Combined Multiplexed Phage Display, High-Throughput Sequencing, and Functional Assays as a Platform for Identifying Modulatory VHHs Targeting the FSHR
Abstract
:1. Introduction
2. Results
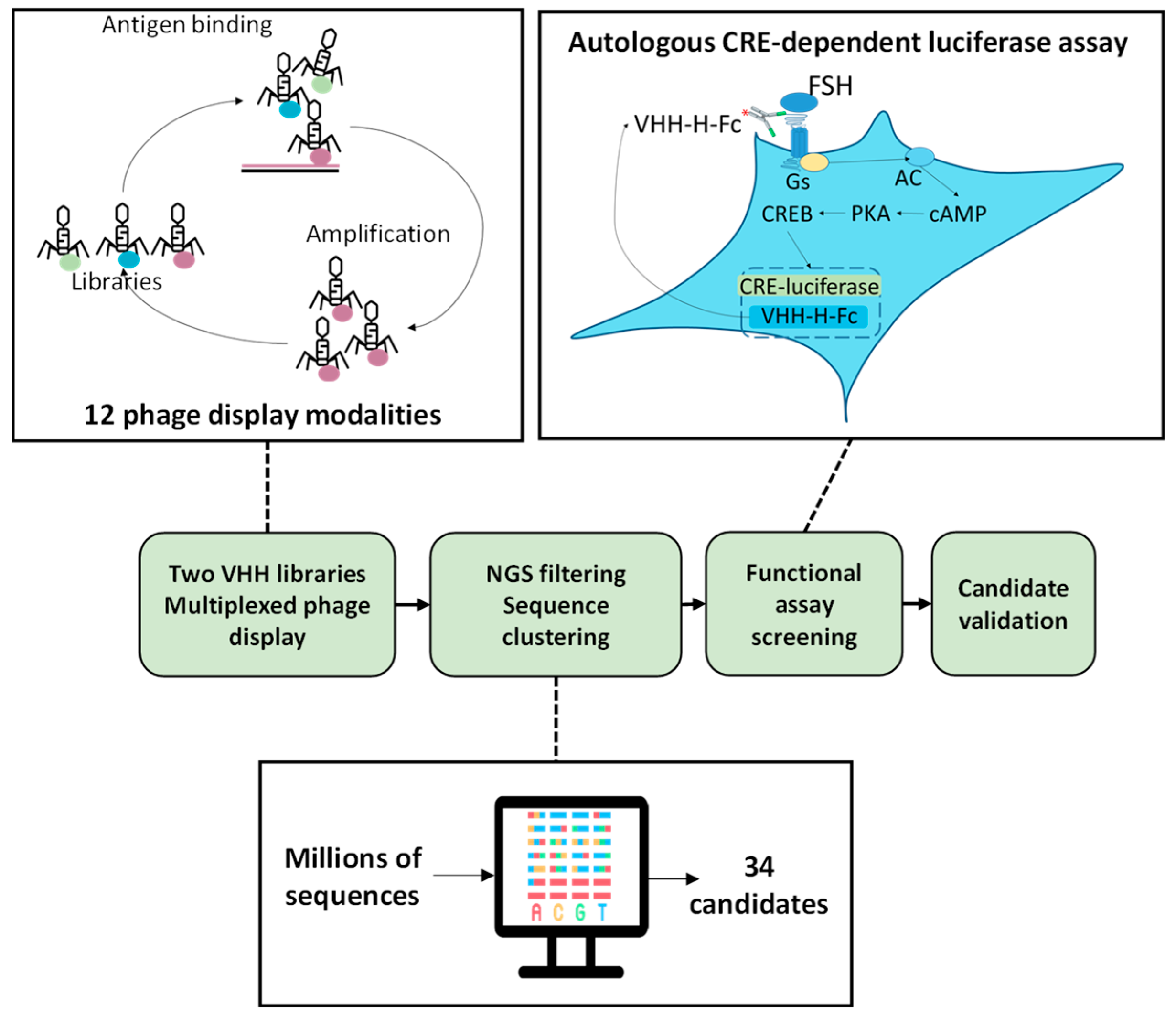
2.1. Multiplexed Phage-Display and NGS Analysis
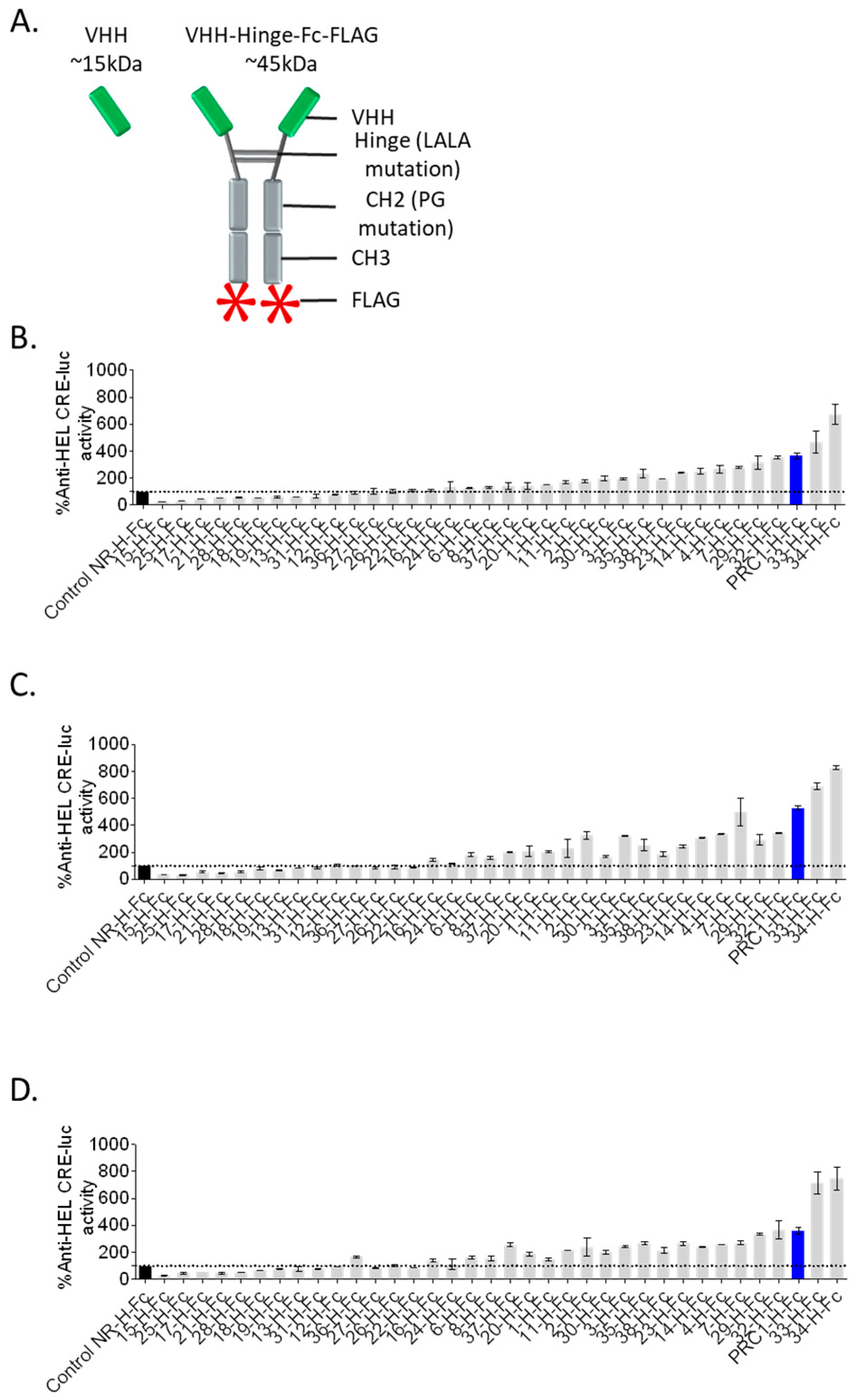
2.2. Autologously Expressed VHH-Hinge-Fc Assayed on CRE-Driven Luciferase Reporter Gene as a Primary Functional Screening
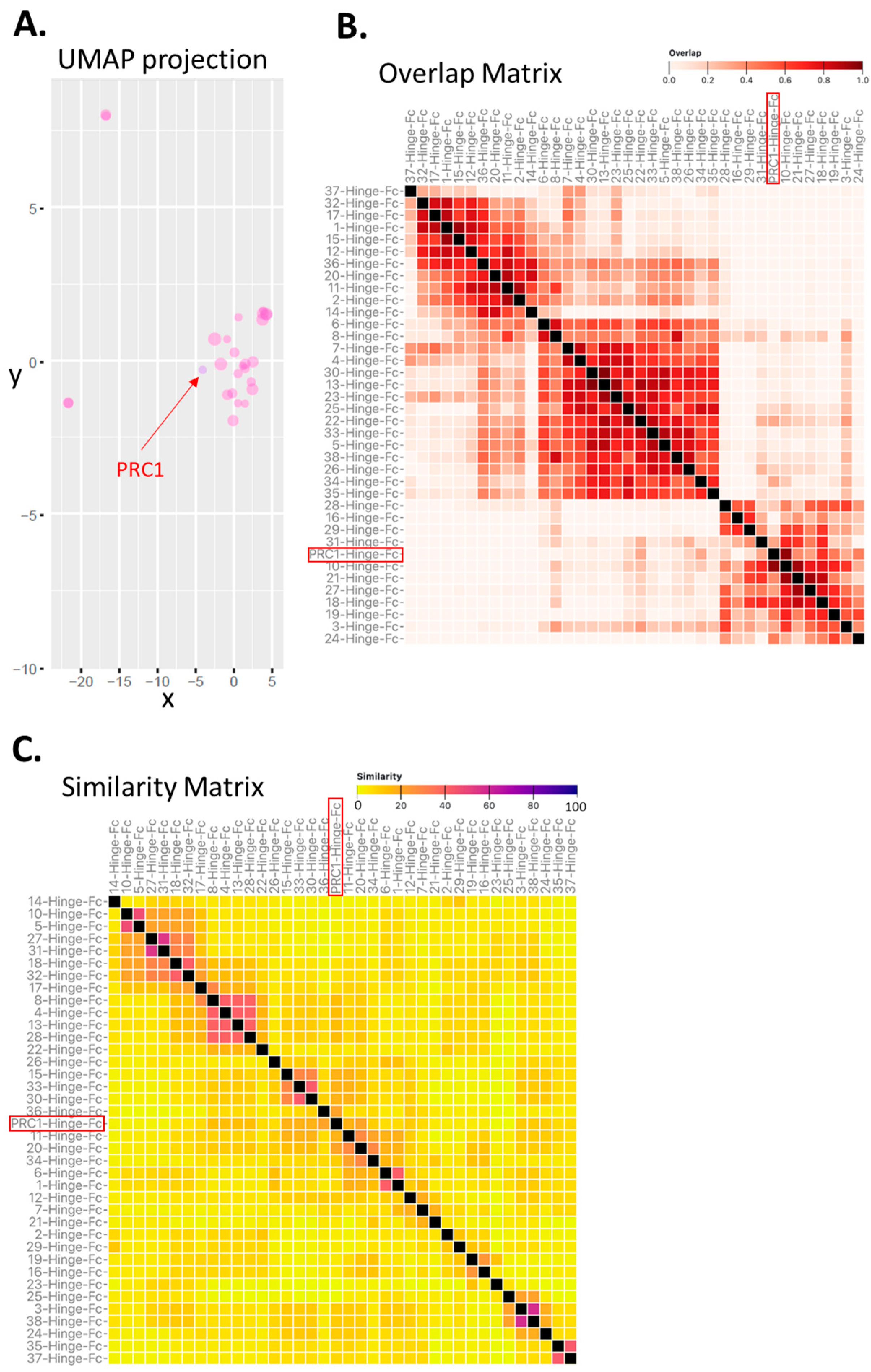
2.3. Clustering of VHH Sequences as a Mean to Assess Epitopic Diversity among Candidate VHHs
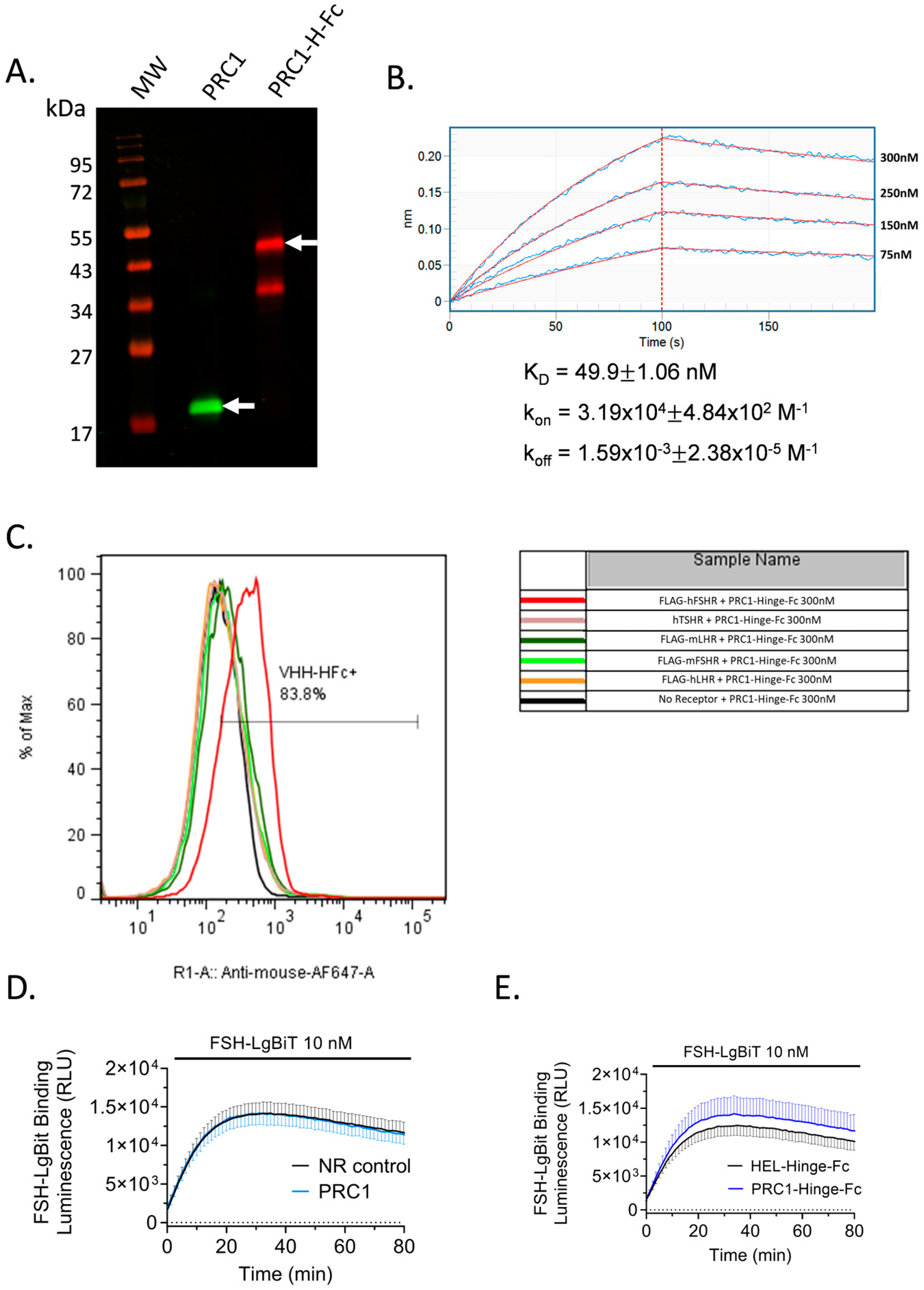
2.4. Validation of PRC1-Hinge-Fc Activity Using expiCHO Extracts and cAMP BRET Assay
2.5. PRC1-Hinge-Fc Binding and Kinetic Rate Constants Determination
2.6. PRC1-Hinge-Fc Binding Specificity and Binding Mode
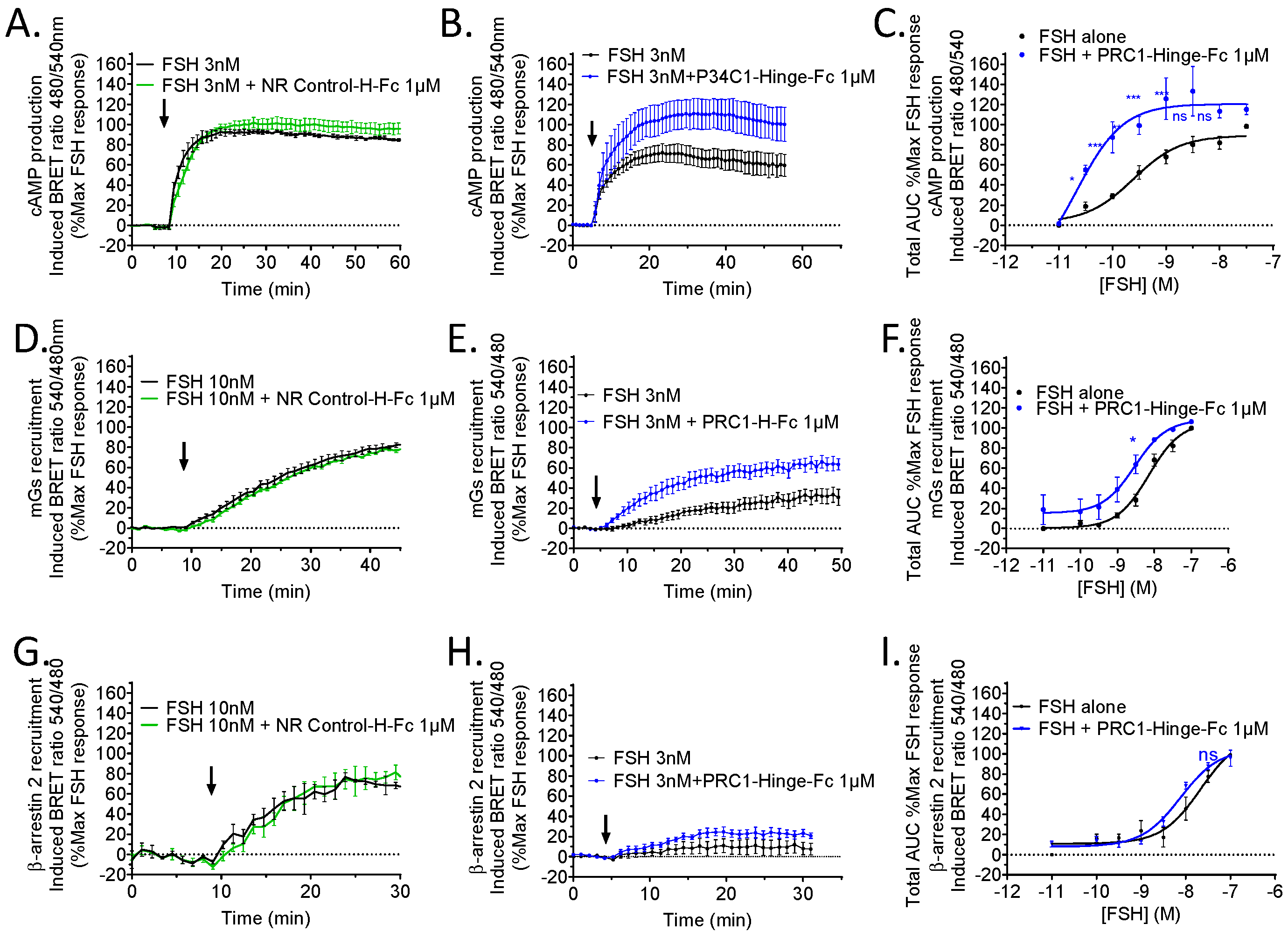
2.7. Pharmacological Profile of PRC1-Hinge-Fc
3. Discussion
4. Materials and Methods
4.1. Ligands and Materials
4.2. Cell Culture and Transfection
4.3. Llama Libraries
4.4. Phage Display
4.5. NGS Analysis by Germinal Lineage
4.6. In Silico Prediction of Epitopes
4.7. Autologous Reporter Gene Assay
4.8. Production of VHH-Hinge-Fc in ExpiCHO Mammalian Cell Expression System
4.9. SDS-PAGE
4.10. BRET Assays
4.11. Flow Cytometry
4.12. Binding Validation with BioLayer Interferometry (BLI)
4.13. NanoBiT FSH Binding Assay
4.14. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Duan, J.; Xu, P.; Zhang, H.; Luan, X.; Yang, J.; He, X.; Mao, C.; Shen, D.-D.; Ji, Y.; Cheng, X.; et al. Mechanism of Hormone and Allosteric Agonist Mediated Activation of Follicle Stimulating Hormone Receptor. Nat. Commun. 2023, 14, 519. [Google Scholar] [CrossRef] [PubMed]
- Richards, J.S.; Pangas, S.A. The Ovary: Basic Biology and Clinical Implications. J. Clin. Investig. 2010, 120, 963–972. [Google Scholar] [CrossRef] [PubMed]
- Huhtaniemi, I. A Short Evolutionary History of FSH-Stimulated Spermatogenesis. Hormones 2015, 14, 468–478. [Google Scholar] [CrossRef] [PubMed]
- Bordoloi, D.; Bhojnagarwala, P.S.; Perales-Puchalt, A.; Kulkarni, A.J.; Zhu, X.; Liaw, K.; O’Connell, R.P.; Park, D.H.; Kulp, D.W.; Zhang, R.; et al. A MAb against Surface-Expressed FSHR Engineered to Engage Adaptive Immunity for Ovarian Cancer Immunotherapy. JCI Insight 2022, 7, e162553. [Google Scholar] [CrossRef]
- Chen, X.; Chen, L.; Wang, Y.; Shu, C.; Zhou, Y.; Wu, R.; Jin, B.; Yang, L.; Sun, J.; Qi, M.; et al. Identification and Characterization of Novel Compound Heterozygous Variants in FSHR Causing Primary Ovarian Insufficiency with Resistant Ovary Syndrome. Front. Endocrinol. 2023, 13, 1013894. [Google Scholar] [CrossRef]
- Laven, J.S.E. Follicle Stimulating Hormone Receptor (FSHR) Polymorphisms and Polycystic Ovary Syndrome (PCOS). Front. Endocrinol. 2019, 10, 23. [Google Scholar] [CrossRef]
- Vloeberghs, V.; Peeraer, K.; Pexsters, A.; D’Hooghe, T. Ovarian Hyperstimulation Syndrome and Complications of ART. Best Pract. Res. Clin. Obstet. Gynaecol. 2009, 23, 691–709. [Google Scholar] [CrossRef]
- Tapanainen, J.S.; Aittomaki, K.; Min, J.; Vaskivuo, T.; Huhtaniemi, I.T. Men Homozygous for an Inactivating Mutation of the Follicle-Stimulating Hormone {FSH) Receptor Gene Present Variable Suppression of Spermatogenesis and Fertility. Nat. Genet. 1997, 15, 205–206. [Google Scholar] [CrossRef]
- Szymańska, K.; Kałafut, J.; Przybyszewska, A.; Paziewska, B.; Adamczuk, G.; Kiełbus, M.; Rivero-Müller, A. FSHR Trans-Activation and Oligomerization. Front. Endocrinol. 2018, 9, 760. [Google Scholar] [CrossRef]
- Landomiel, F.; De Pascali, F.; Raynaud, P.; Jean-Alphonse, F.; Yvinec, R.; Pellissier, L.P.; Bozon, V.; Bruneau, G.; Crépieux, P.; Poupon, A.; et al. Biased Signaling and Allosteric Modulation at the FSHR. Front. Endocrinol. 2019, 10, 48. [Google Scholar] [CrossRef]
- Guo, Y.; Zhao, M.; Bo, T.; Ma, S.; Yuan, Z.; Chen, W.; He, Z.; Hou, X.; Liu, J.; Zhang, Z.; et al. Blocking FSH Inhibits Hepatic Cholesterol Biosynthesis and Reduces Serum Cholesterol. Cell Res. 2019, 29, 151–166. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.; Kang, S.S.; Wang, Z.; Liu, X.; Kuo, T.C.; Korkmaz, F.; Padilla, A.; Miyashita, S.; Chan, P.; Zhang, Z.; et al. FSH Blockade Improves Cognition in Mice with Alzheimer’s Disease. Nature 2022, 603, 470–476. [Google Scholar] [CrossRef] [PubMed]
- Gera, S.; Sant, D.; Haider, S.; Korkmaz, F.; Kuo, T.C.; Mathew, M.; Perez-Pena, H.; Xie, H.; Chen, H.; Batista, R.; et al. First-in-Class Humanized FSH Blocking Antibody Targets Bone and Fat. Proc. Natl. Acad. Sci. USA 2020, 117, 28971–28979. [Google Scholar] [CrossRef] [PubMed]
- Peterson, S.M.; Hutchings, C.J.; Hu, C.F.; Mathur, M.; Salameh, J.W.; Axelrod, F.; Sato, A.K. Discovery and Design of G Protein-Coupled Receptor Targeting Antibodies. Expert Opin. Drug Discov. 2023, 18, 417–428. [Google Scholar] [CrossRef] [PubMed]
- Foster, D.J.; Conn, P.J. Allosteric modulation of GPCRs: New insights and potential utility for treatment of schizophrenia and other CNS disorders. Neuron 2018, 94, 431–446. [Google Scholar] [CrossRef] [PubMed]
- Wold, E.A.; Chen, J.; Cunningham, K.A.; Zhou, J. Allosteric Modulation of Class A GPCRs: Targets, Agents, and Emerging Concepts. J. Med. Chem. 2019, 62, 88–127. [Google Scholar] [CrossRef]
- Reiter, E.; Ahn, S.; Shukla, A.K.; Lefkowitz, R.J. Molecular Mechanism of β-Arrestin-Biased Agonism at Seven-Transmembrane Receptors. Annu. Rev. Pharmacol. Toxicol. 2012, 52, 179–197. [Google Scholar] [CrossRef]
- Hutchings, C.J.; Koglin, M.; Marshall, F.H. Therapeutic Antibodies Directed at G Protein-Coupled Receptors. MAbs 2010, 2, 594–606. [Google Scholar] [CrossRef]
- Jin, B.K.; Odongo, S.; Radwanska, M.; Magez, S. Nanobodies: A Review of Generation, Diagnostics and Therapeutics. Int. J. Mol. Sci. 2023, 24, 5994. [Google Scholar] [CrossRef]
- Ayoub, M.A.; Crépieux, P.; Koglin, M.; Parmentier, M.; Pin, J.P.; Poupon, A.; Reiter, E.; Smit, M.; Steyaert, J.; Watier, H.; et al. Antibodies Targeting G Protein-Coupled Receptors: Recent Advances and Therapeutic Challenges. MAbs 2017, 9, 735–741. [Google Scholar] [CrossRef]
- Steyaert, J.; Kobilka, B.K. Nanobody Stabilization of G Protein-Coupled Receptor Conformational States. Curr. Opin. Struct. Biol. 2011, 21, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Lo, M.; Kim, H.S.; Tong, R.K.; Bainbridge, T.W.; Vernes, J.M.; Zhang, Y.; Lin, Y.L.; Chung, S.; Dennis, M.S.; Zuchero, Y.J.Y.; et al. Effector-Attenuating Substitutions That Maintain Antibody Stability and Reduce Toxicity in Mice. J. Biol. Chem. 2017, 292, 3900–3908. [Google Scholar] [CrossRef]
- Kumar, V.; Vollbrecht, T.; Chernyshev, M.; Mohan, S.; Hanst, B.; Bavafa, N.; Lorenzo, A.; Kumar, N.; Ketteringham, R.; Eren, K.; et al. Long-Read Amplicon Denoising. Nucleic Acids Res. 2019, 47, E104. [Google Scholar] [CrossRef] [PubMed]
- Hanke, L.; Sheward, D.J.; Pankow, A.; Vidakovics, L.P.; Karl, V.; Kim, C.; Urgard, E.; Smith, N.L.; Astorga-Wells, J.; Ekström, S.; et al. Multivariate Mining of an Alpaca Immune Repertoire Identifies Potent Cross-Neutralizing SARS-CoV-2 Nanobodies. Sci. Adv. 2022, 8, eabm0220. [Google Scholar] [CrossRef]
- Bourquard, T.; Musnier, A.; Puard, V.; Tahir, S.; Ayoub, M.A.; Jullian, Y.; Boulo, T.; Gallay, N.; Watier, H.; Bruneau, G.; et al. MAbTope: A Method for Improved Epitope Mapping. J. Immunol. 2018, 201, 3096–3105. [Google Scholar] [CrossRef] [PubMed]
- Musnier, A.; Bourquard, T.; Vallet, A.; Mathias, L.; Bruneau, G.; Ayoub, M.A.; Travert, O.; Corde, Y.; Gallay, N.; Boulo, T.; et al. A New in Silico Antibody Similarity Measure Both Identifies Large Sets of Epitope Binders with Distinct CDRs and Accurately Predicts Off-Target Reactivity. Int. J. Mol. Sci. 2022, 23, 9765. [Google Scholar] [CrossRef] [PubMed]
- Wan, Q.; Okashah, N.; Inoue, A.; Nehme, R.; Carpenter, B.; Tate, C.G.; Lambert, N.A. Mini G Protein Probes for Active G Protein-Coupled Receptors (GPCRs) in Live Cells. J. Biol. Chem. 2018, 293, 7466–7473. [Google Scholar] [CrossRef]
- Liu, L. Pharmacokinetics of Monoclonal Antibodies and Fc-Fusion Proteins. Protein Cell 2018, 9, 15–32. [Google Scholar] [CrossRef]
- Gan, S.K.E.; Phua, S.X.; Yeo, J.Y. Sagacious Epitope Selection for Vaccines, and Both Antibody-Based Therapeutics and Diagnostics: Tips from Virology and Oncology. Antib. Ther. 2022, 5, 63–72. [Google Scholar] [CrossRef]
- Berezovsky, I.N. Thermodynamics of Allostery Paves a Way to Allosteric Drugs. Biochim. Biophys. Acta-Proteins Proteom. 2013, 1834, 830–835. [Google Scholar] [CrossRef]
- Xia, X.; Zhai, T.; Li, J. Preparation and Identification of Anti-Follicle-Stimulating Hormone Receptor Nanobodies. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi 2013, 29, 829–833. [Google Scholar] [PubMed]
- Crepin, R.; Veggiani, G.; Djender, S.; Beugnet, A.; Planeix, F.; Pichon, C.; Moutel, S.; Amigorena, S.; Perez, F.; Ghinea, N.; et al. Whole-Cell Biopanning with a Synthetic Phage Display Library of Nanobodies Enabled the Recovery of Follicle-Stimulating Hormone Receptor Inhibitors. Biochem. Biophys. Res. Commun. 2017, 493, 1567–1572. [Google Scholar] [CrossRef] [PubMed]
- Scholler, P.; Nevoltris, D.; De Bundel, D.; Bossi, S.; Moreno-Delgado, D.; Rovira, X.; Møller, T.C.; El Moustaine, D.; Mathieu, M.; Blanc, E.; et al. Allosteric Nanobodies Uncover a Role of Hippocampal MGlu2 Receptor Homodimers in Contextual Fear Consolidation. Nat. Commun. 2017, 8, 1967. [Google Scholar] [CrossRef] [PubMed]
- Haubrich, J.; Font, J.; Quast, R.B.; Goupil-Lamy, A.; Scholler, P.; Nevoltris, D.; Acher, F.; Chames, P.; Rondard, P.; Prézeau, L.; et al. A Nanobody Activating Metabotropic Glutamate Receptor 4 Discriminates between Homo- And Heterodimers. Proc. Natl. Acad. Sci. USA 2021, 118, 2–7. [Google Scholar] [CrossRef] [PubMed]
- Akiba, H.; Tamura, H.; Kiyoshi, M.; Yanaka, S.; Sugase, K.; Caaveiro, J.M.M.; Tsumoto, K. Structural and Thermodynamic Basis for the Recognition of the Substrate-Binding Cleft on Hen Egg Lysozyme by a Single-Domain Antibody. Sci. Rep. 2019, 9, 4–6. [Google Scholar] [CrossRef] [PubMed]
- Jones, M.L.; Alfaleh, M.A.; Kumble, S.; Zhang, S.; Osborne, G.W.; Yeh, M.; Arora, N.; Hou, J.J.C.; Howard, C.B.; Chin, D.Y.; et al. Targeting Membrane Proteins for Antibody Discovery Using Phage Display. Sci. Rep. 2016, 6, 26240. [Google Scholar] [CrossRef]
- Verheesen, P.; Laeremans, T. Single Domain Antibodies; Saerens, D., Muyldermans, S., Eds.; Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2012; Volume 911, ISBN 978-1-61779-967-9. [Google Scholar]
- Pardon, E.; Laeremans, T.; Triest, S.; Rasmussen, S.G.F.; Wohlkönig, A.; Ruf, A.; Muyldermans, S.; Hol, W.G.J.; Kobilka, B.K.; Steyaert, J. A General Protocol for the Generation of Nanobodies for Structural Biology. Nat. Protoc. 2014, 9, 674–693. [Google Scholar] [CrossRef]
- Ashraf, Y.; Mansouri, H.; Laurent-Matha, V.; Alcaraz, L.B.; Roger, P.; Guiu, S.; Derocq, D.; Robin, G.; Michaud, H.A.; Delpech, H.; et al. Immunotherapy of Triple-Negative Breast Cancer with Cathepsin D-Targeting Antibodies. J. Immunother. Cancer 2019, 7, 29. [Google Scholar] [CrossRef]
- Troispoux, C.; Guillou, F.; Elalouf, J.M.; Firsov, D.; Iacovelli, L.; De Blasi, A.; Combarnous, Y.; Reiter, E. Involvement of G Protein-Coupled Receptor Kinases and Arrestins in Desensitization to Follicle-Stimulating Hormone Action. Mol. Endocrinol. 1999, 13, 1599–1614. [Google Scholar] [CrossRef]
- Ayoub, M.A.; Landomiel, F.; Gallay, N.; Jégot, G.; Poupon, A.; Crépieux, P.; Reiter, E. Assessing Gonadotropin Receptor Function by Resonance Energy Transfer-Based Assays. Front. Endocrinol. 2015, 6, 130. [Google Scholar] [CrossRef]
- Liu, Y.-L.; Guo, Z.-Y. The NanoBiT-Based Homogenous Ligand–Receptor Binding Assay. In Bioluminescence: Methods and Protocols; Springer: New York, NY, USA, 2022; pp. 139–153. [Google Scholar]

| Proteoliposomes | Cells | Maltose-Binding Protein | FSHR Peptide | FSHR Ectodomain | Cell Membranes | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Control | FSHR (R3) | Control | FSHR (R2) | FSHR (R3) | Control | FSHR (R2) | FSHR (R3) | FSHR (R2) | FSHR (R3) | Control | FSHR (R3) | Total | |
| PRC1 | 28,890 | 16,209 | 45,099 | ||||||||||
| 1 | 200 | 952 | 68 | 34 | 228 | ||||||||
| 6 | 560 | 103 | |||||||||||
| 11 | 114 | 49 | 1464 | 500 | 54 | 239 | |||||||
| 15 | 451 | 83 | |||||||||||
| 19 | 757 | 403 | |||||||||||
| 23 | 447 | 238 | |||||||||||
| 27 | 128 | 282 | 160 | ||||||||||
| 31 | 254 | 135 | |||||||||||
| 2 | 197 | 105 | |||||||||||
| 7 | 2074 | 249 | 269 | ||||||||||
| 12 | 1313 | 230 | 185 | ||||||||||
| 16 | 171 | 12018 | 19 | 964 | |||||||||
| 35 | 114 | 8243 | 26 | 668 | |||||||||
| 20 | 95 | 6715 | 533 | ||||||||||
| 24 | 4198 | 327 | |||||||||||
| 36 | 3749 | 191 | 331 | ||||||||||
| 28 | 59 | 6364 | 480 | ||||||||||
| 38 | 4256 | 313 | |||||||||||
| 32 | 504 | 2529 | 288 | ||||||||||
| 3 | 2271 | 167 | |||||||||||
| 8 | 178 | 1972 | 181 | ||||||||||
| 13 | 405 | 1822 | 216 | ||||||||||
| 17 | 1741 | 128 | |||||||||||
| 21 | 1846 | 1469 | 92 | 493 | |||||||||
| 25 | 884 | 1278 | 273 | ||||||||||
| 29 | 592 | 163 | 103 | 151 | |||||||||
| 33 | 504 | 340 | 127 | ||||||||||
| 4 | 494 | 476 | 135 | ||||||||||
| 34 | 776 | 93 | |||||||||||
| 30 | 20 | 4 | |||||||||||
| 14 | 451 | 54 | |||||||||||
| 22 | 42 | 5 | |||||||||||
| 37 | 2863 | 223 | |||||||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zehnaker, A.; Vallet, A.; Gourdon, J.; Sarti, C.; Jugnarain, V.; Haj Hassan, M.; Mathias, L.; Gauthier, C.; Raynaud, P.; Boulo, T.; et al. Combined Multiplexed Phage Display, High-Throughput Sequencing, and Functional Assays as a Platform for Identifying Modulatory VHHs Targeting the FSHR. Int. J. Mol. Sci. 2023, 24, 15961. https://doi.org/10.3390/ijms242115961
Zehnaker A, Vallet A, Gourdon J, Sarti C, Jugnarain V, Haj Hassan M, Mathias L, Gauthier C, Raynaud P, Boulo T, et al. Combined Multiplexed Phage Display, High-Throughput Sequencing, and Functional Assays as a Platform for Identifying Modulatory VHHs Targeting the FSHR. International Journal of Molecular Sciences. 2023; 24(21):15961. https://doi.org/10.3390/ijms242115961
Chicago/Turabian StyleZehnaker, Anielka, Amandine Vallet, Juliette Gourdon, Caterina Sarti, Vinesh Jugnarain, Maya Haj Hassan, Laetitia Mathias, Camille Gauthier, Pauline Raynaud, Thomas Boulo, and et al. 2023. "Combined Multiplexed Phage Display, High-Throughput Sequencing, and Functional Assays as a Platform for Identifying Modulatory VHHs Targeting the FSHR" International Journal of Molecular Sciences 24, no. 21: 15961. https://doi.org/10.3390/ijms242115961
APA StyleZehnaker, A., Vallet, A., Gourdon, J., Sarti, C., Jugnarain, V., Haj Hassan, M., Mathias, L., Gauthier, C., Raynaud, P., Boulo, T., Beauclair, L., Bigot, Y., Casarini, L., Crépieux, P., Poupon, A., Piégu, B., Jean-Alphonse, F., Bruneau, G., & Reiter, É. (2023). Combined Multiplexed Phage Display, High-Throughput Sequencing, and Functional Assays as a Platform for Identifying Modulatory VHHs Targeting the FSHR. International Journal of Molecular Sciences, 24(21), 15961. https://doi.org/10.3390/ijms242115961






