Abstract
Biological membranes, primarily composed of lipids, envelop each living cell. The intricate composition and organization of membrane lipids, including the variety of fatty acids they encompass, serve a dynamic role in sustaining cellular structural integrity and functionality. Typically, modifications in lipid composition coincide with consequential alterations in universally significant signaling pathways. Exploring the various fatty acids, which serve as the foundational building blocks of membrane lipids, provides crucial insights into the underlying mechanisms governing a myriad of cellular processes, such as membrane fluidity, protein trafficking, signal transduction, intercellular communication, and the etiology of certain metabolic disorders. Furthermore, comprehending how alterations in the lipid composition, especially concerning the fatty acid profile, either contribute to or prevent the onset of pathological conditions stands as a compelling area of research. Hence, this review aims to meticulously introduce the intricacies of membrane lipids and their constituent fatty acids in a healthy organism, thereby illuminating their remarkable diversity and profound influence on cellular function. Furthermore, this review aspires to highlight some potential therapeutic targets for various pathological conditions that may be ameliorated through dietary fatty acid supplements. The initial section of this review expounds on the eukaryotic biomembranes and their complex lipids. Subsequent sections provide insights into the synthesis, membrane incorporation, and distribution of fatty acids across various fractions of membrane lipids. The last section highlights the functional significance of membrane-associated fatty acids and their innate capacity to shape the various cellular physiological responses.
1. Introduction
The biological membrane, commonly referred to as the biomembrane, holds paramount importance in both prokaryotic and eukaryotic cells. Its primary function lies in the selective regulation of molecular influx and efflux across the cellular boundary. Furthermore, it plays a crucial role in modulating intercellular communication and is involved in a vast array of complex processes, encompassing cell proliferation, differentiation, secretion, migration, invasion, and phagocytosis. However, the term “biomembrane” extends beyond the plasma membrane, as eukaryotic cells feature membranes within distinct cellular organelles [1,2], such as the endoplasmic reticulum (ER), mitochondria, nucleus, and various intracellular organelles. Additional functions of biomembranes revolve around stabilizing the consistency of cellular activities within the cell and organelles, controlling the trafficking of micromolecules (including O2, CO2, H2O, H+, K+, HCO3−, Mg2+, Ca2+, etc.) and macromolecular compounds, and providing a surface where essential biological events take place. According to Janmey and Kinnunen [3], biomembranes represent heterogeneous, asymmetrical bilayers with complex structures that contribute to the maintenance of cellular homeostasis and functionality. Hence, biomembranes’ systems exhibit considerable structural and dynamic diversity, making them an enduring area of scientific exploration.
The concept of Langmuir films, initially proposed by Langmuir in 1917, represents the earliest paradigm aimed at elucidating membrane systems [4]. Numerous subsequent paradigms have been developed in an attempt to explain membrane systems. The semifluid dynamics of biomembranes are merely determined by their intricate structure. The so-called “fluid mosaic model”, one of the most renowned models in the biological domain, is employed to illustrate the structure and function of membranes. Singer and Nicolson introduced this model in 1972 [5], describing lipids, proteins, and carbohydrates as the primary constituents of the membrane. In light of the fact that proteins do not actually dissolve in membrane lipids, this proposal has undergone several amendments. After 25 years of Singer and Nicolson’s proposal, Simons and Ikonen [6] proposed the “lipid raft” model, predicated on the clustering of sterols (namely, cholesterol in animals) and sphingolipids (SLs) within membranes to form microdomains where membrane-associated proteins are distributed. It has been established that these compartmentalized microdomains limit membrane lateral mobility and actively engage in various cellular events based on their structural arrangements [7]. Following the lipid rafts model, numerous other models have been introduced, which are either focused on revising the fluid mosaic model [8] or explaining the interaction between the similar [9] or distinct molecule classes [10] within membranes. Generally, the complexity of membranes exceeds that of model membranes due to the heterogeneous distribution of building molecules and their complex interactions. The continuous advancement of technology empowers science to delve deeper into the intricate structures of membranes, implying that the cell membrane model will invariably evolve toward increasing complexity, mirroring the progression from initial notions of membrane structure.
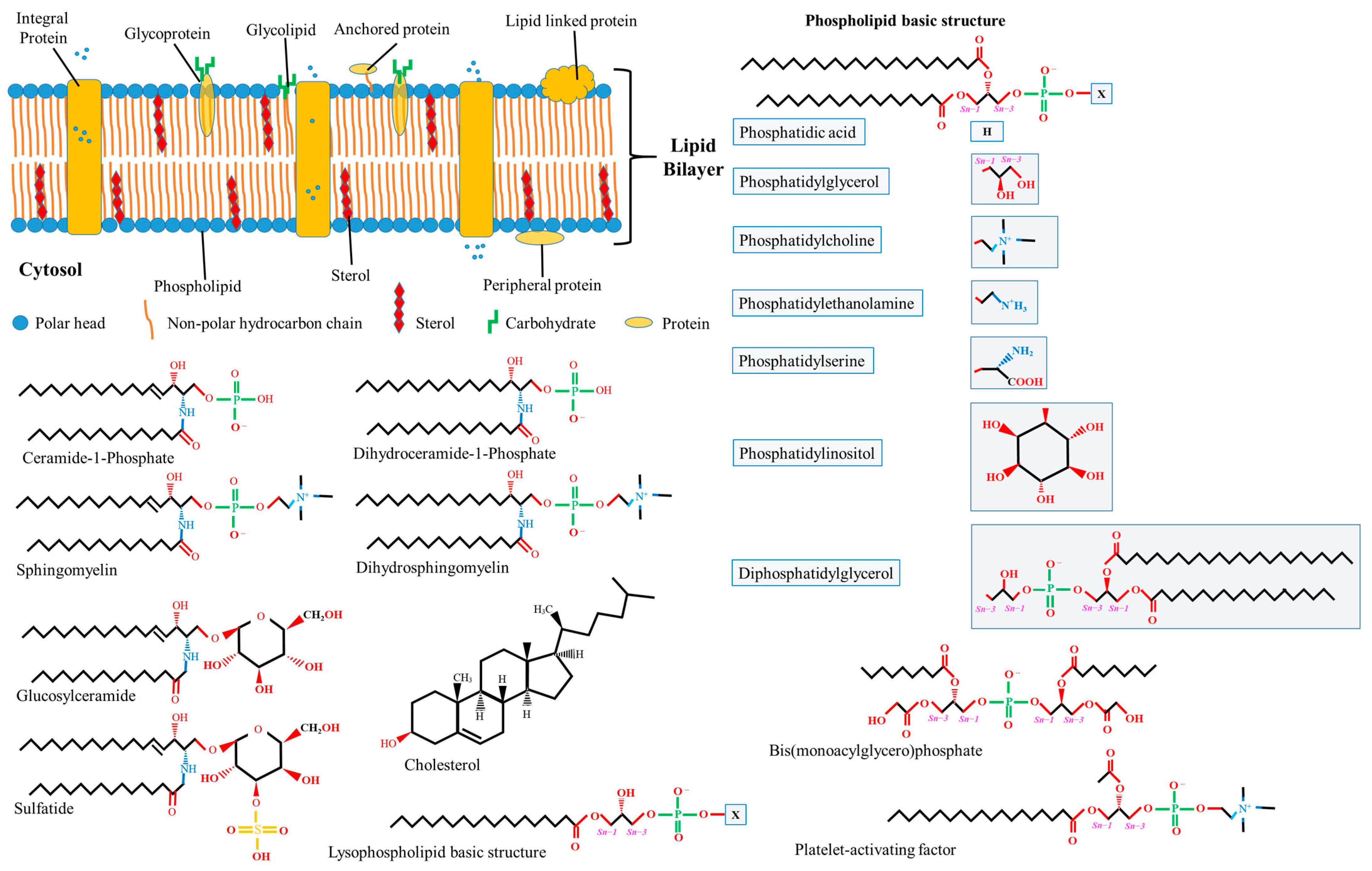
Lipids, proteins, and carbohydrates are pivotal biomolecules within biomembranes, exhibiting heterogeneous dispersion across membranes’ structures (see Figure 1). Membrane lipids, marked by diversity and possessing distinct properties either individually or in conjunction with other moieties, contribute to bilayer development and serve essential functions. Almost 50% of the membrane matrix is composed of proteins, which exist in various structures such as including integral (embedded with lipid bilayers), peripheral (associated with the membrane surface), and anchoring (not directly attached but rather bound to lipid embedded with lipid bilayers) proteins. Hydrophobic forces or ionic interactions mediate the binding of membrane proteins to membrane lipids, forming lateral domains with certain functions such as environmental communication, adhesion, trafficking, and signaling. Carbohydrates form covalent bonds with proteins or lipids within membranes, which only occur at the outward surface of the plasma membrane, yielding glycol-complexes [2,11]. The extant biotic assemblies within biomembranes are postulated to have transited from thermodynamic reactions on analogous abiotic assemblies [12]. The interaction between membrane lipids and proteins may modulate their individual qualities, thereby altering membrane conformation.
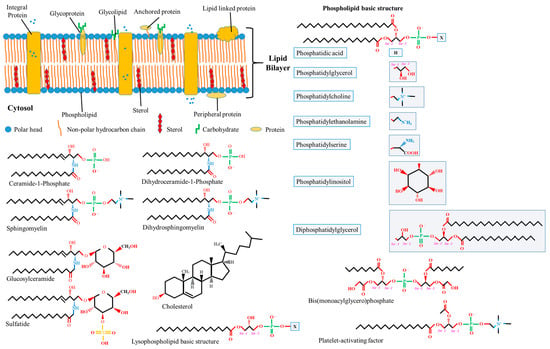
Figure 1.
Schematic representation of biological compartments of the cell membrane and the molecular structure of the different lipids available in membranes. The molecular structures of different lipids have been adapted from the PubChem database (an open database for the public, available at https://pubchem.ncbi.nlm.nih.gov, accessed on 17 September 2023). Abbreviation: Sn, stereospecific numbering in the glycerol; 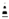 and
and 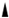 , chiral carbon centers.
, chiral carbon centers.
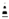 and
and 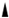 , chiral carbon centers.
, chiral carbon centers.
Among the constituents of biomembranes, fatty acids comprising the lipid portion have gained great focus due to their diverse functions in cellular processes. Understanding the diversity and composition of eukaryotic biomembrane lipids, especially fatty acids, is essential for elucidating the underlying mechanisms controlling cellular functions. Furthermore, it sheds light on the potential roles that particular lipids and fatty acids may play in various physiological and pathological processes, including inflammation and metabolic disorders. The current review primarily focuses on a healthy organism, intending to highlight the enormous diversity of biomembrane lipids and, as a secondary objective, characterize the biological roles of distinct fatty acids embedded into the cellular membranes. In addition, this review enhances our knowledge of fundamental cellular processes and subtly underscores the potential for therapeutic strategies centered on the lipid composition and fatty acid metabolism of biomembranes, which are likely promising foundations for further scientific inquiry.
2. Lipid Bilayer
Lipids have gained recognition and have become a subject of considerable interest among scientists since the original publication of Chevreul’s work [13], which delineated the concept of fatty acids. Lipids are widely acknowledged for their crucial role in forming cellular structures and mediating various physiological and life-sustaining processes. The concept popularity of a lipid layer’s existence on the cell’s surface can be traced back to Overton’s reports between 1885 and 1899, although a comprehensive elucidation of the membrane structure did not emerge until 1925 [14]. It was Gorter and Grendel who, employing a Langmuir monolayer, initially identified the presence of a lipid bilayer within blood chromocytes. Their discovery revealed a distinctive 2:1 ratio between the cellular surfaces covered by lipids and the estimated total cell surface area [15]. Consequently, a lipid bilayer emerges as a supramolecular matrix comprising two leaflets of lipid molecules residing within the biomembrane. Each leaflet necessitates a specific lipid composition characterized by certain physicochemical properties to finely modulate targeted functions.
Despite enduring exposure to changing conditions of temperature, pressure, and solvents, the lipid composition of animal cell membranes remains relatively stable, indicating a relatively confined capacity for drastic alterations in response to external stimuli. Nonetheless, the layers of membranes remain far from static; elements can transfer within (lateral diffusion) and between (vertical or flip-flop diffusion) leaflets. Lipid transporter proteins, namely, flippase, floppase, and scramblase, mediate the movement of lipids across membrane layers. In contrast, the retrograde traffic is responsible for the backward movement of lipids from membranes to organelles [2,16]. The ER, mitochondria, and Golgi apparatus are responsible for biosynthesizing most of the lipid classes in biomembranes, including glycerophospholipids, cholesterols (CHOL), and SLs. Conversely, lipid hydrolysis transpires within the lysosome, specifically the intralysosomal luminal vesicles, where numerous water-soluble hydrolases are active [17,18,19]. Lipids are transported to lysosomes through endocytic and autophagocytic pathways. The products generated from lipid hydrolysis are either utilized within the cell or expelled via exocytosis at the plasma membrane.
Thousands of lipid structures have been identified in mammals [20], with the coexistence of hundreds within a single cell remaining a probable [21]. The chemical properties of membrane lipids are characterized by distinctive features. These include the head-group or backbone structure, hydrocarbon chain length, degree of unsaturation, the presence of chirality, ionization, chelating power, and lipid concentration. Nevertheless, lipid classification is not arbitrary and can be predicated on physical properties, chemical properties, or biosynthetic qualities [22,23]. Within mammalian cell membranes, the preponderant lipid class is glycerophospholipids, also known as phospholipids. Characterized by a hydrophilic head group lining surfaces and a hydrophobic tail interposed in between, this class constitutes the bulk of the membrane lipid matrix. Other minor lipid classes recognized within biomembranes include glycolipids and sterols, with plasma membranes distinctively characterized by a considerable abundance of sterols. A schematic representation delineating the principal lipid classes identified in biomembranes is available in Figure 1. It is well-established that the lipid composition of biomembranes exhibits variations across organelles [24,25] and tissues; it dynamically adapts within the cell in response to specific cellular activities. The distinctive biophysical state of membrane lipids and the fatty acid composition may influence membrane rigidity, serve specific functions, and reveal the cell’s physiological state.
2.1. Glycerophospholipids
In 1811, the pioneering work of Vauquelin led to the identification of phosphorus in cerebral lipid extracts [26], and since then, phosphorus-containing lipids have become an intriguing field of investigation. This class of polar lipids is commonly referred to as ‘glycerophospholipids’ or simply “phospholipids”. It is the most prevalent lipid class in mammalian membranes, accounting for 50–60 mol% of the overall membrane lipid matrix [27]. The foundational structure of phospholipids closely resembles that of diacylglycerol (DAG, featuring a glycerol backbone with two acyl (fatty acid) chains at sn-1 and sn-2 positions); it is further distinguished by the inclusion of a polar phosphorus group at the sn-3 position. Hence, lipids within this class exhibit amphipathic properties, which are characterized by the presence of a hydrophilic head group and two hydrophobic fatty acids.
Over the past century, a multitude of phospholipid types have been identified, with variations in lipid structure playing a profound role in the differentiation of phospholipid varieties. The bulk of phosphate groups are attached to specific molecules or moieties, determining the exact type of phospholipid and its position within the lipid bilayer. Numerous phospholipids have been identified in mammalian membranes, including phosphatidic acid (PA), phosphatidylglycerol (PG), phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylserine (PS), phosphatidylinositol (PI), diphosphatidylglycerol (DPG), bis(monoacylglycero)phosphate (BMP), platelet-activating factor (PAF), and lysophospholipids (LysoP).
2.1.1. Phosphatidic Acid
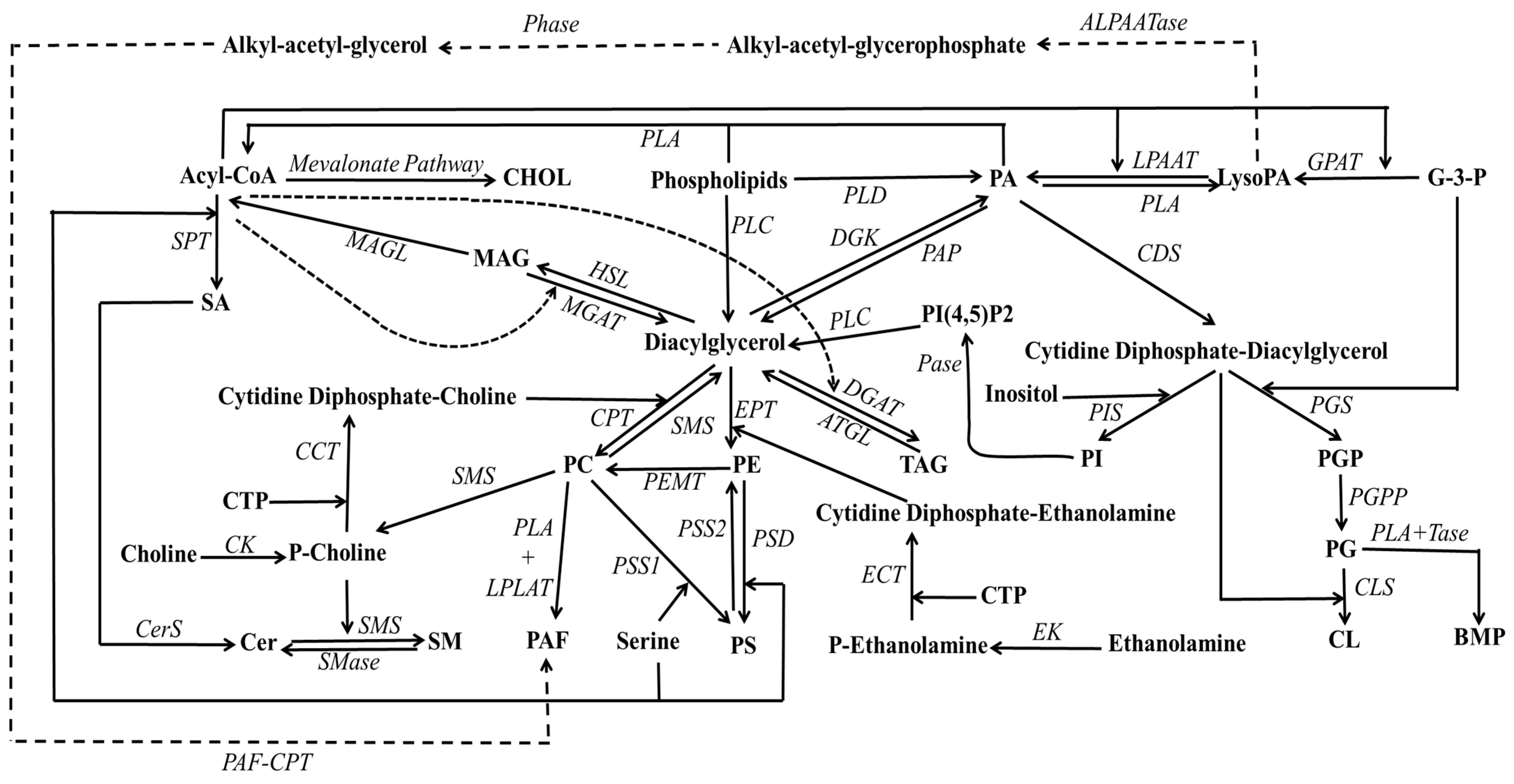
PA, often referred to as phosphatidate (see Figure 1), represents the simplest phospholipid structure and tends to accumulate in membranes in relatively minor proportions, owing to the activity of lipid phosphate phosphohydrolases [28,29]. It was initially identified as a phosphorylated isomer of DAG [30]. PA, therefore, constitutes a non-bilayer lipid characterized by a phosphate group esterified at the sn-3-hydroxyl of the glycerol backbone and two fatty acyl chains occupying the remaining sn-positions. Multiple pathways contribute to PA production (see Figure 2), including the dual acylations of glycerol-3-phosphate, phospholipid hydrolysis pathway (especially involving PC), and DAG phosphorylation [31]. The synthesis of PA from DAG is a reversible process catalyzed by DAG kinase and PA phosphatases (also referred to as lipins).
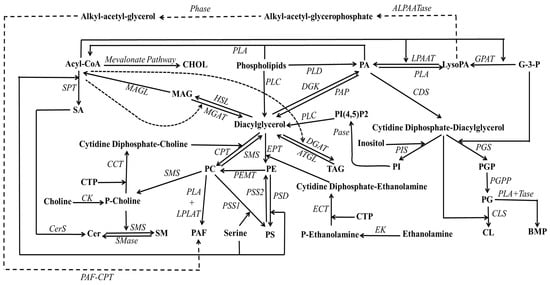
Figure 2.
Schematic representation of the synthesis pathways for various phospholipids available in biomembranes. Abbreviations: ALPAATase, alkyl-acetyltransferase; ATGL, triacylglycerol lipase; CCT, cytidine 5′-triphosphate:phosphocholine cytidylyltransferase; CDP, cytidine diphosphate; CDS, CDP-DAG synthase; Cer, ceramide; CerS, ceramide synthase; CHOL, cholesterol; CK, choline kinase; CL, cardiolipin; CLS, cardiolipin synthase; CPT, CDP-choline:DAG cholinephosphotransferase; CTP, cytidine 5′-triphosphate; DG, diacylgelycerol; DGAT, DAG acyltransferase; DGK, DAG kinase; ECT, cytidine 5′-triphosphate:phosphoethanolamine cytidylyltransferase; EK, ethanolamine kinase; EPT, CDP-ethanolamine:DAG ethanolaminephosphotransferase; G-3-P, glycerol-3-phosphate; GPAT, glycerophosphate acyltransferase; HSL, hormone sensitive lipase; LysoPA, lysophosphatidic acid; LPAAT, lysophosphatidic acid acyltransferase; MAG, monoacylglycerol; MAGL, monoacylglycerol lipase, MGAT, monocylglycerol acyltransferase; P-Choline, phosphocholine; P-ethanolamine, phosphoethanolamine; PA, phosphatidic acid; PAF, platelet activating factor; PAF-CPT, platelet activating factor cholinephosphotransferase; PAP, phosphatidic acid phosphatase; Pases, phosphatases; PC, phosphatidylcholine; PE, phosphatidylethanolamine; PEMT, phosphatidylethanolamine N-methyltransferase; PG, phosphatidylglycerol; PGP, phosphatidylglycerophosphate; PGPP, phosphatidylglycerophosphate phosphatase; PGS, phosphatidylglycerophosphate synthase; Phase, phosphohydrolase; PI, phosphatidylinositol; PIS, phosphatidylinositol synthase; PLA, phospholipase; PLC, phospholipase C; PLD, phospholipase D; PS, phosphatidylserine; PSD, phosphatidylserine decarboxylase; PSS, phosphatidylserine synthase; SA, sphinganine; SM, sphingomyelin; SMase, sphingomyelinase; SMS, sphingomyelin synthase; SPT, serine palmitoyltransferase; TAG, triacylglycerol; Tase, transacylase.
PA is a negatively charged anionic lipid involved in cellular signal transduction and capable of reacting with divalent ions such as Ca2+. Furthermore, its presence within mammalian cells is vital, as it acts as a mediator for phospholipid metabolism, a regulator for glycerolipid metabolism, neuroendocrine cell exocytosis, protein kinases, small G-proteins, and a modulator for membrane fusion and fission machinery [32,33,34,35,36]. Therefore, any alterations in PA levels may indicate disruptions in cellular homeostasis and the onset of metabolic and health-related consequences, as evidenced by Tanguy et al. [31], who linked the high accumulation of PA in cells to metabolic disorders.
2.1.2. Phosphatidylglycerol
When alcohol glycerol esterifies with a phosphate within a phospholipid, the resulting lipid structure is referred to as ‘PG’. Benson and Maruo identified this lipid structure in 1958 [37]; it is characterized by two free hydroxyl groups. Basically, it comprises a glycerol backbone linked with two fatty acyl chains and phosphoglycerol. Within mammals, PG is synthesized in the mitochondria through multiple pathways: (1) it originates from imported PA, which undergoes a series of enzymatic reactions involving intermediates within the cytidine diphosphate-diacyl glycerol pathways in the inner mitochondrial membrane, and (2) from dephosphorylated phosphatidylglycerolphosphate catalyzed by the mitochondrial phosphatase enzyme [38].
Though PG does not constitute a substantial proportion of mammalian membranes (1–2% of membrane polar lipids), it accounts for up to 7–15% of the lipid composition in lung surfactants [39,40]. This heightened presence of PG in the lungs, where it ranks as the second most prevalent phospholipid in the lungs, underscores its crucial role in surfactant activity. Beyond the lung, the PG functionality extends to lipid–protein and lipid–lipid interactions, along with its influence on membrane rigidity. The PG molecular structure relatively resembles that of DPG and BMP, with all of them featuring more than glycerol in their structures. Furthermore, the molecular structure of PG in specific tissues has been considered to be a functional analogue of PI (having an inositol group rather than glycerol) [41]. Thus, these phospholipids may manifest similar activities, such as the inhibition of phosphatidylcholine-dependent kinase activity in swine brain [42]. Elevated levels of PG have been associated with viral infection, as PG can integrate into viral membranes during replication [43,44,45]. In contrast, some reports suggest that PG is involved in regulating innate immunity and suppressing viral infection [46,47,48], potentially including COVID-19 infection [49]. Therefore, further studies are imperative to ascertain the significant biological roles of PG in various mammalian species.
2.1.3. Phosphatidylcholine
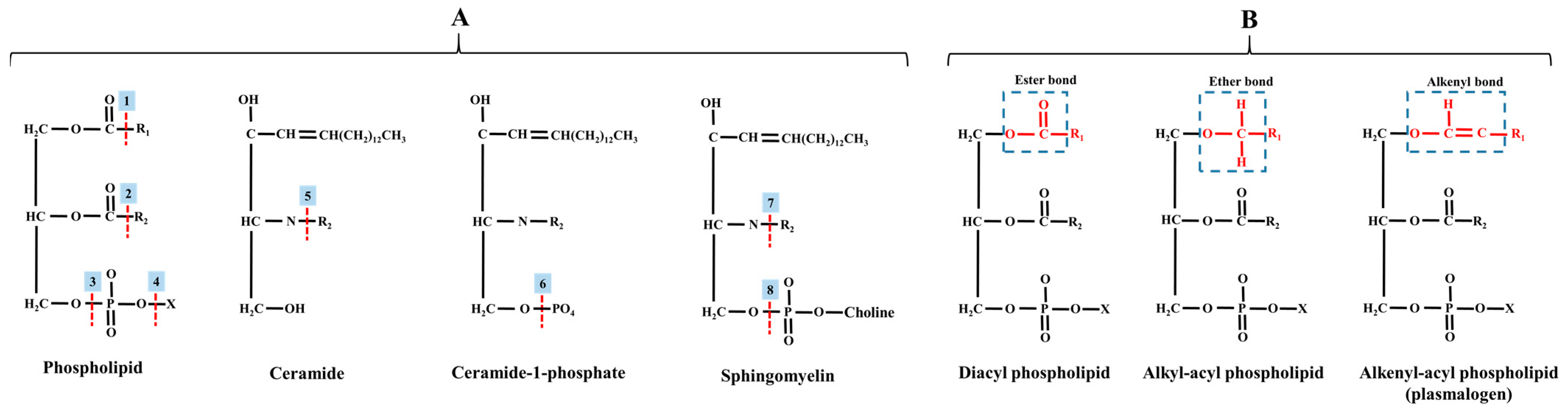
The PC, also known as lecithin, was the first isolated phospholipid in 1850, with choline (a source of the methyl group) serving as the polar head [50]. Herein, it is very self-evident that the PC structure is not entirely endogenous, as choline is an essential nutrient for mammals. PC is a ubiquitous presence in all cell membranes, spanning prokaryotic cells (e.g., bacteria) and eukaryotic cells (i.e., cells of plants and animals). Structurally, PC exhibits two major linkage types in tissues: diacyl-PC (ester bond; most abundant in eukaryotes) and alkyl-PC (featuring an ether bond) [51]. Additionally, the less common isomer of PC is alkenyl-PC (vinyl ether bond), which is referred to as choline plasmalogens and plasmenylcholine. These lipids typically comprise two fatty acids linked to glycerol through ether and ester bonds at sn-1 and sn-2, respectively [52,53].
PC represents the most abundant phospholipid class (constituting nearly 50% of all phospholipids within bilayers), particularly in the pulmonary surfactant, where dipalmitoyl-PC predominates [54,55,56,57]. As a fundamental building block of the membrane bilayer, PC occupies the outer leaflet [58]. Remarkably, approximately 80 to 90% of the lipids in the plasma membrane’s outer leaflet consist of PCs. The preponderance of PC synthesis occurs in the ER, where cytidine 5′-triphosphate (CTP):phosphocholine cytidylyltransfease (PCT) (generally known as CCT) [59,60] catalyzes the rate-limiting step in the cytidine 5′-diphosphocholine (CDP-choline, citicoline or Kennedy) pathway [61]. This CDP moiety is not only involved in PC biosynthesis [62] but in all other phospholipids, with the exception of PA, depending on which moiety replaces choline. A distinctive pathway for PC biosynthesis exclusively takes place in the liver, where PC is generated from PE via sequential methylation [63], facilitated by the phosphatidylethanolamine N-methyltransferase (PEMT) that is found in the mitochondrial-associated membranes (MAM).
It has been believed that PC’s relatively neutral molecular properties (having positive and negative charges but lacking net charge) and its predominance play an essential role in maintaining biomembrane integrity and functionality. Unlike other phospholipids, PC does not exhibit negative charge repulsion. PC serves as a precursor for sphingomyelin (SM) due to its choline molecule [64]. In addition, it acts as a precursor for other polar lipids, such as PA, lysophosphatidylcholines (LysoPC), PS, and PAF. PC plays a crucial part in cell signaling processes and impacts the concentration of circulating lipoproteins [56,65,66]. Furthermore, it is integral to membrane trafficking and molecule transportation. LysoPC composed of C22:6 (at the sn-2 position) has been demonstrated to be more effective than C22:6-free fatty acids in crossing the blood–brain barrier [67].
2.1.4. Phosphatidylethanolamine
Following PC, the second most prevalent phospholipid in mammalian tissues is PE, formerly known as “cephalin”. It was the second discovered phospholipid in cerebral tissue by Thudichum in 1884 [68], constituting approximately 15–25% of the total phospholipids in mammalian cell membranes [69]. In neural tissues, PE can reach even higher levels, up to 45% [70], pointing out its essential role in this tissue. It is profoundly abundant in mitochondrial membranes and is exclusively localized in the cytosolic leaflet of the plasma membrane, in contrast to PC [58]. The structure of PE involves the esterification of the phosphatidyl group to the hydroxyl group of an amino group (namely, the ethanolamine), resulting in a small reactive head group. PE does not form a bilayer independently (on its own) but exhibits an inverted hexagonal phase. This class of lipids features various linkages, including diacyl, alkyl, and alkenyl configurations (see Figure 3). Ethanolamine plasmaologens, also known as plasmenylethanolamine, are more abundant than plasmenylcholine in many tissue types, except for the heart and smooth muscle [52].
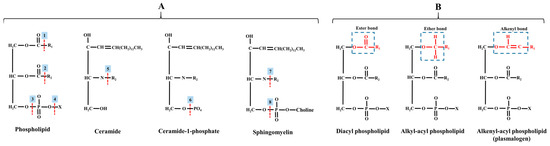
Figure 3.
(A) Site activities of different phospholipases on membrane lipids. (B) Different linkage types in phospholipids. Abbreviations: 1, phospholipase A1; 2, phospholipase A2; 3, phospholipase C; 4, phospholipase D; 5, ceramidase; 6, lipid phosphate phosphatase; 7, sphingomyelin deacylase; 8, sphingomyelinase; R, fatty acid; X, head group.
In eukaryotes, the biosynthesis of PE is an outcome of multiple pathways, notably, the de novo synthesis of PE through CDP-ethanolamine [61] and the salvage pathway involving the decarboxylation of PS by phosphatidylserine decarboxylase (PSD) in the mitochondria [71]. Additional pathways involved in the remodeling of PE, which are also identified in bacteria and plants, include the following: (1) the base-exchange pathway between PE and PS [72]; (2) the degradation of sphingosine-P via sphingosine-P lyase [73]; and (3) the reacylation of Lyso-PE at MAM [74]. Notably, despite the structural resemblance of PE and PC, PE exhibits distinct chemical and biological properties. PE stands apart from PC due to its smaller head group, which manifests less affinity to water. Consequently, PE can form compact aggregation and displays a heightened thermostability [75,76]. These attributes significantly contribute to membrane rigidity, making PE an indispensable component of the membrane’s architecture, permeability, and fluidity.
In terms of membrane rigidity, PE often mimics the behavior of CHOL, particularly in insects [77]. In light of these findings, the PC/PE ratio is likely to exert a substantial influence on the functionality, fluidity, stability [78], and permeability of the membrane. Furthermore, PE plays a vital role in upholding membrane integrity and participating in cellular signaling. Studies have revealed that PE is implicated in various processes, including membrane-to-membrane fusion [79], DAG generation through the involvement of phospholipase C (PLC), and the modification of membrane proteins through the mediation of reactive aldehydes [80]. PE has also been observed to induce negative curvature in biomembranes [81], which is primarily attributed to its diminutive conical head group. In addition, PE serves as a precursor for various other lipids, including N-acyl-phosphatidylethanolamine (NAPE), which serves as a crucial precursor during the biosynthesis of certain essential biological compounds in the brain (e.g., anandamide) [82,83].
2.1.5. Phosphatidylserine
Folch and Schneider identified serine in cephalin components in 1941 [84], marking the beginning of the discovery of PS. PS is a minor class of phospholipids in mammalian cells (2–15% of total phospholipids), which demonstrates a pronounced tendency for accumulation within the cerebral cortex [85,86]. It has also been detected in the membranes of organelles such as mitochondria and ER, where it serves as a substrate for the production of PE. Notably, the distinguishing feature of the PS structure, setting it apart from other phospholipids, is the attachment of the serine residue to the phosphatidyl group, resulting in the formation of a negatively charged head group. This characteristic renders it exceptionally reactive with divalent metals. In contrast to PC and PE, PS exclusively exists in a diacyl isomer, with sn-2 being markedly unsaturated [85].
In contrast to plants [87], yeasts, and prokaryotes [88], mammalian cells lack the de novo CDP-DAG biosynthetic pathway for PS biosynthesis. The biosynthesis of PS in mammalian cells transpires both in the MAM and in the cytosol of the ER and is facilitated by a calcium-dependent base exchange. This pathway is catalyzed by PS synthase-1 and -2 (PSS1 and PSS2, respectively), utilizing PC (catalyzed by PSS2) and PE (catalyzed by PSS2) as the primary precursors at both sites [89]. Subsequent to its production, a fraction of PS translocates to the plasma membrane via passive diffusion. This lipid primarily localizes to the cytosolic leaflet of the plasma membrane [90], although its migration to the outer leaflet is notable during programmed cell death [91] and cancer progression [92].
The externalization of PS on the cell’s outer layer serves as a molecular signal, prompting neighboring cells, including macrophages, to engulf and phagocytose the dying cell. Beyond this role, PS plays a multifaceted biological role within cellular functions. It contributes to the recognition and communication mechanisms between cells. PS existence is crucial during PE biosynthesis, acting as a source pool [71]. Furthermore, PS has been observed to interact with SLs, resulting in elevated interdigitation under the influence of CHOL [93]. PS is also vital for the maintenance of plasma membrane integrity within mammalian cells, exerting modulation over membrane fluidity and permeability, both of which are essential for the regular function of membrane-bound proteins.
PS has been implicated in the activation of protein kinase, prothrombinase, and neuroinflammation signaling pathways, as well as being an essential element of lipid–calcium–phosphate complexes [94,95,96,97]. Consequently, PS facilitates a range of membrane-bound signaling processes, including apoptosis, activation of enzymes, immune regulation, coagulation cascade, and mineral deposition during bone regeneration.
2.1.6. Phosphatidylinositol
The earliest documented report of phosphatidylinositol (PI) traces back to the year 1930 when inositol was initially identified within a lipid extract [98]. It was not until nearly three decades later, in 1959, that Pizer and Ballou elucidated the precise structure of PI [99]. PI, an anionic phospholipid, features a distinctive inositol head group, characterized by a hexa-hydroxy-ring consisting of six carbon atoms. Within the realm of inositol-containing phospholipids (phosphoinositides), PI represents the most elementary form, with the other seven isomers constituting phosphorylated derivatives of the PI structure [100]. In eukaryotic organisms, three primary forms of phosphoinositides prevail: (1) PI, formerly recognized as monophosphoinositide, in which 1′-myo-inositol is linked to PA; (2) PI4P, where a phosphate group esterifies position 4 of inositol, formerly referred to as diphosphoinositide; and (3) PI5P, featuring a phosphate esterifies position 5 of inositol. In eukaryotes, the phosphorylation of positions 2 and 6 of PI is impeded due to steric hindrance. PI can constitute up to 10% of total phospholipids and is ubiquitously present in the cytosol of all cellular membranes and certain organelles (e.g., endoplasmic reticulum and Golgi apparatus) [101,102]. PI of eukaryotic organisms is primarily biosynthesized from PA via a de novo pathway and is catalyzed by the CDP-DAG synthase (which serves as a rate-limiting enzyme [103]) and CDP-DAG myo-inositol 3-phosphatidyltransferase [104]. These enzymes are localized in the ER, where they facilitate the formation of intermediates from PA and the attachment of myo-inositol to these intermediates, respectively. Mammalian cells possess the capability to synthesize inositol de novo from glucose-6-phosphate. Other marked three biosynthetic pathways have been identified in plants and prokaryotes, with the most recent discovery occurring a decade ago [105].
Though PI’s initial discovery was nearly a century ago, our understanding of the biological functions of PI has markedly advanced over the past three decades. PI is not merely a component of bilayer lipids; it is involved in various metabolic processes [106]. Its significance extends to the brain, where it serves critical functions. In addition, it serves as the primary pool of the C20:4 n6 fatty acyl chain in animal cells, frequently occupying the sn-2 position [107,108,109]. This specific acyl chain is of paramount importance for the biosynthesis of eicosanoids, including prostaglandins [101,110]. The enzyme phospholipase A2 (PLA2) is responsible for the removal of C20:4 n6 from PI, resulting in the formation of LysoPI (see Figure 3). Consequently, an accumulation of LysoPI indicates heightened PLA2 activity, implying metabolic alterations and, potentially, the progression of cancer [111].
Furthermore, PI constitutes the major substrate of the signaling DAG in mammalian cells, a process catalyzed by the PLA2 and PLC enzymes. Thus, PI emerges as a dynamic lipid that participates in intracellular signaling, inflammation, and immune regulation. PI also contributes to the formation of glycosyl bridges that facilitate the binding of multiple proteins (known as glycosyl-phosphatidylinositol (GPI)-anchored proteins) to the cellular membrane surface [112]. PI has been shown to engage in regulating protein activities at the cellular interface. The various phosphoinositides, including PI3P, PI4P, PI5P, PI(4,5)P2 (the most abundant PI-phosphorylated structure in mammalian membranes), PI(3,4)P2, PI(3,5)P2, and PI(3,4,5)P3, while accumulating in very low concentrations (1%), significantly contribute to membrane organization. An in-depth discussion has been reviewed by Posor et al. [113]. For instance, PI(4,5)P2 functions as a cofactor for phospholipase D (PLD), an enzyme responsible for the production of PA, which serves as a signaling molecule.
2.1.7. Diphosphatidylglycerol
The DPG, also known as cardiolipin (CL), was initially isolated from bovine hearts by Pangborn in 1942 [114], and the nomenclature “cardio” reflects its association with cardiac tissues. This uncommon tetra-acylated phospholipid is exclusively confined to the inner and outer mitochondrial membranes, with the production site on the matrix side of the mitochondrial inner leaflet [115]. Basically, it can be described as PG with additional phosphatidate groups esterified to glycerol, resulting in two negative charges. The biosynthesis of CL primarily proceeds from the PA substrate [116], which is subsequently transformed into PG within the mitochondria. The conversion of PG species into CL through condensation is facilitated by cardiolipin synthase (CLs). It is postulated that the biosynthesis of CL in eukaryotic cells has evolved from prokaryotic ancestors [117].
CL plays a pivotal role in mitochondrial activity, which is evident through its substantial concentration (15–20%) among the total polar lipids of the mitochondria [118]. Thus, it dynamically contributes to the respiratory chain, interacts with adenosine diphosphate (ADP)/adenosine triphosphate (ATP) and imported complex III and IV proteins, regulates mitochondrial fission and fusion, and controls the release of apoptotic factors [119,120,121]. Therefore, variations in CL concentrations may be associated with mitochondrial dysfunction disorders [119].
2.1.8. Bis(monoacylglycero)phosphate
The BMP is a unique lipid involved in cellular trafficking due to its enrichment in the intracellular membranes of the late endosomes [122,123] and lysosomes [124]. Body and Gary were the first to isolate it from pig lungs in 1967 [125]. While it was initially misidentified as “bisphosphatidic acid” or “lysobisphosphatidic acid”, BMP’s accurate structural characterization was reported by Brotherus and Renkonen in in vitro cultured hamster kidney fibroblast cells [126]. BMP is a negatively charged structural isomer of PG, featuring an unusual sn-1-glycerophospho-sn-1′-glycerol configuration. This lipid structure is related to polyglycerophospholipids, which also encompass PG and DPG [85]. In fact, PG has been identified as the substrate for BMP production [127,128,129], though the precise mechanisms of their production and metabolism continue to be subjects of ongoing research. PG is a fundamental component of mitochondria and ER, and it reaches the lysosome (the BMP biosynthesis site) via autophagy. Herein, the phospholipases are less effective towards BMP, preventing the lysosomal membranes from autodigestion.
The production of BMP involves multiple reactions, including the acylation of fatty acid to glycerol’s hydroxyl moiety and the esterification of phosphoric acid to ethanolamine. Despite BMP constituting a minor fraction of cellular polar lipids, comprising less than 1% of the total [130], elevated levels have been detected in rat splenic tissue [131] and alveolar macrophages [132]. Elevated BMP concentrations have been associated with lipid storage diseases and drug-induced lipidosis [132,133,134]. Studies on BMP have consistently increased over the past 14 years. This interest is attributed to its role in the metabolism of glycosphingolipids (GSLs) and CHOL [19,135], which, in return, influence cellular signaling, vesicle fusion, and membrane integrity.
2.1.9. Platelet-Activating Factor
The PAF is a unique bioactive ether phospholipid with a structural composition of 1-alkyl-2-acetyl-sn-glycero-3-phosphocholine structure, notably lacking the conventional phospholipid sn-1 ester bond [136]. It was initially introduced by Benveniste et al. [137] from rabbit platelets, making PAF the earliest identified phospholipid capable of inciting an inflammatory response. The biosynthesis of PAF occurs within the ER through two primary pathways: the de novo pathway from PC transferred to alkyl acetyl glycerol [138] and a biomembrane remodeling process that involves the substitution of the sn-2 fatty acyl chain with an acetyl group [139]. The latter pathway is catalyzed by PLA2 and acetyltransferase/transacylase.
The accumulation of PAF exhibits variations among cell types, typically representing a negligible fraction of the total phospholipids. This characteristic poses challenges in its precise quantification. The heightened accumulation rate of PAF observed in various tissues correlates with inflammatory responses [140], projecting its major involvement in the regulation of inflammatory and immune responses, as well as physiological processes such as platelet aggregation and thrombosis. In addition, PAF has been documented to exert influence over the activities of multiple physiological systems, including the cardiovascular, nervous, respiratory, excretory, and reproductive apparatuses [141,142,143]. However, alterations in PAF concentrations have been associated with certain diseases, syndromes, and skin cancer [144,145], albeit without serving as a direct mediator.
2.1.10. Lyso-lipids
Shifting the focus to lyso-lipids, this class is alternatively referred to as hydrolyzed lipids. This lipid class is constituted by various isomers originating from the enzymatic cleavage of acyl chains from phospholipids or SLs, which are catalyzed by phospholipase and deacylase enzymes, as illustrated in Figure 3. Thus, lyso-lipids of membranes can be categorized according to their original backbone and further classified into lysoglycerophospholipids (LysoPs) and lysosphingolipids (LsoSLs), respectively. Generally, LysoPs are amphipathic molecules carrying either an alkyl or acyl chain [146,147]. On the other hand, the LysoSLs are distinct due to the absence of the amide-linked fatty acid at the 2-amino position of the sphingoid base [148], setting them apart from their parental structure. Long ago, LysoP isomers were considered intermediates in phospholipid biosynthesis or fragments of disrupted cells. Nevertheless, they display distinct properties and functions compared to parental phospholipids. LysoP contributes to cellular homeostasis by engaging in bilayer remodeling and rigidity. Furthermore, specific LysoP molecules can function as ligands for various G-protein-coupled receptors [149], underscoring their involvement in cellular signaling.
While the current review does not emphasize this category due to its limited prevalence and identification in studies, the most abundant LysoP class is lysophosphatidylcholines/lysolecithins (LysoPC). LysoPC is generated through the hydrolysis PC, mainly catalyzed by PLA2. LysoPC possesses physical properties distinct from PC, forming micelles rather than bilayers. It is typically found in minute proportions and plays a role in the mechanism of the autoimmune response [150]. The accumulation of LysoPC within cells has been associated with metabolic irregularities [150] and phospholipid peroxidation [151,152]. Lysophosphatidic acid (LysoPA), the simplest structure within the LysoP category within mammalian membranes, is biosynthesized in most cells through the activity of lysophospholipase-D on LysoPC or via the actions of phospholipases (phospholipase A1 (PLA1) and PLA2) on PA [153]. LysoPA serves numerous functions, including the regulation of cellular differentiation, growth, proliferation, migration, and apoptosis. In the context of inflammation and cancer, it has gained significant attention, focusing on its structural features and the extent of accumulation [154,155].
2.2. Sterols
This category of membrane lipids is named according to its primary constituent, sterol. Alternatively, it can be referred to as steroid alcohol, distinguishing it from phospholipids. Sterols are characterized by a rigid, always trans tetracyclic hydrocarbon ring, a 3β-hydroxyl group, and a flexible side fatty acyl chain as a tail [156]. Thus, sterols possess both hydrophilic properties (represented by the hydroxyl group) and hydrophobic properties (attributed to the ring and fatty acyl chain). Notably, variations in the floppy tail of sterols account for the structural diversity observed across different biological kingdoms. Sterols are primarily found in membranes of animals (cholesterol), plants (stigmasterol or β-sitosterol), and fungi (ergosterol). It is important to note that most bacterial membranes are devoid of sterols. Among mammalian membranes, CHOL is the most commonly encountered sterol and recognized structure. Despite its widespread presence in various organisms, it is noteworthy that certain insect species lack the genes responsible for its biosynthesis [157].
Cholesterol
CHOL is a sterol isoprenoid characterized by a semi-rigid tetracyclic ring composed of three six-membered rings and one five-membered ring. It features a polar 3β-hydroxyl group and an 8-carbon chain attached to the carbon-17 position, while its side acyl chain exhibits conformational flexibility [158,159]. The polar nature of the CHOL group alone prevents CHOL from forming bilayers. However, when synthetic CHOL is combined with a PC head moiety, bilayer formation occurs [160]. Within bilayers, CHOL is asymmetrically distributed, with the majority of sterols (60–70%) located in the inner leaflet [161,162]. CHOL was initially discovered in gallstones by de La Salle in 1858, but it took another decade for researchers to identify it [163].
CHOL can be obtained from the diet or synthesized by the liver (which contributes 50% to total CHOL synthesis) and the ERs of other cells. The biosynthesis of CHOL is regulated by sterol-responsive element binding protein 2 (SREBP2)-cleavage-activating protein, which senses intracellular CHOL and modulates nuclear transcription. Importantly, cells can also import CHOL from the vascular system, where lysosomes recycle the low-density lipoprotein by transferring CHOL to the ER. The CHOL biosynthesis pathway involves a series of enzyme-catalyzed reactions generating a series of intermediate compounds. Typically, over 20 enzymes are involved in the CHOL biosynthesis pathway, using acetyl-CoA as a substrate. Though animal cholesterol is synthesized in the ER, a relatively higher proportion is found in cellular plasma membranes than in the ER [2,164]. Notably, the plasma membrane contains a significant amount of CHOL (making up to 50% of membrane lipids, primarily in the cytosolic leaflet) as compared to other subcellular organelles [162,165]. In the cytosolic leaflet, the hydroxyl group and the aliphatic chain are oriented towards the aqueous phase and the bilayer’s interior, respectively.
CHOL plays an important role in modulating dynamic membrane activities [156]. Its fused ring structure (exhibiting amphiphilic properties) imparts greater membrane rigidity. Thus, CHOL levels critically influence membranes’ rigidity, fluidity, and permeability [166,167]. The incorporation of CHOL into membranes reduces permeability to non-polar molecules while increasing the hydrophobic barriers to polar molecules. CHOL also has a condensing effect on hydrocarbon chains, thereby reducing the surface area occupied by lipids [168]. Additionally, CHOL participates in the formation of lipid rafts through interactions with various phospholipids, with a notably favorable interaction observed with saturated phospholipids [169]. The solubility of CHOL in membranes depends on the degree of unsaturation of the phospholipids. A high number of unsaturated double bonds has been shown to reduce CHOL solubility [170,171]. Remarkably, even among saturated phospholipids, CHOL affinity was shown to be different. CHOL’s affinity to other lipid complexes relies on various factors, such as the head group tilt structure [172], hydration [173], acyl chain order [174], possible interdigitation of acyl chains [175], and the presence of hydrogen bond acceptor and donor groups [176].
CHOL serves a wide range of signaling functions through its interactions with various cellular molecules and receptors. A recent study indicated that the interaction between cholesterol and lipid-mediated innate immune memory triggers cytokine cascades as associated with COVID-19 [177]. Conversely, an imbalance in membrane-CHOL levels may likely pose severe metabolic consequences, including cancer progression [178,179]. CHOL also serves as a precursor for the biosynthesis of bile acids and steroid hormones [180,181], which mediate crucial roles in biological processes, such as carbohydrate metabolism [182,183,184]. Furthermore, CHOL esters play a critical role in transporting fatty acyl chains via lipoproteins in the bloodstream, and these esters are integral components of amphiphilic plasma lipoproteins [185].
2.3. Sphingolipids
SLs constitute a class of lipids that are present in cells in relatively lower proportions, typically accounting for about 10–20% of total cellular lipids. Despite their relatively lower abundance, SLs exhibit significant signaling activities [27]. The bio-functional roles of SLs can be broadly categorized into three areas: firstly, they modulate the physical properties of biomembranes; secondly, they serve as signaling molecules, acting as second messengers or secreted ligands for cell-surface receptors; and thirdly, they mediate interactions between cells and their external environment [186]. Hence, variations in the ratio of SLs can have a substantial impact on cellular activities and overall cellular survival. The initial identification of SLs can be attributed to Thudichum [68], while the elucidation of their structure, namely, the sphingosine (SO) component, was achieved by Carter [187]. Unlike phospholipids, which are glycerol-based, SLs consist of a long-chain sphingoid base as a backbone, with an amide-linked acyl chain attached instead of an oxygen ester. Notably, the hydrophobic chains (fatty acid) in sphingosine (SO) are structurally constant and non-hydrolyzable, distinguishing SLs from the variable and hydrolyzable fatty acids found in phospholipids. Numerous distinct SL structures have been identified, with structural differences primarily based on variations in backbone structure, hydrophobic chain length, and the level of unsaturation.
2.3.1. Sphingoid Bases
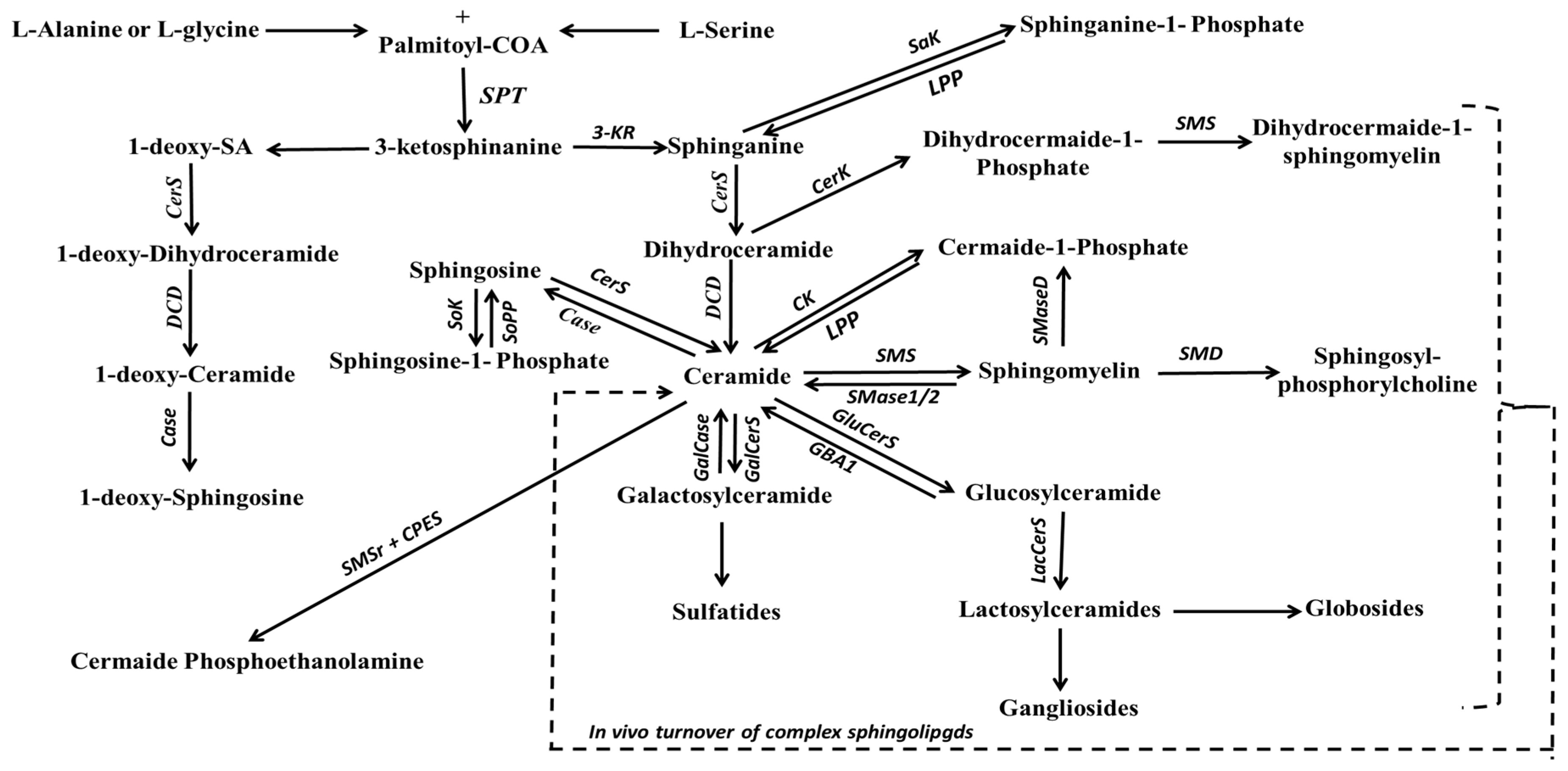
Among the most well-known backbone structures are sphinganine (SA) and SO bases, which serve as the primary reservoir for SL biosynthesis. In the cytosolic side of the ER, serine palmitoyltransferase (SPT) catalyzes the condensation of palmitoyl-Coenzyme A with L-serine, resulting in the formation of 3-ketosphinganine [188]. Subsequently, under the influence of 3-ketosphinganine reductase, 3-ketosphinganine is transformed into SA (as shown in Figure 4). SPT, a pyridoxal 5′-phosphate-dependent enzyme, is the rate-limiting enzyme for SA production [189]. It is worth noting that SPT is not limited to serine alone as a substrate; studies have shown that it can also employ alanine and glycine [190], leading to the production of structures known as 1-deoxysphingolipids. On the other hand, SO is biosynthesized during ceramide (Cer) production or hydrolysis, a process catalyzed by delta-4-desaturase (∆4-desaturase, or D4D) and SPT enzymes, and ceramidase (CDase), respectively. However, it is important to highlight that free SO is not produced via the de novo pathway; rather, it is generated from the hydrolysis of Cer by CDase.
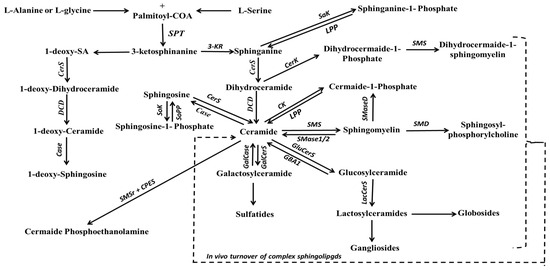
Figure 4.
Schematic representation of the de novo biosynthesis pathway of major complex sphingolipids. Abbreviations: 3-KR, 3-ketosphinanine reductase; Case, ceramidase; CerK, ceramide kinase; CerS, ceramide synthase; CPES, ceramide phosphoethanolamine synthase; DCD, dihydroceramide desaturase; GalCase, galactosylceramidase; GalCerS, galactosylceramide synthase; GBA1, acid β-glucosidase; GluCerS, Glucosylceramide synthase; LacCerS, lactosylcermaide synthase; LPP, lipid phosphate phosphatase; Sak, sphinganine kinase; SMaseD; sphingomyelinaseD; SMD, sphingomyelin deacylase; SMS, sphingomyelin synthase; SMS1/2, sphingomyelin-1 or -2; SMSr, sphingomyelin synthase related proteins; Sok, sphingosine kinase; SoPP, sphingosine phosphate phosphatase; SPT, serine palmitoyltransferase.
SA and SO kinases can phosphorylate SA and SO, generating their respective 1-phosphate forms/derivatives. This pathway is reversible, and sphingoid-1-phosphate produced in this manner can undergo dephosphorylation through sphingoid-1-phosphate phosphatases. Sphingoid-1-phosphate remarkably differs in its activities from free sphingoid bases, serving not only as second messengers but also as first messengers [191]. Sphingoid-1-phosphate also serves as a substrate for phospholipid synthesis, as well as having a universal cellular survival signal [192] that is mediated by its binding to specific G protein-coupled cell surface receptors [193]. Sphingosine phosphate lyase has the capacity to cleave sphingoid-1-phosphate into phosphatidylethanolamine [194,195].
On the other hand, free sphingoid bases are essential secondary mediators, mediating various cellular processes, including growth, proliferation, DNA synthesis, and Cer biosynthesis [196]. These bases can readily traverse membranes, suggesting their potential involvement in stimulus-induced changes in membrane permeability. However, pinpointing the exact signaling functions of sphingoid bases is likely challenging due to their various signals and immense interaction with numerous cellular molecules, such as CHOL, phospholipids, and proteins [197,198,199,200]. It is necessary to highlight that dietary SLs have a proportional direct impact on their detected levels in cellular membranes and tissues [201]. In addition, a number of compounds, such as fumonisin (FUM) mycotoxins and Alternaria toxins [202], share structural similarities with free sphingoid bases, enabling them to interfere with sphingolipid metabolism and alter cellular signaling.
2.3.2. Ceramide
Cer is a non-bilayer-forming lipid characterized by its composition of a sphingosine base and a single acyl chain bonded to an amide group, thus lacking a distinct head group, illustrating its hydrophobic nature. The simple structure of Cer bears a resemblance to DAG. Cer serves as one of the simplest SLs and functions as the core building block for more complex SLs [198,203,204], which have larger molecular dimensions. Cer can be synthesized through multiple pathways: (1) de novo synthesis from SA substrate in the ER (a process catalyzed by N-acyl transferase/ceramide synthase (CerS)) and dihydroceramide desaturase [194]; (2) in vivo turnover of complex SL found in plasma membranes and lysosomes catalyzed by enzymes such as sphingomyelinase (SMase) or non-lysosomal glucosylceramidase) [205]; and (3) the salvage pathway in lysosomes that involves the re-acylation of SO to produce Cer [206].
The key enzyme responsible for de novo Cer generation is CerS, a family of six integral membrane proteins (CerS1–6) located in the ER of mammalian cells and regulated by corresponding six genes situated at distinct chromosomes [207]. The expression of these protein isoforms varies among different tissues [208], leading to variations in Cer proportion and acyl chain lengths. The CerS enzyme is responsible for the formation of dihydroceramide (DCer), which is an intermediate in Cer synthesis. In this step, DCer is formed by acylating a fatty acid to SA, followed by a desaturation reaction catalyzed by DCer desaturase to produce Cer. A decade ago, DCers were commonly considered to be quiescent intermediate metabolites, but recent research has unveiled their distinct functions compared to Cer [209]. Though de novo Cer production takes place in the ER [210], it has been suggested that long-chain bases are acylated in hepatic mitochondria. However, under specific events such as type 2 diabetes and FUM exposure, 1-deoxy-Cer and 1-deoxy-DCer are generated [211,212].
Cer plays a critical role in cellular signaling, regulating cell growth and apoptosis depending on the length of its acyl chain. Specific Cer species, like C16-Cer, have been proposed to be associated with apoptosis rates [213], while C18-Cer has been linked to growth arrest and a proportional decrease during cancer progression [214]. In addition, the ratio between C16 and C24:0/C24:1 Cer species has been identified as a factor in signaling induction, with C16 inducing apoptotic effects and C24:0/C24:1 exhibiting protective effects [215]. Therefore, alterations in the length of the Cer chain may potentially modify signaling, resulting in diverse metabolic effects. Recent review articles have highlighted the connections between Cer acyl chain length and cognitive functions [216] and intracellular lipid regulation [208]. Cer is also known for its ability to cluster apart from membranes, forming ceramide-rich domains with gel-phase properties. These domains are believed to act as platforms for protein–lipid interactions, selectively recruiting or excluding certain membrane components from small transit rafts. Cer-rich domains cluster these components in a stable manner, impeding their in-plane diffusion [217]. Therefore, the high hydrophobicity and complex polymorphic phase behavior of Cer [218] make Cer an important unit in lipid raft formation.
2.3.3. Complex Sphingolipids
In mammals, complex SLs are present in two isomers: phosphosphingolipids (PSLs) and GSLs. Complex SL consists of Cer bonded to complex phosphoryl or carbohydrate moieties, located either in the lumen or at the cytosolic surface of the Golgi apparatus. The transport of Cer between the ER and Golgi organelles is regulated through vesicular and non-vesicular mechanisms, which involve Cer transfer protein [219,220]. This process is primarily coupled by complex SL migrations across membrane leaflets [221] and acts as a rate-limiting factor in the production of complex SLs.
Phosphosphingolipids
In the realm of PSLs, Cer binds to a phosphate group with a polar head group, forming a polar head group mainly composed of choline, ethanolamine, or glycerol. This structural distinction sets PSLs apart from PC in that they not only act as hydrogen bond acceptors but also as hydrogen bond donors. The PSL class includes various subtypes, such as Cer-1-phosphate (Cer1P), DCer-1-phosphate (Dcer1P), Cer phosphoethanolamines (CerPE), sphingomyelins (SM), dihydrosphingomyelins, and LysoPSLs (lacking an attached fatty acyl chain). Among PSLs, SM stands out as the most studied and highlighted class of PSLs in cellular membranes. This review primarily focuses on SM, omitting detailed discussions of other PSLs. However, Cer1P is the simplest PSL with its structure involving the esterification of Cer with the phosphate group. Cer1P serves crucial roles in the regulation of eicosanoids by activating the PLA2 enzyme [198,222].
Sphingomyelin
SM, also referred to as Cer-1-PC, is primarily of animal origin and constitutes a major fraction of SLs in the plasma membrane, accounting for approximately 15% of cerebral lipids [64]. SM is essential for the transmission of nerve impulses. It was initially isolated and described by Thudichum in 1884 [68]. SM is composed of Cer linked to a phosphocholine group [223], a process catalyzed by sphingomyelin synthase (SMS) [224]. Therefore, the overall configuration of SM closely resembles that of PC. SMS is comprised of multiple isomers, including SMS1 and SMS2, each with distinct active sites, with SMS1 situated in the lumen of the Golgi apparatus and SMS2 located on the plasma membrane [225]. SMS is not solely involved in SM production; it also modulates the generation of DAG during de novo synthesis, occurring simultaneously with SM synthesis. SM can also be produced from LysoSM through fatty acid acylation or the straightforward transmission of phosphocholine to Cer [226]. However, the specific enzymes involved in the latter event have yet to be identified.
Similar to PC, SM is primarily located in the outer leaflet of membranes, but it has also been detected in the nuclear envelope membrane [227], mitochondria [228], and liver chromatin [229]. Vesicular transport is the mechanism that facilitates the migration of SM from the Golgi apparatus to the plasma membrane [230], where possible degradation by sphingomyelinase (SMase) may occur, resulting in the generation of Cer. Remarkably, SMS2 catalyzes a contrasting mechanism for SM synthesis in the plasma membrane [224]. Elevated activity of SMase in the plasma has been associated with metabolic dysfunctions and diseases [231]. However, intracellular levels of SM are not exclusively determined by SMS and SMase activities but are also influenced by the dietary uptake of SM. A review by Yang and Chen [232] delves into potential aspects of SM utilization as a dietary supplement.
SM stands apart from PC, despite sharing the same PC head group. Its distinctive characteristics result from a higher proportion of saturated acyl chains and enhanced intermolecular hydrogen bonding capabilities. Unlike PC, SM serves not only as a hydrogen bond acceptor but also as a hydrogen bond donor. Consequently, SM is capable of being involved in various cellular signaling processes, encompassing functions related to proliferation, migration, and apoptosis [233,234]. Previous studies have elucidated how SO and LysoSLs can modulate protein kinase activities [235,236]. Furthermore, both Cer and SM play a role in modulating the uptake of cholesterol esters from high-density lipoprotein (HDL) particles by the target cells [237]. SM also plays a major role in the formation of lipid rafts, engaging in interaction with CHOL to form membrane microdomains [238,239,240], wherein roughly 70% of the cellular total SM is concentrated [241]. This favorable interaction between SM and CHOL can be attributed to the specific attributes of SM molecules, including their elongated saturated chains and reactivity properties (hydrogen donor and acceptor).
Glycosphingolipids
This lipid class closely resembles SM due to their shared origin from Cer. It is commonly referred to as GSL as it distinguishes itself from SM by replacing the complex phosphoryl group with a carbohydrate moiety. GSLs are largely derived from glucose moiety, resulting in the formation of glucosylceramide (GlcCer). In addition, GSLs can also be synthesized from a galactose moiety under the activity of galactosyltransferase, leading to galactosylceramide (GalCer) formation. It is essential to highlight that GSLs vary in their carbohydrate acylation locations. GlcCer is primarily produced at the cytoplasmic surface of the Golgi apparatus, whereas GalCers is made on the luminal side of the ER and is subsequently transported to the Golgi apparatus for further structural modifications to generate various GSLs [242,243]. Within cellular membranes, GSLs are believed to exhibit a preference for partitioning into lipid rafts and are involved in communication with the surrounding environment.
Hundreds of complex GSL structures are currently identified in biological systems, the vast majority of which are gangliosides, which are primarily composed of sialic acid and oligosaccharides [244]. Due to the intricate nature of this lipid class, in-depth classification and discussion have been deliberately avoided. However, two of the simplest GSL structures are glucosylceramide (GlcCer) and GalCer, often referred to as ‘cerebrosides’, featuring either a glucose moiety or a galactose moiety, respectively. The crucial translocation of GlcCer to the luminal leaflet of the Golgi apparatus is an essential step for its subsequent conversion into LacCer, an irreversible pathway involving the addition of a galactose molecule. In addition to these, there exist other GSL complexes such as sulfatides (containing sulfate) and globosides (featuring two or more sugar moieties, typically D-glucose, D-galactose, or N-acetyl-D-galactosamine), which have been identified as GSL derivatives in cellular contexts [245]. For the sake of simplification, scientists have categorized GSLs into two main groups: (1) neutral GSLs, which are characterized by glycosyl groups devoid of acids and remaining unsubstituted, and (2) acidic or amphoteric GSLs, whose glycosyl groups contain one or more sialic acids or a sulfate or phosphate group [246,247].
GlcCer and GalCer function as precursors for numerous complex GSLs, some of which possess additional carbohydrate groups numbering as high as 20 [248]. The addition of these carbohydrate moieties takes place in the Golgi luminal leaflet following the flip-flop translocation of simple GSLs. Majorly, GSLs serve two distinct functions [249]. Firstly, they act as cell receptors to their binding ligands, thereby acting as antigens while facilitating cell adhesion. Secondly, they function as signaling modulators by interacting with other membrane constituents, particularly growth factor receptors. Thus, GSLs play an essential role in immune-cell functions, with a large number of GSL molecules serving as tumor-associated antigens [250,251,252,253].
3. Fatty Acids of Biomembranes
Within the context of biomembranes, the matrix comprises an array of complex molecules, with fatty acids serving as fundamental building blocks. Fatty acids exist typically in two forms: saturated and unsaturated monocarboxylic acids, whereas both are characterized by a terminal carboxyl (-COOH) group and a terminal methyl (-CH3) group designated as carbon 1 (Δ) and omega (ω or n), respectively. Over the past century, numerous nomenclature systems have been proposed, including trivial, systematic, ∆x, n − x, and lipid numbers [22,23]. The trivial nomenclature, though prevalent, lacks systematic patterns. In contrast, the systematic nomenclature adheres to a more regular and structured approach, based on the nomenclature of parent hydrocarbons. It involves adding the suffix “oic” to the hydrocarbon name after removing the terminal “e”. This nomenclature also encompasses the identification of the position of the first double bond from the (n), with the series of fatty acids being named accordingly (e.g., n-3, n-6, n-7, and n-9 series). These distinctions among n-fatty acids lead to variations in their properties, consequently influencing the structure and function of biomembranes [254].
Concerning complex lipids, phospholipids, and SLs addressed in this review, fatty acids play a central role as their primary constituents. Therefore, it is essential to provide a concise overview of their biosynthesis, incorporation into complex molecules, and their biological functions in mammals.
3.1. Synthesis of Fatty Acids
Fatty acids can either be derived from the diet or biosynthesized within the cytosol and ER through an indigenous pathway known as de novo fatty acid synthesis. This synthesis is a complex process influenced by several determinants, including species, transcription genes, dietary composition, age, gender, stored lipids, and both endogenous (metabolic and interactive molecules) and exogenous (environmental) factors. A multitude of genes regulate the synthesis of fatty acids, which can vary among different species. In eukaryotic organisms, nearly 5% of the overall genes are responsible for a significant proportion of lipid structures [255]. Remarkably, the liver X factor (LXR) contributes to the regulation and synthesis of saturated, mono-, and polyunsaturated fatty acids by targeting their transcriptional genes [256]. It also indirectly influences encoding factors involved in lipogenesis, such as sterol regulatory element-binding protein 1c (SREBP1c) [257], peroxisome proliferator-activated receptor gamma (PPAR-γ) [258], and carbohydrate response element-binding protein (ChREBP) [259].
Numerous organisms can produce a wide variety of fatty acids, but only a limited number of molecular structures are synthesized in significant quantities at the natural physiological rate [260,261]. Generally, the synthesis activity of fatty acids is relatively low in normal adult cells, with the exception of certain tissues, including the brain, liver, adipose, and lungs [262,263]. The liver, known as a lipogenic organ, is predominantly responsible for the de novo pathway, although the white adipose tissue (which consists of lipogenic cells) and mammary glands in animals and humans also possess the capability to produce fatty acids through de novo lipogenesis [264,265,266,267]. Under conditions of energy equilibrium, the liver takes up a remarkable proportion (30–50%) of free fatty acids continually absorbed from the diet. These assimilated lipids are either directly incorporated into phospholipids and triglycerides (TAGs) or subjected to modifications (including elongation and/or desaturation) to produce new/modified fatty acids.
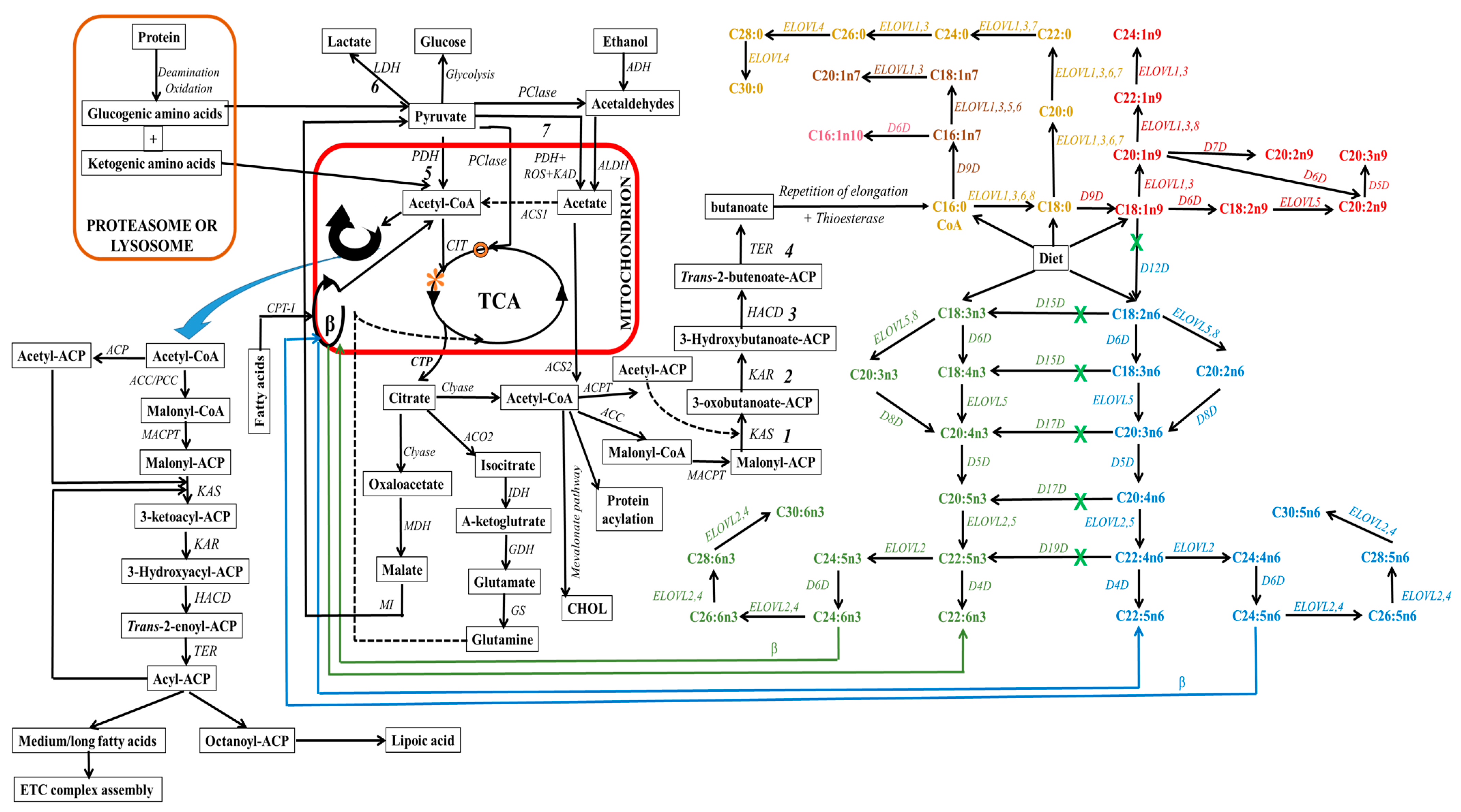
Lipogenic cells can synthesize fatty acids endogenously from non-fat molecules, such as glucose or amino acids (see Figure 5). In this process, pyruvate, a metabolite of glucose, enters the mitochondria, where it undergoes oxidative decarboxylation through the pyruvate dehydrogenase complex to form acetyl-CoA. Within the mitochondria, acetyl-CoA can also be derived from the degradation of ethanol, proteins (deamination and oxidation), and fatty acids undergoing β-oxidation. Subsequently, these produced acetyl-CoA enter the tricarboxylic acid cycle (TCA, citrate cycle, or Szent–Györgyi–Krebs cycle). This event is very crucial since mitochondrial acetyl-CoA molecules are not permeable to mitochondrial membranes. Thus, they are initially located within the mitochondria, whereby they endure a condensation reaction with oxaloacetate to form citrate, a process catalyzed by citrate synthase during TCA [268,269]. The citrate molecule is then expelled from the TCA cycle to the inner mitochondrial membrane, and subsequently to the cytosol. This citrate transporting event requires a dicarboxylate antiporter solute carrier family 25 (SLC25A1) [270].
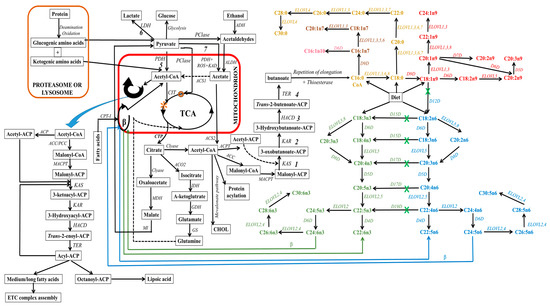
Figure 5.
Schematic depiction of eukaryotic fatty acid biosynthesis and modification pathways, with emphasis on enzyme involvement (in italics). In this schematic, mitochondria are denoted by the red box, while proteasome/ribosome components are indicated by the orange box. However, light orange text delineates the elongation pathway for diverse saturated fatty acids. Within the diagram, both green and blue hues, accompanied by corresponding colored arrows, elucidating the discrete pathways for n3 and n6-fatty acid synthesis, respectively. On the other hand, text with color spectrum transitioning from pink to red designates the pathways for the synthesis of various monounsaturated fatty acids. The green “X” indicates the unattainability of this pathway in mammals, particularly higher eukaryotes, owing to the absence of a specific enzyme. ELOVL8 is a fish-specific elongase. Abbreviations: 1, condensation; 2, reduction; 3, dehydrogenation; 4, reduction; 5, aerobic conditions; 6, hypoxia or anaerobic conditions; 7, aerobic conditions; ACC, acetyl-CoA carboxylase; ACP, acyl carrier protein; ACPT, acyl carrier protein transacylase; ACS1, acetyl-CoA synthetases-1; ACS2, acetyl-CoA synthetases-2; ADH, alcohol dehydrogenase; ALDH, aldehyde dehydrogenase; ACO2, aconitase; β, beta oxidation; CHOL, cholesterol; CIT, citrate synthase; Clyase, citrate lyase; CPT-I, carnitine-palmitoyl transferase-I; CTP, citrate transporter protein, EAR, enoyl-ACP reductase; ETC, electron transport chain; GDH, glutamate dehydrogenase; GS, glutamine synthetase; HACD, β-hydroxyacyl-ACP dehydrase; IDH, isocitrate dehydrogenase; KAD, keto acid dehydrogenase; KAR, β-ketoacyl-ACP reductase; KAS, β-ketoacyl-ACP synthetase; MACPT, malonyl-CoA:ACP transacylase; MDH, malate dehydrogenase, MI, malic enzyme, PCase, pyruvate carboxylase; PCC, propionyl-CoA carboxylase; PDH, pyruvate dehydrogenase; ROS, reactive oxygen species; TCA, tricarboxylic acid cycle; TER, trans-enoyl-ACP reductase; O, oxaloacetate; *, citric acid).
In the cytosol, citrate can undergo distinct metabolic pathways to generate various metabolites (see Figure 5). For instance, ATP-citrate lyase enzymatically cleaves citrate into acetyl-CoA, which leads to the carboxylation of acetyl-CoA and the formation of malonyl-CoA. Within cellular cytoplasm, acetyl-CoA (an active form of acetate) and malonyl-CoA serve as the primary substrates that initiate the process of carbon chain elongation. It is worth noting that propionyl and short-branched acyl units for priming can also be utilized in specific cases. This occurs, for instance, when adipose tissue contains monomethyl-branched fatty acids [271]. Additionally, the elongation process during the synthesis of branched fatty acids in specific glands requires the incorporation of methylmalonyl units [272].
It is imperative to emphasize the critical role played by the acyl carrier protein (ACP), which binds to acetyl-CoA and malonyl-CoA, a process catalyzed by malonyl-CoA:ACP transacylase. This integration event facilitates cytosolic elongation in higher eukaryotes by sequentially transferring these substrates from one enzyme/enzyme domain to another throughout sequential biosynthetic cycles. This active participation of ACP is vital for fatty acid biosynthesis and the functions of fatty acid synthase (FAS), which is a multi-enzyme system regulated by the encoded FASN gene. This cytosolic de novo pathway comprises a series of reactions catalyzed by acetyl-CoA carboxylase (ACC) and FAS, which serve as rate-limiting enzymes [273,274,275]. These reactions include ATP-dependent carboxylation of acetyl-CoA to form malonyl-CoA, Claisen condensation to extend malonyl-ACP and form 3-oxobutanoate, ketoreduction to yield 3-hydroxybutanoate, dehydration to yield butenoate, enoyl reduction to yield butanoate, and repeating elongation reactions (see Figure 5). This process results in the elongation of carbon chains up to the length of C16 or C18 in the cytosol [276,277,278].
ACC, the rate-limiting enzyme in the de novo pathway, facilitates the irreversible decarboxylation of acetyl-CoA through the addition of CO2 to produce malonyl-CoA. The resulting malonyl-CoA attaches to ACP and also serves as a two-carbon donor within a cyclic sequence of reactions facilitated by FAS, leading to the generation of a variety of fatty acid species. The end products (acyl-ACP) of cytosolic de novo synthesis are primarily palmitic acids (C16:0), with lower extents of myristic (C14:0) or stearic (C18:0) acids originating from acetyl-CoA. The determination of chain length during cytosolic de novo biosynthesis involves three enzymes: acyltransferases, ketosynthases, and thioesterases [279]. It is essential to highlight that there are two ACC isoforms: ACC1 (also known as ACCα), which is highly expressed in adipose and hepatic tissues, and ACC2 (also known as ACCβ), which is highly expressed in the heart and skeletal muscles [280]. The mechanism for the conversion of acetyl-CoA to malonyl-CoA by ACC is suggested to differ depending on the ACC type due to their different expression locations [281]. ACC1 is a cytosolic enzyme, whereas ACC2 is located in the outer mitochondrial membrane. On the other hand, FAS enzymes are exclusively cytosolic and can catalyze the formation of C16:0 from acetyl-ACP (substrate) and malonyl-ACP (2-carbon donor) [274,275]. FAS comprises numerous large-multifunctional protein domains (type I FAS) in eukaryotic and specific bacteria, while a monofunctional polypeptide domain is present in the case of major bacteria (type II FAS) [279,282,283,284,285].
In animals, the FAS pathway undergoes termination through a process involving a thioesterase, resulting in the liberation of the free fatty acid as the final product. The termination of the repeating elongation process exhibits the greatest degree of variation in fatty acid biosynthesis. Nevertheless, the conversion of the cytosolic elongated product to the CoA-ester is vital for further biosynthetic pathways that generate new fatty acid structures.
3.1.1. Elongating Fatty Acids through Non-Cytosolic Mechanisms
Generally, the incorporation of the product into lipid structures or its involvement in subsequent elongation and/or desaturation processes is contingent upon the specific requirements of the organism at a given time. Further elongation pathways are not exclusive to fatty acids derived from de novo fatty acid synthesis in the cytosol; they also act on fatty acids derived from the diet, further extending and/or desaturating them to produce longer saturated, monounsaturated, or polyunsaturated fatty acids that are vital for all biomembranes. Following the production of palmitic acid in the cytosol, further modifications of this fatty acid may occur within cell organelles. In mammals, these modifications involve elongation (chains of 18 carbons or longer) and/or desaturation (formation of monenoic/monounsaturated fatty acids).
The ER and mitochondria regulate the modification (elongation) of fatty acids [278] in order to provide sufficient specific signals and functions. Already existing saturated fatty acids are elongated by the sequential addition of two carbon atoms, resulting in the formation of new fatty acids [286]. The principal fatty acid elongation pathway at the cytosolic side of the ER involves a series of four independent reactions: (1) condensation, (2) reduction, (3) dehydration, and (4) a final reduction step [287]. Major enzymes involved in the elongation process include 3-ketoacyl-CoA synthases (elongase enzymes or ELOVLs for reaction 1), 3-ketoacyl-CoA reductase (for reaction 2), hydroxyacyl-CoA dehydratase (for reaction 3), and trans-2,3-enoyl-CoA reductase (for reaction 4). The ELOVL family, sometimes referred to as type III FAS, serves as the rate-limiting enzyme family in the elongation pathway. This family consists of seven subtypes in mice, rats, and humans, and their regulation is governed by ELOVL-encoded genes.
ELOVLs catalyze the condensation of acyl-CoA and malonyl-CoA, which is responsible for elongating fatty acids and determining their carbon chain length, thereby influencing the cell’s fatty acid composition and signaling. ELOVLs exhibit variation in substrate specificity, tissue distribution, and regulation [288]. Based on their final products (see Figure 5), ELOVLs are classified into groups: ELVOLs that elongate saturated and monounsaturated fatty acids (ELOVL1, 3, 6, and 7), ELVOLs that produce very long-chain polyunsaturated fatty acids (ELOVL2 and 4), ELVOL5, which acts on a wide range of substrates with carbon chains ranging from 16 and 22, and ELOVL8, which acts on a wide range of substrates with carbon chains ranging from 16 to 20. ELOVL8 is a distinct subtype that has been recently discovered but is believed to be specific to fish [289]. It is widely acknowledged that these genes are primarily regulated at the transcriptional level; however, additional regulatory mechanisms may exist, including allosteric inhibition. In mammals, ELOVL4 is the sole enzyme capable of catalyzing the formation of fatty acids with more than 26 carbons [290]. These polyunsaturated fatty acids with more than 28 carbon atoms are primarily found in the retina, brain [291,292,293,294], testis [295,296], spermatozoa [297], epidermis [298], meibomian gland [299,300], and Vernix caseosa [301].
An additional pathway for elongating fatty acids occurs in non-cytosolic fatty acid synthesis, especially in the mitochondria. Both animals and yeasts possess mitochondria that contain FAS II enzymes (mtFAS II), which appear to interact with ACP-linked molecules [302,303]. It should be noted that the ACC enzyme has not been identified in the mitochondria of most mammalian species, including humans. However, a recent isoform of ACC1 has been identified in the mitochondria of mice [304]. Thus, isoforms of ACC1, and potentially mitochondrial propionyl-CoA carboxylase [305], are believed to regulate the decarboxylation of acetyl-CoA to yield malonyl-CoA within mammalian mitochondria. Nevertheless, these reactions occur at a limited rate, suggesting that imported malonate may play a role in mitochondrial fatty acid synthesis.
Mitochondrial fatty acid elongation relies on nicotinamide adenine dinucleotide phosphate (NADPH)-dependent enoyl-ACP reductase, with acetyl-ACP and acyl-ACP serving as substrates [306]. This pathway appears to be energetically unfavorable and represents a minor pathway in eukaryotes [278], primarily contributing to the generation of fatty acids used in the lipogenesis of mitochondrial membranes and cellular respiration. Unlike animals, where thioesterase-mediated termination is involved, mitochondrial termination entails channeling the mitochondrial acyl-ACP into the lipid biosynthetic pathway [307]. The primary generated product of mtFAS II activity is an octanoyl chain, which serves as a substrate for lipoic acid synthesis—a vitamin that acts as a scavenger for free radicals [308,309,310] and enhances energy metabolism as a cofactor [311]. Although this pathway can also generate medium and long fatty acids [312], their exact biological roles remain uncertain. However, studies by Nowinski et al. [264] and Angerer et al. [313] have suggested that these mitochondrial long-chain fatty acids are involved in the electron transport chain (ETC) complex assembly.
Furthermore, a similar pathway for fatty acid elongation is proximal fatty acid elongation, which is characterized by reversible β-oxidation. In this pathway, acetyl-CoA acts as the carbon donor, and peroxisomal trans-2-enoyl-CoA reductase substitutes acyl-CoA dehydrogenase to facilitate a thermodynamically favorable reaction [314]. It is important to emphasize that CoA is implicated in the fatty acid catabolism of the reversible pathway, while ACP plays a role in mitochondrial fatty acid elongation. However, the precise functions of fatty acid elongation within peroxisomes remain insufficiently characterized from a scientific standpoint. In practice, the extent of elongation is typically assessed using the elongase estimated index, which is determined by the ratio of C16:0 to C18:0.
3.1.2. Desaturation of Fatty Acids
Within the context of de novo fatty acid synthesis, a process characterized by the removal of two hydrogen atoms to create a double bond often intersects with the fatty acid elongation pathway. This synergy ensures the production of long-chain and very long-chain unsaturated fatty acids, both mono- and polyunsaturated. Notably, the enzymes responsible for fatty acid desaturation, known as fatty acid desaturases, are ubiquitous across all domains of life with the exception of archaea, where they are notably absent [315]. However, it is of significant importance to underscore that the synthesis of polyunsaturated fatty acids can also occur independently of the classical series of desaturase and elongase enzymes. Several studies, including those conducted by Smith and Tsai [316], Kaulmann and Hertweck [317], Napier [318], and Metz et al. [319], have extensively documented an alternative pathway for the biosynthesis of long-chain polyunsaturated fatty acids in both prokaryotes and lower eukaryotes. This alternative route relies on semi-fatty acid synthesis systems, specifically known as polyketide synthases (PKS). PKS employs the same four fundamental reactions as FAS. However, the PKS cycle is frequently condensed, resulting in the formation of highly modified carbon chains featuring numerous keto and hydroxy groups, along with trans-configured double bonds that exhibit various functional roles [320,321].
Among the plethora of desaturase families found in different species, researchers have categorized them into three distinct types, as described by Cerone and Smith [322]. The first family, acyl-acyl carrier protein desaturase, is exclusive to plastids of higher plants. The second family, acyl-lipid desaturases, is found in the ER membranes of higher plants and cyanobacteria. The third family is the family of acyl-CoA desaturases, which can be present in both eukaryotes and prokaryotes, and these enzymes use a cytochrome b5-like system during oxygen reactions [323]. Within the context of this review, with a primary emphasis on mammals, the discussion will be specifically on the acyl-CoA desaturase families. In mammals, a critical in vivo biosynthetic route for the production of long-chain polyunsaturated fatty acids is known as the ‘Sprecher pathway’ [324]. This pathway relies on two fatty acid desaturase enzymes, two ELOVLs, and a peroxisomal β-oxidation process.
The desaturase pathway encompasses diverse enzyme families, such as stearoyl-CoA desaturase (SCD) [325] and fatty acid desaturase enzymes (FADS) [326]. Each of these enzymes acts on distinct substrates. For example, FADS primarily targets polyunsaturated substrates, while SCD predominantly acts on saturated substrates. SCD, also referred to as delta-9 desaturase (∆9-desaturase, or D9D), is an ER enzyme that catalyzes the formation of monounsaturated fatty acids, including palmitoleic (C16:1 n7) and oleic (C18:1 n9) acids, from saturated fatty acids, such as palmitic and stearic acids, respectively. These enzymes exhibit varying specificities and can work on a range of fatty acids with different carbon chain lengths, from 16 to 26 carbon chains. This variation hinges on the specific isoform present, with some species harboring multiple homologues of D9D. For instance, two homologues (D9D-1 and D9D-2) have been identified in marine copepods [327].
Basically, within the D9D reaction, a double bond is introduced at the 9th position from the carboxyl group of the fatty acid. In addition, a multitude of desaturase enzymes present in plants, bacteria, and insects perform the initial double bond introduction on saturated fatty acids at various positions, including ∆3, ∆5, ∆7, and ∆11 [328,329,330,331,332]. The activity of D9D is modulated by dietary fatty acid intake and is subject to up-regulation by the expressions of SCD gene isoforms [333]. Since Bloch’s 1960 pioneering discovery of SCD [315], a plethora of gene isoforms have come to light. In mice, the SCD exhibits four distinct isoforms (SCD-1–4) [334], whereas in humans, only two isoforms (SCD-1 and SCD-5) have been identified [335]. These gene isoforms can vary in substrate preference, ∆ position, and double bond conformation [336]. SCD-1 is the most studied isoform among them [325]. The importance of the SCD pathway is underscored by its involvement in cellular stress, lipid metabolism, inflammation, and autoimmunity. Among the isoforms, SCD-1 is the most extensively studied and is associated with various physiological processes [337,338,339,340,341,342].
Fatty acids featuring a ∆9 double bond are eligible for elongation, but it is important to note that these fatty acids may also be derived from the diet. However, D9D activity is indirectly determined by assessing the ratio between C18:0 and oleic acid. Notably, the de novo elongation of oleic acid results in the formation of longer chains of monounsaturated fatty acids. Furthermore, oleic acid, in particular, may undergo a desaturation process often followed by elongation, where the double bond is introduced before the ∆9 position. Nevertheless, unlike in plants and a limited number of animal species, mammals lack the enzymatic capability to catalyze the introduction of the second double bond in oleic acid, particularly not after the Δ9 position.
Fatty acid desaturase genes (FADs) comprise a gene family responsible for encoding a variety of desaturase enzymes. These enzymes play a significant role in catalyzing the synthesis of polyunsaturated fatty acids by introducing multiple double bonds at positions within fatty acids. Among the genes involved in fatty acid desaturation, mammals have been identified with four distinct FADs [335]: (1) FAD-1, (2) FAD-2, (3) FAD-3, and (4) FAD-6. Each of these FAD types tends to have alternative transcriptions [343,344,345], which can express distinct desaturase activities at different ∆-positions. Generally, FAD enzymes can be categorized into FAD-1, responsible for generating omega-3 (n3) fatty acids; FAD-2, involved in generating omega-x (nx) fatty acids; and FAD-3, which contributes to the formation of omega-6 (n6) fatty acids. While little is known about the substrates of FAD-6 and their major roles have not yet been defined, it appears to be homologous to human FAD-2 [346,347,348] and likely plays a role in the synthesis of polyunsaturated fatty acids.
The transcription factor FAD-6 has been described to possess Δ4-, Δ5-, and Δ8-desaturation activities, with a notable impact on docosahexaenoic acid (C22:6 n3, or DHA) synthesis from n3-docosapentaenoic acid (C22:5 n3, or DPA-n3) in golden pompano fish [349]. On the other hand, Park et al. [345] detailed the existence of nine alternative transcriptions for FAD-3, potentially arising from splicing events. Initially identified through cloning efforts by Marquardt et al. [350], FAD-3 has been traditionally classified as a non-polyunsaturated desaturase, primarily due to its limited substrates, namely, vaccenic acid (C18:1 trans-11) and sphingoid bases. In this regard, it should be noted that FAD-3 may facilitate the unanticipated ∆13-desaturation of trans-vaccenate [351]. These limited substrates give rise to the production of 11E,13Z-octadecadienoic acid [352] and 4E,14Z-sphingodienine [353], respectively.
In contrast to FAD-3, FAD-1 and FAD-2 are the principal contributors to the biosynthesis of polyunsaturated fatty acids in mammals. Nevertheless, mammals lack two crucial desaturases, namely delta-12 desaturase (∆12-desaturase, or D12D) and delta-15 desaturase (∆15-desaturase, or D15D) [354,355,356]. These enzymes are often referred to as “methyl-end desaturases” due to their ability to introduce a new double bond between an existing unsaturated bond and the methyl terminus (–CH3) of the fatty acid. Thus, mammals are de novo incapable of introducing a new double bond after the ∆9 position of oleic acid. Hence, mammals must obtain polyunsaturated fatty acids from their diet, leading to the concept of essential fatty acids. These essential fatty acids, such as linoleic acid (C18:2 n6, or LA) and α-linolenic acid (C18:3 n3, or ALA), cannot be endogenously synthesized by mammals and must be sourced from dietary intake [357,358,359]. Nonetheless, the enzyme activities of FAD-1 (∆5-desaturase, or D5D) and FAD-2 (∆6-desaturase, or D6D) have been identified in mammals. These enzymes are responsible for introducing double bonds at the ∆5 and ∆6 positions, respectively [360,361]. Notably, both D6D and D6D are highly expressed in the liver, with D6D exhibiting particularly high expression levels [362].
In mammals, dietary LA, ALA, and other unsaturated fatty acids, whether from the diet or synthesized endogenously, serve as substrates for the generation of polyunsaturated fatty acids characterized by an increased number of double bonds and longer carbon chains. The enzyme D6D plays a crucial role in the initial steps of synthesizing arachidonic acid (C20:4 n6, or AA), eicosapentaenoic acid (C20:5 n3, or EPA), and DHA by catalyzing the conversion of LA and ALA into γ-linolenic acid (C18:3 n6) and stearidonic acid (C18:4 n3), respectively [362]. However, the biosynthesis of AA and EPA, in particular, involves an additional desaturase enzyme, delta-5 desaturase (D5D), which utilizes the substrates dihomo-γ-linolenic acid (C20:3 n6, or DGLA) and eicosatetraenoic acid (C20:4 n3) to yield AA [360] and EPA [363], respectively. According to Vagner and Santigosa [364], the substrate affinities of D6D appear to exhibit contrasting and debatable characteristics. Consequently, further investigations are imperative to substantiate a definitive conclusion. It is essential to emphasize that the distinctive substrate affinities of D6D play a critical role in determining the ratio of n6- to n3-polyunsaturated fatty acids (referred to as n6/n3 or n6:n3). The role of FAD-6 in determining the ratio of n6/n3-polyunsaturated fatty acids and its influence on the metabolic flux of these fatty acids have been highlighted [365,366].
Remarkably, D6D and D5D are also involved in the synthesis of n9-polyunsaturated fatty acids, specifically, Mead acid (C20:3 n9), which is produced in the absence of LA and ALA [367], when only monounsaturated fatty acids are available as substrates. Elevated levels of Mead acid are matched with the proportional depletion of n6- and n3-polyunsaturated fatty acids and serve as a biomarker for LA and ALA deficiency in diets. For instance, the ratio between trienoic and tetraenoic acids (such as Mead acid:AA) functions as a biomarker for the lack of dietary essential fatty acids [368,369]. Park et al. [370] have identified two pathways for Mead acid production, involving D6D and D5D. These pathways vary in substrates of D6D, which can either utilize oleic acid or gondoic acid (C20:1 n9) to yield C18:2 n9 and C20:2 n9, respectively. D5D catalyzes the direct conversion of C20:2 n9 into Mead acid by introducing a double bond at the ∆5 position. Furthermore, authors have also reported the novel activity of ∆7-desaturase (D7D, regulated by FAD-1), leading to the formation of C20:2 n9 from C20:1 n9.
Notably, AA, EPA, and DHA are biologically essential fatty acids with enormous contributions to membrane phospholipids. In the Sprecher pathway, the primary fatty acid synthetic pathway in mammals, the biosynthesis of DHA from EPA is not direct but rather involves a series of reactions: (1) elongation to a 24-carbon chain, (2) desaturation via D6D, and (3) peroxisomal β-oxidation for chain shortening. However, the synthesis of n6-docosapentaenoic acid (C22:5 n6, or DPA-n6) and DHA in eukaryotes also occurs through ∆4-desaturase (D4D) pathways, depending on the species. In lower eukaryotes, marine vertebrates, and humans, D4D (encoded by the FAD-2 gene), which is expressed to a lesser extent, plays an essential role in introducing a double bond at position ∆4, directly producing DPA-n6 and DHA from docosatetraenoic acid (C22:4 n6, or adrenic acid) and DPA-n3 substrates, respectively [371,372,373,374]. It is noteworthy that this reaction bears resemblance to that facilitated by FAD-6 expression, suggesting the possibility of FAD-6 up-regulating D4D activities.
A further marked expression of FAD-2 identified within mammalian cells is ∆8-desaturase (D8D). This enzyme establishes an autonomous pathway distinct from D6D, introducing an additional double bond to substrates like ALA, LA, and their elongated polyunsaturated fatty acids, resulting in the formation of very long-chain polyunsaturated fatty acids [375,376]. For instance, the emergence of D8D was observed when DGLA and eicosatetraenoic acid were derived from eicosadienoic acid (C20:2 n6) and eicosatrienoic acid (C20:3 n3), respectively [377]. In a study conducted on mouse liver, D8D expressed low activity, which is primarily associated with n3-unsaturated fatty acids, such as eicosatetraenoic acid (C20:4 n3), EPA, DPA-n3, DHA, and nisinic acid (C24:6 n3) [375].
In general, more than a hundred FAD-2-related desaturase enzymes have been identified in various animal species, although they are notably absent in mammals. For instance, the ∆17 (17-desaturase, or D17D) and ∆19 (19-desaturase, or D19D) desaturases have been identified in lower animal families (excluding mammals) and algae [378,379]. These enzymes play an essential role in the production of EPA and docosapentaenoic acid-n3 (C22:5 n3, or DPA-n3) from AA and adrenic acid, respectively. The activities of enzymes involved in lipogenesis are subject to intricate regulation by a matrix of genes and exogenous factors. Notably, polyunsaturated fatty acids have been shown to inhibit the transcription of hepatocellular genes responsible for encoding lipogenic enzymes [380,381]. It is a common practice to assess desaturase enzyme activity indirectly by determining their coefficients based on the ratio of the fatty acids generated to the substrates utilized.
Desaturase enzymes exhibit a broader scope of activities beyond their involvement with fatty acid substrates incorporated into phospholipids, as they are also active in SLs. Ordinarily, SLs are characterized by their predominantly very long saturated or monounsaturated nature. The determination of fatty acid chain length within SLs is intricately governed by the activities of the CerS type [382]. However, it is worth noting that polyunsaturated fatty acids show a slight accumulation in SLs within testes and spermatozoa [383,384] as compared to phospholipids within the same tissues. The classification of desaturases responsible for introducing double bonds into Cer structures has been presented by Nachtschatt et al. [385]. This classification delineates three distinct categories of desaturases: (1) α-hydroxylases [386], (2) D4D and C4-hydroxylases [387], and (3) D8D [388]. These desaturases play a pivotal role in diversifying the composition of SLs, particularly in terms of the introduction of double bonds, which contribute to the functional and structural heterogeneity of these important lipid molecules.
3.2. Incorporation of Fatty Acids into Lipids of Biomembranes
Fatty acid synthesis and their subsequent incorporation into biomembranes represent critical processes in the life of all organisms. The one exception to this rule is the archaea domain, which employs isoprenoids as membrane lipid side chains rather than fatty acids [389]. Understanding how fatty acids are incorporated into membrane lipids is of paramount importance. Fatty acids may become part of phospholipids either through acylation of glycerol-3-phosphate during the biosynthesis of phospholipids or through the action of lysophosphatidic acyltransferases and phospholipases that remodel the structure of pre-existing phospholipids [390,391] (which can be seen in Figure 6).
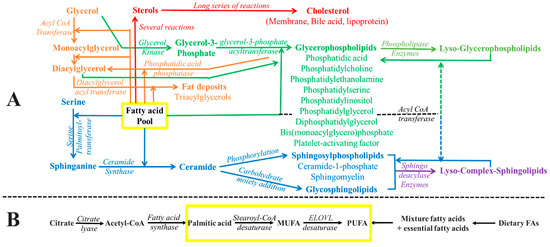
Figure 6.
Schematic representation of (A) fatty acids incorporated into various lipids and (B) the origins of fatty acid pool formation (the de novo pathway and dietary sources). The depiction employs color coding to symbolize distinct metabolic pathways. The yellow box designates the fatty acid pool, signifying its integration into diverse membrane lipids. The brown shade denotes processes related to non-membrane and non-polar lipid formation. The red color represents the metabolic routes responsible for sterol production. Additionally, the green color signifies the integration of fatty acids into various phospholipids, while the blue hue corresponds to the incorporation of fatty acids into diverse sphingolipids.
The vital nature of these processes is underscored by the coexistence of diverse fatty acid structures within complex biomembrane lipids [392]. This structural diversity arises from various factors, including the preferences of sn-positions for particular fatty acids, the substrate preferences of biosynthetic enzymes, and the dynamic lipid remodeling program. Notably, studies by Shindou et al. [393] and Coleman et al. [394] have elucidated the preferences of specific enzymes for distinct fatty acids. For instance, acyl-CoA synthetase long-chain family members 3 and 4 demonstrate preferences for AA and EPA, while 1-acylglycerol-3-phosphate O-acyltransferase-α prefers myristic acid, palmitic acid, and LA. Meanwhile, 1-acylglycerol-3-phosphate O-acyltransferase-β exhibits a preference for AA.
In the context of lipid remodeling, the replacement of fatty acids within existing phospholipids plays a central role. For instance, the incorporation of C20 fatty acids generally takes place post-de novo synthesis, necessitating the employment of the remodeling pathway [395]. Hence, this process involves the conversion of one distinct phospholipid into another [85], highlighting the importance of the Lands cycle in this process. The Lands cycle [396,397] is a central component of the remodeling process, enabling the attainment of specific structural configurations required for the generation of particular cellular signals. In this pathway, phospholipases initiate cleavage (deacylation) of fatty acids from phospholipids, resulting in the formation of free fatty acids and lysophosphatidate. On the other hand, acyltransferases function in a contrasting manner, acylating the requisite fatty acids into lysophosphatidate to generate phospholipids. This process is CoA-dependent, which is primarily due to the extensive utilization of CoA intermediates as substrates. Other remodeling pathways include the CoA-independent transacylation pathway and the direct transacylation pathway, which are specifically suited for highly unsaturated fatty acids such as AA, EPA, and DHA. In these pathways, transacylases catalyze the migration of fatty acids between molecular species of phospholipids. Remarkably, these remodeling pathways exhibit substantial variability across different tissues [395], as comprehensively reviewed [390].
Additionally, it is noteworthy that fatty acids are predominantly incorporated into SLs by the rate-limiting enzyme CerS. However, a remodeling mechanism can also come into play, modulating specific signaling and functional roles. For example, the work of Markham et al. [398] suggests that the accumulation of very long-chain fatty acids contributes to the formation of microdomains through increased hydrophobicity, membrane leaflet interdigitation, and the transition from a fluid to a gel phase. These structural transformations have significant implications for cellular function and signaling within membranes.
The movement of membrane phospholipids between bilayers involves the activities of various transmembrane lipid transporter proteins, namely, flippase, floppase, and scramblase. These proteins play distinct roles in the translocation of lipids and exhibit differential mechanisms of action. For instance, flippase facilitates the translocation of lipids from the exo-leaflet to the inner-leaflet, contrasting with the activity of floppase, which operates in the opposite direction, moving lipids from the inner-leaflet to the exo-leaflet. Notably, both flippase and floppase depend on ATP as an energy source for their functions. Conversely, scramblase functions as an ATP-independent transporter and orchestrates the bidirectional movement of lipids across membranes in a non-specific manner, allowing for the concurrent translocation of lipids from the inner to the outer leaflet and from the outer to the inner leaflet. Comprehensive insights into these membrane fatty acid transporters have been provided by the extensive reviews conducted by Samovski et al. [399] and Glatz et al. [16]. These reviews offer a detailed discussion of the mechanisms and significance of these proteins in lipid translocation processes within biological membranes.
3.3. Fatty Acid Composition in Biomembranes
The fatty acid composition of biomembranes plays a critical role in maintaining cellular homeostasis and ensuring proper functionality. It is worth noting that this composition is not static but can be modified/remodeled in response to homeoviscous adaptation. This process was initially described in algae [400] and has been later extended to non-homeothermic animals. In contrast, mammals are generally considered to have stable biophysical properties and lipid order within cellular membranes, but they can respond to changes in their dietary fatty acid compositions. Furthermore, certain mammalian cells potentially exhibit a lower degree of homeoviscous adaptation in response to the membrane curvature elastic stress [401]. The fact that there is not a single standardized composition for fatty acids in biomembranes should not be viewed as a flaw but rather as an indication of the intricate and dynamic nature of biological systems. The complexity and adaptability of fatty acid composition in biomembranes underscore the potential evolutionary advantage conferred by the ability to alter lipid structures.
The available literature, which will be discussed in subsequent sections, provides extensive data regarding the proportions of specific fatty acids in various lipid complexes within biomembranes. This section merely focuses on the major fatty acids identified in phospholipids and SLs, as these fractions are the central aspects of this review. Regardless, the length and degree of saturation of fatty acyl side chains in CHOL-esters can vary between different cells. This variation is primarily influenced by dietary factors and cell metabolism, and it has a direct impact on the stability and fluidity of the cellular membrane.
3.3.1. Fatty Acid Profile of Phospholipids
Commonly, the acyl chains of phospholipids found at the sn-1 and sn-2 positions of the glycerol backbone are typically composed of a saturated fatty acid (such as C16:0 or C18:0) and an unsaturated fatty acid (with carbon chains of 18, 20, 22, or 24), respectively. Furthermore, these acyl chains exhibit variations in terms of their length, the number of double bonds, and the position of hydrogen atoms adjacent to these double bonds (whether in cis or trans configuration). It is important to note that trans-double bonds are relatively rare within mammalian membranes, while cis-double bonds are frequently abundant. In fact, the occurrence of trans-double bonds in mammals is far less frequent than in bacteria [402,403]. In some instances, identical acyl chains can be found at both sn-1 and sn-2 positions of glycerol [404,405]. It is worth highlighting that even when the number of carbons is the same, a slight mismatch may occur due to the sn-positions. This mismatch is a consequence of the sn-2 chain bending perpendicular to the membrane’s plane [406,407].
In the case of PA, the composition of the two fatty acids can vary across different cell types and constituents. Typically, dietary fatty acids and metabolic disorders play a substantial role in determining the composition of these acyl chains. PA is primarily composed of saturated and monounsaturated fatty acids, with carbon chains ranging from 14 to 24. Concerning PG, this class exhibits variations in the length and number of unsaturation of its fatty acids, depending on the cell type and the dietary fatty acids involved. In the PG of rat lungs, palmitic acid is the most abundant fatty acid, with unsaturated fatty acids constituting a smaller portion [408]. PG is unique among phospholipids due to its higher degree of unsaturation, with unsaturated fatty acids more likely to occupy the sn-1 position. Research by Xie et al. [409] suggests that the degree of unsaturation in PG may have varying effects on mouse keratinocyte proliferation.
Commonly, the fatty acid composition of PCs is typically determined post-synthesis, with various events, such as deacylation and reacylation, taking place during its remodeling [396,397]. These events, part of the Lands cycle, can also impact the composition of other phospholipids due to homeostatic mechanisms or metabolic implications [410]. Generally, PC exhibits variations in its fatty acid composition across species and cell types [411,412,413,414,415,416]. Saturated fatty acids, like palmitic or stearic acids, are typically abundant at the sn-1 position, while C18 unsaturated chains or longer polyunsaturated fatty acids like AA and DHA are more prevalent at the sn-2 position.
The fatty acid composition of PE is highly dependent on the particular cell, tissue, and physiological conditions. In contexts like chicken egg, rat liver, and brain, palmitic and stearic acids tend to occupy the sn-1 position, while AA, oleic, and DHA are more common at the sn-2 position [411,412,417]. Notably, PE in the erythrocyte membrane exhibits a greater tendency for the accumulation/recruitment of polyunsaturated fatty acids compared to PC [418]. Consequently, PE in this context contains more polyunsaturated fatty acids, primarily AA and DHA, at the sn-2 position, despite its diacyl structure bearing similarities to that of PC.
The composition of fatty acids in PS differs depending on the tissue type. Nonetheless, lipid remodeling and selective insertion of fatty acids are common processes that occur along the biosynthetic pathway. Therefore, the fatty acid composition of newly synthesized PS differs from that of its precursors, PE and PC. When PS was isolated from rat liver and cow brain and analyzed regiospecifically [412,414], it was observed that stearic acid was more abundant at the sn-1 position, while the proportion of palmitic acid was lower here. These data underscore the distinct fatty acid incorporation pattern exhibited by PS in comparison to PC and PE. Furthermore, the acylation of fatty acids at the sn-2 position has been shown to be tissue-specific, with high proportions of AA and DHA in the livers of rats and the brains of bovines, respectively. Similar findings regarding DHA in brain regions of mice and rats have been reported by Kim et al. [419] and Hamilton et al. [420]. However, stearic acid was the most abundant fatty acid at the sn-1 position in these cases. Remarkably, the incorporation of these unsaturated fatty acids into PS appears to be more extensive than in the case of PC, as revealed by these studies. The fatty acid composition of PS at its sn-positions plays a crucial role in determining its functional properties to varying degrees [93,421].
In mammalian cells, the composition of PI is characterized by the presence of stearic acid and AA in the sn-1 and sn-2 positions, respectively, as consistently demonstrated in various studies [411,412,422,423]. These two fatty acids collectively account for a substantial portion of PI acyls, typically ranging from 50% to 80% in the brain and liver. Additionally, oleic acid is frequently identified as the third most abundant fatty acid in the rat brain, while palmitic acid and DHA exhibit lower levels of acylation in this phospholipid. However, it is worth noting that Ulmann et al. [424] reported a distinct fatty acid composition in the rat brain, where oleic acid was the most prevalent, followed by stearic and palmitic acids. This variability in PI composition across studies may be attributed to a range of factors, including dietary influences and genetic variations. In general, PI exhibits a strong preference for AA in its acyl composition. Remodeling processes can lead to the deacylation of pre-existing PI, resulting in the formation of lyso-PI via the action of phospholipases. The incorporation of AA into lyso-PI is facilitated by lysophosphatidylinositol acyltransferase 1 [425].
The structure of fatty acids within CL greatly influences its shape and properties. Saturated chains tend to produce lamellar forms of CL, whereas unsaturated chains contribute to non-lamellar structures [38]. Thus, the distinctive fatty acid composition of CL is believed to play a critical role in its diverse biological functions across different cellular and subcellular membranes. CL remodeling is an essential process for CL formation, during which LA and DHA are primarily incorporated, influencing susceptibility to oxidation [426,427,428,429]. LA has been reported as the majority of CL fatty acids, often accounting for 80–90% of the composition [430]. Although C18 polyunsaturated fatty acids are the predominant constituents of CL, dietary fatty acid intake can influence its composition. Increased dietary supplementation levels of C20 polyunsaturated fatty acids and DHA have been shown to elevate their proportions in mammalian CL, as indicated by Berger et al. [431] and Wolff and Entressangles [432].
The structure of BMP exhibits variations in chain length and the degree of fatty acid unsaturation. Oleic acid is frequently identified as the most abundant fatty acid in BMP [126,132,433]. However, in certain cell types, polyunsaturated fatty acids such as LA and/or EPA and DHA have been reported to be highly accumulative [434,435,436,437]. In alveolar macrophages, for instance, Holbrook et al. [438] noted that oleic acid predominates along with either AA or DHA.
In intact tissues (e.g., neural tissue), palmitic acid typically represents the most abundant fatty acid within the alkyl group of PAF. Stearic and oleic acids may also be present but to a relatively lesser extent. The sn-2 position of PAF is often esterified with the acetyl group or other short-chain fatty acids [439]. However, in rat nervous tissue, sn-2 is predominantly occupied by unsaturated fatty acids, primarily AA and adrenic acid (C22:4n6) [440], indicating the profound impact of sn-2 composition on PAF activities. These longer n6-type fatty acids serve as potent precursors for eicosanoids.
3.3.2. Fatty Acid Profile of Sphingolipids
In general, SLs exhibit a prevalent composition of very long-chain saturated and/or monounsaturated fatty acids, typically ranging from 18 to 34 carbon chains. Notably, some sphingolipid structures have been reported to contain odd-numbered fatty acid chains [201,441], suggesting a possible proportional elevation of these chains in the tissues of ruminants and coprophagous species. Despite the fact that the enzyme SPT utilizes palmitoyl-Coenzyme A to produce SA, it has the capability to utilize various other fatty-CoA substrates as well [442]. Sphingoid bases within SLs predominantly feature saturated aliphatic chains, with some instances of mono- and di-unsaturated chains. SO possesses a fixed trans-double bond between carbons 4 and 5. The chain length of sphingoid bases’ fatty acids typically falls within the range of 14 to 32 carbons [443].
Within the context of Cer, the variability in chain length is attributed to the diversity of CerS isoforms. In most scenarios, CerS5 and CerS6 predominantly provide chains with 14–18 carbons, CerS1, CerS4, and CerS2 contribute chains with 18–24 carbons, and CerS3 supplies chains with up to 34 carbons [382]. Cer features an acyl chain linked to an amide that is frequently saturated and considerably longer compared to those found in SLs containing sugar moieties [240]. The length of the Cer fatty acid is determined by ER elongase and desaturase complexes, rather than the Golgi apparatus, which produces complex SLs. Cer and SM are primarily acylated by palmitoleic and oleic acids, in addition to long and very long-chain fatty acids. In most organisms, including mammals, Cer can contain 2-hydroxylated fatty acids [444,445], which is a characteristic favoring interaction with SMS2 over SMS1 [446].
SM structures are characterized by long fatty acids with relatively high chain length inequality, leading to their interdigitation sensitivity. The cause of interdigitation is not fully understood but may involve the regulation and/or induction of proteins in response to specific fatty acids within SM under certain membrane constituents. In comparison to PC, SM typically contains more saturated and longer acyl chains, typically ranging from 16 to 24 carbons [415,447], and may include odd-numbered fatty acids [448]. The palmitic, stearic, behenic (C22:0), lignoceric (C24:0), and nervonic (C24:1 n9) acids are among the most frequently occurring fatty acids in SM [415,447,448,449,450]. Mammalian germ cells, in particular, are abundant in very long-chain fatty acids with up to 34 carbons [297,451], some of which may exist in a 2-hydroxylated form in certain mammals [452]. Double bonds are relatively rare in SM fatty acids, and, when present, they are often located at a distance from the membrane–water interface [447].
GSLs exhibit a greater variation in the chain length of their fatty acids compared to phospholipids, with some GSLs containing fatty acids with more than 16 carbons [453]. Although it is challenging to detect long and/or unsaturated fatty acids in GSL, a few studies have reported the presence of high proportions of long and very long fatty acids. For instance, stearic acid has been found to be more dominant than palmitic acid in the GSLs of the mouse brain [454]. Furthermore, the majority of GalCer in the brain is composed of very long fatty acids [455]. In gangliosides, lignoceric acid (C24:0) is the most abundant acyl chain, constituting up to 70% of total fatty acids in myelin [456], while stearic acid is the predominant component, making up 80% of total fatty acids in the human brain [457].
4. Fatty Acids and Cellular Functionality
The different chemical structures, physicochemical properties, and physiological functions of various fatty acids exhibit remarkable diversity. These distinctions have led to various categorizations, depending on the specific scientific focus. Traditionally, fatty acids have been recognized for their dual biological functions: as structural components of biomembranes and as sources of energy. However, contemporary perspectives on fatty acids have expanded to encompass their roles as bioactive molecules that contribute significantly to overall health. Pioneering work by Burr and Burr [458] and von Euler [459] underscored the diverse biological functions of fatty acids, particularly LA and ALA, highlighting their crucial roles in cellular signaling processes. This section focuses on fatty acids’ impact on membrane properties and avoids delving into their role in energy provision via β-oxidation (primarily relies on TAGs), which is a subject beyond the scope of this review.
4.1. Influence of Fatty Acids on Bilayer Properties
Fatty acids play an essential role as constituents in cellular membrane assembly, profoundly influencing the physicochemical attributes of these membranes. Biomembranes exhibit a discerning preference for incorporating specific fatty acids, particularly favoring long and very long polyunsaturated fatty acids that contribute to the formation of fluidic membranes. A study by Rodriguez-Estrada et al. [460] has associated long-chain lipid metabolites derived from LA and ALA with the preservation of membrane properties. Sensor proteins typically monitor and regulate the physicochemical properties of membranes [7,461,462]. Fatty acids exhibit variations in chain length and degree of unsaturation across different cellular contexts, catering to specialized functions. For instance, the study of Matveyenka et al. [463] has highlighted the correlation between the rate of insulin aggregation and the length and degree of the unsaturation of fatty acids. Therefore, maintaining equilibrium among various fatty acid species (saturated, monounsaturated, and polyunsaturated) within membranes holds a position of critical importance, as any deviations from this equilibrium could lead to modifications in membrane integrity and cellular metabolic signaling. Notably, Baccouch et al. [464], Hashimoto et al. [465], and Ibarguren et al. [466] have reported the effects of fatty acid composition on various aspects of membrane behavior, including fluidity/viscosity/rigidity, thickness, permeability, phase transitions, fusion, lateral pressure, flip-flop dynamics, and structural integrity.
The optimal functionality of membrane-bound enzymes, ion channels, and receptors is intrinsically linked to membrane rigidity and permeability, influencing the diffusion of biomolecules within the lipid bilayer. Incorporating higher proportions of long saturated fatty acids enhances membrane rigidity, as these fatty acids are notably stable, having higher melting points than unsaturated fatty acids [467], and tend to form close clusters [468], particularly at physiological temperatures, resulting in heightened membrane rigidity. In contrast, polyunsaturated fatty acids contribute to greater conformational flexibility in membranes, which is a trait dependent on their chain length, degree of unsaturation, and the positioning of hydrogen atoms relative to the double bond. The presence of unsaturated fatty acids introduces curves/bends (also known as “kinks”) in the hydrocarbon chains, leading to the formation of less densely packed lipids and more fluidic membranes [469]. However, the influence of polyunsaturated fatty acids on membrane fluidity may vary, particularly in different bilayer states [470]. For instance, EPA and DHA have demonstrated negligible effects on fluidity in liquid-crystalline states [471]. In contrast, within different membrane models, polyunsaturated fatty acids with four or more double bonds, specifically AA, EPA, n6-DPA, and DHA, have been reported to decrease membrane thickness [464,472,473,474], increase the tilt angle [472], and elevate membrane fluidity [475,476]. The degree of rigidity contributed by double bonds within fatty acids is contingent on various factors, including their conformation (cis or trans), the degree of unsaturation, and their relative positioning concerning the carboxyl group. According to Roach et al. [477], the membrane properties of fatty acids associated with cis-unsaturated fatty acids were markedly different from those of saturated and trans-unsaturated fatty acids. Typically, cis-isomers exhibit greater polarity and possess relatively higher boiling points compared to trans-isomers, although not as high as those of saturated fatty acids. Notably, the position of the double bond exerts a more substantial impact on boiling points than the number of double bonds [478]. Cis-double bonds have been identified as expanding the spatial area occupied by the fatty acid, thereby increasing membrane fluidity [479,480] and permeability. Moreover, phospholipids containing long-chain n3-fatty acids have shaped more disordered and flexible membrane structures compared to LysoPLs containing n6-fatty acids, underscoring the significant role of n3-fatty acids in shaping membrane integrity.
Unsaturated fatty acids influence biomembrane rigidity not only through their intrinsic molecular structure but also by modifying the proportions of other membrane constituents that contribute to rigidity. For instance, Schumann et al. [481] and Stillwell [482] investigated the role of polyunsaturated fatty acids in modulating raft characteristics, including size, stability, and distribution. Notably, polyunsaturated fatty acids have a reduced affinity for cholesterol (CHOL) compared to their saturated counterparts. Consequently, an increased incorporation of unsaturated fatty acids within biomembranes can result in loosely packed lipid structures. This, in turn, leads to the displacement of raft-associated proteins and the removal of SM and CHOL from lipid rafts. These alterations result in shifts in membrane rigidity and permeability [483,484,485,486,487,488,489,490,491,492]. Therefore, the degree of unsaturation plays a crucial role in modulating the flip-flop rate and the asymmetry/distribution of membranes. Cheng et al. [493] and Armstrong et al. [494] have substantiated a positive correlation between the trans-membrane flip-flop rate and the proportion of unsaturated fatty acids. In assessing membrane rigidity, ratios such as phospholipid/CHOL, PC/PE, and PC/SM (the unsaturation index) have been routinely employed.
It is rational to posit that membranes characterized by inadequate lipid packing correspondingly exhibit elevated permeability. Indeed, a considerable body of research has established a link between polyunsaturated fatty acids and heightened membrane permeability, reflecting the rate at which molecules traverse biomembranes. It has been observed that the incorporation of long and very long polyunsaturated fatty acids, such as ALA, AA, EPA, and DHA, increases the permeability and elasticity of biomembranes [464,469,495,496,497,498,499,500]. This augmentation facilitates the translocation of ions and molecules across the membrane. Mondal et al. [469] attribute the elevated membrane elasticity to the disruption of the robust hydrogen-bond network surrounding the charged lipid head groups by the polyunsaturated fatty acids. The effects of polyunsaturated fatty acids, particularly EPA and DHA, on elasticity (and consequently permeability) can exhibit variability within the same cell [501], depending on the presence of other cellular constituents. Notably, the presence of CHOL can modulate membrane properties [495]. DHA, in particular, elevates permeability more significantly than its precursor, ALA [502], underscoring the pivotal role of the degree of unsaturation and its elevated incorporation levels in the functions of vital cells. DHA promotes heightened hydration within the head group and inter-chain regions, thereby increasing permeability. This is primarily attributed to the elevated number of double bonds. As elucidated by Mitchell and Litman [503], the presence of water within the hydrocarbon bilayer region exhibits a positive correlation with the high number of double bonds.
In addition to its effects on membrane rigidity and permeability, the incorporation of DHA profoundly impacts various aspects of biomembrane dynamics. DHA remarkably alters lipid packing, phase behavior, curvature, elasticity, interleaflet lipid flip-flop rates, lipid phase separations, membrane fusion, and vesicle formation [464,494,504,505,506,507,508]. According to Mitchell and Litman [509], the packing-free volume increases in the following order: 16:0-18:1PC has a lower relative abundance than 16:0-22:6PC, which, in turn, has a lower relative abundance than 22:6-C22:6PC. The potential effect of highly unsaturated fatty acids on phospholipids, which are characterized by loose packing, appears to be closely linked with the promotion of membrane elasticity, vesicle exfoliation (the formation of “blebs”), fusion, and flip-flop processes. In this regard, fatty acids actively participate in the processes of cell fusion and modulate cell phase behavior. During cell fusion, two distinct lipid bilayers merge, resulting in the formation of a continuous bilayer structure and the mixing of the internal contents of the lipid bilayers. Consequently, alterations in fusion processes have been associated with curvature stress in membranes [510,511]. The impact of the degree of unsaturation on membrane fusion has been previously demonstrated by Ahkong et al. [512], Meers et al. [513], Ehringer et al. [502], and, more recently, Li et al. [504]. The configuration of double bonds plays a critical role in determining the extent of biomembrane fusion. According to Creutz [514], AA and oleic acid are particularly effective fusogens, whereas saturated and trans-unsaturated fatty acids exhibit negligible fusogenic activity.
In terms of phase behavior, different fatty acid compositions contribute to various phase transitions, including gel-to-fluid, hexagonal, and liquid phases. Short-chain saturated fatty acids and mono- and polyunsaturated fatty acids result in lower viscosities, contributing to the formation of more fluid membranes compared to long-chain saturated fatty acids [466]. The impact of unsaturated fatty acids is particularly evident in thermal hysteresis, especially the transition between the fluid and hexagonal phases in PE, which is reportedly impeded by oleic acid, LA, and ALA [515,516]. Stearic acid and hydroxylated fatty acids induce a modest shift toward a higher melting temperature (the gel-to-fluid phase transition temperature) in bilayers containing C14:0/C14:0-PC [517]. On the other hand, PCs containing DHA exhibit higher melting points than those containing ALA and AA [518]. Despite DHA’s loose packing property [507], the presence of a saturated fatty acid at the sn-1 position in a PC molecule may affect its packing stability by altering both intra- and intermolecular van der Waals interactions.
4.2. Relative Functional Significance of Polyunsaturated Fatty Acids
The multifaceted role of fatty acids within various membrane lipids, particularly n3-fatty acids, has been documented over the past century. Diets rich in n3-fatty acids have been extensively associated with elevating the proportions of n3-fatty acids in biomembranes, thereby contributing to the maintenance of cardiovascular, vascular, and neural health [519,520]. Moreover, these dietary choices have shown promise in ameliorating conditions such as atherosclerosis, hypercholesterolemia, and cancer [487,521,522]. Fatty acids exhibit a multitude of physicochemical properties that serve diverse purposes by modifying the characteristics of bilayer lipids, thus influencing signal transduction. Notably, the length of a fatty acid exerts a marked influence on cellular signaling and metabolic processes. For instance, SLs containing short-chain fatty acids have been observed to augment susceptibility to apoptosis [523]. Membrane lipids enriched with monounsaturated fatty acids also play specific functional roles. Cao et al. [524] have reported that palmitoleate can function as a lipid-regulating hormone, often referred to as a ‘lipokine’, by enhancing sensitivity to glucose and inhibiting lipogenesis and hepatic inflammation. Furthermore, the well-documented antitumor and apoptotic properties of C18-monounsaturated fatty acids in carcinoma cells [525] underscore their potential utility in anticancer medications.
The acyl chain length of SLs, particularly Cers, significantly influences TAG accumulation and the hepatic uptake of fatty acids, which is attributed to the disruption of CD36/FAT expression [526]. This discovery underscores the role of CerS2 in catalyzing the generation of very long-chained Cers. In a cardiac context, long-saturated and polyunsaturated fatty acids have been shown to up-regulate voltage-dependent calcium release in cardiac myocytes [527], implicating their involvement in cardiac damage. Sassa and Kihara (2014) have presented a comprehensive review detailing the metabolism of very long-chain fatty acids and their contributions to the health and pathophysiology of various tissues, including the skin, meibum, retina, testis, and brain. The extensive body of available literature underscores the remarkable significance of polyunsaturated fatty acids, which have been the subject of substantial research due to their diverse bio-functional roles across various cell types.
The essentiality of LA and ALA in mammals transcends their role as diet-derived fatty acids; they are also fundamental precursors for the synthesis of long and very long polyunsaturated fatty acids. However, it is imperative to note that not all absorbed dietary LA and ALA are available for elongation and desaturation processes, as a fraction of these fatty acids are utilized for generating the energy source ATP during the β-oxidation process. Therefore, a prolonged deficiency in LA and/or ALA can lead to severe consequences, often manifesting as clinical symptoms [528,529,530,531,532]. LA, specifically, serves as a critical substrate for the biosynthesis of arachidonic acid (AA) and adrenic acid, both of which play crucial roles in early brain development [533,534,535]. Furthermore, LA is indispensable for the formation of n-hydroxyceramides, which covalently bond with epidermal proteins, thereby curtailing water loss and bolstering the skin’s barrier function [536]. In addition, it has been demonstrated that LA exhibits antibiotic-like properties, manifesting as an antibacterial effect that inhibits microbial adhesion to cells, a characteristic shared by numerous polyunsaturated fatty acids [537,538,539,540].
In general, polyunsaturated fatty acids exert substantial influence over the epidermis and its barrier properties. Notably, dietary supplementation of γ-linolenic acid has demonstrated anti-inflammatory properties [541] and has proven effective in enhancing skin characteristics in a dry skin model by reinforcing the skin’s barrier function and limiting dehydration [542]. Similar observations have been made with the supplementation of EPA and DHA [543], where an increase in the production of specific Cer families with anti-inflammatory properties was evident. It is worth noting that the effects of these fatty acids varied across distinct skin regions, including the epidermis, dermis, and hypodermis [543]. It is of particular interest that, among n6-fatty acids, γ-linolenic acid and DGLA have gained recognition for their anti-inflammatory attributes, similar to those of EPA and DHA (n3-fatty acids). Notably, γ-linolenic acid is found in inflammatory cells at relatively modest concentrations, and increasing its dietary intake does not necessarily lead to a proportional increase in its intracellular levels [544,545]. Given the efficient conversion of γ-linolenic acid to DGLA in mammals, it is conceivable that DGLA-derived lipid mediators play a role in mediating the anti-inflammatory effects associated with γ-linolenic [546].
The essentiality of ALA initially became apparent through observations of its ability to alleviate symptoms related to LA deficiency [547]. Its significance grew further when it was established that ALA serves as a precursor for EPA and DHA [548,549]. These C20 and C22 n3-polyunsaturated fatty acids are known to constitute a significant portion of the membrane lipids in critical tissues such as the brain [550], retina [551], and testis [552], reflecting their involvement in neurotransmission, visual excitation, and spermium maturation. The implications of n3-polyunsaturated fatty acids, especially DHA, on these tissues have been extensively documented in numerous studies [467,553,554,555,556,557,558,559,560,561,562,563,564,565,566]. These studies have proposed numerous biological functions for n3-polyunsaturated fatty acids, including the modulation of membrane proteins, gene expression, neurogenesis, enhancement of microcirculation, learning processes, and cellular protection. Notably, in neural tissue, the selectivity of PS declines under DHA deficiency [420]. The role of n3-fatty acids incorporated into PS in improving memory [567] and protecting against age-related lipid metabolic disorders, especially in the presence of DHA-enriched PC [567], is well acknowledged. For instance, DHA inhibits the production of amyloid-beta (Aβ) peptides associated with cognitive impairments, thereby mitigating amyloidogenesis, oxidative stress, and apoptosis [520]. The overall impact of polyunsaturated fatty acids on oxidative stress remains a subject of debate, as Shefer-Weinberg et al. [568] found that exposure to polyunsaturated fatty acids elevated oxidative stress biomarkers levels. In this context, it is plausible to hypothesize that the diverse polyunsaturated fatty acids may elicit distinct effects. Nonetheless, DHA has been reported to enhance the fluidity of the synaptic plasma membrane and induce the expression of other memory-related proteins [465]. Consequently, n3-fatty acids, particularly DHA, have gained significant scientific interest, leading to the development of nutraceuticals in the form of dietary supplements that incorporate these fatty acids.
In the preceding sections, the various roles of DHA in the physicochemical properties of membranes have been described. However, DHA also has crucial biological functions within membranes. DHA-enriched membranes have been suggested to influence membrane proteins by inducing curvature stress [569,570,571], affecting membrane thickness [473,570,572], and modulating fatty acid packing free volume [565]. These alterations in membrane properties can lead to modifications in the activity of most cellular proteins, affecting signal propagation. For instance, unsaturated fatty acids have been reported to interact with various proteins, including rhodopsin, ion channels (L-type Ca2+ and Na+), protein kinase C (PKC), apoptosis-associated proteins, PPAR-γ, nuclear receptor Nur77, G-protein coupled receptor 40, mitogen-activated protein kinase, toll-like receptors, and nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB) [573,574,575,576,577,578,579,580,581,582,583,584,585]. However, the major relationships between DHA and cellular protein activities remain ambiguous due to the vast diversity of proteins, the complexity of protein interactions, and the limited number of studies. Despite being highly unsaturated (with six double bonds), DHA exhibits antioxidant properties in the liver [586], brain [587,588], and skeletal muscles [465]. This property is of particular significance for fertility, as Roqueta-Rivera et al. [589] observed that DHA supplementation effectively restored impaired spermatogenesis in male mice.
Both EPA and DHA have demonstrated the ability to counteract pro-inflammatory cytokines by down-regulating the NF-κB signaling pathway [590,591,592], a transcriptional pathway that regulates both innate and adaptive immune responses. In contrast, AA levels have been found to correlate positively with lipid peroxidation [593] and activation of the NF-κB signaling pathway [594], thereby promoting pro-inflammatory stimuli. AA can also up-regulate SMase activity [595], leading to increased levels of Cers, molecules that trigger apoptotic signals, which are derived from SM hydrolysis. Thus, AA is a biologically essential fatty acid, contributing to a wide array of functions either directly or through its bioactive metabolites. Hashidate-Yoshida et al. [596] demonstrated that AA facilitates the transportation of triglycerides to the lumen of the ER in hepatocytes and enterocytes.
The ratio between fatty acids within cellular membranes serves as a reflection of universal cellular signaling and inflammatory responses. Notably, EPA and DHA exhibit distinct signaling profiles compared to AA. Consequently, the ratio of EPA and DHA to AA can serve as an indirect indicator for assessing the inflammatory response and lipid peroxidation. It is worth emphasizing that these fatty acids serve as precursors for numerous bioactive mediators, contributing to a wide array of physiological functions. However, it is also important to recognize that many of the reported findings are likely attributed to the direct alterations of membrane physicochemical properties and membrane-associated proteins [481], along with the unidentified bioactive metabolites they generate. Polyunsaturated fatty acids can undergo chemical reactions with various molecules and cellular components, resulting in the formation of novel compounds with biological activity. Heshmati [597] has described interactions between n3-fatty acids and specific transcription factors in genes. Furthermore, an intriguing observation is the interaction of nitric oxide (NO) with polyunsaturated fatty acids, leading to the formation of nitroalkene derivatives. These plasma-identifiable derivatives have been demonstrated to promote vascular relaxation, inhibit neutrophil cell degranulation and superoxide production, and hinder platelet activation [598,599,600]. Nitroalkene derivatives possess inherent PPAR ligand activity and are known to degrade into NO in the bloodstream. These observations underscore the capacity of polyunsaturated fatty acids to engage in reactions with other non-lipidous cellular constituents, resulting in the formation of novel compounds with specific biological activities.
4.3. Bioactive Lipid Mediators Derived from Fatty Acids
Numerous classes of lipids, including LysoP, SLs, PA, DAG, inositol phosphate, N-acylethanolamine, fatty acids, and oxylipins, are renowned for their bioactive intracellular and extracellular signaling properties, acting as messengers/mediators. On the other hand, certain functions of polyunsaturated fatty acids necessitate their conversion into lipid mediators. These mediators serve as signaling molecules that modulate various biological processes, including the inflammatory response, gene transcription, and signal transduction pathways. For instance, the tissue hormone-like lipids referred to as “eicosanoids”, which were initially identified in the prostate [601], possess the ability to regulate the function of various transcription factors, thus inducing alterations in gene expression. To comprehend the mechanisms underlying the generation of lipid mediators derived from fatty acids, this section elaborates on the cleavage mechanism of membrane fatty acids, the oxygenation mechanisms of deacylated fatty acids, and the biological functions of lipid mediators derived from fatty acids.
4.3.1. Enzyme-Mediated Cleavage of Fatty Acids from Membranes
Polyunsaturated fatty acids are abundant in biomembranes but can be enzymatically cleaved from sn-positions and N-acyl linkages of membrane lipids by lipase-type enzymes. These enzymes encompass PLA2, phospholipase B (PLB, an enzyme with both PLA1 and PLA2 activities), diacylglycerol lipase [602,603], CDase [604], glucosylceramide deacylase [605,606], and sphingomyelin deacylase [605,607]. Other phospholipase enzymes, such as PLA1, PLC, and PLD, play a lesser role in the generation of polyunsaturated fatty acid-derived mediators, as they cleave the highly saturated chains at the sn-1 position [608], the phosphate group at the sn-3 position [609], and the head group from the phosphorus group [610], respectively.
Among these enzymes, PLA2 has received considerable attention due to the biological importance of its substrates. Over recent decades, six isoforms of PLA2, which hydrolyze the ester bond at the sn-2 position, have been identified [611]. Each of these isoforms exhibits selectivity for specific fatty acids on phospholipids. For example, cytosolic PLA2α (cPLA2α) acts on phospholipids rich in AA [612], calcium-independent PLA2β (iPLA2β) acts on phospholipids rich in DHA [613,614], and secretory PLA2 (sPLA2) acts on phospholipids containing various fatty acids, including AA, EPA, and DHA [615,616]. On the other hand, PLB possesses both hydrolase activity, cleaving ester bonds on the sn-1 and sn-2 positions of phospholipids, and acyltransferase activity, acylating fatty acid to form LysoP, and, as a result, may contribute less to the production of oxylipins compared to PLA2. Following the removal of fatty acids from complex membrane lipids, various events, including reacylation and/or oxidation, may occur.
4.3.2. Fatty Acid Oxygenation
CHOL and liberated polyunsaturated fatty acids can undergo oxidation through enzymatic processes and non-enzymatic agents, such as reactive oxygen species (ROS). This oxidative transformation leads to the production of oxysterols and oxylipins, respectively. Notably, these compounds may also originate from dietary sources. It is of particular significance that polyunsaturated fatty acids frequently engage in metabolic competition with each other, a phenomenon specifically mediated by enzymes such as PLA2 and oxidative enzymes. The extent of competition among different fatty acids depends on their respective concentrations within the cell and their relative affinities for oxidative enzymes and reactive molecules [617,618]. These oxidized lipid metabolites serve as pivotal mediators in cell signaling. For instance, oxysterols have the capacity to interact with nuclear receptors and, as a consequence, modulate gene expression [619,620]. This section primarily focuses on the enzymatic pathways involved in generating these bioactive lipid mediators.
Numerous bioactive oxylipins have been identified as products of enzymatic pathways, including those facilitated by cyclooxygenase (COX) and subsequent synthases, lipoxygenase (LOX), and cytochrome P450 (CYP) mixed-function oxidase enzymes [621]. These oxylipins are further categorized based on the chain length of their respective substrates (see Figure 7), resulting in octadecanoids (derived from C18 fatty acids), eicosanoids (derived from C20 fatty acids), docosanoids (derived from C22 fatty acids), and elovanoids (derived from C32 or C34 fatty acids).
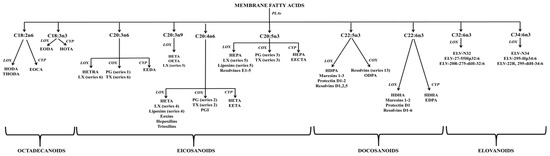
Figure 7.
Diagram illustrating the various lipid mediators (including octadecanoids, eicosanoids, docosanoids, and elovanoids) synthesized from fatty acids such as LA, ALA, DGLA, Mead acid, AA, EPA, DPA-n3, DHA, C32:6n3, and C34:6n3. Abbreviations: COX, cyclooxygenase; EDPA, epoxy-docosapentaenoic acid; EECTA, epoxy-ecosatetraenoic acid; EEDA, epoxy-eicosadienoic acid; EETA, epoxy-eicosatrienoic acid; ELV, elovanoids; EOCA, epoxy-octadecenoic acid; EODA, epoxy-octadecadienoic acid; CYP, cytochrome P450; HDHA, hydroxy-docosahexaenoic acid; HDPA, hydroxy-docosapentaenoic acid; HEPA, hydroxy-eicosapentaenoic acid; HETA, eicosatetraenoic acid; HETRA, hydroxy-eicosatrienoic acid; HODA, hydroxy-octadecadienoic acid; HOTA, hydroxy-octadecatrienoic acid; LOX, lipoxygenase; LX, leukotrienes; ODPA, oxodocosapentaenoic acid; OETA, oxoeicosatetraenoic acid; PG, prostaglandin; PGI, prostacyclins; PLAs, phospholipases; THODA, trihydroxy-octadecenoic acid; TX, thromboxane.
In mammals, COXs, also known as housekeeping enzymes, comprise three isoforms as follows: COX-1, COX-2, and COX-3 isoforms [622], with COX-3 being considered a variant of COX-1 [623]. These enzymes are heme-containing and possess the dual capacity to function as both oxygenases and peroxidases. Notably, these enzymes are constitutively expressed and are subject to modulation by inflammatory signals. Their main role involves catalyzing the oxygenation of various unsaturated fatty acids, culminating in the generation of bioactive end-products collectively referred to as prostanoids. These prostanoids encompass the prostaglandin series (PGD, PGE (dinoprostone), PGF (carboprost), and PGI (prostacy-clins)), thromboxanes, hydroxy fatty acids, resolvins (series 13), and oxo-fatty acids [624,625].
LOXs, which comprise six genes within the human genome, represent a class of non-heme iron-containing dioxygenases. These enzymes possess the capability to oxygenate a broad range of unsaturated fatty acids. It is noteworthy that LOX enzymes typically exist in an inactive form at their base state, necessitating activation facilitated by hydroperoxides. Subsequently, they act on a diverse array of substrates and engage in various modes of action, including dioxygenase activity, functioning as catalysts in processes characterized by the involvement of free radicals [626,627]. These catalytic actions lead to the formation of bioactive end-products recognized as hydroperoxyl fatty acids and their metabolites, including leukotrienes, lipoxins, resolvins, protectins, maresins, and elovanoids [628,629].
On the other hand, CYPs are enzymes encoded by an extensive set of up to 57 genes within the human genome, representing a class of monooxygenases widely distributed in mammals. These enzymes exhibit elevated activity levels in numerous tissues, including but not limited to the liver, brain, kidneys, and lungs [629,630]. CYPs are renowned for their involvement in various modes of action, including hydroxylation, heteroatom oxidation, allylic oxidation reactions, group migration, and various other enzymatic reactions [631,632,633]. They display the capability to act on a diverse range of unsaturated fatty acids and sterols [629], thereby generating a wide array of lipid mediators. In particular, these lipid mediators consist of hydroxyl and epoxy fatty acids, which play critical roles in the induction of various signaling pathways.
4.3.3. Functions of Bioactive Lipid Mediators
Bioactive lipid mediators go beyond being inert components of cellular membranes. Instead, they serve as dynamic signaling agents and are capable of modulating a wide range of signaling pathways, gene regulation, and immune responses. The unique characteristics and functions of these mediators have led to extensive research efforts aimed at harnessing their therapeutic potential for developing innovative treatment/preventive approaches. Therefore, comprehending the functions of bioactive lipid mediators holds great importance in the domains of biological and medical research.
Octadecanoids
Essential fatty acids and their extended metabolites have well-documented diverse biological effects and implications in various disease mechanisms. The effects of lipid mediators can vary depending on the type of cells and metabolic factors, leading to both beneficial and detrimental metabolic outcomes (see Figure 8).
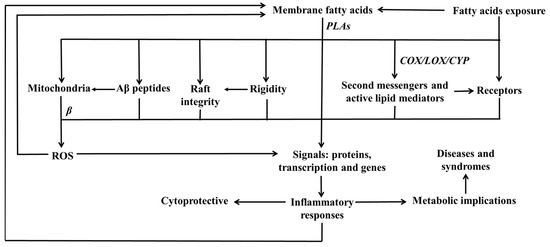
Figure 8.
Illustration of the influence of bilayer-cleaved fatty acids on cellular signaling pathways and inflammation responses. Abbreviations: β, β-oxidation; COX, cyclooxygenase; CYP, cytochrome P450; LOX, lipoxygenase; PLAs, phospholipases; ROS, reactive oxygen species.
LA and ALA play an essential role in the generation of lipid mediators. As essential fatty acids, dietary levels of LA and ALA contribute to their proportions within cellular membranes, potentially playing a crucial role in modulating the extent/degree/severity of inflammation development. Epoxy-octadecadienoic acid and hydroxy-octadecatrienoic acid are lipid mediators derived from ALA through the enzymatic actions of LOX and CYP, respectively [624]. However, further research is needed to fully comprehend the bioactive functions of octadecanoids derived from ALA. Notably, Kumar et al. [634] have suggested that these mediators primarily exert anti-inflammatory effects. On the other hand, oxidized LA metabolites, including hydroxy, trihydroxy, and epoxy fatty acids, are produced under the effects of LOXs and CYPs [546,624]. These metabolites have been implicated in various biological pathways, including brain dysfunction [635], the inhibition of platelet adhesion in endothelial cells [636], the induction of inflammation signals [637,638,639], the maintenance of skin barrier integrity [536], the inhibition of pain thresholds [640,641], and the promotion of metabolic syndromes and cancer [638,642]. Consequently, these LA-derived mediators may greatly contribute to the inflammatory processes and the progression of diseases.
Both LA and ALA serve as essential fatty acids and precursors for extended polyunsaturated fatty acids. Thus, their dietary concentrations can alter the levels of long and very long chain polyunsaturated fatty acids within cellular biomembranes. However, the extent of this influence may vary depending on the specific substrate and metabolic pathways. Notably, a high dietary intake of LA tends to not significantly elevate the proportion of AA or the associated inflammatory cascades in humans [643]. In contrast, a high dietary intake of ALA has been shown to increase EPA and DHA concentrations [644]. Nevertheless, it is essential to recognize that the de novo pathways for elongating essential fatty acids exhibit variations among species. For instance, the conversion rate of ALA to its extended polyunsaturated fatty acids is lower in humans [645] than in marine species. These findings underscore the potential variability in the biological functions of ALA, with specific implications in distinct species to fulfill particular physiological functions.
Eicosanoids
Eicosanoids are bioactive lipid mediators primarily derived from unesterified fatty acids and are characterized by their autocrine/paracrine hormone activities. They mediate local signals and reactions, including processes related to homeostasis, inflammation, and anti-inflammation. Eicosanoids comprise various structures, such as PGs, thromboxanes, leukotrienes, lipoxins, and resolvins. Despite the fact that most mammalian cells are capable of synthesizing eicosanoids, the specific pathways and responses can vary by cell type [646]. DGLA, rapidly extended from γ-linolenic acid, serves as a substrate for enzymes such as COX, which yields series 1 prostaglandins and thromboxanes, 15-LOX, which yields 5-hydroxyeicosatrienoic acid, and CYP, which yields epoxy-eicosadienoic acid. Eicosanoids derived from DGLA are generally considered to be anti-inflammatory [647,648].
On the other hand, beyond the role of AA as a polyene fatty acid, it is unquestionably crucial in biomembranes as it is the primary target for most membrane-modifying effects. The activation of the PLA2 enzyme, which is responsible for cleaving AA from membrane phospholipids, often leads to membrane injury. According to Samuelsson [649], this enzyme rapidly (within seconds to minutes) responds to acute stimuli, releasing AA from membrane lipids. Liberated AA can be utilized as a precursor for the production of eicosanoids under the effects of COX (generating series 2 prostaglandins, prostacyclins, and thromboxanes), LOX (generating leukotrienes, lipoxins, eoxins, hepoxilins, and trioxilins), and CYP (generating hydroxyeicosatetraenoic acid and epoxyeicosatrienoic acid) [546,624]. AA-derived eicosanoids, often referred to as arachidonate or eicosanoid cascades, are involved in multiple systems, including vascular, inflammatory, renal, and neuronal signaling, as well as angiogenesis [650]. For instance, AA-derived eicosanoids have been shown to increase the permeability of the blood–brain barrier in humans [651], revealing the potential for drug modulation of this barrier.
Eicosanoids derived from Mead acid have displayed anti-inflammatory properties. For instance, oxygenated products of Mead acid via 5-lipoxygenase are produced during inflammation, providing potent activities [652]. However, the exact roles of this fatty acid are not yet clearly defined [652,653,654,655], necessitating further research. On the other hand, LOXs oxidize EPA to produce resolvins [656], which are renowned for their anti-inflammatory properties. EPA can also undergo oxygenation via COXs (yielding hydroxy-eicosapentaenoic acid and epoxy-eicosatetraenoic acid) and CYPs (yielding series 3 prostaglandins and leukotrienes) [624,657]. Overall, EPA-derived eicosanoids exhibit anti-inflammatory stimuli, such as the inhibition of platelet aggregation [658].
Eicosanoids play a remarkable role in the regulation of inflammatory responses by modulating pro-inflammatory cytokines, chemokines, and other signaling molecules. They have the potential to influence the recruitment, activation, and function of immune cells. However, it is essential to recognize that eicosanoids can exhibit both pro-inflammatory and anti-inflammatory effects, with the ultimate effect determined by various factors, including mediator concentrations, timing of production, and the sensitivity of targeted cells/tissues [460,659]. Commonly, eicosanoids derived from different fatty acids, such as AA, Mead acid, and EPA, exhibit strikingly distinct biological effects, despite their closely resembling molecular structures.
Eicosanoids derived from n3-fatty acids are well-recognized for their anti-inflammatory properties, while those originating from n6-fatty acids are generally considered to be pro-inflammatory [660,661]. However, it is important to note that not all n6-fatty acids exert pro-inflammatory effects. Some prostanoids (PGs and thromboxanes), n6-fatty acid-derived lipoxins, as well as mediators derived from γ-linolenic and DGLA, along with adrenic acid, have been found to express anti-inflammatory properties and cytoprotective actions [662,663,664,665,666,667,668,669]. Imbalances in the production of eicosanoids have been implicated in numerous pathological processes, including inflammation, autoimmunity, allergy, cancer, atherosclerosis, and metabolic and degenerative diseases [650], by disrupting the normal lipid signaling pathways. In light of this, strategies that involve the suppression of COX, LOX, and CYP enzymes, which are responsible for the synthesis of active lipid mediators, may hold therapeutic potential for the management of disease-related inflammation and oxidative stress.
Docosanoids
DHA, likely the reason for the biological necessity of ALA, is a very long polyunsaturated fatty acid that accumulates abundantly in crucial tissues such as the brain, retina, and testis. Though EPA is known to produce pre-resolving mediators (resolvins), it is DHA that serves as the major precursor for these compounds [656,670]. Specialized pro-resolving mediators (SPMs), known as docosanoids, are primarily derived from the LOX oxidation of DHA and DPA-n3 [670,671,672,673]. However, COX activity on DPA-n3 can also generate SPMs [672], and CYP activity on DHA yields hydroxy-docosapentaenoic acid and epoxy-docosapentaenoic acid [624]. The pre-resolving family comprises various structures, including resolvins, docosatrienes, maresins, and protectins, all of which exhibit anti-inflammatory and pro-resolving properties, countering the effects of pro-inflammatory cascades [659,666,667,671,672,673,674,675,676,677,678,679,680,681,682]. These docosanoids, which are derived from DPA-n3 and DHA, play pivotal roles in the regulation of leukocyte trafficking, suppression of cytokine expression, inhibition of brain ischemia-reperfusion injury, maintenance of cellular homeostasis, mitigation of potential DNA oxidation, normalization of brain-derived neurotrophic factor levels, and promotion of the clearance of apoptotic cells and cellular debris by phagocytes. Thus, these mediators represent a promising therapeutic approach for resolving cellular inflammation and associated diseases.
Furthermore, EPA and DHA are known to limit pro-inflammatory cytokines and reduce inflammation, potentially by increasing peroxisome proliferator-activated receptor alpha (PPAR-α) mRNA and protein activities [683]. Remarkably, alternative lipid mediators with resembling impacts to resolvins have been identified. According to Dalli et al. [684], DPA-n3, an intermediate fatty acid during DHA synthesis, is transformed into novel immunoresolvents similar to resolvins in mice and human leukocytes during inflammation. However, it is important to acknowledge that the resolution of inflammation mediated by docosanoids is characterized by its complexity in restoring cellular homeostasis [656].
Elovanoids
In response to unmitigated oxidative stress, elovanoids exhibit a remarkable ability to enhance the intracellular synthesis of pro-survival signals, owing to their distinctive molecular structures. This class of bioactive lipids, initially discovered by Bazan’s research group in the retinal pigment epithelium in 2017 [685], is derived from mono-hydroxyl-very long polyunsaturated fatty acids formed through the enzymatic activity of ELVOL4 and LOX. It is important to emphasize that very long polyunsaturated fatty acids are prominent constituents of critical tissues such as the brain, testis, and spermatozoa [686]. This observation suggests the potential formation of elovanoids in these tissues, where they might serve as mediators of specific signals. However, while the retina has been a focal point of research on elovanoids, studies examining neural signaling are comparatively limited.
Elovanoids play an indispensable role in the functions of the retina and neural signaling [290,685,687,688,689,690,691]. The protective effects of elovanoids in these tissues are most likely attributed to their role in mitigating the effects of oxidative stress. In events where oxidative stress remains unresolved, elovanoids serve as critical survival signals [685]. These authors have reported that dihydroxylated derivatives of C32:6n-3 and C34:6n-3 effectively protect retinal pigment epithelial cells from apoptosis induced by hydrogen peroxide. These derivatives have been shown to up-regulate the expression of pro-survival proteins, including Bcl-2 and Bcl-xL, while concurrently down-regulating the expression of pro-apoptotic proteins, such as Bax, Bim, and Bid. These findings underscore the ability of elovanoids to mitigate the cytotoxic effects of ROS on photoreceptor cells and contribute to their survival.
5. Conclusions and Future Perspectives
This review intends to provide an in-depth overview of the lipids of eukaryotic cell membrane lipids, with a particular emphasis on fatty acids. It introduces the extensive array of lipids present in biomembranes and delves into their composition within healthy organisms, thereby illustrating the intricate nature of lipid metabolism and its fundamental role within cells. This perspective underscores the remarkable adaptability and flexibility inherent in the fatty acid profiles of biomembranes, enabling organisms to rapidly respond to various stimuli, including alterations in environmental temperature, dietary factors, inflammatory processes, or diseases. Thus, the absence of a universally defined “physiologically normal fatty acid composition” underscores the natural variability in fatty acid composition. This natural phenomenon is, likewise, a continuous process of adaptation. This review further provides an in-depth exploration of fatty acid biosynthesis and post-synthetic modifications, such as elongation and desaturation. In addition, it highlights the preferences of fatty acids for incorporation into diverse complex membrane lipids and their roles in biological systems, encompassing both physicochemical properties and the regulation of biological signaling. This understanding holds significant implications across various disciplines, including lipid-based drug delivery, cell membrane engineering, and the advancement of lipid-based biomaterials. Nevertheless, further research remains essential to unveil the intricate mechanisms and regulatory pathways governing eukaryotic lipid metabolism and fatty acid composition. This includes investigations into the mechanisms underpinning cellular membrane adaptability, with the potential to shed light on the molecular foundations of cellular processes, diverse diseases, and the development of therapeutic strategies for lipid-related disorders.
Evidently, the pivotal role of fatty acids in biomembranes is ascending and is poised to exert a substantial influence across various disciplines, notably within the realms of nutrition and medicine. This review serves to illuminate the multifaceted roles and contributions of distinct membrane lipids, along with their associated fatty acids, with a specific focus on matters pertaining to health and the intricate aspects of inflammatory responses. Enhancing our comprehensive comprehension of the diverse repertoire of membrane lipids stands to be invaluable for assessing the overall health of organisms. The trajectory of the field nutrition is set to emphasize progressively specific fatty acids that are indispensable for organism health. In this context, the n3 and n6-fatty acids are assuming paramount significance due to their critical roles as precursors for bioactive lipids that play a pivotal role in the modulation of inflammatory processes. They also contribute indispensably to the development and sustenance of vital organ functions, exemplified by the brain, heart, lungs, liver, and kidneys. Elevated levels of these fatty acids have been consistently correlated with to the regulation of chronic maladies, encompassing diabetes, cardiovascular disorders, and certain forms of cancer. However, it is crucial to recognize that the optimization of fatty acid biosynthesis, the preservation of their stability, and a comprehensive understanding of their various roles in biological systems continue to remain areas ripe for exploration. Thus, the unwavering dedication to research and development in this domain holds the promise of unveiling the unlocking of novel approaches to incorporate these essential nutrients into the diets of organisms, thereby fostering enduring health and well-being.
Author Contributions
O.A. conceptualized the idea, wrote the manuscript and prepared the figures; A.S. reviewed and edited the manuscript. Both authors actively participated in discussion, manuscript revision, and final version contributions. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the following projects: EFOP-3.6.3-VEKOP-16-2017-00005 and GINOP-2.3.2-15-2016-00046. This work was further funded by the Hungarian Academy of Sciences (HUN-REN-MATE, Mycotoxins in the Food Chain research group) and by the Hungarian National Laboratory project RRF-2.3.1-21-2022-00007.
Acknowledgments
The authors would like to acknowledge András Dinnyés for his insightful suggestions, which improved the present review.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Harayama, T.; Riezman, H. Understanding the diversity of membrane lipid composition. Nat. Rev. Mol. Cell Biol. 2018, 19, 281–296. [Google Scholar] [PubMed]
- Watson, H. Biological membranes. Essays Biochem. 2015, 59, 43–69. [Google Scholar] [PubMed]
- Janmey, P.A.; Kinnunen, P.K.J. Biophysical properties of lipids and dynamic membranes. Trends Cell Biol. 2006, 16, 538–546. [Google Scholar] [PubMed]
- Dynarowicz-Łątka, P.; Dhanabalan, A.; Oliveira, O.N. Modern physicochemical research on Langmuir monolayers. Adv. Colloid Interface Sci. 2001, 91, 221–293. [Google Scholar] [PubMed]
- Singer, S.J.; Nicolson, G.L. The fluid mosaic model of the structure of cell membranes. Science 1972, 175, 720–731. [Google Scholar]
- Simons, K.; Ikonen, E. Functional rafts in cell membranes. Nature 1997, 387, 569–572. [Google Scholar]
- Ballweg, S.; Sezgin, E.; Doktorova, M.; Covino, R.; Reinhard, J.; Wunnicke, D.; Hänelt, I.; Levental, I.; Hummer, G.; Ernst, R. Regulation of lipid saturation without sensing membrane fluidity. Nat. Commun. 2020, 11, 756. [Google Scholar]
- Nicolson, G.L. The fluid—Mosaic model of membrane structure: Still relevant to understanding the structure, function and dynamics of biological membranes after more than 40years. Biochim. Biophys. Acta Biomembr. 2014, 1838, 1451–1466. [Google Scholar]
- Edidin, M. Lipids on the frontier: A century of cell-membrane bilayers. Nat. Rev. Mol. Cell Biol. 2003, 4, 414–418. [Google Scholar]
- Wang, Y.; Gao, J.; Guo, X.; Tong, T.; Shi, X.; Li, L.; Qi, M.; Wang, Y.; Cai, M.; Jiang, J.; et al. Regulation of EGFR nanocluster formation by ionic protein-lipid interaction. Cell Res. 2014, 24, 959–976. [Google Scholar]
- Luckey, M. Membrane Structural Biology, 1st ed.; Cambridge University Press: Cambridge, UK, 2008; ISBN 9780521856553. [Google Scholar]
- Hanczyc, M.M.; Fujikawa, S.M.; Szostak, J.W. Experimental models of primitive cellular compartments: Encapsulation, growth, and division. Science 2003, 302, 618–622. [Google Scholar]
- Chevreul, M.E. Sur plusieurs corps gras et particulièrement sur leurs combinaisons avec les alcalis. [On several fatty substances and particularly on their combinations with alkalis]. Ann. Chim. 1813, 88, 225–261. [Google Scholar]
- Lombard, J. Once upon a time the cell membranes: 175 years of cell boundary research. Biol. Direct 2014, 9, 32. [Google Scholar]
- Gorter, E.; Grendel, F. On bimolecular layers of lipoids on the chromocytes of the blood. J. Exp. Med. 1925, 41, 439–443. [Google Scholar] [CrossRef]
- Glatz, J.F.C.; Luiken, J.J.F.P.; Bonen, A. Membrane fatty acid transporters as regulators of lipid metabolism: Implications for metabolic disease. Physiol. Rev. 2010, 90, 367–417. [Google Scholar] [PubMed]
- Anheuser, S.; Breiden, B.; Sandhoff, K. Membrane lipids and their degradation compounds control GM2 catabolism at intralysosomal luminal vesicles. J. Lipid Res. 2019, 60, 1099–1111. [Google Scholar] [PubMed]
- Settembre, C.; Ballabio, A. Lysosome: Regulator of lipid degradation pathways. Trends Cell Biol. 2014, 24, 743–750. [Google Scholar] [PubMed]
- Kolter, T.; Sandhoff, K. Lysosomal degradation of membrane lipids. FEBS Lett. 2010, 584, 1700–1712. [Google Scholar] [PubMed]
- Van Meer, G. Cellular lipidomics. EMBO J. 2005, 24, 3159–3165. [Google Scholar]
- Sackmann, E. Biological membranes architecture and function. In Structure and Dynamics of Membranes: I. From Cells to Vesicles/II. Generic and Specific Interactions; Lipowsky, R., Sackman, T., Eds.; Elsevier: Amsterdam, The Netherlands, 1995; pp. 1–63. [Google Scholar]
- Liebisch, G.; Fahy, E.; Aoki, J.; Dennis, E.A.; Durand, T.; Ejsing, C.S.; Fedorova, M.; Feussner, I.; Griffiths, W.J.; Köfeler, H.; et al. Update on LIPID MAPS classification, nomenclature, and shorthand notation for MS-derived lipid structures. J. Lipid Res. 2020, 61, 1539–1555. [Google Scholar]
- Fahy, E.; Cotter, D.; Sud, M.; Subramaniam, S. Lipid classification, structures and tools. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2011, 1811, 637–647. [Google Scholar] [CrossRef] [PubMed]
- Bogdanov, M.; Dowhan, W. Functional roles of lipids in biological membranes. In Biochemistry of Lipids, Lipoproteins and Membranes; Ridgway, N.D., McLeod, R.S., Eds.; Elsevier: Amsterdam, The Netherlands, 2021; pp. 1–51. [Google Scholar]
- Drin, G. Topological regulation of lipid balance in cells. Annu. Rev. Biochem. 2014, 83, 51–77. [Google Scholar] [CrossRef] [PubMed]
- Vauquelin, N.L. Analyse de la matière cérébrale de l’homme et de quelques animaux [Translation: Analysis of the Brain Matter of Man and Some Animals]. Ph.D. Thesis, Faculty of Medicine of Paris, Paris, France, 12 August 1811. [Google Scholar]
- Van Meer, G.; Voelker, D.R.; Feigenson, G.W. Membrane lipids: Where they are and how they behave. Nat. Rev. Mol. Cell Biol. 2008, 9, 112–124. [Google Scholar] [PubMed]
- Buckland, A.G.; Wilton, D.C. Anionic phospholipids, interfacial binding and the regulation of cell functions. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2000, 1483, 199–216. [Google Scholar]
- Moolenaar, W.H.; Kruijer, W.; Tilly, B.C.; Verlaan, I.; Bierman, A.J.; de Laat, S.W. Growth factor-like action of phosphatidic acid. Nature 1986, 323, 171–173. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, E.P.; Lehninger, A.L. Oxidation of fatty acids and tricarboxylic acid cycle intermediates by isolated rat liver mitochondria. J. Biol. Chem. 1949, 179, 957–972. [Google Scholar] [CrossRef]
- Tanguy, E.; Wang, Q.; Moine, H.; Vitale, N. Phosphatidic acid: From pleiotropic functions to neuronal pathology. Front. Cell. Neurosci. 2019, 13, 2. [Google Scholar]
- Lee, J.; Ridgway, N.D. Substrate channeling in the glycerol-3-phosphate pathway regulates the synthesis, storage and secretion of glycerolipids. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2020, 1865, 158438. [Google Scholar]
- Tanguy, E.; Costé de Bagneaux, P.; Kassas, N.; Ammar, M.-R.; Wang, Q.; Haeberlé, A.-M.; Raherindratsara, J.; Fouillen, L.; Renard, P.-Y.; Montero-Hadjadje, M.; et al. Mono- and poly-unsaturated phosphatidic acid regulate distinct steps of regulated exocytosis in neuroendocrine cells. Cell Rep. 2020, 32, 108026. [Google Scholar] [CrossRef]
- Zegarlińska, J.; Piaścik, M.; Sikorski, A.F.; Czogalla, A. Phosphatidic acid—A simple phospholipid with multiple faces. Acta Biochim. Pol. 2018, 65, 163–171. [Google Scholar] [CrossRef]
- Ammar, M.-R.; Kassas, N.; Bader, M.-F.; Vitale, N. Phosphatidic acid in neuronal development: A node for membrane and cytoskeleton rearrangements. Biochimie 2014, 107, 51–57. [Google Scholar] [CrossRef] [PubMed]
- Stace, C.; Ktistakis, N. Phosphatidic acid- and phosphatidylserine-binding proteins. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2006, 1761, 913–926. [Google Scholar] [CrossRef]
- Benson, A.A.; Maruo, B. Plant phospholipids I. Identification of the phosphatidyl glycerols. Biochim. Biophys. Acta 1958, 27, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Ridgway, N.D. Phospholipid synthesis in mammalian cells. In Biochemistry of Lipids, Lipoproteins and Membranes; Ridgway, N.D., McLeod, R.S., Eds.; Elsevier: Amsterdam, The Netherlands, 2021; pp. 227–258. [Google Scholar]
- Agassandian, M.; Mallampalli, R.K. Surfactant phospholipid metabolism. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2013, 1831, 612–625. [Google Scholar] [CrossRef]
- Hallman, M.; Enhorning, G.; Possmayer, F. Composition and surface activity of normal and phosphatidylglycerol-deficient lung surfactant. Pediatr. Res. 1985, 19, 286–292. [Google Scholar] [CrossRef]
- Furse, S. Is phosphatidylglycerol essential for terrestrial life? J. Chem. Biol. 2017, 10, 1–9. [Google Scholar] [CrossRef]
- Kanoh, H.; Kondoh, H.; Ono, T. Diacylglycerol kinase from pig brain. Purification and phospholipid dependencies. J. Biol. Chem. 1983, 258, 1767–1774. [Google Scholar] [CrossRef]
- Laurinavičius, S.; Käkelä, R.; Bamford, D.H.; Somerharju, P. The origin of phospholipids of the enveloped bacteriophage phi6. Virology 2004, 326, 182–190. [Google Scholar] [CrossRef]
- Bamford, D.H.; Romantschuk, M.; Somerharju, P.J. Membrane fusion in prokaryotes: Bacteriophage phi 6 membrane fuses with the Pseudomonas syringae outer membrane. EMBO J. 1987, 6, 1467–1473. [Google Scholar] [CrossRef] [PubMed]
- Sands, J.A.; Lowlicht, R.A. Temporal origin of viral phospholipids of the enveloped bacteriophage ϕ6. Can. J. Microbiol. 1976, 22, 154–158. [Google Scholar] [CrossRef]
- Numata, M.; Kandasamy, P.; Nagashima, Y.; Posey, J.; Hartshorn, K.; Woodland, D.; Voelker, D.R. Phosphatidylglycerol suppresses influenza A virus infection. Am. J. Respir. Cell Mol. Biol. 2012, 46, 479–487. [Google Scholar] [CrossRef] [PubMed]
- Kandasamy, P.; Zarini, S.; Chan, E.D.; Leslie, C.C.; Murphy, R.C.; Voelker, D.R. Pulmonary surfactant phosphatidylglycerol inhibits Mycoplasma pneumoniae-stimulated eicosanoid production from Human and mouse macrophages. J. Biol. Chem. 2011, 286, 7841–7853. [Google Scholar] [CrossRef] [PubMed]
- Kuronuma, K.; Mitsuzawa, H.; Takeda, K.; Nishitani, C.; Chan, E.D.; Kuroki, Y.; Nakamura, M.; Voelker, D.R. Anionic pulmonary surfactant phospholipids inhibit inflammatory responses from alveolar macrophages and U937 cells by binding the lipopolysaccharide-interacting proteins CD14 and MD-2. J. Biol. Chem. 2009, 284, 25488–25500. [Google Scholar] [CrossRef] [PubMed]
- Bollag, W.B.; Gonzales, J.N. Phosphatidylglycerol and surfactant: A potential treatment for COVID-19? Med. Hypotheses 2020, 144, 110277. [Google Scholar] [CrossRef]
- Zeisel, S.H. A brief history of choline. Ann. Nutr. Metab. 2012, 61, 254–258. [Google Scholar] [CrossRef]
- Lassègue, B.; Alexander, R.W.; Clark, M.; Akers, M.; Griendling, K.K. Phosphatidylcholine is a major source of phosphatidic acid and diacylglycerol in angiotensin II-stimulated vascular smooth-muscle cells. Biochem. J. 1993, 292, 509–517. [Google Scholar] [CrossRef]
- Bozelli, J.C.; Azher, S.; Epand, R.M. Plasmalogens and chronic inflammatory diseases. Front. Physiol. 2021, 12, 730829. [Google Scholar] [CrossRef]
- Koivuniemi, A. The biophysical properties of plasmalogens originating from their unique molecular architecture. FEBS Lett. 2017, 591, 2700–2713. [Google Scholar] [CrossRef]
- Goss, V.; Hunt, A.N.; Postle, A.D. Regulation of lung surfactant phospholipid synthesis and metabolism. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2013, 1831, 448–458. [Google Scholar] [CrossRef]
- Escribá, P.V.; González-Ros, J.M.; Goñi, F.M.; Kinnunen, P.K.J.; Vigh, L.; Sánchez-Magraner, L.; Fernández, A.M.; Busquets, X.; Horváth, I.; Barceló-Coblijn, G. Membranes: A meeting point for lipids, proteins and therapies. J. Cell. Mol. Med. 2008, 12, 829–875. [Google Scholar] [CrossRef]
- Cui, Z.; Houweling, M. Phosphatidylcholine and cell death. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2002, 1585, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Robertson, B. Lung surfactant for replacement therapy. Clin. Physiol. 1983, 3, 97–110. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Kim, J.; Huang, Z.; St. Clair, J.R.; Brown, D.A.; London, E. Efficient replacement of plasma membrane outer leaflet phospholipids and sphingolipids in cells with exogenous lipids. Proc. Natl. Acad. Sci. USA 2016, 113, 14025–14030. [Google Scholar] [CrossRef] [PubMed]
- Sarri, E.; Sicart, A.; Lázaro-Diéguez, F.; Egea, G. Phospholipid synthesis participates in the regulation of diacylglycerol required for membrane trafficking at the Golgi complex. J. Biol. Chem. 2011, 286, 28632–28643. [Google Scholar] [CrossRef] [PubMed]
- Vance, D.E.; Choy, P.C. How is phosphatidylcholine biosynthesis regulated? Trends Biochem. Sci. 1979, 4, 145–148. [Google Scholar] [CrossRef]
- Kennedy, E.P. The synthesis of cytidine diphosphate choline, cytidine diphosphate ethanolamine, and related compounds. J. Biol. Chem. 1956, 222, 185–191. [Google Scholar] [CrossRef]
- Cornell, R.B.; Ridgway, N.D. CTP: Phosphocholine cytidylyltransferase: Function, regulation, and structure of an amphitropic enzyme required for membrane biogenesis. Prog. Lipid Res. 2015, 59, 147–171. [Google Scholar] [CrossRef]
- Vance, D.E. Glycerolipid biosynthesis in eukaryotes. In Biochemistry of Lipids, Lipoproteins and Membranes; Vance, D.E., Vance, J.E., Eds.; Elsevier: Amsterdam, The Netherlands, 1996; pp. 153–181. [Google Scholar]
- Dawson, G. Measuring brain lipids. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2015, 1851, 1026–1039. [Google Scholar] [CrossRef]
- Van der Veen, J.N.; Lingrell, S.; Vance, D.E. The membrane lipid phosphatidylcholine is an unexpected source of triacylglycerol in the liver. J. Biol. Chem. 2012, 287, 23418–23426. [Google Scholar] [CrossRef]
- Kanno, K.; Wu, M.K.; Scapa, E.F.; Roderick, S.L.; Cohen, D.E. Structure and function of phosphatidylcholine transfer protein (PC-TP)/StarD2. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2007, 1771, 654–662. [Google Scholar] [CrossRef]
- Lagarde, M.; Hachem, M.; Picq, M.; Guichardant, M.; Bernoud-Hubac, N. AceDoPC, a structured phospholipid to target the brain with docosahexaenoic acid. OCL 2016, 23, D102. [Google Scholar] [CrossRef]
- Thudichum, J.L.W. A treatise on the chemical constitution of the brain: Based throughout upon original researches. Glas. Med. J. 1884, 22, 363–364. [Google Scholar]
- Patel, D.; Witt, S.N. Ethanolamine and phosphatidylethanolamine: Partners in health and disease. Oxid. Med. Cell. Longev. 2017, 2017, 4829180. [Google Scholar] [CrossRef] [PubMed]
- Vance, J.E.; Tasseva, G. Formation and function of phosphatidylserine and phosphatidylethanolamine in mammalian cells. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2013, 1831, 543–554. [Google Scholar] [CrossRef]
- Borkenhagen, L.F.; Kennedy, E.P.; Fielding, L. Enzymatic formation and decarboxylation of phosphatidylserine. J. Biol. Chem. 1961, 236, PC28–PC30. [Google Scholar] [CrossRef]
- Sundler, R.; Åkesson, B.; Nilsson, Å. Quantitative role of base exchange in phosphatidylethanolamine synthesis in isolated rat hepatocytes. FEBS Lett. 1974, 43, 303–307. [Google Scholar] [CrossRef]
- Van Veldhoven, P.P.; Gijsbers, S.; Mannaerts, G.P.; Vermeesch, J.R.; Brys, V. Human sphingosine-1-phosphate lyase: cDNA cloning, functional expression studies and mapping to chromosome 10q2211DNA sequence was deposited in the EMBL database (AJ011304). Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2000, 1487, 128–134. [Google Scholar] [CrossRef]
- Riekhof, W.R.; Wu, J.; Jones, J.L.; Voelker, D.R. Identification and characterization of the major lysophosphatidylethanolamine acyltransferase in saccharomyces cerevisiae. J. Biol. Chem. 2007, 282, 28344–28352. [Google Scholar] [CrossRef]
- Bouchet, A.M.; Frías, M.A.; Lairion, F.; Martini, F.; Almaleck, H.; Gordillo, G.; Disalvo, E.A. Structural and dynamical surface properties of phosphatidylethanolamine containing membranes. Biochim. Biophys. Acta Biomembr. 2009, 1788, 918–925. [Google Scholar] [CrossRef][Green Version]
- McIntosh, T.J. Differences in hydrocarbon chain tilt between hydrated phosphatidylethanolamine and phosphatidylcholine bilayers. A molecular packing model. Biophys. J. 1980, 29, 237–245. [Google Scholar] [CrossRef]
- Dawaliby, R.; Trubbia, C.; Delporte, C.; Noyon, C.; Ruysschaert, J.-M.; Van Antwerpen, P.; Govaerts, C. Phosphatidylethanolamine is a key regulator of membrane fluidity in eukaryotic cells. J. Biol. Chem. 2016, 291, 3658–3667. [Google Scholar] [CrossRef] [PubMed]
- Van der Veen, J.N.; Kennelly, J.P.; Wan, S.; Vance, J.E.; Vance, D.E.; Jacobs, R.L. The critical role of phosphatidylcholine and phosphatidylethanolamine metabolism in health and disease. Biochim. Biophys. Acta Biomembr. 2017, 1859, 1558–1572. [Google Scholar] [CrossRef] [PubMed]
- Kreutzberger, A.J.B.; Kiessling, V.; Liang, B.; Yang, S.-T.; Castle, J.D.; Tamm, L.K. Asymmetric phosphatidylethanolamine distribution controls fusion pore lifetime and probability. Biophys. J. 2017, 113, 1912–1915. [Google Scholar] [CrossRef] [PubMed]
- Pohl, E.E.; Jovanovic, O. The role of phosphatidylethanolamine adducts in modification of the activity of membrane proteins under oxidative stress. Molecules 2019, 24, 4545. [Google Scholar] [CrossRef]
- Strandberg, E.; Tiltak, D.; Ehni, S.; Wadhwani, P.; Ulrich, A.S. Lipid shape is a key factor for membrane interactions of amphipathic helical peptides. Biochim. Biophys. Acta Biomembr. 2012, 1818, 1764–1776. [Google Scholar] [CrossRef]
- Tsuboi, K.; Uyama, T.; Okamoto, Y.; Ueda, N. Endocannabinoids and related N-acylethanolamines: Biological activities and metabolism. Inflamm. Regen. 2018, 38, 28. [Google Scholar] [CrossRef]
- Jin, X.-H.; Okamoto, Y.; Morishita, J.; Tsuboi, K.; Tonai, T.; Ueda, N. Discovery and characterization of a Ca2+-independent phosphatidylethanolamine N-acyltransferase generating the anandamide precursor and its congeners. J. Biol. Chem. 2007, 282, 3614–3623. [Google Scholar] [CrossRef]
- Folch, J.; Schneider, H.A. an amino acid constituent of the Ox brain cephalin. J. Biol. Chem. 1941, 137, 51–62. [Google Scholar] [CrossRef]
- Vance, J.E. Historical perspective: Phosphatidylserine and phosphatidylethanolamine from the 1800s to the present. J. Lipid Res. 2018, 59, 923–944. [Google Scholar] [CrossRef]
- Svennerholm, L. Distribution and fatty acid composition of phosphoglycerides in normal human brain. J. Lipid Res. 1968, 9, 570–579. [Google Scholar] [CrossRef]
- Gardner, R.G.; Hampton, R.Y. A highly conserved signal controls degradation of 3-hydroxy-3-methylglutaryl-coenzyme A (HMG-CoA) reductase in eukaryotes. J. Biol. Chem. 1999, 274, 31671–31678. [Google Scholar] [CrossRef] [PubMed]
- Kanfer, J.; Kennedy, E.P. Metabolism and function of bacterial lipids. J. Biol. Chem. 1964, 239, 1720–1726. [Google Scholar] [CrossRef] [PubMed]
- Hübscher, G.; Dils, R.R.; Pover, W.F.R. Studies on the biosynthesis of phosphatidyl serine. Biochim. Biophys. Acta 1959, 36, 518–528. [Google Scholar] [CrossRef]
- Van Meer, G. Dynamic transbilayer lipid asymmetry. Cold Spring Harb. Perspect. Biol. 2011, 3, a004671. [Google Scholar] [CrossRef] [PubMed]
- Fadok, V.A.; Voelker, D.R.; Campbell, P.A.; Cohen, J.J.; Bratton, D.L.; Henson, P.M. Exposure of phosphatidylserine on the surface of apoptotic lymphocytes triggers specific recognition and removal by macrophages. J. Immunol. 1992, 148, 2207–2216. [Google Scholar] [CrossRef] [PubMed]
- Vallabhapurapu, S.D.; Blanco, V.M.; Sulaiman, M.K.; Vallabhapurapu, S.L.; Chu, Z.; Franco, R.S.; Qi, X. Variation in human cancer cell external phosphatidylserine is regulated by flippase activity and intracellular calcium. Oncotarget 2015, 6, 34375–34388. [Google Scholar] [CrossRef]
- Skotland, T.; Sandvig, K. The role of PS 18:0/18:1 in membrane function. Nat. Commun. 2019, 10, 2752. [Google Scholar] [CrossRef]
- Ma, X.; Li, X.; Wang, W.; Zhang, M.; Yang, B.; Miao, Z. Phosphatidylserine, inflammation, and central nervous system diseases. Front. Aging Neurosci. 2022, 14, 975176. [Google Scholar] [CrossRef]
- Naeini, M.B.; Bianconi, V.; Pirro, M.; Sahebkar, A. The role of phosphatidylserine recognition receptors in multiple biological functions. Cell. Mol. Biol. Lett. 2020, 25, 23. [Google Scholar] [CrossRef]
- Merolli, A.; Santin, M. Role of phosphatidyl-serine in bone repair and its technological exploitation. Molecules 2009, 14, 5367–5381. [Google Scholar] [CrossRef]
- Lentz, B.R. Exposure of platelet membrane phosphatidylserine regulates blood coagulation. Prog. Lipid Res. 2003, 42, 423–438. [Google Scholar] [CrossRef]
- Anderson, R.J. The chemistry of the lipoids of Tubercle bacilli. XIV. The occurence of inosite in the phosphatide from human Tubercle bacilli1. J. Am. Chem. Soc. 1930, 52, 1607–1608. [Google Scholar] [CrossRef]
- Pizer, F.L.; Ballou, C.E. Studies on myo-Inositol phosphates of natural origin. J. Am. Chem. Soc. 1959, 81, 915–921. [Google Scholar] [CrossRef]
- Dickson, E.J.; Hille, B. Understanding phosphoinositides: Rare, dynamic, and essential membrane phospholipids. Biochem. J. 2019, 476, 1–23. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, K.; Epand, R.M. Enrichment of phosphatidylinositols with specific acyl chains. Biochim. Biophys. Acta Biomembr. 2014, 1838, 1501–1508. [Google Scholar] [CrossRef] [PubMed]
- Payrastre, B.; Missy, K.; Giuriato, S.; Bodin, S.; Plantavid, M.; Gratacap, M.-P. Phosphoinositides: Key players in cell signalling, in time and space. Cell. Signal. 2001, 13, 377–387. [Google Scholar] [CrossRef]
- Blunsom, N.J.; Cockcroft, S. Phosphatidylinositol synthesis at the endoplasmic reticulum. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2020, 1865, 158471. [Google Scholar] [CrossRef]
- Daniels, C.J.; Palmer, F.B.S.C. Biosynthesis of phosphatidylinositol in Crithidia fasciculata. Biochim. Biophys. Acta Lipids Lipid Metab. 1980, 618, 263–272. [Google Scholar] [CrossRef]
- Jorge, C.D.; Borges, N.; Santos, H. A novel pathway for the synthesis of inositol phospholipids uses cytidine diphosphate (CDP)-inositol as donor of the polar head group. Environ. Microbiol. 2015, 17, 2492–2504. [Google Scholar] [CrossRef]
- Balla, T. Phosphoinositides: Tiny lipids with giant impact on cell regulation. Physiol. Rev. 2013, 93, 1019–1137. [Google Scholar] [CrossRef]
- Anderson, K.E.; Juvin, V.; Clark, J.; Stephens, L.R.; Hawkins, P.T. Investigating the effect of arachidonate supplementation on the phosphoinositide content of MCF10a breast epithelial cells. Adv. Biol. Regul. 2016, 62, 18–24. [Google Scholar] [CrossRef] [PubMed]
- Anderson, K.E.; Kielkowska, A.; Durrant, T.N.; Juvin, V.; Clark, J.; Stephens, L.R.; Hawkins, P.T. Lysophosphatidylinositol-Acyltransferase-1 (LPIAT1) Is Required to Maintain Physiological Levels of PtdIns and PtdInsP2 in the Mouse. PLoS ONE 2013, 8, e58425. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Yang, J.S.; Park, S.M.; Byeon, S.K.; Moon, M.H. On-line high speed lipid extraction for nanoflow liquid chromatography-tandem mass spectrometry. J. Chromatogr. A 2016, 1464, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Yui, K.; Imataka, G.; Nakamura, H.; Ohara, N.; Naito, Y. Eicosanoids derived from arachidonic acid and their family prostaglandins and cyclooxygenase in psychiatric disorders. Curr. Neuropharmacol. 2015, 13, 776–785. [Google Scholar] [CrossRef]
- Piñeiro, R.; Falasca, M. Lysophosphatidylinositol signalling: New wine from an old bottle. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2012, 1821, 694–705. [Google Scholar] [CrossRef]
- Ferguson, M.A.J.; Williams, A.F. Cell-surface anchoring of proteins via glycosyl-phosphatidylinositol structures. Annu. Rev. Biochem. 1988, 57, 285–320. [Google Scholar] [CrossRef]
- Posor, Y.; Jang, W.; Haucke, V. Phosphoinositides as membrane organizers. Nat. Rev. Mol. Cell Biol. 2022, 23, 797–816. [Google Scholar] [CrossRef]
- Pangborn, M.C. Isolation and purification of a serologically active phospholipid from beef heart. J. Biol. Chem. 1942, 143, 247–256. [Google Scholar] [CrossRef]
- Schlame, M.; Haldar, D. Cardiolipin is synthesized on the matrix side of the inner membrane in rat liver mitochondria. J. Biol. Chem. 1993, 268, 74–79. [Google Scholar] [CrossRef]
- Acoba, M.G.; Senoo, N.; Claypool, S.M. Phospholipid ebb and flow makes mitochondria go. J. Cell Biol. 2020, 219, e202003131. [Google Scholar] [CrossRef]
- Luévano-Martínez, L.A.; Duncan, A.L. Origin and diversification of the cardiolipin biosynthetic pathway in the Eukarya domain. Biochem. Soc. Trans. 2020, 48, 1035–1046. [Google Scholar] [CrossRef] [PubMed]
- Hatch, G.M. Cell biology of cardiac mitochondrial phospholipids. Biochem. Cell Biol. 2004, 82, 99–112. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Shen, T.; Huynh, H.; Fang, X.; Han, Z.; Ouyang, K. Cardiolipin regulates mitochondrial ultrastructure and function in mammalian cells. Genes 2022, 13, 1889. [Google Scholar] [CrossRef] [PubMed]
- Poulaki, A.; Giannouli, S. Mitochondrial lipids: From membrane organization to apoptotic facilitation. Int. J. Mol. Sci. 2022, 23, 3738. [Google Scholar] [CrossRef] [PubMed]
- Osman, C.; Voelker, D.R.; Langer, T. Making heads or tails of phospholipids in mitochondria. J. Cell Biol. 2011, 192, 7–16. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Beuchat, M.-H.; Lindsay, M.; Frias, S.; Palmiter, R.D.; Sakuraba, H.; Parton, R.G.; Gruenberg, J. Late endosomal membranes rich in lysobisphosphatidic acid regulate cholesterol transport. Nat. Cell Biol. 1999, 1, 113–118. [Google Scholar] [CrossRef]
- Kobayashi, T.; Stang, E.; Fang, K.S.; de Moerloose, P.; Parton, R.G.; Gruenberg, J. A lipid associated with the antiphospholipid syndrome regulates endosome structure and function. Nature 1998, 392, 193–197. [Google Scholar] [CrossRef]
- Wherrett, J.R.; Huterer, S. Enrichment of Bis-(monoacylglyceryl) phosphate in lysosomes from rat liver. J. Biol. Chem. 1972, 247, 4114–4120. [Google Scholar] [CrossRef]
- Body, D.R.; Gray, G.M. The isolation and characterisation of phosphatidylglycerol and a structural isomer from pig lung. Chem. Phys. Lipids 1967, 1, 254–263. [Google Scholar] [CrossRef]
- Brotherus, J.; Renkonen, O. Isolation and characterisation of bis-phosphatidic acid and its partially deacylated derivatives from cultured BHK-cells. Chem. Phys. Lipids 1974, 13, 11–20. [Google Scholar] [CrossRef]
- Hullin-Matsuda, F.; Kawasaki, K.; Delton-Vandenbroucke, I.; Xu, Y.; Nishijima, M.; Lagarde, M.; Schlame, M.; Kobayashi, T. De novo biosynthesis of the late endosome lipid, bis(monoacylglycero)phosphate. J. Lipid Res. 2007, 48, 1997–2008. [Google Scholar] [CrossRef]
- Waite, M.; Roddick, V.; Thornburg, T.; King, L.; Cochran, F. Conversion of phosphatidylglycerol to lyso(bis)phosphatidic acid by alveolar macrophages. FASEB J. 1987, 1, 318–325. [Google Scholar] [CrossRef]
- Poorthuis, B.J.; Hostetler, K.Y. Conversion of diphosphatidylglycerol to bis(monoacylglyceryl)phosphate by lysosomes. J. Lipid Res. 1978, 19, 309–315. [Google Scholar] [CrossRef]
- Showalter, M.R.; Berg, A.L.; Nagourney, A.; Heil, H.; Carraway, K.L.; Fiehn, O. The emerging and diverse roles of Bis(monoacylglycero) Phosphate lipids in cellular physiology and disease. Int. J. Mol. Sci. 2020, 21, 8067. [Google Scholar] [CrossRef]
- Hostetler, K.Y. Chapter 6 Polyglycerophospholipids: Phosphatidylglycerol, diphosphatidylglycerol and bis (monoacylglycero) phosphate. In New Comprehensive Biochemistry; Hawthorne, N.J., Ansell, G.B., Eds.; Elsevier: Amsterdam, The Netherlands, 1982; Volume 4, pp. 215–261. [Google Scholar]
- Akgoc, Z.; Iosim, S.; Seyfried, T.N. Bis(monoacylglycero)phosphate as a macrophage enriched phospholipid. Lipids 2015, 50, 907–912. [Google Scholar] [CrossRef] [PubMed]
- Matsuzawa, Y.; Hostetler, K.Y. Studies on drug-induced lipidosis: Subcellular localization of phospholipid and cholesterol in the liver of rats treated with chloroquine or 4,4′-bis (diethylaminoethoxy)alpha, beta-diethyldiphenylethane. J. Lipid Res. 1980, 21, 202–214. [Google Scholar] [CrossRef] [PubMed]
- Rouser, G.; Kritchevsky, G.; Knudson, A.G.; Simon, G. Accumulation of a glycerolphospholipid in classical niemann-pick disease. Lipids 1968, 3, 287–290. [Google Scholar] [CrossRef] [PubMed]
- Gallala, H.D.; Sandhoff, K. Biological function of the cellular lipid BMP—BMP as a key activator for cholesterol sorting and membrane digestion. Neurochem. Res. 2011, 36, 1594–1600. [Google Scholar] [CrossRef]
- Snyder, F.; Lee, T.; Wykle, R.L. Ether-linked glycerolipids and their bioactive species: Enzymes and metabolic regulation. In The Enzymes of Biological Membranes; Martonosi, A.N., Ed.; Springer: Boston, MA, USA, 1985; pp. 1–58. [Google Scholar]
- Benveniste, J.; Henson, P.M.; Cochrane, C.G. Leukocyte-dependent histamine release from rabbit platelets. The role of IgE, basophils, and a platelet-activating factor. J. Exp. Med. 1972, 136, 1356–1377. [Google Scholar] [CrossRef]
- Prescott, S.M.; Zimmerman, G.A.; Stafforini, D.M.; McIntyre, T.M. Platelet-activating factor and related lipid mediators. Annu. Rev. Biochem. 2000, 69, 419–445. [Google Scholar] [CrossRef]
- Hishikawa, D.; Hashidate, T.; Shimizu, T.; Shindou, H. Diversity and function of membrane glycerophospholipids generated by the remodeling pathway in mammalian cells. J. Lipid Res. 2014, 55, 799–807. [Google Scholar] [CrossRef] [PubMed]
- Tsoupras, A.; Lordan, R.; Zabetakis, I. Inflammation, not cholesterol, is a cause of chronic disease. Nutrients 2018, 10, 604. [Google Scholar] [CrossRef] [PubMed]
- Roudebush, W.E. Function of platelet-activating factor in spermatozoa motility and fertility potential. Glob. J. Reprod. Med. 2022, 8, 5556749. [Google Scholar] [CrossRef]
- Imaizumi, T.-A.; Stafforini, D.M.; Yamada, Y.; Mcintyre, T.M.; Prescott, S.M.; Zimmerman, G.A. Platelet-activating factor: A mediator for clinicians. J. Intern. Med. 1995, 238, 5–20. [Google Scholar] [CrossRef]
- Schlondorff, D.; Neuwirth, R. Platelet-activating factor and the kidney. Am. J. Physiol. Physiol. 1986, 251, F1–F11. [Google Scholar] [CrossRef]
- Travers, J.B.; Rohan, J.G.; Sahu, R.P. New insights into the pathologic roles of the platelet-activating factor system. Front. Endocrinol. 2021, 12, 624132. [Google Scholar] [CrossRef]
- Zimmerman, G.A.; McIntyre, T.M.; Prescott, S.M.; Stafforini, D.M. The platelet-activating factor signaling system and its regulators in syndromes of inflammation and thrombosis. Crit. Care Med. 2002, 30, S294–S301. [Google Scholar] [CrossRef]
- Makide, K.; Kitamura, H.; Sato, Y.; Okutani, M.; Aoki, J. Emerging lysophospholipid mediators, lysophosphatidylserine, lysophosphatidylthreonine, lysophosphatidylethanolamine and lysophosphatidylglycerol. Prostaglandins Other Lipid Mediat. 2009, 89, 135–139. [Google Scholar] [CrossRef]
- Meyer zu Heringdorf, D.; Jakobs, K.H. Lysophospholipid receptors: Signalling, pharmacology and regulation by lysophospholipid metabolism. Biochim. Biophys. Acta Biomembr. 2007, 1768, 923–940. [Google Scholar] [CrossRef]
- Hannun, Y.A.; Bell, R.M. Lysosphingolipids inhibit protein kinase C: Implications for the sphingolipidoses. Science 1987, 235, 670–674. [Google Scholar] [CrossRef]
- Anliker, B.; Chun, J. Lysophospholipid G protein-coupled receptors. J. Biol. Chem. 2004, 279, 20555–20558. [Google Scholar] [CrossRef] [PubMed]
- Law, S.-H.; Chan, M.-L.; Marathe, G.K.; Parveen, F.; Chen, C.-H.; Ke, L.-Y. An updated review of lysophosphatidylcholine metabolism in human diseases. Int. J. Mol. Sci. 2019, 20, 1149. [Google Scholar] [CrossRef] [PubMed]
- Engel, K.M.; Schiller, J.; Galuska, C.E.; Fuchs, B. Phospholipases and Reactive Oxygen Species Derived Lipid Biomarkers in Healthy and Diseased Humans and Animals—A Focus on Lysophosphatidylcholine. Front. Physiol. 2021, 12, 732319. [Google Scholar] [CrossRef] [PubMed]
- Hellenthal, K.E.M.; Brabenec, L.; Gross, E.R.; Wagner, N.-M. TRP channels as sensors of aldehyde and oxidative stress. Biomolecules 2021, 11, 1401. [Google Scholar] [CrossRef]
- Aoki, J. Two pathways for lysophosphatidic acid production. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2008, 1781, 513–518. [Google Scholar] [CrossRef]
- Lin, Y.-H.; Lin, Y.-C.; Chen, C.-C. Lysophosphatidic acid receptor antagonists and cancer: The current trends, clinical implications, and trials. Cells 2021, 10, 1629. [Google Scholar] [CrossRef]
- Lin, M.-E.; Herr, D.R.; Chun, J. Lysophosphatidic acid (LPA) receptors: Signaling properties and disease relevance. Prostaglandins Other Lipid Mediat. 2010, 91, 130–138. [Google Scholar] [CrossRef]
- Yeagle, P.L. Cholesterol and the cell membrane. Biochim. Biophys. Acta Rev. Biomembr. 1985, 822, 267–287. [Google Scholar] [CrossRef]
- Haas, E.; Kim, Y.; Stanley, D. Why can insects not biosynthesize cholesterol? Arch. Insect Biochem. Physiol. 2023, 112, e21983. [Google Scholar] [CrossRef]
- Bittman, R. Has nature designed the cholesterol side chain for optimal interaction with phospholipids? In Cholesterol: Subcellular Biochemistry; Bittman, R., Ed.; Springer: Boston, MA, USA, 1997; pp. 145–171. [Google Scholar]
- Kroon, P.A.; Kainosho, M.; Chan, S.I. State of molecular motion of cholesterol in lecithin bilayers. Nature 1975, 256, 582–584. [Google Scholar] [CrossRef]
- Lönnfors, M.; Långvik, O.; Björkbom, A.; Slotte, J.P. Cholesteryl phosphocholine—A study on its interactions with ceramides and other membrane lipids. Langmuir 2013, 29, 2319–2329. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, F.; Nemecz, G.; Gibson Wood, W.; Joiner, C.; Morrot, G.; Ayraut-Jarrier, M.; Devaux, P.F. Transmembrane distribution of sterol in the human erythrocyte. Biochim. Biophys. Acta Biomembr. 1991, 1066, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Mondal, M.; Mesmin, B.; Mukherjee, S.; Maxfield, F.R. Sterols are mainly in the cytoplasmic leaflet of the plasma membrane and the endocytic recycling compartment in CHO cells. Mol. Biol. Cell 2009, 20, 581–588. [Google Scholar] [CrossRef] [PubMed]
- Kuijpers, P.M.J.C. History in medicine: The story of cholesterol, lipids and cardiology. e-J. Cardiol. Pract. 2021, 19, 1–5. [Google Scholar]
- Cockcroft, S. Mammalian lipids: Structure, synthesis and function. Essays Biochem. 2021, 65, 813–845. [Google Scholar] [CrossRef]
- Luo, J.; Jiang, L.-Y.; Yang, H.; Song, B.-L. Intracellular cholesterol transport by sterol transfer proteins at membrane contact sites. Trends Biochem. Sci. 2019, 44, 273–292. [Google Scholar] [CrossRef]
- Frallicciardi, J.; Melcr, J.; Siginou, P.; Marrink, S.J.; Poolman, B. Membrane thickness, lipid phase and sterol type are determining factors in the permeability of membranes to small solutes. Nat. Commun. 2022, 13, 1605. [Google Scholar] [CrossRef]
- De Oliveira Andrade, L. Understanding the role of cholesterol in cellular biomechanics and regulation of vesicular trafficking: The power of imaging. Biomed. Spectrosc. Imaging 2016, 5, S101–S117. [Google Scholar] [CrossRef]
- Demel, R.A.; Van Deenen, L.L.M.; Pethica, B.A. Monolayer interactions of phospholipids and cholesterol. Biochim. Biophys. Acta Biomembr. 1967, 135, 11–19. [Google Scholar] [CrossRef]
- Niu, S.-L.; Litman, B.J. Determination of membrane cholesterol partition coefficient using a lipid vesicle–cyclodextrin binary system: Effect of phospholipid acyl chain unsaturation and headgroup composition. Biophys. J. 2002, 83, 3408–3415. [Google Scholar] [CrossRef]
- Shaikh, S.R.; Cherezov, V.; Caffrey, M.; Soni, S.P.; LoCascio, D.; Stillwell, W.; Wassall, S.R. Molecular organization of cholesterol in unsaturated phosphatidylethanolamines: X-ray diffraction and solid state 2 H NMR reveal differences with phosphatidylcholines. J. Am. Chem. Soc. 2006, 128, 5375–5383. [Google Scholar] [CrossRef] [PubMed]
- Brzustowicz, M.R.; Cherezov, V.; Zerouga, M.; Caffrey, M.; Stillwell, W.; Wassall, S.R. Controlling membrane cholesterol content. A role for polyunsaturated (docosahexaenoate) phospholipids. Biochemistry 2002, 41, 12509–12519. [Google Scholar] [CrossRef] [PubMed]
- Siminovitch, D.J.; Jeffrey, K.R. Orientational order in the choline headgroup of sphingomyelin: A 14N-NMR study. Biochim. Biophys. Acta Biomembr. 1981, 645, 270–278. [Google Scholar] [CrossRef] [PubMed]
- Róg, T.; Pasenkiewicz-Gierula, M. Cholesterol-sphingomyelin interactions: A molecular dynamics simulation study. Biophys. J. 2006, 91, 3756–3767. [Google Scholar] [CrossRef] [PubMed]
- Lönnfors, M.; Doux, J.P.F.; Killian, J.A.; Nyholm, T.K.M.; Slotte, J.P. Sterols have higher affinity for sphingomyelin than for phosphatidylcholine bilayers even at equal acyl-chain order. Biophys. J. 2011, 100, 2633–2641. [Google Scholar] [CrossRef]
- Chiantia, S.; London, E. Acyl chain length and saturation modulate interleaflet coupling in asymmetric bilayers: Effects on dynamics and structural order. Biophys. J. 2012, 103, 2311–2319. [Google Scholar] [CrossRef]
- Slotte, J.P. The importance of hydrogen bonding in sphingomyelin’s membrane interactions with co-lipids. Biochim. Biophys. Acta Biomembr. 2016, 1858, 304–310. [Google Scholar] [CrossRef]
- Sohrabi, Y.; Reinecke, H.; Godfrey, R. Altered cholesterol and lipid synthesis Mediates hyperinflammation in COVID-19. Trends Endocrinol. Metab. 2021, 32, 132–134. [Google Scholar] [CrossRef]
- King, R.J.; Singh, P.K.; Mehla, K. The cholesterol pathway: Impact on immunity and cancer. Trends Immunol. 2022, 43, 78–92. [Google Scholar] [CrossRef]
- Kopecka, J.; Trouillas, P.; Gašparović, A.Č.; Gazzano, E.; Assaraf, Y.G.; Riganti, C. Phospholipids and cholesterol: Inducers of cancer multidrug resistance and therapeutic targets. Drug Resist. Updat. 2020, 49, 100670. [Google Scholar] [CrossRef]
- Hu, J.; Zhang, Z.; Shen, W.-J.; Azhar, S. Cellular cholesterol delivery, intracellular processing and utilization for biosynthesis of steroid hormones. Nutr. Metab. 2010, 7, 47. [Google Scholar] [CrossRef] [PubMed]
- Panveliwalla, D.; Lewis, B.; Wootton, I.D.P.; Tabaqchali, S. Determination of individual bile acids in biological fluids by thin-layer chromatography and fluorimetry. J. Clin. Pathol. 1970, 23, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Kuo, T.; McQueen, A.; Chen, T.-C.; Wang, J.-C. Regulation of glucose homeostasis by glucocorticoids. In Glucocorticoid Signaling from Molecules to Mice to Man; Wang, J.-C., Harris, C., Eds.; Springer: New York, NY, USA, 2015; pp. 99–126. [Google Scholar]
- Wollam, J.; Antebi, A. Sterol regulation of metabolism, homeostasis, and development. Annu. Rev. Biochem. 2011, 80, 885–916. [Google Scholar] [CrossRef] [PubMed]
- Christiansen, J.J.; Djurhuus, C.B.; Gravholt, C.H.; Iversen, P.; Christiansen, J.S.; Schmitz, O.; Weeke, J.; Jørgensen, J.O.L.; Møller, N. Effects of cortisol on carbohydrate, lipid, and protein metabolism: Studies of acute cortisol withdrawal in adrenocortical failure. J. Clin. Endocrinol. Metab. 2007, 92, 3553–3559. [Google Scholar] [CrossRef]
- Schwartz, C.C.; VandenBroek, J.M.; Cooper, P.S. Lipoprotein cholesteryl ester production, transfer, and output in vivo in humans. J. Lipid Res. 2004, 45, 1594–1607. [Google Scholar] [CrossRef]
- Breslow, D.K. Sphingolipid homeostasis in the endoplasmic reticulum and beyond. Cold Spring Harb. Perspect. Biol. 2013, 5, a013326. [Google Scholar] [CrossRef]
- Carter, H.E.; Glick, F.J.; Norris, W.P.; Phillips, G.E. Biochemistry of the sphingolipids. J. Biol. Chem. 1947, 170, 285–294. [Google Scholar] [CrossRef]
- Tidhar, R.; Futerman, A.H. The complexity of sphingolipid biosynthesis in the endoplasmic reticulum. Biochim. Biophys. Acta Mol. Cell Res. 2013, 1833, 2511–2518. [Google Scholar] [CrossRef]
- Eliot, A.C.; Kirsch, J.F. Pyridoxal phosphate enzymes: Mechanistic, structural, and evolutionary considerations. Annu. Rev. Biochem. 2004, 73, 383–415. [Google Scholar] [CrossRef]
- Eichler, F.S.; Hornemann, T.; McCampbell, A.; Kuljis, D.; Penno, A.; Vardeh, D.; Tamrazian, E.; Garofalo, K.; Lee, H.-J.; Kini, L.; et al. Overexpression of the Wild-Type SPT1 Subunit Lowers Desoxysphingolipid Levels and Rescues the Phenotype of HSAN1. J. Neurosci. 2009, 29, 14646–14651. [Google Scholar] [CrossRef]
- Rosen, H.; Gonzalez-Cabrera, P.J.; Sanna, M.G.; Brown, S. Sphingosine 1-phosphate receptor signaling. Annu. Rev. Biochem. 2009, 78, 743–768. [Google Scholar] [CrossRef] [PubMed]
- Cuvillier, O.; Rosenthal, D.S.; Smulson, M.E.; Spiegel, S. Sphingosine 1-Phosphate Inhibits Activation of Caspases that Cleave Poly(ADP-ribose) Polymerase and Lamins during Fas- and Ceramide-mediated Apoptosis in Jurkat T Lymphocytes. J. Biol. Chem. 1998, 273, 2910–2916. [Google Scholar] [CrossRef] [PubMed]
- Hait, N.C.; Oskeritzian, C.A.; Paugh, S.W.; Milstien, S.; Spiegel, S. Sphingosine kinases, sphingosine 1-phosphate, apoptosis and diseases. Biochim. Biophys. Acta Biomembr. 2006, 1758, 2016–2026. [Google Scholar] [CrossRef] [PubMed]
- Merrill, A.H. De novo sphingolipid biosynthesis: A necessary, but dangerous, pathway. J. Biol. Chem. 2002, 277, 25843–25846. [Google Scholar] [CrossRef]
- Stoffel, W.; Bauer, E.; Stahl, J. The metabolism of sphingosine bases in Tetrahymena pyriformis. Sphingosine kinase and sphingosine-1-phosphate lyase. Biol. Chem. 1974, 355, 61–74. [Google Scholar] [CrossRef]
- Hannun, Y.A.; Obeid, L.M. Sphingolipids and their metabolism in physiology and disease. Nat. Rev. Mol. Cell Biol. 2018, 19, 175–191. [Google Scholar] [CrossRef]
- Zupancic, E.; Carreira, A.C.; de Almeida, R.F.M.; Silva, L.C. Biophysical implications of sphingosine accumulation in membrane properties at neutral and acidic pH. J. Phys. Chem. B 2014, 118, 4858–4866. [Google Scholar] [CrossRef]
- Merrill, A.H. Sphingolipid and glycosphingolipid metabolic pathways in the era of sphingolipidomics. Chem. Rev. 2011, 111, 6387–6422. [Google Scholar] [CrossRef]
- Katoh, N. Modulation by sphingosine of phosphorylation of substrate proteins by protein kinase C in nuclei from cow mammary gland. J. Vet. Med. Sci. 2004, 66, 1237–1242. [Google Scholar] [CrossRef]
- Natarajan, V.; Jayaram, H.N.; Scribner, W.M.; Garcia, J.G. Activation of endothelial cell phospholipase D by sphingosine and sphingosine-1-phosphate. Am. J. Respir. Cell Mol. Biol. 1994, 11, 221–229. [Google Scholar] [CrossRef]
- Tomonaga, N.; Tsuduki, T.; Manabe, Y.; Sugawara, T. Sphingoid bases of dietary ceramide 2-aminoethylphosphonate, a marine sphingolipid, absorb into lymph in rats. J. Lipid Res. 2019, 60, 333–340. [Google Scholar] [CrossRef] [PubMed]
- Riley, R.T.; Wang, E.; Schroeder, J.J.; Smith, E.R.; Plattner, R.D.; Abbas, H.; Yoo, H.-S.; Merrill, A.H. Evidence for disruption of sphingolipid metabolism as a contributing factor in the toxicity and carcinogenicity of fumonisins. Nat. Toxins 1996, 4, 3–15. [Google Scholar] [CrossRef] [PubMed]
- Iwabuchi, K.; Nakayama, H.; Oizumi, A.; Suga, Y.; Ogawa, H.; Takamori, K. Role of ceramide from glycosphingolipids and Its metabolites in immunological and inflammatory responses in humans. Mediat. Inflamm. 2015, 2015, 120748. [Google Scholar] [CrossRef] [PubMed]
- Fanani, M.L.; Maggio, B. The many faces (and phases) of ceramide and sphingomyelin I—Single lipids. Biophys. Rev. 2017, 9, 589–600. [Google Scholar] [CrossRef]
- Fürst, W.; Sandhoff, K. Activator proteins and topology of lysosomal sphingolipid catabolism. Biochim. Biophys. Acta Lipids Lipid Metab. 1992, 1126, 1–16. [Google Scholar] [CrossRef]
- Kitatani, K.; Idkowiak-Baldys, J.; Hannun, Y.A. The sphingolipid salvage pathway in ceramide metabolism and signaling. Cell. Signal. 2008, 20, 1010–1018. [Google Scholar] [CrossRef]
- Pewzner-Jung, Y.; Ben-Dor, S.; Futerman, A.H. When Do Lasses (Longevity Assurance Genes) Become CerS (Ceramide Synthases)? J. Biol. Chem. 2006, 281, 25001–25005. [Google Scholar] [CrossRef]
- Ho, Q.W.C.; Zheng, X.; Ali, Y. Ceramide acyl chain length and its relevance to intracellular lipid regulation. Int. J. Mol. Sci. 2022, 23, 9697. [Google Scholar] [CrossRef]
- Lachkar, F.; Ferré, P.; Foufelle, F.; Papaioannou, A. Dihydroceramides: Their emerging physiological roles and functions in cancer and metabolic diseases. Am. J. Physiol. Metab. 2021, 320, E122–E130. [Google Scholar] [CrossRef]
- Shimeno, H.; Soeda, S.; Sakamoto, M.; Kouchi, T.; Kowakame, T.; Kihara, T. Partial purification and characterization of sphingosine N-acyltransferase (ceramide synthase) from bovine liver mitochondrion-rich fraction. Lipids 1998, 33, 601–605. [Google Scholar] [CrossRef]
- Zuellig, R.A.; Hornemann, T.; Othman, A.; Hehl, A.B.; Bode, H.; Güntert, T.; Ogunshola, O.O.; Saponara, E.; Grabliauskaite, K.; Jang, J.-H.; et al. Deoxysphingolipids, Novel Biomarkers for Type 2 Diabetes, Are Cytotoxic for Insulin-Producing Cells. Diabetes 2014, 63, 1326–1339. [Google Scholar] [CrossRef] [PubMed]
- Zitomer, N.C.; Mitchell, T.; Voss, K.A.; Bondy, G.S.; Pruett, S.T.; Garnier-Amblard, E.C.; Liebeskind, L.S.; Park, H.; Wang, E.; Sullards, M.C.; et al. Ceramide synthase inhibition by Fumonisin B1 causes accumulation of 1-deoxysphinganine. J. Biol. Chem. 2009, 284, 4786–4795. [Google Scholar] [CrossRef] [PubMed]
- Eto, M.; Bennouna, J.; Hunter, O.C.; Hershberger, P.A.; Kanto, T.; Johnson, C.S.; Lotze, M.T.; Amoscato, A.A. C16 ceramide accumulates following androgen ablation in LNCaP prostate cancer cells. Prostate 2003, 57, 66–79. [Google Scholar] [CrossRef] [PubMed]
- Koybasi, S.; Senkal, C.E.; Sundararaj, K.; Spassieva, S.; Bielawski, J.; Osta, W.; Day, T.A.; Jiang, J.C.; Jazwinski, S.M.; Hannun, Y.A.; et al. Defects in cell growth regulation by C18:0-ceramide and longevity assurance gene 1 in human head and neck squamous cell carcinomas. J. Biol. Chem. 2004, 279, 44311–44319. [Google Scholar] [CrossRef] [PubMed]
- Grösch, S.; Schiffmann, S.; Geisslinger, G. Chain length-specific properties of ceramides. Prog. Lipid Res. 2012, 51, 50–62. [Google Scholar]
- Kalinichenko, L.S.; Gulbins, E.; Kornhuber, J.; Müller, C.P. Sphingolipid control of cognitive functions in health and disease. Prog. Lipid Res. 2022, 86, 101162. [Google Scholar]
- Chiantia, S.; Ries, J.; Chwastek, G.; Carrer, D.; Li, Z.; Bittman, R.; Schwille, P. Role of ceramide in membrane protein organization investigated by combined AFM and FCS. Biochim. Biophys. Acta Biomembr. 2008, 1778, 1356–1364. [Google Scholar]
- Goñi, F.M.; Alonso, A. Effects of ceramide and other simple sphingolipids on membrane lateral structure. Biochim. Biophys. Acta Biomembr. 2009, 1788, 169–177. [Google Scholar] [CrossRef]
- Hanada, K.; Kumagai, K.; Yasuda, S.; Miura, Y.; Kawano, M.; Fukasawa, M.; Nishijima, M. Molecular machinery for non-vesicular trafficking of ceramide. Nature 2003, 426, 803–809. [Google Scholar] [CrossRef]
- Fukasawa, M.; Nishijima, M.; Hanada, K. Genetic Evidence for ATP-dependent Endoplasmic Reticulum-to-Golgi Apparatus Trafficking of Ceramide for Sphingomyelin Synthesis in Chinese Hamster Ovary Cells. J. Cell Biol. 1999, 144, 673–685. [Google Scholar] [CrossRef]
- Blom, T.; Somerharju, P.; Ikonen, E. Synthesis and biosynthetic trafficking of membrane lipids. Cold Spring Harb. Perspect. Biol. 2011, 3, a004713. [Google Scholar] [CrossRef] [PubMed]
- Murakami, M.; Nakatani, Y.; Kudo, I. Type II secretory phospholipase A2 associated with cell surfaces via C-terminal heparin-binding lysine residues augments stimulus-initiated delayed prostaglandin generation. J. Biol. Chem. 1996, 271, 30041–30051. [Google Scholar] [PubMed]
- Ullman, M.D.; Radin, N.S. The enzymatic formation of sphingomyelin from ceramide and lecithin in mouse liver. J. Biol. Chem. 1974, 249, 1506–1512. [Google Scholar]
- Huitema, K.; van den Dikkenberg, J.; Brouwers, J.F.H.M.; Holthuis, J.C.M. Identification of a family of animal sphingomyelin synthases. EMBO J. 2004, 23, 33–44. [Google Scholar] [CrossRef] [PubMed]
- Futerman, A.H. Sphingolipids. In Biochemistry of Lipids, Lipoproteins and Membranes; Elsevier: Amsterdam, The Netherlands, 2021; pp. 281–316. [Google Scholar]
- Bielawski, J.; Pierce, J.S.; Snider, J.; Rembiesa, B.; Szulc, Z.M.; Bielawska, A. Sphingolipid analysis by high performance liquid chromatography-tandem mass spectrometry (HPLC-MS/MS). In Sphingolipids as Signaling and Regulatory Molecules. Advances in Experimental Medicine and Biology; Chalfant, C., Poeta, M.D., Eds.; Springer: New York, NY, USA, 2010; pp. 46–59. [Google Scholar]
- Lucki, N.C.; Sewer, M.B. Nuclear sphingolipid metabolism. Annu. Rev. Physiol. 2012, 74, 131–151. [Google Scholar]
- Birbes, H.; Luberto, C.; Hsu, Y.-T.; El Bawab, S.; Hannun, Y.A.; Obeid, L.M. A mitochondrial pool of sphingomyelin is involved in TNFα-induced Bax translocation to mitochondria. Biochem. J. 2005, 386, 445–451. [Google Scholar] [PubMed]
- Albi, E.; Mersel, M.; Leray, C.; Tomassoni, M.L.; Viola-Magni, M.P. Rat liver chromatin phospholipids. Lipids 1994, 29, 715–719. [Google Scholar] [CrossRef]
- Van Helvoort, A.; Giudici, M.L.; Thielemans, M.; Meer, G. van Transport of sphingomyelin to the cell surface is inhibited by brefeldin A and in mitosis, where C6-NBD-sphingomyelin is translocated across the plasma membrane by a multidrug transporter activity. J. Cell Sci. 1997, 110, 75–83. [Google Scholar]
- Kornhuber, J.; Rhein, C.; Müller, C.P.; Mühle, C. Secretory sphingomyelinase in health and disease. Biol. Chem. 2015, 396, 707–736. [Google Scholar]
- Yang, F.; Chen, G. The nutritional functions of dietary sphingomyelin and its applications in food. Front. Nutr. 2022, 9, 1002574. [Google Scholar]
- Chakraborty, M.; Jiang, X.-C. Sphingomyelin and Its Role in Cellular Signaling. In Lipid-Mediated Protein Signaling: Advances in Experimental Medicine and Biology; Capelluto, D., Ed.; Springer: Dordrecht, The Netherlands, 2013; pp. 1–14. [Google Scholar]
- Slotte, J.P. Biological functions of sphingomyelins. Prog. Lipid Res. 2013, 52, 424–437. [Google Scholar] [PubMed]
- Hannun, Y.A.; Obeid, L.M. Principles of bioactive lipid signalling: Lessons from sphingolipids. Nat. Rev. Mol. Cell Biol. 2008, 9, 139–150. [Google Scholar] [PubMed]
- Hannun, Y.A.; Bell, R.M. Regulation of protein kinase C by sphingosine and lysosphingolipids. Clin. Chim. Acta 1989, 185, 333–345. [Google Scholar]
- Subbaiah, P.V.; Gesquiere, L.R.; Wang, K. Regulation of the selective uptake of cholesteryl esters from high density lipoproteins by sphingomyelin. J. Lipid Res. 2005, 46, 2699–2705. [Google Scholar] [PubMed]
- Dingjan, T.; Futerman, A.H. The role of the ‘sphingoid motif’ in shaping the molecular interactions of sphingolipids in biomembranes. Biochim. Biophys. Acta Biomembr. 2021, 1863, 183701. [Google Scholar] [CrossRef]
- García-Arribas, A.B.; Alonso, A.; Goñi, F.M. Cholesterol interactions with ceramide and sphingomyelin. Chem. Phys. Lipids 2016, 199, 26–34. [Google Scholar] [PubMed]
- Lingwood, D.; Simons, K. Lipid rafts as a membrane-organizing principle. Science 2010, 327, 46–50. [Google Scholar] [PubMed]
- Prinetti, A.; Chigorno, V.; Prioni, S.; Loberto, N.; Marano, N.; Tettamanti, G.; Sonnino, S. Changes in the lipid turnover, composition, and organization, as sphingolipid-enriched membrane domains, in rat cerebellar granule cells developing in vitro. J. Biol. Chem. 2001, 276, 21136–21145. [Google Scholar]
- Futerman, A.H.; Pagano, R.E. Determination of the intracellular sites and topology of glucosylceramide synthesis in rat liver. Biochem. J. 1991, 280, 295–302. [Google Scholar] [CrossRef]
- Carruthers, A.; Carey, E.M. UDP-galactose: Ceramide galactosyl transferase of isolated oligodendroglia. J. Neurochem. 1983, 41, 22–29. [Google Scholar] [CrossRef]
- Yu, R.K.; Yanagisawa, M.; Ariga, T. Glycosphingolipid structures. In Comprehensive Glycoscience: Chemistry to Systems Biology; Kamerling, H., Ed.; Elsevier: Amsterdam, The Netherlands, 2007; pp. 73–122. ISBN 13: 978-0-444-52746-2. [Google Scholar]
- Merrill, A.H.; Sandhoff, K. Chapter 14 Sphingolipids: Metabolism and cell signaling. In Biochemistry of Lipids, Lipoproteins and Membranes; Vance, D.E., Vance, J.E., Eds.; Elsevier: Amsterdam, The Netherlands, 2002; pp. 373–407. [Google Scholar]
- Leray, C. Introduction to Lipidomics: From Human to Bacteria; CRC Press, Taylor and Francis Group: Boca Raton, FL, USA, 2013. [Google Scholar]
- Farwanah, H.; Kolter, T. Lipidomics of glycosphingolipids. Metabolites 2012, 2, 134–164. [Google Scholar] [CrossRef] [PubMed]
- Tettamanti, G. Ganglioside/glycosphingolipid turnover: New concepts. Glycoconj. J. 2003, 20, 301–317. [Google Scholar] [CrossRef] [PubMed]
- Lingwood, C.A. Glycosphingolipid functions. Cold Spring Harb. Perspect. Biol. 2011, 3, a004788. [Google Scholar] [CrossRef] [PubMed]
- He, Q.; Chen, Y.; Wang, Z.; He, H.; Yu, P. Cellular uptake, metabolism and sensing of long-chain fatty acids. Front. Biosci. 2023, 28, 10. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Rodriguez, E.; Bi, C.; Zheng, X.; Suresh, D.; Suh, K.; Li, Z.; Elsebaei, F.; Hage, D.S. High performance affinity chromatography and related separation methods for the analysis of biological and pharmaceutical agents. Analyst 2018, 143, 374–391. [Google Scholar] [CrossRef]
- Rodriguez-Cuenca, S.; Pellegrinelli, V.; Campbell, M.; Oresic, M.; Vidal-Puig, A. Sphingolipids and glycerophospholipids—The “ying and yang” of lipotoxicity in metabolic diseases. Prog. Lipid Res. 2017, 66, 14–29. [Google Scholar] [CrossRef]
- Hakomori, S. Structure and function of glycosphingolipids and sphingolipids: Recollections and future trends. Biochim. Biophys. Acta Gen. Subj. 2008, 1780, 325–346. [Google Scholar] [CrossRef]
- Shaikh, S.R.; Edidin, M. Polyunsaturated fatty acids, membrane organization, T cells, and antigen presentation. Am. J. Clin. Nutr. 2006, 84, 1277–1289. [Google Scholar] [CrossRef]
- Sud, M.; Fahy, E.; Cotter, D.; Brown, A.; Dennis, E.A.; Glass, C.K.; Merrill, A.H.; Murphy, R.C.; Raetz, C.R.H.; Russell, D.W.; et al. LMSD: LIPID MAPS structure database. Nucleic Acids Res. 2007, 35, D527–D532. [Google Scholar] [CrossRef]
- Jalil, A.; Bourgeois, T.; Ménégaut, L.; Lagrost, L.; Thomas, C.; Masson, D. Revisiting the role of LXRs in PUFA metabolism and phospholipid homeostasis. Int. J. Mol. Sci. 2019, 20, 3787. [Google Scholar] [CrossRef]
- Repa, J.J.; Liang, G.; Ou, J.; Bashmakov, Y.; Lobaccaro, J.-M.A.; Shimomura, I.; Shan, B.; Brown, M.S.; Goldstein, J.L.; Mangelsdorf, D.J. Regulation of mouse sterol regulatory element-binding protein-1c gene (SREBP-1c) by oxysterol receptors, LXRα and LXRβ. Genes Dev. 2000, 14, 2819–2830. [Google Scholar] [CrossRef]
- Seo, J.B.; Moon, H.M.; Kim, W.S.; Lee, Y.S.; Jeong, H.W.; Yoo, E.J.; Ham, J.; Kang, H.; Park, M.-G.; Steffensen, K.R.; et al. Activated liver X receptors stimulate adipocyte differentiation through induction of peroxisome proliferator-activated receptor γ expression. Mol. Cell. Biol. 2004, 24, 3430–3444. [Google Scholar] [CrossRef] [PubMed]
- Cha, J.-Y.; Repa, J.J. The liver X receptor (LXR) and hepatic lipogenesis. J. Biol. Chem. 2007, 282, 743–751. [Google Scholar] [CrossRef] [PubMed]
- Schörken, U.; Kempers, P. Lipid biotechnology: Industrially relevant production processes. Eur. J. Lipid Sci. Technol. 2009, 111, 627–645. [Google Scholar]
- Shanklin, J.; Cahoon, E.B. Desaturation and related modifications of fatty acids. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1998, 49, 611–641. [Google Scholar] [CrossRef]
- Jayakumar, A.; Tai, M.H.; Huang, W.Y.; Al-Feel, W.; Hsu, M.; Abu-Elheiga, L.; Chirala, S.S.; Wakil, S.J. Human fatty acid synthase: Properties and molecular cloning. Proc. Natl. Acad. Sci. USA 1995, 92, 8695–8699. [Google Scholar] [CrossRef]
- Semenkovich, C.F.; Coleman, T.; Fiedorek, F.T. Human fatty acid synthase mRNA: Tissue distribution, genetic mapping, and kinetics of decay after glucose deprivation. J. Lipid Res. 1995, 36, 1507–1521. [Google Scholar]
- Nowinski, S.M.; Van Vranken, J.G.; Dove, K.K.; Rutter, J. Impact of mitochondrial fatty acid synthesis on mitochondrial biogenesis. Curr. Biol. 2018, 28, R1212–R1219. [Google Scholar] [CrossRef]
- Strawford, A.; Antelo, F.; Christiansen, M.; Hellerstein, M.K. Adipose tissue triglyceride turnover, de novo lipogenesis, and cell proliferation in humans measured with 2H2O. Am. J. Physiol. Metab. 2004, 286, E577–E588. [Google Scholar]
- Turner, S.M.; Murphy, E.J.; Neese, R.A.; Antelo, F.; Thomas, T.; Agarwal, A.; Go, C.; Hellerstein, M.K. Measurement of TG synthesis and turnover in vivo by 2 H 2 O incorporation into the glycerol moiety and application of MIDA. Am. J. Physiol. Metab. 2003, 285, E790–E803. [Google Scholar] [CrossRef]
- Bauman, D.E.; Mellenberger, R.W.; Derrig, R.G. Fatty acid synthesis in sheep mammary tissue. J. Dairy Sci. 1973, 56, 1312–1318. [Google Scholar] [PubMed]
- Chandel, N.S. Glycolysis. Cold Spring Harb. Perspect. Biol. 2021, 13, a040535. [Google Scholar] [CrossRef] [PubMed]
- McCommis, K.S.; Finck, B.N. Mitochondrial pyruvate transport: A historical perspective and future research directions. Biochem. J. 2015, 466, 443–454. [Google Scholar] [PubMed]
- Martínez-Reyes, I.; Chandel, N.S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 2020, 11, 102. [Google Scholar]
- Wallace, M.; Green, C.R.; Roberts, L.S.; Lee, Y.M.; McCarville, J.L.; Sanchez-Gurmaches, J.; Meurs, N.; Gengatharan, J.M.; Hover, J.D.; Phillips, S.A.; et al. Enzyme promiscuity drives branched-chain fatty acid synthesis in adipose tissues. Nat. Chem. Biol. 2018, 14, 1021–1031. [Google Scholar]
- Bressler, R.; Wakil, S.J. Studies on the mechanism of fatty acid synthesis. J. Biol. Chem. 1962, 237, 1441–1448. [Google Scholar]
- Kuhajda, F.P. Fatty-acid synthase and human cancer: New perspectives on its role in tumor biology. Nutrition 2000, 16, 202–208. [Google Scholar]
- Wakil, S.J. A malonic acid derivative as an intermediate in fatty acid synthesis. J. Am. Chem. Soc. 1958, 80, 6465. [Google Scholar] [CrossRef]
- Wakil, S.J.; Titchener, E.B.; Gibson, D.M. Evidence for the participation of biotin in the enzymic synthesis of fatty acids. Biochim. Biophys. Acta 1958, 29, 225–226. [Google Scholar] [CrossRef]
- O’Neill, L.M.; Miyazaki, M.; Bond, L.M.; Lewis, S.A.; Ding, F.; Liu, Z.; Ntambi, J.M. Fatty acid desaturation and elongation in mammals. In Biochemistry of Lipids, Lipoproteins and Membranes; Ridgway, N.D., McLeod, R.S., Eds.; Elsevier: Amsterdam, The Netherlands, 2021; pp. 201–226. [Google Scholar]
- Wakil, S.J. Fatty acid synthase, a proficient multifunctional enzyme. Biochemistry 1989, 28, 4523–4530. [Google Scholar] [CrossRef]
- Cook, H.W. Fatty acid desaturation and chain elongation in eucaryotes. In Biochemistry of Lipids and Membranes; Vance, D.E., Vance, J.E., Eds.; Elsevier Science: Amsterdam, The Netherlands, 1985; pp. 181–212. [Google Scholar]
- Heil, C.S.; Wehrheim, S.S.; Paithankar, K.S.; Grininger, M. Fatty acid biosynthesis: Chain-length regulation and control. ChemBioChem 2019, 20, 2298–2321. [Google Scholar] [PubMed]
- Kim, K.-H. Regulation of mammalian acetyl-coenzyme A carboxylase. Annu. Rev. Nutr. 1997, 17, 77–99. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yu, W.; Li, S.; Guo, D.; He, J.; Wang, Y. Acetyl-CoA carboxylases and diseases. Front. Oncol. 2022, 12, 836058. [Google Scholar] [PubMed]
- Paiva, P.; Medina, F.E.; Viegas, M.; Ferreira, P.; Neves, R.P.P.; Sousa, J.P.M.; Ramos, M.J.; Fernandes, P.A. Animal fatty acid synthase: A chemical nanofactory. Chem. Rev. 2021, 121, 9502–9553. [Google Scholar]
- Leibundgut, M.; Maier, T.; Jenni, S.; Ban, N. The multienzyme architecture of eukaryotic fatty acid synthases. Curr. Opin. Struct. Biol. 2008, 18, 714–725. [Google Scholar] [CrossRef]
- Smith, S. The animal fatty acid synthase: One gene, one polypeptide, seven enzymes. FASEB J. 1994, 8, 1248–1259. [Google Scholar] [CrossRef]
- Magnuson, K.; Jackowski, S.; Rock, C.O.; Cronan, J.E. Regulation of fatty acid biosynthesis in Escherichia coli. Microbiol. Rev. 1993, 57, 522–542. [Google Scholar] [CrossRef]
- Bazan, H.E.P.; Careaga, M.M.; Sprecher, H.; Bazan, N.G. Chain elongation and desaturation of eicosapentaenoate to docosahexaenoate and phospholipid labeling in the rat retina in vivo. Biochim. Biophys. Acta Lipids Lipid Metab. 1982, 712, 123–128. [Google Scholar] [CrossRef]
- Cinti, D.L.; Cook, L.; Nagi, M.N.; Suneja, S.K. The fatty acid chain elongation system of mammalian endoplasmic reticulum. Prog. Lipid Res. 1992, 31, 1–51. [Google Scholar]
- Jump, D.B. Mammalian fatty acid elongases. In Lipidomics: Methods in Molecular Biology; Armstrong, D., Ed.; Humana Press: Totowa, NJ, USA, 2009; pp. 375–389. [Google Scholar]
- Oboh, A. Investigating the Long-Chain Polyunsaturated Fatty Acid Biosynthesis of the African Catfish Clarias gariepinus (Burchell, 1822). Ph.D. Thesis, University of Stirling, Stirling, UK, 2018. [Google Scholar]
- Yeboah, G.K.; Lobanova, E.S.; Brush, R.S.; Agbaga, M.-P. Very long chain fatty acid-containing lipids: A decade of novel insights from the study of ELOVL4. J. Lipid Res. 2021, 62, 100030. [Google Scholar]
- Robinson, B.S.; Johnson, D.W.; Poulos, A. Unique molecular species of phosphatidylcholine containing very-long-chain (C24-C38) polyenoic fatty acids in rat brain. Biochem. J. 1990, 265, 763–767. [Google Scholar] [CrossRef]
- Řezanka, T. Very-long-chain fatty acids from the animal and plant kingdoms. Prog. Lipid Res. 1989, 28, 147–187. [Google Scholar] [CrossRef]
- Poulos, A.; Sharp, P.; Johnson, D.; Easton, C. The occurrence of polyenoic very long chain fatty acids with greater than 32 carbon atoms in molecular species of phosphatidylcholine in normal and peroxisome-deficient (Zellweger’s syndrome) brain. Biochem. J. 1988, 253, 645–650. [Google Scholar] [CrossRef]
- Aveldaño, M.I.; Sprecher, H. Very long chain (C24 to C36) polyenoic fatty acids of the n-3 and n-6 series in dipolyunsaturated phosphatidylcholines from bovine retina. J. Biol. Chem. 1987, 262, 1180–1186. [Google Scholar] [CrossRef] [PubMed]
- Furland, N.E.; Maldonado, E.N.; Aveldaño, M.I. Very long chain PUFA in murine testicular triglycerides and cholesterol esters. Lipids 2003, 38, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Aveldaño, M.I.; Robinson, B.S.; Johnson, D.W.; Poulos, A. Long and very long chain polyunsaturated fatty acids of the n-6 series in rat seminiferous tubules. Active desaturation of 24:4n-6 to 24:5n-6 and concomitant formation of odd and even chain tetraenoic and pentaenoic fatty acids up to C32. J. Biol. Chem. 1993, 268, 11663–11669. [Google Scholar] [CrossRef] [PubMed]
- Furland, N.E.; Zanetti, S.R.; Oresti, G.M.; Maldonado, E.N.; Aveldaño, M.I. Ceramides and sphingomyelins with high proportions of very long-chain polyunsaturated fatty acids in mammalian germ cells. J. Biol. Chem. 2007, 282, 18141–18150. [Google Scholar] [CrossRef]
- Torrissen, M.; Ytteborg, E.; Svensen, H.; Stoknes, I.; Nilsson, A.; Østbye, T.-K.; Berge, G.M.; Bou, M.; Ruyter, B. Investigation of the functions of n-3 very-long-chain PUFAs in skin using in vivo Atlantic salmon and in vitro human and fish skin models. Br. J. Nutr. 2023, 1–17. [Google Scholar] [CrossRef]
- Butovich, I.A. Cholesteryl esters as a depot for very long chain fatty acids in human meibum. J. Lipid Res. 2009, 50, 501–513. [Google Scholar] [CrossRef]
- Butovich, I.A.; Uchiyama, E.; McCulley, J.P. Lipids of human meibum: Mass-spectrometric analysis and structural elucidation. J. Lipid Res. 2007, 48, 2220–2235. [Google Scholar] [CrossRef]
- Rissmann, R.; Groenink, H.W.W.; Weerheim, A.M.; Hoath, S.B.; Ponec, M.; Bouwstra, J.A. New insights into ultrastructure, lipid composition and organization of Vernix Caseosa. J. Investig. Dermatol. 2006, 126, 1823–1833. [Google Scholar] [CrossRef] [PubMed]
- Hiltunen, J.K.; Schonauer, M.S.; Autio, K.J.; Mittelmeier, T.M.; Kastaniotis, A.J.; Dieckmann, C.L. Mitochondrial fatty acid synthesis type II: More than just fatty acids. J. Biol. Chem. 2009, 284, 9011–9015. [Google Scholar] [CrossRef] [PubMed]
- Chuman, L.; Brody, S. Acyl carrier protein is present in the mitochondria of plants and eucaryotic micro-organisms. Eur. J. Biochem. 1989, 184, 643–649. [Google Scholar] [CrossRef] [PubMed]
- Monteuuis, G.; Suomi, F.; Kerätär, J.M.; Masud, A.J.; Kastaniotis, A.J. A conserved mammalian mitochondrial isoform of acetyl-CoA carboxylase ACC1 provides the malonyl-CoA essential for mitochondrial biogenesis in tandem with ACSF3. Biochem. J. 2017, 474, 3783–3797. [Google Scholar] [CrossRef]
- Wongkittichote, P.; Ah Mew, N.; Chapman, K.A. Propionyl-CoA carboxylase—A review. Mol. Genet. Metab. 2017, 122, 145–152. [Google Scholar] [CrossRef]
- Seubert, W.; Podack, E.R. Mechanisms and physiological roles of fatty acid chain elongation in microsomes and mitochondria. Mol. Cell. Biochem. 1973, 1, 29–40. [Google Scholar] [CrossRef]
- Lu, Y.-J.; Zhang, Y.-M.; Grimes, K.D.; Qi, J.; Lee, R.E.; Rock, C.O. Acyl-phosphates initiate membrane phospholipid synthesis in gram-positive pathogens. Mol. Cell 2006, 23, 765–772. [Google Scholar] [CrossRef]
- Rezaei Zonooz, S.; Hasani, M.; Morvaridzadeh, M.; Beatriz Pizarro, A.; Heydari, H.; Yosaee, S.; Rezamand, G.; Heshmati, J. Effect of alpha-lipoic acid on oxidative stress parameters: A systematic review and meta-analysis. J. Funct. Foods 2021, 87, 104774. [Google Scholar] [CrossRef]
- Petersen Shay, K.; Moreau, R.F.; Smith, E.J.; Hagen, T.M. Is α-lipoic acid a scavenger of reactive oxygen speciesin vivo? Evidence for its initiation of stress signaling pathways that promote endogenous antioxidant capacity. IUBMB Life 2008, 60, 362–367. [Google Scholar] [CrossRef]
- Stoll, S.; Hartmann, H.; Cohen, S.A.; Müller, W.E. The potent free radical scavenger α-lipoic acid improves memory in aged mice: Putative relationship to NMDA receptor deficits. Pharmacol. Biochem. Behav. 1993, 46, 799–805. [Google Scholar] [CrossRef]
- Capece, U.; Moffa, S.; Improta, I.; Di Giuseppe, G.; Nista, E.C.; Cefalo, C.M.A.; Cinti, F.; Pontecorvi, A.; Gasbarrini, A.; Giaccari, A.; et al. Alpha-lipoic acid and glucose metabolism: A comprehensive update on biochemical and therapeutic features. Nutrients 2022, 15, 18. [Google Scholar] [CrossRef]
- Hiltunen, J.K.; Autio, K.J.; Schonauer, M.S.; Kursu, V.A.S.; Dieckmann, C.L.; Kastaniotis, A.J. Mitochondrial fatty acid synthesis and respiration. Biochim. Biophys. Acta Bioenerg. 2010, 1797, 1195–1202. [Google Scholar] [CrossRef] [PubMed]
- Angerer, H.; Schönborn, S.; Gorka, J.; Bahr, U.; Karas, M.; Wittig, I.; Heidler, J.; Hoffmann, J.; Morgner, N.; Zickermann, V. Acyl modification and binding of mitochondrial ACP to multiprotein complexes. Biochim. Biophys. Acta Mol. Cell Res. 2017, 1864, 1913–1920. [Google Scholar] [CrossRef] [PubMed]
- Das, A.K.; Uhler, M.D.; Hajra, A.K. Molecular cloning and expression of mammalian peroxisomaltrans-2-enoyl-coenzyme A reductase cDNAs. J. Biol. Chem. 2000, 275, 24333–24340. [Google Scholar] [CrossRef] [PubMed]
- Buist, P.H. Fatty acid desaturases: Selecting the dehydrogenation channel. Nat. Prod. Rep. 2004, 21, 249. [Google Scholar] [CrossRef]
- Smith, S.; Tsai, S.-C. The type I fatty acid and polyketide synthases: A tale of two megasynthases. Nat. Prod. Rep. 2007, 24, 1041. [Google Scholar] [CrossRef]
- Kaulmann, U.; Hertweck, C. Biosynthesis of polyunsaturated fatty acids by polyketide synthases. Angew. Chem. Int. Ed. 2002, 41, 1866. [Google Scholar] [CrossRef]
- Napier, J. Plumbing the depths of PUFA biosynthesis: A novel polyketide synthase-like pathway from marine organisms. Trends Plant Sci. 2002, 7, 51–54. [Google Scholar] [CrossRef]
- Metz, J.G.; Roessler, P.; Facciotti, D.; Levering, C.; Dittrich, F.; Lassner, M.; Valentine, R.; Lardizabal, K.; Domergue, F.; Yamada, A.; et al. Production of polyunsaturated fatty acids by polyketide synthases in both prokaryotes and eukaryotes. Science 2001, 293, 290–293. [Google Scholar] [CrossRef]
- Chen, Z.; Chen, H.; Li, X.; Yuan, Q.; Su, J.; Yang, L.; Ning, L.; Lei, H. Fumonisin B1 damages the barrier functions of porcine intestinal epithelial cells in vitro. J. Biochem. Mol. Toxicol. 2019, 33, e22397. [Google Scholar] [CrossRef]
- Bentley, R.; Bennett, J.W. Constructing polyketides: From collie to combinatorial biosynthesis. Annu. Rev. Microbiol. 1999, 53, 411–446. [Google Scholar] [CrossRef] [PubMed]
- Cerone, M.; Smith, T.K. Desaturases: Structural and mechanistic insights into the biosynthesis of unsaturated fatty acids. IUBMB Life 2022, 74, 1036–1051. [Google Scholar] [CrossRef] [PubMed]
- Gostinčar, C.; Turk, M.; Gunde-Cimerman, N. The evolution of fatty acid desaturases and cytochrome b5 in eukaryotes. J. Membr. Biol. 2010, 233, 63–72. [Google Scholar] [CrossRef] [PubMed]
- Sprecher, H.; Luthria, D.L.; Mohammed, B.S.; Baykousheva, S.P. Reevaluation of the pathways for the biosynthesis of polyunsaturated fatty acids. J. Lipid Res. 1995, 36, 2471–2477. [Google Scholar] [CrossRef]
- Paton, C.M.; Ntambi, J.M. Biochemical and physiological function of stearoyl-CoA desaturase. Am. J. Physiol. Metab. 2009, 297, E28–E37. [Google Scholar] [CrossRef]
- Nakamura, M.T.; Nara, T.Y. Structure, function, and dietary regulation of delta6, delta5, and delta9 desaturases. Annu. Rev. Nutr. 2004, 24, 345–376. [Google Scholar] [CrossRef]
- Meesapyodsuk, D.; Qiu, X. Structure Determinants for the Substrate Specificity of Acyl-CoA Δ9 Desaturases from a Marine Copepod. ACS Chem. Biol. 2014, 9, 922–934. [Google Scholar] [CrossRef]
- Haritos, V.S.; Horne, I.; Damcevski, K.; Glover, K.; Gibb, N. Unexpected functional diversity in the fatty acid desaturases of the flour beetle Tribolium castaneum and identification of key residues determining activity. Insect Biochem. Mol. Biol. 2014, 51, 62–70. [Google Scholar] [CrossRef]
- Bonamore, A.; Macone, A.; Colotti, G.; Matarese, R.M.; Boffi, A. The desaturase from Bacillus subtilis, a promising tool for the selective olefination of phospholipids. J. Biotechnol. 2006, 121, 49–53. [Google Scholar] [CrossRef]
- Heilmann, I.; Mekhedov, S.; King, B.; Browse, J.; Shanklin, J. Identification of the Arabidopsis Palmitoyl-Monogalactosyldiacylglycerol Δ7-Desaturase Gene FAD5, and Effects of Plastidial Retargeting of Arabidopsis Desaturases on the fad5 Mutant Phenotype. Plant Physiol. 2004, 136, 4237–4245. [Google Scholar] [CrossRef]
- Knipple, D.C.; Rosenfield, C.-L.; Miller, S.J.; Liu, W.; Tang, J.; Ma, P.W.K.; Roelofs, W.L. Cloning and functional expression of a cDNA encoding a pheromone gland-specific acyl-CoA Δ11-desaturase of the cabbage looper moth, Trichoplusia ni. Proc. Natl. Acad. Sci. USA 1998, 95, 15287–15292. [Google Scholar] [CrossRef] [PubMed]
- Ohnishi, M.; Thompson, G.A. Biosynthesis of the unique trans-Δ3-hexadecenoic acid component of chloroplast phosphatidylglycerol: Evidence concerning its site and mechanism of formation. Arch. Biochem. Biophys. 1991, 288, 591–599. [Google Scholar] [CrossRef] [PubMed]
- Weiss-Hersh, K.; Garcia, A.L.; Marosvölgyi, T.; Szklenár, M.; Decsi, T.; Rühl, R. Saturated and monounsaturated fatty acids in membranes are determined by the gene expression of their metabolizing enzymes SCD1 and ELOVL6 regulated by the intake of dietary fat. Eur. J. Nutr. 2020, 59, 2759–2769. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, M.; Jacobson, M.J.; Man, W.C.; Cohen, P.; Asilmaz, E.; Friedman, J.M.; Ntambi, J.M. Identification and characterization of murine SCD4, a novel heart-specific stearoyl-CoA desaturase isoform regulated by leptin and dietary factors. J. Biol. Chem. 2003, 278, 33904–33911. [Google Scholar] [CrossRef]
- Human Gene Nomenclature Containing (HGNC) Gene Group: Fatty Acid Desaturases (FADS). Available online: https://www.genenames.org/data/genegroup/#!/group/553 (accessed on 15 September 2023).
- Nagao, K.; Murakami, A.; Umeda, M. Structure and Function of Δ9-Fatty Acid Desaturase. Chem. Pharm. Bull. 2019, 67, 327–332. [Google Scholar] [CrossRef]
- Grajchen, E.; Loix, M.; Baeten, P.; Côrte-Real, B.F.; Hamad, I.; Vanherle, S.; Haidar, M.; Dehairs, J.; Broos, J.Y.; Ntambi, J.M.; et al. Fatty acid desaturation by stearoyl-CoA desaturase-1 controls regulatory T cell differentiation and autoimmunity. Cell. Mol. Immunol. 2023, 20, 666–679. [Google Scholar] [CrossRef]
- Tang, B.; Qiu, J.; Hu, S.; Li, L.; Wang, J. Role of stearyl-coenzyme A desaturase 1 in mediating the effects of palmitic acid on endoplasmic reticulum stress, inflammation, and apoptosis in goose primary hepatocytes. Anim. Biosci. 2021, 34, 1210–1220. [Google Scholar] [CrossRef]
- Koeberle, A.; Löser, K.; Thürmer, M. Stearoyl-CoA desaturase-1 and adaptive stress signaling. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2016, 1861, 1719–1726. [Google Scholar] [CrossRef]
- Kucharski, M.; Kaczor, U. Stearoyl-CoA desaturase—The lipid metabolism regulator. Adv. Hyg. Exp. Med. 2014, 68, 334–342. [Google Scholar] [CrossRef]
- Liu, X.; Strable, M.S.; Ntambi, J.M. Stearoyl CoA Desaturase 1: Role in Cellular Inflammation and Stress. Adv. Nutr. 2011, 2, 15–22. [Google Scholar] [CrossRef]
- Dobrzyn, P.; Dobrzyn, A.; Miyazaki, M.; Cohen, P.; Asilmaz, E.; Hardie, D.G.; Friedman, J.M.; Ntambi, J.M. Stearoyl-CoA desaturase 1 deficiency increases fatty acid oxidation by activating AMP-activated protein kinase in liver. Proc. Natl. Acad. Sci. USA 2004, 101, 6409–6414. [Google Scholar] [CrossRef] [PubMed]
- Park, W.J.; Kothapalli, K.S.D.; Reardon, H.T.; Lawrence, P.; Qian, S.-B.; Brenna, J.T. A novel FADS1 isoform potentiates FADS2-mediated production of eicosanoid precursor fatty acids. J. Lipid Res. 2012, 53, 1502–1512. [Google Scholar] [CrossRef] [PubMed]
- Park, W.J.; Reardon, H.T.; Tyburczy, C.; Kothapalli, K.S.D.; Brenna, J.T. Alternative splicing generates a novel FADS2 alternative transcript in baboons. Mol. Biol. Rep. 2010, 37, 2403–2406. [Google Scholar] [CrossRef] [PubMed]
- Park, W.J.; Kothapalli, K.S.D.; Reardon, H.T.; Kim, L.Y.; Brenna, J.T. Novel fatty acid desaturase 3 (FADS3) transcripts generated by alternative splicing. Gene 2009, 446, 28–34. [Google Scholar] [CrossRef] [PubMed]
- Srikanth, K.; Kwan, A.; Lee, E.; Kim, S.; Lim, Y.; Chung, H. Associations of single nucleotide polymorphisms in the bovine FADS6 gene with fatty acid composition in Hanwoo (Korean Cattle). Open J. Genet. 2015, 05, 137–144. [Google Scholar] [CrossRef]
- Chen, H.; Hao, G.; Wang, L.; Wang, H.; Gu, Z.; Liu, L.; Zhang, H.; Chen, W.; Chen, Y.Q. Identification of a critical determinant that enables efficient fatty acid synthesis in oleaginous fungi. Sci. Rep. 2015, 5, 11247. [Google Scholar] [CrossRef]
- Stroud, C.K.; Nara, T.Y.; Roqueta-Rivera, M.; Radlowski, E.C.; Lawrence, P.; Zhang, Y.; Cho, B.H.; Segre, M.; Hess, R.A.; Brenna, J.T.; et al. Disruption of FADS2 gene in mice impairs male reproduction and causes dermal and intestinal ulceration. J. Lipid Res. 2009, 50, 1870–1880. [Google Scholar] [CrossRef]
- Zhu, K.-C.; Song, L.; Guo, H.-Y.; Guo, L.; Zhang, N.; Liu, B.-S.; Jiang, S.-G.; Zhang, D.-C. Identification of fatty acid desaturase 6 in Golden Pompano Trachinotus Ovatus (Linnaeus 1758) and its regulation by the PPARαb transcription factor. Int. J. Mol. Sci. 2018, 20, 23. [Google Scholar] [CrossRef]
- Marquardt, A.; Stöhr, H.; White, K.; Weber, B.H.F. cDNA cloning, genomic structure, and chromosomal localization of three members of the human fatty acid desaturase family. Genomics 2000, 66, 175–183. [Google Scholar] [CrossRef]
- Rioux, V.; Pédrono, F.; Blanchard, H.; Duby, C.; Boulier-Monthéan, N.; Bernard, L.; Beauchamp, E.; Catheline, D.; Legrand, P. Trans-vaccenate is Δ13-desaturated by FADS3 in rodents. J. Lipid Res. 2013, 54, 3438–3452. [Google Scholar] [CrossRef]
- Zhang, J.Y.; Qin, X.; Liang, A.; Kim, E.; Lawrence, P.; Park, W.J.; Kothapalli, K.S.D.; Brenna, J.T. Fads3 modulates docosahexaenoic acid in liver and brain. Prostaglandins Leukot. Essent. Fat. Acids 2017, 123, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Karsai, G.; Lone, M.; Kutalik, Z.; Brenna, J.T.; Li, H.; Pan, D.; von Eckardstein, A.; Hornemann, T. FADS3 is a Δ14Z sphingoid base desaturase that contributes to gender differences in the human plasma sphingolipidome. J. Biol. Chem. 2020, 295, 1889–1897. [Google Scholar] [CrossRef] [PubMed]
- Castro, L.F.C.; Tocher, D.R.; Monroig, O. Long-chain polyunsaturated fatty acid biosynthesis in chordates: Insights into the evolution of Fads and Elovl gene repertoire. Prog. Lipid Res. 2016, 62, 25–40. [Google Scholar] [CrossRef]
- Lee, J.; Lee, H.; Kang, S.; Park, W. Fatty acid desaturases, polyunsaturated fatty acid regulation, and biotechnological advances. Nutrients 2016, 8, 23. [Google Scholar] [CrossRef] [PubMed]
- Guillou, H.; Zadravec, D.; Martin, P.G.P.; Jacobsson, A. The key roles of elongases and desaturases in mammalian fatty acid metabolism: Insights from transgenic mice. Prog. Lipid Res. 2010, 49, 186–199. [Google Scholar] [CrossRef] [PubMed]
- Spector, A.A.; Kim, H.-Y. Discovery of essential fatty acids. J. Lipid Res. 2015, 56, 11–21. [Google Scholar] [CrossRef]
- Kaur, N.; Chugh, V.; Gupta, A.K. Essential fatty acids as functional components of foods—A review. J. Food Sci. Technol. 2014, 51, 2289–2303. [Google Scholar] [CrossRef]
- Burr, G.O.; Burr, M.M. On the nature and role of the fatty acids essential in nutrition. J. Biol. Chem. 1930, 86, 587–621. [Google Scholar] [CrossRef]
- Cho, H.P.; Nakamura, M.; Clarke, S.D. Cloning, expression, and fatty acid regulation of the human Δ-5 desaturase. J. Biol. Chem. 1999, 274, 37335–37339. [Google Scholar] [CrossRef]
- Cho, H.P.; Nakamura, M.T.; Clarke, S.D. Cloning, expression, and nutritional regulation of the mammalian Δ-6 desaturase. J. Biol. Chem. 1999, 274, 471–477. [Google Scholar] [CrossRef]
- Scott, B.L.; Bazan, N.G. Membrane docosahexaenoate is supplied to the developing brain and retina by the liver. Proc. Natl. Acad. Sci. USA 1989, 86, 2903–2907. [Google Scholar] [CrossRef] [PubMed]
- De Antueno, R.J.; Knickle, L.C.; Smith, H.; Elliot, M.L.; Allen, S.J.; Nwaka, S.; Winther, M.D. Activity of human Δ5 and Δ6 desaturases on multiple n-3 and n-6 polyunsaturated fatty acids. FEBS Lett. 2001, 509, 77–80. [Google Scholar] [CrossRef] [PubMed]
- Vagner, M.; Santigosa, E. Characterization and modulation of gene expression and enzymatic activity of delta-6 desaturase in teleosts: A review. Aquaculture 2011, 315, 131–143. [Google Scholar] [CrossRef]
- Arshad, Z.; Rezapour-Firouzi, S.; Ebrahimifar, M.; Mosavi-Jarrahi, A.; Mohammadian, M. Association of delta-6-desaturase expression with aggressiveness of cancer, diabetes mellitus, and multiple sclerosis: A narrative review. Asian Pac. J. Cancer Prev. 2019, 20, 1005–1018. [Google Scholar] [CrossRef]
- Tosi, F.; Sartori, F.; Guarini, P.; Olivieri, O.; Martinelli, N. Delta-5 and delta-6 desaturases: Crucial enzymes in polyunsaturated fatty acid-related pathways with pleiotropic influences in health and disease. In Oxidative Stress and Inflammation in Non-Communicable Diseases—Molecular Mechanisms and Perspectives in Therapeutics: Advances in Experimental Medicine and Biology; Camps, J., Ed.; Springer: Cham, Switzerland, 2014; pp. 61–81. [Google Scholar]
- Mead, J.F.; Slaton, W.H. Metabolism of essential fatty acids. J. Biol. Chem. 1956, 219, 705–709. [Google Scholar] [CrossRef]
- Siguel, E.N.; Chee, K.M.; Gong, J.X.; Schaefer, E.J. Criteria for essential fatty acid deficiency in plasma as assessed by capillary column gas-liquid chromatography. Clin. Chem. 1987, 33, 1869–1873. [Google Scholar] [CrossRef]
- Holman, R.T. The ratio of trienoic: Tetraenoic acids in tissue lipids as a measure of essential fatty acid requirement. J. Nutr. 1960, 70, 405–410. [Google Scholar] [CrossRef]
- Park, H.G.; Engel, M.G.; Vogt-Lowell, K.; Lawrence, P.; Kothapalli, K.S.; Brenna, J.T. The role of fatty acid desaturase (FADS) genes in oleic acid metabolism: FADS1 Δ7 desaturates 11-20:1 to 7,11-20:2. Prostaglandins Leukot. Essent. Fat. Acids 2018, 128, 21–25. [Google Scholar] [CrossRef]
- Park, H.G.; Park, W.J.; Kothapalli, K.S.D.; Brenna, J.T. The fatty acid desaturase 2 (FADS2) gene product catalyzes Δ4 desaturation to yield n-3 docosahexaenoic acid and n-6 docosapentaenoic acid in human cells. FASEB J. 2015, 29, 3911–3919. [Google Scholar] [CrossRef]
- Li, Y.; Monroig, O.; Zhang, L.; Wang, S.; Zheng, X.; Dick, J.R.; You, C.; Tocher, D.R. Vertebrate fatty acyl desaturase with Δ4 activity. Proc. Natl. Acad. Sci. USA 2010, 107, 16840–16845. [Google Scholar] [CrossRef]
- Martinez, M.; Ichaso, N.; Setien, F.; Durany, N.; Qiu, X.; Roesler, W. The Δ4-desaturation pathway for DHA biosynthesis is operative in the human species: Differences between normal controls and children with the Zellweger syndrome. Lipids Health Dis. 2010, 9, 98. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Hong, H.; MacKenzie, S.L. Identification of a Δ4 fatty acid desaturase from Thraustochytrium sp. involved in the biosynthesis of docosahexanoic acid by Heterologous Expression in Saccharomyces cerevisiae and Brassica juncea. J. Biol. Chem. 2001, 276, 31561–31566. [Google Scholar] [CrossRef] [PubMed]
- Schenck, P.A.; Rakoff, H.; Emken, E.A. δ8 desaturationin vivo of deuterated eicosatrienoic acid by mouse liver. Lipids 1996, 31, 593–600. [Google Scholar] [CrossRef]
- Cook, H.; Byers, D.; Palmer, F.; Spence, M.; Rakoff, H.; Duval, S.; Emken, E. Alternate pathways in the desaturation and chain elongation of linolenic acid, 18:3(n-3), in cultured glioma cells. J. Lipid Res. 1991, 32, 1265–1273. [Google Scholar] [CrossRef]
- Park, W.J.; Kothapalli, K.S.D.; Lawrence, P.; Tyburczy, C.; Brenna, J.T. An alternate pathway to long-chain polyunsaturates: The FADS2 gene product Δ8-desaturates 20:2n-6 and 20:3n-3. J. Lipid Res. 2009, 50, 1195–1202. [Google Scholar] [CrossRef] [PubMed]
- Kabeya, N.; Fonseca, M.M.; Ferrier, D.E.K.; Navarro, J.C.; Bay, L.K.; Francis, D.S.; Tocher, D.R.; Castro, L.F.C.; Monroig, Ó. Genes for de novo biosynthesis of omega-3 polyunsaturated fatty acids are widespread in animals. Sci. Adv. 2018, 4, eaar6849. [Google Scholar] [CrossRef]
- Xue, Z.; He, H.; Hollerbach, D.; Macool, D.J.; Yadav, N.S.; Zhang, H.; Szostek, B.; Zhu, Q. Identification and characterization of new Δ-17 fatty acid desaturases. Appl. Microbiol. Biotechnol. 2013, 97, 1973–1985. [Google Scholar] [CrossRef]
- Jump, D.B.; Clarke, S.D.; Thelen, A.; Liimatta, M.; Ren, B.; Badin, M. Dietary polyunsaturated fatty acid regulation of gene transcription. Prog. Lipid Res. 1996, 35, 227–241. [Google Scholar] [CrossRef]
- Blake, W.L.; Clarke, S.D. Suppression of rat hepatic fatty acid synthase and S14 gene transcription by dietary polyunsaturated fat. J. Nutr. 1990, 120, 1727–1729. [Google Scholar] [CrossRef]
- Mullen, T.D.; Hannun, Y.A.; Obeid, L.M. Ceramide synthases at the centre of sphingolipid metabolism and biology. Biochem. J. 2012, 441, 789–802. [Google Scholar] [CrossRef]
- Brush, R.S.; Tran, J.-T.A.; Henry, K.R.; McClellan, M.E.; Elliott, M.H.; Mandal, M.N.A. Retinal sphingolipids and their very-long-chain fatty acid–containing species. Investig. Opthalmology Vis. Sci. 2010, 51, 4422. [Google Scholar] [CrossRef]
- Sandhoff, R. Very long chain sphingolipids: Tissue expression, function and synthesis. FEBS Lett. 2010, 584, 1907–1913. [Google Scholar] [CrossRef] [PubMed]
- Nachtschatt, M.; Okada, S.; Speight, R. Integral membrane fatty acid desaturases: A review of biochemical, structural, and biotechnological advances. Eur. J. Lipid Sci. Technol. 2020, 122, 2000181. [Google Scholar] [CrossRef]
- Zhu, G.; Koszelak-Rosenblum, M.; Connelly, S.M.; Dumont, M.E.; Malkowski, M.G. The crystal structure of an integral membrane fatty acid α-hydroxylase. J. Biol. Chem. 2015, 290, 29820–29833. [Google Scholar] [CrossRef] [PubMed]
- Vacchina, P.; Tripodi, K.E.J.; Escalante, A.M.; Uttaro, A.D. Characterization of bifunctional sphingolipid Δ4-desaturases/C4-hydroxylases of trypanosomatids by liquid chromatography–electrospray tandem mass spectrometry. Mol. Biochem. Parasitol. 2012, 184, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Markham, J.E.; Cahoon, E.B. Sphingolipid Δ8 unsaturation is important for glucosylceramide biosynthesis and low-temperature performance in Arabidopsis. Plant J. 2012, 69, 769–781. [Google Scholar] [PubMed]
- López-Lara, I.M.; Soto, M.J. Fatty acid synthesis and regulation. In Biogenesis of Fatty Acids, Lipids and Membranes; Springer International Publishing: Cham, Switzerland, 2019; pp. 391–407. [Google Scholar]
- Yamashita, A.; Hayashi, Y.; Nemoto-Sasaki, Y.; Ito, M.; Oka, S.; Tanikawa, T.; Waku, K.; Sugiura, T. Acyltransferases and transacylases that determine the fatty acid composition of glycerolipids and the metabolism of bioactive lipid mediators in mammalian cells and model organisms. Prog. Lipid Res. 2014, 53, 18–81. [Google Scholar] [PubMed]
- Kent, C. Eukaryotic phospholipid biosynthesis. Annu. Rev. Biochem. 1995, 64, 315–343. [Google Scholar] [CrossRef] [PubMed]
- Quehenberger, O.; Armando, A.M.; Brown, A.H.; Milne, S.B.; Myers, D.S.; Merrill, A.H.; Bandyopadhyay, S.; Jones, K.N.; Kelly, S.; Shaner, R.L.; et al. Lipidomics reveals a remarkable diversity of lipids in human plasma. J. Lipid Res. 2010, 51, 3299–3305. [Google Scholar] [CrossRef]
- Shindou, H.; Hishikawa, D.; Harayama, T.; Yuki, K.; Shimizu, T. Recent progress on acyl CoA: Lysophospholipid acyltransferase research. J. Lipid Res. 2009, 50, S46–S51. [Google Scholar] [CrossRef]
- Coleman, R.A.; Lewin, T.M.; Van Horn, C.G.; Gonzalez-Baró, M.R. Do long-chain acyl-CoA synthetases regulate fatty acid Entry into Synthetic Versus degradative pathways? J. Nutr. 2002, 132, 2123–2126. [Google Scholar] [PubMed]
- MacDonald, J.I.S.; Sprecher, H. Phospholipid fatty acid remodeling in mammalian cells. Biochim. Biophys. Acta Lipids Lipid Metab. 1991, 1084, 105–121. [Google Scholar]
- Lands, W.E.M. Stories about acyl chains. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2000, 1483, 1–14. [Google Scholar]
- Lands, W.E.M.; Merkl, I. Metabolism of glycerolipids. J. Biol. Chem. 1963, 238, 898–904. [Google Scholar] [CrossRef]
- Markham, J.E.; Molino, D.; Gissot, L.; Bellec, Y.; Hématy, K.; Marion, J.; Belcram, K.; Palauqui, J.-C.; Satiat-JeuneMaître, B.; Faure, J.-D. Sphingolipids containing very-long-chain fatty acids define a secretory pathway for specific polar plasma membrane protein targeting in arabidopsis. Plant Cell 2011, 23, 2362–2378. [Google Scholar] [CrossRef]
- Samovski, D.; Jacome-Sosa, M.; Abumrad, N.A. Fatty Acid transport and signaling: Mechanisms and physiological implications. Annu. Rev. Physiol. 2023, 85, 317–337. [Google Scholar] [CrossRef]
- Farkas, T. Adaptation of fatty acid compositions to temperature—A study on planktonic crustaceans. Comp. Biochem. Physiol. Part B Comp. Biochem. 1979, 64, 71–76. [Google Scholar] [CrossRef]
- Dymond, M.K. Mammalian phospholipid homeostasis: Evidence that membrane curvature elastic stress drives homeoviscous adaptation in vivo. J. R. Soc. Interface 2016, 13, 20160228. [Google Scholar] [CrossRef]
- Kulig, W.; Pasenkiewicz-Gierula, M.; Róg, T. Cis and trans unsaturated phosphatidylcholine bilayers: A molecular dynamics simulation study. Chem. Phys. Lipids 2016, 195, 12–20. [Google Scholar]
- Gillan, F.T.; Johns, R.B.; Verheyen, T.V.; Volkman, J.K.; Bavor, H.J. Trans-monounsaturated acids in a marine bacterial isolate. Appl. Environ. Microbiol. 1981, 41, 849–856. [Google Scholar] [CrossRef]
- Hanahan, D.J.; Brockerhoff, H.; Barron, E.J. The Site of Attack of Phospholipase (Lecithinase) A on Lecithin: A Re-evaluation. J. Biol. Chem. 1960, 235, 1917–1923. [Google Scholar] [CrossRef] [PubMed]
- Tattrie, N.H. Positional distribution of saturated and unsaturated fatty acids on egg lecithin. J. Lipid Res. 1959, 1, 60–65. [Google Scholar] [CrossRef]
- Pearson, R.H.; Pascher, I. The molecular structure of lecithin dihydrate. Nature 1979, 281, 499–501. [Google Scholar] [CrossRef] [PubMed]
- Büldt, G.; Gally, H.U.; Seelig, J.; Zaccai, G. Neutron diffraction studies on phosphatidylcholine model membranes. J. Mol. Biol. 1979, 134, 673–691. [Google Scholar] [CrossRef] [PubMed]
- Sanders, R.L.; Longmore, W.J. Phosphatidylglycerol in rat lung. II. Comparison of occurrence, composition, and metabolism in surfactant and residual lung fractions. Biochemistry 1975, 14, 835–840. [Google Scholar] [CrossRef]
- Xie, D.; Seremwe, M.; Edwards, J.G.; Podolsky, R.; Bollag, W.B. Distinct Effects of Different Phosphatidylglycerol Species on Mouse Keratinocyte Proliferation. PLoS ONE 2014, 9, e107119. [Google Scholar]
- O’Donnell, V.B. New appreciation for an old pathway: The Lands Cycle moves into new arenas in health and disease. Biochem. Soc. Trans. 2022, 50, 1–11. [Google Scholar]
- Nakanishi, H.; Iida, Y.; Shimizu, T.; Taguchi, R. Separation and quantification of sn-1 and sn-2 fatty acid positional isomers in phosphatidylcholine by RPLC-ESIMS/MS. J. Biochem. 2010, 147, 245–256. [Google Scholar] [CrossRef]
- Wood, R.; Harlow, R.D. Structural studies of neutral glycerides and phosphoglycerides of rat liver. Arch. Biochem. Biophys. 1969, 131, 495–501. [Google Scholar] [CrossRef]
- Kuksis, A.; Breckenridge, W.C.; Marai, L.; Stachnyk, O. Molecular species of lecithins of rat heart, kidney, and plasma. J. Lipid Res. 1969, 10, 25–32. [Google Scholar] [CrossRef]
- Yabuuchi, H.; O’Brien, J.S. Positional distribution of fatty acids in glycerophosphatides of bovine gray matter. J. Lipid Res. 1968, 9, 65–67. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.S.; Rouser, G. The fatty acid composition of brain sphingolipids: Sphingomyelin, ceramide, cerebroside, and cerebroside sulfate. J. Lipid Res. 1964, 5, 339–342. [Google Scholar] [CrossRef] [PubMed]
- Kuksis, A.; Marai, L. Determination of the complete structure of natural lecithins. Lipids 1967, 2, 217–224. [Google Scholar] [CrossRef]
- Holub, B.J.; Kuksis, A. Molecular species of phosphatidyl ethanolamine from egg yolk. Lipids 1969, 4, 466–472. [Google Scholar] [CrossRef]
- Doğru Pekiner, B. Fatty acid composition of red blood cell membrane phosphatidylethanolamine and phosphatidylcholine in the rat, rabbit, human and dog: Sıçan, tavşan, insan ve köpek eritrosit membranı fosfatidiletanol. Ankara Univ. Eczac. Fak. Derg. 2002, 31, 169–182. [Google Scholar] [CrossRef]
- Kim, H.-Y.; Huang, B.X.; Spector, A.A. Phosphatidylserine in the brain: Metabolism and function. Prog. Lipid Res. 2014, 56, 1–18. [Google Scholar] [PubMed]
- Hamilton, J.; Greiner, R.; Salem, N.; Kim, H.-Y. n-3 Fatty acid deficiency decreases phosphatidylserine accumulation selectively in neuronal tissues. Lipids 2000, 35, 863–869. [Google Scholar] [CrossRef]
- Clark, S.R.; Thomas, C.P.; Hammond, V.J.; Aldrovandi, M.; Wilkinson, G.W.; Hart, K.W.; Murphy, R.C.; Collins, P.W.; O’Donnell, V.B. Characterization of platelet aminophospholipid externalization reveals fatty acids as molecular determinants that regulate coagulation. Proc. Natl. Acad. Sci. USA 2013, 110, 5875–5880. [Google Scholar] [CrossRef]
- Thompson, W.; MacDonald, G. Isolation and characterization of cytidine diphosphate diglyceride from beef liver. J. Biol. Chem. 1975, 250, 6779–6785. [Google Scholar] [CrossRef]
- Holub, B.J.; Kuksis, A.; Thompson, W. Molecular species of mono-, di-, and triphosphoinositides of bovine brain. J. Lipid Res. 1970, 11, 558–564. [Google Scholar] [CrossRef]
- Ulmann, L.; Mimouni, V.; Roux, S.; Porsolt, R.; Poisson, J.-P. Brain and hippocampus fatty acid composition in phospholipid classes of aged-relative cognitive deficit rats. Prostaglandins Leukot. Essent. Fat. Acids 2001, 64, 189–195. [Google Scholar]
- Gijón, M.A.; Riekhof, W.R.; Zarini, S.; Murphy, R.C.; Voelker, D.R. Lysophospholipid Acyltransferases and Arachidonate Recycling in Human Neutrophils. J. Biol. Chem. 2008, 283, 30235–30245. [Google Scholar] [PubMed]
- Jia, D.; Zhang, J.; Nie, J.; Andersen, J.-P.; Rendon, S.; Zheng, Y.; Liu, X.; Tian, Z.; Shi, Y. Cardiolipin remodeling by ALCAT1 links hypoxia to coronary artery disease by promoting mitochondrial dysfunction. Mol. Ther. 2021, 29, 3498–3511. [Google Scholar] [PubMed]
- Pennington, E.R.; Funai, K.; Brown, D.A.; Shaikh, S.R. The role of cardiolipin concentration and acyl chain composition on mitochondrial inner membrane molecular organization and function. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2019, 1864, 1039–1052. [Google Scholar]
- Li, J.; Romestaing, C.; Han, X.; Li, Y.; Hao, X.; Wu, Y.; Sun, C.; Liu, X.; Jefferson, L.S.; Xiong, J.; et al. Cardiolipin remodeling by ALCAT1 links oxidative stress and mitochondrial dysfunction to obesity. Cell Metab. 2010, 12, 154–165. [Google Scholar]
- Stefanyk, L.E.; Coverdale, N.; Roy, B.D.; Peters, S.J.; LeBlanc, P.J. Skeletal muscle type comparison of subsarcolemmal mitochondrial membrane phospholipid fatty acid composition in rat. J. Membr. Biol. 2010, 234, 207–215. [Google Scholar]
- Schlame, M.; Ren, M.; Xu, Y.; Greenberg, M.L.; Haller, I. Molecular symmetry in mitochondrial cardiolipins. Chem. Phys. Lipids 2005, 138, 38–49. [Google Scholar]
- Berger, A.; Gershwin, M.E.; German, J.B. Effects of various dietary fats on cardiolipin acyl composition during ontogeny of mice. Lipids 1992, 27, 605–612. [Google Scholar]
- Wolff, R.L.; Entressangles, B. Compositional changes of fatty acids in the 1(1″)- and 2(2″)-positions of cardiolipin from liver, heart, and kidney mitochondria of rats fed a low-fat diet. Biochim. Biophys. Acta Lipids Lipid Metab. 1991, 1082, 136–142. [Google Scholar]
- Anderson, D.M.G.; Ablonczy, Z.; Koutalos, Y.; Hanneken, A.M.; Spraggins, J.M.; Calcutt, M.W.; Crouch, R.K.; Caprioli, R.M.; Schey, K.L. Bis(monoacylglycero)phosphate lipids in the retinal pigment epithelium implicate lysosomal/endosomal dysfunction in a model of Stargardt disease and human retinas. Sci. Rep. 2017, 7, 17352. [Google Scholar]
- Besson, N.; Hullin-Matsuda, F.; Makino, A.; Murate, M.; Lagarde, M.; Pageaux, J.-F.; Kobayashi, T.; Delton-Vandenbroucke, I. Selective incorporation of docosahexaenoic acid into lysobisphosphatidic acid in cultured THP-1 macrophages. Lipids 2006, 41, 189–196. [Google Scholar] [PubMed]
- Luquain, C.; Dolmazon, R.; Enderlin, J.M.; Laugier, C.; Lagarde, M.; Pageaux, J.F. Bis(monoacylglycerol) phosphate in rat uterine stromal cells: Structural characterization and specific esterification of docosahexaenoic acid. Biochem. J. 2000, 351 Pt 3, 795–804. [Google Scholar] [PubMed]
- Huterer, S.; Wherrett, J. Metabolism of bis(monoacylglycero)phosphate in macrophages. J. Lipid Res. 1979, 20, 966–973. [Google Scholar] [PubMed]
- Wherrett, J.R.; Huterer, S. Bis-(monoacylglyceryl)-phosphate of rat and human liver: Fatty acid composition and NMR spectroscopy. Lipids 1973, 8, 531–533. [Google Scholar]
- Holbrook, P.G.; Pannell, L.K.; Murata, Y.; Daly, J.W. Bis(monoacylglycero) phosphate from PC12 cells, a phospholipid that can comigrate with phosphatidic acid: Molecular species analysis by fast atom bombardment mass spectrometry. Biochim. Biophys. Acta Lipids Lipid Metab. 1992, 1125, 330–334. [Google Scholar] [CrossRef]
- Ryan, S.D.; Harris, C.S.; Carswell, C.L.; Baenziger, J.E.; Bennett, S.A.L. Heterogeneity in the sn-1 carbon chain of platelet-activating factor glycerophospholipids determines pro- or anti-apoptotic signaling in primary neurons. J. Lipid Res. 2008, 49, 2250–2258. [Google Scholar]
- Nakagawa, Y.; Horrocks, L.A. Different metabolic rates for arachidonoyl molecular species of ethanolamine glycerophospholipids in rat brain. J. Lipid Res. 1988, 27, 629–636. [Google Scholar]
- Pruett, S.T.; Bushnev, A.; Hagedorn, K.; Adiga, M.; Haynes, C.A.; Sullards, M.C.; Liotta, D.C.; Merrill, A.H. Thematic review series: Sphingolipids. Biodiversity of sphingoid bases (“sphingosines”) and related amino alcohols. J. Lipid Res. 2008, 49, 1621–1639. [Google Scholar]
- Merrill, A.H.; Nixon, D.W.; Williams, R.D. Activities of serine palmitoyltransferase (3-ketosphinganine synthase) in microsomes from different rat tissues. J. Lipid Res. 1985, 26, 617–622. [Google Scholar]
- Ardail, D.; Popa, I.; Alcantara, K.; Pons, A.; Zanetta, J.; Louisot, P.; Thomas, L.; Portoukalian, J. Occurrence of ceramides and neutral glycolipids with unusual long-chain base composition in purified rat liver mitochondria. FEBS Lett. 2001, 488, 160–164. [Google Scholar] [CrossRef]
- Eckhardt, M. Fatty Acid 2-Hydroxylase and 2-Hydroxylated Sphingolipids: Metabolism and Function in Health and Diseases. Int. J. Mol. Sci. 2023, 24, 4908. [Google Scholar] [CrossRef] [PubMed]
- Marquês, J.T.; Marinho, H.S.; de Almeida, R.F.M. Sphingolipid hydroxylation in mammals, yeast and plants—An integrated view. Prog. Lipid Res. 2018, 71, 18–42. [Google Scholar] [CrossRef] [PubMed]
- Sessa, L.; Nardiello, A.M.; Santoro, J.; Concilio, S.; Piotto, S. Hydroxylated fatty acids: The role of the sphingomyelin synthase and the origin of selectivity. Membranes 2021, 11, 787. [Google Scholar] [CrossRef] [PubMed]
- Barenholz, Y. Sphingomyelin-lecithin balance in membranes: Composition, structure, and function relationships. In Physiology of Membrane Fluidity; Shinitzky, M., Ed.; CRC Press: Boca Raton, FL, USA, 1984; Volume 1, pp. 131–173. [Google Scholar]
- Ramstedt, B.; Leppimäki, P.; Axberg, M.; Slotte, J.P. Analysis of natural and synthetic sphingomyelins using high-performance thin-layer chromatography. Eur. J. Biochem. 1999, 266, 997–1002. [Google Scholar] [CrossRef]
- Karlsson, A.Å.; Michélsen, P.; Odham, G. Molecular species of sphingomyelin: Determination by high-performance liquid chromatography/mass spectrometry with electrospray and high-performance liquid chromatography/tandem mass spectrometry with atmospheric pressure chemical ionization. J. Mass Spectrom. 1998, 33, 1192–1198. [Google Scholar] [CrossRef]
- Calhoun, W.I.; Shipley, G.G. Fatty acid composition and thermal behavior of natural sphingomyelins. Biochim. Biophys. Acta Biomembr. 1979, 555, 436–441. [Google Scholar] [CrossRef]
- Poulos, A.; Sharp, P.; Johnson, D.; White, I.; Fellenberg, A. The occurrence of polyenoic fatty acids with greater than 22 carbon atoms in mammalian spermatozoa. Biochem. J. 1986, 240, 891–895. [Google Scholar] [CrossRef]
- Robinson, B.S.; Johnson, D.W.; Poulos, A. Novel molecular species of sphingomyelin containing 2-hydroxylated polyenoic very-long-chain fatty acids in mammalian testes and spermatozoa. J. Biol. Chem. 1992, 267, 1746–1751. [Google Scholar] [CrossRef]
- Skotland, T.; Sandvig, K. Need for more focus on lipid species in studies of biological and model membranes. Prog. Lipid Res. 2022, 86, 101160. [Google Scholar] [CrossRef]
- Fujiwara, Y.; Hama, K.; Yokoyama, K. Mass spectrometry in combination with a chiral column and multichannel-MRM allows comprehensive analysis of glycosphingolipid molecular species from mouse brain. Carbohydr. Res. 2020, 490, 107959. [Google Scholar] [CrossRef]
- Baumann, N.; Pham-Dinh, D. Biology of oligodendrocyte and myelin in the mammalian central nervous system. Physiol. Rev. 2001, 81, 871–927. [Google Scholar] [CrossRef] [PubMed]
- Klenk, E.; Hendricks, U.W.; Gielen, W. β-D-Galaktosido-(1->3)-N-acetyl-D-galaktosamin, ein kristallisiertes Disaccharid aus menschlichen Gehirngangliosiden. Biol. Chem. 1962, 330, 140–144. [Google Scholar] [CrossRef] [PubMed]
- Sastry, P.S. Lipids of nervous tissue: Composition and metabolism. Prog. Lipid Res. 1985, 24, 69–176. [Google Scholar] [PubMed]
- Burr, G.O.; Burr, M.M. A new deficiency disease produced by the rigid exclusion of fat from the diet. J. Biol. Chem. 1929, 82, 345–367. [Google Scholar] [CrossRef]
- von Euler, U.S. History and development of prostaglandins. Gen. Pharmacol. Vasc. Syst. 1983, 14, 3–6. [Google Scholar] [CrossRef]
- Rodriguez-Estrada, M.T.; Tatay, A.C.; Cardenia, V.; Garcia-Llatas, G. Fats and sterols. In Reference Module in Biomedical Sciences; Elsevier: Amsterdam, The Netherlands, 2014. [Google Scholar]
- Covino, R.; Ballweg, S.; Stordeur, C.; Michaelis, J.B.; Puth, K.; Wernig, F.; Bahrami, A.; Ernst, A.M.; Hummer, G.; Ernst, R. A Eukaryotic sensor for membrane lipid saturation. Mol. Cell 2016, 63, 49–59. [Google Scholar] [CrossRef]
- Radhakrishnan, A.; Goldstein, J.L.; McDonald, J.G.; Brown, M.S. Switch-like control of SREBP-2 transport triggered by small changes in ER cholesterol: A delicate Balance. Cell Metab. 2008, 8, 512–521. [Google Scholar] [CrossRef]
- Matveyenka, M.; Rizevsky, S.; Kurouski, D. Length and unsaturation of fatty acids of phosphatidic acid determines the aggregation rate of insulin and modifies the structure and toxicity of insulin aggregates. ACS Chem. Neurosci. 2022, 13, 2483–2489. [Google Scholar] [CrossRef]
- Baccouch, R.; Shi, Y.; Vernay, E.; Mathelié-Guinlet, M.; Taib-Maamar, N.; Villette, S.; Feuillie, C.; Rascol, E.; Nuss, P.; Lecomte, S.; et al. The impact of lipid polyunsaturation on the physical and mechanical properties of lipid membranes. Biochim. Biophys. Acta Biomembr. 2023, 1865, 184084. [Google Scholar] [CrossRef]
- Hashimoto, M.; Hossain, S.; Al Mamun, A.; Matsuzaki, K.; Arai, H. Docosahexaenoic acid: One molecule diverse functions. Crit. Rev. Biotechnol. 2017, 37, 579–597. [Google Scholar] [CrossRef]
- Ibarguren, M.; López, D.J.; Escribá, P.V. The effect of natural and synthetic fatty acids on membrane structure, microdomain organization, cellular functions and human health. Biochim. Biophys. Acta Biomembr. 2014, 1838, 1518–1528. [Google Scholar] [CrossRef] [PubMed]
- Rustan, A.C.; Drevon, C.A. Fatty acids: Structures and properties. In eLS; John Wiley & Sons: Hoboken, NJ, USA, 2005. [Google Scholar]
- Small, D.M. Lateral chain packing in lipids and membranes. J. Lipid Res. 1984, 25, 1490–1500. [Google Scholar] [CrossRef] [PubMed]
- Mondal, D.; Dutta, R.; Banerjee, P.; Mukherjee, D.; Maiti, T.K.; Sarkar, N. Modulation of membrane fluidity performed on model phospholipid membrane and live cell membrane: Revealing through spatiotemporal approaches of FLIM, FAIM, and TRFS. Anal. Chem. 2019, 91, 4337–4345. [Google Scholar] [CrossRef] [PubMed]
- Goñi, F.M. The basic structure and dynamics of cell membranes: An update of the Singer–Nicolson model. Biochim. Biophys. Acta Biomembr. 2014, 1838, 1467–1476. [Google Scholar] [CrossRef] [PubMed]
- De Santis, A.; Varela, Y.; Sot, J.; D’Errico, G.; Goñi, F.M.; Alonso, A. Omega-3 polyunsaturated fatty acids do not fluidify bilayers in the liquid-crystalline state. Sci. Rep. 2018, 8, 16240. [Google Scholar] [CrossRef]
- Lor, C.; Hirst, L. Effects of low concentrations of docosahexaenoic acid on the structure and phase behavior of model lipid membranes. Membranes 2015, 5, 857–874. [Google Scholar] [CrossRef]
- Killian, J.A. Hydrophobic mismatch between proteins and lipids in membranes. Biochim. Biophys. Acta Rev. Biomembr. 1998, 1376, 401–416. [Google Scholar] [CrossRef]
- Salmon, A.; Dodd, S.W.; Williams, G.D.; Beach, J.M.; Brown, M.F. Configurational statistics of acyl chains in polyunsaturated lipid bilayers from deuterium NMR. J. Am. Chem. Soc. 1987, 109, 2600–2609. [Google Scholar] [CrossRef]
- Yang, X.; Sheng, W.; Sun, G.Y.; Lee, J.C.-M. Effects of fatty acid unsaturation numbers on membrane fluidity and α-secretase-dependent amyloid precursor protein processing. Neurochem. Int. 2011, 58, 321–329. [Google Scholar] [CrossRef]
- Straume, M.; Litman, B.J. Equilibrium and dynamic structure of large, unilamellar, unsaturated acyl chain phosphatidylcholine vesicles. Higher order analysis of 1,6-diphenyl-1,3,5-hexatriene and 1-[4-(trimethylammonio)phenyl]-6-phenyl-1,3,5-hexatriene anisotropy decay. Biochemistry 1987, 26, 5113–5120. [Google Scholar] [CrossRef]
- Roach, C.; Feller, S.E.; Ward, J.A.; Shaikh, S.R.; Zerouga, M.; Stillwell, W. Comparison of cis and trans fatty acid containing phosphatidylcholines on membrane properties. Biochemistry 2004, 43, 6344–6351. [Google Scholar] [CrossRef]
- Christie, W.W.; Holman, R.T. Synthesis and characterization of the complete series of methylene-interrupted cis,cis-octadecadienoic acids. Chem. Phys. Lipids 1967, 1, 407–423. [Google Scholar] [CrossRef]
- Tyler, A.I.I.; Greenfield, J.L.; Seddon, J.M.; Brooks, N.J.; Purushothaman, S. Coupling phase behavior of fatty acid containing membranes to membrane bio-mechanics. Front. Cell Dev. Biol. 2019, 7, 187. [Google Scholar] [CrossRef]
- Brenner, R. Effect of unsaturated acids on membrane structure and enzyme kinetics. Prog. Lipid Res. 1984, 23, 69–96. [Google Scholar] [CrossRef]
- Schumann, J.; Leichtle, A.; Thiery, J.; Fuhrmann, H. Fatty acid and peptide profiles in plasma membrane and membrane rafts of PUFA supplemented RAW264.7 macrophages. PLoS ONE 2011, 6, e24066. [Google Scholar] [CrossRef]
- Stillwell, W. The role of polyunsaturated lipids in membrane raft function. Scand. J. Food Nutr. 2006, 50, 107–113. [Google Scholar] [CrossRef]
- Bennett, W.F.D.; Shea, J.-E.; Tieleman, D.P. Phospholipid chain interactions with cholesterol drive domain formation in lipid membranes. Biophys. J. 2018, 114, 2595–2605. [Google Scholar] [CrossRef]
- Martinez-Seara, H.; Róg, T.; Karttunen, M.; Vattulainen, I.; Reigada, R. Cholesterol induces specific spatial and orientational order in cholesterol/phospholipid membranes. PLoS ONE 2010, 5, e11162. [Google Scholar] [CrossRef]
- Ye, S.; Tan, L.; Ma, J.; Shi, Q.; Li, J. Polyunsaturated docosahexaenoic acid suppresses oxidative stress induced endothelial cell calcium influx by altering lipid composition in membrane caveolar rafts. Prostaglandins Leukot. Essent. Fat. Acids 2010, 83, 37–43. [Google Scholar] [CrossRef]
- Wassall, S.R.; Stillwell, W. Polyunsaturated fatty acid–cholesterol interactions: Domain formation in membranes. Biochim. Biophys. Acta Biomembr. 2009, 1788, 24–32. [Google Scholar] [CrossRef]
- Chen, W.; Jump, D.B.; Esselman, W.J.; Busik, J.V. Inhibition of cytokine Signaling in human retinal endothelial cells through modification of caveolae/lipid rafts by docosahexaenoic acid. Investig. Opthalmology Vis. Sci. 2007, 48, 18. [Google Scholar] [CrossRef] [PubMed]
- Pitman, M.C.; Suits, F.; MacKerell, A.D.; Feller, S.E. Molecular-level organization of saturated and polyunsaturated fatty acids in a phosphatidylcholine bilayer containing cholesterol. Biochemistry 2004, 43, 15318–15328. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, M.; Hossain, M.S.; Shimada, T.; Yamasaki, H.; Fujii, Y.; Shido, O. Effects of docosahexaenoic acid on annular lipid fluidity of the rat bile canalicular plasma membrane. J. Lipid Res. 2001, 42, 1160–1168. [Google Scholar] [CrossRef] [PubMed]
- Stulnig, T.M.; Huber, J.; Leitinger, N.; Imre, E.-M.; Angelisová, P.; Nowotny, P.; Waldhäusl, W. Polyunsaturated eicosapentaenoic acid displaces proteins from membrane rafts by altering raft lipid composition. J. Biol. Chem. 2001, 276, 37335–37340. [Google Scholar] [CrossRef]
- Hashimoto, M.; Hossain, M.S.; Yamasaki, H.; Yazawa, K.; Masumura, S. Effects of eicosapentaenoic acid and docosahexaenoic acid on plasma membrane fluidity of aortic endothelial cells. Lipids 1999, 34, 1297–1304. [Google Scholar] [CrossRef]
- Van Blitterswijk, W.J.; Van der Meer, B.W.; Hilkmann, H. Quantitative contributions of cholesterol and the individual classes of phospholipids and their degree of fatty acyl (un)saturation to membrane fluidity measured by fluorescence polarization. Biochemistry 1987, 26, 1746–1756. [Google Scholar] [CrossRef]
- Cheng, V.; Rallabandi, R.; Gorusupudi, A.; Lucas, S.; Rognon, G.; Bernstein, P.S.; Rainier, J.D.; Conboy, J.C. Influence of very-long-chain polyunsaturated fatty acids on membrane structure and dynamics. Biophys. J. 2022, 121, 2730–2741. [Google Scholar] [CrossRef]
- Armstrong, V.T.; Brzustowicz, M.R.; Wassall, S.R.; Jenski, L.J.; Stillwell, W. Rapid flip-flop in polyunsaturated (docosahexaenoate) phospholipid membranes. Arch. Biochem. Biophys. 2003, 414, 74–82. [Google Scholar] [CrossRef]
- Jacobs, M.L.; Faizi, H.A.; Peruzzi, J.A.; Vlahovska, P.M.; Kamat, N.P. EPA and DHA differentially modulate membrane elasticity in the presence of cholesterol. Biophys. J. 2021, 120, 2317–2329. [Google Scholar] [CrossRef]
- Rawicz, W.; Olbrich, K.C.; McIntosh, T.; Needham, D.; Evans, E. Effect of chain length and unsaturation on elasticity of lipid bilayers. Biophys. J. 2000, 79, 328–339. [Google Scholar] [CrossRef]
- Beck, R.; Bertolino, S.; Abbot, S.E.; Aaronson, P.I.; Smirnov, S.V. Modulation of arachidonic acid release and membrane fluidity by Albumin in Vascular Smooth muscle and endothelial cells. Circ. Res. 1998, 83, 923–931. [Google Scholar] [CrossRef] [PubMed]
- Treen, M.; Uauy, R.D.; Jameson, D.M.; Thomas, V.L.; Hoffman, D.R. Effect of docosahexaenoic acid on membrane fluidity and function in intact cultured Y-79 retinoblastoma cells. Arch. Biochem. Biophys. 1992, 294, 564–570. [Google Scholar] [CrossRef] [PubMed]
- Hendriks, T.; Klompmakers, A.A.; Daemen, F.J.M.; Bonting, S.L. Biochemical aspects of the visual process XXXII. Movement of sodium ions through bilayers composed of retinal and rod outer segment lipids. Biochim. Biophys. Acta Biomembr. 1976, 433, 271–281. [Google Scholar] [CrossRef]
- Demel, R.A.; Geurts van Kessel, W.S.M.; van Deenen, L.L.M. The properties of polyunsaturated lecithins in monolayers and liposomes and the interactions of these lecithins with cholesterol. Biochim. Biophys. Acta Biomembr. 1972, 266, 26–40. [Google Scholar] [CrossRef]
- Sherratt, S.C.R.; Juliano, R.A.; Copland, C.; Bhatt, D.L.; Libby, P.; Mason, R.P. EPA and DHA containing phospholipids have contrasting effects on membrane structure. J. Lipid Res. 2021, 62, 100106. [Google Scholar] [CrossRef]
- Ehringer, W.; Belcher, D.; Wassall, S.R.; Stillwell, W. A comparison of the effects of linolenic (18:3Ω3) and docosahexaenoic (22:6Ω3) acids on phospholipid bilayers. Chem. Phys. Lipids 1990, 54, 79–88. [Google Scholar] [CrossRef]
- Mitchell, D.C.; Litman, B.J. Effect of cholesterol on molecular order and dynamics in highly polyunsaturated phospholipid bilayers. Biophys. J. 1998, 75, 896–908. [Google Scholar] [CrossRef]
- Li, X.; Zhou, S.; Lin, X. Molecular view on the impact of DHA molecules on the physical properties of a model cell membrane. J. Chem. Inf. Model. 2022, 62, 2421–2431. [Google Scholar] [CrossRef]
- Pinot, M.; Vanni, S.; Pagnotta, S.; Lacas-Gervais, S.; Payet, L.-A.; Ferreira, T.; Gautier, R.; Goud, B.; Antonny, B.; Barelli, H. Polyunsaturated phospholipids facilitate membrane deformation and fission by endocytic proteins. Science 2014, 345, 693–697. [Google Scholar] [CrossRef]
- Stillwell, W.; Wassall, S.R. Docosahexaenoic acid: Membrane properties of a unique fatty acid. Chem. Phys. Lipids 2003, 126, 1–27. [Google Scholar] [CrossRef]
- Williams, E.E.; Jenski, L.J.; Stillwell, W. Docosahexaenoic acid (DHA) alters the structure and composition of membranous vesicles exfoliated from the surface of a murine leukemia cell line. Biochim. Biophys. Acta Biomembr. 1998, 1371, 351–362. [Google Scholar] [CrossRef] [PubMed]
- Koenig, B.W.; Strey, H.H.; Gawrisch, K. Membrane lateral compressibility determined by NMR and x-ray diffraction: Effect of acyl chain polyunsaturation. Biophys. J. 1997, 73, 1954–1966. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, D.C.; Litman, B.J. Modulation of receptor signaling by phospholipid acyl chain composition. In Fatty Acids: Nutrition and Health; Mostofsky, D.I., Yehuda, S., Salem, N., Eds.; Humana Press: Totowa, NJ, USA, 2001; pp. 23–40. ISBN 978-1-59259-119-0. [Google Scholar]
- Joardar, A.; Pattnaik, G.P.; Chakraborty, H. Mechanism of membrane fusion: Interplay of lipid and peptide. J. Membr. Biol. 2022, 255, 211–224. [Google Scholar] [CrossRef] [PubMed]
- Ellens, H.; Siegel, D.P.; Alford, D.; Yeagle, P.L.; Boni, L.; Lis, L.J.; Quinn, P.J.; Bentz, J. Membrane fusion and inverted phases. Biochemistry 1989, 28, 3692–3703. [Google Scholar] [CrossRef] [PubMed]
- Ahkong, Q.F.; Fisher, D.; Tampion, W.; Lucy, J.A. The fusion of erythrocytes by fatty acids, esters, retinol and α-tocopherol. Biochem. J. 1973, 136, 147–155. [Google Scholar] [CrossRef]
- Meers, P.; Hong, K.; Papahadjopoulos, D. Free fatty acid enhancement of cation-induced fusion of liposomes: Synergism with synexin and other promoters of vesicle aggregation. Biochemistry 1988, 27, 6784–6794. [Google Scholar] [CrossRef]
- Creutz, C.E. cis-unsaturated fatty acids induce the fusion of chromaffin granules aggregated by synexin. J. Cell Biol. 1981, 91, 247–256. [Google Scholar] [CrossRef]
- Funari, S.S.; Barceló, F.; Escribá, P.V. Effects of oleic acid and its congeners, elaidic and stearic acids, on the structural properties of phosphatidylethanolamine membranes. J. Lipid Res. 2003, 44, 567–575. [Google Scholar] [CrossRef]
- Prades, J.; Funari, S.S.; Escribá, P.V.; Barceló, F. Effects of unsaturated fatty acids and triacylglycerols on phosphatidylethanolamine membrane structure. J. Lipid Res. 2003, 44, 1720–1727. [Google Scholar] [CrossRef]
- Jenske, R.; Lindström, F.; Gröbner, G.; Vetter, W. Impact of free hydroxylated and methyl-branched fatty acids on the organization of lipid membranes. Chem. Phys. Lipids 2008, 154, 26–32. [Google Scholar] [CrossRef]
- Niebylski, C.D.; Salem, N. A calorimetric investigation of a series of mixed-chain polyunsaturated phosphatidylcholines: Effect of sn-2 chain length and degree of unsaturation. Biophys. J. 1994, 67, 2387–2393. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Monteiro, J.; Leslie, M.; Moghadasian, M.H.; Arendt, B.M.; Allard, J.P.; Ma, D.W.L. The role of n-6 and n-3 polyunsaturated fatty acids in the manifestation of the metabolic syndrome in cardiovascular disease and non-alcoholic fatty liver disease. Food Funct. 2014, 5, 426. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, M.; Hossain, S. Neuroprotective and ameliorative actions of polyunsaturated fatty acids against neuronal diseases: Beneficial effect of docosahexaenoic acid on cognitive decline in Alzheimer’s disease. J. Pharmacol. Sci. 2011, 116, 150–162. [Google Scholar] [CrossRef] [PubMed]
- Eltweri, A.M.; Thomas, A.L.; Metcalfe, M.; Calder, P.C.; Dennison, A.R.; Bowrey, D.J. Potential applications of fish oils rich in omega-3 polyunsaturated fatty acids in the management of gastrointestinal cancer. Clin. Nutr. 2017, 36, 65–78. [Google Scholar] [CrossRef]
- Dyerberg, J. Linolenate-derived polyunsaturated fatty acids and prevention of atherosclerosis. Nutr. Rev. 1986, 44, 125–134. [Google Scholar] [CrossRef]
- Sassa, T.; Suto, S.; Okayasu, Y.; Kihara, A. A shift in sphingolipid composition from C24 to C16 increases susceptibility to apoptosis in HeLa cells. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2012, 1821, 1031–1037. [Google Scholar] [CrossRef]
- Cao, H.; Gerhold, K.; Mayers, J.R.; Wiest, M.M.; Watkins, S.M.; Hotamisligil, G.S. Identification of a lipokine, a lipid hormone linking adipose tissue to systemic metabolism. Cell 2008, 134, 933–944. [Google Scholar] [CrossRef]
- Carrillo, C.; Cavia, M.D.M.; Alonso-Torre, S.R. Antitumor effect of oleic acid; mechanisms of action. A review. Nutr. Hosp. 2012, 27, 1860–1865. [Google Scholar]
- Park, W.-J.; Park, J.-W.; Merrill, A.H.; Storch, J.; Pewzner-Jung, Y.; Futerman, A.H. Hepatic fatty acid uptake is regulated by the sphingolipid acyl chain length. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2014, 1841, 1754–1766. [Google Scholar] [CrossRef]
- Huang, J.M.; Xian, H.; Bacaner, M. Long-chain fatty acids activate calcium channels in ventricular myocytes. Proc. Natl. Acad. Sci. USA 1992, 89, 6452–6456. [Google Scholar] [CrossRef]
- Chang, J.P.-C.; Su, K.-P.; Mondelli, V.; Pariante, C.M. Omega-3 Polyunsaturated Fatty Acids in Youths with Attention Deficit Hyperactivity Disorder: A Systematic Review and Meta-Analysis of Clinical Trials and Biological Studies. Neuropsychopharmacology 2018, 43, 534–545. [Google Scholar] [CrossRef] [PubMed]
- Watkins, B. Bioactive fatty acids: Role in bone biology and bone cell function. Prog. Lipid Res. 2001, 40, 125–148. [Google Scholar] [CrossRef] [PubMed]
- Holman, R.; Johnson, S.; Hatch, T. A case of human linolenic acid deficiency involving neurological abnormalities. Am. J. Clin. Nutr. 1982, 35, 617–623. [Google Scholar] [CrossRef] [PubMed]
- Brown, W.R.; Hansen, A.E.; Burr, G.O.; McQuarrie, I. Effects of prolonged use of extremely low-fat diet on an adult human subject. J. Nutr. 1938, 16, 511–524. [Google Scholar] [CrossRef]
- Burr, G.O.; Burr, M.M.; Miller, E.S. On the fatty acids essential in nutrition. III. J. Biol. Chem. 1932, 97, 1–9. [Google Scholar] [CrossRef]
- Sambra, V.; Echeverria, F.; Valenzuela, A.; Chouinard-Watkins, R.; Valenzuela, R. Docosahexaenoic and arachidonic acids as neuroprotective nutrients throughout the life cycle. Nutrients 2021, 13, 986. [Google Scholar] [CrossRef]
- Hadley, K.; Ryan, A.; Forsyth, S.; Gautier, S.; Salem, N. The essentiality of arachidonic acid in infant development. Nutrients 2016, 8, 216. [Google Scholar] [CrossRef]
- Martinez, M. Tissue levels of polyunsaturated fatty acids during early human development. J. Pediatr. 1992, 120, S129–S138. [Google Scholar] [CrossRef]
- Zheng, Y.; Yin, H.; Boeglin, W.E.; Elias, P.M.; Crumrine, D.; Beier, D.R.; Brash, A.R. Lipoxygenases mediate the effect of essential fatty acid in skin barrier formation. J. Biol. Chem. 2011, 286, 24046–24056. [Google Scholar] [CrossRef]
- Casillas-Vargas, G.; Ocasio-Malavé, C.; Medina, S.; Morales-Guzmán, C.; Del Valle, R.G.; Carballeira, N.M.; Sanabria-Ríos, D.J. Antibacterial fatty acids: An update of possible mechanisms of action and implications in the development of the next-generation of antibacterial agents. Prog. Lipid Res. 2021, 82, 101093. [Google Scholar] [CrossRef]
- Das, U.N. Essential fatty acids: Biochemistry, physiology and pathology. Biotechnol. J. 2006, 1, 420–439. [Google Scholar] [CrossRef] [PubMed]
- Giamarellos-Bourboulis, E.J.; Mouktaroudi, M.; Adamis, T.; Koussoulas, V.; Baziaka, F.; Perrea, D.; Karayannacos, P.E.; Giamarellou, H. n-6 polyunsaturated fatty acids enhance the activities of ceftazidime and amikacin in experimental sepsis caused by multidrug-resistant Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2004, 48, 4713–4717. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.Q.; O’Connor, C.J.; Roberton, A.M. Antibacterial actions of fatty acids and monoglycerides against Helicobacter pylori. FEMS Immunol. Med. Microbiol. 2003, 36, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, R.; Huang, Y.-S. Gamma linolenic acid: An antiinflammatory omega-6 fatty acid. Curr. Pharm. Biotechnol. 2006, 7, 531–534. [Google Scholar] [CrossRef] [PubMed]
- Kawamura, A.; Ooyama, K.; Kojima, K.; Kachi, H.; Abe, T.; Amano, K.; Aoyama, T. Dietary supplementation of Gamma-linolenic acid improves skin parameters in subjects with dry skin and mild atopic dermatitis. J. Oleo Sci. 2011, 60, 597–607. [Google Scholar] [CrossRef]
- Kendall, A.C.; Kiezel-Tsugunova, M.; Brownbridge, L.C.; Harwood, J.L.; Nicolaou, A. Lipid functions in skin: Differential effects of n-3 polyunsaturated fatty acids on cutaneous ceramides, in a human skin organ culture model. Biochim. Biophys. Acta Biomembr. 2017, 1859, 1679–1689. [Google Scholar] [CrossRef]
- Thies, F.; Nebe-von-Caron, G.; Powell, J.R.; Yaqoob, P.; Newsholme, E.A.; Calder, P.C. Dietary supplementation with γ-linolenic acid or fish oil decreases T lymphocyte proliferation in healthy older humans. J. Nutr. 2001, 131, 1918–1927. [Google Scholar] [CrossRef]
- Yaqoob; Pala; Cortina-Borja; Newsholme; Calder. Encapsulated fish oil enriched in α-tocopherol alters plasma phospholipid and mononuclear cell fatty acid compositions but not mononuclear cell functions. Eur. J. Clin. Investig. 2000, 30, 260–274. [Google Scholar] [CrossRef]
- Innes, J.K.; Calder, P.C. Omega-6 fatty acids and inflammation. Prostaglandins Leukot. Essent. Fat. Acids 2018, 132, 41–48. [Google Scholar] [CrossRef]
- Hume, E.M.; Nunn, L.C.A.; Smedley-Maclean, I.; Smith, H.H. Studies of the essential unsaturated fatty acids in their relation to the fat-deficiency disease of rats. Biochem. J. 1938, 32, 2162–2177. [Google Scholar] [CrossRef]
- Rieckehoff, I.G.; Holman, R.T.; Burr, G.O. Polyethenoid Fatty Acid Metabolism. Effect of Dietary Fat on Polyethenoid Fatty Acids of Rat Tissues. Nutr. Rev. 1980, 38, 247–250. [Google Scholar] [CrossRef] [PubMed]
- Klenk, E.; Bongard, W. Die Konstitution der ungesättigten C20- und C22-Fettsäuren der Glycerinphosphatide des Gehirns [Constitution of the unsaturated C20 and C22 fatty acids of the glycophosphatides of the brain]. Biol. Chem. 1952, 291, 104–118. [Google Scholar] [CrossRef]
- Biran, L.A.; Bartley, W. Distribution of fatty acids in lipids of rat brain, brain mitochondria and microsomes. Biochem. J. 1961, 79, 159–176. [Google Scholar] [CrossRef]
- Benolken, R.M.; Anderson, R.E.; Wheeler, T.G. Membrane fatty acids associated with the electrical response in visual excitation. Science 1973, 182, 1253–1254. [Google Scholar] [CrossRef] [PubMed]
- Bieri, J.G.; Prival, E.L. Lipid composition of testes from various species. Comp. Biochem. Physiol. 1965, 15, 275–282. [Google Scholar] [CrossRef]
- Schnebelen-Berthier, C.; Acar, N.; Simon, E.; Thabuis, C.; Bourdillon, A.; Mathiaud, A.; Dauchet, L.; Delcourt, C.; Benlian, P.; Crochet, M.; et al. The ALGOVUE clinical trial: Effects of the daily consumption of eggs enriched with lutein and docosahexaenoic acid on plasma composition and macular pigment optical density. Nutrients 2021, 13, 3347. [Google Scholar] [CrossRef]
- Hishikawa, D.; Valentine, W.J.; Iizuka-Hishikawa, Y.; Shindou, H.; Shimizu, T. Metabolism and functions of docosahexaenoic acid-containing membrane glycerophospholipids. FEBS Lett. 2017, 591, 2730–2744. [Google Scholar] [CrossRef]
- Antonny, B.; Vanni, S.; Shindou, H.; Ferreira, T. From zero to six double bonds: Phospholipid unsaturation and organelle function. Trends Cell Biol. 2015, 25, 427–436. [Google Scholar] [CrossRef]
- Soubias, O.; Gawrisch, K. The role of the lipid matrix for structure and function of the GPCR rhodopsin. Biochim. Biophys. Acta Biomembr. 2012, 1818, 234–240. [Google Scholar] [CrossRef]
- Eckert, G.P.; Chang, S.; Eckmann, J.; Copanaki, E.; Hagl, S.; Hener, U.; Müller, W.E.; Kögel, D. Liposome-incorporated DHA increases neuronal survival by enhancing non-amyloidogenic APP processing. Biochim. Biophys. Acta Biomembr. 2011, 1808, 236–243. [Google Scholar] [CrossRef]
- Roqueta-Rivera, M.; Abbott, T.L.; Sivaguru, M.; Hess, R.A.; Nakamura, M.T. Deficiency in the omega-3 fatty acid pathway results in failure of acrosome biogenesis in mice. Biol. Reprod. 2011, 85, 721–732. [Google Scholar] [CrossRef] [PubMed]
- Sato, H.; Taketomi, Y.; Isogai, Y.; Miki, Y.; Yamamoto, K.; Masuda, S.; Hosono, T.; Arata, S.; Ishikawa, Y.; Ishii, T.; et al. Group III secreted phospholipase A2 regulates epididymal sperm maturation and fertility in mice. J. Clin. Investig. 2010, 120, 1400–1414. [Google Scholar] [CrossRef] [PubMed]
- Bennett, M.P.; Mitchell, D.C. Regulation of membrane proteins by dietary lipids: Effects of cholesterol and docosahexaenoic acid acyl chain-containing phospholipids on rhodopsin stability and function. Biophys. J. 2008, 95, 1206–1216. [Google Scholar] [CrossRef]
- Fedorova, I.; Hussein, N.; Di Martino, C.; Moriguchi, T.; Hoshiba, J.; Majchrzak, S.; Salem, N. An n-3 fatty acid deficient diet affects mouse spatial learning in the Barnes circular maze. Prostaglandins Leukot. Essent. Fat. Acids 2007, 77, 269–277. [Google Scholar] [CrossRef] [PubMed]
- Kawakita, E.; Hashimoto, M.; Shido, O. Docosahexaenoic acid promotes neurogenesis in vitro and in vivo. Neuroscience 2006, 139, 991–997. [Google Scholar] [CrossRef] [PubMed]
- Rejraji, H.; Sion, B.; Prensier, G.; Carreras, M.; Motta, C.; Frenoux, J.-M.; Vericel, E.; Grizard, G.; Vernet, P.; Drevet, J.R. Lipid remodeling of murine epididymosomes and spermatozoa during epididymal maturation1. Biol. Reprod. 2006, 74, 1104–1113. [Google Scholar] [CrossRef]
- Kitajka, K.; Puskás, L.G.; Zvara, Á.; Hackler, L.; Barceló-Coblijn, G.; Yeo, Y.K.; Farkas, T. The role of n-3 polyunsaturated fatty acids in brain: Modulation of rat brain gene expression by dietary n-3 fatty acids. Proc. Natl. Acad. Sci. USA 2002, 99, 2619–2624. [Google Scholar] [CrossRef]
- Litman, B.J.; Mitchell, D.C. A role for phospholipid polyunsaturation in modulating membrane protein function. Lipids 1996, 31, S193–S197. [Google Scholar] [CrossRef]
- Miljanich, G.P.; Sklar, L.A.; White, D.L.; Dratz, E.A. Disaturated and dipolyunsaturated phospholipids in the bovine retinal rod outer segment disk membrane. Biochim. Biophys. Acta Biomembr. 1979, 552, 294–306. [Google Scholar] [CrossRef]
- Ding, L.; Zhang, T.; Che, H.; Zhang, L.; Xue, C.; Chang, Y.; Wang, Y. DHA-enriched phosphatidylcholine and DHA-enriched phosphatidylserine improve age-related lipid metabolic disorder through different metabolism in the senescence-accelerated mouse. Eur. J. Lipid Sci. Technol. 2018, 120, 1700490. [Google Scholar] [CrossRef]
- Shefer-Weinberg, D.; Sasson, S.; Schwartz, B.; Argov-Argaman, N.; Tirosh, O. Deleterious effect of n-3 polyunsaturated fatty acids in non-alcoholic steatohepatitis in the fat-1 mouse model. Clin. Nutr. Exp. 2017, 12, 37–49. [Google Scholar] [CrossRef]
- Epand, R.M.; D’Souza, K.; Berno, B.; Schlame, M. Membrane curvature modulation of protein activity determined by NMR. Biochim. Biophys. Acta Biomembr. 2015, 1848, 220–228. [Google Scholar] [CrossRef] [PubMed]
- Andersen, O.S.; Koeppe, R.E. Bilayer thickness and membrane protein function: An energetic perspective. Annu. Rev. Biophys. Biomol. Struct. 2007, 36, 107–130. [Google Scholar] [CrossRef] [PubMed]
- Gruner, S.M. Intrinsic curvature hypothesis for biomembrane lipid composition: A role for nonbilayer lipids. Proc. Natl. Acad. Sci. USA 1985, 82, 3665–3669. [Google Scholar] [CrossRef] [PubMed]
- Johannsson, A.; Smith, G.A.; Metcalfe, J.C. The effect of bilayer thickness on the activity of (Na+ + K+)-ATPase. Biochim. Biophys. Acta Biomembr. 1981, 641, 416–421. [Google Scholar] [CrossRef]
- Jalili, M.; Hekmatdoost, A. Dietary ω-3 fatty acids and their influence on inflammation via Toll-like receptor pathways. Nutrition 2021, 85, 111070. [Google Scholar] [CrossRef]
- Cordero-Morales, J.F.; Vásquez, V. How lipids contribute to ion channel function, a fat perspective on direct and indirect interactions. Curr. Opin. Struct. Biol. 2018, 51, 92–98. [Google Scholar] [CrossRef]
- Elsherbiny, M.; Chen, H.; Emara, M.; Godbout, R. ω-3 and ω-6 fatty acids modulate conventional and atypical protein kinase C activities in a brain fatty acid binding protein dependent manner in glioblastoma multiforme. Nutrients 2018, 10, 454. [Google Scholar] [CrossRef]
- Senapati, S.; Gragg, M.; Samuels, I.S.; Parmar, V.M.; Maeda, A.; Park, P.S.-H. Effect of dietary docosahexaenoic acid on rhodopsin content and packing in photoreceptor cell membranes. Biochim. Biophys. Acta Biomembr. 2018, 1860, 1403–1413. [Google Scholar] [CrossRef]
- Esposito, G.; Scuderi, C.; Valenza, M.; Togna, G.I.; Latina, V.; De Filippis, D.; Cipriano, M.; Carratù, M.R.; Iuvone, T.; Steardo, L. Cannabidiol reduces Aβ-induced neuroinflammation and promotes hippocampal neurogenesis through PPARγ involvement. PLoS ONE 2011, 6, e28668. [Google Scholar] [CrossRef]
- Vinayavekhin, N.; Saghatelian, A. Discovery of a protein–metabolite interaction between unsaturated fatty acids and the nuclear receptor Nur77 using a metabolomics approach. J. Am. Chem. Soc. 2011, 133, 17168–17171. [Google Scholar] [CrossRef] [PubMed]
- Akhtar Khan, N. Polyunsaturated fatty acids in the modulation of T-cell signalling. Prostaglandins Leukot. Essent. Fat. Acids 2010, 82, 179–187. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Zhang, M.; Larsen, C.P.; Xu, F.; Hua, W.; Yamashima, T.; Mao, Y.; Zhou, L. DHA promotes the neuronal differentiation of rat neural stem cells transfected with GPR40 gene. Brain Res. 2010, 1330, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Grossfield, A.; Feller, S.E.; Pitman, M.C. Contribution of omega-3 fatty acids to the thermodynamics of membrane protein solvation. J. Phys. Chem. B 2006, 110, 8907–8909. [Google Scholar] [CrossRef]
- Kang, J.X.; Leaf, A. Evidence that free polyunsaturated fatty acids modify Na+ channels by directly binding to the channel proteins. Proc. Natl. Acad. Sci. USA 1996, 93, 3542–3546. [Google Scholar] [CrossRef]
- Mosior, M.; Golini, E.S.; Epand, R.M. Chemical specificity and physical properties of the lipid bilayer in the regulation of protein kinase C by anionic phospholipids: Evidence for the lack of a specific binding site for phosphatidylserine. Proc. Natl. Acad. Sci. USA 1996, 93, 1907–1912. [Google Scholar] [CrossRef]
- Pepe, S.; Bogdanov, K.; Hallaq, H.; Spurgeon, H.; Leaf, A.; Lakatta, E. Omega 3 polyunsaturated fatty acid modulates dihydropyridine effects on L-type Ca2+ channels, cytosolic Ca2+, and contraction in adult rat cardiac myocytes. Proc. Natl. Acad. Sci. USA 1994, 91, 8832–8836. [Google Scholar] [CrossRef]
- Xiao, Y.-F.; Gomez, A.M.; Morgan, J.P.; Lederer, W.J.; Leaf, A. Suppression of voltage-gated L-type Ca2+ currents by polyunsaturated fatty acids in adult and neonatal rat ventricular myocytes. Proc. Natl. Acad. Sci. USA 1997, 94, 4182–4187. [Google Scholar] [CrossRef]
- Nanji, A.A.; Griniuviene, B.; Sadrzadeh, S.M.; Levitsky, S.; McCully, J.D. Effect of type of dietary fat and ethanol on antioxidant enzyme mRNA induction in rat liver. J. Lipid Res. 1995, 36, 736–744. [Google Scholar] [CrossRef]
- Casañas-Sánchez, V.; Pérez, J.A.; Fabelo, N.; Quinto-Alemany, D.; Díaz, M.L. Docosahexaenoic (DHA) modulates phospholipid-hydroperoxide glutathione peroxidase (Gpx4) gene expression to ensure self-protection from oxidative damage in hippocampal cells. Front. Physiol. 2015, 6, 203. [Google Scholar] [CrossRef]
- Hossain, M.S.; Hashimoto, M.; Gamoh, S.; Masumura, S. Antioxidative effects of docosahexaenoic acid in the cerebrum versus cerebellum and brainstem of aged hypercholesterolemic rats. J. Neurochem. 2008, 72, 1133–1138. [Google Scholar] [CrossRef]
- Roqueta-Rivera, M.; Stroud, C.K.; Haschek, W.M.; Akare, S.J.; Segre, M.; Brush, R.S.; Agbaga, M.-P.; Anderson, R.E.; Hess, R.A.; Nakamura, M.T. Docosahexaenoic acid supplementation fully restores fertility and spermatogenesis in male delta-6 desaturase-null mice. J. Lipid Res. 2010, 51, 360–367. [Google Scholar] [CrossRef]
- Hernández-Rodas, M.C.; Valenzuela, R.; Echeverría, F.; Rincón-Cervera, M.Á.; Espinosa, A.; Illesca, P.; Muñoz, P.; Corbari, A.; Romero, N.; Gonzalez-Mañan, D.; et al. Supplementation with docosahexaenoic acid and extra virgin olive oil prevents liver steatosis induced by a high-fat diet in mice through PPAR-α and Nrf2 upregulation with concomitant SREBP-1c and NF-kB downregulation. Mol. Nutr. Food Res. 2017, 61, 1700479. [Google Scholar] [CrossRef]
- Allam-Ndoul, B.; Guénard, F.; Barbier, O.; Vohl, M.-C. Effect of n-3 fatty acids on the expression of inflammatory genes in THP-1 macrophages. Lipids Health Dis. 2016, 15, 69. [Google Scholar] [CrossRef]
- Mullen, A.; Loscher, C.E.; Roche, H.M. Anti-inflammatory effects of EPA and DHA are dependent upon time and dose-response elements associated with LPS stimulation in THP-1-derived macrophages. J. Nutr. Biochem. 2010, 21, 444–450. [Google Scholar] [CrossRef]
- Bondy, S.C.; Guo, S.X. Effect of ethanol treatment on indices of cumulative oxidative stress. Eur. J. Pharmacol. Environ. Toxicol. Pharmacol. 1994, 270, 349–355. [Google Scholar] [CrossRef]
- Camandola, S.; Leonarduzzi, G.; Musso, T.; Varesio, L.; Carini, R.; Scavazza, A.; Chiarpotto, E.; Baeuerle, P.A.; Poli, G. Nuclear factor kB is activated by arachidonic acid but not by eicosapentaenoic Acid. Biochem. Biophys. Res. Commun. 1996, 229, 643–647. [Google Scholar] [CrossRef]
- Jayadev, S.; Linardic, C.M.; Hannun, Y.A. Identification of arachidonic acid as a mediator of sphingomyelin hydrolysis in response to tumor necrosis factor alpha. J. Biol. Chem. 1994, 269, 5757–5763. [Google Scholar] [CrossRef]
- Hashidate-Yoshida, T.; Harayama, T.; Hishikawa, D.; Morimoto, R.; Hamano, F.; Tokuoka, S.M.; Eto, M.; Tamura-Nakano, M.; Yanobu-Takanashi, R.; Mukumoto, Y.; et al. Fatty acid remodeling by LPCAT3 enriches arachidonate in phospholipid membranes and regulates triglyceride transport. eLife 2015, 4, e06328. [Google Scholar] [CrossRef]
- Heshmati, J. Effect of omega-3 fatty acid supplementation on gene expression of inflammation, oxidative stress and cardiometabolic parameters: Systematic review and meta-analysis. J. Funct. Foods 2021, 85, 104619. [Google Scholar] [CrossRef]
- Baker, P.R.S.; Lin, Y.; Schopfer, F.J.; Woodcock, S.R.; Groeger, A.L.; Batthyany, C.; Sweeney, S.; Long, M.H.; Iles, K.E.; Baker, L.M.S.; et al. Fatty acid transduction of nitric oxide signaling. J. Biol. Chem. 2005, 280, 42464–42475. [Google Scholar] [CrossRef]
- Lima, É.S.; Bonini, M.G.; Augusto, O.; Barbeiro, H.V.; Souza, H.P.; Abdalla, D.S.P. Nitrated lipids decompose to nitric oxide and lipid radicals and cause vasorelaxation. Free Radic. Biol. Med. 2005, 39, 532–539. [Google Scholar] [CrossRef]
- Coles, B.; Bloodsworth, A.; Clark, S.R.; Lewis, M.J.; Cross, A.R.; Freeman, B.A.; O’Donnell, V.B. Nitrolinoleate inhibits superoxide generation, degranulation, and integrin expression by human neutrophils. Circ. Res. 2002, 91, 375–381. [Google Scholar] [CrossRef]
- von Euler, U.S. On the specific vaso-dilating and plain muscle stimulating substances from accessory genital glands in man and certain animals (prostaglandin and vesiglandin). J. Physiol. 1936, 88, 213–234. [Google Scholar] [CrossRef]
- Tang, X.; Edwards, E.M.; Holmes, B.B.; Falck, J.R.; Campbell, W.B. Role of phospholipase C and diacylglyceride lipase pathway in arachidonic acid release and acetylcholine-induced vascular relaxation in rabbit aorta. Am. J. Physiol. Circ. Physiol. 2006, 290, H37–H45. [Google Scholar] [CrossRef]
- Abraham, R.T.; McKinney, M.M.; Forray, C.; Shipley, G.D.; Handwerger, B.S. Stimulation of arachidonic acid release and eicosanoid biosynthesis in an interleukin 2-dependent T cell Line. Immunopharmacol. Immunotoxicol. 1986, 8, 165–204. [Google Scholar] [CrossRef]
- Yada, Y.; Higuchi, K.; Imokawa, G. Purification and biochemical characterization of membrane-bound epidermal ceramidases from Guinea pig skin. J. Biol. Chem. 1995, 270, 12677–12684. [Google Scholar] [CrossRef]
- Imokawa, G. A possible mechanism underlying the ceramide deficiency in atopic dermatitis: Expression of a deacylase enzyme that cleaves the N-acyl linkage of sphingomyelin and glucosylceramide. J. Dermatol. Sci. 2009, 55, 1–9. [Google Scholar] [CrossRef]
- Ishibashi, M.; Arikawa, J.; Okamoto, R.; Kawashima, M.; Takagi, Y.; Ohguchi, K.; Imokawa, G. Abnormal expression of the novel epidermal enzyme, glucosylceramide deacylase, and the accumulation of its enzymatic reaction product, glucosylsphingosine, in the skin of patients with atopic dermatitis. Lab. Investig. 2003, 83, 397–408. [Google Scholar] [CrossRef]
- Imokawa, G.; Takagi, Y.; Higuchi, K.; Kondo, H.; Yada, Y. Sphingosylphosphorylcholine is a potent inducer of intercellular adhesion molecule-1 expression in human keratinocytes. J. Investig. Dermatol. 1999, 112, 91–96. [Google Scholar] [CrossRef]
- Richmond, G.S.; Smith, T.K. Phospholipases A1. Int. J. Mol. Sci. 2011, 12, 588–612. [Google Scholar] [CrossRef]
- Goñi, F.M.; Montes, L.-R.; Alonso, A. Phospholipases C and sphingomyelinases: Lipids as substrates and modulators of enzyme activity. Prog. Lipid Res. 2012, 51, 238–266. [Google Scholar] [CrossRef]
- Yang, S.F.; Freer, S.; Benson, A.A. Transphosphatidylation by phospholipase D. J. Biol. Chem. 1967, 242, 477–484. [Google Scholar] [CrossRef]
- Dennis, E.A.; Cao, J.; Hsu, Y.-H.; Magrioti, V.; Kokotos, G. Phospholipase A 2 enzymes: Physical structure, biological function, disease implication, chemical inhibition, and therapeutic intervention. Chem. Rev. 2011, 111, 6130–6185. [Google Scholar] [CrossRef]
- Clark, J.D.; Lin, L.-L.; Kriz, R.W.; Ramesha, C.S.; Sultzman, L.A.; Lin, A.Y.; Milona, N.; Knopf, J.L. A novel arachidonic acid-selective cytosolic PLA2 contains a Ca2+-dependent translocation domain with homology to PKC and GAP. Cell 1991, 65, 1043–1051. [Google Scholar] [CrossRef]
- Cheon, Y.; Kim, H.-W.; Igarashi, M.; Modi, H.R.; Chang, L.; Ma, K.; Greenstein, D.; Wohltmann, M.; Turk, J.; Rapoport, S.I.; et al. Disturbed brain phospholipid and docosahexaenoic acid metabolism in calcium-independent phospholipase A2-VIA (iPLA2β)-knockout mice. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2012, 1821, 1278–1286. [Google Scholar] [CrossRef]
- Strokin, M.; Sergeeva, M.; Reiser, G. Docosahexaenoic acid and arachidonic acid release in rat brain astrocytes is mediated by two separate isoforms of phospholipase A 2 and is differently regulated by cyclic AMP and Ca2+. Br. J. Pharmacol. 2003, 139, 1014–1022. [Google Scholar] [CrossRef]
- Murase, R.; Sato, H.; Yamamoto, K.; Ushida, A.; Nishito, Y.; Ikeda, K.; Kobayashi, T.; Yamamoto, T.; Taketomi, Y.; Murakami, M. Group X secreted phospholipase A2 releases ω3 polyunsaturated fatty acids, suppresses colitis, and promotes sperm fertility. J. Biol. Chem. 2016, 291, 6895–6911. [Google Scholar] [CrossRef]
- Fonteh, A.N.; Bass, D.A.; Marshall, L.A.; Seeds, M.; Samet, J.M.; Chilton, F.H. Evidence that secretory phospholipase A2 plays a role in arachidonic acid release and eicosanoid biosynthesis by mast cells. J. Immunol. 1994, 152, 5438–5446. [Google Scholar] [CrossRef]
- Wada, M.; DeLong, C.J.; Hong, Y.H.; Rieke, C.J.; Song, I.; Sidhu, R.S.; Yuan, C.; Warnock, M.; Schmaier, A.H.; Yokoyama, C.; et al. Enzymes and receptors of prostaglandin pathways with arachidonic acid-derived versus eicosapentaenoic acid-derived substrates and products. J. Biol. Chem. 2007, 282, 22254–22266. [Google Scholar] [CrossRef]
- Chan, J.K.; McDonald, B.E.; Gerrard, J.M.; Bruce, V.M.; Weaver, B.J.; Holub, B.J. Effect of dietary α-linolenic acid and its ratio to linoleic acid on platelet and plasma fatty acids and thrombogenesis. Lipids 1993, 28, 811–817. [Google Scholar] [CrossRef]
- Luu, W.; Sharpe, L.J.; Capell-Hattam, I.; Gelissen, I.C.; Brown, A.J. Oxysterols: Old tale, new twists. Annu. Rev. Pharmacol. Toxicol. 2016, 56, 447–467. [Google Scholar] [CrossRef]
- Schroepfer, G.J. Oxysterols: Modulators of cholesterol metabolism and other processes. Physiol. Rev. 2000, 80, 361–554. [Google Scholar] [CrossRef]
- Phillis, J.W.; Horrocks, L.A.; Farooqui, A.A. Cyclooxygenases, lipoxygenases, and epoxygenases in CNS: Their role and involvement in neurological disorders. Brain Res. Rev. 2006, 52, 201–243. [Google Scholar] [CrossRef]
- Bazan, N.G.; Fletcher, B.S.; Herschman, H.R.; Mukherjee, P.K. Platelet-activating factor and retinoic acid synergistically activate the inducible prostaglandin synthase gene. Proc. Natl. Acad. Sci. USA 1994, 91, 5252–5256. [Google Scholar] [CrossRef]
- Chandrasekharan, N.V.; Dai, H.; Roos, K.L.T.; Evanson, N.K.; Tomsik, J.; Elton, T.S.; Simmons, D.L. COX-3, a cyclooxygenase-1 variant inhibited by acetaminophen and other analgesic/antipyretic drugs: Cloning, structure, and expression. Proc. Natl. Acad. Sci. USA 2002, 99, 13926–13931. [Google Scholar] [CrossRef]
- Simard, M.; Morin, S.; Ridha, Z.; Pouliot, R. Current knowledge of the implication of lipid mediators in psoriasis. Front. Immunol. 2022, 13, 961107. [Google Scholar] [CrossRef]
- Alhouayek, M.; Muccioli, G.G. COX-2-derived endocannabinoid metabolites as novel inflammatory mediators. Trends Pharmacol. Sci. 2014, 35, 284–292. [Google Scholar] [CrossRef]
- Ivanov, I.; Kuhn, H.; Heydeck, D. Structural and functional biology of arachidonic acid 15-lipoxygenase-1 (ALOX15). Gene 2015, 573, 1–32. [Google Scholar] [CrossRef]
- Lehnert, N.; Solomon, E.I. Density-functional investigation on the mechanism of H-atom abstraction by lipoxygenase. JBIC J. Biol. Inorg. Chem. 2003, 8, 294–305. [Google Scholar] [CrossRef]
- Dyall, S.C.; Balas, L.; Bazan, N.G.; Brenna, J.T.; Chiang, N.; da Costa Souza, F.; Dalli, J.; Durand, T.; Galano, J.-M.; Lein, P.J.; et al. Polyunsaturated fatty acids and fatty acid-derived lipid mediators: Recent advances in the understanding of their biosynthesis, structures, and functions. Prog. Lipid Res. 2022, 86, 101165. [Google Scholar] [CrossRef] [PubMed]
- Hajeyah, A.A.; Griffiths, W.J.; Wang, Y.; Finch, A.J.; O’Donnell, V.B. The biosynthesis of enzymatically oxidized lipids. Front. Endocrinol. 2020, 11, 591819. [Google Scholar] [CrossRef] [PubMed]
- Nebert, D.W.; Wikvall, K.; Miller, W.L. Human cytochromes P450 in health and disease. Philos. Trans. R. Soc. B Biol. Sci. 2013, 368, 20120431. [Google Scholar] [CrossRef]
- Davis, C.M.; Liu, X.; Alkayed, N.J. Cytochrome P450 eicosanoids in cerebrovascular function and disease. Pharmacol. Ther. 2017, 179, 31–46. [Google Scholar] [CrossRef]
- Lamb, D.C.; Waterman, M.R. Unusual properties of the cytochrome P450 superfamily. Philos. Trans. R. Soc. B Biol. Sci. 2013, 368, 20120434. [Google Scholar] [CrossRef]
- Isin, E.M.; Guengerich, F.P. Complex reactions catalyzed by cytochrome P450 enzymes. Biochim. Biophys. Acta Gen. Subj. 2007, 1770, 314–329. [Google Scholar] [CrossRef]
- Kumar, N.; Gupta, G.; Anilkumar, K.; Fatima, N.; Karnati, R.; Reddy, G.V.; Giri, P.V.; Reddanna, P. 15-Lipoxygenase metabolites of α-linolenic acid, [13-(S)-HPOTrE and 13-(S)-HOTrE], mediate anti-inflammatory effects by inactivating NLRP3 inflammasome. Sci. Rep. 2016, 6, 31649. [Google Scholar] [CrossRef]
- Budowski, P.; Bartov, I.; Dror, Y.; Frankel, E.N. Lipid oxidation products and chick nutritional encephalopathy. Lipids 1979, 14, 768–772. [Google Scholar] [CrossRef]
- Buchanan, M.R.; Haas, T.A.; Lagarde, M.; Guichardant, M. 13-Hydroxyoctadecadienoic acid is the vessel wall chemorepellant factor, LOX. J. Biol. Chem. 1985, 260, 16056–16059. [Google Scholar] [CrossRef]
- Shearer, G.C.; Walker, R.E. An overview of the biologic effects of omega-6 oxylipins in humans. Prostaglandins Leukot. Essent. Fat. Acids 2018, 137, 26–38. [Google Scholar] [CrossRef]
- Vangaveti, V.N.; Jansen, H.; Kennedy, R.L.; Malabu, U.H. Hydroxyoctadecadienoic acids: Oxidised derivatives of linoleic acid and their role in inflammation associated with metabolic syndrome and cancer. Eur. J. Pharmacol. 2016, 785, 70–76. [Google Scholar] [CrossRef] [PubMed]
- Moghaddam, M.F.; Grant, D.F.; Cheek, J.M.; Greene, J.F.; Williamson, K.C.; Hammock, B.D. Bioactivation of leukotoxins to their toxic diols by epoxide hydrolase. Nat. Med. 1997, 3, 562–566. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, J.J.; Domenichiello, A.F.; Jensen, J.R.; Keyes, G.S.; Maiden, K.M.; Davis, J.M.; Ramsden, C.E.; Mishra, S.K. Endogenous derivatives of linoleic acid and their stable analogs are potential pain mediators. JID Innov. 2023, 3, 100177. [Google Scholar] [CrossRef] [PubMed]
- Alsalem, M.; Wong, A.; Millns, P.; Arya, P.H.; Chan, M.S.L.; Bennett, A.; Barrett, D.A.; Chapman, V.; Kendall, D.A. The contribution of the endogenous TRPV1 ligands 9-HODE and 13-HODE to nociceptive processing and their role in peripheral inflammatory pain mechanisms. Br. J. Pharmacol. 2013, 168, 1961–1974. [Google Scholar] [CrossRef] [PubMed]
- Biswas, P.; Datta, C.; Rathi, P.; Bhattacharjee, A. Fatty acids and their lipid mediators in the induction of cellular apoptosis in cancer cells. Prostaglandins Other Lipid Mediat. 2022, 160, 106637. [Google Scholar] [CrossRef]
- Johnson, G.H.; Fritsche, K. Effect of dietary linoleic acid on markers of inflammation in healthy persons: A systematic review of randomized controlled trials. J. Acad. Nutr. Diet. 2012, 112, 1029–1041.e15. [Google Scholar] [CrossRef]
- Takic, M.; Pokimica, B.; Petrovic-Oggiano, G.; Popovic, T. Effects of Dietary α-Linolenic Acid Treatment and the Efficiency of Its Conversion to Eicosapentaenoic and Docosahexaenoic Acids in Obesity and Related Diseases. Molecules 2022, 27, 4471. [Google Scholar] [CrossRef]
- Arterburn, L.M.; Hall, E.B.; Oken, H. Distribution, interconversion, and dose response of n-3 fatty acids in humans. Am. J. Clin. Nutr. 2006, 83, 1467S–1476S. [Google Scholar] [CrossRef]
- Shimizu, T. Lipid mediators in health and disease: Enzymes and receptors as therapeutic targets for the regulation of immunity and inflammation. Annu. Rev. Pharmacol. Toxicol. 2009, 49, 123–150. [Google Scholar] [CrossRef]
- Sergeant, S.; Rahbar, E.; Chilton, F.H. Gamma-linolenic acid, Dihommo-gamma linolenic, Eicosanoids and Inflammatory Processes. Eur. J. Pharmacol. 2016, 785, 77–86. [Google Scholar] [CrossRef]
- Miller, C.C.; McCreedy, C.A.; Jones, A.D.; Ziboh, V.A. Oxidative metabolism of dihomogammalinolenic acid by guinea pig epidermis: Evidence of generation of anti-inflammatory products. Prostaglandins 1988, 35, 917–938. [Google Scholar] [CrossRef]
- Samuelsson, B. From studies of biochemical mechanism to novel biological mediators: Prostaglandin endoperoxides, thromboxanes, and leukotriences. Biosci. Rep. 1983, 3, 791–813. [Google Scholar] [CrossRef] [PubMed]
- Buczynski, M.W.; Dumlao, D.S.; Dennis, E.A. Thematic review series: Proteomics. An integrated omics analysis of eicosanoid biology. J. Lipid Res. 2009, 50, 1015–1038. [Google Scholar] [CrossRef]
- Dalvi, S.; Nguyen, H.H.; On, N.; Mitchell, R.W.; Aukema, H.M.; Miller, D.W.; Hatch, G.M. Exogenous arachidonic acid mediates permeability of human brain microvessel endothelial cells through prostaglandin E 2 activation of EP 3 and EP 4 receptors. J. Neurochem. 2015, 135, 867–879. [Google Scholar] [CrossRef]
- Patel, P.; Cossette, C.; Anumolu, J.R.; Gravel, S.; Lesimple, A.; Mamer, O.A.; Rokach, J.; Powell, W.S. Structural requirements for activation of the 5-Oxo-6E, 8Z, 11Z, 14Z-eicosatetraenoic acid (5-Oxo-ETE) receptor: Identification of a Mead acid metabolite with potent agonist activity. J. Pharmacol. Exp. Ther. 2008, 325, 698–707. [Google Scholar] [CrossRef] [PubMed]
- Balestrieri, B.; Di Costanzo, D.; Dwyer, D.F. Macrophage-Mediated Immune Responses: From Fatty Acids to Oxylipins. Molecules 2021, 27, 152. [Google Scholar] [CrossRef]
- Farag, M.A.; Gad, M.Z. Omega-9 fatty acids: Potential roles in inflammation and cancer management. J. Genet. Eng. Biotechnol. 2022, 20, 48. [Google Scholar] [CrossRef]
- Hamazaki, T.; Hamazaki, K. What Are the Physiological Roles of Mead Acid (5,8,11-Eicosatrienoic Acid)? In Handbook of Lipids in Human Function; Elsevier: Amsterdam, The Netherlands, 2016; pp. 483–497. [Google Scholar]
- Duvall, M.G.; Levy, B.D. DHA- and EPA-derived resolvins, protectins, and maresins in airway inflammation. Eur. J. Pharmacol. 2016, 785, 144–155. [Google Scholar] [CrossRef]
- Bergström, S.; Danielsson, H.; Klenberg, D.; Samuelsson, B. The enzymatic conversion of essential fatty acids into prostaglandins. J. Biol. Chem. 1964, 239, PC4006–PC4008. [Google Scholar] [CrossRef]
- Dyerberg, J. Eicosapentaenoic acid and prevention of thrombosis and atherosclerosis? Lancet 1978, 312, 117–119. [Google Scholar] [CrossRef]
- Levy, B.D.; Clish, C.B.; Schmidt, B.; Gronert, K.; Serhan, C.N. Lipid mediator class switching during acute inflammation: Signals in resolution. Nat. Immunol. 2001, 2, 612–619. [Google Scholar] [CrossRef] [PubMed]
- Schunck, W.-H.; Konkel, A.; Fischer, R.; Weylandt, K.-H. Therapeutic potential of omega-3 fatty acid-derived epoxyeicosanoids in cardiovascular and inflammatory diseases. Pharmacol. Ther. 2018, 183, 177–204. [Google Scholar] [CrossRef] [PubMed]
- Calder, P.C.; Yaqoob, P.; Thies, F.; Wallace, F.A.; Miles, E.A. Fatty acids and lymphocyte functions. Br. J. Nutr. 2002, 87, S31–S48. [Google Scholar] [CrossRef] [PubMed]
- Mustonen, A.-M.; Nieminen, P. Dihomo-γ-linolenic acid (20:3n-6)—Metabolism, derivatives, and potential significance in chronic inflammation. Int. J. Mol. Sci. 2023, 24, 2116. [Google Scholar] [CrossRef]
- Baker, E.J.; Valenzuela, C.A.; Dooremalen, W.T.M.; Martínez-Fernández, L.; Yaqoob, P.; Miles, E.A.; Calder, P.C. Gamma-linolenic and pinolenic acids exert anti-inflammatory effects in cultured human endothelial cells through their elongation products. Mol. Nutr. Food Res. 2020, 64, 2000382. [Google Scholar] [CrossRef]
- Brouwers, H.; Jónasdóttir, H.S.; Kuipers, M.E.; Kwekkeboom, J.C.; Auger, J.L.; Gonzalez-Torres, M.; López-Vicario, C.; Clària, J.; Freysdottir, J.; Hardardottir, I.; et al. Anti-inflammatory and proresolving effects of the omega-6 polyunsaturated fatty acid adrenic acid. J. Immunol. 2020, 205, 2840–2849. [Google Scholar] [CrossRef]
- Poorani, R.; Bhatt, A.; Das, U. Modulation of pro-inflammatory and pro-resolution mediators by γ-linolenic acid: An important element in radioprotection against ionizing radiation. Arch. Med. Sci. 2020, 16, 1448–1456. [Google Scholar]
- Serhan, C.N.; Levy, B.D. Resolvins in inflammation: Emergence of the pro-resolving superfamily of mediators. J. Clin. Investig. 2018, 128, 2657–2669. [Google Scholar] [CrossRef]
- Farooqui, A.A.; Horrocks, L.A.; Farooqui, T. Modulation of inflammation in brain: A matter of fat. J. Neurochem. 2006, 101, 577–599. [Google Scholar] [CrossRef]
- Chiang, N.; Gronert, K.; Clish, C.B.; O’Brien, J.A.; Freeman, M.W.; Serhan, C.N. Leukotriene B4 receptor transgenic mice reveal novel protective roles for lipoxins and aspirin-triggered lipoxins in reperfusion. J. Clin. Investig. 1999, 104, 309–316. [Google Scholar] [CrossRef]
- Clària, J.; Serhan, C.N. Aspirin triggers previously undescribed bioactive eicosanoids by human endothelial cell-leukocyte interactions. Proc. Natl. Acad. Sci. USA 1995, 92, 9475–9479. [Google Scholar] [CrossRef] [PubMed]
- Weylandt, K.-H. Docosapentaenoic acid derived metabolites and mediators—The new world of lipid mediator medicine in a nutshell. Eur. J. Pharmacol. 2016, 785, 108–115. [Google Scholar] [CrossRef] [PubMed]
- Serhan, C.N.; Gupta, S.S.S.K.; Perretti, M.; Godson, C.; Brennan, E.; Li, Y.; Soehnlein, O.; Shimizu, T.; Werz, O.; Chiurchiù, V.; et al. The atlas of inflammation resolution (AIR). Mol. Asp. Med. 2020, 74, 100894. [Google Scholar] [CrossRef]
- Drouin, G.; Rioux, V.; Legrand, P. The n-3 docosapentaenoic acid (DPA): A new player in the n-3 long chain polyunsaturated fatty acid family. Biochimie 2019, 159, 36–48. [Google Scholar] [CrossRef]
- Serhan, C.N.; Dalli, J.; Colas, R.A.; Winkler, J.W.; Chiang, N. Protectins and maresins: New pro-resolving families of mediators in acute inflammation and resolution bioactive metabolome. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2015, 1851, 397–413. [Google Scholar] [CrossRef]
- Terranova, L.; Risé, P.; Gramegna, A.; Pinna, C.; Agostoni, C.; Syrén, M.-L.; Turolo, S.; Marchisio, P.; Amati, F.; Aliberti, S.; et al. Pro-resolving and pro-inflammatory fatty acid-derived mediators in sputum of stable state bronchiectasis patients. Respir. Res. 2022, 23, 363. [Google Scholar] [CrossRef]
- Tang, S.; Wan, M.; Huang, W.; Stanton, R.C.; Xu, Y. Maresins: Specialized proresolving lipid mediators and their potential role in inflammatory-related diseases. Mediat. Inflamm. 2018, 2018, 2380319. [Google Scholar] [CrossRef]
- Balas, L.; Durand, T. Dihydroxylated E,E,Z-docosatrienes. An overview of their synthesis and biological significance. Prog. Lipid Res. 2016, 61, 1–18. [Google Scholar] [CrossRef]
- Serhan, C.N.; Chiang, N.; Van Dyke, T.E. Resolving inflammation: Dual anti-inflammatory and pro-resolution lipid mediators. Nat. Rev. Immunol. 2008, 8, 349–361. [Google Scholar] [CrossRef]
- Serhan, C.N.; Arita, M.; Hong, S.; Gotlinger, K. Resolvins, docosatrienes, and neuroprotectins, novel omega-3-derived mediators, and their endogenous aspirin-triggered epimers. Lipids 2004, 39, 1125–1132. [Google Scholar] [CrossRef]
- Wu, A.; Ying, Z.; Gomez-Pinilla, F. Dietary omega-3 fatty acids normalize BDNF levels, reduce oxidative damage, and counteract learning disability after traumatic brain injury in rats. J. Neurotrauma 2004, 21, 1457–1467. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.; Gronert, K.; Devchand, P.R.; Moussignac, R.-L.; Serhan, C.N. Novel docosatrienes and 17S-resolvins generated from docosahexaenoic acid in murine brain, human blood, and glial cells. J. Biol. Chem. 2003, 278, 14677–14687. [Google Scholar] [CrossRef] [PubMed]
- Marcheselli, V.L.; Hong, S.; Lukiw, W.J.; Tian, X.H.; Gronert, K.; Musto, A.; Hardy, M.; Gimenez, J.M.; Chiang, N.; Serhan, C.N.; et al. Novel docosanoids inhibit brain ischemia-reperfusion-mediated leukocyte infiltration and pro-inflammatory gene expression. J. Biol. Chem. 2003, 278, 43807–43817. [Google Scholar] [CrossRef]
- Serhan, C.N.; Hong, S.; Gronert, K.; Colgan, S.P.; Devchand, P.R.; Mirick, G.; Moussignac, R.-L. Resolvins. J. Exp. Med. 2002, 196, 1025–1037. [Google Scholar] [CrossRef] [PubMed]
- Li, X.-A.; Everson, W.V.; Smart, E.J. Caveolae, lipid rafts, and vascular disease. Trends Cardiovasc. Med. 2005, 15, 92–96. [Google Scholar] [CrossRef]
- Dalli, J.; Colas, R.A.; Serhan, C.N. Novel n-3 Immunoresolvents: Structures and Actions. Sci. Rep. 2013, 3, 1940. [Google Scholar] [CrossRef]
- Jun, B.; Mukherjee, P.K.; Asatryan, A.; Kautzmann, M.-A.; Heap, J.; Gordon, W.C.; Bhattacharjee, S.; Yang, R.; Petasis, N.A.; Bazan, N.G. Elovanoids are novel cell-specific lipid mediators necessary for neuroprotective signaling for photoreceptor cell integrity. Sci. Rep. 2017, 7, 5279. [Google Scholar] [CrossRef]
- Kyselová, L.; Vítová, M.; Řezanka, T. Very long chain fatty acids. Prog. Lipid Res. 2022, 87, 101180. [Google Scholar] [CrossRef]
- Asatryan, A.; Calandria, J.M.; Kautzmann, M.-A.I.; Jun, B.; Gordon, W.C.; Do, K.V.; Bhattacharjee, S.; Pham, T.L.; Bermúdez, V.; Mateos, M.V.; et al. New retinal pigment epithelial cell model to unravel neuroprotection sensors of neurodegeneration in retinal disease. Front. Neurosci. 2022, 16, 3894. [Google Scholar] [CrossRef]
- Bazan, N.G. Overview of how N32 and N34 elovanoids sustain sight by protecting retinal pigment epithelial cells and photoreceptors. J. Lipid Res. 2021, 62, 100058. [Google Scholar] [CrossRef]
- Do, K.V.; Kautzmann, M.-A.I.; Jun, B.; Gordon, W.C.; Nshimiyimana, R.; Yang, R.; Petasis, N.A.; Bazan, N.G. Elovanoids counteract oligomeric β-amyloid-induced gene expression and protect photoreceptors. Proc. Natl. Acad. Sci. USA 2019, 116, 24317–24325. [Google Scholar] [CrossRef] [PubMed]
- Bazan, N.G. Docosanoids and elovanoids from omega-3 fatty acids are pro-homeostatic modulators of inflammatory responses, cell damage and neuroprotection. Mol. Asp. Med. 2018, 64, 18–33. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharjee, S.; Jun, B.; Belayev, L.; Heap, J.; Kautzmann, M.-A.; Obenaus, A.; Menghani, H.; Marcell, S.J.; Khoutorova, L.; Yang, R.; et al. Elovanoids are a novel class of homeostatic lipid mediators that protect neural cell integrity upon injury. Sci. Adv. 2017, 3, e1700735. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).