Unlocking Neural Function with 3D In Vitro Models: A Technical Review of Self-Assembled, Guided, and Bioprinted Brain Organoids and Their Applications in the Study of Neurodevelopmental and Neurodegenerative Disorders
Abstract
1. Introduction
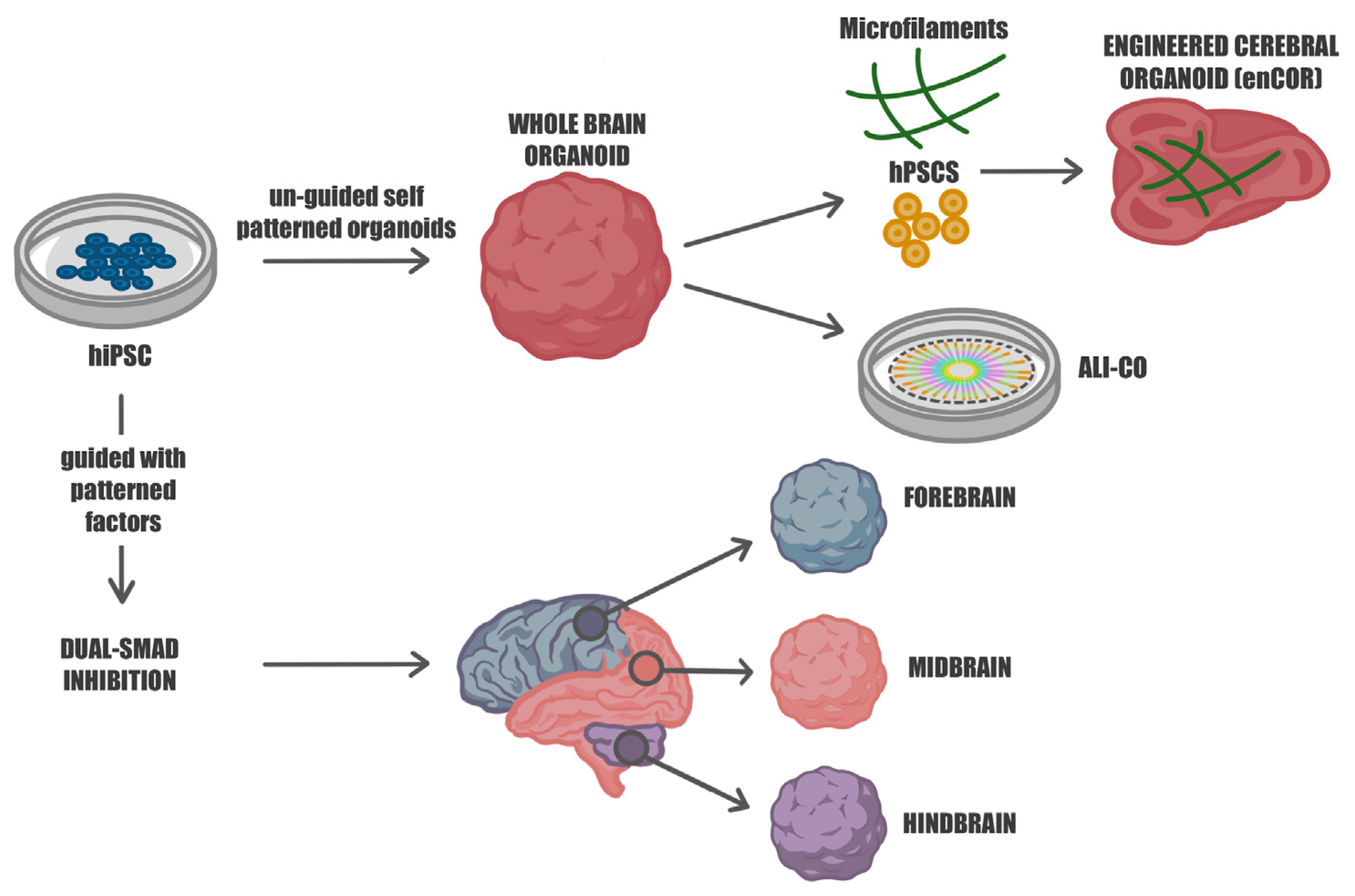
2. From 2D to 3D: The Rise of Cerebral Organoids for Studying Human Brain Development
3. Beyond Batch Syndrome: Guiding Cellular Fate in Brain Organoid Development
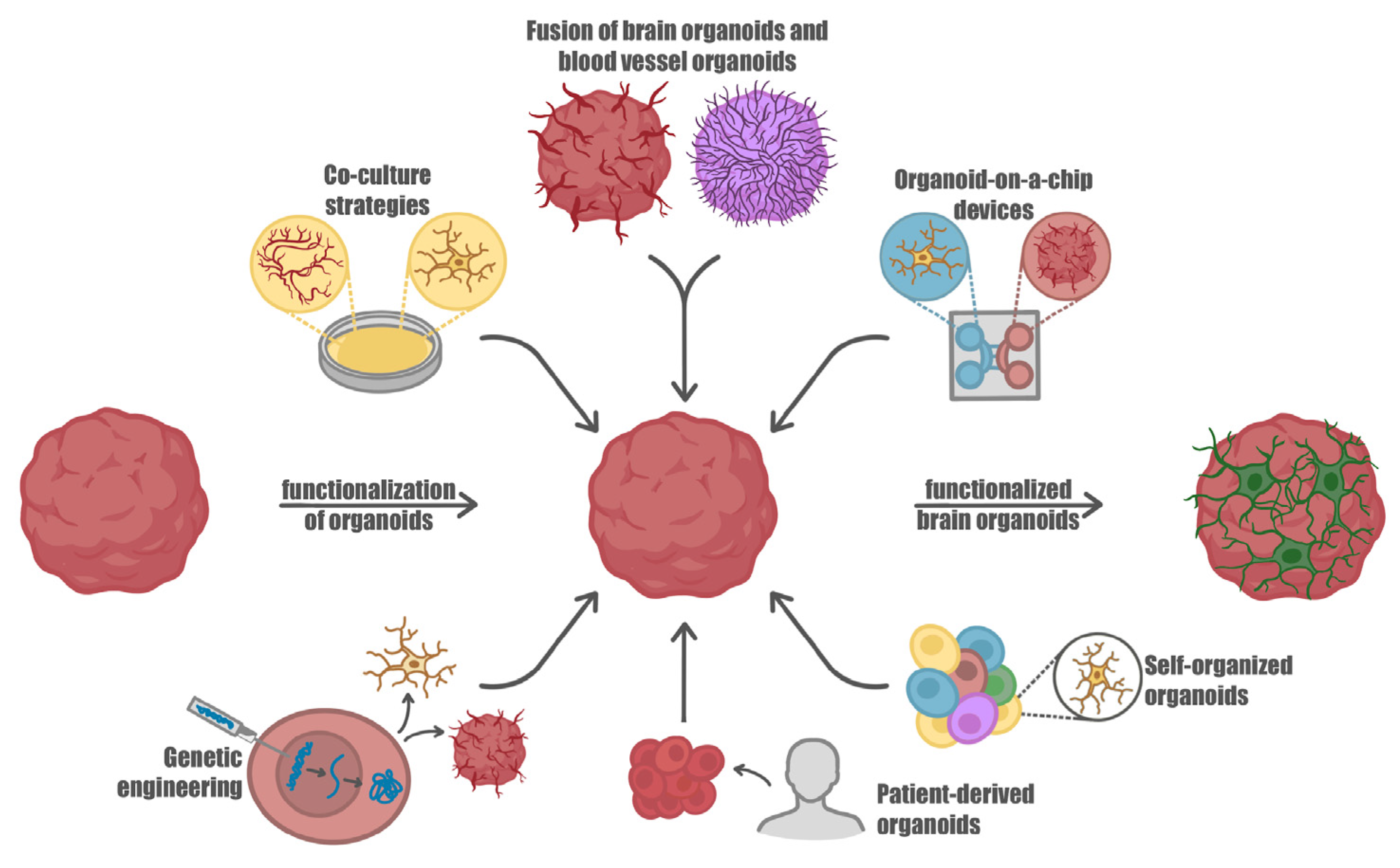
4. Improving Brain Organoids by Introducing Microglia and Vascularization
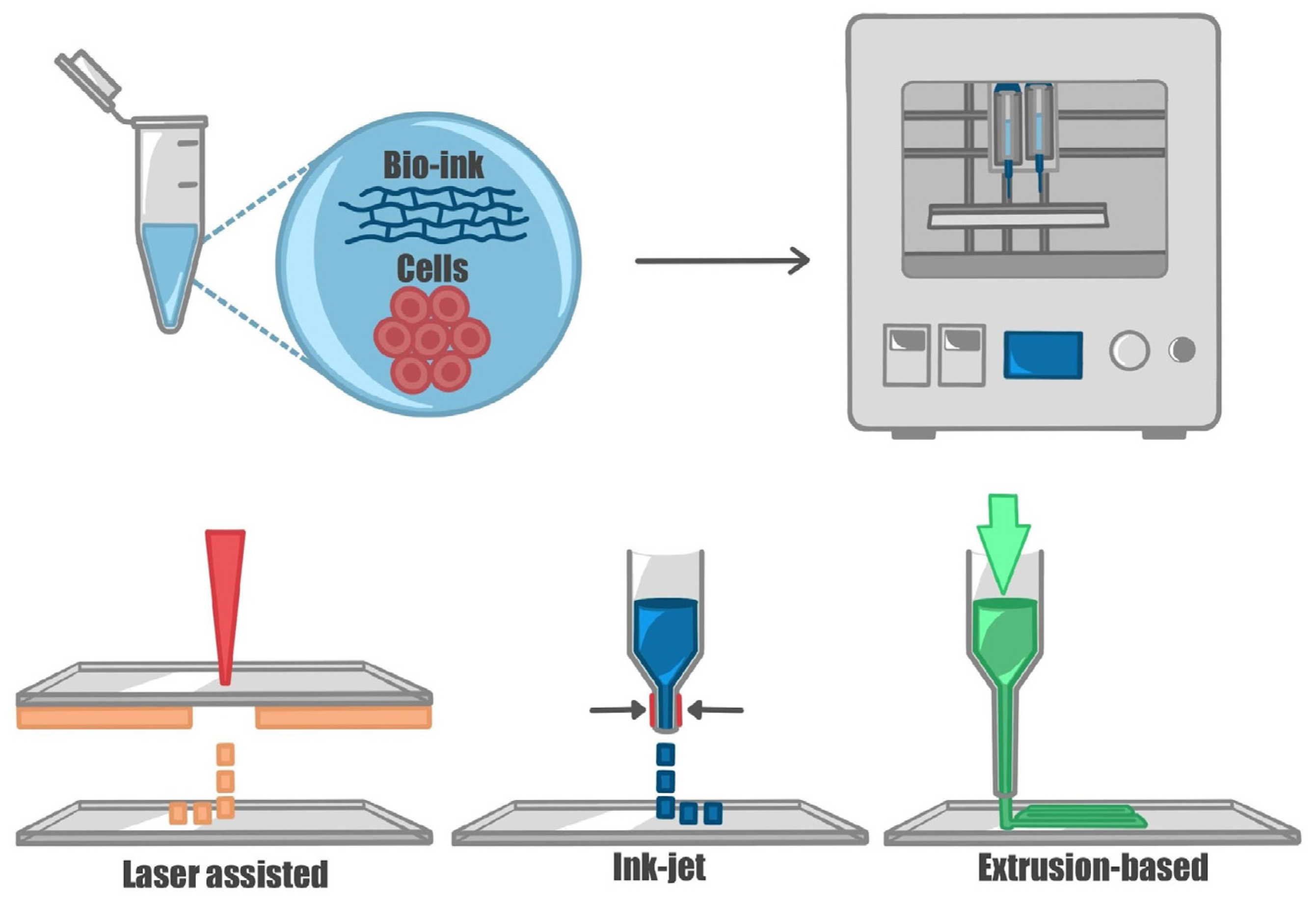
5. Assembling Neural Tissue: Hydrogels and Techniques Used in 3D Bioprinting and Organ-on-a-Chip Technologies
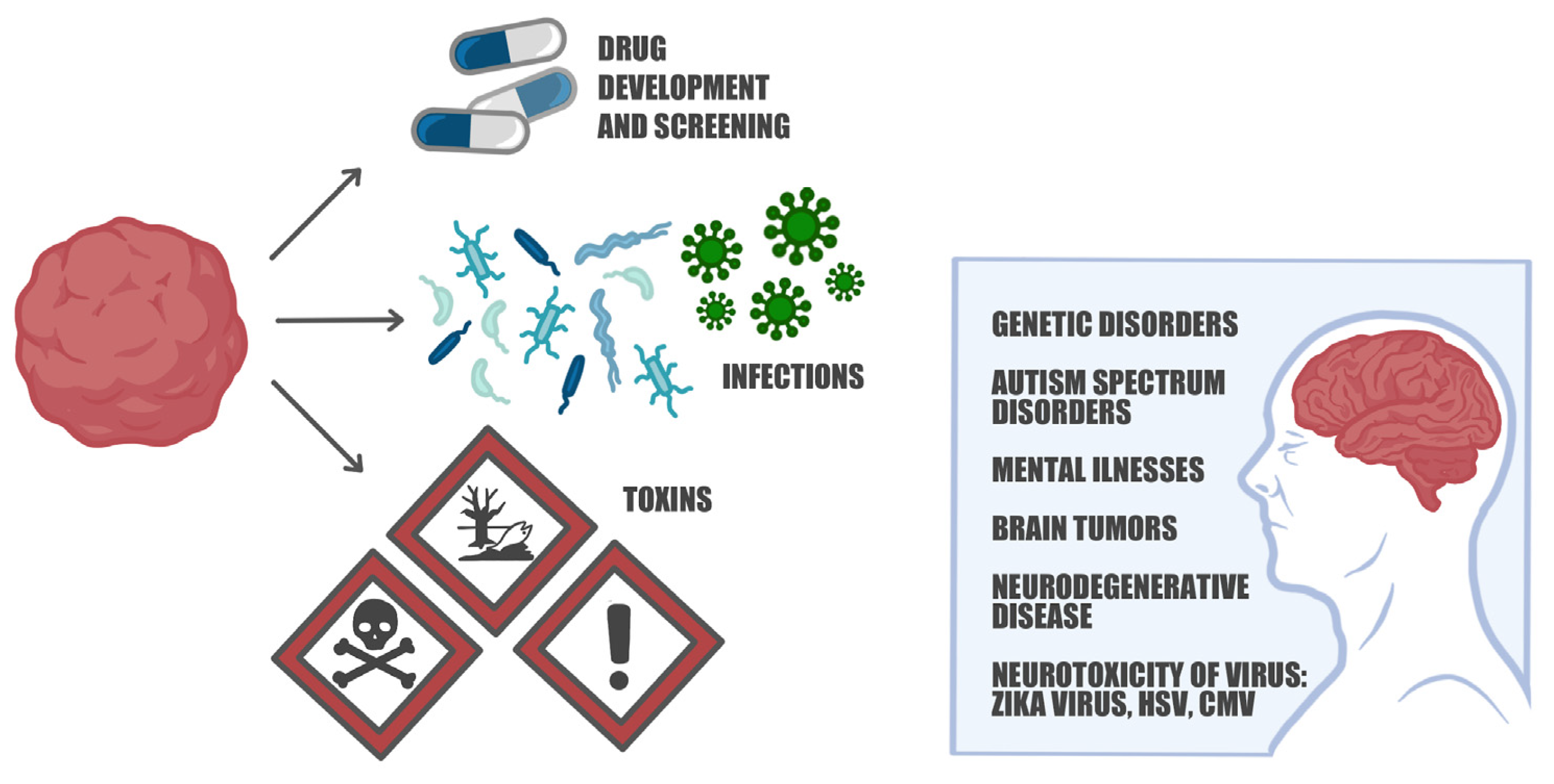
6. The Versatility of 3D Brain Organoids: Modeling Neurodevelopmental and Neurodegenerative Disorders Unraveling Pathophysiological Mechanisms
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Steimberg, N.; Chiono, V.; Era, P.D.; Di Angelantonio, S. iPS, Organoids and 3D Models as Advanced Tools for In Vitro Toxicology. ALTEX Altern. Anim. Exp. 2020, 37, 136–140. [Google Scholar] [CrossRef]
- Muñoz-sanjuán, I.; Brivanlou, A.H. Neural Induction, The Default Model and Embryonic Stem Cells. Nat. Rev. Neurosci. 2002, 3, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Eiraku, M.; Takata, N.; Ishibashi, H.; Kawada, M.; Sakakura, E.; Okuda, S. Self-Organizing Optic-Cup Morphogenesis in Three-Dimensional Culture. Nature 2011, 472, 51–56. [Google Scholar] [CrossRef] [PubMed]
- Eiraku, M.; Watanabe, K.; Matsuo-takasaki, M.; Kawada, M.; Yonemura, S.; Matsumura, M.; Wataya, T.; Nishiyama, A.; Muguruma, K.; Sasai, Y. Article Self-Organized Formation of Polarized Cortical Tissues from ESCs and Its Active Manipulation by Extrinsic Signals. Stem Cell 2008, 3, 519–532. [Google Scholar] [CrossRef]
- Lancaster, M.A.; Renner, M.; Martin, C.A.; Wenzel, D.; Bicknell, L.S.; Hurles, M.E.; Homfray, T.; Penninger, J.M.; Jackson, A.P.; Knoblich, J.A. Cerebral Organoids Model Human Brain Development and Microcephaly. Nature 2013, 501, 373–379. [Google Scholar] [CrossRef]
- Del Dosso, A.; Urenda, J.; Nguyen, T.; Quadrato, G. Primer Upgrading the Physiological Relevance of Human Brain Organoids. Neuron 2020, 107, 1014–1028. [Google Scholar] [CrossRef]
- Di Lullo, E.; Kriegstein, A.R. The Use of Brain Organoids To investigate neural development and disease. Nat. Rev. Neurosci. 2017, 18, 573–584. [Google Scholar] [CrossRef]
- Kelava, I.; Lancaster, M.A. Protocol Review Stem Cell Models of Human Brain Development. Stem Cell 2016, 18, 736–748. [Google Scholar] [CrossRef]
- Qian, X.; Song, H.; Ming, G.L. Brain Organoids: Advances, Applications and Challenges. Development 2019, 146, dev166074. [Google Scholar] [CrossRef]
- Quadrato, G.; Brown, J.; Arlotta, P. The Promises and Challenges of Human Brain Organoids as Models of Neuropsychiatric Disease. Nat. Med. 2016, 22, 1220–1228. [Google Scholar] [CrossRef]
- Xiang, Y.; Cakir, B.; Park, I. Seminars in Cell and Developmental Biology Deconstructing and Reconstructing the Human Brain with Regionally Specified Brain Organoids. Semin. Cell Dev. Biol. 2021, 111, 40–51. [Google Scholar] [CrossRef]
- Pasca, A.M.; Sloan, S.A.; Clarke, L.E.; Tian, Y.; Makinson, C.D.; Huber, N.; Kim, C.H.; Park, J.Y.; O’Rourke, N.A.; Nguyen, K.D.; et al. Functional Cortical Neurons and Astrocytes from Human Pluripotent Stem Cells in 3D Culture. Nat. Methods 2015, 12, 671–678. [Google Scholar] [CrossRef]
- Kadoshima, T.; Sakaguchi, H.; Nakano, T.; Soen, M.; Ando, S.; Eiraku, M.; Sasai, Y. Self-Organization of Axial Polarity, inside-out Layer Pattern, and Species-Specific Progenitor Dynamics in Human ES Cell-Derived Neocortex. Proc. Natl. Acad. Sci. USA 2013, 110, 20284–20289. [Google Scholar] [CrossRef]
- Muguruma, K.; Nishiyama, A.; Kawakami, H.; Hashimoto, K.; Sasai, Y. Self-Organization of Polarized Cerebellar Tissue in 3D Culture of Human Pluripotent Stem Cells. Cell Rep. 2015, 10, 537–550. [Google Scholar] [CrossRef]
- Sakaguchi, H.; Kadoshima, T.; Soen, M.; Narii, N.; Ishida, Y.; Ohgushi, M.; Takahashi, J.; Eiraku, M.; Sasai, Y. Generation of Functional Hippocampal Neurons from Self-Organizing Human Embryonic Stem Cell-Derived Dorsomedial Telencephalic Tissue. Nat. Commun. 2015, 6, 8896. [Google Scholar] [CrossRef]
- Ozone, C.; Suga, H.; Eiraku, M.; Kadoshima, T.; Yonemura, S.; Takata, N.; Oiso, Y.; Tsuji, T.; Sasai, Y. Functional Anterior Pituitary Generated in Self-Organizing Culture of Human Embryonic Stem Cells. Nat. Commun. 2016, 7, 1–10. [Google Scholar] [CrossRef]
- Qian, X.; Nguyen, H.N.; Song, M.M.; Hadiono, C.; Ogden, S.C.; Hammack, C.; Yao, B.; Hamersky, G.R.; Jacob, F.; Zhong, C.; et al. Brain-Region-Specific Organoids Using Mini-Bioreactors for Modeling ZIKV Exposure. Cell 2016, 165, 1238–1254. [Google Scholar] [CrossRef]
- Wataya, T.; Ando, S.; Muguruma, K.; Ikeda, H.; Watanabe, K.; Eiraku, M.; Kawada, M.; Takahashi, J.; Hashimoto, N.; Sasai, Y. Minimization of Exogenous Signals in ES Cell Culture Induces Rostral Hypothalamic Differentiation. Proc. Natl. Acad. Sci. USA 2008, 105, 11796–11801. [Google Scholar] [CrossRef]
- Ogura, T.; Sakaguchi, H.; Miyamoto, S.; Takahashi, J. Three-Dimensional Induction of Dorsal, Intermediate and Ventral Spinal Cord Tissues from Human Pluripotent Stem Cells. Development 2018, 145, dev162214. [Google Scholar] [CrossRef]
- Xiang, Y.; Cakir, B.; Park, I.H. Generation of Regionally Specified Human Brain Organoids Resembling Thalamus Development. STAR Protoc. 2020, 1, 100001. [Google Scholar] [CrossRef]
- Cakir, B.; Xiang, Y.; Tanaka, Y.; Kural, M.H.; Parent, M.; Kang, Y.J.; Chapeton, K.; Patterson, B.; Yuan, Y.; He, C.S.; et al. Engineering of Human Brain Organoids with a Functional Vascular-like System. Nat. Methods 2019, 16, 1169–1175. [Google Scholar] [CrossRef] [PubMed]
- Pellegrini, L.; Bonfio, C.; Chadwick, J.; Begum, F.; Skehel, M.; Lancaster, M.A. Human CNS Barrier-Forming Organoids with Cerebrospinal Fluid Production. Science 2020, 369, eaaz5626. [Google Scholar] [CrossRef] [PubMed]
- Miura, Y.; Li, M.Y.; Birey, F.; Ikeda, K.; Revah, O.; Thete, M.V.; Park, J.Y.; Puno, A.; Lee, S.H.; Porteus, M.H.; et al. Generation of Human Striatal Organoids and Cortico-Striatal Assembloids from Human Pluripotent Stem Cells. Nat. Biotechnol. 2020, 38, 1421–1430. [Google Scholar] [CrossRef] [PubMed]
- Nakano, T.; Ando, S.; Takata, N.; Kawada, M.; Muguruma, K.; Sekiguchi, K.; Saito, K.; Yonemura, S.; Eiraku, M.; Sasai, Y. Self-Formation of Optic Cups and Storable Stratified Neural Retina from Human ESCs. Cell Stem Cell 2012, 10, 771–785. [Google Scholar] [CrossRef] [PubMed]
- Cordella, F.; Ferrucci, L.; D’Antoni, C.; Ghirga, S.; Brighi, C.; Soloperto, A.; Gigante, Y.; Ragozzino, D.; Bezzi, P.; DiAngelantonio, S. Human iPSC-Derived Cortical Neurons Display Homeostatic Plasticity. Life 2022, 12, 1884. [Google Scholar] [CrossRef]
- Garone, M.G.; Birsa, N.; Rosito, M.; Salaris, F.; Mochi, M.; de Turris, V.; Nair, R.R.; Cunningham, T.J.; Fisher, E.M.C.; Morlando, M.; et al. ALS-Related FUS Mutations Alter Axon Growth in Motoneurons and Affect HuD/ELAVL4 and FMRP Activity. Commun. Biol. 2021, 4, 1025. [Google Scholar] [CrossRef]
- Garone, M.G.; D’Antoni, C.; Rosa, A. Culture of Human IPSC-Derived Motoneurons in Compartmentalized Microfluidic Devices and Quantitative Assays for Studying Axonal Phenotypes BT—Stem Cell Assays: Methods and Protocols. In Stem Cell Assays; Kannan, N., Beer, P., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2022; pp. 189–199. ISBN 978-1-0716-1979-7. [Google Scholar]
- Brighi, C.; Salaris, F.; Soloperto, A.; Cordella, F.; Ghirga, S.; de Turris, V.; Rosito, M.; Porceddu, P.F.; D’Antoni, C.; Reggiani, A.; et al. Novel Fragile X Syndrome 2D and 3D Brain Models Based on Human Isogenic FMRP-KO IPSCs. Cell Death Dis. 2021, 12. [Google Scholar] [CrossRef]
- Shi, Y.; Inoue, H.; Wu, J.C.; Yamanaka, S. Induced Pluripotent Stem Cell Technology: A Decade of Progress. Nat. Rev. Drug Discov. 2017, 16, 115–130. [Google Scholar] [CrossRef]
- Mulder, L.A.; Depla, J.A.; Sridhar, A.; Wolthers, K.; Pajkrt, D.; Vieira de Sá, R. A Beginner’s Guide on the Use of Brain Organoids for Neuroscientists: A Systematic Review. Stem Cell Res. Ther. 2023, 14, 87. [Google Scholar] [CrossRef]
- Reynolds, B.A.; Weiss, S. Generation of Neurons and Astrocytes from Isolated Cells of the Adult Mammalian Central Nervous System. Science 1992, 255, 1707–1710. [Google Scholar] [CrossRef]
- Jensen, J.B.; Parmar, M. Strengths and Limitations of the Neurosphere Culture System. Mol. Neurobiol. 2006, 34, 153–161. [Google Scholar] [CrossRef]
- Soares, R.; Ribeiro, F.; Lourenço, D.M.; Rodrigues, R.; Moreira, J.; Sebastião, A.; Morais, V.; Xapelli, S. The Neurosphere Assay: An Effective in Vitro Technique to Study Neural Stem Cells. Neural Regen. Res. 2021, 16, 2229–2231. [Google Scholar] [CrossRef]
- Singec, I.; Knoth, R.; Meyer, R.P.; Maciaczyk, J.; Volk, B.; Nikkhah, G.; Frotscher, M.; Snyder, E.Y. Defining the Actual Sensitivity and Specificity of the Neurosphere Assay in Stem Cell Biology. Nat. Methods 2006, 3, 801–806. [Google Scholar] [CrossRef]
- Gil-Perotín, S.; Duran-Moreno, M.; Cebrián-Silla, A.; Ramírez, M.; García-Belda, P.; García-Verdugo, J.M. Adult Neural Stem Cells from the Subventricular Zone: A Review of the Neurosphere Assay. Anat. Rec. 2013, 296, 1435–1452. [Google Scholar] [CrossRef]
- Merzaban, J.S.; Imitola, J.; Starossom, S.C.; Zhu, B.; Wang, Y.; Lee, J.; Ali, A.J.; Olah, M.; Abuelela, A.F.; Khoury, S.J.; et al. Cell Surface Glycan Engineering of Neural Stem Cells Augments Neurotropism and Improves Recovery in a Murine Model of Multiple Sclerosis. Glycobiology 2015, 25, 1392–1409. [Google Scholar] [CrossRef]
- Rasmussen, S.; Imitola, J.; Ayuso-Sacido, A.; Wang, Y.; Starossom, S.C.; Kivisäkk, P.; Zhu, B.; Meyer, M.; Bronson, R.T.; Garcia-Verdugo, J.M.; et al. Reversible Neural Stem Cell Niche Dysfunction in a Model of Multiple Sclerosis. Ann. Neurol. 2011, 69, 878–891. [Google Scholar] [CrossRef]
- Sheen, V.L.; Ganesh, V.S.; Topcu, M.; Sebire, G.; Bodell, A.; Hill, R.S.; Grant, P.E.; Shugart, Y.Y.; Imitola, J.; Khoury, S.J.; et al. Mutations in ARFGEF2 Implicate Vesicle Trafficking in Neural Progenitor Proliferation and Migration in the Human Cerebral Cortex. Nat. Genet. 2004, 36, 69–76. [Google Scholar] [CrossRef]
- Ganapathee, D.S.; Gunz, P. Insights into Brain Evolution through the Genotype-Phenotype Connection. Prog. Brain Res. 2023, 275, 73–92. [Google Scholar] [CrossRef]
- Lancaster, M.A.; Knoblich, J.A. Generation of Cerebral Organoids from Human Pluripotent Stem Cells. Nat. Protoc. 2014, 9, 2329–2340. [Google Scholar] [CrossRef]
- Renner, M.; Lancaster, M.A.; Bian, S.; Choi, H.; Ku, T.; Peer, A.; Chung, K.; Knoblich, J.A. Self-organized Developmental Patterning and Differentiation in Cerebral Organoids. EMBO J. 2017, 36, 1316–1329. [Google Scholar] [CrossRef]
- Camp, J.G.; Badsha, F.; Florio, M.; Kanton, S.; Gerber, T.; Wilsch-Bräuninger, M.; Lewitus, E.; Sykes, A.; Hevers, W.; Lancaster, M.; et al. Human Cerebral Organoids Recapitulate Gene Expression Programs of Fetal Neocortex Development. Proc. Natl. Acad. Sci. USA 2015, 112, 15672–15677. [Google Scholar] [CrossRef] [PubMed]
- Quadrato, G.; Nguyen, T.; Macosko, E.Z.; Sherwood, J.L.; Yang, S.M.; Berger, D.R.; Maria, N.; Scholvin, J.; Goldman, M.; Kinney, J.P.; et al. Cell Diversity and Network Dynamics in Photosensitive Human Brain Organoids. Nature 2017, 545, 48–53. [Google Scholar] [CrossRef]
- Giandomenico, S.L.; Mierau, S.B.; Gibbons, G.M.; Wenger, L.M.D.; Masullo, L.; Sit, T.; Sutcliffe, M.; Boulanger, J.; Tripodi, M.; Derivery, E.; et al. Cerebral Organoids at the Air–Liquid Interface Generate Diverse Nerve Tracts with Functional Output. Nat. Neurosci. 2019, 22, 669–679. [Google Scholar] [CrossRef] [PubMed]
- Lancaster, M.A.; Corsini, N.S.; Wolfinger, S.; Gustafson, E.H.; Phillips, A.W.; Burkard, T.R.; Otani, T.; Livesey, F.J.; Knoblich, J.A. Guided Self-Organization and Cortical Plate Formation in Human Brain Organoids. Nat. Biotechnol. 2017, 35, 659–666. [Google Scholar] [CrossRef] [PubMed]
- Giandomenico, S.L.; Sutcliffe, M.; Lancaster, M.A. Generation and Long-Term Culture of Advanced Cerebral Organoids for Studying Later Stages of Neural Development. Nat. Protoc. 2021, 16, 579–602. [Google Scholar] [CrossRef]
- Tchieu, J.; Zimmer, B.; Fattahi, F.; Amin, S.; Zeltner, N.; Chen, S.; Studer, L. A Modular Platform for Differentiation of Human PSCs into All Major Ectodermal Lineages. Cell Stem Cell 2017, 21, 399–410.e7. [Google Scholar] [CrossRef]
- Xiang, Y.; Tanaka, Y.; Patterson, B.; Kang, Y.J.; Govindaiah, G.; Roselaar, N.; Cakir, B.; Kim, K.Y.; Lombroso, A.P.; Hwang, S.M.; et al. Fusion of Regionally Specified HPSC-Derived Organoids Models Human Brain Development and Interneuron Migration. Cell Stem Cell 2017, 21, 383–398.e7. [Google Scholar] [CrossRef]
- Allen, N.J.; Eroglu, C. Cell Biology of Astrocyte-Synapse Interactions. Neuron 2017, 96, 697–708. [Google Scholar] [CrossRef]
- Tan, C.X.; Burrus Lane, C.J.; Eroglu, C. Role of Astrocytes in Synapse Formation and Maturation. In Molecular Mechanisms of Neural Development and Insights into Disease, 1st ed.; Elsevier: Amsterdam, The Netherlands, 2021; Volume 142, pp. 371–407. [Google Scholar]
- Zehnder, T.; Petrelli, F.; Romanos, J.; Polleux, F.; Santello, M.; Bezzi, P.; Zehnder, T.; Petrelli, F.; Romanos, J.; Figueiredo, E.C.D.O.; et al. Article Mitochondrial Biogenesis in Developing Astrocytes Regulates Astrocyte Maturation and Synapse Formation Graphical Abstract Ll Mitochondrial Biogenesis in Developing Astrocytes Regulates Astrocyte Maturation and Synapse Formation. Cell Rep. 2021, 35. [Google Scholar] [CrossRef]
- Stogsdill, J.A.; Ramirez, J.; Liu, D.; Kim, Y.H.; Baldwin, K.T.; Enustun, E.; Ejikeme, T.; Ji, R.R.; Eroglu, C. Astrocytic Neuroligins Control Astrocyte Morphogenesis and Synaptogenesis. Nature 2017, 551, 192–197. [Google Scholar] [CrossRef]
- Yakoub, A.M.; Sadek, M. Development and Characterization of Human Cerebral Organoids: An Optimized Protocol. Cell Transplant. 2018, 27, 393–406. [Google Scholar] [CrossRef]
- Pasca, S.P. Assembling Human Brain Organoids. Science 2019, 363, 126–127. [Google Scholar] [CrossRef]
- Basilico, B.; Ferrucci, L.; Ratano, P.; Golia, M.T.; Grimaldi, A.; Rosito, M.; Ferretti, V.; Reverte, I.; Sanchini, C.; Marrone, M.C.; et al. Microglia Control Glutamatergic Synapses in the Adult Mouse Hippocampus. Glia 2022, 70, 173–195. [Google Scholar] [CrossRef] [PubMed]
- Rosito, M.; Sanchini, C.; Gosti, G.; Moreno, M.; De Panfilis, S.; Giubettini, M.; Debellis, D.; Catalano, F.; Peruzzi, G.; Marotta, R.; et al. Microglia Reactivity Entails Microtubule Remodeling from Acentrosomal to Centrosomal Arrays. Cell Rep. 2023, 42, 112104. [Google Scholar] [CrossRef]
- Cordella, F.; Sanchini, C.; Rosito, M.; Ferrucci, L.; Pediconi, N.; Cortese, B.; Guerrieri, F.; Pascucci, G.R.; Antonangeli, F.; Peruzzi, G.; et al. Antibiotics Treatment Modulates Microglia–Synapses Interaction. Cells 2021, 10, 2648. [Google Scholar] [CrossRef]
- Basilico, B.; Pagani, F.; Grimaldi, A.; Cortese, B.; Di Angelantonio, S.; Weinhard, L.; Gross, C.; Limatola, C.; Maggi, L.; Ragozzino, D. Microglia Shape Presynaptic Properties at Developing Glutamatergic Synapses. Glia 2019, 67, 53–67. [Google Scholar] [CrossRef]
- Pagani, F.; Paolicelli, R.C.; Murana, E.; Cortese, B.; Di Angelantonio, S.; Zurolo, E.; Guiducci, E.; Ferreira, T.A.; Garofalo, S.; Catalano, M.; et al. Defective Microglial Development in the Hippocampus of Cx3cr1 Deficient Mice. Front. Cell. Neurosci. 2015, 9, 111. [Google Scholar] [CrossRef]
- Paolicelli, R.C.; Bolasco, G.; Pagani, F.; Maggi, L.; Scianni, M.; Panzanelli, P.; Giustetto, M.; Ferreira, T.A.; Guiducci, E.; Dumas, L.; et al. Synaptic Pruning by Microglia Is Necessary for Normal Brain Development. Science 2011, 333, 1456–1458. [Google Scholar] [CrossRef]
- Salter, M.W.; Stevens, B. Microglia Emerge as Central Players in Brain Disease. Nat. Med. 2017, 23, 1018–1027. [Google Scholar] [CrossRef]
- Ormel, P.R.; Vieira de Sá, R.; van Bodegraven, E.J.; Karst, H.; Harschnitz, O.; Sneeboer, M.A.M.; Johansen, L.E.; van Dijk, R.E.; Scheefhals, N.; Berdenis van Berlekom, A.; et al. Microglia Innately Develop within Cerebral Organoids. Nat. Commun. 2018, 9, 4167. [Google Scholar] [CrossRef]
- Abud, E.M.; Ramirez, R.N.; Martinez, E.S.; Healy, L.M.; Nguyen, C.H.H.; Newman, S.A.; Yeromin, A.V.; Scarfone, V.M.; Marsh, S.E.; Fimbres, C.; et al. IPSC-Derived Human Microglia-like Cells to Study Neurological Diseases. Neuron 2017, 94, 278–293.e9. [Google Scholar] [CrossRef]
- Song, L.; Yuan, X.; Jones, Z.; Vied, C.; Miao, Y.; Marzano, M.; Hua, T.; Sang, Q.X.A.; Guan, J.; Ma, T.; et al. Functionalization of Brain Region-Specific Spheroids with Isogenic Microglia-like Cells. Sci. Rep. 2019, 9, 11055. [Google Scholar] [CrossRef] [PubMed]
- Ao, Z.; Cai, H.; Wu, Z.; Song, S.; Karahan, H.; Kim, B.; Lu, H.C.; Kim, J.; Mackie, K.; Guo, F. Tubular Human Brain Organoids to Model Microglia-Mediated Neuroinflammation. Lab Chip 2021, 21, 2751–2762. [Google Scholar] [CrossRef] [PubMed]
- Cakir, B.; Tanaka, Y.; Kiral, F.R.; Xiang, Y.; Dagliyan, O.; Wang, J.; Lee, M.; Greaney, A.M.; Yang, W.S.; duBoulay, C.; et al. Expression of the Transcription Factor PU.1 Induces the Generation of Microglia-like Cells in Human Cortical Organoids. Nat. Commun. 2022, 13, 430. [Google Scholar] [CrossRef]
- Pham, M.T.; Pollock, K.M.; Rose, M.D.; Cary, W.A.; Stewart, H.R.; Zhou, P.; Nolta, J.A.; Waldau, B. Generation of Human Vascularized Brain Organoids. Neuroreport 2018, 29, 588–593. [Google Scholar] [CrossRef]
- Shi, Y.; Sun, L.; Wang, M.; Liu, J.; Zhong, S.; Li, R.; Li, P.; Guo, L.; Fang, A.; Chen, R.; et al. Vascularized Human Cortical Organoids (VOrganoids) Model Cortical Development In Vivo. PLoS Biol. 2020, 18, e3000705. [Google Scholar] [CrossRef]
- Ahn, Y.; An, J.H.; Yang, H.J.; Lee, D.G.; Kim, J.; Koh, H.; Park, Y.H.; Song, B.S.; Sim, B.W.; Lee, H.J.; et al. Human Blood Vessel Organoids Penetrate Human Cerebral Organoids and Form a Vessel-like System. Cells 2021, 10, 2036. [Google Scholar] [CrossRef]
- Sun, X.Y.; Ju, X.C.; Li, Y.; Zeng, P.M.; Wu, J.; Zhou, Y.Y.; Shen, L.B.; Dong, J.; Chen, Y.J.; Luo, Z.G. Generation of Vascularized Brain Organoids to Study Neurovascular Interactions. eLife 2022, 11, e76707. [Google Scholar] [CrossRef]
- Salmon, I.; Grebenyuk, S.; Abdel Fattah, A.R.; Rustandi, G.; Pilkington, T.; Verfaillie, C.; Ranga, A. Engineering Neurovascular Organoids with 3D Printed Microfluidic Chips. Lab Chip 2022, 22, 1615–1629. [Google Scholar] [CrossRef]
- Lee, S.; Park, C.; Han, J.W.; Kim, J.Y.; Cho, K.; Kim, E.J.; Kim, S.; Lee, S.J.; Oh, S.Y.; Tanaka, Y.; et al. Direct Reprogramming of Human Dermal Fibroblasts into Endothelial Cells Using ER71/ETV2. Circ. Res. 2017, 120, 848–861. [Google Scholar] [CrossRef]
- Iafrate, L.; Benedetti, M.C.; Donsante, S.; Rosa, A.; Corsi, A.; Oreffo, R.O.C.; Riminucci, M.; Ruocco, G.; Scognamiglio, C.; Cidonio, G. Modelling Skeletal Pain Harnessing Tissue Engineering. Vitr. Model. 2022, 1, 289–307. [Google Scholar] [CrossRef]
- Reid, J.A.; Mollica, P.A.; Johnson, G.D.; Ogle, R.C.; Bruno, R.D.; Sachs, P.C. Accessible Bioprinting: Adaptation of a Low-Cost 3D-Printer for Precise Cell Placement and Stem Cell Differentiation. Biofabrication 2016, 8, 025017. [Google Scholar] [CrossRef] [PubMed]
- Gu, Q.; Tomaskovic-Crook, E.; Lozano, R.; Chen, Y.; Kapsa, R.M.; Zhou, Q.; Wallace, G.G.; Crook, J.M. Functional 3D Neural Mini-Tissues from Printed Gel-Based Bioink and Human Neural Stem Cells. Adv. Healthc. Mater. 2016, 5, 1429–1438. [Google Scholar] [CrossRef]
- Nguyen, D.; Hgg, D.A.; Forsman, A.; Ekholm, J.; Nimkingratana, P.; Brantsing, C.; Kalogeropoulos, T.; Zaunz, S.; Concaro, S.; Brittberg, M.; et al. Cartilage Tissue Engineering by the 3D Bioprinting of IPS Cells in a Nanocellulose/Alginate Bioink. Sci. Rep. 2017, 7, 658. [Google Scholar] [CrossRef] [PubMed]
- De la Vega, L.; Abelseth, L.; Sharma, R.; Triviño-Paredes, J.; Restan, M.; Willerth, S.M. 3D Bioprinting Human-Induced Pluripotent Stem Cells and Drug-Releasing Microspheres to Produce Responsive Neural Tissues. Adv. NanoBiomed Res. 2021, 1, 2000077. [Google Scholar] [CrossRef]
- Koch, L.; Deiwick, A.; Franke, A.; Schwanke, K.; Haverich, A.; Zweigerdt, R.; Chichkov, B. Laser Bioprinting of Human Induced Pluripotent Stem Cells—The Effect of Printing and Biomaterials on Cell Survival, Pluripotency, and Differentiation. Biofabrication 2018, 10, 035005. [Google Scholar] [CrossRef]
- Faulkner-Jones, A.; Fyfe, C.; Cornelissen, D.J.; Gardner, J.; King, J.; Courtney, A.; Shu, W. Bioprinting of Human Pluripotent Stem Cells and Their Directed Differentiation into Hepatocyte-like Cells for the Generation of Mini-Livers in 3D. Biofabrication 2015, 7, 044102. [Google Scholar] [CrossRef]
- Ma, X.; Qu, X.; Zhu, W.; Li, Y.S.; Yuan, S.; Zhang, H.; Liu, J.; Wang, P.; Lai, C.S.E.; Zanella, F.; et al. Deterministically Patterned Biomimetic Human IPSC-Derived Hepatic Model via Rapid 3D Bioprinting. Proc. Natl. Acad. Sci. USA 2016, 113, 2206–2211. [Google Scholar] [CrossRef]
- Yu, C.; Ma, X.; Zhu, W.; Wang, P.; Miller, K.L.; Stupin, J.; Koroleva-Maharajh, A.; Hairabedian, A.; Chen, S. Scanningless and Continuous 3D Bioprinting of Human Tissues with Decellularized Extracellular Matrix. Biomaterials 2019, 194, 1–13. [Google Scholar] [CrossRef]
- Ong, C.S.; Fukunishi, T.; Zhang, H.; Huang, C.Y.; Nashed, A.; Blazeski, A.; Disilvestre, D.; Vricella, L.; Conte, J.; Tung, L.; et al. Biomaterial-Free Three-Dimensional Bioprinting of Cardiac Tissue Using Human Induced Pluripotent Stem Cell Derived Cardiomyocytes. Sci. Rep. 2017, 7, 2–12. [Google Scholar] [CrossRef]
- Moldovan, L.; Barnard, A.; Gil, C.H.; Lin, Y.; Grant, M.B.; Yoder, M.C.; Prasain, N.; Moldovan, N.I. IPSC-Derived Vascular Cell Spheroids as Building Blocks for Scaffold-Free Biofabrication. Biotechnol. J. 2017, 12, 1700444. [Google Scholar] [CrossRef]
- Sorkio, A.; Koch, L.; Koivusalo, L.; Deiwick, A.; Miettinen, S.; Chichkov, B.; Skottman, H. Human Stem Cell Based Corneal Tissue Mimicking Structures Using Laser-Assisted 3D Bioprinting and Functional Bioinks. Biomaterials 2018, 171, 57–71. [Google Scholar] [CrossRef]
- Joung, D.; Truong, V.; Neitzke, C.C.; Guo, S.Z.; Walsh, P.J.; Monat, J.R.; Meng, F.; Park, S.H.; Dutton, J.R.; Parr, A.M.; et al. 3D Printed Stem-Cell Derived Neural Progenitors Generate Spinal Cord Scaffolds. Adv. Funct. Mater. 2018, 28, 1801850. [Google Scholar] [CrossRef] [PubMed]
- De la Vega, L.; Gómez, D.A.R.; Abelseth, E.; Abelseth, L.; da Silva, V.A.; Willerth, S.M. 3D Bioprinting Human Induced Pluripotent Stem Cell-Derived Neural Tissues Using a Novel Lab-on-a-Printer Technology. Appl. Sci. 2018, 8, 2414. [Google Scholar] [CrossRef]
- Kador, K.E.; Grogan, S.P.; Dorthé, E.W.; Venugopalan, P.; Malek, M.F.; Goldberg, J.L.; D’Lima, D.D. Control of Retinal Ganglion Cell Positioning and Neurite Growth: Combining 3D Printing with Radial Electrospun Scaffolds. Tissue Eng. Part A 2016, 22, 286–294. [Google Scholar] [CrossRef] [PubMed]
- Lozano, R.; Stevens, L.; Thompson, B.C.; Gilmore, K.J.; Gorkin, R.; Stewart, E.M.; in het Panhuis, M.; Romero-Ortega, M.; Wallace, G.G. 3D Printing of Layered Brain-like Structures Using Peptide Modified Gellan Gum Substrates. Biomaterials 2015, 67, 264–273. [Google Scholar] [CrossRef]
- Lee, W.; Pinckney, J.; Lee, V.; Lee, J.H.; Fischer, K.; Polio, S.; Park, J.K.; Yoo, S.S. Three-Dimensional Bioprinting of Rat Embryonic Neural Cells. Neuroreport 2009, 20, 798–803. [Google Scholar] [CrossRef]
- Zhu, W.; Harris, B.T.; Zhang, L.G. Gelatin Methacrylamide Hydrogel with Graphene Nanoplatelets for Neural Cell-Laden 3D Bioprinting. In Proceedings of the 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Orlando, FL, USA, 16–20 August 2016; pp. 4185–4188. [Google Scholar] [CrossRef]
- Salaris, F.; Colosi, C.; Brighi, C.; Soloperto, A.; de Turris, V.; Benedetti, M.C.; Ghirga, S.; Rosito, M.; Di Angelantonio, S.; Rosa, A. 3D Bioprinted Human Cortical Neural Constructs Derived from Induced Pluripotent Stem Cells. J. Clin. Med. 2019, 8, 1595. [Google Scholar] [CrossRef]
- Dao, D.T.; Mahon, P.B.; Cai, X.; Kovacsics, C.E.; Blackwell, R.A.; Arad, M.; Shi, J.; Zandi, P.P.; Donnell, P.O.; Genome, B.; et al. Mood Disorder Susceptibility Gene CACNA1C Modifies Mood-Related Behaviors in Mice and Interacts with Sex to Influence Behavior in Mice and Diagnosis in Humans. Biol. Psychiatry 2021, 68, 801–810. [Google Scholar] [CrossRef]
- Nishiyama, Y.; Nakamura, M.; Henmi, C.; Yamaguchi, K.; Mochizuki, S.; Nakagawa, H.; Takiura, K. Development of a Three-Dimensional Bioprinter: Construction of Cell Supporting Structures Using Hydrogel and State-of-the-Art Inkjet Technology. J. Biomech. Eng. 2009, 131, 1–6. [Google Scholar] [CrossRef]
- Hewes, S.; Wong, A.D.; Searson, P.C. Bioprinting Microvessels Using an Inkjet Printer. Bioprinting 2017, 7, 14–18. [Google Scholar] [CrossRef]
- Solis, L.H.; Ayala, Y.; Portillo, S.; Varela-Ramirez, A.; Aguilera, R.; Boland, T. Thermal Inkjet Bioprinting Triggers the Activation of the VEGF Pathway in Human Microvascular Endothelial Cells In Vitro. Biofabrication 2019, 11, 045005. [Google Scholar] [CrossRef]
- Liu, W.; Zhang, Y.S.; Heinrich, M.A.; De Ferrari, F.; Jang, H.L.; Bakht, S.M.; Alvarez, M.M.; Yang, J.; Li, Y.C.; Trujillo-de Santiago, G.; et al. Rapid Continuous Multimaterial Extrusion Bioprinting. Adv. Mater. 2017, 29, 1604630. [Google Scholar] [CrossRef]
- Andersen, T.; Auk-Emblem, P.; Dornish, M. 3D Cell Culture in Alginate Hydrogels. Microarrays 2015, 4, 133–161. [Google Scholar] [CrossRef]
- Cidonio, G.; Glinka, M.; Dawson, J.I.; Oreffo, R.O.C. The Cell in the Ink: Improving Biofabrication by Printing Stem Cells for Skeletal Regenerative Medicine. Biomaterials 2019, 209, 10–24. [Google Scholar] [CrossRef]
- Colosi, C.; Shin, S.R.; Manoharan, V.; Massa, S.; Costantini, M.; Barbetta, A.; Dokmeci, M.R.; Dentini, M.; Khademhosseini, A. Microfluidic Bioprinting of Heterogeneous 3D Tissue Constructs Using Low-Viscosity Bioink. Adv. Mater. 2016, 28, 677–684a. [Google Scholar] [CrossRef]
- Costantini, M.; Colosi, C.; Świȩszkowski, W.; Barbetta, A. Co-Axial Wet-Spinning in 3D Bioprinting: State of the Art and Future Perspective of Microfluidic Integration. Biofabrication 2019, 11, 012001. [Google Scholar] [CrossRef]
- Hsieh, F.Y.; Lin, H.H.; Hsu, S. hui 3D Bioprinting of Neural Stem Cell-Laden Thermoresponsive Biodegradable Polyurethane Hydrogel and Potential in Central Nervous System Repair. Biomaterials 2015, 71, 48–57. [Google Scholar] [CrossRef]
- Davoodi, E.; Sarikhani, E.; Montazerian, H.; Ahadian, S.; Costantini, M.; Swieszkowski, W.; Willerth, S.M.; Walus, K.; Mofidfar, M.; Toyserkani, E.; et al. Extrusion and Microfluidic-Based Bioprinting to Fabricate Biomimetic Tissues and Organs. Adv. Mater. Technol. 2020, 5, 1901044. [Google Scholar] [CrossRef]
- Silvani, G.; Scognamiglio, C.; Caprini, D.; Marino, L.; Chinappi, M.; Sinibaldi, G.; Peruzzi, G.; Kiani, M.F.; Casciola, C.M. Reversible Cavitation-Induced Junctional Opening in an Artificial Endothelial Layer. Small 2019, 15, 1905375. [Google Scholar] [CrossRef]
- Grisanti, G.; Caprini, D.; Sinibaldi, G.; Scognamiglio, C.; Silvani, G.; Peruzzi, G.; Casciola, C.M. A Microfluidic Platform for Cavitation-Enhanced Drug Delivery. Micromachines 2021, 12, 658. [Google Scholar] [CrossRef] [PubMed]
- Serex, L.; Sharma, K.; Rizov, V.; Bertsch, A.; McKinney, J.D.; Renaud, P. Microfluidic-Assisted Bioprinting of Tissues and Organoids at High Cell Concentrations. Biofabrication 2021, 13, 025006. [Google Scholar] [CrossRef] [PubMed]
- Marcotulli, M.; Tirelli, M.C.; Volpi, M.; Jaroszewicz, J.; Scognamiglio, C.; Kasprzycki, P.; Karnowski, K.; Święszkowski, W.; Ruocco, G.; Costantini, M.; et al. Microfluidic 3D Printing of Emulsion Ink for Engineering Porous Functionally Graded Materials. Adv. Mater. Technol. 2023, 8, 2201244. [Google Scholar] [CrossRef]
- Van der Heide, D.; Cidonio, G.; Stoddart, M.J.; D’Este, M. 3D Printing of Inorganic-Biopolymer Composites for Bone Regeneration. Biofabrication 2022, 14, 042003. [Google Scholar] [CrossRef]
- Okesola, B.O.; Mendoza-Martinez, A.K.; Cidonio, G.; Derkus, B.; Boccorh, D.K.; Osuna De La Peña, D.; Elsharkawy, S.; Wu, Y.; Dawson, J.I.; Wark, A.W.; et al. De Novo Design of Functional Coassembling Organic-Inorganic Hydrogels for Hierarchical Mineralization and Neovascularization. ACS Nano 2021, 15, 11202–11217. [Google Scholar] [CrossRef]
- Cidonio, G.; Costantini, M.; Pierini, F.; Scognamiglio, C.; Agarwal, T.; Barbetta, A. 3D Printing of Biphasic Inks: Beyond Single-Scale Architectural Control. J. Mater. Chem. C 2021, 9, 12489–12508. [Google Scholar] [CrossRef]
- Antill-O’Brien, N.; Bourke, J.; O’Connell, C.D. Layer-by-Layer: The Case for 3D Bioprinting Neurons to Create Patient-Specific Epilepsy Models. Materials 2019, 12, 3218. [Google Scholar] [CrossRef]
- Qiu, B.; Bessler, N.; Figler, K.; Buchholz, M.B.; Rios, A.C.; Malda, J.; Levato, R.; Caiazzo, M. Bioprinting Neural Systems to Model Central Nervous System Diseases. Adv. Funct. Mater. 2020, 30, 1910250. [Google Scholar] [CrossRef]
- Brassard, J.A.; Nikolaev, M.; Hübscher, T.; Hofer, M.; Lutolf, M.P. Recapitulating Macro-Scale Tissue Self-Organization through Organoid Bioprinting. Nat. Mater. 2021, 20, 22–29. [Google Scholar] [CrossRef]
- Lawlor, K.T.; Vanslambrouck, J.M.; Higgins, J.W.; Chambon, A.; Bishard, K.; Arndt, D.; Er, P.X.; Wilson, S.B.; Howden, S.E.; Tan, K.S.; et al. Cellular Extrusion Bioprinting Improves Kidney Organoid Reproducibility and Conformation. Nat. Mater. 2021, 20, 260–271. [Google Scholar] [CrossRef]
- Clark, C.C.; Yoo, K.M.; Sivakumar, H.; Strumpf, K.; Laxton, A.W.; Tatter, S.B.; Strowd, R.E.; Skardal, A. Immersion Bioprinting of Hyaluronan and Collagen Bioink-Supported 3D Patient-Derived Brain Tumor Organoids. Biomed. Mater. 2023, 18, 015014. [Google Scholar] [CrossRef]
- Layrolle, P.; Payoux, P.; Chavanas, S. Message in a Scaffold: Natural Biomaterials for Three-Dimensional (3D) Bioprinting of Human Brain Organoids. Biomolecules 2023, 13, 25. [Google Scholar] [CrossRef]
- Song, J.; Bang, S.; Choi, N.; Kim, H.N. Brain Organoid-on-a-Chip: A next-Generation Human Brain Avatar for Recapitulating Human Brain Physiology and Pathology. Biomicrofluidics 2022, 16, 061301. [Google Scholar] [CrossRef] [PubMed]
- Castiglione, H.; Vigneron, P.A.; Baquerre, C.; Yates, F.; Rontard, J.; Honegger, T. Human Brain Organoids-on-Chip: Advances, Challenges, and Perspectives for Preclinical Applications. Pharmaceutics 2022, 14, 2301. [Google Scholar] [CrossRef] [PubMed]
- Karzbrun, E.; Kshirsagar, A.; Cohen, S.R.; Hanna, J.H.; Reiner, O. Human Brain Organoids on a Chip Reveal the Physics of Folding. Nat. Phys. 2018, 14, 515–522. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, L.; Zhu, Y.; Qin, J. Human Brain Organoid-on-a-Chip to Model Prenatal Nicotine Exposure. Lab Chip 2018, 18, 851–860. [Google Scholar] [CrossRef]
- Chen, H.I.; Song, H.; Ming, G. li Applications of Human Brain Organoids to Clinical Problems. Dev. Dyn. 2019, 248, 53–64. [Google Scholar] [CrossRef]
- Eichmüller, O.L.; Knoblich, J.A. Human Cerebral Organoids—A New Tool for Clinical Neurology Research. Nat. Rev. Neurol. 2022, 18, 661–680. [Google Scholar] [CrossRef]
- Zhou, T.; Tan, L.; Cederquist, G.Y.; Fan, Y.; Hartley, B.J.; Mukherjee, S.; Tomishima, M.; Brennand, K.J.; Zhang, Q.; Schwartz, R.E.; et al. High-Content Screening in HPSC-Neural Progenitors Identifies Drug Candidates That Inhibit Zika Virus Infection in Fetal-like Organoids and Adult Brain. Cell Stem Cell 2017, 21, 274–283.e5. [Google Scholar] [CrossRef]
- Shou, Y.; Liang, F.; Xu, S.; Li, X. The Application of Brain Organoids: From Neuronal Development to Neurological Diseases. Front. Cell Dev. Biol. 2020, 8, 579659. [Google Scholar] [CrossRef]
- Tang, X.Y.; Wu, S.; Wang, D.; Chu, C.; Hong, Y.; Tao, M.; Hu, H.; Xu, M.; Guo, X.; Liu, Y. Human Organoids in Basic Research and Clinical Applications. Signal Transduct. Target. Ther. 2022, 7, 168. [Google Scholar] [CrossRef]
- Ravi, K.; Franson, A.; Homan, M.J.; Roberts, H.; Pai, M.P.; Miklja, Z.; He, M.; Wen, B.; Benitez, L.L.; Perissinotti, A.J.; et al. Comparative Pharmacokinetic Analysis of the Blood-Brain Barrier Penetration of Dasatinib and Ponatinib in Mice. Leuk. Lymphoma 2021, 62, 1990–1994. [Google Scholar] [CrossRef]
- Eichmüller, O.L.; Corsini, N.S.; Vértesy, Á.; Morassut, I.; Scholl, T.; Gruber, V.E.; Peer, A.M.; Chu, J.; Novatchkova, M.; Hainfellner, J.A.; et al. Amplification of Human Interneuron Progenitors Promotes Brain Tumors and Neurological Defects. Science 2022, 375, eabf5546. [Google Scholar] [CrossRef]
- Li, R.; Sun, L.; Fang, A.; Li, P.; Wu, Q.; Wang, X. Recapitulating Cortical Development with Organoid Culture In Vitro and Modeling Abnormal Spindle-like (ASPM Related Primary) Microcephaly Disease. Protein Cell 2017, 8, 823–833. [Google Scholar] [CrossRef]
- Hernández, D.; Rooney, L.A.; Daniszewski, M.; Gulluyan, L.; Liang, H.H.; Cook, A.L.; Hewitt, A.W.; Pébay, A. Culture Variabilities of Human IPSC-Derived Cerebral Organoids Are a Major Issue for the Modelling of Phenotypes Observed in Alzheimer’s Disease. Stem Cell Rev. Rep. 2022, 718–731. [Google Scholar] [CrossRef]
- Li, Y.; Muffat, J.; Omer, A.; Bosch, I.; Lancaster, M.A.; Sur, M.; Gehrke, L.; Knoblich, J.A.; Jaenisch, R. Induction of Expansion and Folding in Human Cerebral Organoids. Cell Stem Cell 2017, 20, 385–396.e3. [Google Scholar] [CrossRef]
- Garcez, P.P.; Loiola, E.C.; Madeiro da Costa, R.; Higa, L.M.; Trindade, P.; Delvecchio, R.; Nascimento, J.M.; Brindeiro, R.; Tanuri, A.; Rehen, S.K. Zika Virus Impairs Growth in Human Neurospheres and Brain Organoids. Science 2016, 352, 816–818. [Google Scholar] [CrossRef]
- D’Aiuto, L.; Bloom, D.C.; Naciri, J.N.; Smith, A.; Edwards, T.G.; McClain, L.; Callio, J.A.; Jessup, M.; Wood, J.; Chowdari, K.; et al. Modeling Herpes Simplex Virus 1 Infections in Human Central Nervous System Neuronal Cells Using Two- and Three-Dimensional Cultures Derived from Induced Pluripotent Stem Cells. J. Virol. 2019, 93, e00111-19. [Google Scholar] [CrossRef]
- Krenn, V.; Bosone, C.; Burkard, T.R.; Spanier, J.; Kalinke, U.; Calistri, A.; Salata, C.; Rilo Christoff, R.; Pestana Garcez, P.; Mirazimi, A.; et al. Organoid Modeling of Zika and Herpes Simplex Virus 1 Infections Reveals Virus-Specific Responses Leading to Microcephaly. Cell Stem Cell 2021, 28, 1362–1379.e7. [Google Scholar] [CrossRef]
- Qiao, H.; Guo, M.; Shang, J.; Zhao, W.; Wang, Z.; Liu, N.; Li, B.; Zhou, Y.; Wu, Y.; Chen, P. Herpes Simplex Virus Type 1 Infection Leads to Neurodevelopmental Disorder-Associated Neuropathological Changes. PLoS Pathog. 2020, 16, e1008899. [Google Scholar] [CrossRef]
- Qiao, H.; Zhao, W.; Guo, M.; Zhu, L.; Chen, T.; Wang, J.; Xu, X.; Zhang, Z.; Wu, Y.; Chen, P. Cerebral Organoids for Modeling of HSV-1-Induced-Amyloid β Associated Neuropathology and Phenotypic Rescue. Int. J. Mol. Sci. 2022, 23, 5981. [Google Scholar] [CrossRef]
- Sun, G.; Chiuppesi, F.; Chen, X.; Wang, C.; Tian, E.; Nguyen, J.; Kha, M.; Trinh, D.; Zhang, H.; Marchetto, M.C.; et al. Modeling Human Cytomegalovirus-Induced Microcephaly in Human IPSC-Derived Brain Organoids. Cell Rep. Med. 2020, 1, 100002. [Google Scholar] [CrossRef] [PubMed]
- Brown, R.M.; Rana, P.S.J.B.; Jaeger, H.K.; O’Dowd, J.M.; Balemba, O.B.; Fortunato, E.A. Human Cytomegalovirus Compromises Development of Cerebral Organoids. J. Virol. 2019, 93, e00957-19. [Google Scholar] [CrossRef]
- Russo, M.; Levine, S.Z.; Demjaha, A.; Di Forti, M.; Bonaccorso, S.; Fearon, P.; Dazzan, P.; Pariante, C.M.; David, A.S.; Morgan, C.; et al. Association Between Symptom Dimensions and Categorical Diagnoses of Psychosis: A Cross-Sectional and Longitudinal Investigation. Schizophr. Bull. 2014, 40, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Davis, J.; Eyre, H.; Jacka, F.N.; Dodd, S.; Dean, O.; McEwen, S.; Debnath, M.; McGrath, J.; Maes, M.; Amminger, P.; et al. A Review of Vulnerability and Risks for Schizophrenia: Beyond the Two Hit Hypothesis. Neurosci. Biobehav. Rev. 2016, 65, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, P.F. Puzzling over Schizophrenia: Schizophrenia as a Pathway Disease. Nat. Med. 2012, 18, 210–211. [Google Scholar] [CrossRef]
- Rees, E.; Creeth, H.D.J.; Hwu, H.; Chen, W.J.; Tsuang, M.; Glatt, S.J.; Rey, R.; Kirov, G.; Walters, J.T.R.; Holmans, P.; et al. Schizophrenia, Autism Spectrum Disorders and Developmental Disorders Share Specific Disruptive Coding Mutations. Nat. Commun. 2021, 12, 5353. [Google Scholar] [CrossRef] [PubMed]
- Schmock, H.; Vangkilde, A.; Larsen, K.M.; Fischer, E.; Birknow, M.R.; Richardt, J.; Jepsen, M.; Olesen, C.; Skovby, F.; Plessen, K.J.; et al. The Danish 22q11 Research Initiative. BMC Psychiatry 2015, 15, 1–12. [Google Scholar] [CrossRef]
- Karayiorgou, M.; Morris, M.A.; Morrow, B.; Shprintzeni, R.J.; Goldberg, R.; Borrowii, J.; Gost, A.; Nestadt, G.; Wolyniec, P.S.; Lasseter, V.K.; et al. Schizophrenia Susceptibility Associated with Interstitial Deletions of Chromosome 22qll. Proc. Natl. Acad. Sci. USA 1995, 92, 7612–7616. [Google Scholar] [CrossRef]
- Karayiorgou, M.; Simon, T.J.; Gogos, J.A. 22q11.2 microdeletions: Linking DNA structural variation to brain dysfunction and schizophrenia. Nat. Rev. Neurosci. 2010, 11, 402–416. [Google Scholar] [CrossRef]
- Mcdonald, T.V.; Choi, C.H.; Schoenfeld, B.P.; Bell, A.J.; Hinchey, J.; Hinchey, P.; Kollaros, M.; Ferrick, N.J.; Chambers, D.B.; Langer, S. Multiple Drug Treatments That Increase CAMP Signaling Restore Long-Term Memory and Aberrant Signaling in Fragile X Syndrome Models. Front. Behav. Neurosci. 2016, 10, 136. [Google Scholar] [CrossRef]
- De Oliveira Figueiredo, E.C.; Bondiolotti, B.M.; Laugeray, A.; Bezzi, P. Synaptic Plasticity Dysfunctions in the Pathophysiology of 22q11 Deletion Syndrome: Is There a Role for Astrocytes? Int. J. Mol. Sci. 2022, 23, 4412. [Google Scholar] [CrossRef]
- Zinkstok, J.R.; Boot, E.; Bassett, A.S.; Hiroi, N.; Butcher, N.J.; Vingerhoets, C.; Vorstman, J.A.S.; van Amelsvoort, T.A.M.J. Neurobiological Perspective of 22q11.2 Deletion Syndrome. Lancet. Psychiatry 2019, 6, 951–960. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira Figueiredo, E.C.; Calì, C.; Petrelli, F.; Bezzi, P. Emerging Evidence for Astrocyte Dysfunction in Schizophrenia. Glia 2022, 70, 1585–1604. [Google Scholar] [CrossRef]
- Chun, S.; Westmoreland, J.J.; Bayazitov, I.T.; Eddins, D.; Pani, A.K.; Smeyne, R.J.; Yu, J.; Blundon, J.A.; Zakharenko, S.S. Specific Disruption of Thalamic Inputs to the Auditory Cortex in Schizophrenia Models. Science 2014, 344, 1178–1182. [Google Scholar] [CrossRef]
- Sigurdsson, T.; Stark, K.L.; Karayiorgou, M.; Gogos, J.A.; Gordon, J.A. Impaired Hippocampal–Prefrontal Synchrony in a Genetic Mouse Model of Schizophrenia. Nature 2010, 464, 763–767. [Google Scholar] [CrossRef]
- Mukherjee, A.; Carvalho, F.; Eliez, S.; Caroni, P.; Mukherjee, A.; Carvalho, F.; Eliez, S.; Caroni, P. Long-Lasting Rescue of Network and Cognitive Dysfunction in a Genetic Schizophrenia Model Article Long-Lasting Rescue of Network and Cognitive Dysfunction. Cell 2019, 178, 1387–1402.e14. [Google Scholar] [CrossRef]
- Fernandez, A.; Meechan, D.W.; Karpinski, B.A.; Elizabeth, M.; Bryan, C.A.; Rutz, H.L.; Radin, E.A.; Lubin, N.; Erin, R.; Popratiloff, A.; et al. Mitochondrial Dysfunction Leads to Cortical Under-Connectivity and Cognitive Impairment. Neuron 2020, 102, 1127–1142.E3. [Google Scholar] [CrossRef]
- Devaraju, P.; Zakharenko, S.S. Mitochondria in Complex Psychiatric Disorders: Lessons from Mouse Models of 22q11.2 Deletion Syndrome. Bioessays 2016, 39, 1600177. [Google Scholar] [CrossRef]
- Steullet, P.; Cabungcal, J.; Coyle, J.; Didriksen, M.; Gill, K.; Grace, A.A.; Hensch, T.K.; Lamantia, A.; Lindemann, L.; Maynard, T.M. Oxidative Stress-Driven Parvalbumin Interneuron Impairment as a Common Mechanism in Models of Schizophrenia. Mol. Psychiatry 2017, 22, 936–943. [Google Scholar] [CrossRef]
- Zhao, D.; Lin, M.; Chen, J.; Pedrosa, E.; Hrabovsky, A.; Fourcade, H.M.; Zheng, D.; Lachman, H.M. MicroRNA Profiling of Neurons Generated Using Induced Pluripotent Stem Cells Derived from Patients with Schizophrenia and Schizoaffective Disorder, and 22q11.2 Del. PLoS ONE 2015, 10, e0132387. [Google Scholar] [CrossRef] [PubMed]
- Toyoshima, M.; Akamatsu, W.; Okada, Y.; Ohnishi, T.; Balan, S.; Hisano, Y.; Iwayama, Y.; Toyota, T.; Matsumoto, T.; Itasaka, N. Analysis of Induced Pluripotent Stem Cells Carrying 22q11. 2 Deletion. Transl. Psychiatry 2016, 6, e934. [Google Scholar] [CrossRef]
- Lin, M.; Pedrosa, E.; Hrabovsky, A.; Chen, J.; Puliafito, B.R.; Gilbert, S.R.; Zheng, D.; Lachman, H.M. Integrative Transcriptome Network Analysis of IPSC-Derived Neurons from Schizophrenia and Schizoaffective Disorder Patients with 22q11.2 Deletion. BMC Syst. Biol. 2016, 10, 105. [Google Scholar] [CrossRef] [PubMed]
- Moskvina, V.; Craddock, N.; Holmans, P.; Nikolov, I.; Pahwa, J.S.; Green, E.; Owen, M.J.; O’Donovan, M.C.; Consortium, W.T.C.C. Gene-Wide Analyses of Genome-Wide Association Data Sets: Evidence for Multiple Common Risk Alleles for Schizophrenia and Bipolar Disorder and for Overlap in Genetic Risk. Mol. Psychiatry 2009, 14, 252–260. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.S.; Wu, H.; Krzisch, M.; Wu, X.; Graef, J.; Muffat, J.; Hnisz, D.; Li, C.H.; Yuan, B.; Xu, C.; et al. Rescue of Fragile X Syndrome Neurons by DNA Methylation Editing of the FMR1 Gene. Cell 2018, 172, 979–992.e6. [Google Scholar] [CrossRef]
- Bagni, C.; Zukin, R.S. A Synaptic Perspective of Fragile X Syndrome and Autism Spectrum Disorders. Neuron 2022, 101, 1070–1088. [Google Scholar] [CrossRef]
- Boland, M.J.; Nazor, K.L.; Tran, H.T.; Szücs, A.; Lynch, C.L.; Paredes, R.; Tassone, F.; Sanna, P.P.; Hagerman, R.J.; Loring, J.F. Molecular Analyses of Neurogenic Defects in a Human Pluripotent Stem Cell Model of Fragile X Syndrome. Brain 2017, 140, 582–598. [Google Scholar] [CrossRef] [PubMed]
- Halevy, T.; Czech, C.; Benvenisty, N. Molecular Mechanisms Regulating the Defects in Fragile X Syndrome Neurons Derived from Human Pluripotent Stem Cells. Stem Cell Rep. 2015, 4, 37–46. [Google Scholar] [CrossRef]
- Sunamura, N.; Iwashita, S.; Enomoto, K.; Kadoshima, T.; Isono, F. Loss of the Fragile X Mental Retardation Protein Causes Aberrant Differentiation in Human Neural Progenitor Cells. Sci. Rep. 2018, 8, 11585. [Google Scholar] [CrossRef]
- Utami, K.H.; Skotte, N.H.; Colaço, A.R.; Amirah, N.; Mohammad, B.; Sim, B.; Yeo, X.Y.; Bae, H.; Garcia-miralles, M.; Radulescu, C.I.; et al. Integrative Analysis Identi Fi Es Key Molecular Signatures Underlying Neurodevelopmental De Fi Cits in Fragile X Syndrome. Biol. Psychiatry 2020, 88, 500–511. [Google Scholar] [CrossRef]
- Raj, N.; McEachin, Z.T.; Harousseau, W.; Zhou, Y.; Zhang, F.; Merritt-Garza, M.E.; Taliaferro, J.M.; Kalinowska, M.; Marro, S.G.; Hales, C.M.; et al. Cell-Type-Specific Profiling of Human Cellular Models of Fragile X Syndrome Reveal PI3K-Dependent Defects in Translation and Neurogenesis. Cell Rep. 2021, 35, 108991. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.; Zhou, Y.; Li, Y.; Han, Y.; Xu, J.; Niu, W.; Li, Z.; Liu, S.; Feng, H.; Huang, W.; et al. A Human Forebrain Organoid Model of Fragile X Syndrome Exhibits Altered Neurogenesis and Highlights New Treatment Strategies. Nat. Neurosci. 2021, 24, 1377–1391. [Google Scholar] [CrossRef] [PubMed]
- Bassell, G.J.; Gross, C. Reducing Glutamate Signaling Pays off in Fragile X. Nat. Med. 2008, 14, 249–250. [Google Scholar] [CrossRef] [PubMed]
- Lozano, R.; Rosero, C.A.; Hagerman, R.J. Fragile X Spectrum Disorders. Intractable Rare Dis. Res. 2014, 3, 134–146. [Google Scholar] [CrossRef]
- Choi, H.; Song, J.; Park, G.; Kim, J. Modeling of Autism Using Organoid Technology. Mol. Neurobiol. 2017, 54, 7789–7795. [Google Scholar] [CrossRef]
- Mariani, J.; Coppola, G.; Zhang, P.; Abyzov, A.; Provini, L.; Tomasini, L.; Amenduni, M.; Szekely, A.; Palejev, D.; Wilson, M.; et al. FOXG1-Dependent Dysregulation of GABA/Glutamate Neuron Differentiation in Autism Spectrum Disorders. Cell 2015, 162, 375–390. [Google Scholar] [CrossRef]
- Wang, P.; Mokhtari, R.; Pedrosa, E.; Kirschenbaum, M.; Bayrak, C.; Zheng, D.; Lachman, H.M. CRISPR/Cas9-Mediated Heterozygous Knockout of the Autism Gene CHD8 and Characterization of Its Transcriptional Networks in Cerebral Organoids Derived from IPS Cells. Mol. Autism 2017, 8, 1–17. [Google Scholar] [CrossRef]
- Schafer, S.T.; Paquola, A.C.M.; Stern, S.; Gosselin, D.; Ku, M.; Pena, M.; Kuret, T.J.M.; Liyanage, M.; Mansour, A.A.F.; Jaeger, B.N.; et al. Pathological Priming Causes Developmental Gene Network Heterochronicity in Autistic Subject-Derived Neurons. Nat. Neurosci. 2019, 22, 243–255. [Google Scholar] [CrossRef]
- Wang, L.; Wang, Y.; Zhou, S.; Yang, L.; Shi, Q.; Li, Y.; Zhang, K.; Yang, L.; Zhao, M.; Yang, Q. Imbalance between Glutamate and GABA in Fmr1 Knockout Astrocytes Influences Neuronal Development. Genes 2016, 7, 45. [Google Scholar] [CrossRef]
- Hagerman, R.J. Lessons from Fragile X Regarding Neurobiology, Autism, and Neurodegeneration. J. Dev. Behav. Pediatr. 2006, 27, 63–74. [Google Scholar] [CrossRef]
- Buscemi, L.; Ginet, V.; Lopatar, J.; Montana, V.; Pucci, L.; Spagnuolo, P.; Zehnder, T.; Grubi, V.; Truttman, A.; Sala, C.; et al. Homer1 Scaffold Proteins Govern Ca2+ Dynamics in Normal and Reactive Astrocytes. Cereb. Cortex 2017, 23, 2365–2384. [Google Scholar] [CrossRef]
- Han, Q.; Kim, Y.H.; Wang, X.; Liu, D.; Zhang, Z.-J.; Bey, A.L.; Lay, M.; Chang, W.; Berta, T.; Zhang, Y.; et al. SHANK3 Deficiency Impairs Heat Hyperalgesia and TRPV1 Signaling in Primary Sensory Neurons. Neuron 2016, 92, 1279–1293. [Google Scholar] [CrossRef] [PubMed]
- Petrelli, F.; Bezzi, P. ScienceDirect MGlu5-Mediated Signalling in Developing Astrocyte and the Pathogenesis of Autism Spectrum Disorders. Curr. Opin. Neurobiol. 2018, 48, 139–145. [Google Scholar] [CrossRef] [PubMed]
- Morel, L.; Higashimori, H.; Tolman, M.; Yang, Y. VGluT1+ Neuronal Glutamatergic Signaling Regulates Postnatal Developmental Maturation of Cortical Protoplasmic Astroglia. J. Neurosci. 2014, 34, 10950–10962. [Google Scholar] [CrossRef] [PubMed]
- Bajaj, S.; Bagley, J.A.; Sommer, C.; Vertesy, A.; Knoblich, J.A.; Wong, S.N.; Krenn, V.; Julie, L. Neurotransmitter Signaling Regulates Distinct Phases of Multimodal Human Interneuron Migration. EMBO J. 2021, 40, e108714. [Google Scholar] [CrossRef]
- Splawski, I.; Timothy, K.W.; Sharpe, L.M.; Decher, N.; Kumar, P.; Bloise, R.; Napolitano, C.; Schwartz, P.J.; Joseph, R.M.; Condouris, K.; et al. CaV1.2 Calcium Channel Dysfunction Causes a Multisystem Disorder Including Arrhythmia and Autism. Cell 2004, 119, 19–31. [Google Scholar] [CrossRef]
- Birey, F.; Andersen, J.; Makinson, C.D.; Islam, S.; Wei, W.; Huber, N.; Fan, H.C.; Metzler, K.R.C.; Panagiotakos, G.; Thom, N.; et al. Assembly of Functionally Integrated Human Forebrain Spheroids. Nature 2017, 545, 54–59. [Google Scholar] [CrossRef]
- Revah, O.; Gore, F.; Kelley, K.W.; Andersen, J.; Sakai, N.; Chen, X.; Li, M.Y.; Birey, F.; Yang, X.; Saw, N.L.; et al. Maturation and Circuit Integration of Transplanted Human Cortical Organoids. Nature 2022, 610, 319–326. [Google Scholar] [CrossRef]
- Brady, M.V.; Mariani, J.; Koca, Y.; Szekely, A.; King, R.A.; Bloch, M.H.; Landeros-Weisenberger, A.; Leckman, J.F.; Vaccarino, F.M. Mispatterning and Interneuron Deficit in Tourette Syndrome Basal Ganglia Organoids. Mol. Psychiatry 2022, 27, 5007–5019. [Google Scholar] [CrossRef]
- Welch, J.M.; Lu, J.; Rodriguiz, R.M.; Trotta, N.C.; Peca, J.; Ding, J.-D.; Feliciano, C.; Chen, M.; Adams, J.P.; Luo, J.; et al. Cortico-Striatal Synaptic Defects and OCD-like Behaviours in Sapap3-Mutant Mice. Nature 2007, 448, 894–900. [Google Scholar] [CrossRef]
- Ahmari, S.E.; Spellman, T.; Douglass, N.L.; Kheirbek, M.A.; Simpson, H.B.; Deisseroth, K.; Gordon, J.A.; Hen, R. Repeated Cortico-Striatal Stimulation Generates Persistent OCD-Like Behavior. Science 2013, 340, 1234–1239. [Google Scholar] [CrossRef]
- Wang, Z.; Maia, T.V.; Marsh, R.; Colibazzi, T.; Gerber, A.; Peterson, B.S. The Neural Circuits That Generate Tics in Tourette’s Syndrome. Am. J. Psychiatry 2011, 168, 1326–1337. [Google Scholar] [CrossRef]
- Nikolaus, S.; Antke, C.; Beu, M.; Müller, H.W. Cortical GABA, Striatal Dopamine and Midbrain Serotonin as the Key Players in Compulsive and Anxiety Disorders—Results from In Vivo Imaging Studies. Rev. Neurosci. 2010, 21, 119–140. [Google Scholar] [CrossRef]
- Olver, J.S.; O’Keefe, G.; Jones, G.R.; Burrows, G.D.; Tochon-Danguy, H.J.; Ackermann, U.; Scott, A.M.; Norman, T.R. Dopamine D1 Receptor Binding in the Anterior Cingulate Cortex of Patients with Obsessive–Compulsive Disorder. Psychiatry Res. Neuroimaging 2010, 183, 85–88. [Google Scholar] [CrossRef]
- Kalueff, A.V.; Stewart, A.M.; Song, C.; Berridge, K.C.; Graybiel, A.M.; Fentress, J.C. Neurobiology of Rodent Self-Grooming and Its Value for Translational Neuroscience. Nat. Rev. Neurosci. 2016, 17, 45–59. [Google Scholar] [CrossRef]
- Petrelli, F.; Zehnder, T.; Laugeray, A.; Mondoloni, S.; Calì, C.; Pucci, L.; Molinero Perez, A.; Bondiolotti, B.M.; De Oliveira Figueiredo, E.; Dallerac, G.; et al. Disruption of Astrocyte-Dependent Dopamine Control in the Developing Medial Prefrontal Cortex Leads to Excessive Grooming in Mice. Biol. Psychiatry 2023, 93, 966–975. [Google Scholar] [CrossRef]
- Goodman, W.K.; Price, L.H.; Delgado, P.L.; Palumbo, J.; Krystal, J.H.; Nagy, L.M.; Rasmussen, S.A.; Heninger, G.R.; Charney, D.S. Specificity of Serotonin Reuptake Inhibitors in the Treatment of Obsessive-Compulsive Disorder: Comparison of Fluvoxamine and Desipramine. Arch. Gen. Psychiatry 1990, 47, 577–585. [Google Scholar] [CrossRef]
- Ducasse, D.; Boyer, L.; Michel, P.; Loundou, A.; Macgregor, A.; Micoulaud-Franchi, J.-A.; Courtet, P.; Abbar, M.; Leboyer, M.; Fond, G. D2 and D3 Dopamine Receptor Affinity Predicts Effectiveness of Antipsychotic Drugs in Obsessive-Compulsive Disorders: A Metaregression Analysis. Psychopharmacology 2014, 231, 3765–3770. [Google Scholar] [CrossRef]
- Kofuji, P.; Araque, A. Astrocytes and Behavior. Annu. Rev. Neurosci. 2021, 44, 49–67. [Google Scholar] [CrossRef]
- Peterson, B.; Riddle, M.A.; Cohen, D.J.; Katz, L.D.; Smith, J.C.; Hardin, M.T.; Leckman, J.F. Reduced Basal Ganglia Volumes in Tourette’s Syndrome Using Three-dimensional Reconstruction Techniques from Magnetic Resonance Images. Neurology 1993, 43, 941. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.H.; Kim, Y.H.; Hebisch, M.; Sliwinski, C.; Lee, S.; Avanzo, C.D.; Chen, H.; Hooli, B.; Asselin, C.; Muffat, J.; et al. Model of Alzheimer’s Disease. Nature 2014, 515, 274–278. [Google Scholar] [CrossRef] [PubMed]
- Jorfi, M.; Avanzo, C.D.; Tanzi, R.E.; Kim, D.Y.; Irimia, D. Human Neurospheroid Arrays for In Vitro Studies of Alzheimer’s Disease. Sci. Rep. 2018, 8, 2450. [Google Scholar] [CrossRef] [PubMed]
- Garcia-reitboeck, P.; Anichtchik, O.; Dalley, J.W.; Ninkina, N.; Tofaris, G.K.; Buchman, V.L.; Grazia, M. Endogenous Alpha-Synuclein in Fl Uences the Number of Dopaminergic Neurons in Mouse Substantia Nigra. Exp. Neurol. 2013, 248, 541–545. [Google Scholar] [CrossRef] [PubMed]
- Le Grand, J.N.; Gonzalez-Cano, L.; Pavlou, M.A.; Schwamborn, J.C. Neural Stem Cells in Parkinson’s Disease: A Role for Neurogenesis Defects in Onset and Progression. Cell. Mol. Life Sci. 2015, 72, 773–797. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.H.; Choi, S.H.; D’Avanzo, C.; Hebisch, M.; Sliwinski, C.; Bylykbashi, E.; Washicosky, K.J.; Klee, J.B.; Brüstle, O.; Tanzi, R.E.; et al. A 3D Human Neural Cell Culture System for Modeling Alzheimer’s Disease. Nat. Protoc. 2015, 10, 985–1006. [Google Scholar] [CrossRef] [PubMed]
- Cairns, D.M.; Rouleau, N.; Parker, R.N.; Walsh, K.G.; Gehrke, L.; Kaplan, D.L. A 3D Human Brain—Like Tissue Model of Herpes-Induced Alzheimer’s Disease. Sci. Adv. 2020, 6, eaay8828. [Google Scholar] [CrossRef]
- Gonzalez, C.; Armijo, E.; Bravo-alegria, J.; Becerra-Calixto, A.; Mays, C.E.; Soto, C. Modeling Amyloid Beta and Tau Pathology in Human Cerebral Organoids. Mol. Psychiatry 2018, 23, 2363–2374. [Google Scholar] [CrossRef]
- Alić, I.; Goh, P.A.; Murray, A.; Portelius, E.; Gkanatsiou, E.; Gough, G.; Mok, K.Y.; Koschut, D.; Brunmeir, R.; Yeap, Y.J.; et al. Patient-Specific Alzheimer-like Pathology in Trisomy 21 Cerebral Organoids Reveals BACE2 as a Gene Dose-Sensitive AD Suppressor in Human Brain. Mol. Psychiatry 2021, 26, 5766–5788. [Google Scholar] [CrossRef]
- Park, J.; Wetzel, I.; Marriott, I.; Dréau, D.; D’Avanzo, C.; Kim, D.Y.; Tanzi, R.E.; Cho, H. A 3D Human Triculture System Modeling Neurodegeneration and Neuroinflammation in Alzheimer’s Disease. Nat. Neurosci. 2018, 21, 941–951. [Google Scholar] [CrossRef]
- Yin, J.; Vandongen, A.M. Enhanced Neuronal Activity and Asynchronous Calcium Transients Revealed in a 3D Organoid Model of Alzheimer’s Disease. ACS Biomater. Sci. Eng. 2021, 7, 254–264. [Google Scholar] [CrossRef]
- Lin, Y.T.; Seo, J.; Gao, F.; Feldman, H.M.; Wen, H.L.; Penney, J.; Cam, H.P.; Gjoneska, E.; Raja, W.K.; Cheng, J.; et al. APOE4 Causes Widespread Molecular and Cellular Alterations Associated with Alzheimer’s Disease Phenotypes in Human IPSC-Derived Brain Cell Types. Neuron 2018, 98, 1141–1154.e7. [Google Scholar] [CrossRef]
- Kim, S.; Kwon, S.-H.; Kam, T.-I.; Panicker, N.; Karuppagounder, S.S.; Lee, S.; Lee, J.H.; Kim, W.R.; Kook, M.; Foss, C.A.; et al. Transneuronal Propagation of Pathologic α-Synuclein from the Gut to the Brain Models Parkinson’s Disease. Neuron 2019, 103, 627–641.e7. [Google Scholar] [CrossRef]
- Smits, L.M.; Reinhardt, L.; Reinhardt, P.; Glatza, M.; Monzel, A.S.; Stanslowsky, N.; Rosato-Siri, M.D.; Zanon, A.; Antony, P.M.; Bellmann, J.; et al. Modeling Parkinson’s Disease in Midbrain-like Organoids. npj Park. Dis. 2019, 5, 5. [Google Scholar] [CrossRef]
- Kano, M.; Takanashi, M.; Oyama, G.; Yoritaka, A.; Hatano, T.; Shiba-Fukushima, K.; Nagai, M.; Nishiyama, K.; Hasegawa, K.; Inoshita, T.; et al. Reduced Astrocytic Reactivity in Human Brains and Midbrain Organoids with PRKN Mutations. npj Park. Dis. 2020, 6, 33. [Google Scholar] [CrossRef]
- Wulansari, N.; Darsono, W.H.W.; Woo, H.J.; Chang, M.Y.; Kim, J.; Bae, E.J.; Sun, W.; Lee, J.H.; Cho, I.J.; Shin, H.; et al. Neurodevelopmental Defects and Neurodegenerative Phenotypes in Human Brain Organoids Carrying Parkinson’s Disease-Linked DNAJC6 Mutations. Sci. Adv. 2021, 7, eabb1540. [Google Scholar] [CrossRef]
- Escartin, C.; Galea, E.; Lakatos, A.; O’Callaghan, J.P.; Petzold, G.C.; Serrano-Pozo, A.; Steinhäuser, C.; Volterra, A.; Carmignoto, G.; Agarwal, A.; et al. Reactive Astrocyte Nomenclature, Definitions, and Future Directions. Nat. Neurosci. 2021, 24, 312–325. [Google Scholar] [CrossRef]
- Kelly, Á.M. Exercise-Induced Modulation of Neuroinflammation in Models of Alzheimer’s Disease. Brain Plast. 2018, 4, 81–94. [Google Scholar] [CrossRef]
- Cvetkovic, C.; Patel, R.; Shetty, A.; Hogan, M.K.; Anderson, M.; Basu, N.; Aghlara-Fotovat, S.; Ramesh, S.; Sardar, D.; Veiseh, O.; et al. Assessing Gq-GPCR–Induced Human Astrocyte Reactivity Using Bioengineered Neural Organoids. J. Cell Biol. 2022, 221, e202107135. [Google Scholar] [CrossRef]
- Bezzi, P. Tweaking Neural Organoids to Model Human Reactive Astrocytes. J. Cell Biol. 2022, 221, e202202026. [Google Scholar] [CrossRef]
- Mansour, A.A.; Gonçalves, J.T.; Bloyd, C.W.; Li, H.; Fernandes, S.; Quang, D.; Johnston, S.; Parylak, S.L.; Jin, X.; Gage, F.H. An In Vivo Model of Functional and Vascularized Human Brain Organoids. Nat. Biotechnol. 2018, 36, 432–441. [Google Scholar] [CrossRef]
- Lavazza, A.; Massimini, M. Cerebral Organoids: Ethical Issues and Consciousness Assessment. J. Med. Ethics 2018, 44, 606–610. [Google Scholar] [CrossRef] [PubMed]
| Pros of Neurospheres | Pros of Brain Organoids |
|---|---|
|
|
|
|
|
|
|
|
|
|
| Cells | Hydrogel Precursor | Bioprinting Technology | Features | Ref. |
|---|---|---|---|---|
| Human neural stem cells |
| Extrusion |
| [75] |
| iPSC derived NPC |
| Microfluidic-assisted extrusion |
| [77] |
| Primary rat cortical neurons |
| Extrusion |
| [88] |
| Rat embryonic neurons and astrocytes |
| Microvalve |
| [89] |
| Neural stem cells |
| Stereolithography |
| [90] |
| sNPC and OPC |
| Extrusion |
| [85] |
| iPSC-derived cortical neurons and glial cells. |
| Extrusion |
| [92] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
D’Antoni, C.; Mautone, L.; Sanchini, C.; Tondo, L.; Grassmann, G.; Cidonio, G.; Bezzi, P.; Cordella, F.; Di Angelantonio, S. Unlocking Neural Function with 3D In Vitro Models: A Technical Review of Self-Assembled, Guided, and Bioprinted Brain Organoids and Their Applications in the Study of Neurodevelopmental and Neurodegenerative Disorders. Int. J. Mol. Sci. 2023, 24, 10762. https://doi.org/10.3390/ijms241310762
D’Antoni C, Mautone L, Sanchini C, Tondo L, Grassmann G, Cidonio G, Bezzi P, Cordella F, Di Angelantonio S. Unlocking Neural Function with 3D In Vitro Models: A Technical Review of Self-Assembled, Guided, and Bioprinted Brain Organoids and Their Applications in the Study of Neurodevelopmental and Neurodegenerative Disorders. International Journal of Molecular Sciences. 2023; 24(13):10762. https://doi.org/10.3390/ijms241310762
Chicago/Turabian StyleD’Antoni, Chiara, Lorenza Mautone, Caterina Sanchini, Lucrezia Tondo, Greta Grassmann, Gianluca Cidonio, Paola Bezzi, Federica Cordella, and Silvia Di Angelantonio. 2023. "Unlocking Neural Function with 3D In Vitro Models: A Technical Review of Self-Assembled, Guided, and Bioprinted Brain Organoids and Their Applications in the Study of Neurodevelopmental and Neurodegenerative Disorders" International Journal of Molecular Sciences 24, no. 13: 10762. https://doi.org/10.3390/ijms241310762
APA StyleD’Antoni, C., Mautone, L., Sanchini, C., Tondo, L., Grassmann, G., Cidonio, G., Bezzi, P., Cordella, F., & Di Angelantonio, S. (2023). Unlocking Neural Function with 3D In Vitro Models: A Technical Review of Self-Assembled, Guided, and Bioprinted Brain Organoids and Their Applications in the Study of Neurodevelopmental and Neurodegenerative Disorders. International Journal of Molecular Sciences, 24(13), 10762. https://doi.org/10.3390/ijms241310762









