Δ8-THC Induces Up-Regulation of Glutamatergic Pathway Genes in Differentiated SH-SY5Y: A Transcriptomic Study
Abstract
1. Introduction
2. Results
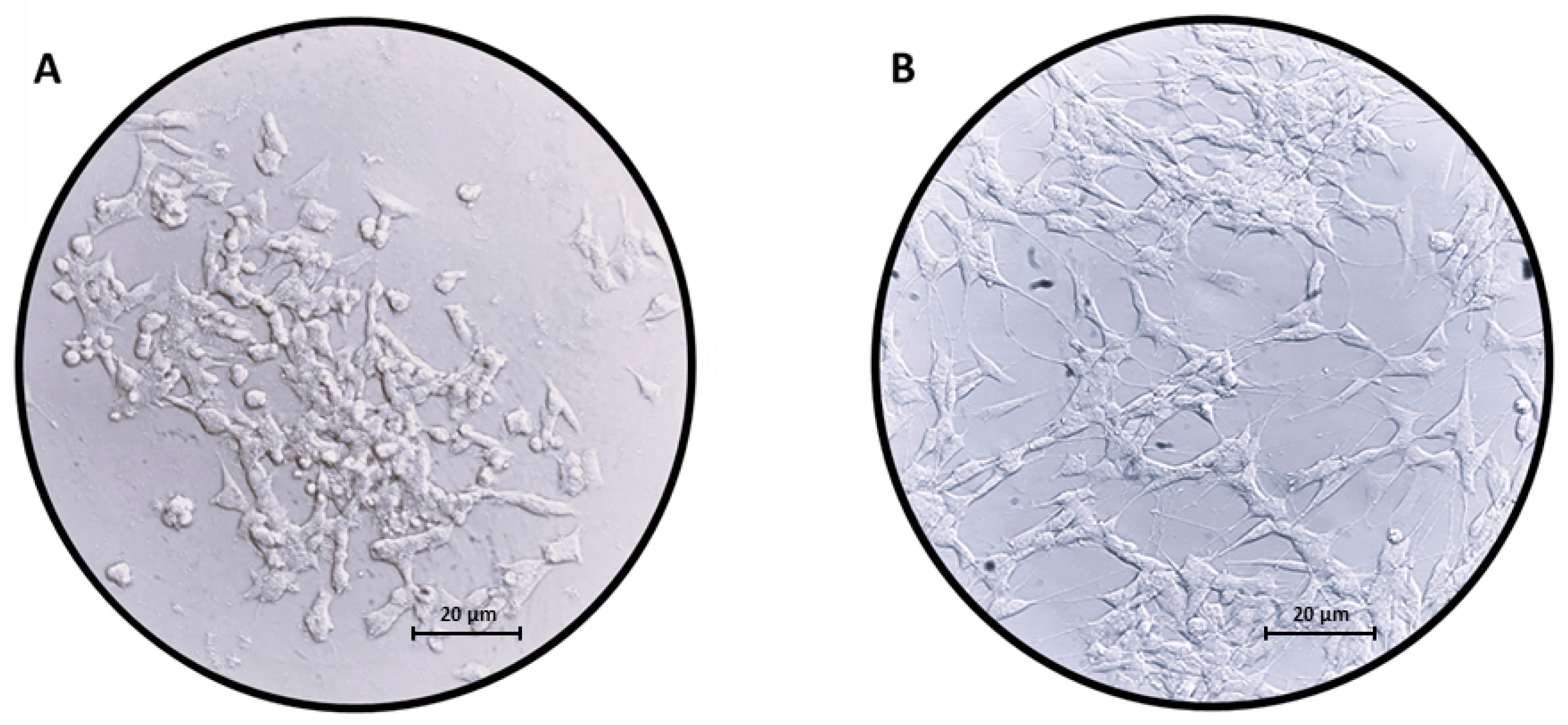
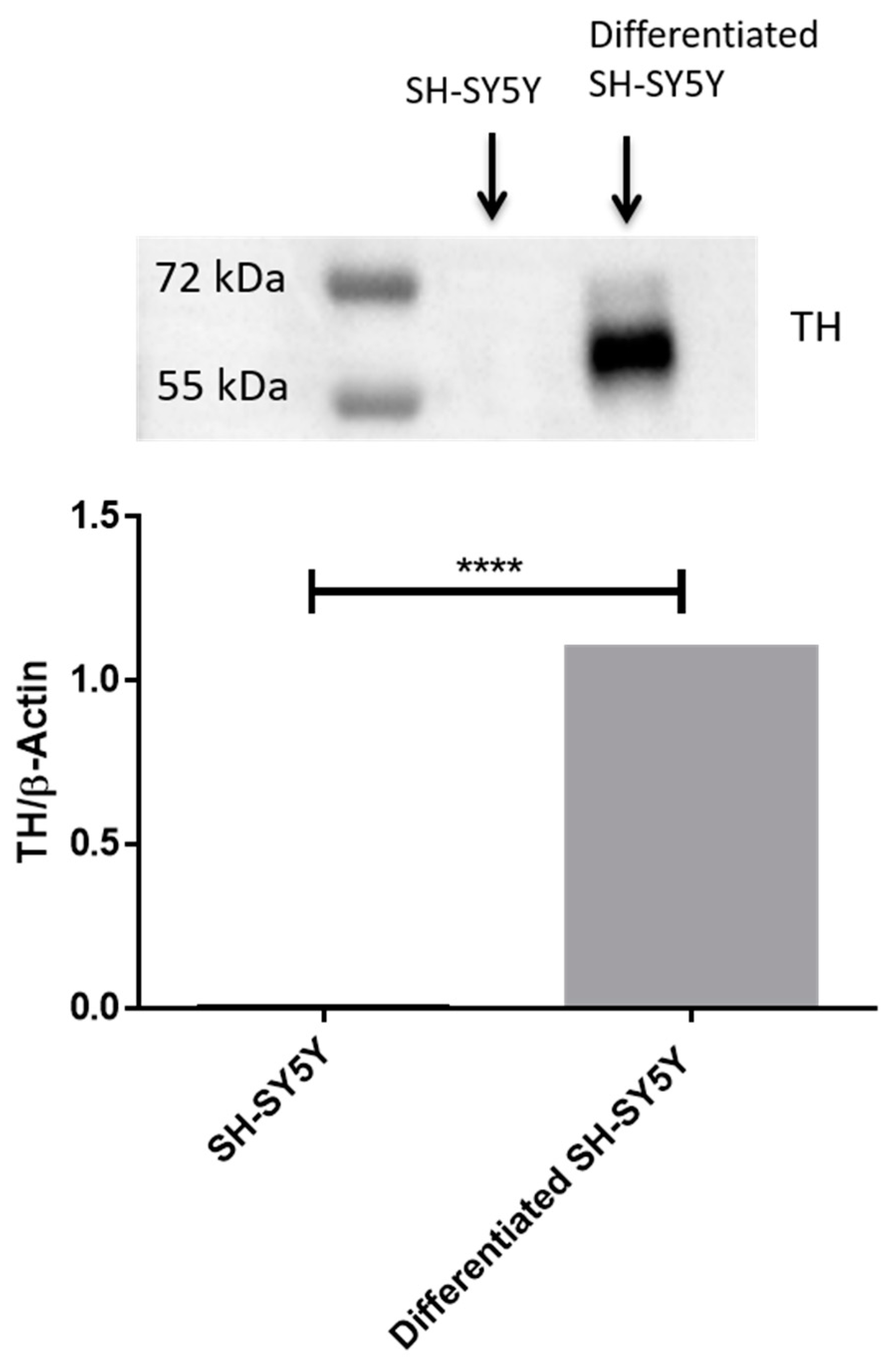
2.1. Differentiation of SH-SY5Y Cells with RA
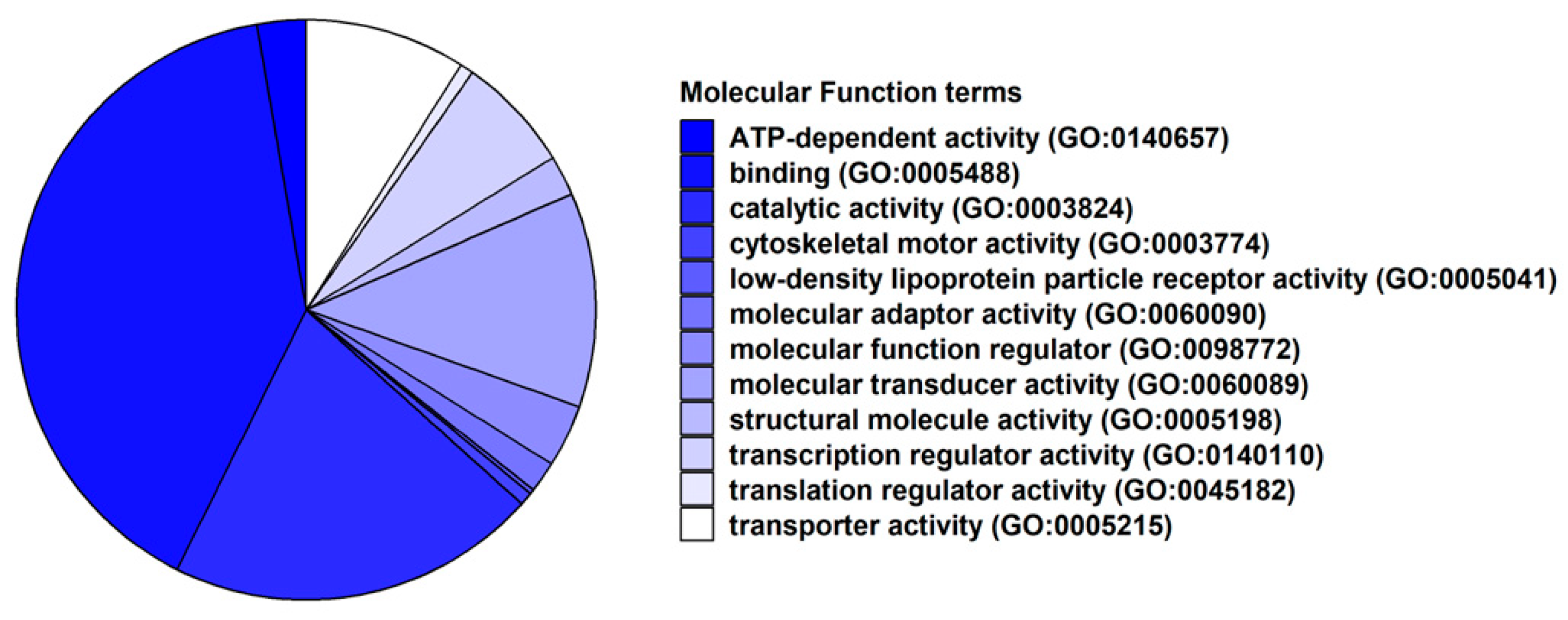
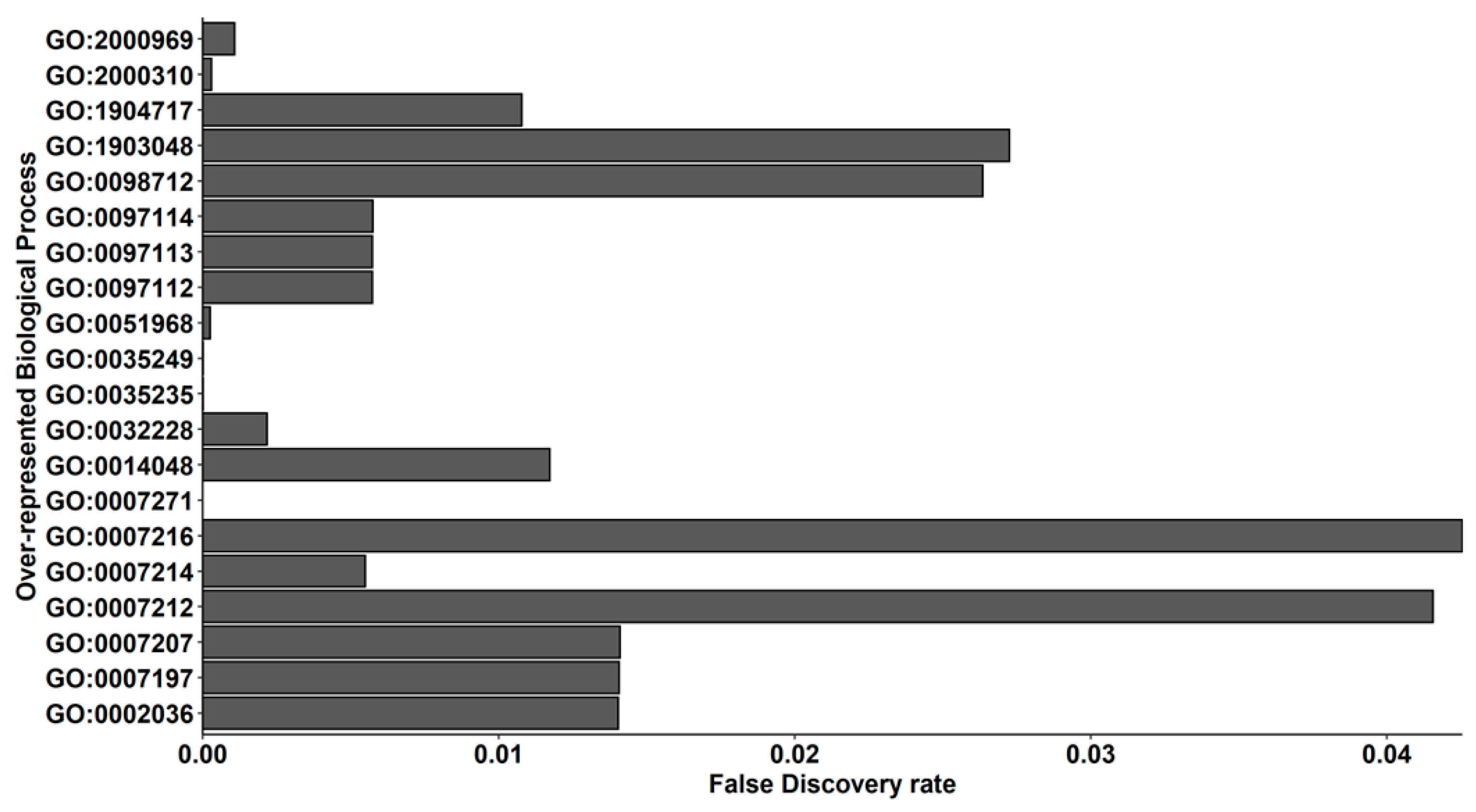
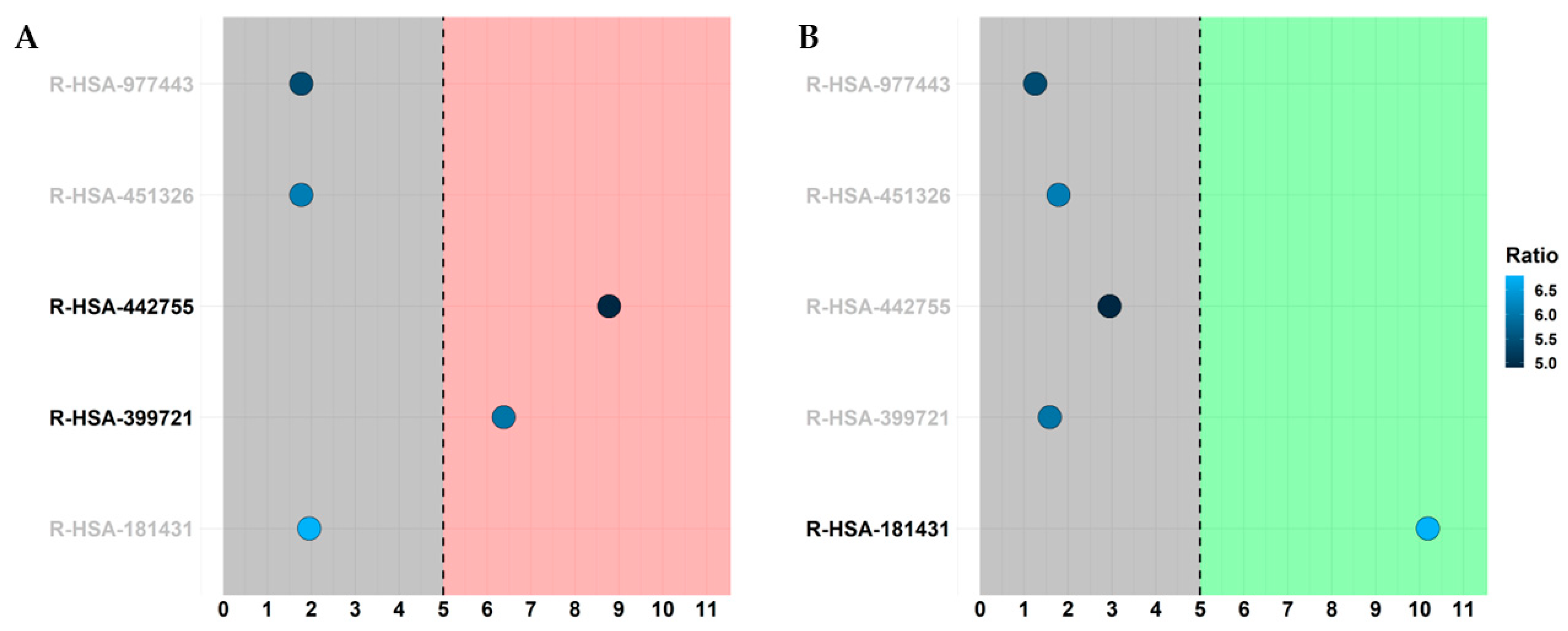
2.2. Transcriptomic Analysis
3. Discussion
4. Materials and Methods
4.1. Synthesis and Purification of Δ8-THC
4.2. Cell Culture, Differentiation and Treatment with Δ8-THC
4.3. Western Blot Analysis for Tyrosine Hydroxylase
4.4. RNA Extraction and Library Preparation
4.5. Sequencing Analysis
4.6. Comparative Analysis and In Silico Inspection
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pellati, F.; Borgonetti, V.; Brighenti, V.; Biagi, M.; Benvenuti, S.; Corsi, L. Cannabis sativa L. and Nonpsychoactive Cannabinoids: Their Chemistry and Role against Oxidative Stress, Inflammation, and Cancer. BioMed Res. Int. 2018, 2018, 1691428. [Google Scholar] [CrossRef]
- Pertwee, R.G. Pharmacological actions of Cannabinoids. Handb. Exp. Pharmacol. 2005, 168, 1–51. [Google Scholar] [CrossRef]
- Appendino, G.; Chianese, G.; Taglialatela-Scafati, O. Cannabinoids: Occurrence and medicinal chemistry. Curr. Med. Chem. 2011, 18, 1085–1099. [Google Scholar] [CrossRef] [PubMed]
- Hanus, L.O.; Meyer, S.M.; Munoz, E.; Taglialatela-Scafati, O.; Appendino, G. Phytocannabinoids: A unified critical inventory. Nat. Prod. Rep. 2016, 33, 1357–1392. [Google Scholar] [CrossRef]
- Tagen, M.; Klumpers, L.E. Review of delta-8-tetrahydrocannabinol (Δ8-THC): Comparative pharmacology with Δ9-THC. Br. J. Pharmacol. 2022, 179, 3915–3933. [Google Scholar] [CrossRef]
- LoParco, C.R.; Rossheim, M.E.; Walters, S.T.; Zhou, Z.; Olsson, S.; Sussman, S.Y. Delta-8 tetrahydrocannabinol: A scoping review and commentary. Addiction 2023, 118, 1011–1028. [Google Scholar] [CrossRef] [PubMed]
- Gugliandolo, A.; Blando, S.; Salamone, S.; Caprioglio, D.; Pollastro, F.; Mazzon, E.; Chiricosta, L. Delta(8)-THC Protects against Amyloid Beta Toxicity Modulating ER Stress In Vitro: A Transcriptomic Analysis. Int. J. Mol. Sci. 2023, 24, 6598. [Google Scholar] [CrossRef] [PubMed]
- Szabo, B.; Schlicker, E. Effects of Cannabinoids on neurotransmission. Handb. Exp. Pharmacol. 2005, 168, 327–365. [Google Scholar] [CrossRef]
- Hoffman, A.F.; Lupica, C.R. Synaptic targets of Delta9-tetrahydrocannabinol in the central nervous system. Cold Spring Harb. Perspect. Med. 2013, 3, a039743. [Google Scholar] [CrossRef]
- Kennedy, M.B. Synaptic Signaling in Learning and Memory. Cold Spring Harb. Perspect. Biol. 2013, 8, a016824. [Google Scholar] [CrossRef]
- Gugliandolo, A.; Silvestro, S.; Chiricosta, L.; Pollastro, F.; Bramanti, P.; Mazzon, E. The Transcriptomic Analysis of NSC-34 Motor Neuron-Like Cells Reveals That Cannabigerol Influences Synaptic Pathways: A Comparative Study with Cannabidiol. Life 2020, 10, 227. [Google Scholar] [CrossRef] [PubMed]
- Moretto, E.; Murru, L.; Martano, G.; Sassone, J.; Passafaro, M. Glutamatergic synapses in neurodevelopmental disorders. Prog. Neuropsychopharmacol. Biol. Psychiatry 2018, 84, 328–342. [Google Scholar] [CrossRef]
- Pittenger, C.; Bloch, M.H.; Williams, K. Glutamate abnormalities in obsessive compulsive disorder: Neurobiology, pathophysiology, and treatment. Pharmacol. Ther. 2011, 132, 314–332. [Google Scholar] [CrossRef] [PubMed]
- Traynelis, S.F.; Wollmuth, L.P.; McBain, C.J.; Menniti, F.S.; Vance, K.M.; Ogden, K.K.; Hansen, K.B.; Yuan, H.; Myers, S.J.; Dingledine, R. Glutamate receptor ion channels: Structure, regulation, and function. Pharmacol. Rev. 2010, 62, 405–496. [Google Scholar] [CrossRef] [PubMed]
- Miladinovic, T.; Nashed, M.G.; Singh, G. Overview of Glutamatergic Dysregulation in Central Pathologies. Biomolecules 2015, 5, 3112–3141. [Google Scholar] [CrossRef]
- Kobayashi, K. Role of catecholamine signaling in brain and nervous system functions: New insights from mouse molecular genetic study. J. Investig. Dermatol. Symp. Proc. 2001, 6, 115–121. [Google Scholar] [CrossRef]
- Fernandez-Ruiz, J.; Hernandez, M.; Ramos, J.A. Cannabinoid-dopamine interaction in the pathophysiology and treatment of CNS disorders. CNS Neurosci. Ther. 2010, 16, e72–e91. [Google Scholar] [CrossRef]
- Teleanu, R.I.; Niculescu, A.G.; Roza, E.; Vladacenco, O.; Grumezescu, A.M.; Teleanu, D.M. Neurotransmitters-Key Factors in Neurological and Neurodegenerative Disorders of the Central Nervous System. Int. J. Mol. Sci. 2022, 23, 5954. [Google Scholar] [CrossRef]
- Domino, E.F. Cannabinoids and the cholinergic system. J. Clin. Pharmacol. 1981, 21, 249S–255S. [Google Scholar] [CrossRef]
- Vanin, A.P.; Tamagno, W.A.; Alves, C.; Mesacasa, L.; Santin, L.F.; Sutorillo, N.T.; Bilibio, D.; Muller, C.; Galon, L.; Kaizer, R.R. Neuroprotective potential of Cannabis sativa-based oils in Caenorhabditis elegans. Sci. Rep. 2022, 12, 15376. [Google Scholar] [CrossRef]
- Pundir, C.S.; Chauhan, N. Acetylcholinesterase inhibition-based biosensors for pesticide determination: A review. Anal. Biochem. 2012, 429, 19–31. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhyaya, B. Molecular mechanisms underlying activity-dependent GABAergic synapse development and plasticity and its implications for neurodevelopmental disorders. Neural Plast. 2011, 2011, 734231. [Google Scholar] [CrossRef] [PubMed]
- Lu, W.; Bromley-Coolidge, S.; Li, J. Regulation of GABAergic synapse development by postsynaptic membrane proteins. Brain Res. Bull. 2017, 129, 30–42. [Google Scholar] [CrossRef] [PubMed]
- Lotan, R. Retinoids in cancer chemoprevention. FASEB J. 1996, 10, 1031–1039. [Google Scholar] [CrossRef]
- Melino, G.; Thiele, C.J.; Knight, R.A.; Piacentini, M. Retinoids and the control of growth/death decisions in human neuroblastoma cell lines. J. Neuro-Oncol. 1997, 31, 65–83. [Google Scholar] [CrossRef]
- Khwanraj, K.; Phruksaniyom, C.; Madlah, S.; Dharmasaroja, P. Differential Expression of Tyrosine Hydroxylase Protein and Apoptosis-Related Genes in Differentiated and Undifferentiated SH-SY5Y Neuroblastoma Cells Treated with MPP+. Neurol. Res. Int. 2015, 2015, 734703. [Google Scholar] [CrossRef]
- Daubner, S.C.; Le, T.; Wang, S. Tyrosine hydroxylase and regulation of dopamine synthesis. Arch. Biochem. Biophys. 2011, 508, 1–12. [Google Scholar] [CrossRef]
- Gerdeman, G.; Lovinger, D.M. CB1 cannabinoid receptor inhibits synaptic release of glutamate in rat dorsolateral striatum. J. Neurophysiol. 2001, 85, 468–471. [Google Scholar] [CrossRef]
- Morera-Herreras, T.; Ruiz-Ortega, J.A.; Ugedo, L. Two opposite effects of Delta(9)-tetrahydrocannabinol on subthalamic nucleus neuron activity: Involvement of GABAergic and glutamatergic neurotransmission. Synapse 2010, 64, 20–29. [Google Scholar] [CrossRef]
- Kelley, B.G.; Thayer, S.A. Delta 9-tetrahydrocannabinol antagonizes endocannabinoid modulation of synaptic transmission between hippocampal neurons in culture. Neuropharmacology 2004, 46, 709–715. [Google Scholar] [CrossRef]
- Bara, A.; Ferland, J.N.; Rompala, G.; Szutorisz, H.; Hurd, Y.L. Cannabis and synaptic reprogramming of the developing brain. Nat. Rev. Neurosci. 2021, 22, 423–438. [Google Scholar] [CrossRef] [PubMed]
- Petroff, O.A. GABA and glutamate in the human brain. Neuroscientist 2002, 8, 562–573. [Google Scholar] [CrossRef]
- Filpa, V.; Moro, E.; Protasoni, M.; Crema, F.; Frigo, G.; Giaroni, C. Role of glutamatergic neurotransmission in the enteric nervous system and brain-gut axis in health and disease. Neuropharmacology 2016, 111, 14–33. [Google Scholar] [CrossRef] [PubMed]
- Panther, E.J.; Dodd, W.; Clark, A.; Lucke-Wold, B. Gastrointestinal Microbiome and Neurologic Injury. Biomedicines 2022, 10, 500. [Google Scholar] [CrossRef]
- Zhou, Y.; Danbolt, N.C. GABA and Glutamate Transporters in Brain. Front. Endocrinol. 2013, 4, 165. [Google Scholar] [CrossRef] [PubMed]
- Luscher, C.; Xia, H.; Beattie, E.C.; Carroll, R.C.; von Zastrow, M.; Malenka, R.C.; Nicoll, R.A. Role of AMPA receptor cycling in synaptic transmission and plasticity. Neuron 1999, 24, 649–658. [Google Scholar] [CrossRef]
- Gambrill, A.C.; Storey, G.P.; Barria, A. Dynamic regulation of NMDA receptor transmission. J. Neurophysiol. 2011, 105, 162–171. [Google Scholar] [CrossRef] [PubMed]
- Sanderson, D.J.; Sprengel, R.; Seeburg, P.H.; Bannerman, D.M. Deletion of the GluA1 AMPA receptor subunit alters the expression of short-term memory. Learn. Memmory 2011, 18, 128–131. [Google Scholar] [CrossRef]
- Islam, M.S. Calcium Signaling: From Basic to Bedside. Adv. Exp. Med. Biol. 2020, 1131, 1–6. [Google Scholar] [CrossRef]
- Lipton, S.A. Paradigm shift in neuroprotection by NMDA receptor blockade: Memantine and beyond. Nat. Rev. Drug Discov. 2006, 5, 160–170. [Google Scholar] [CrossRef]
- Naisbitt, S.; Kim, E.; Tu, J.C.; Xiao, B.; Sala, C.; Valtschanoff, J.; Weinberg, R.J.; Worley, P.F.; Sheng, M. Shank, a novel family of postsynaptic density proteins that binds to the NMDA receptor/PSD-95/GKAP complex and cortactin. Neuron 1999, 23, 569–582. [Google Scholar] [CrossRef] [PubMed]
- Ha, H.T.T.; Leal-Ortiz, S.; Lalwani, K.; Kiyonaka, S.; Hamachi, I.; Mysore, S.P.; Montgomery, J.M.; Garner, C.C.; Huguenard, J.R.; Kim, S.A. Shank and Zinc Mediate an AMPA Receptor Subunit Switch in Developing Neurons. Front. Mol. Neurosci. 2018, 11, 405. [Google Scholar] [CrossRef]
- Yang, M.; Bozdagi, O.; Scattoni, M.L.; Wohr, M.; Roullet, F.I.; Katz, A.M.; Abrams, D.N.; Kalikhman, D.; Simon, H.; Woldeyohannes, L.; et al. Reduced excitatory neurotransmission and mild autism-relevant phenotypes in adolescent Shank3 null mutant mice. J. Neurosci. 2012, 32, 6525–6541. [Google Scholar] [CrossRef] [PubMed]
- Castelli, V.; Brancato, A.; Cavallaro, A.; Lavanco, G.; Cannizzaro, C. Homer2 and Alcohol: A Mutual Interaction. Front. Psychiatry 2017, 8, 268. [Google Scholar] [CrossRef] [PubMed]
- Petralia, R.S.; Wang, Y.X.; Sans, N.; Worley, P.F.; Hammer, J.A., 3rd; Wenthold, R.J. Glutamate receptor targeting in the postsynaptic spine involves mechanisms that are independent of myosin Va. Eur. J. Neurosci. 2001, 13, 1722–1732. [Google Scholar] [CrossRef] [PubMed]
- Maher, B.J.; Mackinnon, R.L., 2nd; Bai, J.; Chapman, E.R.; Kelly, P.T. Activation of postsynaptic Ca(2+) stores modulates glutamate receptor cycling in hippocampal neurons. J. Neurophysiol. 2005, 93, 178–188. [Google Scholar] [CrossRef]
- Neyman, S.; Manahan-Vaughan, D. Metabotropic glutamate receptor 1 (mGluR1) and 5 (mGluR5) regulate late phases of LTP and LTD in the hippocampal CA1 region in vitro. Eur. J. Neurosci. 2008, 27, 1345–1352. [Google Scholar] [CrossRef]
- Chen, J.S.; Exton, J.H. Regulation of phospholipase D2 activity by protein kinase C alpha. J. Biol. Chem. 2004, 279, 22076–22083. [Google Scholar] [CrossRef]
- Raben, D.M.; Barber, C.N. Phosphatidic acid and neurotransmission. Adv. Biol. Regul. 2017, 63, 15–21. [Google Scholar] [CrossRef]
- Yogeswara, I.B.A.; Maneerat, S.; Haltrich, D. Glutamate Decarboxylase from Lactic Acid Bacteria-A Key Enzyme in GABA Synthesis. Microorganisms 2020, 8, 1923. [Google Scholar] [CrossRef]
- Wisden, W.; Yu, X.; Franks, N.P. GABA Receptors and the Pharmacology of Sleep. Handb. Exp. Pharmacol. 2019, 253, 279–304. [Google Scholar] [CrossRef]
- Zhang, D.; Pan, Z.H.; Awobuluyi, M.; Lipton, S.A. Structure and function of GABA(C) receptors: A comparison of native versus recombinant receptors. Trends Pharm. Sci. 2001, 22, 121–132. [Google Scholar] [CrossRef] [PubMed]
- Ghit, A.; Assal, D.; Al-Shami, A.S.; Hussein, D.E.E. GABA(A) receptors: Structure, function, pharmacology, and related disorders. J. Genet. Eng. Biotechnol. 2021, 19, 123. [Google Scholar] [CrossRef] [PubMed]
- Picciotto, M.R.; Higley, M.J.; Mineur, Y.S. Acetylcholine as a neuromodulator: Cholinergic signaling shapes nervous system function and behavior. Neuron 2012, 76, 116–129. [Google Scholar] [CrossRef] [PubMed]
- Hogg, R.C.; Raggenbass, M.; Bertrand, D. Nicotinic acetylcholine receptors: From structure to brain function. Rev. Physiol. Biochem. Pharmacol. 2003, 147, 1–46. [Google Scholar] [CrossRef] [PubMed]
- O’Grady, G.L.; Verschuuren, C.; Yuen, M.; Webster, R.; Menezes, M.; Fock, J.M.; Pride, N.; Best, H.A.; Benavides Damm, T.; Turner, C.; et al. Variants in SLC18A3, vesicular acetylcholine transporter, cause congenital myasthenic syndrome. Neurology 2016, 87, 1442–1448. [Google Scholar] [CrossRef]
- Dani, J.A. Neuronal Nicotinic Acetylcholine Receptor Structure and Function and Response to Nicotine. Int. Rev. Neurobiol. 2015, 124, 3–19. [Google Scholar] [CrossRef]
- Tarnawski, L.; Reardon, C.; Caravaca, A.S.; Rosas-Ballina, M.; Tusche, M.W.; Drake, A.R.; Hudson, L.K.; Hanes, W.M.; Li, J.H.; Parrish, W.R.; et al. Adenylyl Cyclase 6 Mediates Inhibition of TNF in the Inflammatory Reflex. Front. Immunol. 2018, 9, 2648. [Google Scholar] [CrossRef]
- Shen, J.X.; Yakel, J.L. Nicotinic acetylcholine receptor-mediated calcium signaling in the nervous system. Acta Pharmacol. Sin. 2009, 30, 673–680. [Google Scholar] [CrossRef]
- Resende, R.R.; Adhikari, A. Cholinergic receptor pathways involved in apoptosis, cell proliferation and neuronal differentiation. Cell Commun. Signal. 2009, 7, 20. [Google Scholar] [CrossRef]
- Klein, M.O.; Battagello, D.S.; Cardoso, A.R.; Hauser, D.N.; Bittencourt, J.C.; Correa, R.G. Dopamine: Functions, Signaling, and Association with Neurological Diseases. Cell. Mol. Neurobiol. 2019, 39, 31–59. [Google Scholar] [CrossRef] [PubMed]
- Guenter, J.; Lenartowski, R. Molecular characteristic and physiological role of DOPA-decarboxylase. Postep. Hig. Med. Dosw. 2016, 70, 1424–1440. [Google Scholar] [CrossRef] [PubMed]
- Beaulieu, J.M.; Gainetdinov, R.R. The physiology, signaling, and pharmacology of dopamine receptors. Pharmacol. Rev. 2011, 63, 182–217. [Google Scholar] [CrossRef]
- Ha, C.M.; Park, D.; Han, J.K.; Jang, J.I.; Park, J.Y.; Hwang, E.M.; Seok, H.; Chang, S. Calcyon forms a novel ternary complex with dopamine D1 receptor through PSD-95 protein and plays a role in dopamine receptor internalization. J. Biol. Chem. 2012, 287, 31813–31822. [Google Scholar] [CrossRef]
- Kalinovskii, A.P.; Osmakov, D.I.; Koshelev, S.G.; Lubova, K.I.; Korolkova, Y.V.; Kozlov, S.A.; Andreev, Y.A. Retinoic Acid-Differentiated Neuroblastoma SH-SY5Y Is an Accessible In Vitro Model to Study Native Human Acid-Sensing Ion Channels 1a (ASIC1a). Biology 2022, 11, 167. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Lazareva-Ulitsky, B.; Loo, R.; Kejariwal, A.; Vandergriff, J.; Rabkin, S.; Guo, N.; Muruganujan, A.; Doremieux, O.; Campbell, M.J.; et al. The PANTHER database of protein families, subfamilies, functions and pathways. Nucleic Acids Res. 2005, 33, D284–D288. [Google Scholar] [CrossRef]
- Carbon, S.; Ireland, A.; Mungall, C.J.; Shu, S.; Marshall, B.; Lewis, S.; Ami, G.O.H.; Web Presence Working, G. AmiGO: Online access to ontology and annotation data. Bioinformatics 2009, 25, 288–289. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, M.; Jassal, B.; Stephan, R.; Milacic, M.; Rothfels, K.; Senff-Ribeiro, A.; Griss, J.; Sevilla, C.; Matthews, L.; Gong, C.; et al. The reactome pathway knowledgebase 2022. Nucleic Acids Res. 2022, 50, D687–D692. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.H.; Hazekamp, A.; Peltenburg-Looman, A.M.; Frederich, M.; Erkelens, C.; Lefeber, A.W.; Verpoorte, R. NMR assignments of the major cannabinoids and cannabiflavonoids isolated from flowers of Cannabis sativa. Phytochem. Anal. PCA 2004, 15, 345–354. [Google Scholar] [CrossRef] [PubMed]
- Gaoni, Y.; Mechoulam, R. The isolation and structure of delta-1-tetrahydrocannabinol and other neutral cannabinoids from hashish. J. Am. Chem. Soc. 1971, 93, 217–224. [Google Scholar] [CrossRef]


| Gene | CTRL | Δ8-THC | log2 Fold Change | q-Value | Protein |
|---|---|---|---|---|---|
| GLS2 | 120.39 | 151.17 | 0.33 | 2.38 × 10−2 | GLS |
| GRIA1 | 494.82 | 561.49 | 0.18 | 1.19 × 10−2 | AMPA receptor |
| GRIA2 | 1928.70 | 2098.12 | 0.12 | 8.32 × 10−4 | |
| GRIA3 | 1727.64 | 1915.39 | 0.15 | 8.36 × 10−5 | |
| GRIA4 | 541.77 | 745.06 | 0.46 | 1.46 × 10−13 | |
| GRIN2C | 2.41 | 13.29 | 2.46 | 4.22 × 10−3 | NMDA receptor |
| HOMER2 | 816.27 | 1007.53 | 0.30 | 6.95 × 10−9 | Homer |
| SHANK3 | 1790.25 | 1910.40 | 0.09 | 1.52 × 10−2 | SHANK |
| ITPR1 | 9393.09 | 9664.99 | 0.04 | 1.53 × 10−2 | IP3R |
| ITPR2 | 1542.24 | 1645.44 | 0.09 | 2.57 × 10−2 | |
| PLCB1 | 1167.82 | 1269.17 | 0.12 | 1.16 × 10−2 | PLC |
| PLD2 | 1390.54 | 1577.33 | 0.18 | 1.25 × 10−5 | PLD |
| PRKCA | 8513.01 | 9226.42 | 0.12 | 1.58 × 10−12 | PKC |
| PRKCG | 87.89 | 136.22 | 0.63 | 6.35 × 10−5 |
| Gene | CTRL | Δ8-THC | log2 Fold Change | q-Value | Synapsis |
|---|---|---|---|---|---|
| GABRA3 | 551.40 | 616.31 | 0.16 | 2.03 × 10−2 | Gabaergic |
| GABRG3 | 50.57 | 98.84 | 0.97 | 1.12 × 10−6 | Gabaergic |
| GABRR2 | 30.10 | 17.44 | −0.79 | 2.27 × 10−2 | Gabaergic |
| CHRNA3 | 3521.51 | 3364.80 | −0.07 | 2.01 × 10−2 | Cholinergic |
| CHRNA6 | 15.65 | 1.66 | −3.24 | 1.60 × 10−4 | Cholinergic |
| CHRNA7 | 2073.17 | 1844.79 | −0.17 | 2.26 × 10−6 | Cholinergic |
| CHRNB2 | 9219.72 | 8367.57 | −0.14 | 5.31 × 10−18 | Cholinergic |
| PRKACB | 3247.01 | 3042.53 | −0.09 | 1.17 × 10−3 | Cholinergic |
| MAPK1 | 1856.47 | 1716.04 | −0.11 | 3.28 × 10−3 | Cholinergic |
| CREB3L3 | 13.24 | 4.98 | −1.41 | 1.67 × 10−2 | Cholinergic |
| CREB3L4 | 1438.70 | 1281.63 | −0.17 | 1.24 × 10−4 | Cholinergic |
| CREB5 | 2677.55 | 2544.16 | −0.07 | 2.34 × 10−2 | Cholinergic |
| SLC18A3 | 280.52 | 213.47 | −0.39 | 1.11 × 10−4 | Cholinergic |
| DRD4 | 108.35 | 54.82 | −0.98 | 3.20 × 10−8 | Dopaminergic |
| CALY | 7.22 | 1.66 | −2.12 | 2.62 × 10−2 | Dopaminergic |
| DDC | 13,066.29 | 12,497.37 | −0.06 | 4.46 × 10−6 | Dopaminergic |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Anchesi, I.; Schepici, G.; Chiricosta, L.; Gugliandolo, A.; Salamone, S.; Caprioglio, D.; Pollastro, F.; Mazzon, E. Δ8-THC Induces Up-Regulation of Glutamatergic Pathway Genes in Differentiated SH-SY5Y: A Transcriptomic Study. Int. J. Mol. Sci. 2023, 24, 9486. https://doi.org/10.3390/ijms24119486
Anchesi I, Schepici G, Chiricosta L, Gugliandolo A, Salamone S, Caprioglio D, Pollastro F, Mazzon E. Δ8-THC Induces Up-Regulation of Glutamatergic Pathway Genes in Differentiated SH-SY5Y: A Transcriptomic Study. International Journal of Molecular Sciences. 2023; 24(11):9486. https://doi.org/10.3390/ijms24119486
Chicago/Turabian StyleAnchesi, Ivan, Giovanni Schepici, Luigi Chiricosta, Agnese Gugliandolo, Stefano Salamone, Diego Caprioglio, Federica Pollastro, and Emanuela Mazzon. 2023. "Δ8-THC Induces Up-Regulation of Glutamatergic Pathway Genes in Differentiated SH-SY5Y: A Transcriptomic Study" International Journal of Molecular Sciences 24, no. 11: 9486. https://doi.org/10.3390/ijms24119486
APA StyleAnchesi, I., Schepici, G., Chiricosta, L., Gugliandolo, A., Salamone, S., Caprioglio, D., Pollastro, F., & Mazzon, E. (2023). Δ8-THC Induces Up-Regulation of Glutamatergic Pathway Genes in Differentiated SH-SY5Y: A Transcriptomic Study. International Journal of Molecular Sciences, 24(11), 9486. https://doi.org/10.3390/ijms24119486







