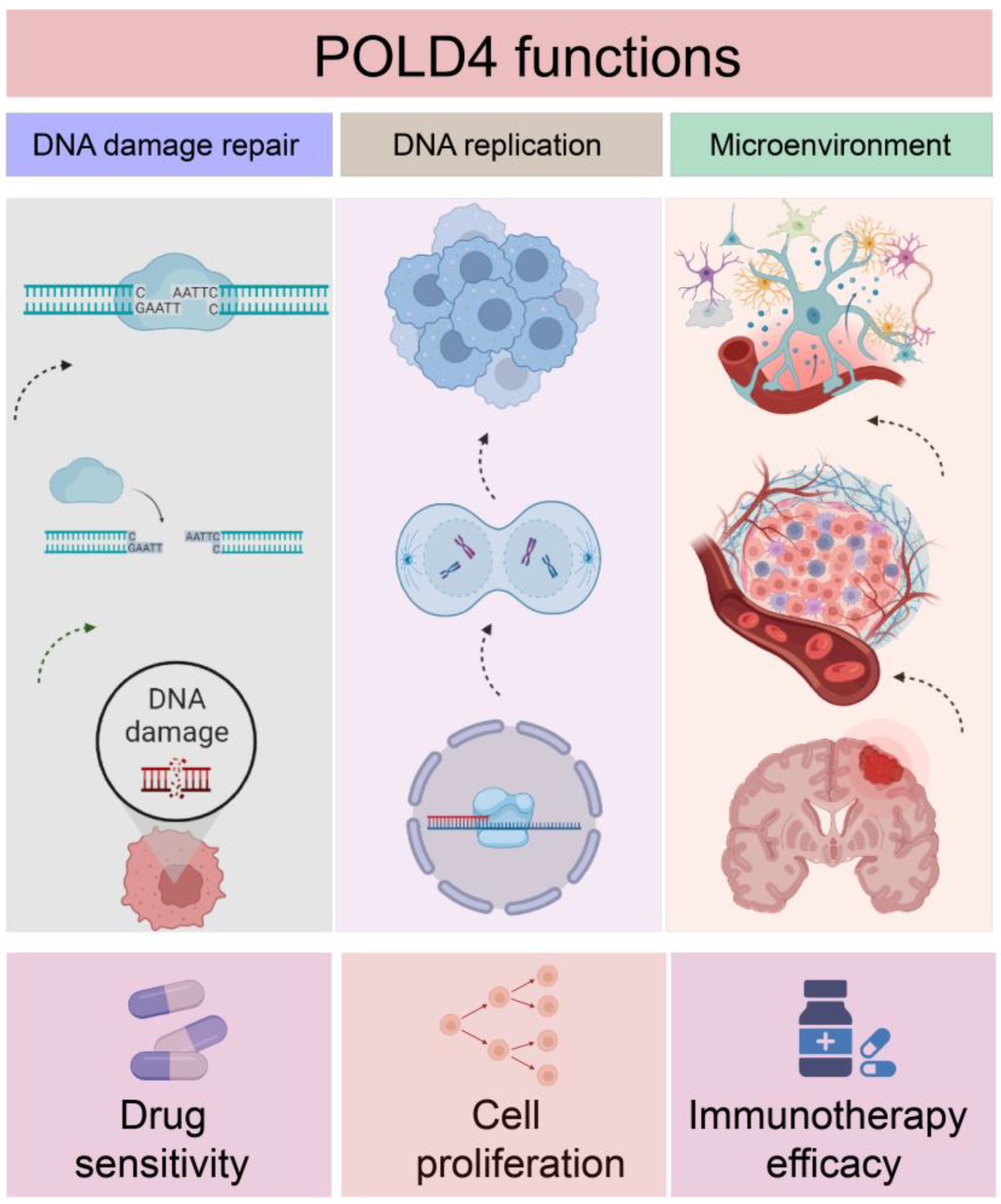
1. Introduction
Cancer is a devastating disease characterized by uncontrolled cell growth and proliferation, posing a significant global health burden. Despite remarkable strides in medical technology and the expanding array of treatment alternatives for cancer, a significant proportion of patients receive diagnoses at an advanced stage, rendering conventional therapeutic strategies impractical and culminating in fatal outcomes associated with the disease [
1,
2]. Recognizing the limitations of existing treatment methods, there is a pressing requirement to encourage the development of fresh targets for cancer therapy, intended to enhance precise diagnosis and successful treatment approaches [
3].
Up until now, although it has been suggested that abnormalities in DNA replication contribute to the progression of cancer, there has been limited identification of core DNA replication proteins as commonly altered entities in cancer [
4,
5]. We conducted extensive bioinformatics analyses, screening for DNA replication genes of significant biological importance across different cancer types. POLD4 stood out due to its intriguing expression patterns, correlation with prognosis, and functional enrichment in multiple cancer types, which piqued our interest. The essential DNA replication complex known as pol δ in eukaryotes is made up of four distinct subunits: POLD1, POLD2, POLD3, and POLD4. With its exonuclease activity, this enzyme significantly enhances proofreading capabilities, crucially upholding the precision of DNA replication [
6]. Additionally, it aids in the restoration of DNA damage resulting from mutagen exposure [
7]. Over the past decade, an increasing body of research established a connection between mutations in Pol δ and various human pathologies [
8]. Playing a fundamental role in DNA replication and repair processes, POLD4 serves as the smallest subunit of Pol δ [
9]. It acts as a crucial mediator of faithful DNA synthesis and maintenance of genomic stability. Currently, research on POLD4 in cancer is primarily focused on lung cancer. Research has indicated that a significant tobacco carcinogen, the benzopyrene analogue 4NQO, downregulated the expression level of the POLD4 gene, resulting in a decreased nucleotide excision repair and further leading to genomic instability, ultimately elevating the risk of lung cancer formation [
10]. Huang and colleagues discovered that reducing POLD4 levels in lung cancer cells not only delays the progression via the G1-S cell cycle transition but also results in heightened genomic instability [
11]. Subsequently, the team conducted additional research and found that the reduction in POLD4 in Calu6 cells leads to G1-S blockage by inhibiting the Akt-Skp2-p27 pathway [
12]. The reduction in POLD4 mediated by shRNA resulted in a significant decrease in the colony formation activity of Calu6, ACC-LC-319, and PC-10 cells [
13]. Additionally, evidence indicates that POLD4 may serve as a promising novel target for lung carcinoma, given its selective impact on cancer cells without affecting normal cells [
9]. However, the exploration of POLD4 in other cancer types and the in-depth mechanisms of its actions remain limited, and its complete function has yet to be fully clarified.
Integrated bioinformatics and public databases are indispensable in cancer research, providing a centralized platform for aggregating diverse datasets. These resources encompass genomic, gene expression, clinical, and drug response data, enabling a holistic comprehension of cancer biology. Additionally, they promote global collaboration, empowering researchers to share and analyze data, and facilitating meta-analyses, pattern identification, and biomarker discovery. These databases offer critical analysis tools, from basic sequence alignment to advanced machine learning, aiding in data interpretation. Moreover, they drive personalized medicine by identifying genetic factors influencing cancer predisposition, prognosis, and treatment response.
By integrating computational analysis with experimental data, our goal is to provide valuable insights into the potential therapeutic targeting of POLD4 and the modulation of the immune microenvironment in cancers.
3. Discussion
The DNA polymerase delta (Pol δ) is a crucial player in DNA replication and genomic replication processes [
21]. This enzyme has been linked to the recovery from replication fork collapse via various pathways, such as error-free template switch, break-induced replication, and error-prone translesion DNA synthesis [
22]. Additionally, it has been observed that all subunits of DNA Pol δ are frequently overexpressed in human cancers [
23]. However, the specific mechanism underlying DNA Pol δ amplification remains unclear, awaiting further investigation to discern its direct role in oncogenesis or its potential function as a defensive mechanism activated to bolster tolerance in the face of replication stress.
The POLD4 subunit has gained significant prominence as a pivotal factor in governing the regulation of Pol δ [
24,
25]. In response to DNA damage and during the transition into the S-phase, POLD4 undergoes transient degradation. This degradation leads to the transformation of Pol δ4 into a trimer, designated as Pol δ3. Cells knocked out for POLD4 exhibit a deficiency in homologous recombination (HR) repair, highlighting the requirement of Pol δ4, rather than Pol δ3, for effective HR. Furthermore, POLD4 knockout H1299 cells exhibit increased sensitivity to PARP inhibitors [
26]. The inhibition of POLD4 in lung cancer cells was discovered to impede the progression of the G1-S cell cycle transition, thereby causing heightened genomic instability. Furthermore, the downregulation of POLD4 in Calu6 cells induces a blockade during the G1-S phase by suppressing the Akt-Skp2-p27 pathway [
11,
12]. Despite these studies reporting the role and mechanisms of POLD4 in relevant tumors, there is still vast research potential for POLD4 in cancer. Therefore, further research is needed to unveil the veil of POLD4 in tumors.
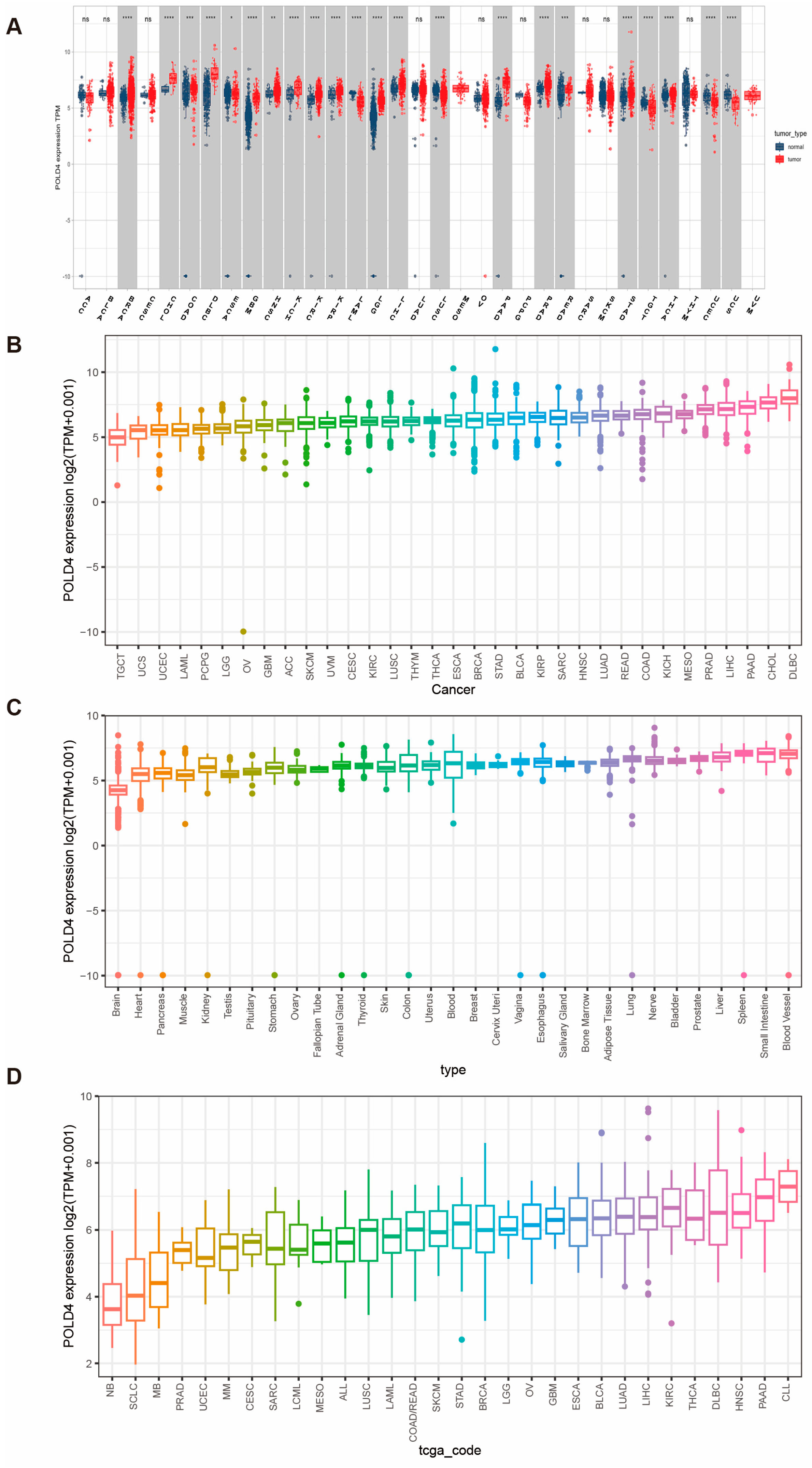
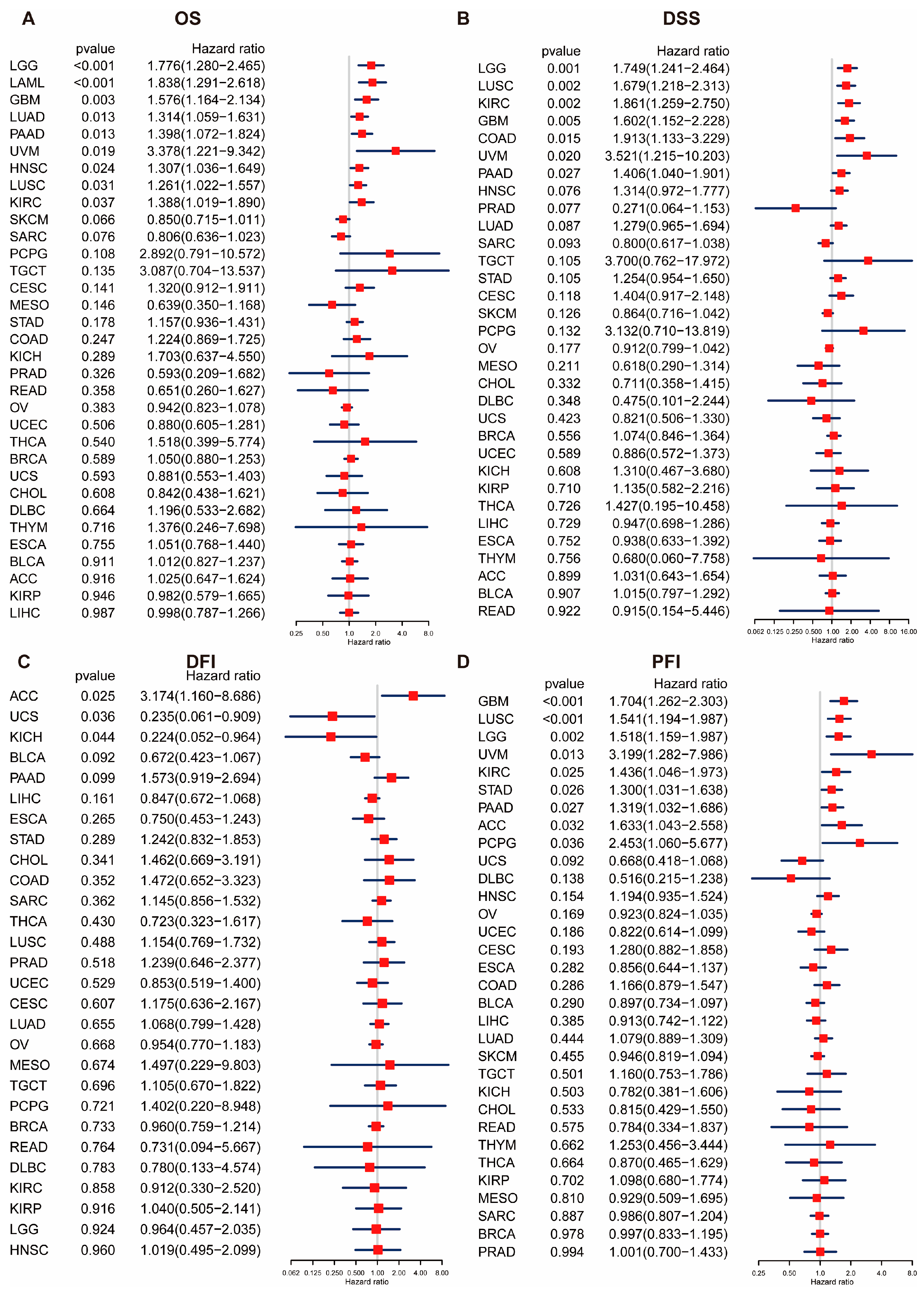
Our research findings indicate that the POLD4 gene is significantly overexpressed in 17 types of cancer compared to adjacent normal tissues. Utilizing the Univariate Cox regression analyses, we established a strong association between upregulated POLD4 expression and poorer overall survival in LGG, LAML, GBM, LUAD, PAAD, and UVM. Furthermore, a noteworthy correlation between POLD4 expression and CNA was observed. These compelling findings emphasize the significance of POLD4 as a noteworthy prognostic marker in the aforementioned tumor types.
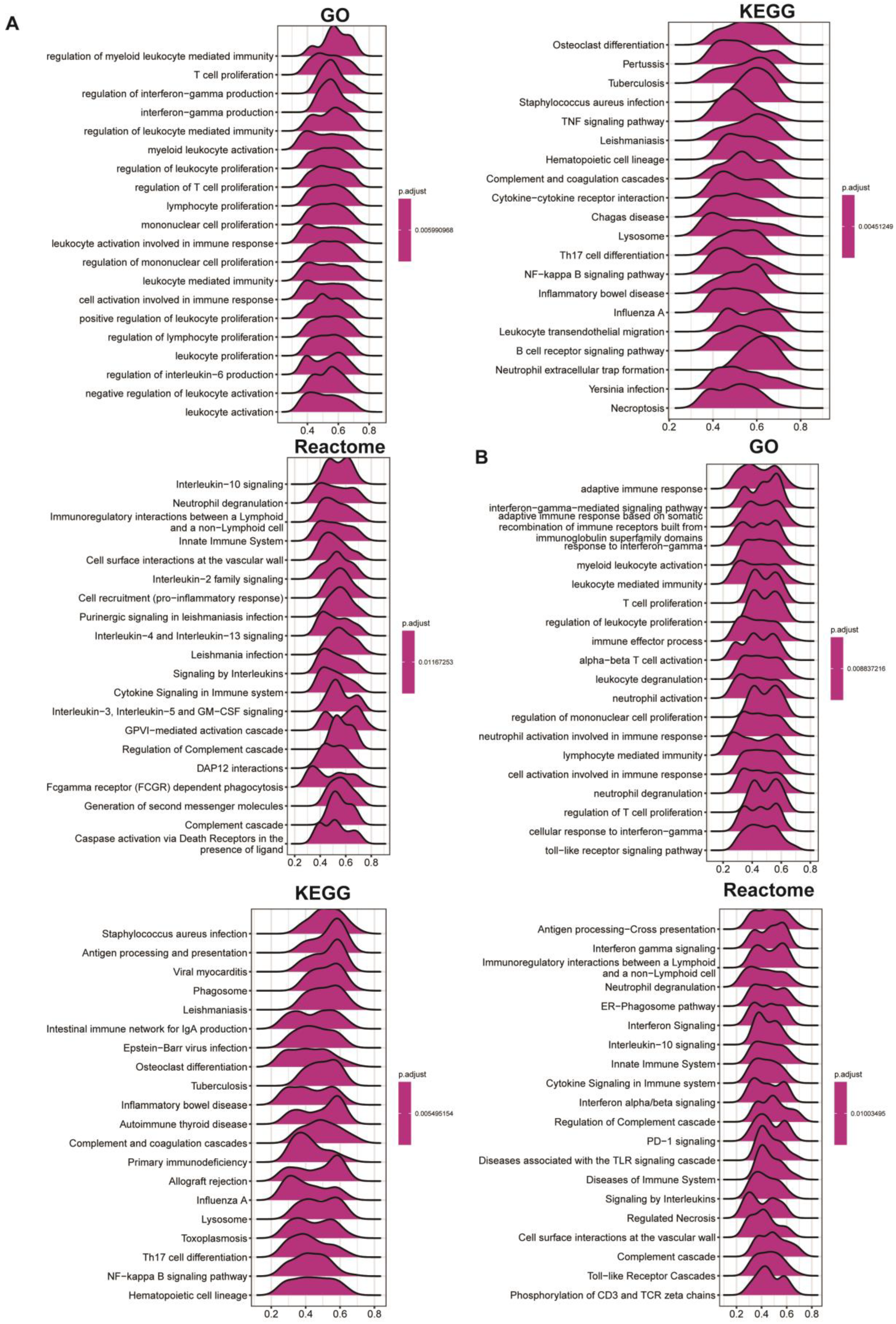
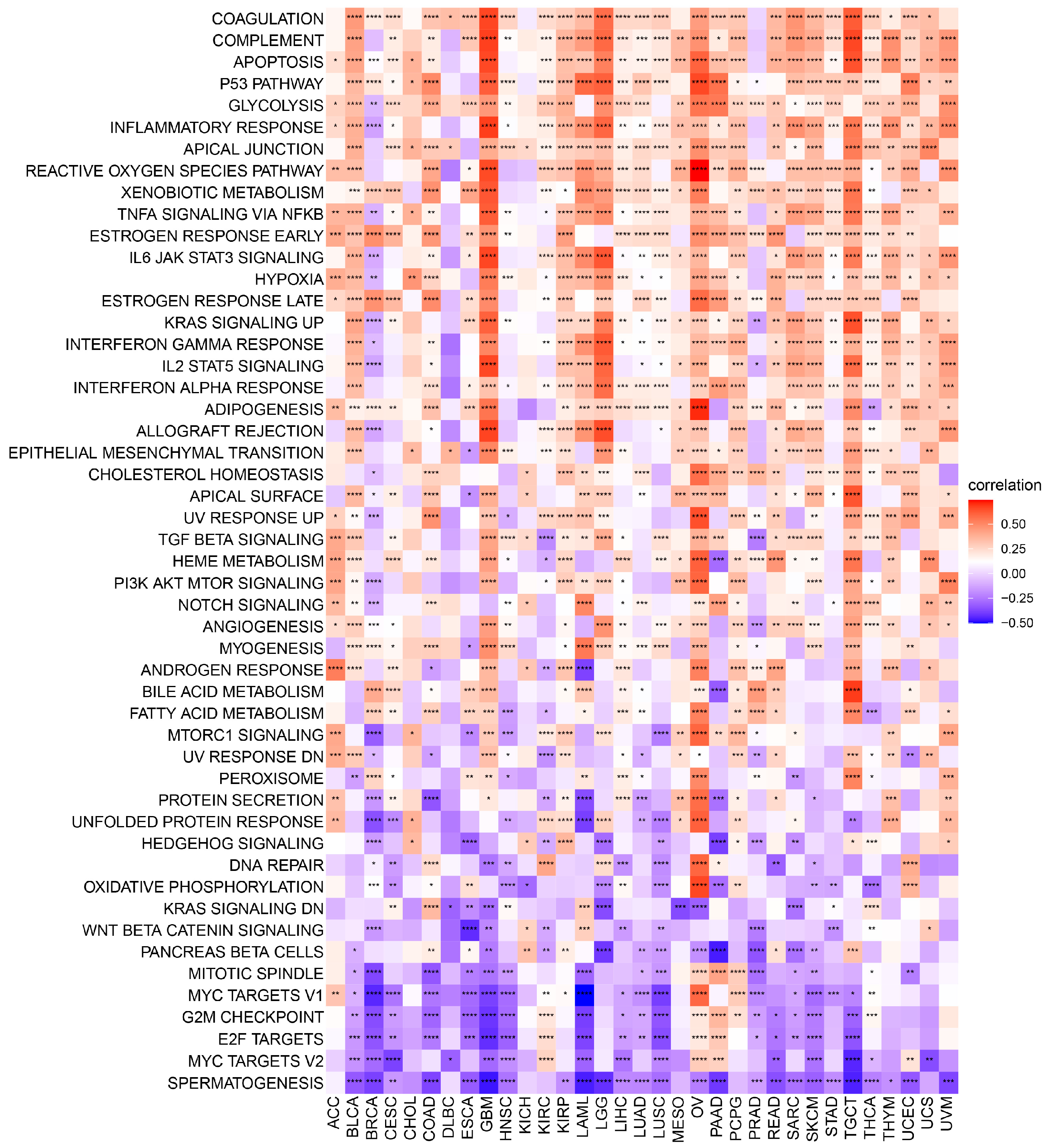
The exact molecular mechanisms of POLD4 in cancer initiation and progression remain incompletely understood and require further research. However, the associations between POLD4 and some potential tumor-related pathways may provide valuable clues for understanding its unique functional mechanisms in cancer. Functional results from GSEA demonstrate that POLD4 has the potential to influence cancer development and progression via various mechanisms, including the regulation of immune responses such as modulating myeloid cell-mediated immunity, T-cell proliferation, leukocyte-mediated immune regulation, and involvement in immune response cell activation. Additionally, our Gene Set Variation Analysis (GSVA) reveals the extensive involvement of POLD4 in multiple pathways. Notably, POLD4 plays a role in stress-induced tumor responses, such as apoptosis pathways, P53 pathways, reactive oxygen species pathways, and hypoxia pathways, which might be correlated with the literature-reported involvement of POLD4 in DNA damage repair. These cumulative findings indicate POLD4’s multifaceted engagement in cancer development.
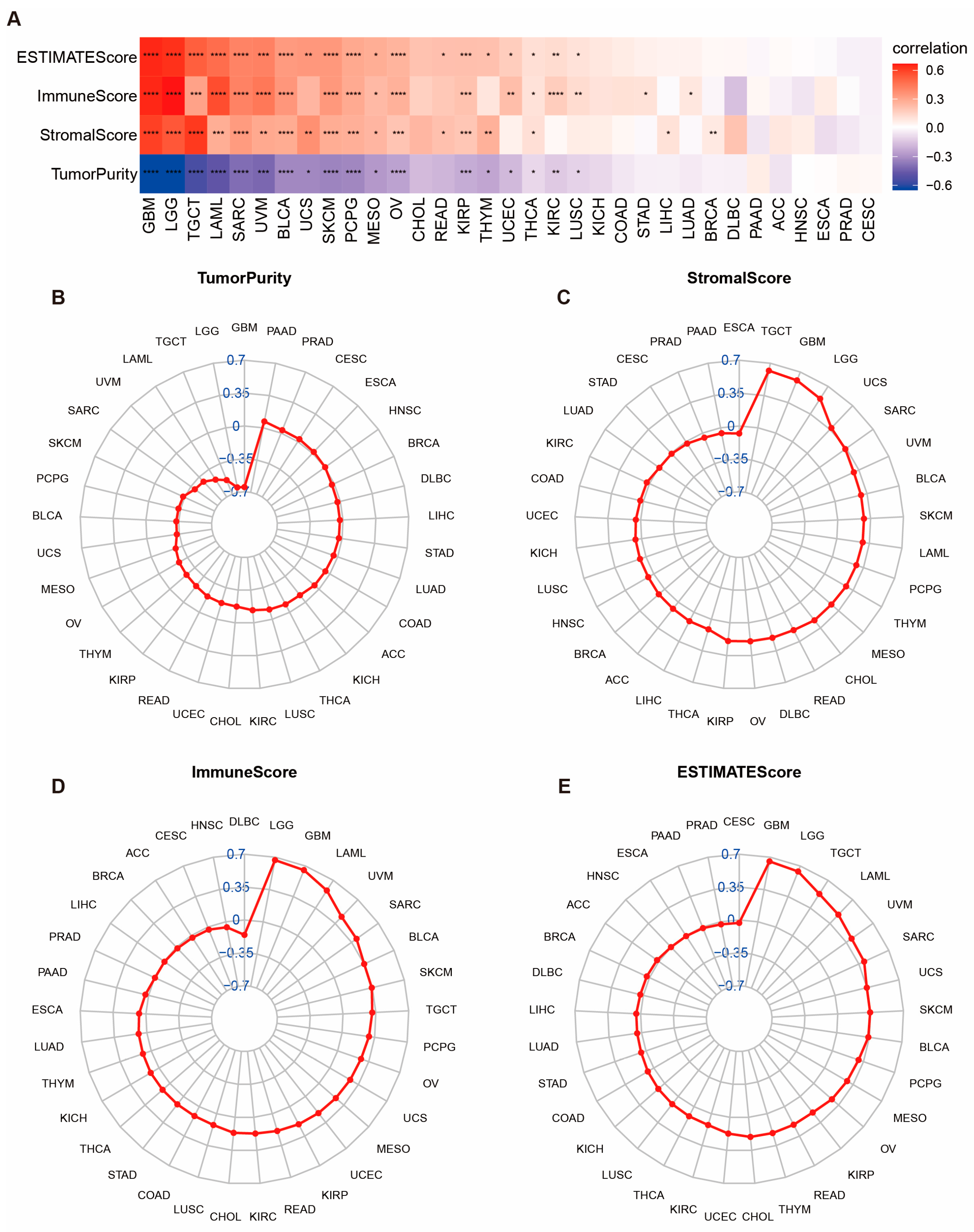
Our research highlights the essential role of POLD4 in cancer immunity, where the characteristics of the tumor microenvironment (TME) serve as valuable indicators for evaluating tumor cell responses to immunotherapy and influencing clinical outcomes. By utilizing ESTIMATE scores, we established positive correlations between POLD4 expression and the presence of stromal and immune cells in the TME across various types of cancer. Immunocyte populations, which play a pivotal role in the tumor immune network, significantly contribute to the development and occurrence of malignancies [
27,
28,
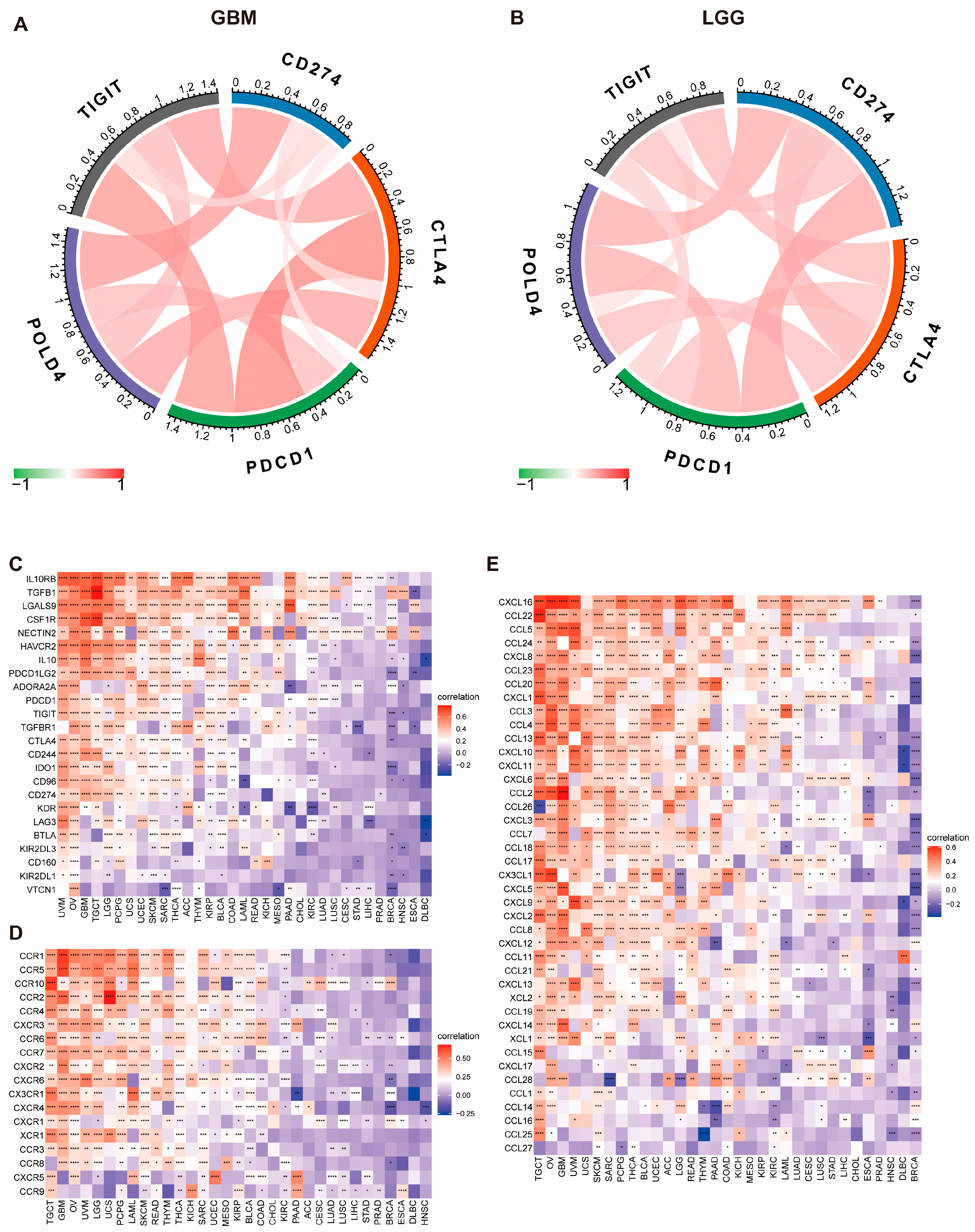
29]. By investigating the ImmuCellAI and TIMER2 databases, we observed the significant impact of POLD4 on immunocyte infiltration in various cancer types, especially in LGG and GBM. Additionally, our study revealed the co-expression of POLD4 with genes encoding immunosuppressive factors, chemokine receptors, and chemokines. These comprehensive findings strongly suggest a close association between POLD4 expression and immune infiltration within tumor cells, consequently influencing patient prognosis and serving as a potential therapeutic target for the development of immunosuppressive treatments.
Cancer cells heavily depend on exhausted T cells to evade the immune system, as these T cells exhibit diminished cytokine levels and impaired effector function [
30,
31,
32]. Numerous studies have demonstrated that targeting the underlying causes of T-cell exhaustion presents a promising therapeutic approach to enhance antitumor immunity [
33,
34]. Therapeutic methods involving blocking antibodies for PD-1/PD-L1 have proven effective in reactivating dormant T cells [
35,
36].
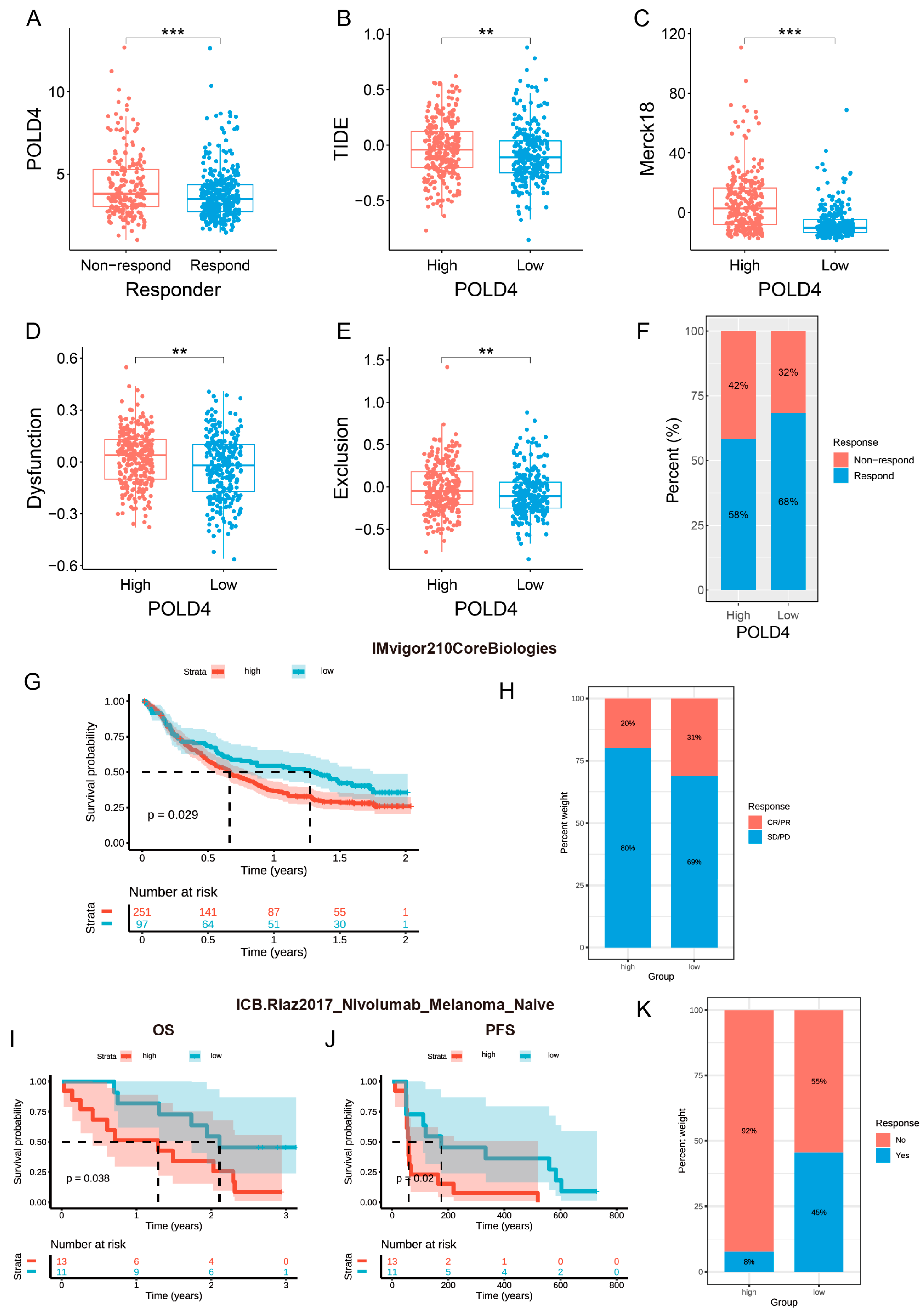
In our study, we discovered a positive correlation between the expression level of POLD4 and the gene markers associated with immune checkpoints. We utilized three datasets, TCGA, CGGA_mRNAseq_325, and CGGA_mRNAseq_693, and performed TIDE analysis to predict the efficacy of immune checkpoint inhibitors in glioma patients. To ensure the reliability of our conclusions, we further corroborated our findings using two clinical datasets, IMvigor210 and GSE91061. Our consistent results demonstrate that patients with low POLD4 expression exhibit a better response to immune checkpoint inhibitors. This finding suggests that high POLD4 expression may create an immune microenvironment that is less responsive to immunotherapy. Consequently, reducing the expression of POLD4 may be a viable combination strategy to enhance the effectiveness of checkpoint inhibitors by alleviating T cell exhaustion, thereby bolstering the antineoplastic response.
According to reports, POLD4 has been found to be associated with DNA damage repair [
9,
26], a critical process that influences the development of drug resistance [
37,
38]. In light of this, our investigation aimed to explore the potential correlation between POLD4 expression and drug sensitivity. The Cancer Therapeutics Response Portal (CTRP) connects the cellular characteristics of cancer cell lines with small molecule sensitivity, supporting the discovery of drugs that can be matched to patients based on predictive biomarkers. Through the Gene Set Cancer Analysis (GSCA) project, we utilized CTRP data online to perform a correlation analysis between POLD4 and drug sensitivity. Our findings indicate that POLD4 is associated with sensitivity to various drugs, particularly positively correlated with the IC50 values of multiple drugs. This intriguing finding suggests that POLD4 may play a pivotal role in mediating resistance to these drugs. Our investigation also revealed a noteworthy negative correlation between the expression of POLD4 and the IC50 values of certain drugs, including PD318088, selumetinib, pluripotin, vandetanib, IC-87114, trametinib, saracatinib, and dasatinib. Such a revelation indicates that heightened POLD4 expression might heighten cellular responsiveness to these drugs, implying a potential avenue for enhancing their therapeutic efficacy for tumor patients with elevated POLD4 levels.
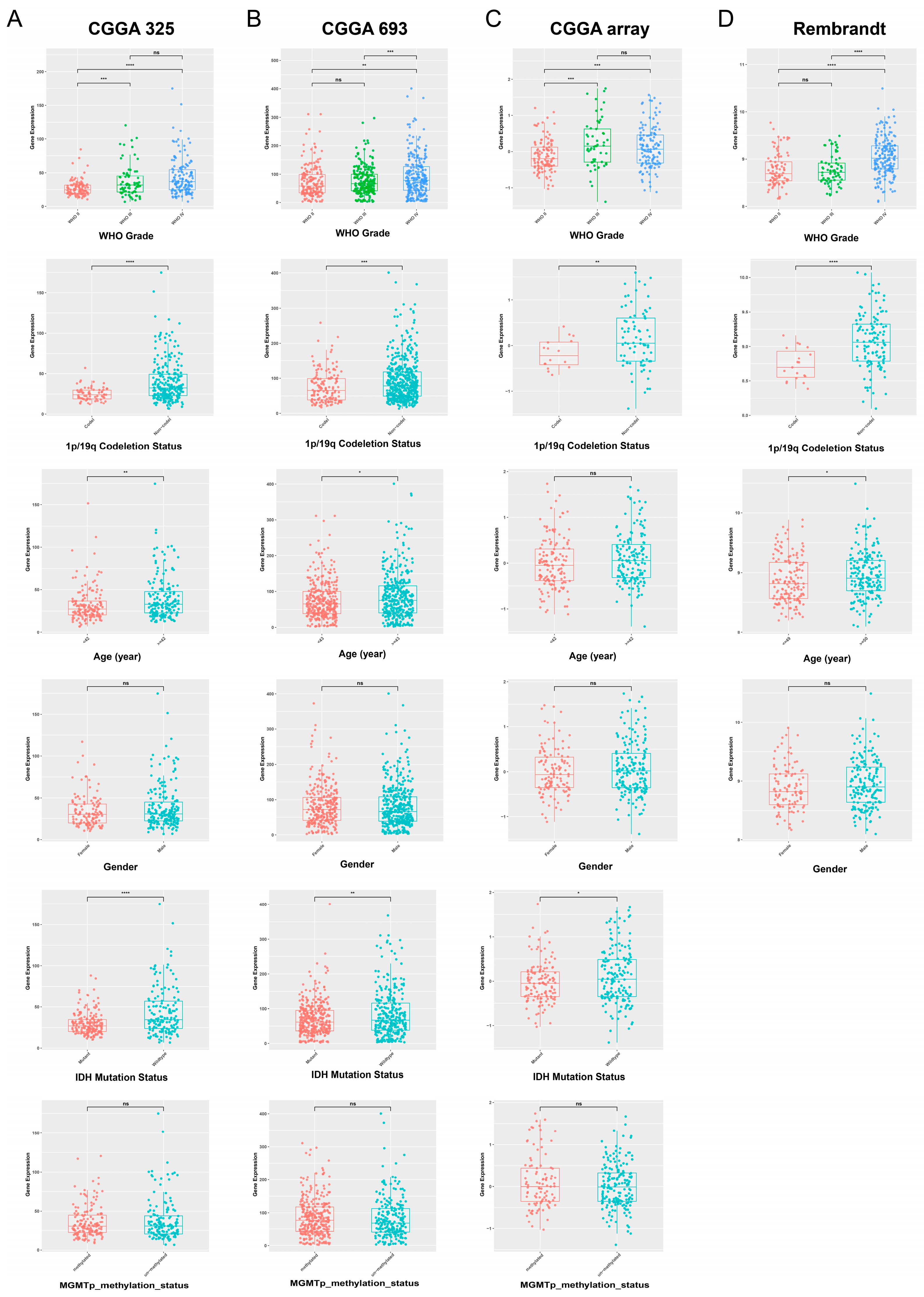
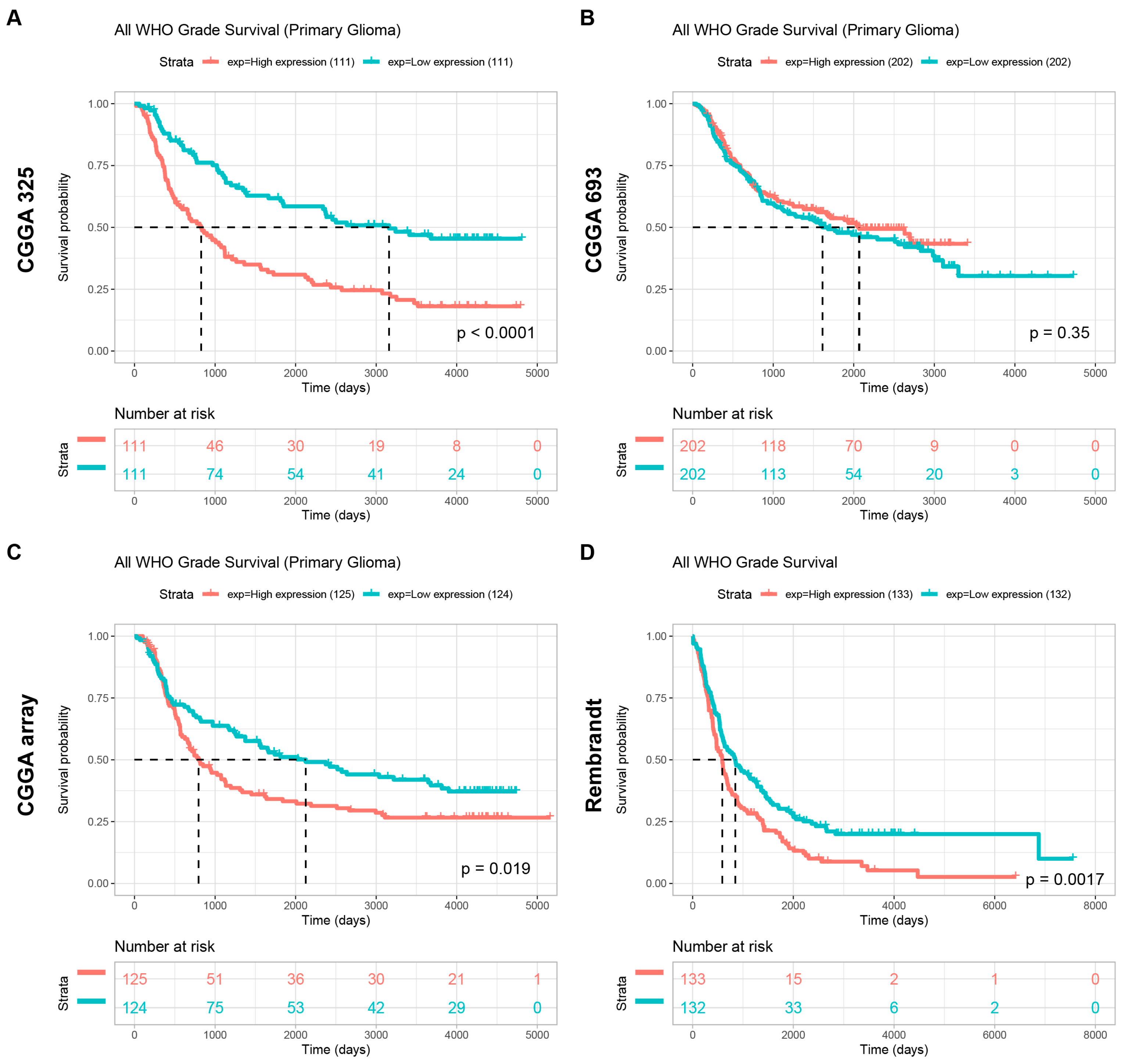
To further explore the potential of POLD4 in glioma, we conducted further analysis on the expression levels of POLD4 in glioma patients with different clinical characteristics using the CGGA_mRNAseq_325, CGGA_mRNAseq_693, CGGA_array_301, and Rembrandt datasets. Additionally, we examined the differences in survival among glioma patients with high and low expression levels of POLD4. Our analysis results demonstrate a significant association between POLD4 expression levels and various malignant clinical features, as well as an impact on patient survival prognosis.
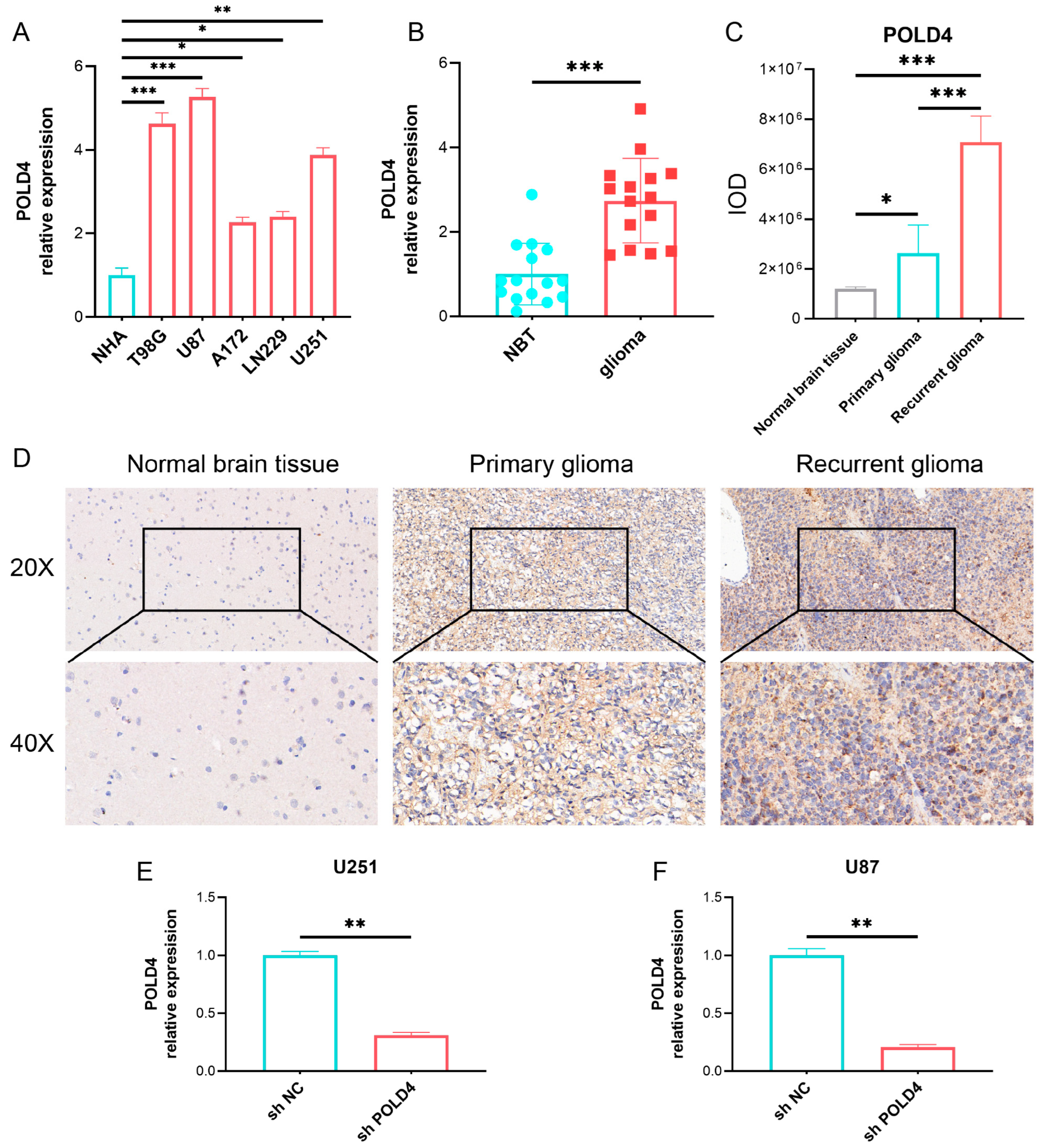
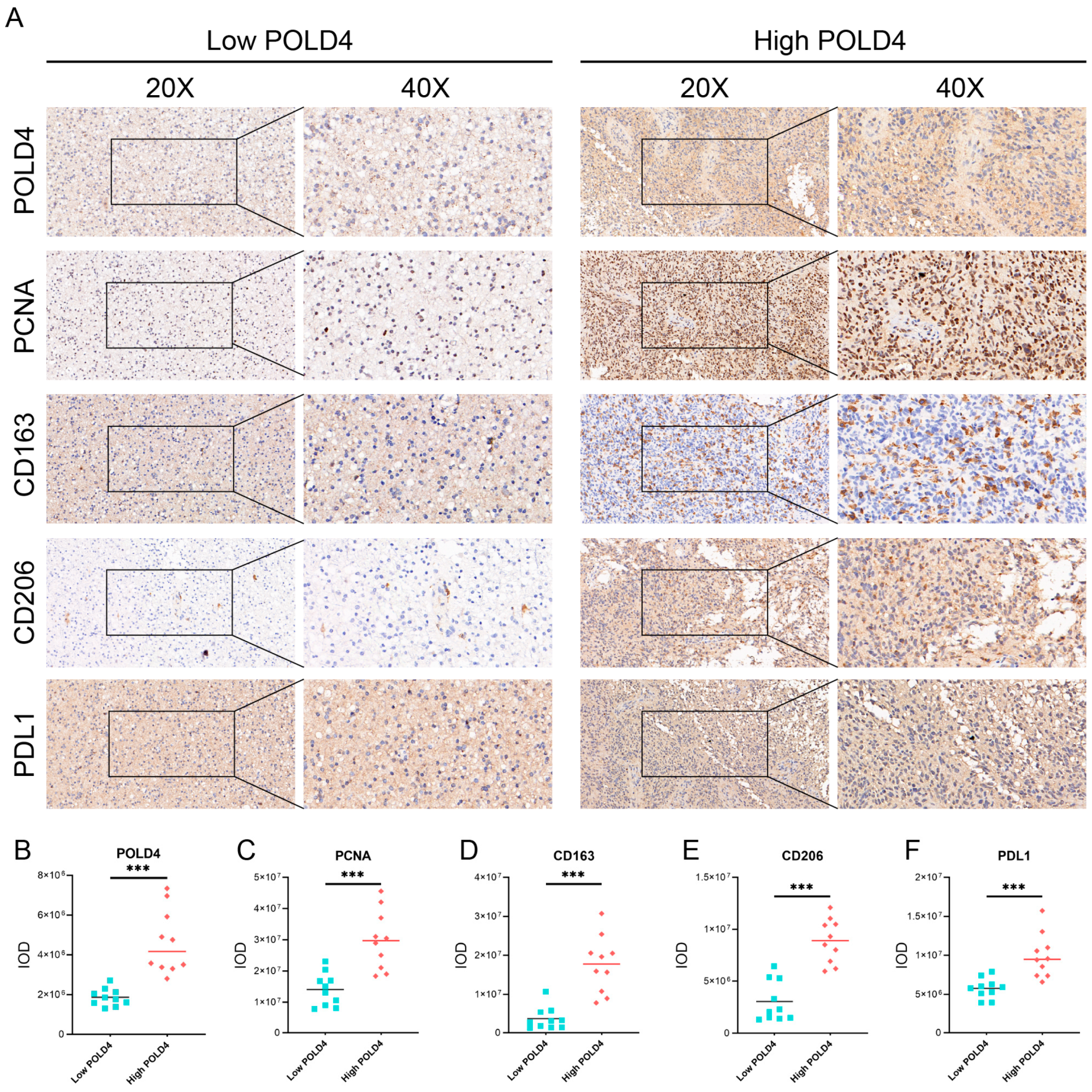
Based on the results of bioinformatic analysis, we proceeded to validate the expression of POLD4 in glioma tissues and cell lines. We found that, compared to normal astrocyte cell lines, POLD4 was expressed at higher levels in various glioma cell lines. Furthermore, compared to normal brain tissues adjacent to the tumor, POLD4 expression was significantly elevated in glioma tissues, particularly in recurrent gliomas. This substantial increase in expression suggests that POLD4 may not only be involved in glioma tumor growth but also play a role in the glioma recurrence process. By conducting knockdown experiments targeting POLD4 in glioma cell lines U251 and U87, we have unraveled compelling evidence underscoring the pivotal role of POLD4 in governing the proliferation of glioma cells. The knockdown of POLD4 exerts a profound effect on cell viability, and the clonogenic potential of glioma cells is markedly reduced upon POLD4 inhibition. Furthermore, the EDU incorporation assay found that POLD4 knockdown led to a significant reduction in EDU uptake, corroborating its role as a player in the regulation of glioma cell proliferation. POLD4 plays a crucial role in DNA replication, and we speculate that this may be a significant mechanism through which POLD4 influences the proliferation of glioma cells. Finally, we conducted a study on the correlation of POLD4 expression with cell proliferation markers PCNA, tumor-associated macrophage markers (CD163, CD206), and the immune checkpoint PDL1 in glioma tissue. Twenty glioma tissue specimens were collected, and based on POLD4 expression, they were divided into the low POLD4 expression group and the high POLD4 expression group. We found that compared to the low POLD4 expression group, the high POLD4 expression group exhibited higher levels of PCNA, CD163, CD206, and PDL1 expression in glioma tissue. This research on tissue specimens further supports the role of POLD4 in glioma cell proliferation and its impact on the glioma immune microenvironment.
In this study, an extensive investigation was conducted to examine the differences in POLD4 expression, prognostic significance, and its underlying functions across 33 different types of human cancers. The results demonstrated a robust link between increased POLD4 expression and adverse clinical outcomes in a diverse range of cancer types. Moreover, the study unearthed noteworthy associations between deviant POLD4 expression and an array of factors, encompassing the tumor microenvironment, efficacy of immunotherapy, drug sensitivity, and glioma cell proliferation (
Figure 15).
To summarize, the elevated expression of POLD4 serves as an indicator of malignant characteristics in tumors such as gliomas, suggesting an unfavorable prognosis and contributing to the formation of an immunosuppressive microenvironment within the tumor. Utilizing a combination of POLD4-targeted cancer immunotherapy methods may offer higher efficacy compared to relying solely on a single treatment approach.
This study has several limitations. Firstly, our analysis heavily depended on publicly available datasets, which may introduce biases and limitations. Secondly, while we conducted experimental validation, we admit that the exact molecular mechanisms underlying POLD4’s role in glioma cell proliferation and the modulation of the immune microenvironment remain incompletely understood. Lastly, the extension of our findings to other tumor types is preliminary, and further research is needed to ascertain the broader implications of POLD4 in cancer biology.
4. Materials and Methods
4.1. Data Collection
The TCGA database via UCSC Xena (accessible at
https://xena.ucsc.edu/, accessed on 5 December 2022) provided us with RNA sequences and clinical data from 11069 samples representing 33 different cancer types. Rembrandt (accessible via
http://gliovis.bioinfo.cnio.es/, accessed on 5 December 2022) and CGGA (accessible via
http://www.cgga.org.cn/, accessed on 5 December 2022) databases are also used to obtain transcriptomic and clinical information of glioma patients. We obtained gene profile data of normal human tissues from GTEx, which can be accessed through
https://commonfund.nih.gov/GTEx, accessed on 5 December 2022. The POLD4 mRNA expression data for numerous cancer cell lines was acquired from the Cancer Cell Line Encyclopedia (CCLE) database, available at
https://portals.broadinstitute.org/ccle/data, accessed on 5 December 2022. The DNA copy number retrieved originates from the cBioPortal database, which can be accessed at
https://www.cbioportal.org/, accessed on 5 December 2022.
4.2. POLD4 Expression Analysis
By integrating and analyzing GTEx and TCGA databases, we conducted a comparison of POLD4 expression in different tumor types and their corresponding normal tissues. In order to obtain a thorough examination of POLD4 expression patterns, data from the TCGA and GTEx databases were used to evaluate POLD4 gene expression in a diverse set of samples, encompassing 31 normal tissues and 33 tumor tissues. We employed the CCLE database to examine POLD4 expression in tumor cell lines. Moreover, we investigated its expression discrepancies between tumors and adjacent normal tissues using data only from the TCGA database.
4.3. Exploration into the Copy Number Alterations
The copy number alterations (CNA) data of POLD4 in patients with various types of cancer were obtained from the cBioPortal database. We analyzed the correlation between the expression and copy number of the POLD4 gene in various types of cancer and presented the results using bar plots. The plotting was performed using the ggplot2 package. Our specific focus was on examining the correlation between POLD4 gene expression and copy number in GBM and LGG cases.
4.4. Prognosis Analysis
The prognostic value of POLD4 in predicting survival indicators across 33 types of cancers was assessed using Univariate Cox regression analyses, and the results are displayed in a forest plot. Moreover, to assess the prognostic significance of POLD4, the survival disparity between the high- and low-expression groups was examined utilizing Kaplan–Meier analysis.
4.5. Function Analysis
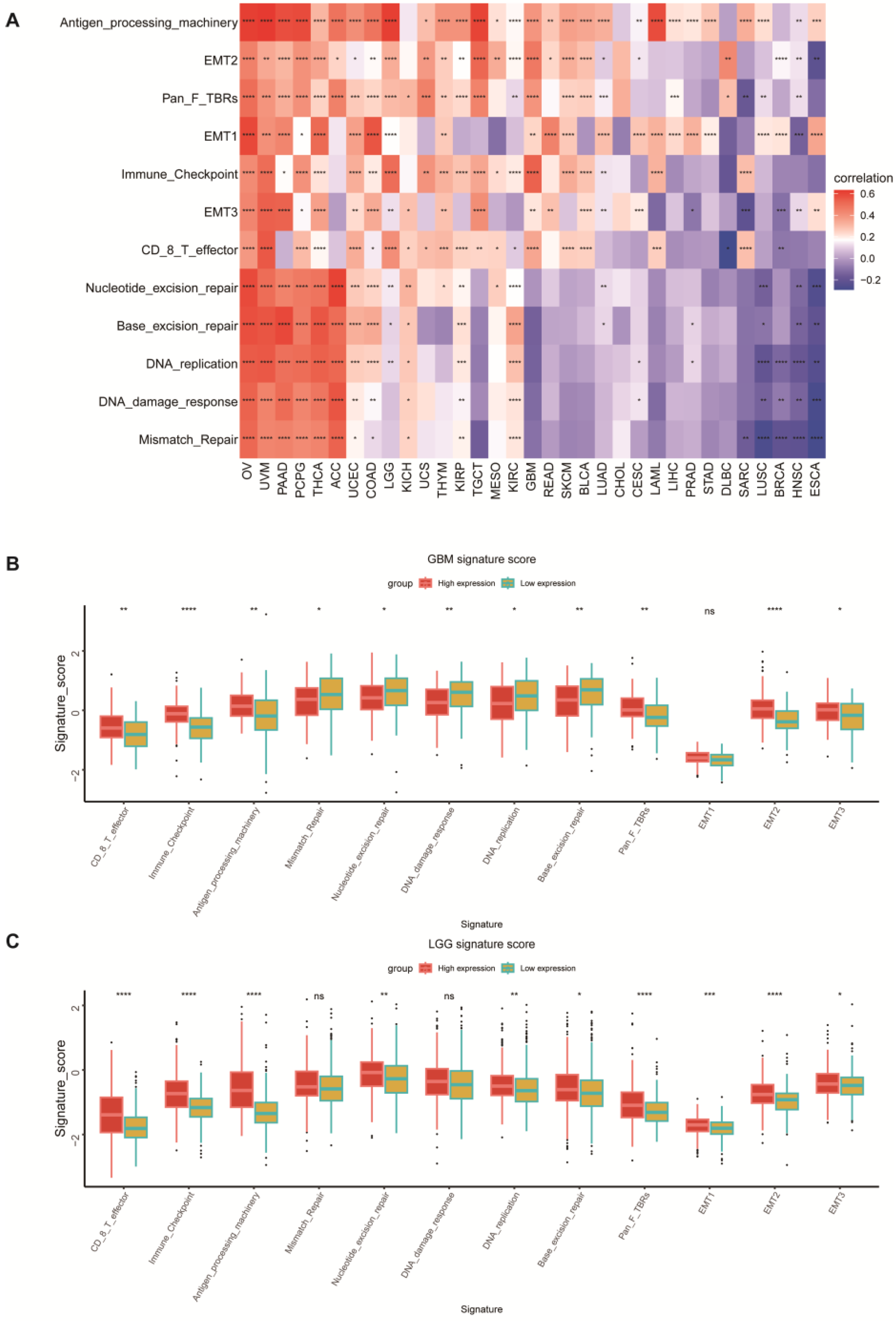
We conducted a correlation analysis between POLD4 and all genes in GBM and LGG separately. Heatmaps were utilized to display the expression of the top 50 genes, both positively and negatively correlated. After conducting correlation analysis, we proceeded with GSEA analysis on the GO, KEGG, and Reactome pathways in GBM and LGG. In addition, a comprehensive GSVA enrichment analysis was performed across various types of cancers. The R language “GSVA” package was utilized to derive the Hallmark pathway scores for each cancer. Visual representation of the correlation between Hallmark pathway scores and POLD4 gene expression was achieved through the use of a heatmap. To present the results in a clear manner, bar charts were employed to analyze the correlation coefficient between POLD4 gene expression and each Hallmark pathway score in GBM and LGG. Moreover, we analyzed the correlation between POLD4 gene expression and immune-relevant signature, mismatch-relevant signature, and stromal-relevant signature, which were derived from a previous literature report [
14]. Principal component analysis was conducted and the first principal component (PC1) was extracted as the signature score representing corresponding biological functions. We explored the impact of POLD4 on the corresponding functions by analyzing the correlation between signature scores and POLD4 expression levels.
4.6. Correlation between POLD4 Expression and Tumor Microenvironment
Using the R “ESTIMATE” package, we calculated the TME score for each patient across 33 cancer types, incorporating the stromal score, immune score, ESTIMATE score, and tumor purity score. To delve deeper into our analysis, we investigated the relationship between the POLD4 expression and these aforementioned scores. Later, we acquired immune cell infiltration scores for TCGA pan-cancer patients using the ImmuCellAI platform (
http://bioinfo.life.hust.edu.cn/web/ImmuCellAI/, accessed on 25 December 2022). We examined the relationship between POLD4 expression and these scores, employing the R “ggplot2” package to generate a visually informative heatmap. Finally, we assessed the relationship between POLD4 gene expression and diverse immune cell infiltrations using the TIMER2 database (
http://timer.comp-genomics.org/, accessed on 25 December 2022), obtaining pan-cancer immune infiltration data.
4.7. Correlation between POLD4 Expression and Immunotherapy Response
We initiated our investigation into the impact of POLD4 expression on immunotherapy by investigating its association with crucial factors, including immune checkpoints, immunosuppressive genes, chemokine receptors, chemokines, and MHC gene expression. These factors have well-established roles in influencing the effectiveness of immunotherapy. Afterwards, we assessed the correlation between POLD4 expression and essential indicators for immunotherapy, such as mDNAsi score, mRNAsi score, tumor mutational burden (TMB), and microsatellite instability (MSI). Additionally, we utilized the Tumor Immune Dysfunction and Exclusion (TIDE) score to evaluate the response of glioma patients to immunotherapy [
39]. In addition, our investigation encompassed an in-depth analysis of a previously documented cohort of clinical studies (IMvigor210 and GSE91061) [
40,
41], aiming to elucidate any plausible association between the levels of POLD4 expression and the response to immunotherapy.
4.8. Drug Sensitivity Analysis
In order to examine the associations between POLD4 and various small-molecule drugs, we leveraged the Cancer Therapeutics Response Portal (CTRP, accessible at
https://portals.broadinstitute.org/ctrp/, accessed on 6 January 2023) datasets to calculate the Pearson correlation coefficients. These coefficients were used to evaluate the relationship between POLD4 expression and the corresponding drug sensitivity.
4.9. Clinical Sample Collection
Clinical samples were collected from glioma patients who underwent surgical resection. Informed consent was obtained from all participants, and the study was conducted following the guidelines and approval of Union Hospital, Tongji Medical College, Huazhong University of Science and Technology.
4.10. Quantitative Real-Time PCR (qPCR)
RNAiso Plus (Takara, 9109) was used to isolate total RNA from the experimental samples, following the manufacturer’s instructions. Subsequently, the extracted RNA was employed to synthesize complementary DNA (cDNA) via a reverse transcription kit (Vazyme, R323). For qPCR, a SYBR Green-based master mix (Vazyme, Q311-02/03) was utilized. The 2−ΔΔCT method was applied to quantify the gene expression levels, with GAPDH serving as the internal control for data normalization. Primers for POLD4 (forward, TGTGAAGAGGAGGGAGGGG, reverse, TGCCAGGCCAGGTCAAACT) and GAPDH (forward, AAAAGCATCACCCGGAGGAGAA, reverse, AAGGAAATGAATGGGCAGCCG) were used.
4.11. Immunohistochemistry
After deparaffinization and rehydration of tissue sections, antigen retrieval was performed. To quench endogenous peroxidase activity, a solution of methanol containing 3% hydrogen peroxide was utilized, while 3% bovine serum albumin was employed to block non-specific binding. Primary antibodies specific to the target proteins were applied to the sections, and after incubation, a secondary antibody (HRP polymer) was used for detection. The stained tissue sections were examined under a microscope.
4.12. Cell Culture and shRNA Interference
Cell lines relevant to the study were cultured in DMEM supplemented with serum and antibiotics. Incubation was carried out in a humidified incubator at 37 °C with 5% CO2. Lentiviral particles containing targeted ShRNA constructs for POLD4 or non-targeting control ShRNA were transduced into the cells for ShRNA interference experiments. Stable knockdown cell lines were established using puromycin selection. Knockdown efficiency was subsequently validated using qPCR.
4.13. Cell Proliferation Assays
CCK-8, colony formation, and EdU staining assays were performed to explore the effect of POLD4 on cell proliferation. Cell viability was assessed using CCK-8 assays (Biosharp, Hefei, China, BS350) following the manufacturer’s instructions. Cells were seeded in 96-well plates and treated with different conditions as required. The absorbance at the specified wavelength was measured using a microplate reader. In the colony formation assay, cells were plated at a low density of 1000 cells/well and allowed to grow for 2 weeks. Afterward, cells were fixed, and stained, and the number of colonies was counted. For EdU staining assays, cells were cultured with EDU, a nucleoside analog of thymidine, which is incorporated into actively replicating DNA. After 1–3 h, the cells were fixed, and the incorporated EDU was detected using fluorescence microscopy.
4.14. Xenograft Mouse Model
To establish an orthotopic glioma model, the researchers used six-week-old female BALB/c nude mice. In brief, we injected a total of 5 × 105 U87MG cells, which stably expressed firefly luciferase (Fluc), into the mouse brain utilizing a stereotactic instrument. Subsequently, the progression of the tumor was monitored on the 30th day post-tumor implantation using bioluminescence imaging (Bruker Corporation, Billerica, MA, USA).
4.15. Statistical Analysis
The gene expression data underwent normalization using a log2 transformation with the formula [TPM (Transcripts per million) + 1]. To compare the two groups, the statistical analysis employed Student’s t-test. For comparisons involving more than two groups, the non-parametric Kruskal–Wallis test and the parametric one-way ANOVA test were utilized. Spearman’s test was employed to analyze the correlation between the two variables. The R software (Version 4.2.2) was used for bioinformatics analysis, while GraphPad Prism 9.0 software was utilized for analyzing the experimental data. A significance level below p < 0.05 was considered statistically significant.