Abstract
Diabetes is a metabolic disorder characterized by lower responsiveness of tissues to insulin and consequent large variations in circulating levels of glucose. This fluctuation has harmful effects as both hyperglycemia and hypoglycemia can be very injurious. The causes of diabetes are varied but the consequences are rather uniform. Dietary factors are important especially in adult onset type 2 diabetes (T2D) while type 1 diabetes (T1D) is characterized by having a stronger heritable component and involving autoimmune attach on pancreatic beta cells. This review is focused on the relation of the bacterial components found within the intestine, to the establishment and maintenance of diabetes. The precise composition of the gut microbiome is increasingly recognized as a factor in organismic health and its interaction with a variety of disease states has been described. This is especially marked in the case of diabetes since the nature of the diet is an important factor in establishing both the microbiome and the incidence of diabetes. The bidirectional nature of this relationship is discussed. The effects of disease that lead to altered microbiomal composition together with aberrant metabolic changes are also included. Emphasis is given to the important role of short chain fatty acids (SCFAs) as mediators of the microbiome-diabetes relation.
1. Introduction
The incidence of Type 2 diabetes (T2D) is growing rapidly worldwide [1]. To a large degree, this reflects the transition from consumption of less refined native foodstuffs toward a modern industrially produced diet rich in rapidly assimilable processed materials. This shift has also resulted in widespread gains in body weight, so that while in the past, obesity was associated with affluence, it is now more prevalent among lower income groups. While Type 1 diabetes (T1D) is generally found among a much younger age group, it is increasing equally rapidly as T2D. An over 100-fold variation of the extent of its incidence exists between various countries, being highest in Scandinavian countries and lowest in least developed countries [2]. This suggests that T1D prevalence is also strongly affected by environmental factors.
While the various forms of diabetes have a significant overlap in some aspects, the distinctions between them are briefly discussed in the next section.
The components of the diet, are a key determinant of a predisposition to diabetes. Diet is also an important factor in establishing the nature of the bacterial, viral and fungal elements that constitute the intestinal microbiome. The elements within the heterogeneous collection of microbiota are variable and in constant flux. Inappropriate composition of the gut microbiota and resulting abnormal content of soluble metabolic factors have been related to increases in adiposity, chronic inflammation and insulin resistance and imbalance of microbiome composition is common in pathological states such as obesity and metabolic syndrome, and diabetes [3].
The gut microbiomal composition both influences and is sensitive to, the state of health of the host. This review is focused on how the microbiome can influence the development and progression of diabetes, and conversely, how the disease may have an impact on the microbiome. The molecular processes that may underlie this reciprocal relationship are considered.
2. Type 1 and 2 Diabetes, Similarities and Distinctions
The two major types of diabetes have been classically distinguished in several general ways. Type 1 diabetes (T1D) is described as a disease with a large genetic component that involves an auto-immune attack on insulin producing pancreatic beta cells. It therefore tends to develop early in life and involves failure of insulin synthesis. T1D involves irreversible destruction of insulin producing cells, and autoantibodies to pancreatic b-cells can to detected long before the onset of clinical disease [4]. This disruption leads to wide fluctuations of levels of circulating glucose. Both hyperglycemia and hypoglycemia can rapidly lead to dangerous toxic effects. Disease onset can be very sudden. As T1D is essentially a deficiency disease, the most effective treatment is replacement therapy by administration of insulin and regular monitoring of blood glucose levels.
Type 2 diabetes (T2D) develops more slowly and progressively, generally later in life and may reflect lifestyle patterns of diet and exercise. In this disease insulin may still be produced but tissues become insulin resistant and do not respond to this hormone by adequately increasing uptake of circulating glucose. T2D is associated with excess body weight, physical inactivity, but additionally, has a genetic component. Epigenetic changes that can be regulated by metabolites produced by intestinal bacteria are likely important to establishing the onset of diabetes. The epigenetic profile determines the spectrum of proteins produced and this can influence the promotion or retardation of onset of the disorder. For example, histone deacetylases can be inhibited by butyrate produced by bacterial fermentation and can increase the degree of transcription of several genes associated with production of anti-inflammatory factors [5]. Likewise, expression of the microRNAs, miR-181a and miR-181b, which are associated with development of obesity and insulin resistance, is down-regulated by bacterially produced beneficial tryptophan metabolites [6].
In recent years, the degree of overlap between T1D and T2D has been increasingly recognized. Both have an association with the gene expression profile superimposed on which are environmental factors. Even among identical twins the extent of concordance of T1D is only 16% [7], while in the case of T2D it is 6% [8]. A degree of autoimmune attack on pancreatic insulin producing b-cells which is a key feature of T1D, is also found in T2D [9]. The age of onset of T1D is typically in childhood but can occur in adult life while T2D is also occasionally found in children. Other commonalities include the increased risk of either T1D or T2D following preterm birth [10] and genetic and molecular susceptibilities shared between the two variants [11]. Both variants of the disorder are also characterized by increased intestinal permeability and diminished bacterial diversity in the gut (discussed below).
This emerging convergence of perspectives on these variants of diabetes has led to identification of a new disorder reflecting a combination of their characteristics, latent autoimmune diabetes of adults (LADA) [12]. The identification of such an intermediate form has led to perception of diabetes as somewhat of a graded disease continuum.
3. Composition of the Human Microbiome and Relevance to Diabetes
The human intestinal tract has a surface area of around 300 m3, presenting a large interface with exogenous factors originating in the environment. A complex symbiotic relationship has evolved between these microbiota of which there are are over 2000 individual species, and the host. These vary considerably in composition during stages of maturation and also between geographic regions [13]. The breadth and variation of the intestinal microbiome is influence by many environmental factors. While the 12 phyla known to exist with the microbiome can be subdivided into many genera and a host of species, the focus of this list is upon those predominant phyla, members of which often bear an underlying similarity of their characteristics. Some distinctive and important species are also reviewed.
3.1. Firmicutes
The most predominant phylum in the adult human intestine are the Firmicutes. This phylum includes Clostridium, Lactobacillus and Fecalibacterium species, many of which are associated with production of butyrate and other short chain fatty acids (SCFA) which have been reported to attenuate inflammatory and allergenic activity [14].
Relevance to Diabetes
The content of differing Lactobacillus species in the microbiome in T2D can be differ dramatically. While the content of L. acidophilus, L. gasseri, L. salivarius are all increased in T2D, L. amylovorus is decreased [15]. Generalizations involving even a single genus cannot always be made but overall levels of butyrate -producing Firmicutes species are broadly decreased in T2D [16].
3.2. Bacteriodetes
The second most dominant phylum are Bacteriodetes which include several anaerobic Bacteriodes species such as B. dorei, B. fragilis and B. thetaiotaomicron. Bacteroides fragilis is a common cause of intra-abdominal infections in humans.
Prevotella is another genus found within this phylum which tends to promote inflammatory changes. T1D is associated with an elevated presence of Bacteriodetes
Non-vegetararians have a large excess of Bacteriodetes over Firmicutes species but this is ratio reversed in vegetarians [17].
Relevance to Diabetes
While butyrate is principally produced by Firmicutes, propionate and acetate are largely produced by Bacteroidetes [18]. Butyrate seems to be the most potent SCFA in attenuating inflammatory responses by reducing the accumulation of neutrophils and pro-inflammatory macrophages in the M1 conformation [19].
Levels of B.dorei have been used as s predictor for T1D onset [20].
Bacteriodetes and Actinobacteria are overrepresented in the microbiome of T2D [15] (Gurung et al., 2020). B. thetaiotaomicron reduced plasma glutamate concentrations in mice, and lowered diet-induced excess weight gain in mice [21].
3.3. Actinobacteria
The most abundant genus within actinobacteria is the Bifidobacteria. This genus constitutes around 5% of the adult microbiome but is over 90% of that of the infants under 2 years old. Species such as Bifidobacterium adolescentis can promote digestion of milk constituents and synthesize several B vitamins. Its administration has found therapeutic utility since 1899. Bifidobacteria are able to digest fiber to produce butyrate and also seem to suppress autoimmune activity. Lactobacillus and various Firmicutes (Roseburia and Eubacterium) also promote butyrate production [22].
Relevance to Diabetes
Inflammation is an important etiological factor for insulin resistance, which may lead to the development of T2DM [23] and several strains of Bifidobacteria have been found to inhibit inflammation by blocking NF-κB activation [24].
Fecal L. acidophilus content is elevated in uncontrolled diabetic patients [25]. Probiotic supplements that contain Bifidobacterium together with Lactobacillus strains may improve insulin sensitivity and reduces fasting levels of plasma glucose in diabetes [26].
3.4. Verrucomicrobiota
The phylum in represented in the gut by a sole species, Akkermansia muciniphila which colonizes the mucosal layer of the gut and is able to degrade mucin to acetate. This species is relatively prevalent in that it constitutes 0.5–5% of the total intestinal bacterial population.
Relevance to Diabetes
Obesity leads to a range of disease consequences among which diabetes is a major hazard. There is a clear association between obesity and T2D. Increased intestinal levels of Akkermansia muciniphila lead to inhibition of diet induced obesity and diabetes in mice [27]. Conversely, decreased abundance of this bacterium impairs glucose homeostasis [28]. The attenuation of diabetes in the presence of this this bacterium has recently been reviewed [27].
3.5. Proteobacteria
This phylum which constitutes less than 1% of the healthy human microbiome, includes the family Enterobacteriacea. An increased abundance of members belonging to this family, especially Escherichia coli a pathogen that has been associated with several inflammatory disease states including metabolic disorders and inflammatory bowel disease [29].
Relevance to Diabetes
An excess intestinal presence of E. coli at the expense of anti-inflammatory butyrate producing bacteria in T2D and in pre-diabetics has been reported on several occasions [30]. It has also been found in excessive amounts in T1D [31].
3.6. Bacillota
This phylum of anaerobes includes the Blauta genus, members of which have inflammation-suppressing qualities and antibiotic activity against specific pathogens [32] and Ruminococcus is important for digestion of resistant starches [33].
Relevance to Diabetes
Levels of these bacterial species within this phylum are elevated in T2D [15].
The content of butyrate-producing bacteria Lactobacillus (from the Bacillota phylum) is reduced in patients with T2D. These bacteria have also been reported to have anti-allergic effects [34] and to modulate gene expression in a desirable direction tending toward the alleviation of diabetes [35].
4. Relative Prevalence of Differing Bacterial Classes
An enrichment of pro-inflammatory bacteria (e.g., Escherichia coli) at the expense of anti-inflammatory bacteria (e.g., Fecalibacterium prausnitzii) in T2D has been reported on several occasions [30]. While it is clear that the predominance of various microbial species is altered in T2D, the causal relationships are unclear and whether these changes lead to T2D or are a response to the disorder is not well established. Furthermore, if the second possibility is the case, it is not firmly established if changes in microbiome composition represent a positive defensive response to the disorder or have a more damaging consequence [36]). Causality requires that when all confounders are allowed for, inducing change in one parameter should lead to modifications of a different parameter in a reproducible manner. These can only be shown by using delineated interventional experiments.
Much use has been made of the degree of prevalence of members of the Firmicutes phylum relative to that of those within Bacteriodetes. This widely used ratio (F/B) is reported as low in diabetes but high in obesity [37]. An increase in the F/B ratio has also been seen in obese mice [14]. Obesity and diabetes seem to be linked with different groups of intestinal microbiota [38]. In lean patients with T2D the F/B ratio is higher than in those with the more common form of the disease associated with excess fat deposition [39].
Since the severity of T2D in obese subjects is generally improved after weight loss, T2D in lean people may be a different disease from the diabetes associated with obesity. Malnutrition-related diabetes has several features that distinguish it from obesity-related diabetes [40]. Malnutrition is known to limit diversity of the gut microbiome but any association of this with diabetes has not been studied. The major defect in T2D of lean patients may involve a reduced capacity for insulin secretion rather than increasing insulin resistance [41,42].
Additionally, the number of contradictory reports concerning this use of the F/B ratio with respect to diabetes may be attributable to the existence of many potentially confounding factors such as selection of subjects, extent of physical activity, presence or absence of contemporaneous obesity [15]. In addition, disparities may exist due to methodological variations used. Taxonomic study of gut bacteria is often carried out performed by sequencing of 16S rRNA to allow precise characterization of microbial species. There can be variation of primers used in amplification of the 16S rRNA region, the sequencing technique, and analysis of data [43].
5. Bacterial Diversity
The appearance of diabetes-associated autoantibodies in T1D is preceded by an overall reduced diversity of the spectrum of intestinal microbiota [44]. Especially prominent is an elevated proportion of Bacteroides species [45,46] and a profound depression of Akkermansia [44]. The reduced overall palette of bacterial diversity combined with the disproportionate presence of Bacteriodetes may lead to abnormal development of the immune system and the appearance of immune related abnormalities including a trend toward increased autoimmune and inflammatory activity [47]. A similar situation whereby imbalance of specific bacterial species within the microbiome can promote autoimmune and inflammatory responses may exist in T2D [48]. Factors released from certain bacterial classes within the microbiome can be recognized and responded to by epithelial cells and immune cells, leading to a defensive inflammatory response by effector T lymphocytes. The extent of this reaction is delimited by regulatory T lymphocytes, induced by butyrate produced by several bacterial species especially those within the Firmicutes phylum. A distortion of the normal composition of the microbiome resulting from diminution of such Firmicutes, can lead to excess inflammatory activity. This information may reach the hypothalamus by way of the enteric nervous system and thereby lead to systemic hyperglycemia and insulin-resistance [49].
6. Bacterial Regulation of Intestinal Permeability
An increased intestinal permeability due to diminished integrity of tight junctions, enables dietary antigens and immune stimulants such as microbial lipopolysaccharides, to become systemically distributed. This can promote general inflammatory activity and specific autoimmune attack on the pancreatic β-cells. These damaging events are found in both T1D and T2D and are inhibited by butyrate [44,50]. This is enabled by stimulation of increased expression of a range of tight junction proteins. Inhibition of the activation of NLRP3 inflammasome and autophagy induced by LPS are inhibited by all SCFAs [51]. However, the potentially protective role of propionate and acetate remains uncertain since they have also been reported to have an opposite effect by impairing the assembly of tight junction proteins [44].
Increasing evidence suggests that gut leakiness is a cause rather than a consequence of the diabetic condition. Such leakiness has been reported to occur prior to the onset of T1D [52]. The hyperglycemia that often precedes overt T2D has been found to disrupt the tight adherence and barrier function of the intestinal epithelial cells [53].
7. Modulation of the Microbiome as a Means of Treating Diabetes
This strategy may be of especial value in T1D where pancreatic b-cells are lost before the onset of clinical symptoms of the disease [54]. Since T1D has a major genetic component, early detection of pre-diabetes can give a long period for initiation of attempted mitigation by modulation of the microbiome profile. T2D has also been improved by alteration in the intestinal microbiota [55]. Several means of enabling such changes are described below.
7.1. Diet
The constitution of the microbiome can rapidly by altered as a result of changes in the composition of the diet [56].
Dietary inclusion of modified starches can alter the numbers and activities of both autoreactive T cells and in this manner, may provide protection against autoimmune diabetes [45]. A causal relation between these events is likely but remains to be unambiguously demonstrated.
This diet rich in fibre and some plants compounds was associated with higher levels of Akkermansia muciniphila and Faecalibacterium prausnitzii, providing a reduction in endotoxemia in patients with T2DM [57]. The presence of non-digestible fibers, and complexes of starch and amylose in the diet are needed to serve as substrates to allow fermentation to butyrate. The useful qualities of butyrate are many fold and have been thought to include its acting as a histone deacetylase (HDAC) inhibitor or signaling through several G protein–coupled receptors (GPCRs) and consequent inhibition of NF-kB [58]. However, at lower concentrations, SCFA have recently been reported to activate a specific HDAC, acetyltransferase p300 [59]. Elevated levels of butyrate are associated with inhibition of NF-kB-associated histone deacetylases, leading to reduced activity of this inflammatory signaling pathway [60]. Short chain fatty acids including butyric acid (SCFA) are also reported to improve the function of the intestinal barrier which is impaired in both T1D and T2D. It may be that this is the major positive attribute of bacteria whose fermentation products include SCFA [61,62].
7.2. Exercise
Extended and regular exercise is known to improve glycemic control and to reduce systemic inflammation [63]. This is accompanied by reduced intestinal fungal mycetes overgrowth and intestinal permeability [64]. While exercise can increase microbiomal diversity [65], overall diet seems to be more important than exercise in establishing the composition and diversity of gut microbiota [66].
7.3. Pharmacological Agents
Metformin is an orally administered drug that is commonly used in T2D treatment. This agents acts by inhibition of hepatic gluconeogenesis. However, the molecular events underlying this are unclear [67]. Metformin reaches 300-fold higher concentrations in the gut than in the serum. Metformin treatment appears able to increase the proportion of Akkermansia muciniphila as well as butyrate-producing Bifidobacterium species [68], while decreasing the abundance of Bacteriodetes [56]. Its metabolic effects are likely to be mediated in part its actions in the intestine. It has been suggested that the predominant effect of metformin in regulating glucose levels, may involve decreasing the levels of B. fragilis and thus increasing content of the bile acid glycoursodeoxycholic acid leading to inhibition farnesoid X receptor (FXR) signaling [69]. T2D is characterized by microbiomal changes away from butyrate-producing taxa. The beneficial effects of metformin on T2D may be largely by increasing the relative abundance of microbes producing short-chain fatty acids [70].
The use of metformin has been sufficiently promising in studies on animals, that human studies are in progress [71].
A range of other pharmacological agents is emerging that involve indirect regulation of glucose levels. However, a significant portion of their utility is also likely to be based on their modification of the microbiome in an overall desirable direction [72]. These include the sulfonylureas which enhance insulin secretion by blocking ATP-sensitive K-channels, thiazolidinediones which increase sensitivity to insulin, alpha-glucosidase Inhibitors which retard the rate of oligosaccharide breakdown, and glucagon-like peptide-1 (GLP-1) receptor agonists which slows stomach emptying and increase insulin secretion. All of these also seem to drive the microbiome composition of the gut in a beneficial direction [72].
7.4. Probiotics
Many changes in the inflammatory profile, insulin resistance and bacterial composition in the gut of experimental animals have been reported following oral introduction of beneficial microorganisms such as Lactobacillus, Bifidobacterium or Akkermansia sp. into the diet. However, results from clinical studies have been more equivocal [73]. This is reminiscent of Alzheimer’s disease (AD) where rodent models of the disorder are easier to treat than are AD patients. The incomplete transference of success in animal models to humans may reflect the multifactorial nature of clinical disease and the much longer interval before manifestation of symptoms. These features probably result in the emergence of a more complicated disease profile in humans.
7.5. Fecal Transplantation
The introduction of bacterial species from healthy human to diabetic or pre-diabetic patients by this route attempts to improve the composition of the microbiome biome so as to create a profile with a more normal spectrum. Promising results have been obtained in experimental animals [74] and in human gestational diabetes [26]. Overall, there is growing evidence that the establishment of key bacterial species in the gut microbiome can lead an increased presence of SCFA as a result of fermentative activity [55]. The use of a single bacterial species (Akkermansia muciniphila) or even a specific membrane protein thereof, has been suggested as a more focused means of utilizing the desirable qualities of gut bacteria [75].
8. Summary
While this review has focused on beneficial products, notably SCFA, produced by the gut microbiome, it should be kept in mind that various classes of intestinal bacteria produce a range of harmful metabolites from dietary constituents and these are associated with obesity and diabetes. These include trimethylamine N-oxide produced from the choline and carnitine content of red meat and eggs, metabolites from the kyenurenine pathway of tryptophan breakdown, and imidazole propionate derived from histidine catabolism [76].
The close relation and many interaction between the diabetic state and the bacterial composition of contents of the intestine (summarized in Figure 1), offers a range of means of amelioration of this disorder. The incidence of diabetes can vary between different nations from as high as 30% of the population down to 5%. In addition, both childhood ands adult onset T1D also have a sharp variance between global regions [2]. This is somewhat surprising as genetic makeup plays a large role inT1D. It must be surmised that environmental factors are critical in determining the development of much T1D. Cultural and social determinants form the basis of most T2D. Thus, an inexpensive yet challenging means to dramatically alter the extent of the incidence of both types of diabetes can be by addressing and modifying extrinsic factors. The disease generally progresses slowly, giving much time for early interventions not involving sophisticated clinical medicine. The power of the intestinal microbiota in determining the state of organismic health is well illustrated in a recent report on kwashiorkor. Using animal models of this condition, a probiotic formulation served to optimize overall health despite severe malnutrition, as judged by reduction of weight loss, protection of immune function, improved resistance to infection and behavioral decline [77]. Addressing diabetes by way of revision of the microbiome spectrum is relatively non-invasive, and inexpensive. Furthermore, it does not involve extended drug usage which can at length induce further disruption of glucose homeostasis by provoking systemic responses contrary to those desired. However, improvements are only likely to be sustained by the patient’s development of a fresh lifestyle.
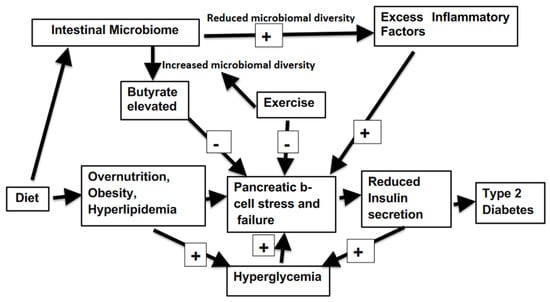
Figure 1.
Factors influencing microbiomal interaction with diabetes progression.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The author declares no conflict of interest with any of the studies described.
References
- Yang, G.; Wei, J.; Liu, P.; Zhang, Q.; Tian, Y.; Hou, G.; Meng, L.; Xin, Y.; Jiang, X. Role of the gut microbiota in type 2 diabetes and related diseases. Metabolism 2021, 117, 154712. [Google Scholar] [CrossRef] [PubMed]
- IDF Diabetes Atlas 2021–10th Edition. Available online: https://idf.org/ (accessed on 12 November 2022).
- Ortega, M.A.; Fraile-Martínez, O.; Naya, I.; García-Honduvilla, N.; Álvarez-Mon, M.; Buján, J.; Asúnsolo; De La Torre, B. Type 2 Diabetes Mellitus Associated with Obesity (Diabesity). The Central Role of Gut Microbiota and Its Translational Applications. Nutrients 2020, 12, 2749. [Google Scholar] [CrossRef] [PubMed]
- Ilonen, J.; Lempainen, J.; Veijola, R. The heterogeneous pathogenesis of type 1 diabetes mellitus. Nat. Rev. Endocrinol. 2019, 15, 635–650. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.H. Microbiota or short-chain fatty acids: Which regulates diabetes? Cell Mol. Immunol. 2017, 15, 88–91. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Li, Y.; Stoll, M.L.; Tollefsbol, T.O. The Epigenetic Connection Between the Gut Microbiome in Obesity and Diabetes. Front. Genet. 2020, 10, 1329. [Google Scholar] [CrossRef]
- Triolo, T.M.; Fouts, A.; Pyle, L.; Yu, L.; Gottlieb, P.A.; Steck, A.K.; the Type 1 Diabetes TrialNet Study Group; Greenbaum, C.J.; Atkinson, M.; Baidal, D.; et al. Identical and Nonidentical Twins: Risk and Factors Involved in Development of Islet Autoimmunity and Type 1 Diabetes. Diabetes Care 2018, 42, 192–199. [Google Scholar] [CrossRef]
- Willemsen, G.; Ward, K.J.; Bell, C.; Christensen, K.; Bowden, J.; Dalgård, C.; Harris, J.R.; Kaprio, J.; Lyle, R.; Magnusson, P.; et al. The Concordance and Heritability of Type 2 Diabetes in 34,166 Twin Pairs from International Twin Registers: The Discordant Twin (DISCOTWIN) Consortium. Twin Res. Hum. Genet. 2015, 18, 762–771. [Google Scholar] [CrossRef]
- de Candia, P.; Prattichizzo, F.; Garavelli, S.; De Rosa, V.; Galgani, M.; Di Rella, F.; Spagnuolo, M.I.; Colamatteo, A.; Fusco, C.; Micillo, T.; et al. Type 2 Diabetes: How Much of an Autoimmune Disease? Front. Endocrinol. 2019, 10, 451. [Google Scholar] [CrossRef]
- Li, S.; Zhang, M.; Tian, H.; Liu, Z.; Yin, X.; Xi, B. Preterm birth and risk of type 1 and type 2 diabetes: Systematic review and meta-analysis. Obes. Rev. 2014, 15, 804–811. [Google Scholar] [CrossRef]
- Ikegami, H.; Babaya, N.; Noso, S. β-Cell failure in diabetes: Common susceptibility and mechanisms shared between type 1 and type 2 diabetes. J. Diabetes Investig. 2021, 12, 1526–1539. [Google Scholar] [CrossRef]
- Jones, A.G.; McDonald, T.J.; Shields, B.M.; Hagopian, W.; Hattersley, A.T. Latent Autoimmune Diabetes of Adults (LADA) Is Likely to Represent a Mixed Population of Autoimmune (Type 1) and Nonautoimmune (Type 2) Diabetes. Diabetes Care 2021, 44, 1243–1251. [Google Scholar] [CrossRef] [PubMed]
- Thursby, E.; Juge, N. Introduction to the human gut microbiota. Biochem. J. 2017, 474, 1823–1836. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Pitocco, D.; DI Leo, M.; Tartaglione, L.; De Leva, F.; Petruzziello, C.; Saviano, A.; Pontecorvi, A.; Ojetti, V. The role of gut microbiota in mediating obesity and diabetes mellitus. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 1548–1562. [Google Scholar] [PubMed]
- Gurung, M.; Li, Z.; You, H.; Rodrigues, R.; Jump, D.B.; Morgun, A.; Shulzhenko, N. Role of gut microbiota in type 2 diabetes pathophysiology. EBioMedicine 2020, 51, 102590. [Google Scholar] [CrossRef] [PubMed]
- James, M.M.; Pal, N.; Sharma, P.; Kumawat, M.; Shubham, S.; Verma, V.; Tiwari, R.R.; Singh, B.; Nagpal, R.; Sarma, D.K.; et al. Role of butyrogenic Firmicutes in type-2 diabetes. J. Diabetes Metab. Disord. 2022, 21, 1873–1882. [Google Scholar] [CrossRef] [PubMed]
- Bamola, V.D.; Ghosh, A.; Kapardar, R.K.; Lal, B.; Cheema, S.; Sarma, P.; Chaudhry, R. Gut microbial diversity in health and disease: Experience of healthy Indian subjects, and colon carcinoma and inflammatory bowel disease patients. Microb. Ecol. Health Dis. 2017, 28, 1322447. [Google Scholar] [CrossRef]
- Tingirikari, J.M.R. Microbiota-accessible pectic poly- and oligosaccharides in gut health. Food Funct. 2018, 9, 5059–5073. [Google Scholar] [CrossRef]
- Cholan, P.M.; Han, A.; Woodie, B.R.; Watchon, M.; Kurz, A.R.; Laird, A.S.; Britton, W.J.; Ye, L.; Holmes, Z.C.; McCann, J.R.; et al. Conserved anti-inflammatory effects and sensing of butyrate in zebrafish. Gut Microbes 2020, 12, 1824563. [Google Scholar] [CrossRef]
- Zheng, P.; Li, Z.; Zhou, Z. Gut microbiome in type 1 diabetes: A comprehensive review. Diabetes/Metab. Res. Rev. 2018, 34, e3043. [Google Scholar] [CrossRef]
- Liu, L.; Zhang, J.; Cheng, Y.; Zhu, M.; Xiao, Z.; Ruan, G.; Wei, Y. Gut microbiota: A new target for T2DM prevention and treatment. Front. Endocrinol. 2022, 13, 958218. [Google Scholar] [CrossRef]
- Coppola, S.; Avagliano, C.; Calignano, A.; Canani, R.B. The Protective Role of Butyrate against Obesity and Obesity-Related Diseases. Molecules 2021, 26, 682. [Google Scholar] [CrossRef] [PubMed]
- Halim, M.; Halim, A. The effects of inflammation, aging and oxidative stress on the pathogenesis of diabetes mellitus (type 2 diabetes). Diabetes Metab. Syndr. Clin. Res. Rev. 2019, 13, 1165–1172. [Google Scholar] [CrossRef]
- Riedel, C.U.; Foata, F.; Philippe, D.; Adolfsson, O.; Eikmanns, B.J.; Blum, S. Anti-inflammatory effects of bifidobacteria by inhibition of LPS-induced NF-κB activation. World J. Gastroenterol. 2006, 12, 3729–3735. [Google Scholar] [CrossRef] [PubMed]
- Halawa, M.R.; El-Salam, M.A.; Mostafa, B.M.; Sallout, S.S. The Gut Microbiome, Lactobacillus acidophilus; Relation with Type 2 Diabetes Mellitus. Curr. Diabetes Rev. 2019, 15, 480–485. [Google Scholar] [CrossRef]
- Kijmanawat, A.; Panburana, P.; Reutrakul, S.; Tangshewinsirikul, C. Effects of probiotic supplements on insulin resistance in gestational diabetes mellitus: A double-blind randomized controlled trial. J. Diabetes Investig. 2018, 10, 163–170. [Google Scholar] [CrossRef]
- Rodrigues, V.F.; Elias-Oliveira, J.; Pereira, Í.S.; Pereira, J.A.; Barbosa, S.C.; Machado, M.S.G.; Carlos, D. Akkermansia muciniphila and Gut Immune System: A Good Friendship That Attenuates Inflammatory Bowel Disease, Obesity, and Diabetes. Front. Immunol. 2022, 13, 934695. [Google Scholar] [CrossRef]
- Zhang, J.; Ni, Y.; Qian, L.; Fang, Q.; Zheng, T.; Zhang, M.; Gao, Q.; Ni, J.; Hou, X.; Bao, Y.; et al. Decreased Abundance of Akkermansia muciniphila Leads to the Impairment of Insulin Secretion and Glucose Homeostasis in Lean Type 2 Diabetes. Adv. Sci. 2021, 8, e2100536. [Google Scholar] [CrossRef]
- Matsuoka, K.; Kanai, T. The gut microbiota and inflammatory bowel disease. Semin. Immunopathol. 2015, 37, 47–55. [Google Scholar] [CrossRef]
- Scheithauer, T.P.M.; Rampanelli, E.; Nieuwdorp, M.; Vallance, B.A.; Verchere, C.B.; Van Raalte, D.H.; Herrema, H. Gut Microbiota as a Trigger for Metabolic Inflammation in Obesity and Type 2 Diabetes. Front. Immunol. 2020, 11, 571731. [Google Scholar] [CrossRef]
- Soyucen, E.; Gulcan, A.; Aktuglu-Zeybek, A.C.; Onal, H.; KIYKIM, E.; Aydin, A. Differences in the gut microbiota of healthy children and those with type 1 diabetes. Pediatr. Int. 2014, 56, 336–343. [Google Scholar] [CrossRef]
- Liu, X.; Mao, B.; Gu, J.; Wu, J.; Cui, S.; Wang, G.; Zhao, J.; Zhang, H.; Chen, W. Blautia-a new functional genus with potential probiotic properties? Gut Microbes 2021, 13, 1875796. [Google Scholar] [CrossRef] [PubMed]
- Bojarczuk, A.; Skąpska, S.; Khaneghah, A.M.; Marszałek, K. Health benefits of resistant starch: A review of the literature. J. Funct. Foods 2022, 93, 105094. [Google Scholar] [CrossRef]
- Cheng, S.H.; Yang, T.Y.; Hsu, C.C.; Wei, Y.H.; Wu, C.C.; Tsai, Y.C. Lactobacillus paragasseri BBM171 Ameliorates Allergic Airway Inflammation Induced by Ovalbumin in Mice via Modulating the Th1/Th2 Balance. Microorganisms 2022, 10, 2041. [Google Scholar] [CrossRef] [PubMed]
- Gomes, A.C.; Bueno, A.A.; de Souza, R.G.M.; Mota, J.F. Gut microbiota, probiotics and diabetes. Nutr. J. 2014, 13, 60. [Google Scholar] [CrossRef]
- Rinninella, E.; Raoul, P.; Cintoni, M.; Franceschi, F.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, M.C. What Is the Healthy Gut Microbiota Composition? A Changing Ecosystem across Age, Environment, Diet, and Diseases. Microorganisms 2019, 7, 14. [Google Scholar] [CrossRef]
- Stojanov, S.; Berlec, A.; Štrukelj, B. The Influence of Probiotics on the Firmicutes/Bacteroidetes Ratio in the Treatment of Obesity and Inflammatory Bowel disease. Microorganisms 2020, 8, 1715. [Google Scholar] [CrossRef]
- Larsen, N.; Vogensen, F.K.; Van Den Berg, F.W.J.; Nielsen, D.S.; Andreasen, A.S.; Pedersen, B.K.; Al-Soud, W.A.; Sørensen, S.J.; Hansen, L.H.; Jakobsen, M. Gut Microbiota in Human Adults with Type 2 Diabetes Differs from Non-Diabetic Adults. PLoS ONE 2010, 5, e9085. [Google Scholar] [CrossRef]
- Hung, W.-C.; Tsai, H.-J.; Chang, C.-C.; Chiu, Y.-W.; Hwang, S.-J.; Kuo, M.-C.; Chen, S.-C.; Dai, C.-Y.; Tsai, Y.-C. The Association of Targeted Gut Microbiota with Body Composition in Type 2 Diabetes Mellitus. Int. J. Med Sci. 2021, 18, 511–519. [Google Scholar] [CrossRef]
- Lontchi-Yimagou, E.; Dasgupta, R.; Anoop, S.; Kehlenbrink, S.; Koppaka, S.; Goyal, A.; Venkatesan, P.; Livingstone, R.; Ye, K.; Chapla, A.; et al. An Atypical Form of Diabetes Among Individuals with Low BMI. Diabetes Care 2022, 45, 1428–1437. [Google Scholar] [CrossRef]
- George, A.M.; Jacob, A.G.; Fogelfeld, L. Lean diabetes mellitus: An emerging entity in the era of obesity. World J. Diabetes 2015, 6, 613–620. [Google Scholar] [CrossRef]
- Koh, H.-C.E.; Cao, C.; Mittendorfer, B. Insulin Clearance in Obesity and Type 2 Diabetes. Int. J. Mol. Sci. 2022, 23, 596. [Google Scholar] [CrossRef] [PubMed]
- Magne, F.; Gotteland, M.; Gauthier, L.; Zazueta, A.; Pesoa, S.; Navarrete, P.; Balamurugan, R. The Firmicutes/Bacteroidetes Ratio: A Relevant Marker of Gut Dysbiosis in Obese Patients? Nutrients 2020, 12, 1474. [Google Scholar] [CrossRef] [PubMed]
- Mokhtari, P.; Metos, J.; Babu, P.V.A. Impact of type 1 diabetes on the composition and functional potential of gut microbiome in children and adolescents: Possible mechanisms, current knowledge, and challenges. Gut Microbes 2021, 13, 1926841. [Google Scholar] [CrossRef] [PubMed]
- Knip, M.; Honkanen, J. Modulation of Type 1 Diabetes Risk by the Intestinal Microbiome. Curr. Diabetes Rep. 2017, 17, 105. [Google Scholar] [CrossRef]
- Zhou, H.; Zhao, X.; Sun, L.; Liu, Y.; Lv, Y.; Gang, X.; Wang, G. Gut Microbiota Profile in Patients with Type 1 Diabetes Based on 16S rRNA Gene Sequencing: A Systematic Review. Dis. Markers 2020, 2020, 3936247. [Google Scholar] [CrossRef]
- Siljander, H.; Honkanen, J.; Knip, M. Microbiome and type 1 diabetes. Ebiomedicine 2019, 46, 512–521. [Google Scholar] [CrossRef]
- Moffa, S.; Mezza, T.; Cefalo, C.M.A.; Cinti, F.; Impronta, F.; Sorice, G.P.; Santoro, A.; Di Giuseppe, G.; Pontecorvi, A.; Giaccari, A. The Interplay between Immune System and Microbiota in Diabetes. Mediat. Inflamm. 2019, 2019, 9367404. [Google Scholar] [CrossRef]
- Bessac, A.; Cani, P.D.; Meunier, E.; Dietrich, G.; Knauf, C. Inflammation and Gut-Brain Axis During Type 2 Diabetes: Focus on the Crosstalk Between Intestinal Immune Cells and Enteric Nervous System. Front. Neurosci. 2018, 12, 725. [Google Scholar] [CrossRef]
- Liu, Y.; Lou, X. Type 2 diabetes mellitus-related environmental factors and the gut microbiota: Emerging evidence and challenges. Clinics 2020, 75, e1277. [Google Scholar] [CrossRef]
- Feng, Y.; Wang, Y.; Wang, P.; Huang, Y.; Wang, F. Short-Chain Fatty Acids Manifest Stimulative and Protective Effects on Intestinal Barrier Function Through the Inhibition of NLRP3 Inflammasome and Autophagy. Cell Physiol. Biochem. 2018, 49, 190–205. [Google Scholar] [CrossRef]
- Ho, J.; Nicolucci, A.C.; Virtanen, H.; Schick, A.; Meddings, J.; Reimer, R.A.; Huang, C. Effect of Prebiotic on Microbiota, Intestinal Permeability, and Glycemic Control in Children with Type 1 Diabetes. J. Clin. Endocrinol. Metab. 2019, 104, 4427–4440. [Google Scholar] [CrossRef] [PubMed]
- Thaiss, C.A.; Levy, M.; Grosheva, I.; Zheng, D.; Soffer, E.; Blacher, E.; Braverman, S.; Tengeler, A.C.; Barak, O.; Elazar, M.; et al. Hyperglycemia drives intestinal barrier dysfunction and risk for enteric infection. Science 2018, 359, 1376–1383. [Google Scholar] [CrossRef] [PubMed]
- Chiarelli, F.; Giannini, C.; Primavera, M. Prediction and prevention of type 1 diabetes in children. Clin. Pediatr. Endocrinol. 2019, 28, 43–57. [Google Scholar] [CrossRef] [PubMed]
- Huda, M.N.; Kim, M.; Bennett, B.J. Modulating the Microbiota as a Therapeutic Intervention for Type 2 Diabetes. Front. Endocrinol. 2021, 12, 632335. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Wang, K.; Wang, X.; Pang, Y.; Jiang, C. The role of the gut microbiome and its metabolites in metabolic diseases. Protein Cell 2021, 12, 360–373. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.K.; Chang, H.-W.; Yan, D.; Lee, K.M.; Ucmak, D.; Wong, K.; Abrouk, M.; Farahnik, B.; Nakamura, M.; Zhu, T.H.; et al. Influence of diet on the gut microbiome and implications for human health. J. Transl. Med. 2017, 15, 73. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, J.; He, T.; Becker, S.; Zhang, G.; Li, D.; Ma, X. Butyrate: A Double-Edged Sword for Health? Adv. Nutr. 2018, 9, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Thomas, S.P.; Denu, J.M. Short-chain fatty acids activate acetyltransferase p300. eLife 2021, 10, e72171. [Google Scholar] [CrossRef]
- Pedersen, S.S.; Prause, M.; Williams, K.; Barrès, R.; Billestrup, N. Butyrate inhibits IL-1β-induced inflammatory gene expression by suppression of NF-κB activity in pancreatic beta cells. J. Biol. Chem. 2022, 298, 102312. [Google Scholar] [CrossRef]
- Blaak, E.E.; Canfora, E.E.; Theis, S.; Frost, G.; Groen, A.K.; Mithieux, G.; Nauta, A.; Scott, K.; Stahl, B.; Van Harsselaar, J.; et al. Short chain fatty acids in human gut and metabolic health. Benef. Microbes 2020, 11, 411–455. [Google Scholar] [CrossRef]
- Harbison, J.E.; Roth-Schulze, A.J.; Giles, L.; Tran, C.D.; Ngui, K.M.; Penno, M.; Thomson, R.L.; Wentworth, J.M.; Colman, P.G.; Craig, M.E.; et al. Gut microbiome dysbiosis and increased intestinal permeability in children with islet autoimmunity and type 1 diabetes: A prospective cohort study. Pediatr. Diabetes 2019, 20, 574–583. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, S.-X.; Cai, Y.; Xie, K.-L.; Zhang, W.-L.; Zheng, F. Effects of combined aerobic and resistance training on the glycolipid metabolism and inflammation levels in type 2 diabetes mellitus. J. Phys. Ther. Sci. 2015, 27, 2365–2371. [Google Scholar] [CrossRef] [PubMed]
- Pasini, E.; Corsetti, G.; Assanelli, D.; Testa, C.; Romano, C.; Dioguardi, F.S.; Aquilani, R. Effects of chronic exercise on gut microbiota and intestinal barrier in human with type 2 diabetes. Minerva Med. 2019, 110, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Monda, V.; Villano, I.; Messina, A.; Valenzano, A.; Esposito, T.; Moscatelli, F.; Viggiano, A.; Cibelli, G.; Chieffi, S.; Monda, M.; et al. Exercise Modifies the Gut Microbiota with Positive Health Effects. Oxidative Med. Cell. Longev. 2017, 2017, 3831972. [Google Scholar] [CrossRef] [PubMed]
- Yun, E.-J.; Imdad, S.; Jang, J.; Park, J.; So, B.; Kim, J.-H.; Kang, C. Diet Is a Stronger Covariate than Exercise in Determining Gut Microbial Richness and Diversity. Nutrients 2022, 14, 2507. [Google Scholar] [CrossRef] [PubMed]
- LaMoia, T.E.; Shulman, G.I. Cellular and Molecular Mechanisms of Metformin Action. Endocr. Rev. 2020, 42, 77–96. [Google Scholar] [CrossRef] [PubMed]
- Vallianou, N.G.; Stratigou, T.; Tsagarakis, S. Metformin and gut microbiota: Their interactions and their impact on diabetes. Hormones (Athens) 2019, 18, 141–144. [Google Scholar] [CrossRef]
- Sun, L.; Xie, C.; Wang, G.; Wu, Y.; Wu, Q.; Wang, X.; Liu, J.; Deng, Y.; Xia, J.; Chen, B.; et al. Gut microbiota and intestinal FXR mediate the clinical benefits of metformin. Nat. Med. 2018, 24, 1919–1929. [Google Scholar] [CrossRef]
- Forslund, K.; Hildebrand, F.; Nielsen, T.; Falony, G.; Le Chatelier, E.; Sunagawa, S.; Prifti, E.; Vieira-Silva, S.; Gudmundsdottir, V.; Krogh Pedersen, H.; et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 2015, 528, 262–266. [Google Scholar] [CrossRef]
- Lee, C.; Chae, S.; Jo, S.; Jerng, U.; Bae, S. The Relationship between the Gut Microbiome and Metformin as a Key for Treating Type 2 Diabetes Mellitus. Int. J. Mol. Sci. 2021, 22, 3566. [Google Scholar] [CrossRef]
- Craciun, C.I.; Neag, M.A.; Catinean, A.; Mitre, A.O.; Rusu, A.; Bala, C.; Roman, G.; Buzoianu, A.D.; Muntean, D.M.; Craciun, A.E. The Relationships between Gut Microbiota and Diabetes Mellitus, and Treatments for Diabetes Mellitus. Biomedicines 2022, 10, 308. [Google Scholar] [CrossRef] [PubMed]
- Salles, B.I.M.; Cioffi, D.; Ferreira, S.R.G. Probiotics supplementation and insulin resistance: A systematic review. Diabetol. Metab. Syndr. 2020, 12, 98. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Lu, Y.; Yan, Y.; Tian, S.; Zheng, D.; Leng, D.; Wang, C.; Jiao, J.; Wang, Z.; Bai, Y. Promising Treatment for Type 2 Diabetes: Fecal Microbiota Transplantation Reverses Insulin Resistance and Impaired Islets. Front. Cell. Infect. Microbiol. 2020, 9, 455. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; De Vos, W.M. Next-Generation Beneficial Microbes: The Case of Akkermansia muciniphila. Front. Microbiol. 2017, 8, 1765. [Google Scholar] [CrossRef]
- Agus, A.; Clément, K.; Sokol, H. Gut microbiota-derived metabolites as central regulators in metabolic disorders. Gut 2020, 70, 1174–1182. [Google Scholar] [CrossRef]
- Michael, H.; Amimo, J.O.; Rajashekara, G.; Saif, L.J.; Vlasova, A.N. Mechanisms of Kwashiorkor-Associated Immune Suppression: Insights from Human, Mouse, and Pig Studies. Front. Immunol. 2022, 13, 826268. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).