Structures and Applications of Nucleic Acid-Based Micelles for Cancer Therapy
Abstract
1. Introduction
2. Nucleic Acid-Based Micelles
2.1. DNA Micelles
2.2. RNA Micelles
3. Cancer Therapeutic Applications
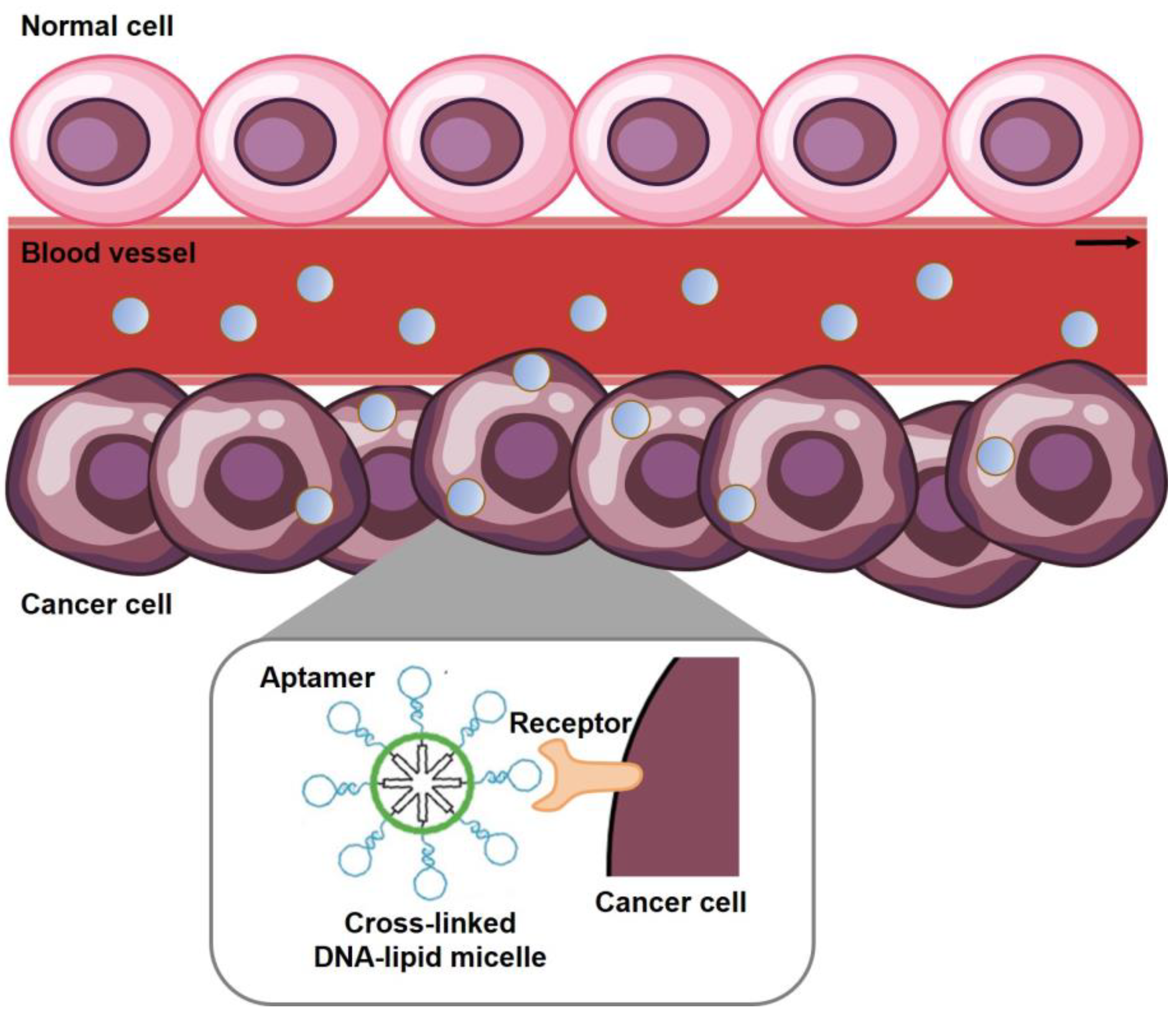
3.1. Cancer Targeting
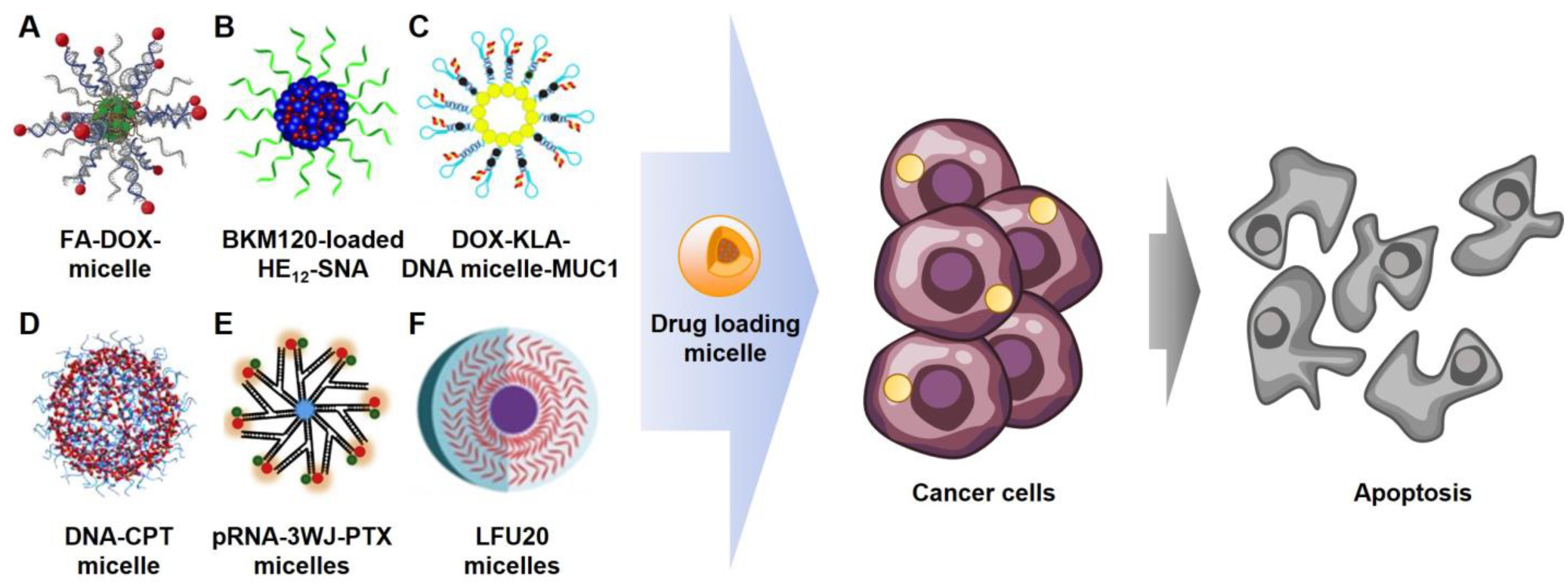
3.2. Chemotherapy
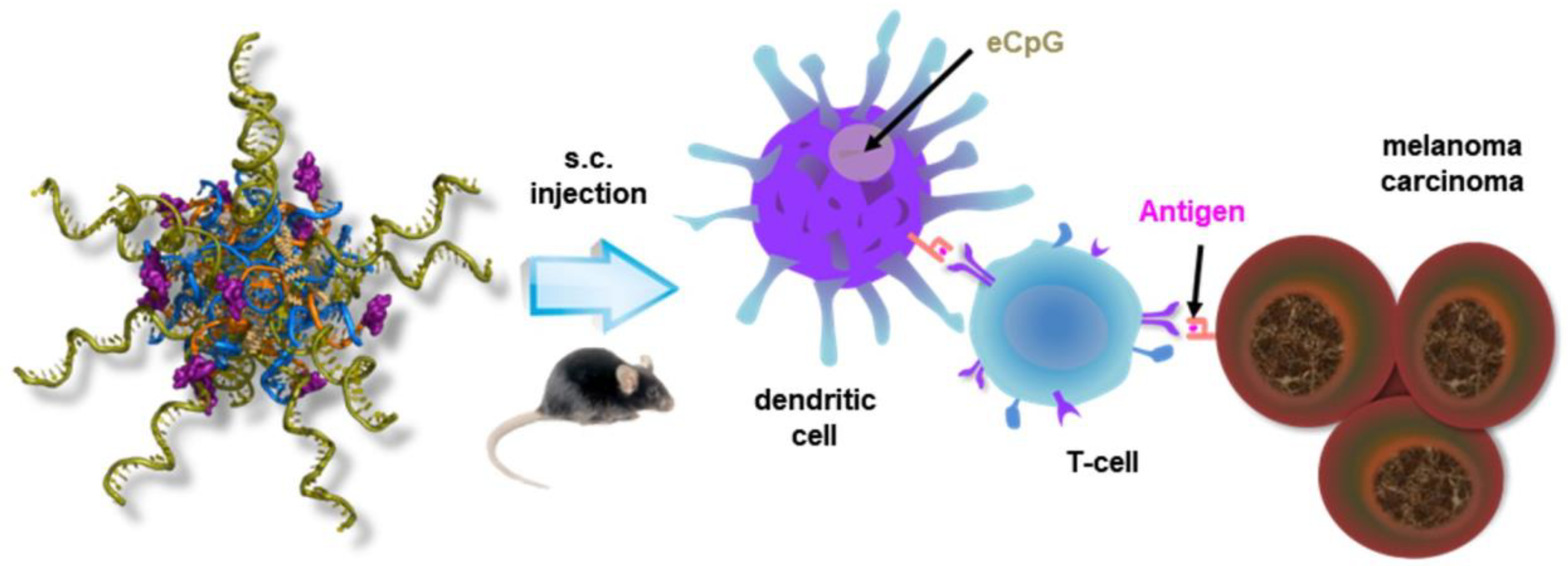
3.3. Immunotherapy
3.4. Gene Silencing Therapy
4. Conclusions and Outlook
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Watson, J.D.; Crick, F.H.C. Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid. Nature 1953, 171, 737–738. [Google Scholar] [CrossRef] [PubMed]
- Neidle, S.; Sanderson, M. Principles of Nucleic Acid Structure; Academic Press: Cambridge, MA, USA, 2021. [Google Scholar]
- Pedersen, R.; Marchi, A.N.; Majikes, J.; Nash, J.A.; Estrich, N.A.; Courson, D.S.; Hall, C.K.; Craig, S.L.; LaBean, T.H. Properties of DNA. In Handbook of Nanomaterials Properties; Bhushan, B., Luo, D., Schricker, S.R., Sigmund, W., Zauscher, S., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 1125–1157. [Google Scholar] [CrossRef]
- Watson, J.D.; Crick, F.H. A Structure for Deoxyribose Nucleic Acid; University of Chicago Press: Chicago, IL, USA, 1953. [Google Scholar]
- Jaeger, L.; Chworos, A. The architectonics of programmable RNA and DNA nanostructures. Curr. Opin. Struct. Biol. 2006, 16, 531–543. [Google Scholar] [CrossRef] [PubMed]
- Ramezani, H.; Dietz, H. Building machines with DNA molecules. Nat. Rev. Genet. 2020, 21, 5–26. [Google Scholar] [CrossRef]
- Seeman, N.C.; Sleiman, H.F. DNA nanotechnology. Nat. Rev. Mater. 2017, 3, 17068. [Google Scholar] [CrossRef]
- Gothelf, K.V.; LaBean, T.H. DNA-programmed assembly of nanostructures. Org. Biomol. Chem. 2005, 3, 4023–4037. [Google Scholar] [CrossRef] [PubMed]
- Brucale, M.; Zuccheri, G.; Samorì, B. Mastering the complexity of DNA nanostructures. Trends Biotechnol. 2006, 24, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Han, Y.; Yang, Y.; de la Fuente, J.M.; Cui, D.; Wang, X. Application of DNA nanostructures in cancer therapy. Appl. Mater. Today 2020, 21, 100861. [Google Scholar] [CrossRef]
- Wang, D.-X.; Wang, J.; Wang, Y.-X.; Du, Y.-C.; Huang, Y.; Tang, A.-N.; Cui, Y.-X.; Kong, D.-M. DNA nanostructure-based nucleic acid probes: Construction and biological applications. Chem. Sci. 2021, 12, 7602–7622. [Google Scholar] [CrossRef] [PubMed]
- Nicolson, F.; Ali, A.; Kircher, M.F.; Pal, S. DNA Nanostructures and DNA-Functionalized Nanoparticles for Cancer Theranostics. Adv. Sci. 2020, 7, 2001669. [Google Scholar] [CrossRef]
- Bujold, K.E.; Lacroix, A.; Sleiman, H.F. DNA Nanostructures at the Interface with Biology. Chem 2018, 4, 495–521. [Google Scholar] [CrossRef]
- Lacroix, A.; Sleiman, H.F. DNA Nanostructures: Current Challenges and Opportunities for Cellular Delivery. ACS Nano 2021, 15, 3631–3645. [Google Scholar] [CrossRef] [PubMed]
- Advancing Cancer Therapy. Nat. Cancer 2021, 2, 245–246. [CrossRef]
- The global challenge of cancer. Nat. Cancer 2020, 1, 1–2. [CrossRef] [PubMed]
- Qin, S.-Y.; Zhang, A.-Q.; Cheng, S.-X.; Rong, L.; Zhang, X.-Z. Drug self-delivery systems for cancer therapy. Biomaterials 2017, 112, 234–247. [Google Scholar] [CrossRef]
- Sun, T.; Zhang, Y.S.; Pang, B.; Hyun, D.C.; Yang, M.; Xia, Y. Engineered nanoparticles for drug delivery in cancer therapy. In Nanomaterials and Neoplasms; Jenny Stanford Publishing: New Delhi, India, 2021; pp. 31–142. [Google Scholar]
- Peer, D.; Karp, J.M.; Hong, S.; Farokhzad, O.C.; Margalit, R.; Langer, R. Nanocarriers as an emerging platform for cancer therapy. In Nano-Enabled Medical Applications; Jenny Stanford Publishing: New Delhi, India, 2020; pp. 61–91. [Google Scholar]
- Shin, D.H.; Tam, Y.T.; Kwon, G.S. Polymeric micelle nanocarriers in cancer research. Front. Chem. Sci. Eng. 2016, 10, 348–359. [Google Scholar] [CrossRef]
- Gong, J.; Chen, M.; Zheng, Y.; Wang, S.; Wang, Y. Polymeric micelles drug delivery system in oncology. J. Control. Release 2012, 159, 312–323. [Google Scholar] [CrossRef] [PubMed]
- Aliabadi, H.M.; Lavasanifar, A. Polymeric micelles for drug delivery. Expert Opin. Drug Deliv. 2006, 3, 139–162. [Google Scholar] [CrossRef]
- Qiu, L.; Zhang, T.; Jiang, J.; Wu, C.; Zhu, G.; You, M.; Chen, X.; Zhang, L.; Cui, C.; Yu, R.; et al. Cell Membrane-Anchored Biosensors for Real-Time Monitoring of the Cellular Microenvironment. J. Am. Chem. Soc. 2014, 136, 13090–13093. [Google Scholar] [CrossRef]
- Huo, S.; Li, H.; Boersma, A.J.; Herrmann, A. DNA Nanotechnology Enters Cell Membranes. Adv. Sci. 2019, 6, 1900043. [Google Scholar] [CrossRef]
- Liu, K.; Zheng, L.; Liu, Q.; de Vries, J.W.; Gerasimov, J.Y.; Herrmann, A. Nucleic Acid Chemistry in the Organic Phase: From Functionalized Oligonucleotides to DNA Side Chain Polymers. J. Am. Chem. Soc. 2014, 136, 14255–14262. [Google Scholar] [CrossRef]
- Caruthers, M.H. Gene Synthesis Machines: DNA Chemistry and Its Uses. Science 1985, 230, 281–285. [Google Scholar] [CrossRef] [PubMed]
- Kosuri, S.; Church, G.M. Large-scale de novo DNA synthesis: Technologies and applications. Nat. Methods 2014, 11, 499–507. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.; Li, B.B.; Lu, X.; Jia, F.; Santori, C.; Menon, P.; Li, H.; Zhang, B.; Zhao, J.J.; Zhang, K. Light-Triggered, Self-Immolative Nucleic Acid-Drug Nanostructures. J. Am. Chem. Soc. 2015, 137, 6112–6115. [Google Scholar] [CrossRef] [PubMed]
- Trinh, T.; Chidchob, P.; Bazzi, H.S.; Sleiman, H.F. DNA micelles as nanoreactors: Efficient DNA functionalization with hydrophobic organic molecules. Chem. Commun. 2016, 52, 10914–10917. [Google Scholar] [CrossRef]
- Zhang, C.; Hao, L.; Calabrese, C.M.; Zhou, Y.; Choi, C.H.J.; Xing, H.; Mirkin, C.A. Biodegradable DNA-Brush Block Copolymer Spherical Nucleic Acids Enable Transfection Agent-Free Intracellular Gene Regulation. Small 2015, 11, 5360–5368. [Google Scholar] [CrossRef]
- Alemdaroglu, F.E.; Ding, K.; Berger, R.; Herrmann, A. DNA-Templated Synthesis in Three Dimensions: Introducing a Micellar Scaffold for Organic Reactions. Angew. Chem. Int. Ed. 2006, 45, 4206–4210. [Google Scholar] [CrossRef]
- Wilks, T.R.; Bath, J.; de Vries, J.W.; Raymond, J.E.; Herrmann, A.; Turberfield, A.J.; O’Reilly, R.K. “Giant Surfactants” Created by the Fast and Efficient Functionalization of a DNA Tetrahedron with a Temperature-Responsive Polymer. ACS Nano 2013, 7, 8561–8572. [Google Scholar] [CrossRef]
- Jeong, J.H.; Park, T.G. Novel Polymer−DNA Hybrid Polymeric Micelles Composed of Hydrophobic Poly(d,l-lactic-co-glycolic Acid) and Hydrophilic Oligonucleotides. Bioconjug. Chem. 2001, 12, 917–923. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, Y.; Fullhart, P.; Mirkin, C.A. Reversible and Chemically Programmable Micelle Assembly with DNA Block-Copolymer Amphiphiles. Nano Lett. 2004, 4, 1055–1058. [Google Scholar] [CrossRef]
- Banga, R.J.; Meckes, B.; Narayan, S.P.; Sprangers, A.J.; Nguyen, S.T.; Mirkin, C.A. Cross-Linked Micellar Spherical Nucleic Acids from Thermoresponsive Templates. J. Am. Chem. Soc. 2017, 139, 4278–4281. [Google Scholar] [CrossRef]
- Liu, H.; Zhu, Z.; Kang, H.; Wu, Y.; Sefan, K.; Tan, W. DNA-Based Micelles: Synthesis, Micellar Properties and Size-Dependent Cell Permeability. Chem. Eur. J. 2010, 16, 3791–3797. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Jin, C.; Wang, R.; Kuai, H.; Zhang, L.; Zhang, X.; Li, J.; Qiu, L.; Tan, W. Fluorinated DNA Micelles: Synthesis and Properties. Anal. Chem. 2018, 90, 6843–6850. [Google Scholar] [CrossRef]
- Godeau, G.; Arnion, H.; Brun, C.; Staedel, C.; Barthélémy, P. Fluorocarbon oligonucleotide conjugates for nucleic acids delivery. MedChemComm 2010, 1, 76–78. [Google Scholar] [CrossRef]
- Charbgoo, F.; Alibolandi, M.; Taghdisi, S.M.; Abnous, K.; Soltani, F.; Ramezani, M. MUC1 aptamer-targeted DNA micelles for dual tumor therapy using doxorubicin and KLA peptide. Nanomed. Nanotechnol. Biol. Med. 2018, 14, 685–697. [Google Scholar] [CrossRef]
- Godeau, G.; Staedel, C.; Barthélémy, P. Lipid-Conjugated Oligonucleotides via “Click Chemistry” Efficiently Inhibit Hepatitis C Virus Translation. J. Med. Chem. 2008, 51, 4374–4376. [Google Scholar] [CrossRef]
- Liu, H.; Kwong, B.; Irvine, D.J. Membrane Anchored Immunostimulatory Oligonucleotides for In Vivo Cell Modification and Localized Immunotherapy. Angew. Chem. Int. Ed. 2011, 50, 7052–7055. [Google Scholar] [CrossRef]
- Kim, H.; Zhang, W.; Hwang, J.; An, E.-K.; Choi, Y.K.; Moon, E.; Loznik, M.; Huh, Y.H.; Herrmann, A.; Kwak, M.; et al. Carrier-free micellar CpG interacting with cell membrane for enhanced immunological treatment of HIV-1. Biomaterials 2021, 277, 121081. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.-O.; Park, H.; Zhang, W.; de Vries, J.W.; Gruszka, A.; Lee, M.W.; Ahn, D.-R.; Herrmann, A.; Kwak, M. Modular delivery of CpG-incorporated lipid-DNA nanoparticles for spleen DC activation. Biomaterials 2017, 115, 81–89. [Google Scholar] [CrossRef]
- Anaya, M.; Kwak, M.; Musser, A.J.; Müllen, K.; Herrmann, A. Tunable Hydrophobicity in DNA Micelles: Design, Synthesis, and Characterization of a New Family of DNA Amphiphiles. Chem. Eur. J. 2010, 16, 12852–12859. [Google Scholar] [CrossRef] [PubMed]
- Alemdaroglu, F.E.; Alemdaroglu, N.C.; Langguth, P.; Herrmann, A. DNA Block Copolymer Micelles—A Combinatorial Tool for Cancer Nanotechnology. Adv. Mater. 2008, 20, 899–902. [Google Scholar] [CrossRef]
- Shu, Y.; Yin, H.; Rajabi, M.; Li, H.; Vieweger, M.; Guo, S.; Shu, D.; Guo, P. RNA-based micelles: A novel platform for paclitaxel loading and delivery. J. Control. Release 2018, 276, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Zipkin, M. Big pharma buys into exosomes for drug delivery. Nat. Biotechnol. 2020, 38, 1226–1228. [Google Scholar] [CrossRef] [PubMed]
- Zuckerman, J.E.; Gritli, I.; Tolcher, A.; Heidel, J.D.; Lim, D.; Morgan, R.; Chmielowski, B.; Ribas, A.; Davis, M.E.; Yen, Y. Correlating animal and human phase Ia/Ib clinical data with CALAA-01, a targeted, polymer-based nanoparticle containing siRNA. Proc. Natl. Acad. Sci. USA 2014, 111, 11449–11454. [Google Scholar] [CrossRef] [PubMed]
- Jiang, T.; Qiao, Y.; Ruan, W.; Zhang, D.; Yang, Q.; Wang, G.; Chen, Q.; Zhu, F.; Yin, J.; Zou, Y.; et al. Cation-Free siRNA Micelles as Effective Drug Delivery Platform and Potent RNAi Nanomedicines for Glioblastoma Therapy. Adv. Mater. 2021, 33, 2104779. [Google Scholar] [CrossRef]
- Yoon, P.O.; Park, J.W.; Lee, C.-M.; Kim, S.H.; Kim, H.-N.; Ko, Y.; Bae, S.J.; Yun, S.; Park, J.H.; Kwon, T.; et al. Self-assembled Micelle Interfering RNA for Effective and Safe Targeting of Dysregulated Genes in Pulmonary Fibrosis. J. Biol. Chem. 2016, 291, 6433–6446. [Google Scholar] [CrossRef]
- Wu, Y.; Sefah, K.; Liu, H.; Wang, R.; Tan, W. DNA aptamer–micelle as an efficient detection/delivery vehicle toward cancer cells. Proc. Natl. Acad. Sci. USA 2010, 107, 5–10. [Google Scholar] [CrossRef]
- Li, X.; Figg, C.A.; Wang, R.; Jiang, Y.; Lyu, Y.; Sun, H.; Liu, Y.; Wang, Y.; Teng, I.-T.; Hou, W.; et al. Cross-Linked Aptamer–Lipid Micelles for Excellent Stability and Specificity in Target-Cell Recognition. Angew. Chem. Int. Ed. 2018, 57, 11589–11593. [Google Scholar] [CrossRef] [PubMed]
- Bousmail, D.; Amrein, L.; Fakhoury, J.J.; Fakih, H.H.; Hsu, J.C.C.; Panasci, L.; Sleiman, H.F. Precision spherical nucleic acids for delivery of anticancer drugs. Chem. Sci. 2017, 8, 6218–6229. [Google Scholar] [CrossRef]
- Jin, C.; Zhang, H.; Zou, J.; Liu, Y.; Zhang, L.; Li, F.; Wang, R.; Xuan, W.; Ye, M.; Tan, W. Floxuridine Homomeric Oligonucleotides “Hitchhike” with Albumin In Situ for Cancer Chemotherapy. Angew. Chem. Int. Ed. 2018, 57, 8994–8997. [Google Scholar] [CrossRef]
- Jin, J.-O.; Kim, H.; Huh, Y.H.; Herrmann, A.; Kwak, M. Soft matter DNA nanoparticles hybridized with CpG motifs and peptide nucleic acids enable immunological treatment of cancer. J. Control. Release 2019, 315, 76–84. [Google Scholar] [CrossRef]
- Yin, H.; Wang, H.; Li, Z.; Shu, D.; Guo, P. RNA Micelles for the Systemic Delivery of Anti-miRNA for Cancer Targeting and Inhibition without Ligand. ACS Nano 2019, 13, 706–717. [Google Scholar] [CrossRef] [PubMed]
- Rosenblum, D.; Joshi, N.; Tao, W.; Karp, J.M.; Peer, D. Progress and challenges towards targeted delivery of cancer therapeutics. Nat. Commun. 2018, 9, 1410. [Google Scholar] [CrossRef] [PubMed]
- Dai, L.; Liu, J.; Luo, Z.; Li, M.; Cai, K. Tumor therapy: Targeted drug delivery systems. J. Mater. Chem. B 2016, 4, 6758–6772. [Google Scholar] [CrossRef] [PubMed]
- Mun, E.J.; Babiker, H.M.; Weinberg, U.; Kirson, E.D.; Von Hoff, D.D. Tumor-Treating Fields: A Fourth Modality in Cancer Treatment. Clin. Cancer Res. 2018, 24, 266–275. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Kim, D.-M.; Kim, K.-S.; Jung, W.; Kim, D.-E. Applications of Cancer Cell-Specific Aptamers in Targeted Delivery of Anticancer Therapeutic Agents. Molecules 2018, 23, 830. [Google Scholar] [CrossRef] [PubMed]
- Gao, F.; Yin, J.; Chen, Y.; Guo, C.; Hu, H.; Su, J. Recent advances in aptamer-based targeted drug delivery systems for cancer therapy. Front. Bioeng. Biotechnol. 2022, 10, 972933. [Google Scholar] [CrossRef]
- Zhang, Y.; Hong, H.; Cai, W. Tumor-Targeted Drug Delivery with Aptamers. Curr. Med. Chem. 2011, 18, 4185–4194. [Google Scholar] [CrossRef]
- Anand, U.; Dey, A.; Chandel, A.K.S.; Sanyal, R.; Mishra, A.; Pandey, D.K.; De Falco, V.; Upadhyay, A.; Kandimalla, R.; Chaudhary, A.; et al. Cancer chemotherapy and beyond: Current status, drug candidates, associated risks and progress in targeted therapeutics. Genes Dis. 2022. [Google Scholar] [CrossRef]
- Chabner, B.A.; Roberts, T.G. Chemotherapy and the war on cancer. Nat. Rev. Cancer 2005, 5, 65–72. [Google Scholar] [CrossRef]
- Schirrmacher, V. From chemotherapy to biological therapy: A review of novel concepts to reduce the side effects of systemic cancer treatment (Review). Int. J. Oncol. 2019, 54, 407–419. [Google Scholar] [CrossRef]
- Li, Z.; Tan, S.; Li, S.; Shen, Q.; Wang, K. Cancer drug delivery in the nano era: An overview and perspectives (Review). Oncol. Rep. 2017, 38, 611–624. [Google Scholar] [CrossRef] [PubMed]
- Senapati, S.; Mahanta, A.K.; Kumar, S.; Maiti, P. Controlled drug delivery vehicles for cancer treatment and their performance. Signal Transduct. Target. Ther. 2018, 3, 7. [Google Scholar] [CrossRef] [PubMed]
- Dang, Y.; Guan, J. Nanoparticle-based drug delivery systems for cancer therapy. Smart Mater. Med. 2020, 1, 10–19. [Google Scholar] [CrossRef]
- Nussbaumer, S.; Bonnabry, P.; Veuthey, J.-L.; Fleury-Souverain, S. Analysis of anticancer drugs: A review. Talanta 2011, 85, 2265–2289. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Blum, N.T.; Lin, J.; Shi, J.; Zhang, C.; Huang, P. Chemotherapeutic drug–DNA hybrid nanostructures for anti-tumor therapy. Mater. Horiz. 2021, 8, 78–101. [Google Scholar] [CrossRef]
- Chaires, J.B. Calorimetry and Thermodynamics in Drug Design. Annu. Rev. Biophys. 2008, 37, 135–151. [Google Scholar] [CrossRef] [PubMed]
- Agudelo, D.; Bourassa, P.; Bérubé, G.; Tajmir-Riahi, H.-A. Intercalation of antitumor drug doxorubicin and its analogue by DNA duplex: Structural features and biological implications. Int. J. Biol. Macromol. 2014, 66, 144–150. [Google Scholar] [CrossRef] [PubMed]
- Jawad, B.; Poudel, L.; Podgornik, R.; Ching, W.-Y. Thermodynamic Dissection of the Intercalation Binding Process of Doxorubicin to dsDNA with Implications of Ionic and Solvent Effects. J. Phys. Chem. B 2020, 124, 7803–7818. [Google Scholar] [CrossRef]
- Rider, B.J. 5 Fluorodeoxyuridine. In xPharm: The Comprehensive Pharmacology Reference; Enna, S.J., Bylund, D.B., Eds.; Elsevier: New York, NY, USA, 2007; pp. 1–4. [Google Scholar] [CrossRef]
- Longley, D.B.; Harkin, D.P.; Johnston, P.G. 5-Fluorouracil: Mechanisms of action and clinical strategies. Nat. Rev. Cancer 2003, 3, 330–338. [Google Scholar] [CrossRef] [PubMed]
- Efficacy of adjuvant fluorouracil and folinic acid in colon cancer: International Multicentre Pooled Analysis of Colon Cancer Trials (IMPACT) investigators. Lancet 1995, 345, 939–944. [CrossRef]
- Schuster, M.; Nechansky, A.; Kircheis, R. Cancer immunotherapy. Biotechnol. J. 2006, 1, 138–147. [Google Scholar] [CrossRef] [PubMed]
- Waldman, A.D.; Fritz, J.M.; Lenardo, M.J. A guide to cancer immunotherapy: From T cell basic science to clinical practice. Nat. Rev. Immunol. 2020, 20, 651–668. [Google Scholar] [CrossRef] [PubMed]
- Dougan, M.; Dranoff, G. Immune Therapy for Cancer. Annu. Rev. Immunol. 2009, 27, 83–117. [Google Scholar] [CrossRef] [PubMed]
- Chuang, Y.-C.; Tseng, J.-C.; Huang, L.-R.; Huang, C.-M.; Huang, C.-Y.F.; Chuang, T.-H. Adjuvant Effect of Toll-like Receptor 9 Activation on Cancer Immunotherapy Using Checkpoint Blockade. Front. Immunol. 2020, 11, 1075. [Google Scholar] [CrossRef]
- Krieg, A.M. Toll-like receptor 9 (TLR9) agonists in the treatment of cancer. Oncogene 2008, 27, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Dongye, Z.; Li, J.; Wu, Y. Toll-like receptor 9 agonists and combination therapies: Strategies to modulate the tumour immune microenvironment for systemic anti-tumour immunity. Br. J. Cancer 2022, 127, 1584–1594. [Google Scholar] [CrossRef]
- Riley, R.S.; June, C.H.; Langer, R.; Mitchell, M.J. Delivery technologies for cancer immunotherapy. Nat. Rev. Drug Discov. 2019, 18, 175–196. [Google Scholar] [CrossRef]
- Chi, Q.; Yang, Z.; Xu, K.; Wang, C.; Liang, H. DNA Nanostructure as an Efficient Drug Delivery Platform for Immunotherapy. Front. Pharmacol. 2020, 10, 1585. [Google Scholar] [CrossRef]
- Mukalel, A.J.; Riley, R.S.; Zhang, R.; Mitchell, M.J. Nanoparticles for nucleic acid delivery: Applications in cancer immunotherapy. Cancer Lett. 2019, 458, 102–112. [Google Scholar] [CrossRef]
- Tian, Z.; Liang, G.; Cui, K.; Liang, Y.; Wang, Q.; Lv, S.; Cheng, X.; Zhang, L. Insight Into the Prospects for RNAi Therapy of Cancer. Front. Pharmacol. 2021, 12, 644718. [Google Scholar] [CrossRef]
- Mansoori, B.; Sandoghchian Shotorbani, S.; Baradaran, B. RNA Interference and its Role in Cancer Therapy. Adv. Pharm. Bull. 2014, 4, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Wang, C.C.; Choy, K.W.; Du, Q.; Chen, J.; Wang, Q.; Li, L.; Chung, T.K.H.; Tang, T. Therapeutic potentials of gene silencing by RNA interference: Principles, challenges, and new strategies. Gene 2014, 538, 217–227. [Google Scholar] [CrossRef] [PubMed]
- Xin, Y.; Huang, M.; Guo, W.W.; Huang, Q.; Zhang, L.z.; Jiang, G. Nano-based delivery of RNAi in cancer therapy. Mol. Cancer. 2017, 16, 134. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Gao, C.; Kuang, M.; Xu, M.; Wang, B.; Luo, Y.; Teng, L.; Xie, J. Nanoparticles as Drug Delivery Systems of RNAi in Cancer Therapy. Molecules 2021, 26, 2380. [Google Scholar] [CrossRef]
- Kara, G.; Calin, G.A.; Ozpolat, B. RNAi-based therapeutics and tumor targeted delivery in cancer. Adv. Drug Delivery Rev. 2022, 182, 114113. [Google Scholar] [CrossRef]
- Letsinger, R.L.; Zhang, G.R.; Sun, D.K.; Ikeuchi, T.; Sarin, P.S. Cholesteryl-conjugated oligonucleotides: Synthesis, properties, and activity as inhibitors of replication of human immunodeficiency virus in cell culture. Proc. Natl. Acad. Sci. USA 1989, 86, 6553–6556. [Google Scholar] [CrossRef]
- Ezzat, K.; Aoki, Y.; Koo, T.; McClorey, G.; Benner, L.; Coenen-Stass, A.; O’Donovan, L.; Lehto, T.; Garcia-Guerra, A.; Nordin, J.; et al. Self-Assembly into Nanoparticles Is Essential for Receptor Mediated Uptake of Therapeutic Antisense Oligonucleotides. Nano Lett. 2015, 15, 4364–4373. [Google Scholar] [CrossRef]
- Boutorin, A.S.; Gus’kova, L.V.; Ivanova, E.M.; Kobetz, N.D.; Zarytova, V.F.; Ryte, A.S.; Yurchenko, L.V.; Vlassov, V.V. Synthesis of alkylating oligonucleotide derivatives containing cholesterol or phenazinium residues at their 3′-terminus and their interaction with DNA within mammalian cells. FEBS Lett. 1989, 254, 129–132. [Google Scholar] [CrossRef]
- Li, X.; Chen, Y.; Wang, M.; Ma, Y.; Xia, W.; Gu, H. A mesoporous silica nanoparticle—EI—Fusogenic peptide system for siRNA delivery in cancer therapy. Biomaterials 2013, 34, 1391–1401. [Google Scholar] [CrossRef] [PubMed]
- Rupaimoole, R.; Slack, F.J. MicroRNA therapeutics: Towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 2017, 16, 203–222. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, C.M.; Echeverria, D.; Hassler, M.; Ly, S.; Khvorova, A. Cell Type Impacts Accessibility of mRNA to Silencing by RNA Interference. Mol. Ther. Nucleic 2020, 21, 384–393. [Google Scholar] [CrossRef]
- de Fougerolles, A.; Vornlocher, H.-P.; Maraganore, J.; Lieberman, J. Interfering with disease: A progress report on siRNA-based therapeutics. Nat. Rev. Drug Discov. 2007, 6, 443–453. [Google Scholar] [CrossRef] [PubMed]
- Watts, J.K.; Corey, D.R. Silencing disease genes in the laboratory and the clinic. J. Pathol. 2012, 226, 365–379. [Google Scholar] [CrossRef]
- Zhu, S.; Zhang, T.; Zheng, L.; Liu, H.; Song, W.; Liu, D.; Li, Z.; Pan, C.-x. Combination strategies to maximize the benefits of cancer immunotherapy. J. Hematol. Oncol. 2021, 14, 156. [Google Scholar] [CrossRef] [PubMed]
- Mokhtari, R.B.; Homayouni, T.S.; Baluch, N.; Morgatskaya, E.; Kumar, S.; Das, B.; Yeger, H. Combination therapy in combating cancer. Oncotarget 2017, 8, 38022. [Google Scholar] [CrossRef] [PubMed]
- Kutova, O.M.; Guryev, E.L.; Sokolova, E.A.; Alzeibak, R.; Balalaeva, I.V. Targeted Delivery to Tumors: Multidirectional Strategies to Improve Treatment Efficiency. Cancers 2019, 11, 68. [Google Scholar] [CrossRef]
- Shin, M.D.; Shukla, S.; Chung, Y.H.; Beiss, V.; Chan, S.K.; Ortega-Rivera, O.A.; Wirth, D.M.; Chen, A.; Sack, M.; Pokorski, J.K.; et al. COVID-19 vaccine development and a potential nanomaterial path forward. Nat. Nanotechnol. 2020, 15, 646–655. [Google Scholar] [CrossRef]
- Ojasalo, S.; Piskunen, P.; Shen, B.; Kostiainen, M.A.; Linko, V. Hybrid Nanoassemblies from Viruses and DNA Nanostructures. Nanomaterials 2021, 11, 1413. [Google Scholar] [CrossRef]
- Tian, T.; Li, Y.; Lin, Y. Prospects and challenges of dynamic DNA nanostructures in biomedical applications. Bone Res. 2022, 10, 40. [Google Scholar] [CrossRef]
- Zimmerman, M.I.; Porter, J.R.; Ward, M.D.; Singh, S.; Vithani, N.; Meller, A.; Mallimadugula, U.L.; Kuhn, C.E.; Borowsky, J.H.; Wiewiora, R.P.; et al. SARS-CoV-2 simulations go exascale to predict dramatic spike opening and cryptic pockets across the proteome. Nat. Chem. 2021, 13, 651–659. [Google Scholar] [CrossRef]
- Guevara, M.L.; Persano, F.; Persano, S. Advances in Lipid Nanoparticles for mRNA-Based Cancer Immunotherapy. Front. Chem. 2020, 8, 589959. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.; Zaks, T.; Langer, R.; Dong, Y. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 2021, 6, 1078–1094. [Google Scholar] [CrossRef] [PubMed]
- Tenchov, R.; Bird, R.; Curtze, A.E.; Zhou, Q. Lipid Nanoparticles─From Liposomes to mRNA Vaccine Delivery, a Landscape of Research Diversity and Advancement. ACS Nano 2021, 15, 16982–17015. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines—A new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279. [Google Scholar] [CrossRef]
| Nucleic Acid | Hydrophobic Moiety | Size (nm) | Therapeutic Agent | Therapy | Cell or Tumor Type | Ref. |
|---|---|---|---|---|---|---|
| DNA | Diacyl chain | 68 ± 13 | TDO5 aptamer | Cancer targeting | Ramos cells | [51] |
| DNA | Diacyl lipid | 17 ± 2 | sgc8 aptamer | Cancer targeting | CCRF-CEM cells | [52] |
| DNA | PPO | 10.8 ± 2.2 | Doxorubicin (Dox), Folic acid (FA) | Chemotherapy Cancer targeting | Caco-2 cells | [45] |
| DNA | Dodecane | 11.8 ± 0.4 | BKM120 | Chemotherapy | HeLa cells | [53] |
| DNA | Cholesterol | 371.3 ± 3.1 | DOX, anti-MUC1, KLA peptide | Chemotherapy Cancer targeting | MCF-7 cells, C26-tumor | [39] |
| DNA | CPT | 32 ± 13 | CPT | Chemotherapy | SK-BR-3 cells | [28] |
| RNA | Cholesterol | 118.7 ± 14.50 | Paclitaxel (PTX) | Chemotherapy | Human KB cells, KB tumor | [46] |
| DNA | Octadecyl chain | 43 | Fluorouracil | Chemotherapy | HeLa cells | [54] |
| DNA | DPPE lipid | 25 ± 5 | ODN 1826, ODN 7909 | Immunotherapy | HEK-Blue™-mTLR9 cells, Ramos-Blue cells | [35] |
| DNA | Dodec-1-ynyl uracil | 10.10 ± 0.68 | ODN 1826, OVA antigen | Immunotherapy | B16-OVA cells, B16 melanoma, CT-26 carcinoma | [55] |
| RNA | Cholesterol | 21.51 ± 6.136 | anti-miR21, folate | Gene silencing therapy Cancer targeting | KB cells, HT29 cells, KB tumor | [56] |
| RNA | poly(N-isopropylacrylamide) | 40~150 | siPLK1, temozolomide (TMZ) | Gene silencing therapy Chemotherapy | U87MG cells, U251 cells | [49] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, H.; Kwak, M. Structures and Applications of Nucleic Acid-Based Micelles for Cancer Therapy. Int. J. Mol. Sci. 2023, 24, 1592. https://doi.org/10.3390/ijms24021592
Kim H, Kwak M. Structures and Applications of Nucleic Acid-Based Micelles for Cancer Therapy. International Journal of Molecular Sciences. 2023; 24(2):1592. https://doi.org/10.3390/ijms24021592
Chicago/Turabian StyleKim, Haejoo, and Minseok Kwak. 2023. "Structures and Applications of Nucleic Acid-Based Micelles for Cancer Therapy" International Journal of Molecular Sciences 24, no. 2: 1592. https://doi.org/10.3390/ijms24021592
APA StyleKim, H., & Kwak, M. (2023). Structures and Applications of Nucleic Acid-Based Micelles for Cancer Therapy. International Journal of Molecular Sciences, 24(2), 1592. https://doi.org/10.3390/ijms24021592






