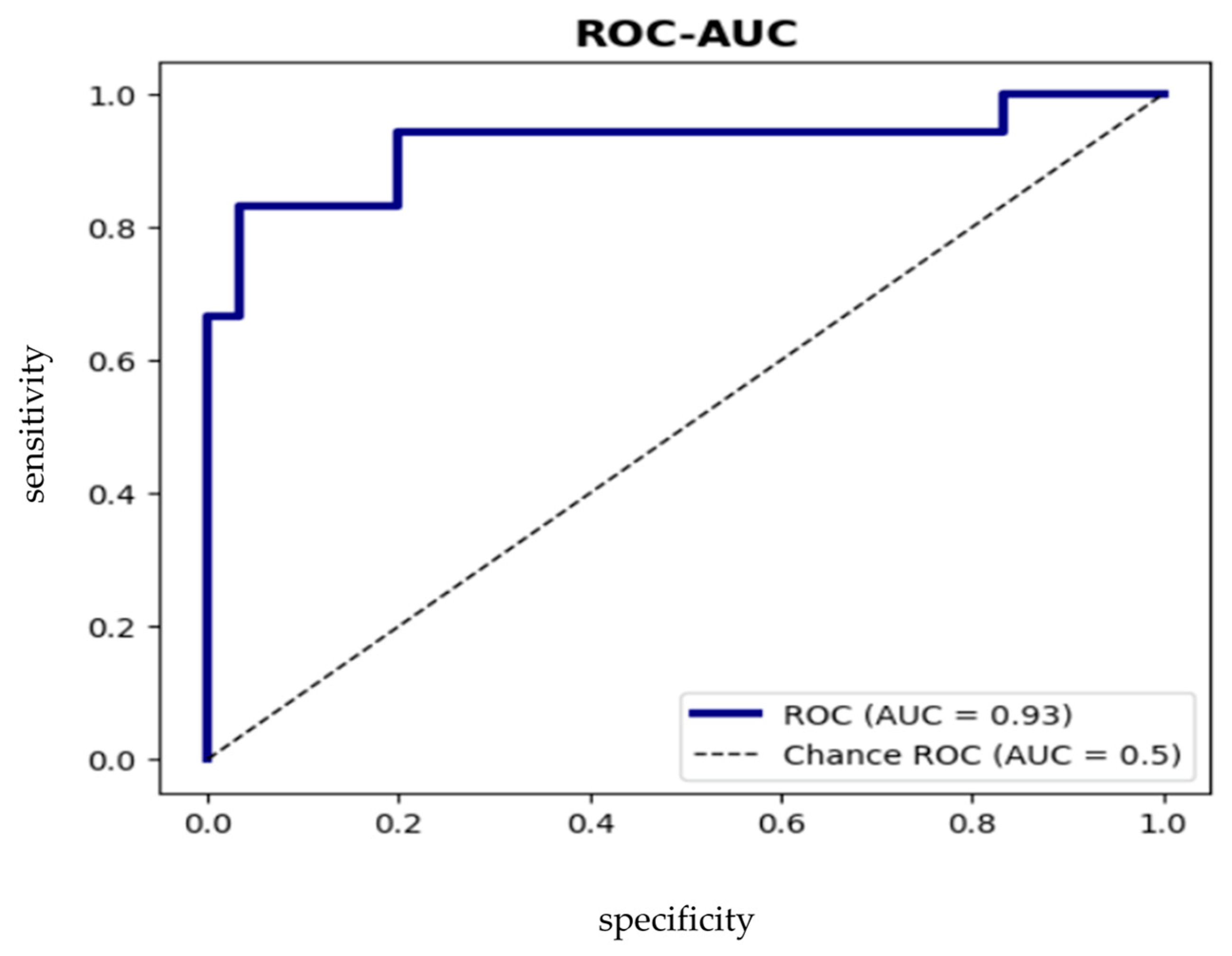
2.4. Correlations of WNT Proteins with Circulating miRNAs
3. Discussion
4. Materials and Methods
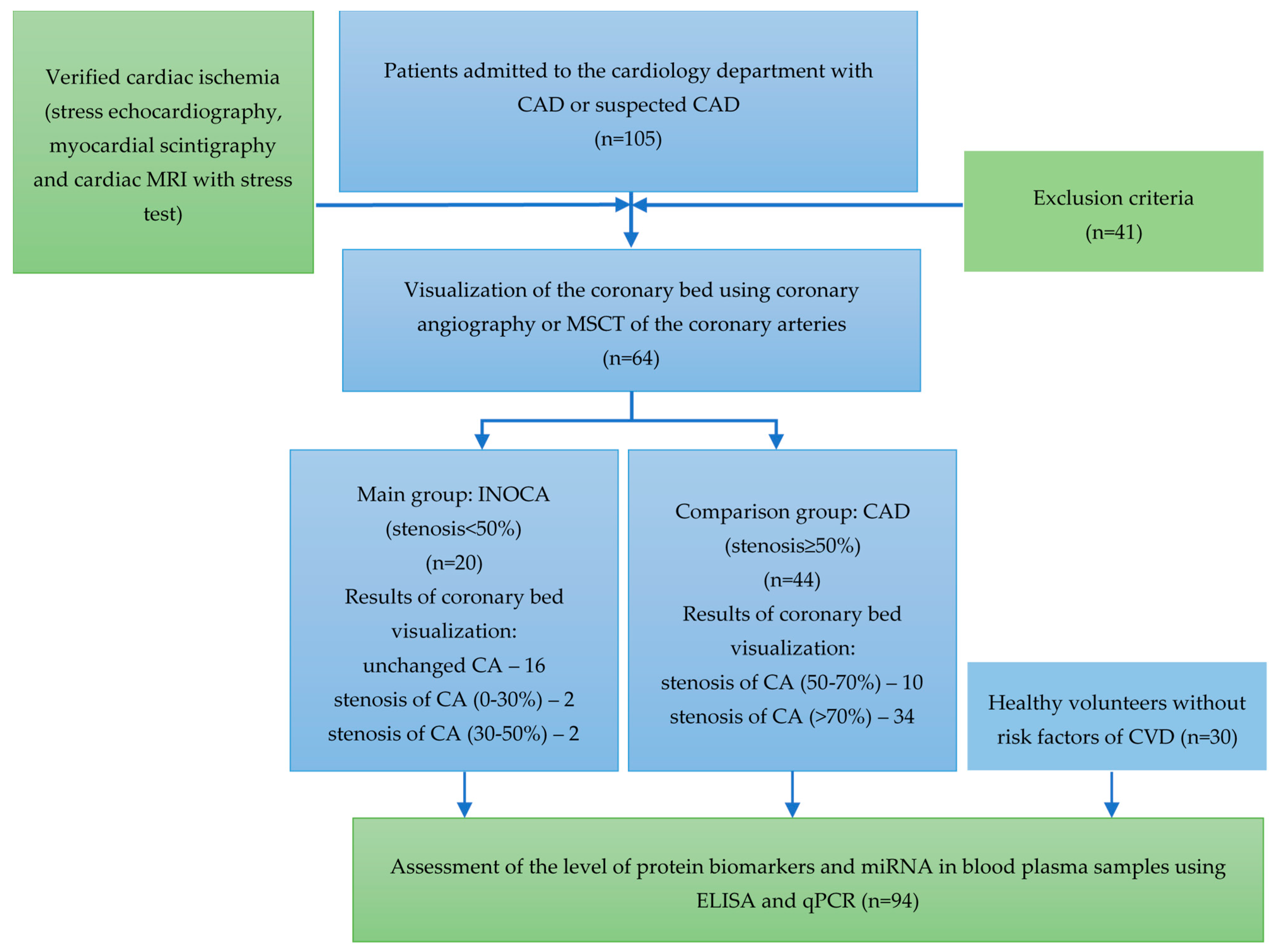
4.1. Patient Population
4.2. Collection of Blood Samples and ELISA
4.3. RNA Extraction and Reverse Transcription-Polymerase Chain Reaction (RT-PCR) Assay
4.4. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- The Top 10 Causes of Death. Available online: https://www.who.int/news-room/fact-sheets/detail/the-top-10-causes-of-death (accessed on 8 December 2023).
- Tsao, C.W.; Aday, A.W.; Almarzooq, Z.I.; Anderson, C.A.M.; Arora, P.; Avery, C.L.; Baker-Smith, C.M.; Beaton, A.Z.; Boehme, A.K.; Buxton, A.E.; et al. Heart Disease and Stroke Statistics-2023 Update: A Report From the American Heart Association. Circulation 2023, 147, e93–e621. [Google Scholar] [CrossRef] [PubMed]
- Gevaert, A.B.; Wood, N.; Boen, J.R.A.; Davos, C.H.; Hansen, D.; Hanssen, H.; Krenning, G.; Moholdt, T.; Osto, E.; Paneni, F.; et al. Epigenetics in the Primary and Secondary Prevention of Cardiovascular Disease: Influence of Exercise and Nutrition. Eur. J. Prev. Cardiol. 2022, 29, 2183–2199. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Kamato, D.; Little, P.J.; Nakagawa, S.; Pelisek, J.; Jin, Z.G. Targeting Epigenetics and Non-Coding RNAs in Atherosclerosis: From Mechanisms to Therapeutics. Pharmacol. Ther. 2019, 196, 15–43. [Google Scholar] [CrossRef] [PubMed]
- Singla, B.; Lin, H.-P.; Chen, A.; Ahn, W.; Ghoshal, P.; Cherian-Shaw, M.; White, J.; Stansfield, B.K.; Csányi, G. Role of R-Spondin 2 in Arterial Lymphangiogenesis and Atherosclerosis. Cardiovasc. Res. 2021, 117, 1489–1509. [Google Scholar] [CrossRef] [PubMed]
- Rashdan, N.A.; Sim, A.M.; Cui, L.; Phadwal, K.; Roberts, F.L.; Carter, R.; Ozdemir, D.D.; Hohenstein, P.; Hung, J.; Kaczynski, J.; et al. Osteocalcin Regulates Arterial Calcification Via Altered Wnt Signaling and Glucose Metabolism. J. Bone Miner. Res. 2020, 35, 357–367. [Google Scholar] [CrossRef] [PubMed]
- Weerackoon, N.; Gunawardhana, K.L.; Mani, A. Wnt Signaling Cascades and Their Role in Coronary Artery Health and Disease. J. Cell. Signal. 2021, 2, 52–62. [Google Scholar] [CrossRef] [PubMed]
- Manolagas, S.C.; Almeida, M. Gone with the Wnts: Beta-Catenin, T-Cell Factor, Forkhead Box O, and Oxidative Stress in Age-Dependent Diseases of Bone, Lipid, and Glucose Metabolism. Mol. Endocrinol. 2007, 21, 2605–2614. [Google Scholar] [CrossRef] [PubMed]
- Anthony, C.C.; Robbins, D.J.; Ahmed, Y.; Lee, E. Nuclear Regulation of Wnt/β-Catenin Signaling: It’s a Complex Situation. Genes 2020, 11, 886. [Google Scholar] [CrossRef]
- Liu, J.; Xiao, Q.; Xiao, J.; Niu, C.; Li, Y.; Zhang, X.; Zhou, Z.; Shu, G.; Yin, G. Wnt/β-Catenin Signalling: Function, Biological Mechanisms, and Therapeutic Opportunities. Signal Transduct. Target. Ther. 2022, 7, 3. [Google Scholar] [CrossRef]
- Yousefi, F.; Shabaninejad, Z.; Vakili, S.; Derakhshan, M.; Movahedpour, A.; Dabiri, H.; Ghasemi, Y.; Mahjoubin-Tehran, M.; Nikoozadeh, A.; Savardashtaki, A.; et al. TGF-β and WNT Signaling Pathways in Cardiac Fibrosis: Non-Coding RNAs Come into Focus. Cell Commun. Signal. 2020, 18, 87. [Google Scholar] [CrossRef]
- Foulquier, S.; Daskalopoulos, E.P.; Lluri, G.; Hermans, K.C.M.; Deb, A.; Blankesteijn, W.M. WNT Signaling in Cardiac and Vascular Disease. Pharmacol. Rev. 2018, 70, 68–141. [Google Scholar] [CrossRef] [PubMed]
- Duan, J.; Gherghe, C.; Liu, D.; Hamlett, E.; Srikantha, L.; Rodgers, L.; Regan, J.N.; Rojas, M.; Willis, M.; Leask, A.; et al. Wnt1/Βcatenin Injury Response Activates the Epicardium and Cardiac Fibroblasts to Promote Cardiac Repair. EMBO J. 2012, 31, 429–442. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.; Hu, R.; Zhu, N.; Yao, H.-L.; Lei, X.-Y.; Li, S.-X.; Liao, D.-F.; Zheng, X.-L. The Novel Role and Underlying Mechanism of Wnt5a in Regulating Cellular Cholesterol Accumulation. Clin. Exp. Pharmacol. Physiol. 2014, 41, 671–678. [Google Scholar] [CrossRef]
- Skaria, T.; Bachli, E.; Schoedon, G. Wnt5A/Ryk Signaling Critically Affects Barrier Function in Human Vascular Endothelial Cells. Cell Adhes. Migr. 2017, 11, 24–38. [Google Scholar] [CrossRef] [PubMed]
- Mill, C.; Monk, B.A.; Williams, H.; Simmonds, S.J.; Jeremy, J.Y.; Johnson, J.L.; George, S.J. Wnt5a-Induced Wnt1-Inducible Secreted Protein-1 Suppresses Vascular Smooth Muscle Cell Apoptosis Induced by Oxidative Stress. Arterioscler. Thromb. Vasc. Biol. 2014, 34, 2449–2456. [Google Scholar] [CrossRef] [PubMed]
- Schaale, K.; Neumann, J.; Schneider, D.; Ehlers, S.; Reiling, N. Wnt Signaling in Macrophages: Augmenting and Inhibiting Mycobacteria-Induced Inflammatory Responses. Eur. J. Cell Biol. 2011, 90, 553–559. [Google Scholar] [CrossRef] [PubMed]
- Masckauchán, T.N.H.; Agalliu, D.; Vorontchikhina, M.; Ahn, A.; Parmalee, N.L.; Li, C.-M.; Khoo, A.; Tycko, B.; Brown, A.M.C.; Kitajewski, J. Wnt5a Signaling Induces Proliferation and Survival of Endothelial Cells in Vitro and Expression of MMP-1 and Tie-2. Mol. Biol. Cell 2006, 17, 5163–5172. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.; Yeh, J.; Fan, T.-P.; Smith, S.K.; Charnock-Jones, D.S. Wnt5a-Mediated Non-Canonical Wnt Signalling Regulates Human Endothelial Cell Proliferation and Migration. Biochem. Biophys. Res. Commun. 2008, 365, 285–290. [Google Scholar] [CrossRef]
- Go, G. Low-Density Lipoprotein Receptor-Related Protein 6 (LRP6) Is a Novel Nutritional Therapeutic Target for Hyperlipidemia, Non-Alcoholic Fatty Liver Disease, and Atherosclerosis. Nutrients 2015, 7, 4453–4464. [Google Scholar] [CrossRef]
- Jeong, W.; Jho, E.-H. Regulation of the Low-Density Lipoprotein Receptor-Related Protein LRP6 and Its Association With Disease: Wnt/β-Catenin Signaling and Beyond. Front. Cell Dev. Biol. 2021, 9, 714330. [Google Scholar] [CrossRef]
- Mineo, C. Lipoprotein Receptor Signalling in Atherosclerosis. Cardiovasc. Res. 2020, 116, 1254–1274. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Atanasov, A.G. The MicroRNAs Regulating Vascular Smooth Muscle Cell Proliferation: A Minireview. Int. J. Mol. Sci. 2019, 20, 324. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Shi, L.; Huang, Y.; Shen, L.; Peng, H.; Zhu, X.; Zhou, G. Nanoparticle Delivery of Wnt-1 SiRNA Enhances Photodynamic Therapy by Inhibiting Epithelial-Mesenchymal Transition for Oral Cancer. Biomater. Sci. 2017, 5, 494–501. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Yang, B.; Yang, H.; Wang, L.; Li, H.; Chen, S.; Lu, X.; Gu, D. MicroRNA-320b Modulates Cholesterol Efflux and Atherosclerosis. J. Atheroscler. Thromb. 2022, 29, 200–220. [Google Scholar] [CrossRef] [PubMed]
- Canfrán-Duque, A.; Rotllan, N.; Zhang, X.; Fernández-Fuertes, M.; Ramírez-Hidalgo, C.; Araldi, E.; Daimiel, L.; Busto, R.; Fernández-Hernando, C.; Suárez, Y. Macrophage Deficiency of MiR-21 Promotes Apoptosis, Plaque Necrosis, and Vascular Inflammation during Atherogenesis. EMBO Mol. Med. 2017, 9, 1244–1262. [Google Scholar] [CrossRef]
- Zhou, J.; Wang, K.-C.; Wu, W.; Subramaniam, S.; Shyy, J.Y.-J.; Chiu, J.-J.; Li, J.Y.-S.; Chien, S. MicroRNA-21 Targets Peroxisome Proliferators-Activated Receptor-Alpha in an Autoregulatory Loop to Modulate Flow-Induced Endothelial Inflammation. Proc. Natl. Acad. Sci. USA 2011, 108, 10355–10360. [Google Scholar] [CrossRef]
- Noone, S.; Schubert, R.; Fichtlscherer, S.; Hilberg, T.; Alesci, S.; Miesbach, W.; Klophaus, N.; Wehmeier, U.F. Endothelial Dysfunction and Atherosclerosis Related MiRNA-Expression in Patients with Haemophilia. Haemophilia 2023, 29, 61–71. [Google Scholar] [CrossRef]
- Sharma, A.R.; Sharma, G.; Bhattacharya, M.; Lee, S.-S.; Chakraborty, C. Circulating MiRNA in Atherosclerosis: A Clinical Biomarker and Early Diagnostic Tool. Curr. Mol. Med. 2022, 22, 250–262. [Google Scholar] [CrossRef]
- Ward, Z.; Schmeier, S.; Pearson, J.; Cameron, V.A.; Frampton, C.M.; Troughton, R.W.; Doughty, R.N.; Richards, A.M.; Pilbrow, A.P. Identifying Candidate Circulating RNA Markers for Coronary Artery Disease by Deep RNA-Sequencing in Human Plasma. Cells 2022, 11, 3191. [Google Scholar] [CrossRef]
- Huang, W.; Shang, W.; Wang, H.; Wu, W.; Hou, S. Sirt1 Overexpression Protects Murine Osteoblasts against TNF-α-Induced Injury in Vitro by Suppressing the NF-ΚB Signaling Pathway. Acta Pharmacol. Sin. 2012, 33, 668–674. [Google Scholar] [CrossRef]
- Wu, B.-W.; Wu, M.-S.; Liu, Y.; Lu, M.; Guo, J.-D.; Meng, Y.-H.; Zhou, Y.-H. SIRT1-Mediated Deacetylation of NF-ΚB Inhibits the MLCK/MLC2 Pathway and the Expression of ET-1, Thus Alleviating the Development of Coronary Artery Spasm. Am. J. Physiol. Heart Circ. Physiol. 2021, 320, H458–H468. [Google Scholar] [CrossRef] [PubMed]
- Donato, A.J.; Morgan, R.G.; Walker, A.E.; Lesniewski, L.A. Cellular and Molecular Biology of Aging Endothelial Cells. J. Mol. Cell. Cardiol. 2015, 89, 122–135. [Google Scholar] [CrossRef] [PubMed]
- Mattagajasingh, I.; Kim, C.-S.; Naqvi, A.; Yamamori, T.; Hoffman, T.A.; Jung, S.-B.; DeRicco, J.; Kasuno, K.; Irani, K. SIRT1 Promotes Endothelium-Dependent Vascular Relaxation by Activating Endothelial Nitric Oxide Synthase. Proc. Natl. Acad. Sci. USA 2007, 104, 14855–14860. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Thavarajah, T.; Gu, W.; Cai, J.; Xu, Q. Impact of MiRNA in Atherosclerosis. Arterioscler. Thromb. Vasc. Biol. 2018, 38, e159–e170. [Google Scholar] [CrossRef] [PubMed]
- Alcendor, R.R.; Gao, S.; Zhai, P.; Zablocki, D.; Holle, E.; Yu, X.; Tian, B.; Wagner, T.; Vatner, S.F.; Sadoshima, J. Sirt1 Regulates Aging and Resistance to Oxidative Stress in the Heart. Circ. Res. 2007, 100, 1512–1521. [Google Scholar] [CrossRef] [PubMed]
- Stein, S.; Schäfer, N.; Breitenstein, A.; Besler, C.; Winnik, S.; Lohmann, C.; Heinrich, K.; Brokopp, C.E.; Handschin, C.; Landmesser, U.; et al. SIRT1 Reduces Endothelial Activation without Affecting Vascular Function in ApoE−/− Mice. Aging 2010, 2, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Luo, G.; Jian, Z.; Zhu, Y.; Zhu, Y.; Chen, B.; Ma, R.; Tang, F.; Xiao, Y. Sirt1 Promotes Autophagy and Inhibits Apoptosis to Protect Cardiomyocytes from Hypoxic Stress. Int. J. Mol. Med. 2019, 43, 2033–2043. [Google Scholar] [CrossRef]
- Jansen, F.; Schäfer, L.; Wang, H.; Schmitz, T.; Flender, A.; Schueler, R.; Hammerstingl, C.; Nickenig, G.; Sinning, J.-M.; Werner, N. Kinetics of Circulating MicroRNAs in Response to Cardiac Stress in Patients With Coronary Artery Disease. J. Am. Heart Assoc. 2017, 6, e005270. [Google Scholar] [CrossRef]
- Zhong, Z.; Zhong, W.; Zhang, Q.; Zhang, Q.; Yu, Z.; Wu, H. Circulating MicroRNA Expression Profiling and Bioinformatics Analysis of Patients with Coronary Artery Disease by RNA Sequencing. J. Clin. Lab. Anal. 2020, 34, e23020. [Google Scholar] [CrossRef]
- Wang, T.; Cheng, Z.; Zhao, R.; Cheng, J.; Ren, H.; Zhang, P.; Liu, P.; Hao, Q.; Zhang, Q.; Yu, X.; et al. Sirt6 Enhances Macrophage Lipophagy and Improves Lipid Metabolism Disorder by Regulating the Wnt1/β-Catenin Pathway in Atherosclerosis. Lipids Health Dis. 2023, 22, 156. [Google Scholar] [CrossRef]
- Brown, B.A.; Connolly, G.M.; Mill, C.E.J.; Williams, H.; Angelini, G.D.; Johnson, J.L.; George, S.J. Aging Differentially Modulates the Wnt Pro-Survival Signalling Pathways in Vascular Smooth Muscle Cells. Aging Cell 2019, 18, e12844. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.-G.; Tang, X.; Wang, J.-J.; Liu, J.; Chen, P.; Sun, Y. A Circular RNA, CircUSP36, Accelerates Endothelial Cell Dysfunction in Atherosclerosis by Adsorbing MiR-637 to Enhance WNT4 Expression. Bioengineered 2021, 12, 6759–6770. [Google Scholar] [CrossRef] [PubMed]
- Tsaousi, A.; Williams, H.; Lyon, C.A.; Taylor, V.; Swain, A.; Johnson, J.L.; George, S.J. Wnt4/β-Catenin Signaling Induces VSMC Proliferation and Is Associated with Intimal Thickening. Circ. Res. 2011, 108, 427–436. [Google Scholar] [CrossRef] [PubMed]
- Shang, X.; Böker, K.O.; Taheri, S.; Hawellek, T.; Lehmann, W.; Schilling, A.F. The Interaction between MicroRNAs and the Wnt/β-Catenin Signaling Pathway in Osteoarthritis. Int. J. Mol. Sci. 2021, 22, 9887. [Google Scholar] [CrossRef] [PubMed]
- Butz, H.; Mészáros, K.; Likó, I.; Patocs, A. Wnt-Signaling Regulated by Glucocorticoid-Induced MiRNAs. Int. J. Mol. Sci. 2021, 22, 11778. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Hu, C.; Li, J.; Liu, L.; Jing, W.; Tang, W.; Tian, W.; Long, J. Effect of MiR-26a-5p on the Wnt/Ca(2+) Pathway and Osteogenic Differentiation of Mouse Adipose-Derived Mesenchymal Stem Cells. Calcif Tissue Int 2016, 99, 174–186. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liu, J.; Cui, J.; Sun, M.; Du, W.; Chen, T.; Ming, X.; Zhang, L.; Tian, J.; Li, J.; et al. MiR218 Modulates Wnt Signaling in Mouse Cardiac Stem Cells by Promoting Proliferation and Inhibiting Differentiation through a Positive Feedback Loop. Sci. Rep. 2016, 6, 20968. [Google Scholar] [CrossRef]
- Bazan, H.A.; Hatfield, S.A.; O’Malley, C.B.; Brooks, A.J.; Lightell, D.J.; Woods, T.C. Acute Loss of MiR-221 and MiR-222 in the Atherosclerotic Plaque Shoulder Accompanies Plaque Rupture. Stroke 2015, 46, 3285–3287. [Google Scholar] [CrossRef]
- Knoka, E.; Trusinskis, K.; Mazule, M.; Briede, I.; Crawford, W.; Jegere, S.; Kumsars, I.; Narbute, I.; Sondore, D.; Lejnieks, A.; et al. Circulating Plasma MicroRNA-126, MicroRNA-145, and MicroRNA-155 and Their Association with Atherosclerotic Plaque Characteristics. J. Clin. Transl. Res. 2020, 5, 60–67. [Google Scholar]
- Gao, H.; Guddeti, R.R.; Matsuzawa, Y.; Liu, L.-P.; Su, L.-X.; Guo, D.; Nie, S.-P.; Du, J.; Zhang, M. Plasma Levels of MicroRNA-145 Are Associated with Severity of Coronary Artery Disease. PLoS ONE 2015, 10, e0123477. [Google Scholar] [CrossRef]
- O’Sullivan, J.F.; Neylon, A.; McGorrian, C.; Blake, G.J. MiRNA-93-5p and Other MiRNAs as Predictors of Coronary Artery Disease and STEMI. Int. J. Cardiol. 2016, 224, 310–316. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, Y.; Xue, S.; Ding, H.; Wang, Y.; Qi, H.; Wang, Y.; Zhu, W.; Li, P. Clinical Significance of Circulating MicroRNAs as Diagnostic Biomarkers for Coronary Artery Disease. J. Cell. Mol. Med. 2020, 24, 1146–1150. [Google Scholar] [CrossRef] [PubMed]
- Kumar, D.; Narang, R.; Sreenivas, V.; Rastogi, V.; Bhatia, J.; Saluja, D.; Srivastava, K. Circulatory MiR-133b and MiR-21 as Novel Biomarkers in Early Prediction and Diagnosis of Coronary Artery Disease. Genes 2020, 11, 164. [Google Scholar] [CrossRef] [PubMed]
- Kumar, D.; Narang, R.; Saluja, D.; Srivastava, K. Functional Association of MiR-133b and MiR-21 Through Novel Gene Targets ATG5, LRP6 and SGPP1 in Coronary Artery Disease. Mol. Diagn. Ther. 2022, 26, 655–664. [Google Scholar] [CrossRef]
- Busk, P.K. A Tool for Design of Primers for MicroRNA-Specific Quantitative RT-QPCR. BMC Bioinforma. 2014, 15, 29. [Google Scholar] [CrossRef]
| All CAD (n = 64) | INOCA (n = 20) | Obstructive CAD (n = 44) | Control (n = 30) | p-Value | |
|---|---|---|---|---|---|
| Men (%) | 33 (51.6) | 5 (25) | 28 (63.6) | 10 (33.3) | 0.004 * p INOCA–obstructive CAD = 0.012 p obstructive CAD–Control = 0.021 |
| Women (%) | 31 (48.4) | 15 (75) | 16 (36.4) | 20 (66.7) | |
| Age (year) | 65 [59; 71] | 66.5 [62.8; 71.2] | 64.0 [56.5; 71] | 28.5 [26; 39.2] | <0.001 * p control–INOCA < 0.001 p control–obstructive CAD < 0.001 |
| BMI (kg/m2) | 26.7 [24.9; 28.8] | 26.20 [25.67; 30.40] | 26.23 [24.68; 28.68] | 21.95 [20.75; 25.23] | <0.001 * p control–INOCA = 0.003 p control–obstructive CAD = 0.003 |
| Smoking (%) | 9 (14) | 3 (15.0) | 6 (13.6) | - | 0.879 |
| Hemoglobin (g/L) | 142 [133; 152] | 142 [134; 151] | 144 [133; 152] | 136 [129; 152] | 0.459 |
| Glucose (mmol/L) | 5.5 [5.17; 5.8] | 5.60 [5.2; 6.21] | 5.40 [5.1; 5.63] | 4.9 [4.59; 5.35] | <0.001 |
| Creatinine (µmol/L) | 89 [78.2; 99.2] | 83.4 [74.8; 96.3] | 89.80 [81; 101.8] | 82 [77.7; 87] | 0.106 |
| Total cholesterol (mmol/L) | 4.39 ± 1.36 | 5.11 ± 1.51 | 3.85 ± 0.95 | 4.87 ± 0.77 | <0.001 * p INOCA– obstructive CAD = 0.005 p obstructive CAD–Control < 0.001 |
| LDL (mmol/L) | 2.36 [1.85; 2.97] | 2.89 [4.34; 3.62] | 2.12 [1.79; 2.48] | 2.54 [2.28; 3.21] | <0.001 * p obstructive CAD–INOCA < 0.001 p control–obstructive CAD = 0.049 |
| HDL (mmol/L) | 1.15 [1.02; 1.36] | 1.27 [1.06; 1.37] | 1.11 [1.02; 1.33] | 1.62 [1.35; 1.9] | <0.001 * p control–INOCA = 0.015 p control–obstructive CAD < 0.001 |
| INOCA | Obstructive CAD | p-Value | |
|---|---|---|---|
| ACE inhibitors | 9 (45.0) | 27 (62.8) | 0.184 |
| ARB II | 6 (30.0) | 10 (23.3) | 0.567 |
| Beta-blocker | 17 (85.0) | 34 (79.1) | 0.737 |
| Calcium channel blockers | 11 (55.0) | 18 (41.9) | 0.33 |
| Antiaggregants | 16 (80.0) | 40 (93.0) | 0.195 |
| Antiarrhythmic drugs | 3 (10.3) | 6 (20.0) | 0.237 |
| HMG-CoA reductase inhibitors | 19 (95.0) | 44 (100.0) | 0.323 |
| Anticoagulants | 4 (15) | 6 (13.6) | 1.000 |
| Proteins | Groups | Concentration (Me [Q1–Q3]) | p-Value |
|---|---|---|---|
| LRP6, ng/mL | INOCA | 13.02 [12.05–13.7] | 0.075 |
| Obstructive CAD | 11.60 [10.5–12.88] | ||
| Control | 12.55 [10.28–14.17] | ||
| WNT1, ng/mL | INOCA | 0.15 [0.15–0.16] | <0.001 * p obstructive CAD–INOCA < 0.001 p control–INOCA = 0.036 p control–CAD < 0.001 |
| Obstructive CAD | 0.189 [0.184–0.193] | ||
| Control | 0.15 [0.15–0.184] | ||
| WNT3a, ng/mL | INOCA | 0.115 [0.07–0.16] | <0.001 * p obstructive CAD–INOCA < 0.001 p control–INOCA < 0.001 |
| Obstructive CAD | 0.227 [0.181–0.252] | ||
| Control | 0.25 [0.162–0.37] | ||
| WNT4, ng/mL | INOCA | 0.345 [0.278–0.492] | 0.015 * p obstructive CAD–INOCA = 0.025 p control–obstructive CAD = 0.047 |
| Obstructive CAD | 0.203 [0.112–0.378] | ||
| Control | 0.345 [0.232–0.528] | ||
| WNT5a, ng/mL | INOCA | 0.17 [0.16–0.17] | <0.001 * p obstructive CAD–INOCA < 0.001 p control–obstructive CAD = 0.001 |
| Obstructive CAD | 0.01 [0.007–0.018] | ||
| Control | 0.16 [0.015–0.17] | ||
| SIRT1, ng/mL | INOCA | 1.09 [1.09–1.1] | <0.001 * p obstructive CAD–INOCA < 0.001 p control–INOCA = 0.012 p control–obstructive CAD = 0.007 |
| Obstructive CAD | 0.079 [0.066–0.104] | ||
| Control | 1.09 [0.035–1.1] |
| Factor/Predictor | B | OR (95%CI) Exp (B) [95%CI] | p-Value |
|---|---|---|---|
| miR-21a (REU) | −2.568 | 0.077 [0.001. 5.757] | p = 0.244 |
| miR-145 (REU) | −2.02 | 0.133 [0.0. 62.492] | p = 0.520 |
| miR-221 (REU) | −1.122 | 0.326 [0.0. 283.87] | p = 0.745 |
| LRP6 (ng/mL) | 0.475 | 1.608 [1.037. 2.494] | p = 0.034 * |
| WNT1 (ng/mL) | −1808.16 | 0.0 [0.0. inf] | p = 0.999 |
| WNT3A (ng/mL) | −30.917 | 0.0 [0.0. 0.00005] | p = 0.001 * |
| WNT4 (ng/mL) | −0.321 | 0.725 [0.368. 1.429] | p = 0.354 |
| WNT5a (ng/mL) | 2.466 | 11.775 [0.298. 465.105] | p = 0.188 |
| SIRT1 (ng/mL) | 64.169 | 7.38e + 27 [0.0. inf] | p = 0.994 |
| Age (years) | 0.036 | 1.037 [0.969. 1.11] | p = 0.297 |
| Smoking (n) | 0.111 | 1.117 [0.25. 5.005] | p = 0.884 |
| Gender (male/female) | −1.658 | 0.191 [0.058. 0.622] | p = 0.006 * |
| BMI (kg/m2) | 0.041 | 1.042 [0.891. 1.218] | p = 0.605 |
| Hypertension (n) | 0.329 | 1.39 [0.136. 14.255] | p = 0.781 |
| Dyslipidemia (n) | 0.329 | 1.39 [0.136. 14.255] | p = 0.781 |
| Angina pain (n) | 0.542 | 1.719 [0.417. 7.084] | p = 0.454 |
| Myocardial infarction (n) | −1.923 | 0.146 [0.03. 0.708] | p = 0.017 * |
| ACE inhibitors | −0.724 | 0.485 [0.165. 1.422] | p = 0.187 |
| ARB II | 0.347 | 1.415 [0.43. 4.647] | p = 0.568 |
| Beta blockers | 0.405 | 1.499 [0.359. 6.271] | p = 0.579 |
| Calcium channel blockers | 0.529 | 1.697 [0.583. 4.945] | p = 0.332 |
| Antiaggregant | −1.204 | 0.3 [0.06. 1.494] | p = 0.142 |
| Statins | −21.683 | 0.0 [0.0. inf] | p = 0.999 |
| Fasting glucose (mmol/l) | −0.021 | 0.979 [0.898. 1.067] | p = 0.635 |
| Variables | Coef (B) | Exp (B) | p-Value |
|---|---|---|---|
| LRP6, ng/mL | 0.451 | 1.57 [1.17. 2.09] | p = 0.002 * |
| WNT3a, ng/mL | −34.4454 | 0.0 [0.0. 0.00001] | p < 0.001 |
| Primer | Sequence |
|---|---|
| miR-21a | 5′-TAGCTTATCAGACTGATGTTGAAAA-3′ |
| miR-145 | 5′-TCCAGTTTTCCCAGGAATCCCT-3′ |
| miR-221 | 5′-GACCTGGCATACAATGTAGATTTAAA-3′ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Iusupova, A.O.; Pakhtusov, N.N.; Slepova, O.A.; Belenkov, Y.N.; Privalova, E.V.; Bure, I.V.; Vetchinkina, E.A.; Nemtsova, M.V. MiRNA-21a, miRNA-145, and miRNA-221 Expression and Their Correlations with WNT Proteins in Patients with Obstructive and Non-Obstructive Coronary Artery Disease. Int. J. Mol. Sci. 2023, 24, 17613. https://doi.org/10.3390/ijms242417613
Iusupova AO, Pakhtusov NN, Slepova OA, Belenkov YN, Privalova EV, Bure IV, Vetchinkina EA, Nemtsova MV. MiRNA-21a, miRNA-145, and miRNA-221 Expression and Their Correlations with WNT Proteins in Patients with Obstructive and Non-Obstructive Coronary Artery Disease. International Journal of Molecular Sciences. 2023; 24(24):17613. https://doi.org/10.3390/ijms242417613
Chicago/Turabian StyleIusupova, Alfiya Oskarovna, Nikolay Nikolaevich Pakhtusov, Olga Alexandrovna Slepova, Yuri Nikitich Belenkov, Elena Vitalievna Privalova, Irina Vladimirovna Bure, Ekaterina Alexandrovna Vetchinkina, and Marina Vyacheslavovna Nemtsova. 2023. "MiRNA-21a, miRNA-145, and miRNA-221 Expression and Their Correlations with WNT Proteins in Patients with Obstructive and Non-Obstructive Coronary Artery Disease" International Journal of Molecular Sciences 24, no. 24: 17613. https://doi.org/10.3390/ijms242417613






