Circulating miRNA Expression Profiles and Machine Learning Models in Association with Response to Irinotecan-Based Treatment in Metastatic Colorectal Cancer
Abstract
1. Introduction
2. Results
2.1. miRNA Differential Expression
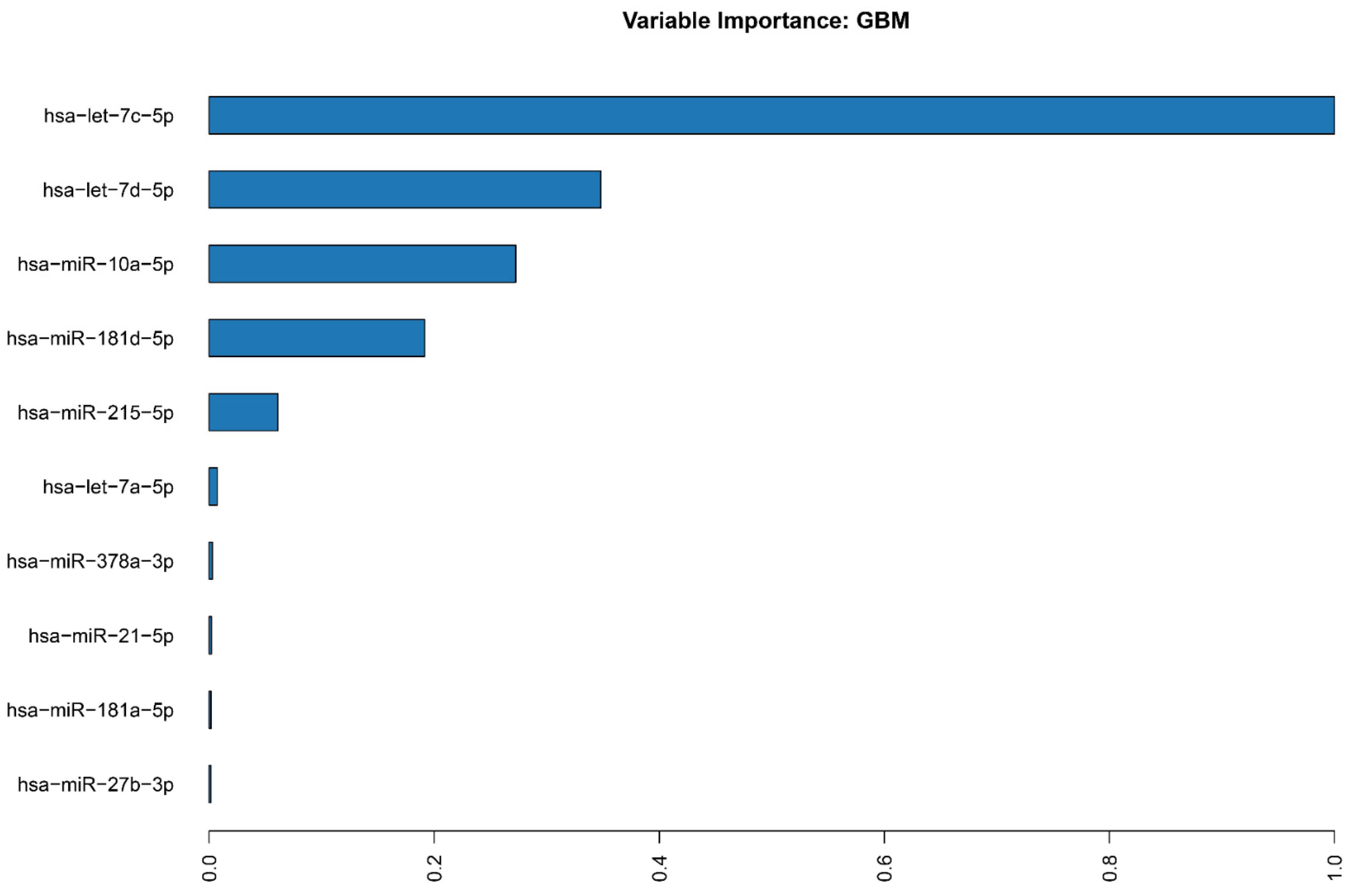
2.2. miRNA Machine Learning Model
3. Discussion
4. Materials and Methods
4.1. Patients and Blood Samples
4.2. MiRNA Expression
4.3. Machine Learning Model
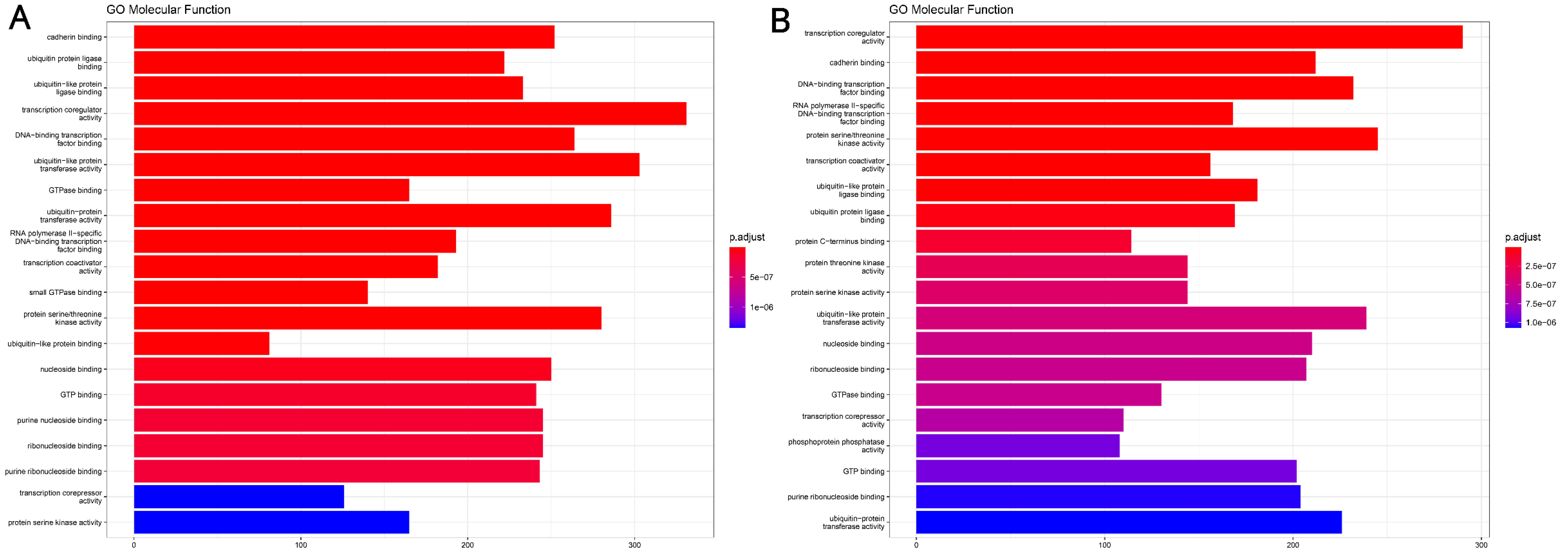
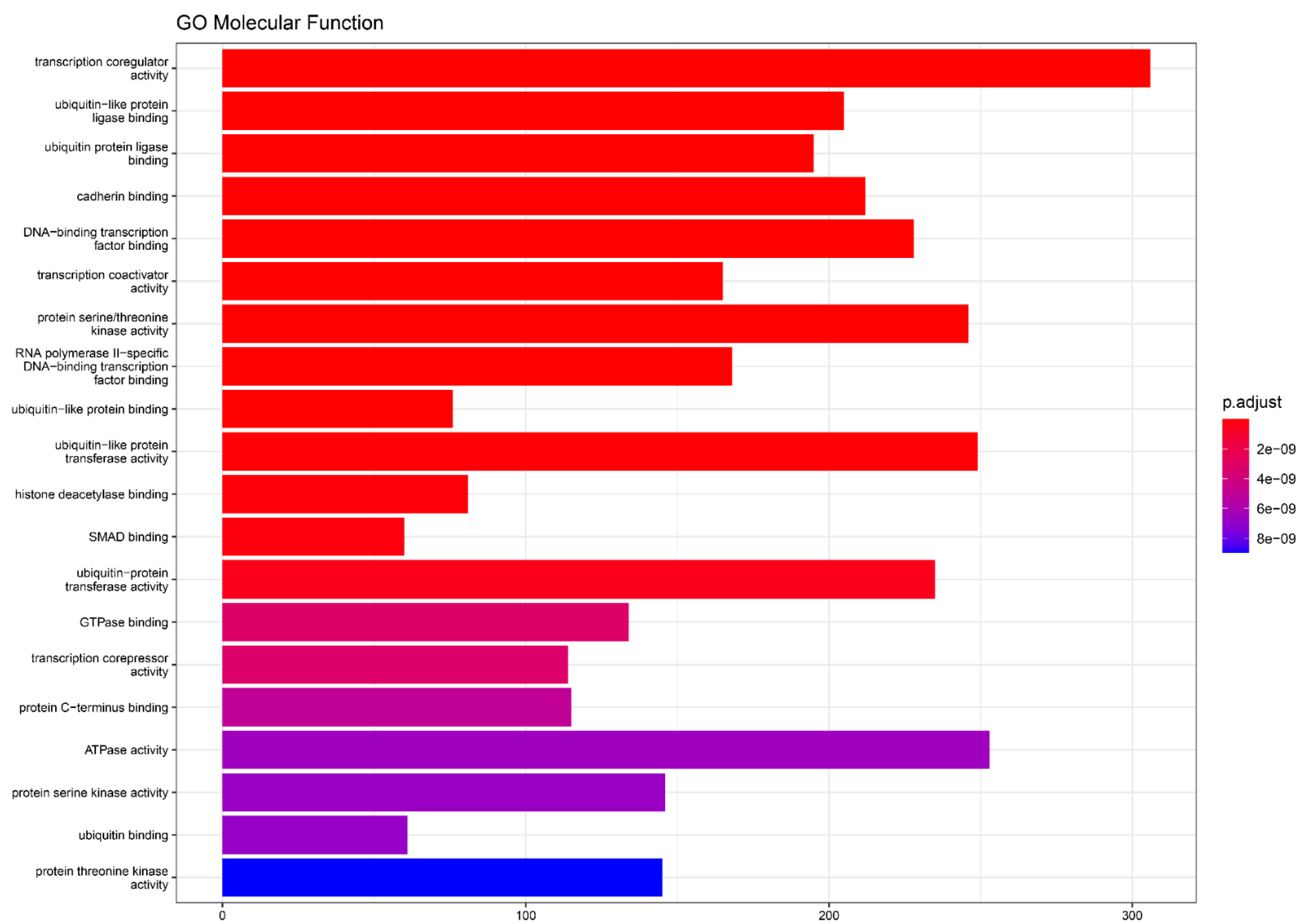
4.4. miRNA Target Genes and Molecular Function Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ismail, I.; Chan, H.K.; Aiman, S.S.; Muhammad Radzi, A.H. A 10-year registry-based incidence, mortality, and survival analysis of colorectal cancer in Northern Malaysia. J. Cancer Res. Ther. 2022, 18, 931–938. [Google Scholar] [CrossRef] [PubMed]
- Kopetz, S.; Chang, G.J.; Overman, M.J.; Eng, C.; Sargent, D.J.; Larson, D.W.; Grothey, A.; Vauthey, J.N.; Nagorney, D.M.; McWilliams, R.R. Improved survival in metastatic colorectal cancer is associated with adoption of hepatic resection and improved chemotherapy. J. Clin. Oncol. 2009, 27, 3677–3683. [Google Scholar] [CrossRef] [PubMed]
- Van Cutsem, E.; Cervantes, A.; Adam, R.; Sobrero, A.; Van Krieken, J.H.; Aderka, D.; Aranda Aguilar, E.; Bardelli, A.; Benson, A.; Bodoky, G.; et al. ESMO consensus guidelines for the management of patients with metastatic colorectal cancer. Ann. Oncol. 2016, 27, 1386–1422. [Google Scholar] [CrossRef] [PubMed]
- Dansero, L.; Ricceri, F.; De Marco, L.; Fiano, V.; Nesi, G.; Padroni, L.; Milani, L.; Caini, S.; Masala, G.; Agnoli, C.; et al. Investigating the Role of Circulating miRNAs as Biomarkers in Colorectal Cancer: An Epidemiological Systematic Review. Biomedicines 2022, 10, 2224. [Google Scholar] [CrossRef] [PubMed]
- Baharudin, R.; Rus Bakarurraini, N.Q.; Ismail, I.; Lee, L.H.; Ab Mutalib, N.S. MicroRNA Methylome Signature and Their Functional Roles in Colorectal Cancer Diagnosis, Prognosis, and Chemoresistance. Int. J. Mol. Sci. 2022, 23, 7281. [Google Scholar] [CrossRef]
- Wei, S.; Hu, W.; Feng, J.; Geng, Y. Promotion or remission: A role of noncoding RNAs in colorectal cancer resistance to anti-EGFR therapy. Cell Commun. Signal. 2022, 20, 150. [Google Scholar] [CrossRef]
- Wei, L.; Wang, X.; Lv, L.; Zheng, Y.; Zhang, N.; Yang, M. The emerging role of noncoding RNAs in colorectal cancer chemoresistance. Cell Oncol. 2019, 42, 757–768. [Google Scholar] [CrossRef]
- Pouya, F.D.; Gazouli, M.; Rasmi, Y.; Lampropoulou, D.I.; Nemati, M. MicroRNAs and drug resistance in colorectal cancer with special focus on 5-fluorouracil. Mol. Biol. Rep. 2022, 49, 5165–5178. [Google Scholar] [CrossRef]
- Ozawa, S.; Miura, T.; Terashima, J.; Habano, W. Cellular irinotecan resistance in colorectal cancer and overcoming irinotecan refractoriness through various combination trials including DNA methyltransferase inhibitors: A review. Cancer Drug Resist. 2021, 4, 946–964. [Google Scholar] [CrossRef]
- Kannathasan, T.; Kuo, W.W.; Chen, M.C.; Viswanadha, V.P.; Shen, C.Y.; Tu, C.C.; Yeh, Y.L.; Bharath, M.; Shibu, M.A.; Huang, C.Y. Chemoresistance-Associated Silencing of miR-4454 Promotes Colorectal Cancer Aggression through the GNL3L and NF-κB Pathway. Cancers 2020, 12, 1231. [Google Scholar] [CrossRef]
- Poorebrahim, M.; Sadeghi, S.; Ghanbarian, M.; Kalhor, H.; Mehrtash, A.; Teimoori-Toolabi, L. Identification of candidate genes and miRNAs for sensitizing resistant colorectal cancer cells to oxaliplatin and irinotecan. Cancer Chemother. Pharmacol. 2020, 85, 153–171. [Google Scholar] [CrossRef] [PubMed]
- Larrañaga, P.; Calvo, B.; Santana, R.; Bielza, C.; Galdiano, J.; Inza, I.; Lozano, J.A.; Armañanzas, R.; Santafé, G.; Pérez, A.; et al. Machine learning in bioinformatics. Brief. Bioinform. 2006, 7, 86–112. [Google Scholar] [CrossRef] [PubMed]
- Nagy, M.; Radakovich, N.; Nazha, A. Machine learning in oncology: What should clinicians know? JCO Clin. Cancer. Inform. 2020, 4, 799–810. [Google Scholar] [CrossRef]
- Bzdok, D.; Altman, N.; Krzywinski, M. Statistics versus machine learning. Nat. Meth. 2018, 15, 233–234. [Google Scholar] [CrossRef]
- Abbas, M.; El-Manzalawy, Y. Machine learning based refined differential gene expression analysis of pediatric sepsis. BMC Med. Genom. 2020, 13, 122. [Google Scholar] [CrossRef]
- Adam, G.; Rampasek, L.; Safikhani, Z.; Smirnov, P.; Haibe-Kains, B.; Goldenberg, A. Machine learning approaches to drug response prediction: Challenges and recent progress. NPJ Precis. Oncol. 2020, 4, 19. [Google Scholar] [CrossRef] [PubMed]
- Cuocolo, R.; Caruso, M.; Perillo, T.; Ugga, L.; Petretta, M. Machine Learning in oncology: A clinical appraisal. Cancer Lett. 2020, 481, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Tanoli, Z.; Vaha-Koskela, M.; Aittokallio, T. Artificial intelligence, machine learning, and drug repurposing in cancer. Expert Opin. Drug Discov. 2021, 16, 977–989. [Google Scholar] [CrossRef]
- Rauschert, S.; Raubenheimer, K.; Melton, P.E.; Huang, R.C. Machine learning and clinical epigenetics: A review of challenges for diagnosis and classification. Clin. Epigenet. 2020, 12, 51. [Google Scholar] [CrossRef]
- Rafique, R.; Islam, S.M.R.; Kazi, J.U. Machine learning in the prediction of cancer therapy. Comput. Struct. Biotechnol. J. 2021, 19, 4003–4017. [Google Scholar] [CrossRef]
- Rupaimoole, R.; Slack, F.J. MicroRNA therapeutics: Towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 2017, 16, 203–222. [Google Scholar] [CrossRef]
- Anastasiadou, E.; Jacob, L.S.; Slack, F.J. Non-coding RNA networks in cancer. Nat. Rev. Cancer. 2018, 18, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Sevignani, C.; Dumitru, C.D.; Hyslop, T.; Noch, E.; Yendamuri, S.; Shimizu, M.; Rattan, S.; Bullrich, F.; Negrini, M.; et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl. Acad. Sci. USA 2004, 101, 2999–3004. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhang, X.; Ren, X.P.; Chen, J.; Liu, H.; Yang, J.; Medvedovic, M.; Hu, Z.; Fan, G.C. MicroRNA-494 targeting both pro-apoptotic and anti-apoptotic proteins protects against ischemia/reperfusion-induced cardiac injury. Circulation 2010, 122, 1308–1318. [Google Scholar] [CrossRef]
- Carroll, A.P.; Tooney, P.A.; Cairns, M.J. Context-specific microRNA function in developmental complexity. J. Mol. Cell Biol. 2013, 5, 73–84. [Google Scholar] [CrossRef]
- Dhawan, A.; Scott, J.G.; Harris, A.L.; Buffa, F.M. Pan-cancer characterisation of microRNA across cancer hallmarks reveals microRNA-mediated downregulation of tumour suppressors. Nat. Commun. 2018, 9, 5228. [Google Scholar] [CrossRef] [PubMed]
- Moradi, S.; Kamal, A.; Aboulkheyr, E.; Farhadi, F.; Ebrahimi, M.; Chitsaz, H.; Sharifi-Zarchi, A.; Baharvand, H. Pan-cancer analysis of microRNA expression profiles highlights microRNAs enriched in normal body cells as effective suppressors of multiple tumor types: A study based on TCGA database. PLoS ONE. 2022, 17, e0267291. [Google Scholar] [CrossRef]
- Sun, X.; Chen, J.; Chen, X.; Gao, Q.; Chen, W.; Zou, X.; Zhang, F.; Gao, S.; Qiu, S.; Yue, X.; et al. A Systematic Review of Clinical Validated and Potential miRNA Markers Related to the Efficacy of Fluoropyrimidine Drugs. Dis Markers 2022, 2022, 1360954. [Google Scholar] [CrossRef]
- Tsai, H.L.; Tsai, Y.C.; Chen, Y.C.; Huang, C.W.; Chen, P.J.; Li, C.C.; Su, W.C.; Chang, T.K.; Yeh, Y.S.; Yin, T.C.; et al. MicroRNA-148a induces apoptosis and prevents angiogenesis with bevacizumab in colon cancer through direct inhibition of ROCK1/c-Met via HIF-1α under hypoxia. Aging 2022, 14, 6668–6688. [Google Scholar] [CrossRef]
- Meng, T.; Lan, Z.; Zhao, X.; Niu, L.; Chen, C.; Zhang, W. Comprehensive bioinformatics analysis of functional molecules in colorectal cancer. J. Gastrointest. Oncol. 2022, 13, 231–245. [Google Scholar] [CrossRef]
- Kutilin, D.S. Regulation of Gene Expression of Cancer/Testis Antigens in Colorectal Cancer Patients]. Mol. Biol. 2020, 54, 580–595. [Google Scholar] [CrossRef]
- Pellatt, D.F.; Stevens, J.R.; Wolff, R.K.; Mullany, L.E.; Herrick, J.S.; Samowitz, W.; Slattery, M.L. Expression Profiles of miRNA Subsets Distinguish Human Colorectal Carcinoma and Normal Colonic Mucosa. Clin. Transl. Gastroenterol. 2016, 7, e152. [Google Scholar] [CrossRef] [PubMed]
- Eneh, S.; Heikkinen, S.; Hartikainen, J.M.; Kuopio, T.; Mecklin, J.P.; Kosma, V.M.; Mannermaa, A. MicroRNAs Associated With Biological Pathways of Left- and Right-sided Colorectal Cancer. Anticancer Res. 2020, 40, 3713–3722. [Google Scholar] [CrossRef]
- Yang, L.; Li, L.; Ma, J.; Yang, S.; Zou, C.; Yu, X. miRNA and mRNA Integration Network Construction Reveals Novel Key Regulators in Left-Sided and Right-Sided Colon Adenocarcinoma. BioMed Res. Int. 2019, 2019, 7149296. [Google Scholar] [CrossRef]
- Liu, Y.; Hu, X.; Hu, L.; Xu, C.; Liang, X. Let-7i-5p enhances cell proliferation, migration and invasion of ccRCC by targeting HABP4. BMC Urol. 2021, 21, 49. [Google Scholar] [CrossRef] [PubMed]
- Mullany, L.E.; Slattery, M.L. The functional role of miRNAs in colorectal cancer: Insights from a large population-based study. Cancer Biol. Med. 2019, 16, 211–219. [Google Scholar] [CrossRef]
- Pan, S.; Deng, Y.; Fu, J.; Zhang, Y.; Zhang, Z.; Qin, X. N6 methyladenosine upregulates miR 181d 5p in exosomes derived from cancer associated fibroblasts to inhibit 5 FU sensitivity by targeting NCALD in colorectal cancer. Int. J. Oncol. 2022, 60, 14. [Google Scholar] [CrossRef]
- Chen, S.; Liu, Y.; Wang, Y.; Xue, Z. LncRNA CCAT1 Promotes Colorectal Cancer Tumorigenesis Via A miR-181b-5p/TUSC3 Axis. Onco Targets Ther. 2019, 12, 9215–9225. [Google Scholar] [CrossRef]
- Cheng, Y.; Yu, C.; Li, W.; He, Y.; Bao, Y. Matrine Inhibits Proliferation, Invasion, and Migration and Induces Apoptosis of Colorectal Cancer Cells Via miR-10b/PTEN Pathway. Cancer Biother. Radiopharm. 2020, 37, 871–881. [Google Scholar] [CrossRef]
- Tie, Y.; Chen, C.; Yang, Y.; Qian, Z.; Yuan, H.; Wang, H.; Tang, H.; Peng, Y.; Du, X.; Liu, B. Upregulation of let-7f-5p promotes chemotherapeutic resistance in colorectal cancer by directly repressing several pro-apoptotic proteins. Oncol. Lett. 2018, 15, 8695–8702. [Google Scholar] [CrossRef]
- Zhang, H.; Zhu, M.; Shan, X.; Zhou, X.; Wang, T.; Zhang, J.; Tao, J.; Cheng, W.; Chen, G.; Li, J.; et al. A panel of seven-miRNA signature in plasma as potential biomarker for colorectal cancer diagnosis. Gene 2019, 687, 246–254. [Google Scholar] [CrossRef] [PubMed]
- Lukosevicius, R.; Juzenas, S.; Salteniene, V.; Kulokiene, U.; Arstikyte, J.; Hemmrich-Stanisak, G.; Franke, A.; Link, A.; Ruzgys, P.; Satkauskas, S.; et al. miRNome Profiling and Functional Analysis Reveal Involvement of hsa-miR-1246 in Colon Adenoma-Carcinoma Transition by Targeting AXIN2 and CFTR. Int. J. Mol. Sci. 2022, 23, 2107. [Google Scholar] [CrossRef] [PubMed]
- Romero-Lorca, A.; Novillo, A.; Gaibar, M.; Gilsanz, M.F.; Galán, M.; Beltrán, L.; Antón, B.; Malón, D.; Moreno, A.; Fernández-Santander, A. miR-7, miR-10a and miR-143 Expression May Predict Response to Bevacizumab Plus Chemotherapy in Patients with Metastatic Colorectal Cancer. Pharmgenom. Pers. Med. 2021, 14, 1263–1273. [Google Scholar] [CrossRef] [PubMed]
- Bahnassy, A.A.; Salem, S.E.; El-Sayed, M.; Khorshid, O.; Abdellateif, M.S.; Youssef, A.S.; Mohanad, M.; Hussein, M.; Zekri, A.N.; Ali, N.M. MiRNAs as molecular biomarkers in stage II egyptian colorectal cancer patients. Exp. Mol. Pathol. 2018, 105, 260–267. [Google Scholar] [CrossRef]
- Nersisyan, S.; Galatenko, A.; Chekova, M.; Tonevitsky, A. Hypoxia-Induced miR-148a Downregulation Contributes to Poor Survival in Colorectal Cancer. Front. Genet. 2021, 12, 662468. [Google Scholar] [CrossRef]
- Despotovic, J.; Dragicevic, S.; Nikolic, A. Effects of Chemotherapy for Metastatic Colorectal Cancer on the TGF-β Signaling and Related miRNAs hsa-miR-17-5p, hsa-miR-21-5p and hsa-miR-93-5p. Cell Biochem. Biophys. 2021, 79, 757–767. [Google Scholar] [CrossRef]
- Li, X.; Han, J.; Zhu, H.; Peng, L.; Chen, Z. miR 181b 5p mediates TGF β1-induced epithelial-to-mesenchymal transition in non-small cell lung cancer stem-like cells derived from lung adenocarcinoma A549 cells. Int. J. Oncol. 2017, 51, 158–168. [Google Scholar] [CrossRef]
- Liu, Y.; Uzair-Ur, R.; Guo, Y.; Liang, H.; Cheng, R.; Yang, F.; Hong, Y.; Zhao, C.; Liu, M.; Yu, M.; et al. miR-181b functions as an oncomiR in colorectal cancer by targeting PDCD4. Protein Cell 2016, 7, 722–734. [Google Scholar]
- Zhou, Q.; Zheng, X.; Chen, L.; Xu, B.; Yang, X.; Jiang, J.; Wu, C. Smad2/3/4 Pathway Contributes to TGF-β-Induced MiRNA-181b Expression to Promote Gastric Cancer Metastasis by Targeting Timp3. Cell Physiol. Biochem. 2016, 39, 453–466. [Google Scholar] [CrossRef]
- Qin, Y.; Zheng, Y.; Huang, C.; Li, Y.; Gu, M.; Wu, Q. Downregulation of miR-181b-5p Inhibits the Viability, Migration, and Glycolysis of Gallbladder Cancer by Upregulating PDHX Under Hypoxia. Front. Oncol. 2021, 11, 683725. [Google Scholar] [CrossRef]
- Yan, W.; Song, L.; Wang, H.; Yang, W.; Hu, L.; Yang, Y. Extracellular vesicles carrying miRNA-181b-5p affects the malignant progression of acute lymphoblastic leukemia. J. Transl. Med. 2021, 19, 511. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Sun, Y.; Zhang, Y.; Han, S. Downregulation of miR 181b inhibits human colon cancer cell proliferation by targeting CYLD and inhibiting the NF κB signaling pathway. Int. J. Mol. Med. 2020, 46, 1755–1764. [Google Scholar] [CrossRef] [PubMed]
- Rezaei, T.; Amini, M.; Hashemi, Z.S.; Mansoori, B.; Rezaei, S.; Karami, H.; Mosafer, J.; Mokhtarzadeh, A.; Baradaran, B. microRNA-181 serves as a dual-role regulator in the development of human cancers. Free Radic. Biol. Med. 2020, 152, 432–454. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Yu, J.; Zhao, C.; Ren, H.; Yuan, Z.; Zhang, B.; Zhuang, J.; Wang, J.; Feng, B. MiR-181b-5p modulates chemosensitivity of glioma cells to temozolomide by targeting Bcl-2. Biomed. Pharmacother. 2019, 109, 2192–2202. [Google Scholar] [CrossRef]
- Cai, B.; An, Y.; Lv, N.; Chen, J.; Tu, M.; Sun, J.; Wu, P.; Wei, J.; Jiang, K.; Miao, Y. miRNA-181b increases the sensitivity of pancreatic ductal adenocarcinoma cells to gemcitabine in vitro and in nude mice by targeting BCL-2. Oncol. Rep. 2013, 29, 1769–1776. [Google Scholar] [CrossRef]
- Hosseini, F.; Shanehbandi, D.; Soleimanpour, J.; Yousefi, B.; Alemi, F. Melatonin Increases the Sensitivity of Osteosarcoma Cells to Chemotherapy Drug Cisplatin. Drug Res. 2022, 72, 312–318. [Google Scholar] [CrossRef] [PubMed]
- Schlick, K.; Hohla, F.; Hamacher, F.; Hackl, H.; Hufnagl, C.; Markus, S.; Magnes, T.; Gampenrieder, S.P.; Melchardt, T.; Stättner, S.; et al. Overcoming negative predictions of microRNA expressions to gemcitabine response with FOLFIRINOX in advanced pancreatic cancer patients. Future Sci. OA 2020, 7, FSO644. [Google Scholar] [CrossRef]
- Ouyang, M.; Li, Y.; Ye, S.; Ma, J.; Lu, L.; Lv, W.; Chang, G.; Li, X.; Li, Q.; Wang, S.; et al. MicroRNA profiling implies new markers of chemoresistance of triple-negative breast cancer. PLoS ONE. 2014, 9, e96228. [Google Scholar] [CrossRef]
- Han, P.; Li, J.W.; Zhang, B.M.; Lv, J.C.; Li, Y.M.; Gu, X.Y.; Yu, Z.W.; Jia, Y.H.; Bai, X.F.; Li, L.; et al. The lncRNA CRNDE promotes colorectal cancer cell proliferation and chemoresistance via miR-181a-5p-mediated regulation of Wnt/β-catenin signaling. Mol. Cancer 2017, 16, 9. [Google Scholar] [CrossRef]
- Pop-Bica, C.; Pintea, S.; Cojocneanu-Petric, R.; Del Sal, G.; Piazza, S.; Wu, Z.H.; Alencar, A.J.; Lossos, I.S.; Berindan-Neagoe, I.; Calin, G.A. MiR-181 family-specific behavior in different cancers: A meta-analysis view. Cancer Metastasis Rev. 2018, 37, 17–32. [Google Scholar] [CrossRef]
- Ren, Z.; Wang, Z.; Gu, D.; Ma, H.; Zhu, Y.; Cai, M.; Zhang, J. Genome Instability and Long Noncoding RNA Reveal Biomarkers for Immunotherapy and Prognosis and Novel Competing Endogenous RNA Mechanism in Colon Adenocarcinoma. Front. Cell Dev. Biol. 2021, 9, 740455. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, Y.; Zhu, H.; Sun, X.; Wang, X.; Wu, P.; Xu, X. Overexpression of miR-301a-3p promotes colorectal cancer cell proliferation and metastasis by targeting deleted in liver cancer-1 and runt-related transcription factor 3. J. Cell Biochem. 2019, 120, 6078–6089. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Zhang, K.; Luo, G.; Cen, G.; Cao, J.; Huang, K.; Qiu, Z. Downregulation of miR-301a-3p sensitizes pancreatic cancer cells to gemcitabine treatment via PTEN. Am. J. Transl. Res. 2017, 9, 1886–1895. [Google Scholar]
- Hu, J.L.; Wang, W.; Lan, X.L.; Zeng, Z.C.; Liang, Y.S.; Yan, Y.R.; Song, F.Y.; Wang, F.F.; Zhu, X.H.; Liao, W.J.; et al. CAFs secreted exosomes promote metastasis and chemotherapy resistance by enhancing cell stemness and epithelial-mesenchymal transition in colorectal cancer. Mol. Cancer 2019, 18, 91. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Yao, S.; Hu, Y.; Feng, Y.; Li, M.; Bian, Z.; Zhang, J.; Qin, Y.; Qi, X.; Zhou, L.; et al. The Immune-microenvironment Confers Chemoresistance of Colorectal Cancer through Macrophage-Derived IL6. Clin. Cancer Res. 2017, 23, 7375–7387. [Google Scholar] [CrossRef]
- La, X.; Zhang, L.; Li, Z.; Li, H.; Yang, Y. (-)-Epigallocatechin Gallate (EGCG) Enhances the Sensitivity of Colorectal Cancer Cells to 5-FU by Inhibiting GRP78/NF-κB/miR-155-5p/MDR1 Pathway. J. Agric. Food Chem. 2019, 67, 2510–2518. [Google Scholar] [CrossRef]
- Liu, B.; Hu, J.; Zhao, H.; Zhao, L.; Pan, S. MicroRNA-155-5p Contributes to 5-Fluorouracil Resistance Through Down-Regulating TP53INP1 in Oral Squamous Cell Carcinoma. Front. Oncol. 2022, 11, 706095. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, L.; Dong, Z.; Xu, H.; Yan, L.; Wang, W.; Yang, Q.; Chen, C. MicroRNA-155-5p promotes tumor progression and contributes to paclitaxel resistance via TP53INP1 in human breast cancer. Pathol. Res. Pract. 2021, 220, 153405. [Google Scholar] [CrossRef]
- Herichova, I.; Reis, R.; Hasakova, K.; Vician, M. Downregulation of miR-30c-5p expression in colorectal cancer tissue is sex-dependent. Physiol. Res. 2020, 69 (Suppl. S3), S479–S487. [Google Scholar] [CrossRef]
- Guo, Y.; Guo, Y.; Chen, C.; Fan, D.; Wu, X.; Zhao, L.; Shao, B.; Sun, Z.; Ji, Z. Circ3823 contributes to growth, metastasis and angiogenesis of colorectal cancer: Involvement of miR-30c-5p/TCF7 axis. Mol. Cancer. 2021, 20, 93. [Google Scholar] [CrossRef]
- Niculae, A.M.; Dobre, M.; Herlea, V.; Manuc, T.E.; Trandafir, B.; Milanesi, E.; Hinescu, M.E. Let-7 microRNAs Are Possibly Associated with Perineural Invasion in Colorectal Cancer by Targeting IGF Axis. Life 2022, 12, 1638. [Google Scholar] [CrossRef] [PubMed]
- Boyerinas, B.; Park, S.M.; Shomron, N.; Hedegaard, M.M.; Vinther, J.; Andersen, J.S.; Feig, C.; Xu, J.; Burge, C.B.; Peter, M.E. Identification of Let-7–regulated oncofetal genes. Cancer Res. 2008, 68, 2587–2591. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.H.; Kuwano, Y.; Srikantan, S.; Lee, E.K.; Martindale, J.L.; Gorospe, M. HuR recruits let-7/RISC to repress c-Myc expression. Genes Dev. 2009, 23, 1743–1748. [Google Scholar] [CrossRef] [PubMed]
- Johnson, S.M.; Grosshans, H.; Shingara, J.; Byrom, M.; Jarvis, R.; Cheng, A.; Labourier, E.; Reinert, K.L.; Brown, D.; Slack, F.J. RAS is regulated by the let-7 mi-croRNA family. Cell 2005, 120, 635–647. [Google Scholar] [CrossRef]
- Danac, J.M.C.; Garcia, R.L. CircPVT1 attenuates negative regulation of NRAS by let-7 and drives cancer cells towards oncogenicity. Sci. Rep. 2021, 11, 9021. [Google Scholar] [CrossRef]
- Pidíkova, P.; Reis, R.; Herichova, I. miRNA Clusters with Down-Regulated Expression in Human Colorectal Cancer and Their Regulation. Int. J. Mol. Sci. 2020, 21, 4633. [Google Scholar] [CrossRef]
- Han, H.B.; Gu, J.; Zuo, H.J.; Chen, Z.G.; Zhao, W.; Li, M.; Ji, D.B.; Lu, Y.Y.; Zhang, Z.Q. Let-7c functions as a metastasis suppressor by targeting MMP11 and PBX3 in colorectal can-cer. J. Pathol. 2012, 226, 544–555. [Google Scholar] [CrossRef]
- Yu, J.; Zhang, W.; Ding, D.; Hu, Y.; Guo, G.; Wang, J.; Han, Y. Bioinformatics Analysis Combined With Experiments Predicts PUDP as a Potential Prognostic Biomarker for Hepatocellular Carcinoma Through Its Interaction With Tumor Microenvironment. Front. Oncol. 2022, 12, 830174. [Google Scholar] [CrossRef]
- Yuan, Y.; Jiang, X.; Tang, L.; Yang, H.; Wang, J.; Zhang, D.; Duan, L. Comprehensive Analyses of the Immunological and Prognostic Roles of an IQGAP3AR/let-7c-5p/IQGAP3 Axis in Different Types of Human Cancer. Front. Mol. Biosci. 2022, 9, 763248. [Google Scholar] [CrossRef]
- Uhr, K.; Prager-van der Smissen, W.J.C.; Heine, A.A.J.; Ozturk, B.; van Jaarsveld, M.T.M.; Boersma, A.W.M.; Jager, A.; Wiemer, E.A.C.; Smid, M.; Foekens, J.A.; et al. MicroRNAs as possible indicators of drug sensitivity in breast cancer cell lines. PLoS ONE 2019, 14, e0216400. [Google Scholar] [CrossRef]
- Wang, F.; Zhang, P.; Ma, Y.; Yang, J.; Moyer, M.P.; Shi, C.; Peng, J.; Qin, H. NIRF is frequently upregulated in colorectal cancer and its oncogenicity can be suppressed by let-7a microRNA. Cancer Lett. 2012, 314, 223–231. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Chen, P.; Chang, Y.; Qi, J.; Fu, H.; Guo, H. Let-7a inhibits tumor cell growth and metastasis by directly targeting RTKN in human colon cancer. Biochem. Biophys. Res. Commun. 2016, 478, 739–745. [Google Scholar] [CrossRef] [PubMed]
- Sureban, S.M.; May, R.; Ramalingam, S.; Subramaniam, D.; Natarajan, G.; Anant, S.; Houchen, C.W. Selective blockade of DCAMKL-1 results in tumor growth arrest by a Let-7a MicroRNA-dependent mechanism. Gastroenterology 2009, 137, 649–659. [Google Scholar] [CrossRef] [PubMed]
- Sakatani, A.; Sonohara, F.; Goel, A. Melatonin-mediated downregulation of thymidylate synthase as a novel mechanism for overcoming 5-fluorouracil associated chemoresistance in colorectal cancer cells. Carcinogenesis 2019, 40, 422–431. [Google Scholar] [CrossRef]
- Ciszewski, W.M.; Chmielewska-Kassassir, M.; Wozniak, L.A.; Sobierajska, K. Thymidylate Synthase Overexpression Drives the Invasive Phenotype in Colon Cancer Cells. Biomedicines 2022, 10, 1267. [Google Scholar] [CrossRef] [PubMed]
- He, Y.Z.; Yu, S.L.; Li, X.N.; Bai, X.H.; Li, H.T.; Liu, Y.C.; Lv, B.L.; Zhao, X.M.; Wei, D.; Zhang, H.L.; et al. Curcumin increases crizotinib sensitivity through the inactivation of autophagy via epigenetic modulation of the miR-142-5p/Ulk1 axis in non-small cell lung cancer. Cancer Biomark. 2022, 34, 297–307. [Google Scholar] [CrossRef]
- Li, H.; Gai, L.; Wu, Z.; Li, F. Maternal embryonic leucine zipper kinase serves as a potential prognostic marker and leads to sorafenib chemoresistance modified by miR-142-5p in hepatocellular carcinoma. Mol. Biol. Rep. 2022, 49, 3015–3024. [Google Scholar] [CrossRef]
- Shadbad, M.A.; Asadzadeh, Z.; Derakhshani, A.; Hosseinkhani, N.; Mokhtarzadeh, A.; Baghbanzadeh, A.; Hajiasgharzadeh, K.; Brunetti, O.; Argentiero, A.; Racanelli, V.; et al. A scoping review on the potentiality of PD-L1-inhibiting microRNAs in treating colorectal cancer: Toward single-cell sequencing-guided biocompatible-based delivery. Biomed. Pharmacother. 2021, 143, 112213. [Google Scholar] [CrossRef]
- Zhang, J.; Pang, X.; Lei, L.; Zhang, J.; Zhang, X.; Chen, Z.; Zhu, J.; Jiang, Y.; Chen, G.; Wu, Y.; et al. LncRNA CRART16/miR-122-5p/FOS axis promotes angiogenesis of gastric cancer by upregulating VEGFD expression. Aging 2022, 14, 4137–4157. [Google Scholar] [CrossRef]
- Jepsen, R.K.; Novotny, G.W.; Klarskov, L.L.; Bang-Berthelsen, C.H.; Haakansson, I.T.; Hansen, A.; Christensen, I.J.; Riis, L.B.; Høgdall, E. Early metastatic colorectal cancers show increased tissue expression of miR-17/92 cluster members in the invasive tumor front. Hum. Pathol. 2018, 80, 231–238. [Google Scholar] [CrossRef]
- Mastropasqua, F.; Marzano, F.; Valletti, A.; Aiello, I.; Di Tullio, G.; Morgano, A.; Liuni, S.; Ranieri, E.; Guerrini, L.; Gasparre, G.; et al. TRIM8 restores p53 tumour suppressor function by blunting N-MYC activity in chemo-resistant tumours. Mol. Cancer. 2017, 16, 67. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, A.; Ramalhete, S.V.; Mestre, A.; Pires das Neves, R.; Marreiros, A.; Castelo-Branco, P.; Roberto, V.P. Identification of colorectal cancer associated biomarkers: An integrated analysis of miRNA expression. Aging 2021, 13, 21991–22029. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Xu, Y.; Liu, S.; Qiao, L.; Sun, J.; Zhao, Q. MicroRNAs Associated With Colon Cancer: New Potential Prognostic Markers and Targets for Therapy. Front. Bioeng Biotechnol. 2020, 8, 176. [Google Scholar] [CrossRef]
- Baretti, M.; Azad, N.S. The role of epigenetic therapies in colorectal cancer. Curr. Probl. Cancer. 2018, 42, 530–547. [Google Scholar] [CrossRef]
- Skarkova, V.; Skarka, A.; Manethova, M.; Stefanidi, A.A.; Rudolf, E. Silencing of E-cadherin expression leads to increased chemosensitivity to irinotecan and oxaliplatin in colorectal cancer cell lines. Hum. Exp. Toxicol. 2021, 40, 2063–2073. [Google Scholar] [CrossRef] [PubMed]
- Druzhkova, I.; Ignatova, N.; Prodanets, N.; Kiselev, N.; Zhukov, I.; Shirmanova, M.; Zagainov, V.; Zagaynova, E. E-Cadherin in Colorectal Cancer: Relation to Chemosensitivity. Clin. Color. Cancer. 2019, 18, e74–e86. [Google Scholar] [CrossRef]
- Wäsch, R.; Engelbert, D. Anaphase-promoting complex-dependent proteolysis of cell cycle regulators and genomic instability of cancer cells. Oncogene 2005, 24, 1–10. [Google Scholar] [CrossRef]
- Wang, H.; Yang, W.; Qin, Q.; Yang, X.; Yang, Y.; Liu, H.; Lu, W.; Gu, S.; Cao, X.; Feng, D.; et al. E3 ubiquitin ligase MAGI3 degrades c-Myc and acts as a predictor for chemotherapy response in colorectal cancer. Mol. Cancer. 2022, 21, 151. [Google Scholar] [CrossRef]
- Tomicic, M.T.; Steigerwald, C.; Rasenberger, B.; Brozovic, A.; Christmann, M. Functional mismatch repair and inactive p53 drive sensitization of colorectal cancer cells to irinotecan via the IAP antagonist BV6. Arch. Toxicol 2019, 93, 2265–2277. [Google Scholar] [CrossRef]
- Dervenis, C.; Xynos, E.; Sotiropoulos, G.; Gouvas, N.; Boukovinas, I.; Agalianos, C.; Androulakis, N.; Athanasiadis, A.; Christodoulou, C.; Chrysou, E.; et al. Clinical practice guidelines for the management of metastatic colorectal cancer: A consensus statement of the Hellenic Society of Medical Oncologists (HeSMO). Ann. Gastroenterol. 2016, 29, 390–416. [Google Scholar] [CrossRef]
- Eisenhauer, E.A.; Therasse, P.; Bogaerts, J.; Schwartz, L.H.; Sargent, D.; Ford, R.; Jänne, P.A.; Johnson, B.E.; Van den Abbeele, A.D. New response evaluation criteria in solid tumors: Revised RECIST guideline (version 1.1). Eur. J. Cancer 2009, 45, 228–247. [Google Scholar] [CrossRef] [PubMed]
- LeDell, E.; Gill, N.; Aiello, S.; Fu, A.; Candel, A.; Click, C.; Kraljevic, T.; Nykodym, T.; Aboyoun, P.; Kurka, M.; et al. H2O: R Interface for the ‘H2O’ Scalable Machine Learning Platform, 2020. Available online: https://github.com/h2oai/h2o-3 (accessed on 12 November 2022).
- Ru, Y.; Kechris, K.J.; Tabakoff, B.; Hoffman, P.; Radcliffe, R.A.; Bowler, R.; Mahaffey, S.; Rossi, S.; Calin, G.A.; Bemis, L. The multiMiR R package and database: Integration of microRNA–target interactions along with their disease and drug associations. Nucleic Acids Res. 2014, 42, e133. [Google Scholar] [CrossRef] [PubMed]
- Xiao, F.; Zuo, Z.; Cai, G.; Kang, S.; Gao, X.; Li, T. miRecords: An integrated resource for microRNA–target interactions. Nucleic Acids Res. 2009, 37, D105–D110. [Google Scholar] [CrossRef]
- Huang, H.Y.; Lin, Y.C.D.; Li, J.; Huang, K.Y.; Shrestha, S.; Hong, H.C.; Tang, Y.; Chen, Y.G.; Jin, C.N.; Yu, Y.; et al. miRTarBase 2020: Updates to the experimentally validated microRNA-target interaction database. Nucleic Acids Res. 2020, 48, D148–D154. [Google Scholar] [CrossRef]
- Paraskevopoulou, M.D.; Vlachos, I.S.; Hatzigeorgiou, A.G. DIANA-TarBase and DIANA suite tools: Studying experimentally supported microRNA targets. Curr. Protoc. Bioinform. 2016, 55, 12.14.1–12.14.18. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene Ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef]
| miRNA | Fold Regulation | p-Value | p-Adjusted | Fold Change | 95% CI |
|---|---|---|---|---|---|
| hsa-miR-181b-5p | 116.66 | 0.00000 | 0.00000 | 116.66 | (40.84, 192.47) |
| hsa-miR-10b-5p | 47.20 | 0.00000 | 0.00000 | 47.20 | (16.08, 78.31) |
| hsa-let-7f-5p | 24.32 | 0.00188 | 0.00295 | 24.32 | (9.28, 39.36) |
| hsa-miR-181a-5p | 24.03 | 0.00000 | 0.00000 | 24.03 | (13.52, 34.53) |
| hsa-miR-181d-5p | 23.33 | 0.00000 | 0.00000 | 23.33 | (12.72, 33.94) |
| hsa-miR-301a-3p | 23.21 | 0.00000 | 0.00000 | 23.21 | (14.58, 31.83) |
| hsa-miR-92a-3p | 6.11 | 0.00000 | 0.00000 | 6.11 | (3.00, 9.22) |
| hsa-miR-155-5p | 6.11 | 0.00000 | 0.00000 | 6.11 | (3.00, 9.22) |
| hsa-miR-30c-5p | 4.06 | 0.00000 | 0.00000 | 4.06 | (2.19, 5.94) |
| hsa-let-7i-5p | 3.78 | 0.03537 | 0.04421 | 3.78 | (1.51, 6.04) |
| hsa-miR-29a-3p | 3.77 | 0.00009 | 0.00019 | 3.77 | (2.17, 5.37) |
| hsa-miR-138-5p | 3.70 | 0.00000 | 0.00000 | 3.70 | (2.46, 4.95) |
| hsa-miR-193b-3p | 3.51 | 0.00083 | 0.00143 | 3.51 | (1.70, 5.32) |
| hsa-miR-133b | 2.52 | 0.00010 | 0.00020 | 2.52 | (0.99, 4.05) |
| hsa-miR-218-5p | 2.28 | 0.00059 | 0.00112 | 2.28 | (1.45, 3.11) |
| hsa-miR-150-5p | 2.10 | 0.01732 | 0.02215 | 2.10 | (1.29, 2.91) |
| hsa-miR-196a-5p | −2.18 | 0.00797 | 0.01124 | 0.46 | (0.17, 0.74) |
| hsa-miR-20a-5p | −2.85 | 0.00004 | 0.00009 | 0.35 | (0.22, 0.49) |
| hsa-miR-27b-3p | −4.10 | 0.00075 | 0.00133 | 0.24 | (0.14, 0.35) |
| hsa-miR-200c-3p | −4.28 | 0.00000 | 0.00000 | 0.23 | (0.14, 0.32) |
| hsa-miR-193a-5p | −6.74 | 0.00000 | 0.00000 | 0.15 | (0.10, 0.20) |
| hsa-miR-134-5p | −7.02 | 0.00000 | 0.00000 | 0.14 | (0.09, 0.20) |
| hsa-miR-20b-5p | −7.34 | 0.00000 | 0.00000 | 0.14 | (0.07, 0.20) |
| hsa-miR-124-3p | −7.56 | 0.00000 | 0.00000 | 0.13 | (0.09, 0.18) |
| hsa-miR-17-5p | −8.35 | 0.00000 | 0.00000 | 0.12 | (0.07, 0.17) |
| hsa-miR-122-5p | −8.71 | 0.00000 | 0.00000 | 0.11 | (0.06, 0.17) |
| hsa-miR-148a-3p | −14.62 | 0.00000 | 0.00000 | 0.07 | (0.03, 0.11) |
| hsa-miR-142-5p | −16.53 | 0.00001 | 0.00003 | 0.06 | (0.03, 0.09) |
| hsa-miR-10a-5p | −17.65 | 0.00006 | 0.00013 | 0.06 | (0.04, 0.08) |
| hsa-let-7a-5p | −19.54 | 0.00000 | 0.00000 | 0.05 | (0.03, 0.07) |
| hsa-miR-143-3p | −23.44 | 0.00001 | 0.00003 | 0.04 | (0.02, 0.07) |
| hsa-miR-215-5p | −26.06 | 0.00002 | 0.00005 | 0.04 | (0.02, 0.05) |
| hsa-let-7c-5p | −36.94 | 0.00000 | 0.00000 | 0.03 | (0.01, 0.04) |
| hsa-let-7d-5p | −54.23 | 0.00000 | 0.00000 | 0.02 | (0.01, 0.03) |
| Characteristics | Number of Patients | |
|---|---|---|
| Gender | Male | 58 |
| Female | 37 | |
| Surgery on primary | Yes | 83 |
| No | 12 | |
| Primary Location | Cecum | 9 |
| Ascending colon | 5 | |
| Ascending/Transverse colon | 1 | |
| Transverse colon | 3 | |
| Splenic Flexure | 3 | |
| Descending colon | 3 | |
| Descending colon/Rectum | 1 | |
| Sigmoid | 26 | |
| Rectosigmoid | 21 | |
| Rectum | 23 | |
| Metastasis site | Lung | 23 |
| Liver | 37 | |
| Peritoneum | 7 | |
| Adrenal gland | 1 | |
| Multiple sites | 20 | |
| Local Recurrence | 7 | |
| Therapeutics | Panitumumab, irinotecan and 5-FU | 10 |
| Panitumumab, irinotecan and capecitabine | 8 | |
| Bevacizumab, irinotecan and 5-FU | 21 | |
| Bevacizumab, irinotecan and capecitabine | 27 | |
| Aflibercept, irinotecan and 5-FU | 3 | |
| Cetuximab, irinotecan and 5-FU | 4 | |
| Bevacizumab, irinotecan, 5-FU and oxaliplatin | 1 | |
| Bevacizumab, irinotecan, capecitabine and oxaliplatin | 1 | |
| Bevacizumab, irinotecan and oxaliplatin | 1 | |
| Capecitabine and irinotecan | 10 | |
| Oxaliplatin and irinotecan | 3 | |
| Irinotecan and 5-FU | 5 | |
| Panitumumab and irinotecan | 1 | |
| Response status | Stable Disease (SD) | 47 |
| Progressive Disease (PD) | 48 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pliakou, E.; Lampropoulou, D.I.; Dovrolis, N.; Chrysikos, D.; Filippou, D.; Papadimitriou, C.; Vezakis, A.; Aravantinos, G.; Gazouli, M. Circulating miRNA Expression Profiles and Machine Learning Models in Association with Response to Irinotecan-Based Treatment in Metastatic Colorectal Cancer. Int. J. Mol. Sci. 2023, 24, 46. https://doi.org/10.3390/ijms24010046
Pliakou E, Lampropoulou DI, Dovrolis N, Chrysikos D, Filippou D, Papadimitriou C, Vezakis A, Aravantinos G, Gazouli M. Circulating miRNA Expression Profiles and Machine Learning Models in Association with Response to Irinotecan-Based Treatment in Metastatic Colorectal Cancer. International Journal of Molecular Sciences. 2023; 24(1):46. https://doi.org/10.3390/ijms24010046
Chicago/Turabian StylePliakou, Evangelia, Dimitra Ioanna Lampropoulou, Nikolas Dovrolis, Dimosthenis Chrysikos, Dimitrios Filippou, Christos Papadimitriou, Antonios Vezakis, Gerasimos Aravantinos, and Maria Gazouli. 2023. "Circulating miRNA Expression Profiles and Machine Learning Models in Association with Response to Irinotecan-Based Treatment in Metastatic Colorectal Cancer" International Journal of Molecular Sciences 24, no. 1: 46. https://doi.org/10.3390/ijms24010046
APA StylePliakou, E., Lampropoulou, D. I., Dovrolis, N., Chrysikos, D., Filippou, D., Papadimitriou, C., Vezakis, A., Aravantinos, G., & Gazouli, M. (2023). Circulating miRNA Expression Profiles and Machine Learning Models in Association with Response to Irinotecan-Based Treatment in Metastatic Colorectal Cancer. International Journal of Molecular Sciences, 24(1), 46. https://doi.org/10.3390/ijms24010046









