Transcriptome Profiling Provides New Insights into the Molecular Mechanism Underlying the Sensitivity of Cotton Varieties to Mepiquat Chloride
Abstract
:1. Introduction
2. Results
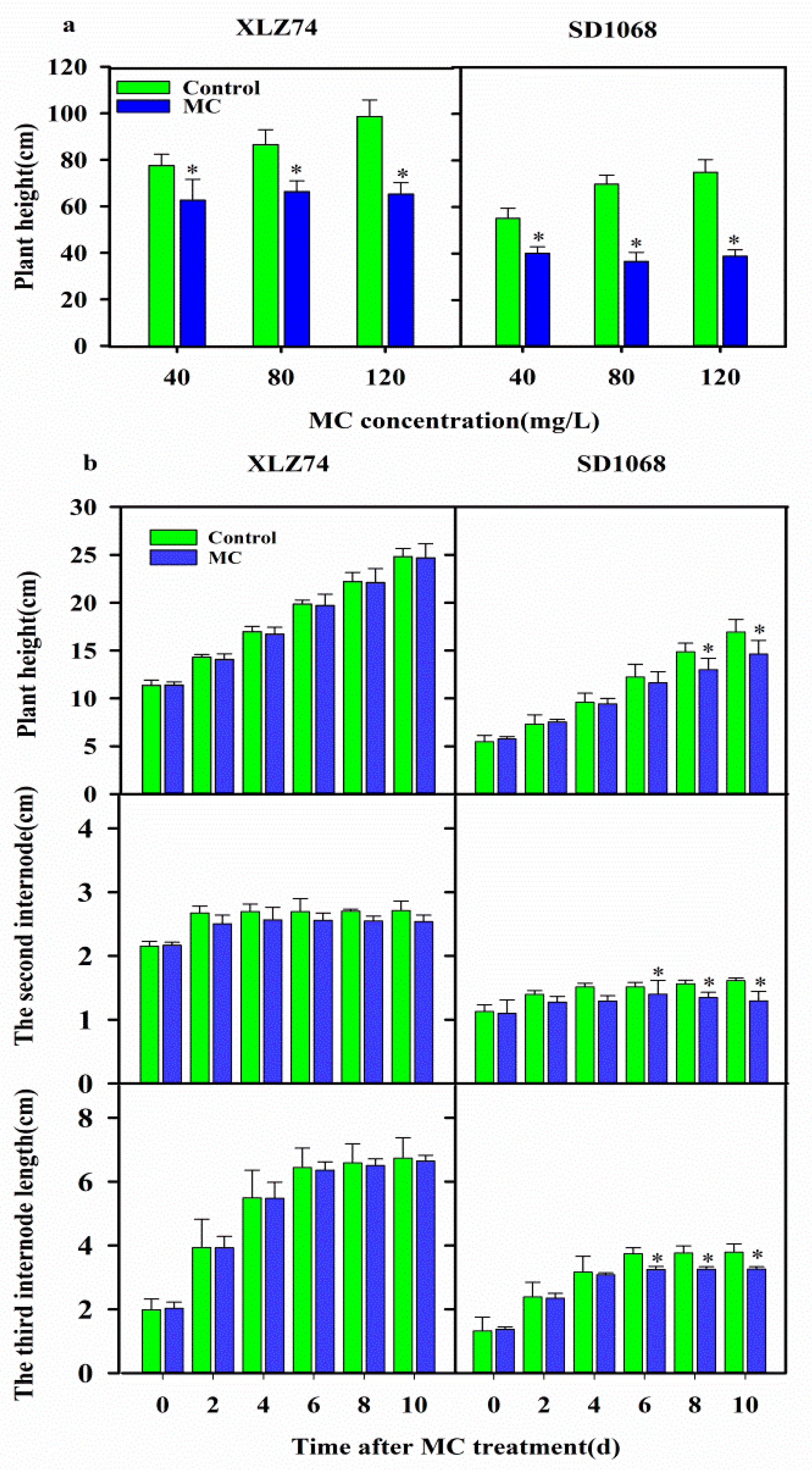
2.1. The Sensitivity of Cotton Varieties to MC
2.2. An Overview of the RNA-Seq Data
2.3. Analyses of Differentially Expressed Genes (DEGs)
2.4. GO and KEGG Analyses
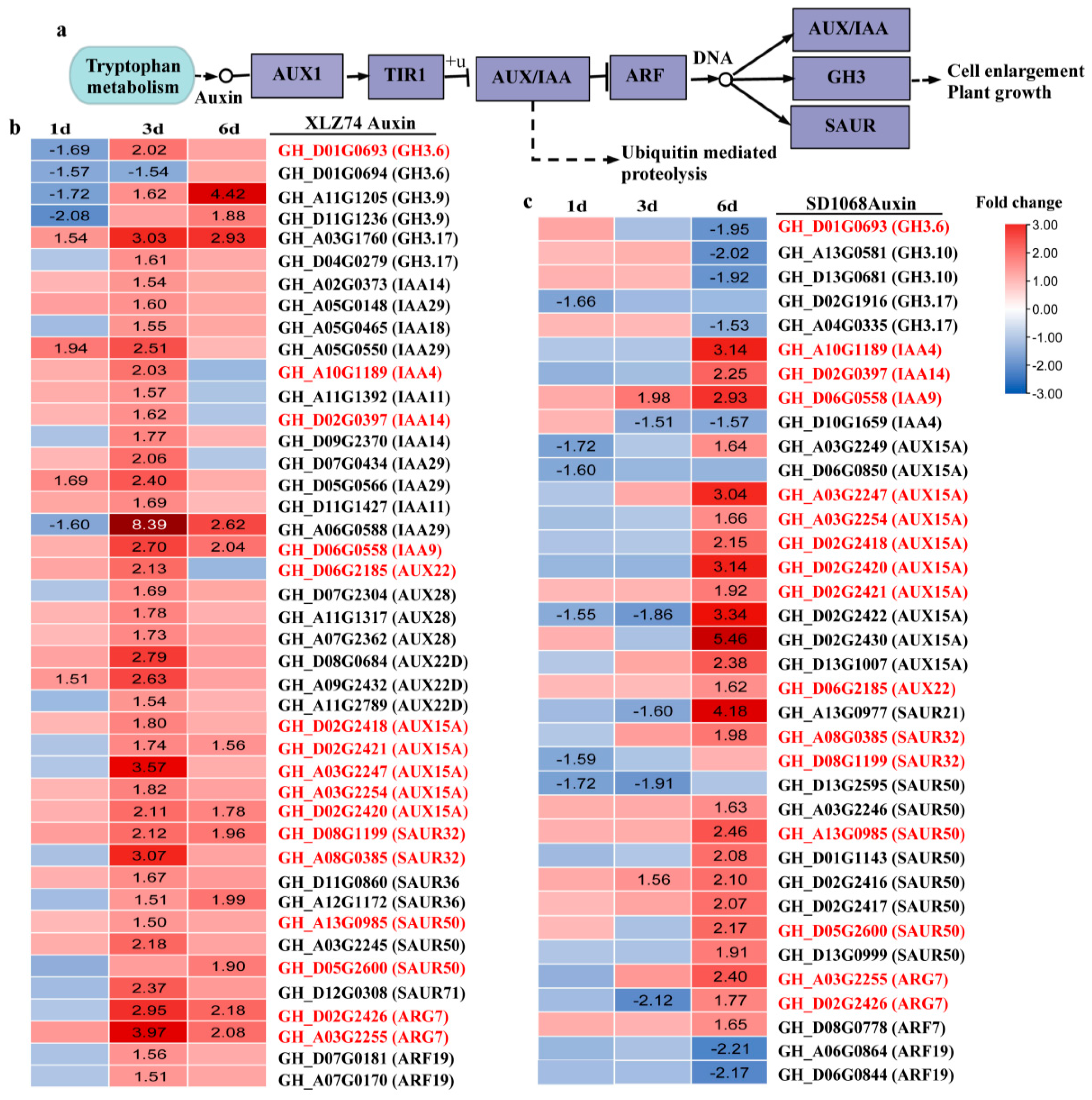
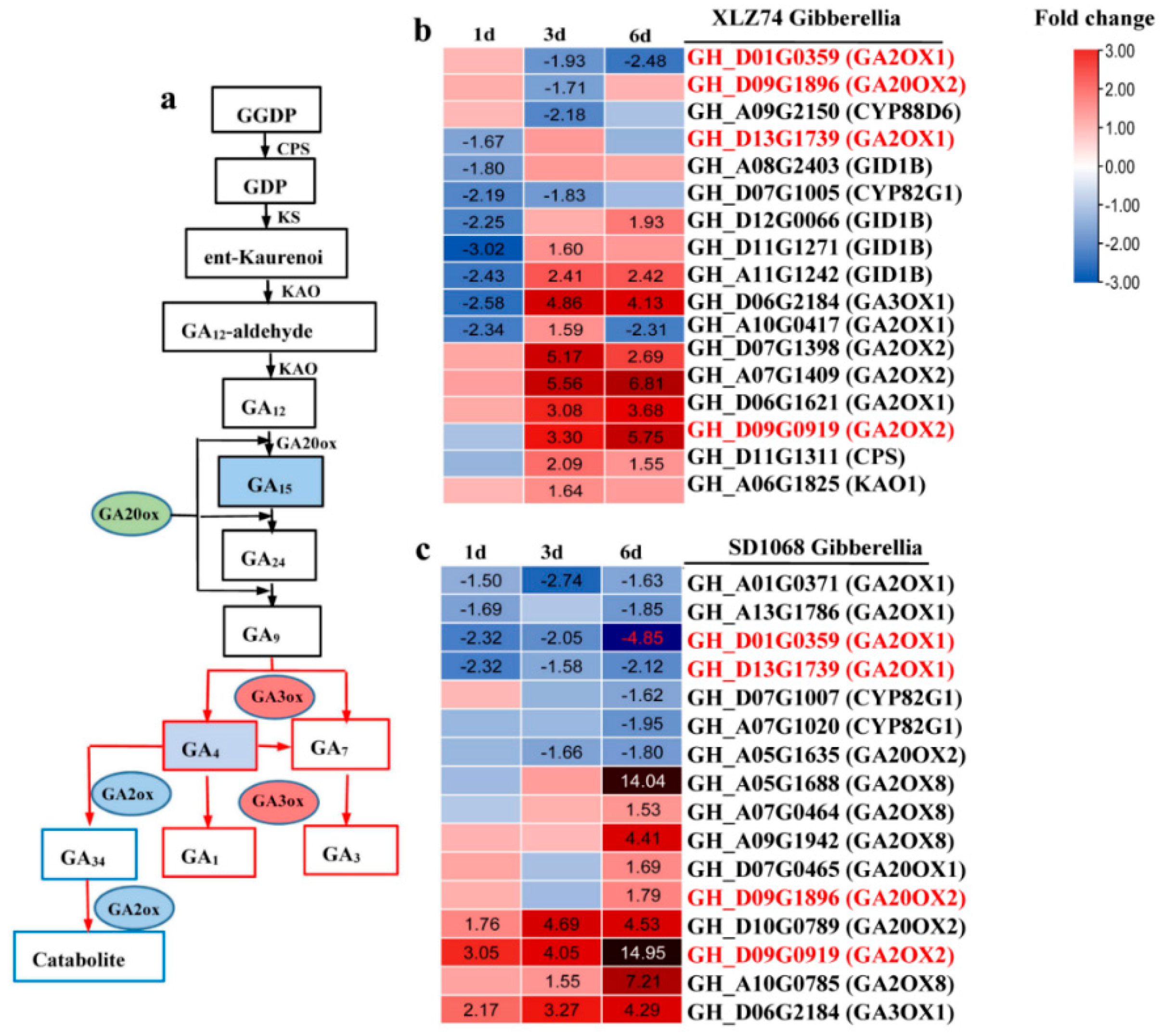
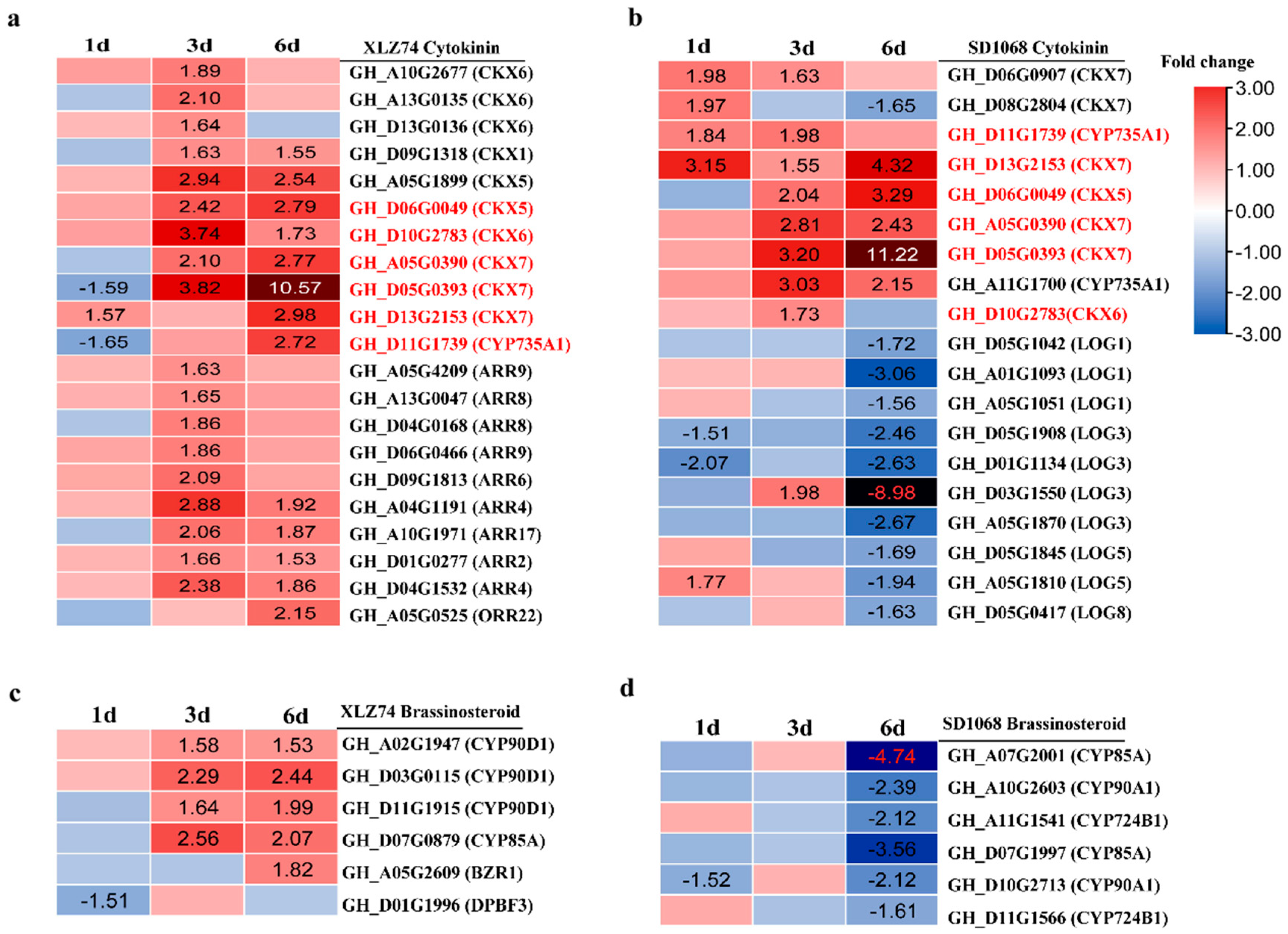
2.5. DEGs Related to Hormone Metabolism and Signaling Pathways
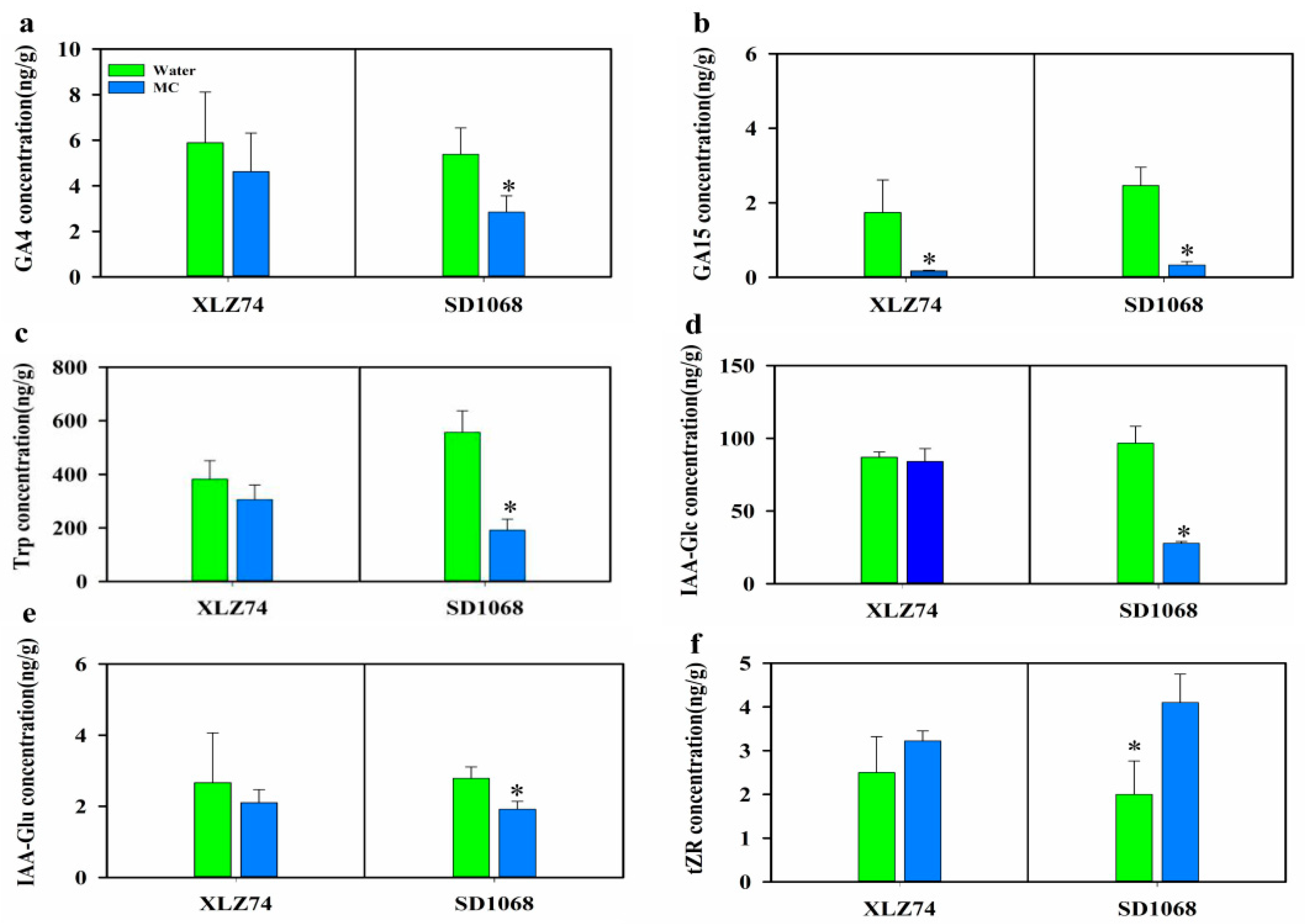
2.6. The Effect of MC on the Contents of Endogenous Hormones in the Two Varieties
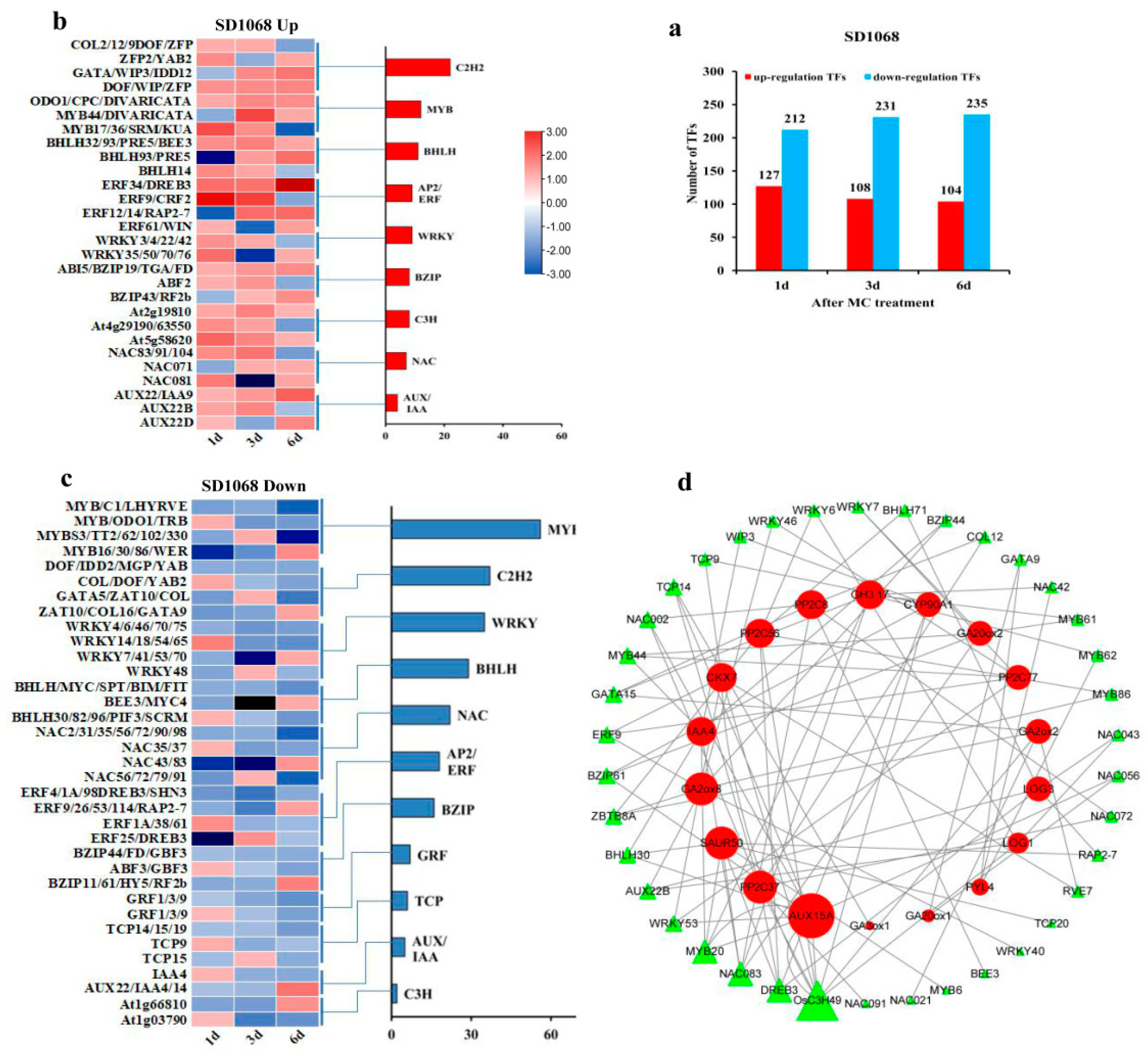
2.7. Differentially Expressed Transcription Factors
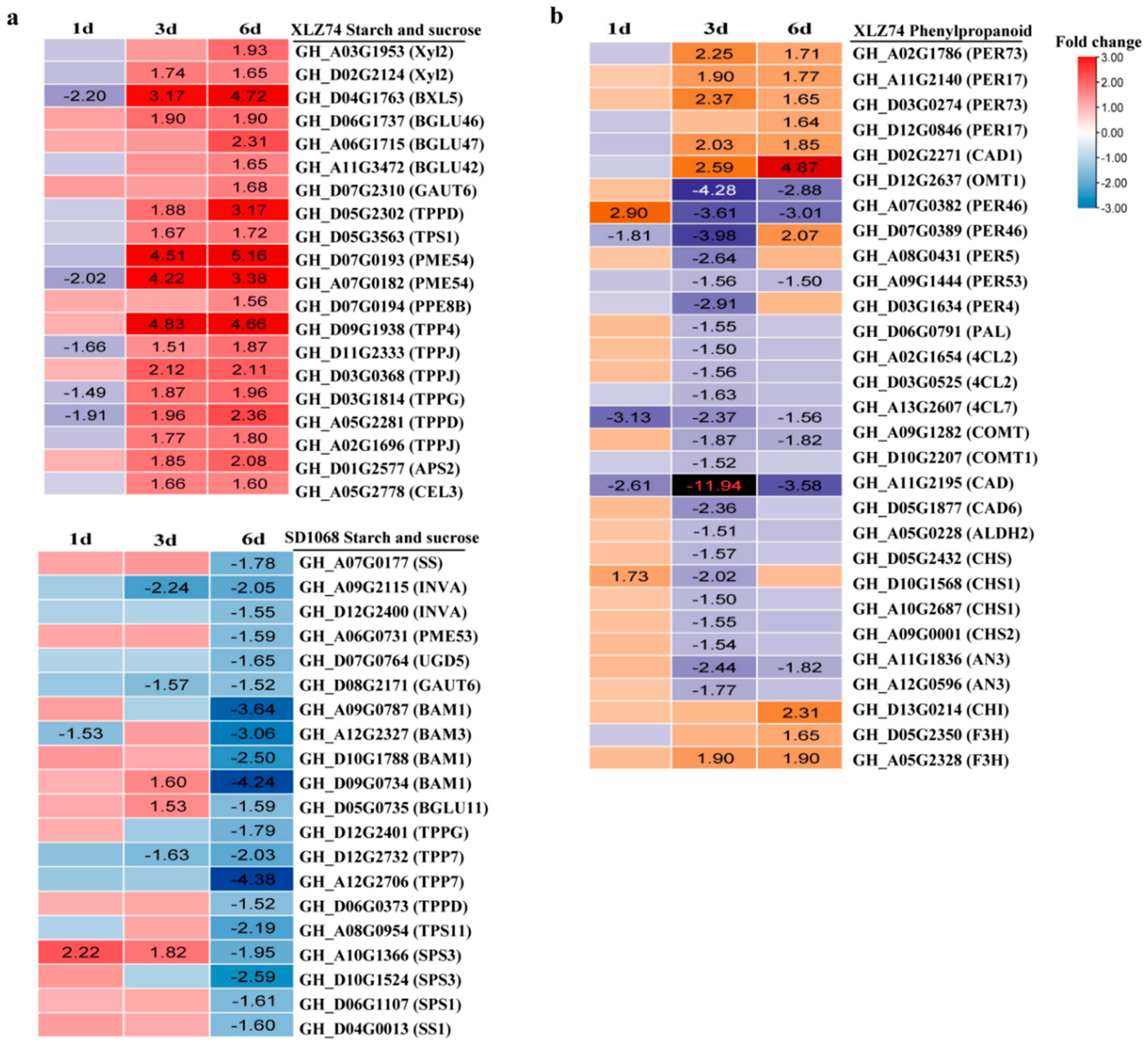
2.8. DEGs Related to Metabolism of Starch, Sucrose and Phenylpropanoid
3. Discussion
3.1. MC Treatment Induces Significant Upregulation of GA Catabolic Genes GA2ox, Especially in SD1068
3.2. MC-Induced Upregulation of Auxin Signaling Genes Is Delayed in SD1068
3.3. MC-Induced Upregulation of CTK Signaling Genes Is Earlier in SD1068 Than in XLZ74
3.4. MC Treatment Suppresses BR Biosynthesis-Related Genes Only in SD1068
3.5. MC Treatment Induces Different Profiles of TFs in XLZ74 and SD1068
3.6. MC Treatment Suppresses Sugar and Starch Pathway in SD1068
4. Materials and Methods
4.1. Plant Materials and MC Treatments
4.2. RNA Extraction and Sequencing
4.3. RNA-seq Analysis and Identification of Differentially Expressed Genes
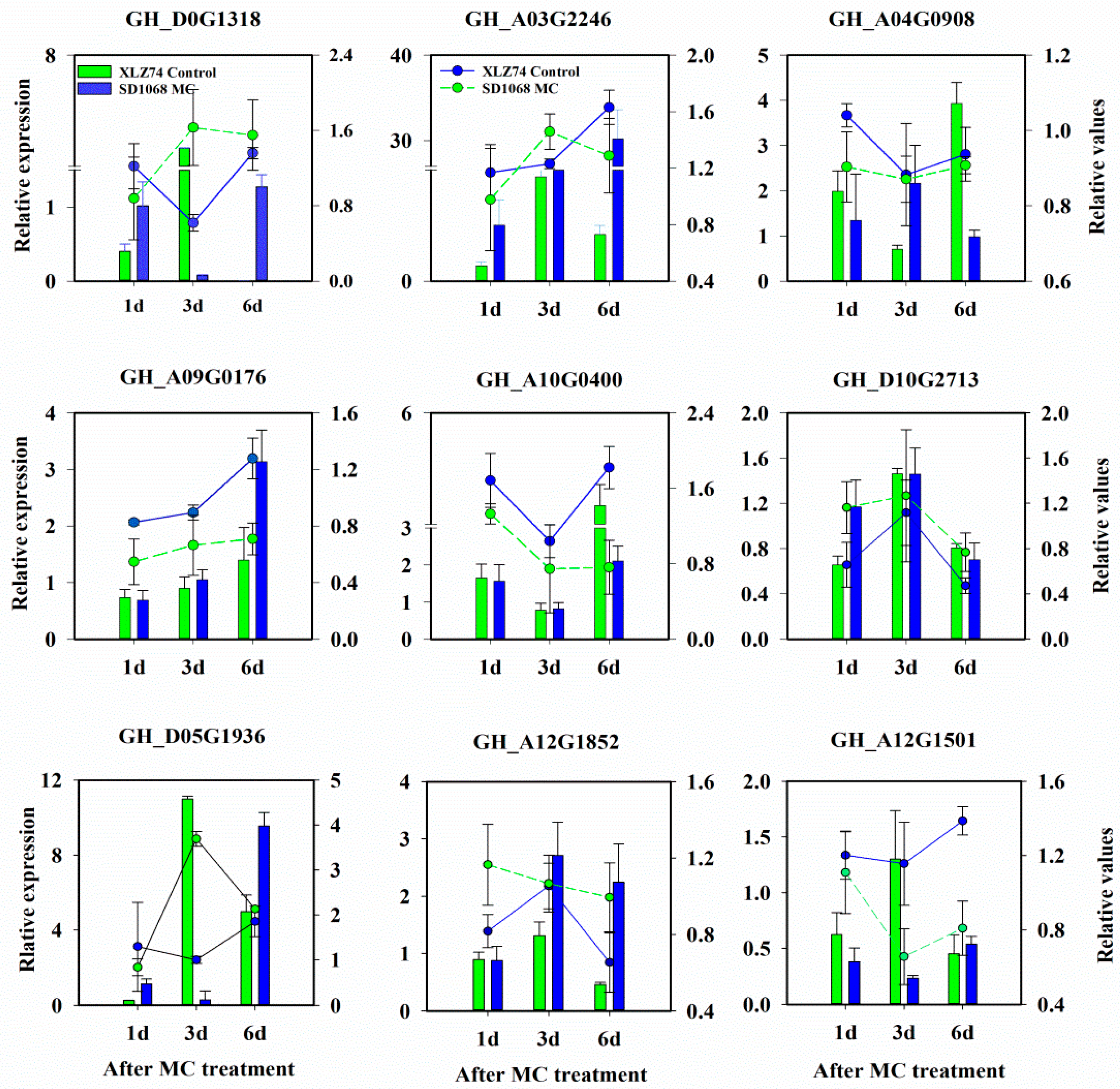
4.4. qRT-PCR Analysis
4.5. Hormone Quantification
4.6. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhao, D.; Oosterhuis, D.M. Pix plus and mepiquat chloride effects on physiology, growth, and yield of field-grown cotton. J. Plant Growth Regul. 2000, 19, 415–422. [Google Scholar] [CrossRef]
- Cook, D.R.; Kennedy, W. Early flower bud loss and mepiquat chloride effects on cotton yield distribution. Crop Sci. 2000, 40, 1678–1684. [Google Scholar] [CrossRef]
- Wang, L.; Mu, C.; Du, M.; Chen, Y.; Tian, X.; Zhang, M.; Li, Z. The effect of mepiquat chloride on elongation of cotton (Gossypium hirsutum L.) internode is associated with low concentration of gibberellic acid. Plant Sci. 2014, 225, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Reddy, V.R.; Baker, D.N.; Hodges, H.F. Temperature and mepiquat chloride effects on cotton canopy architecture. Agron. J. 1990, 82, 190–195. [Google Scholar] [CrossRef]
- Almeida, A.Q.D.; Rosolem, C.A. Cotton root and shoot growth as affected by application of mepiquat chloride to cotton seeds. Acta Sci.-Agron. 2012, 34, 61–65. [Google Scholar] [CrossRef]
- Ren, X.; Zhang, L.; Du, M.; Jochem, B.E.; Wopke, V.D.W.; Xiao, L.; Tian, X.; Li, Z. Managing mepiquat chloride and plant density for optimal yield and quality of cotton. Field Crops Res. 2013, 149, 1–10. [Google Scholar] [CrossRef]
- Wu, Q.; Du, M.; Wu, J.; Wang, N.; Wang, B.; Li, F.; Tian, X.; Li, Z. Mepiquat chloride promotes cotton lateral root formation by modulating plant hormone homeostasis. BMC Plant Biol. 2019, 19, 573. [Google Scholar] [CrossRef]
- Olszewski, N.; Sun, T.P.; Gubler, F. Gibberellin signaling: Biosynthesis, catabolism, and response pathways. Plant Cell. 2002, 14, 61–80. [Google Scholar] [CrossRef] [Green Version]
- Hedden, P. Gibberellin metabolism and its regulation. J. Plant Growth Regul. 2002, 20, 317–318. [Google Scholar] [CrossRef]
- Rademacher, W. Growth retardants: Effects on gibberellin biosynthesis and other metabolic pathways. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2000, 51, 501–531. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Yin, Y.; Wang, L.; Wang, M.; Zhao, M.; Tian, Y.; Li, Y. Transcriptome profiling of the elongating internode of cotton (Gossypium hirsutum L.) seedlings in response to mepiquat chloride. Front. Plant Sci. 2020, 10, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Tung, S.A.; Huang, Y.; Ali, S.; Hafeez, A.; Shah, A.N.; Song, X.; Ma, X.; Luo, D.; Yang, G. Mepiquat chloride application does not favor leaf photosynthesis and carbohydrate metabolism as well as lint yield in late-planted cotton at high plant density. Field Crop Res. 2018, 221, 108–118. [Google Scholar] [CrossRef]
- Matsoukis, A.; Gasparatos, D.; Chronopoulou, S.A. Mepiquat chloride and shading effects on specific leaf and K, P, Ca, Fe and Mn content of Lantana camara L. Emir. J. Food Agric. 2015, 27, 121–125. [Google Scholar] [CrossRef]
- Bogiani, J.C.; Rosolem, C.A. Sensibility of cotton varieties to mepiquat chloride. Pesq. Agrop. Bras. 2009, 44, 1246–1253. [Google Scholar] [CrossRef]
- Gencsoylu, I. Effect of plant growth regulators on agronomic characteristics, lint quality, pests, and predators in cotton. J. Plant Growth Regul. 2009, 28, 147–153. [Google Scholar] [CrossRef]
- Tung, S.A.; Huang, Y.; Hafeez, A.; Ali, S.; Khan, A.; Souliyanonh, B.; Song, X.; Liu, A.; Yang, G. Mepiquat chloride effects on cotton yield and biomass accumulation under late sowing and high density. Field Crops Res. 2018, 215, 59–65. [Google Scholar] [CrossRef]
- Woodward, A.W.; Bartel, B. Auxin: Regulation, action, and interaction. Ann. Bot. 2005, 95, 70–735. [Google Scholar] [CrossRef] [Green Version]
- Jones, R.J.; Schreiber, B.M.M. Role and function of cytokinin oxidase in plants. Plant Growth Regul. 1997, 23, 123–134. [Google Scholar] [CrossRef]
- Kieber, J.J.; Schaller, G.E. Cytokinin signaling in plant development. Development 2018, 145, dev.149344. [Google Scholar] [CrossRef] [Green Version]
- Kurakawa, T.; Ueda, N.; Maekawa, M.; Kobayashi, K.; Kojima, M.; Nagato, Y.; Sakakibara, H.; Kyozuka, J. Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature. 2007, 445, 652–655. [Google Scholar] [CrossRef]
- Kuroha, T.; Tokunaga, H.; Kojima, M.; Ueda, N.; Ishida, T.; Nagawa, S.; Fukuda, H.; Sugimoto, K.; Sakakibara, H. Functional analyses of LONELY GUY cytokinin-activating enzymes reveal the importance of the direct activation pathway in Arabidopsis. Plant Cell 2009, 21, 3152–3169. [Google Scholar] [CrossRef] [Green Version]
- Cheon, J.; Fujioka, S.; Dilkes, B.P.; Choe, S. Brassionosteroids regulate plant growth through distinct signaling pathways in Selaginella and Arabidopsis. PLoS ONE 2013, 8, e81938. [Google Scholar] [CrossRef] [Green Version]
- Yeates, S.; Constable, G.; McCumstie, T. Developing management options for mepiquat chloride in tropical winter season cotton. Field Crop Res. 2002, 74, 217–230. [Google Scholar] [CrossRef]
- Chen, R.; Fan, Y.; Zhou, H.; Mo, S.; Zhou, Z.; Yan, H.; Luo, T.; Huang, X.; Weng, M.; Lakshmanan, P.; et al. Global transcriptome changes of elongating internode of sugarcane in response to mepiquat chloride. BMC Genom. 2021, 22, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Sonia, S.C.; Nancy, M.M.; Jenny, G.S.; Rocío, P.T.; Rebeca, D.M.C. Gibberellin biosynthesis and metabolism: A convergent route for plants, fungi and bacteria. Microbiol. Res. 2018, 208, 85–89. [Google Scholar]
- Thomas, S.G.; Rieu, I.; Steber, C.M. Gibberellin metabolism and signaling. Vitam. Horm. 2005, 72, 289–338. [Google Scholar]
- Rieu, I.; Eriksson, S.; Powers, S.J.; Stephen, J.; Gong, F.; Griffiths, J.; Woolley, L.; Benlloch, R.; Nilsson, O.; Hedden, P.; et al. Genetic analysis reveals that C19-GA2-Oxidation is a major gibberellin inactivation pathway in Arabidopsis. Plant Cell. 2008, 20, 2420–2436. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lo, S.F.; Yang, S.Y.; Chen, K.T.; Hsing, Y.L.; Zeevaart, J.A.D.; Chen, L.J.; Yu, S.M. A novel class of 2-oxidases control semidwarfism, tillering, and root development in rice. Plant Cell. 2008, 20, 2603–2618. [Google Scholar] [CrossRef] [Green Version]
- Dayan, J.; Schwarzkopf, M.; Avni, A.; Aloin, K. Enhancing plant growth and fiber production by silencing GA2-oxidase. Plant Biothchnol. J. 2010, 8, 425–435. [Google Scholar] [CrossRef]
- Huang, J.; Tang, D.; Shen, Y.; Qin, B.X.; Hong, L.; You, A.Q.; Li, M.; Wang, X.; Yu, H.; Gu, M.; et al. Activation of gibberel-lin 2-oxidase 6 decreases active gibberellin levels and creates a donimant semi dwarf phenotype in rice (Oryza sativa L). J. Genet. Genom. 2010, 37, 23–36. [Google Scholar] [CrossRef]
- Ford, B.A.; Foo, E.; Sharwood, R.; Karafiatova, M.; Vrana, J.; MacMillan, C.; Nichols, D.S.; Steuernagel, B.; Uauy, C.; Dolezel, J.; et al. Rht18 semidwarfism in wheat is due to increased GA2-oxidaseA9 expression and reduced GA content. Plant Physiol. 2018, 177, 168–180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, J.; Liao, X.; He, R.; Zhong, M.; Feng, P.; Li, X.; Tang, D.; Liu, X.; Zhao, X. Ectopic expression of GA2-oxidase 6 from rapeseed (Brassica napus L.) causes dwarfism, late flowering and enhanced chlorophyll accumulation in Arabidopsis thaliana. Plant Physiol. Bioch. 2017, 111, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Otani, M.; Meguro, S.; Gondaira, H.; Hayashi, M.; Saito, M.; Han, D.; Inthima, P.; Supaibulwatana, K.; Mori, S.; Jikumaru, K.Y.; et al. Overexpression of the gibberellin 2-oxidase gene from torenia fournieri induces dwarf phenotypes in the liliaceous monocotyledon Tricyrtis sp. J. Plant Physiol. 2013, 170, 1416–1423. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Xiang, F.J.; Yang, P.; Li, X.; Zhong, M.; He, R.; Li, X.; Peng, W.; Liu, X.; Zhao, X.Y. Overexpression of BnGA2ox2, a rapeseed gibberellin 2-oxidase, causes dwarfism and increased chlorophyll and anthocyanin accumulation in Arabidopsis and repeseed. Plant Growth Regul. 2021, 93, 65–77. [Google Scholar] [CrossRef]
- Song, Y.L.; You, J.; Xiong, L.Z. Characterization of OsIAA1 gene, a member of rice Aux/IAA family involved in auxin and brassinosteroid hormone responses and plant morphogenesis. Plant Mol. Biol. 2009, 70, 297–309. [Google Scholar] [CrossRef]
- Kloosterman, B.; Visser, R.G.F.; Bachem, C.W.B. Isolation and characterization of a novel potato AuxinIndole-3-Acetic Acid family member (StIAA2) that is involved in petiole hyponasty and shoot morphogenesis. Plant Physiol. Bioch. 2006, 44, 766–775. [Google Scholar] [CrossRef]
- Kant, S.; Rothstein, S. Auxin-responsive SAUR39 gene modulates auxin level in rice. Plant Signal Behav. 2009, 4, 1174–1175. [Google Scholar] [CrossRef] [Green Version]
- Kant, S.; Bi, Y.M.; Zhu, T.; Rothstein, S.J. SAUR39, a small auxin-Up RNA gene, acts as a negative regulator of auxin synthesis and transport in rice. Plant Physiol. 2009, 151, 691–701. [Google Scholar] [CrossRef] [Green Version]
- Schülling, T.; Werner, T.; Riefler, M.; Krupková, E.; Bartrinay, M.I. Structure and function of cytokinin oxidase/dehydrogenase genes of maize, rice, Arabidopsis and other species. J. Plant Res. 2003, 116, 241–252. [Google Scholar] [CrossRef]
- Bhargava, A.; Clabaugh, I.; To, J.P.; Maxwell, B.B.; Chiang, Y.H.; Schaller, G.E.; Loraine, A.; Kieber, J.J. Identification of cytokinin responsive genes using microarray meta-analysis and RNA-seq in Arobidopsis. Plant Physiol. 2013, 162, 272–294. [Google Scholar] [CrossRef] [Green Version]
- Sakakibara, H. Cytokinins: Activity, biosynthesis, and translocation. Annu. Rev. Plant Biol. 2006, 57, 431–449. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bishop, G.J. Brassinosteroid Mutants of Crops. J. Plant Growth Regul. 2003, 22, 325–335. [Google Scholar] [CrossRef] [PubMed]
- Byrne, M.E.; Barley, R.; Curtis, M.; Arroyo, J.M.; Dunham, M.; Hudson, A.; Martienssen, R.A. Asymmetric leaves mediates leaf patterning and stem cell function in Arabidopsis. Nature. 2000, 408, 967–971. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.; Nishizawa, T.; Higashitani, A.; Suge, H.; Wakui, Y.; Takeda, K.; Takahashi, A. A variety of wheat tolerant to deepseeding conditions: Elongation of the first internode depends on the response to gibberellin and potassium. Plant Cell Environ. 2001, 24, 469–476. [Google Scholar] [CrossRef]
- Zheng, X.; Zhao, Y.; Shan, D.; Shi, K.; Wang, L.; Li, Q.; Wang, N.; Zhou, J.; Yao, J.; Xue, Y.; et al. Mdwrky 9 overexpression confers intersive dwarfing in the M26 rootstock of apple by directly inhibiting brassinosteroid synthetase Mddwf4 expression. New Phytol. 2018, 217, 1086–1098. [Google Scholar] [CrossRef] [Green Version]
- Hibara, K.I.; Karim, M.R.; Takada, S.; Taoka, K.I.; Furutani, M.; Aida, M.; Tasaka, M. Arabidopsis Cup-shaped cotyledon3 regulates postembryonic shoot meristem and organ boundary formation. Plant Cell 2006, 18, 2946–2957. [Google Scholar] [CrossRef] [Green Version]
- Zimmermann, R.; Werr, W. Pattern formation in the monocot embryo as revealed by NAM and CUC3 orthologues from Zea mays L. Plant Mol. Bio. 2005, 58, 669–685. [Google Scholar] [CrossRef]
- Wu, H.; Ren, Z.; Zheng, L.; Guo, M.; Yang, J.; Hou, L.; Qanmber, G.; Li, F.; Yang, Z. The bHLH transcription factor GhPAS1 medi ates BR signaling to regulate plant development and architecture in cotton. Crop J. 2021, 9, 1049–1059. [Google Scholar] [CrossRef]
- Cabib, B.E.; Leloir, L.F. The biosynthesis of trehalose phosphate. J BIOL CHEM. 1958, 259–275. [Google Scholar] [CrossRef]
- Schluepmann, H.; Pellny, T.; Dijken, A.V.; Smeekens, S.; Paul, M. Trehalose 6-phosphate is indispensable for carbohydrate utilization and growth in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2002, 100, 6849–6854. [Google Scholar] [CrossRef] [Green Version]
- Pellny, T.K.; Ghannoum, O.; Conroy, J.P.; Schluepmann, H.; Smeekens, S.; Andralojc, J.; Kranse, K.P.; Goddijn, O.; Paul, M.J. Genetic modification of photosynthesis with E. coli genes for trehalose synthesis. Plant Biotechnol. 2004, 2, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Dijken, A.; Schluepmann, H.; Smeekens, S. Arabidopsis trehalose-6-phosphate synthase is essential for normal vegetative growth and transition to flowering. Plant Physiol. 2004, 135, 969–977. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schluepmann, H.; Dijken, A.V.; Aghdasi, M.; Wobbes, B.; Paul, M.; Smeekens, S. Trehalose mediated growth inhibition of arabidop sis seedlings is due to trehalose-6-phosphate accumulation. Plant Physiol. 2004, 135, 879–890. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, Y.; Chen, J.D.; Fang, L.; Zhang, Z.Y.; Ma, W.; Niu, Y.C.; Ju, L.Z.; Deng, J.Q.; Zhao, T.; Lian, J.; et al. Gossypium barbadense and Gossypium hirsutum genomes provide insights into the origin and evolution of allotetraploid cotton. Nat. Genet. 2019, 51, 739–748. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mortazavi, A.; Williams, B.A.; Mccue, K.; Schaeffer, L. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; Van Baren, M.J.; Salzberg, S.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by rna-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [Green Version]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [Green Version]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Li, Y.; Zhou, C.; Yan, X.; Zhang, J.; Xu, J. Simultaneous analysis of ten phytohormones in Sargassum horneri by high-performance liquid chromatography with electrospray ionization tandem mass spectrometry. J. Sep. Sci. 2016, 39, 1804–1813. [Google Scholar] [CrossRef]
- Floková, K.; Tarkowská, D.; Miersch, O.; Strnad, M.; Wasternack, C.; Novák, O. UHPLC-MS/MS based target profiling of profiling stress-induced phytohormones. Phytochemistry. 2014, 105, 147–157. [Google Scholar] [CrossRef] [PubMed]
- Cai, B.D.; Zhu, J.; Gao, Q.; Luo, D.; Yuan, B.; Feng, Y. Rapid and high-throughput determination of endogenous cytokinins in Oryza sativa by bare Fe3O4 nanoparticles-based magnetic solid-phase extraction. J. Chromatogr. A 2014, 1340, 146–150. [Google Scholar] [CrossRef] [PubMed]
- Niu, Q.; Zong, Y.; Qian, M.; Yang, F.; Teng, Y. Simultaneous quantitative determination of major plant hormones in pear flowers and fruit by UPLC/ESI-MS/MS. Anal. Methods 2014, 6, 1766–1773. [Google Scholar] [CrossRef]
- Pan, X.; Welti, R.; Wang, X.M. Quantitative analysis of major plant hormones in crude plant extracts by high-performance liquid chromatography-mass spectrometry. Nat. Protoc. 2010, 6, 986–992. [Google Scholar] [CrossRef] [PubMed]
- Cui, K.Y.; Lin, Y.Y.; Zhou, X.; Li, S.C.; Liu, H.; Zeng, F.; Zhu, F.; Ouyang, G.F.; Zeng, Z.X. Comparison of sample pretreatment methods for the determination of multiple phytohormones in plant samples by liquid chromatography-electrospray ionization-tandem mass spectrometry. Microchem. J. 2015, 121, 25–31. [Google Scholar] [CrossRef]





Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Z.; Li, Y.; Zhu, Q.; Tian, L.; Liu, F.; Zhang, X.; Sun, J. Transcriptome Profiling Provides New Insights into the Molecular Mechanism Underlying the Sensitivity of Cotton Varieties to Mepiquat Chloride. Int. J. Mol. Sci. 2022, 23, 5043. https://doi.org/10.3390/ijms23095043
Wang Z, Li Y, Zhu Q, Tian L, Liu F, Zhang X, Sun J. Transcriptome Profiling Provides New Insights into the Molecular Mechanism Underlying the Sensitivity of Cotton Varieties to Mepiquat Chloride. International Journal of Molecular Sciences. 2022; 23(9):5043. https://doi.org/10.3390/ijms23095043
Chicago/Turabian StyleWang, Zhijun, Yanjun Li, Qianhao Zhu, Liwen Tian, Feng Liu, Xinyu Zhang, and Jie Sun. 2022. "Transcriptome Profiling Provides New Insights into the Molecular Mechanism Underlying the Sensitivity of Cotton Varieties to Mepiquat Chloride" International Journal of Molecular Sciences 23, no. 9: 5043. https://doi.org/10.3390/ijms23095043
APA StyleWang, Z., Li, Y., Zhu, Q., Tian, L., Liu, F., Zhang, X., & Sun, J. (2022). Transcriptome Profiling Provides New Insights into the Molecular Mechanism Underlying the Sensitivity of Cotton Varieties to Mepiquat Chloride. International Journal of Molecular Sciences, 23(9), 5043. https://doi.org/10.3390/ijms23095043






