Exosomal miRNAs as a Promising Source of Biomarkers in Colorectal Cancer Progression
Abstract
:1. Introduction
2. Colorectal Cancer Incidence and Diagnosis
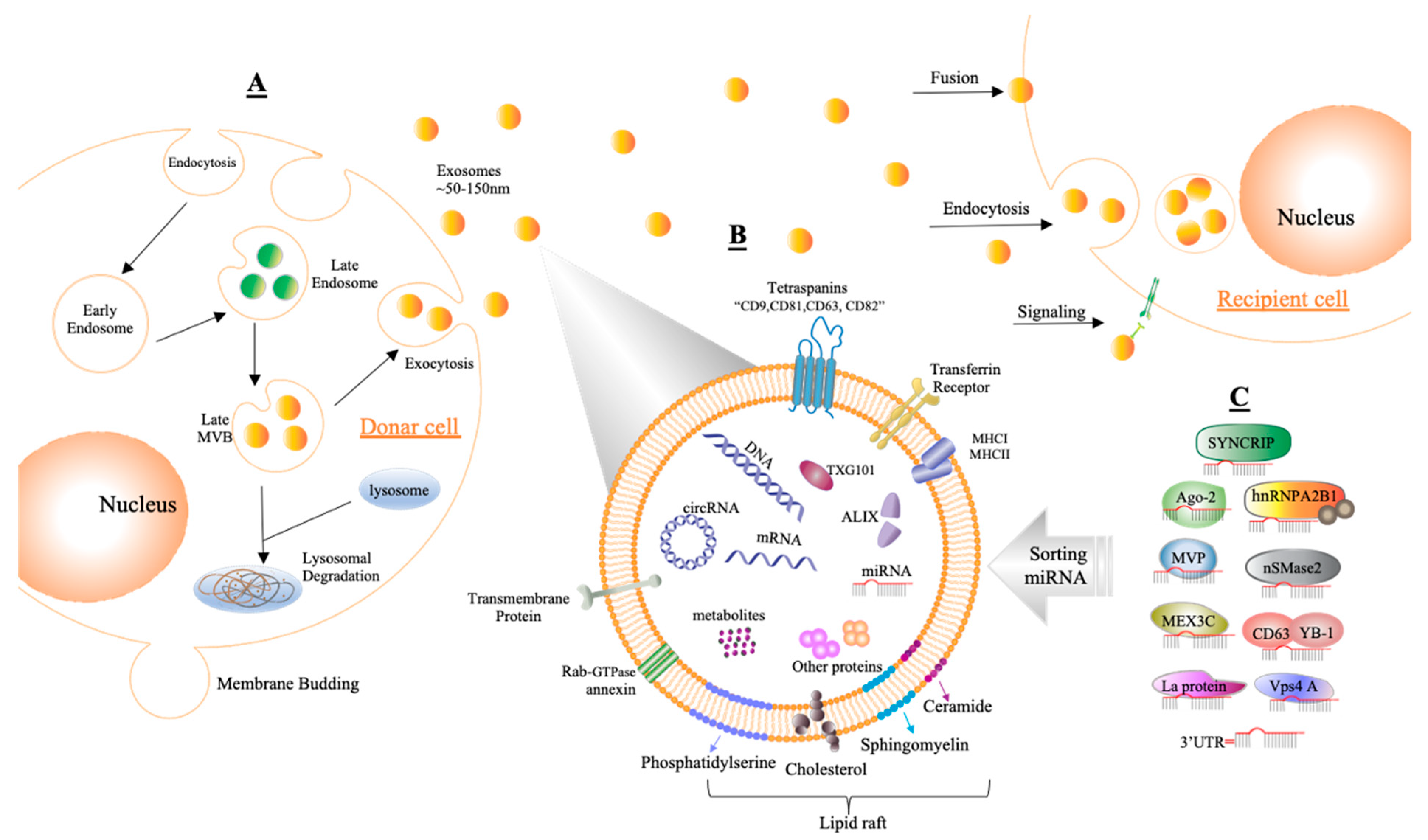
3. Tumor Microenvironment and Exosomes
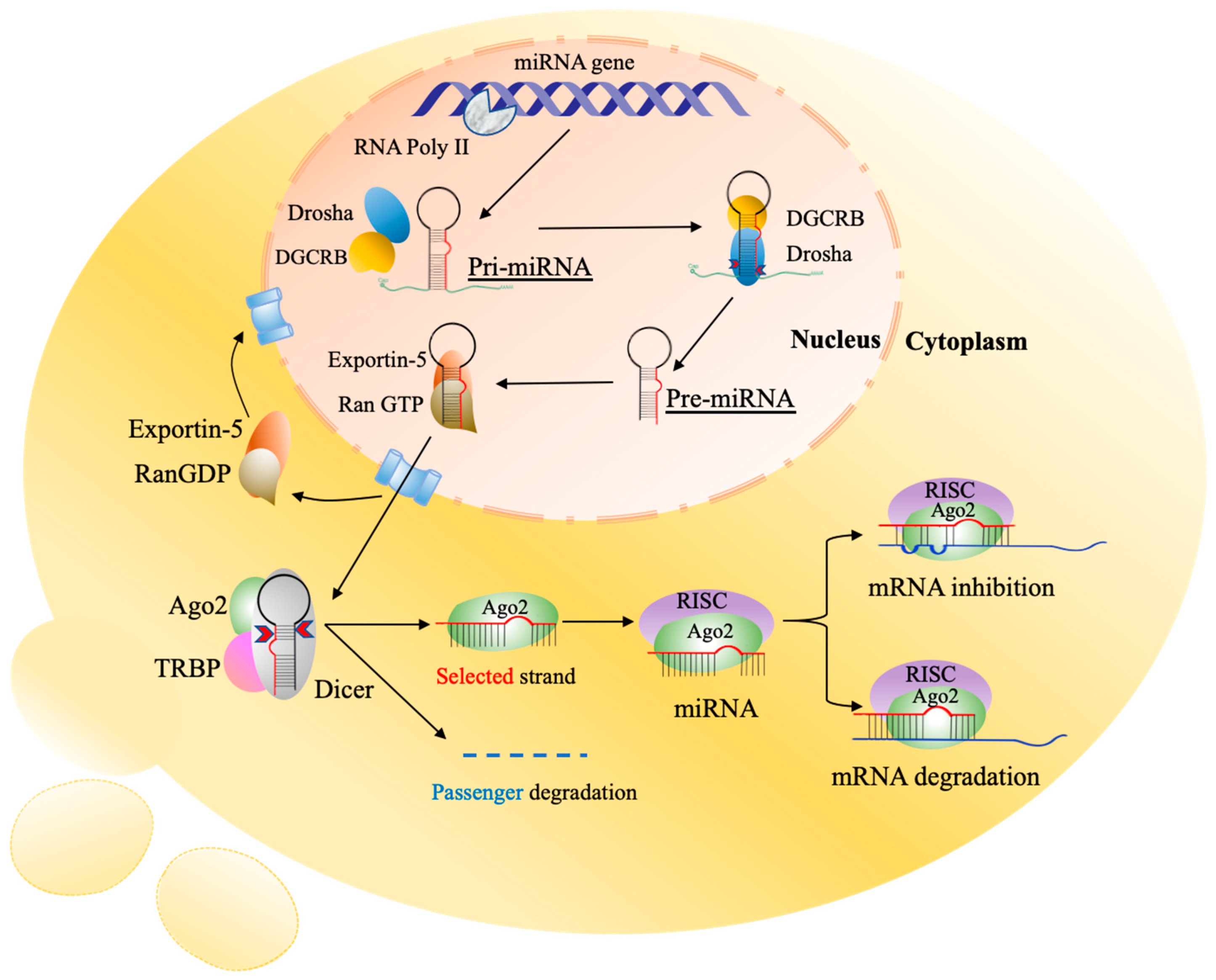
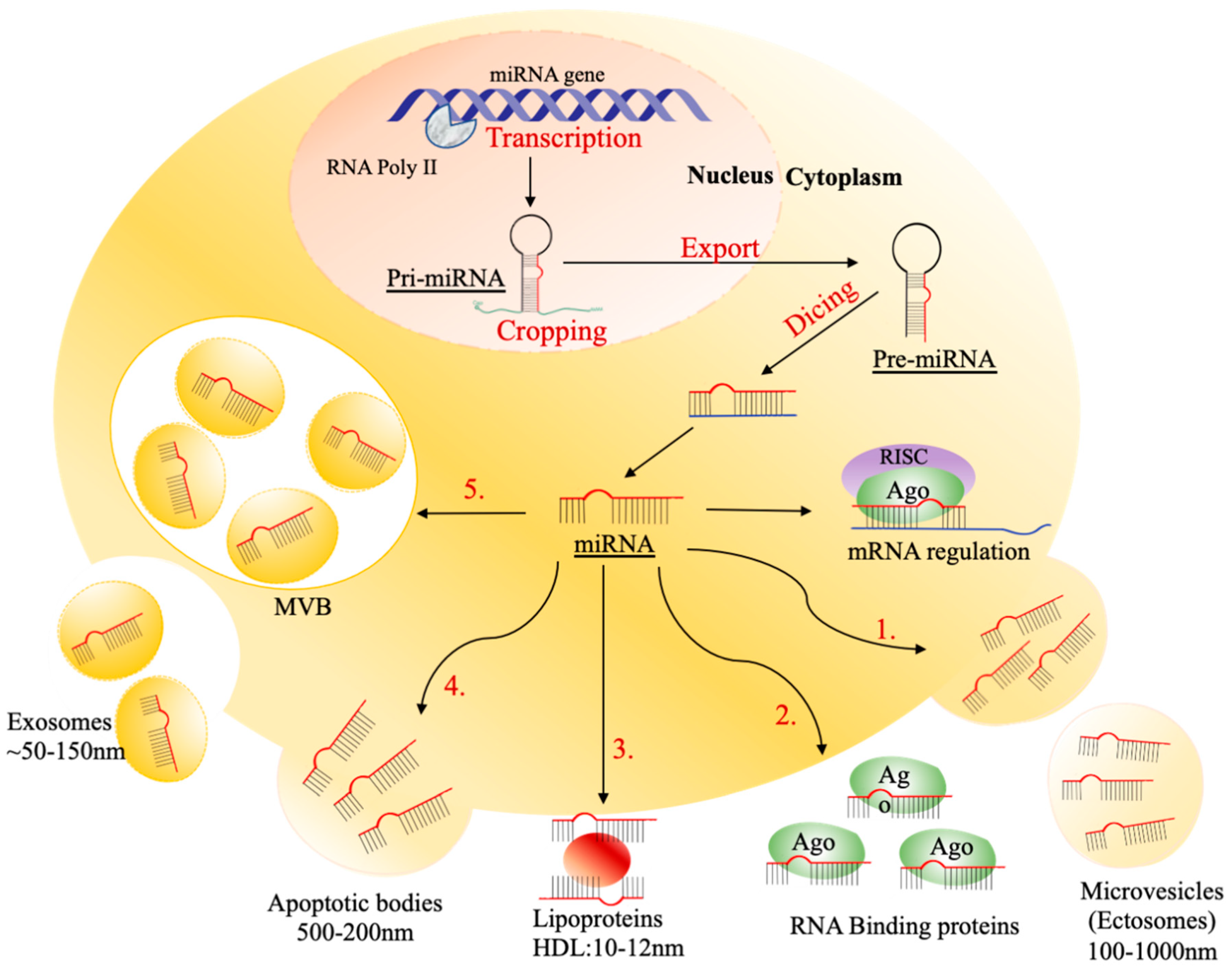
4. MicroRNAs
4.1. Packaging and Sorting miRNAs into Exosomes
4.2. The Role of Exosomal miRNAs in CRC Progression and Drug Resistance
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Ding, L.; Lan, Z.; Xiong, X.; Ao, H.; Feng, Y.; Gu, H.; Yu, M.; Cui, Q. The Dual Role of MicroRNAs in Colorectal Cancer Progression. Int. J. Mol. Sci. 2018, 19, 2791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Augestad, K.; Bakaki, P.; Rose, J.; Crawshaw, B.; Lindsetmo, R.; Dørum, L.; Koroukian, S.; Delaney, C. Metastatic spread pattern after curative colorectal cancer surgery. A retrospective, longitudinal analysis. Cancer Epidemiol. 2015, 39, 734–744. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Toiyama, Y.; Okugawa, Y.; Fleshman, J.; Boland, C.R.; Goel, A. MicroRNAs as potential liquid biopsy biomarkers in colorectal cancer: A systematic review. Biochim. Biophys. Acta Rev. Cancer 2018, 1870, 274–282. [Google Scholar] [CrossRef] [PubMed]
- Swiderska, M.; Choromańska, B.; Dąbrowska, E.; Konarzewska-Duchnowska, E.; Choromańska, K.; Szczurko, G.; Myśliwiec, P.; Dadan, J.; Ladny, J.R.; Zwierz, K. The diagnostics of colorectal cancer. Contemp. Oncol. 2014, 18, 1–6. [Google Scholar] [CrossRef]
- Tan, S.; Xia, L.; Yi, P.; Han, Y.; Tang, L.; Pan, Q.; Tian, Y.; Rao, S.; Oyang, L.; Liang, J. Exosomal miRNAs in tumor microenvironment. J. Exp. Clin. Cancer Res. 2020, 39, 67. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.; Deng, C.-X. Current progresses of exosomes as cancer diagnostic and prognostic biomarkers. Int. J. Biol. Sci. 2019, 15, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Ingenito, F.; Roscigno, G.; Affinito, A.; Nuzzo, S.; Scognamiglio, I.; Quintavalle, C.; Condorelli, G. The role of exo-miRNAs in cancer: A focus on therapeutic and diagnostic applications. Int. J. Mol. Sci. 2019, 20, 4687. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alzahrani, F.A.; El-Magd, M.A.; Abdelfattah-Hassan, A.; Saleh, A.A.; Saadeldin, I.M.; El-Shetry, E.S.; Badawy, A.A.; Alkarim, S. Potential Effect of Exosomes Derived from Cancer Stem Cells and MSCs on Progression of DEN-Induced HCC in Rats. Stem Cells Int. 2018, 2018, 8058979. [Google Scholar] [CrossRef] [Green Version]
- Badawy, A.A.; El-Magd, M.A.; AlSadrah, S.A. Therapeutic Effect of Camel Milk and Its Exosomes on MCF7 Cells In Vitro and In Vivo. Integr. Cancer Ther. 2018, 7, 1235–1246. [Google Scholar] [CrossRef] [Green Version]
- Yang, E.; Wang, X.; Gong, Z.; Yu, M.; Wu, H.; Zhang, D. Exosome-mediated metabolic reprogramming: The emerging role in tumor microenvironment remodeling and its influence on cancer progression. Signal Transduct. Target. Ther. 2020, 5, 242. [Google Scholar] [CrossRef] [PubMed]
- Vaidya, M.; Sugaya, K. Differential sequences and single nucleotide polymorphism of exosomal SOX2 DNA in cancer. PLoS ONE 2020, 15, e0229309. [Google Scholar] [CrossRef] [PubMed]
- Dai, J.; Su, Y.; Zhong, S.; Cong, L.; Liu, B.; Yang, J.; Tao, Y.; He, Z.; Chen, C.; Jiang, Y. Exosomes: Key players in cancer and potential therapeutic strategy. Signal Transduct. Target. Ther. 2020, 5, 145. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Luo, Y.; He, W.; Zhao, Y.; Kong, Y.; Liu, H.; Zhong, G.; Li, Y.; Li, J.; Huang, J. Exosomal long noncoding RNA LNMAT2 promotes lymphatic metastasis in bladder cancer. J. Clin. Investig. 2020, 130, 404–421. [Google Scholar] [CrossRef]
- WHO. 2020. Available online: http://www.who.int/cancer/en/ (accessed on 3 April 2020).
- Zhu, M.; Huang, Z.; Zhu, D.; Zhou, X.; Shan, X.; Qi, L.-W.; Wu, L.; Cheng, W.; Zhu, J.; Zhang, L. A panel of microRNA signature in serum for colorectal cancer diagnosis. Oncotarget 2017, 8, 17081. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.G.; Gu, J. Serum microRNA-29a is a promising novel marker for early detection of colorectal liver metastasis. Cancer Epidemiol. 2012, 36, e61–e67. [Google Scholar] [CrossRef] [PubMed]
- Alomar, S.Y.; Mansour, L.; Abuderman, A.; Alkhuriji, A.; Arafah, M.; Alwasel, S.; Harrath, A.H.; Almutairi, M.; Trayhyrn, P.; Dar, J.A. β-Catenin accumulation and S33F mutation of CTNNB1 gene in colorectal cancer in Saudi Arabia. Pol. J. Pathol. 2016, 67, 156–162. [Google Scholar] [CrossRef] [Green Version]
- Strubberg, A.M.; Madison, B.B. MicroRNAs in the etiology of colorectal cancer: Pathways and clinical implications. Dis. Models Mech. 2017, 10, 197–214. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zanutto, S.; Ciniselli, C.M.; Belfiore, A.; Lecchi, M.; Masci, E.; Delconte, G.; Primignani, M.; Tosetti, G.; Dal Fante, M.; Fazzini, L. Plasma miRNA-based signatures in CRC screening programs. Int. J. Cancer 2019, 146, 1164–1173. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alsanea, N.; Almadi, M.A.; Abduljabbar, A.S.; Alhomoud, S.; Alshaban, T.A.; Alsuhaibani, A.; Alzahrani, A.; Batwa, F.; Hassan, A.-H.; Hibbert, D. National Guidelines for Colorectal Cancer Screening in Saudi Arabia with strength of recommendations and quality of evidence: Tripartite Task Force from Saudi Society of Colon & Rectal Surgery, Saudi Gastroenterology Association and Saudi Oncology Society. Ann. Saudi Med. 2015, 35, 189–195. [Google Scholar]
- Kim, N.H.; Lee, M.Y.; Park, J.H.; Park, D.I.; Sohn, C.I.; Choi, K.; Jung, Y.S. Serum CEA and CA 19-9 levels are associated with the presence and severity of colorectal neoplasia. Yonsei Med. J. 2017, 58, 918–924. [Google Scholar] [CrossRef]
- Vega, P.; Valentín, F.; Cubiella, J. Colorectal cancer diagnosis: Pitfalls and opportunities. World J. Gastrointest. Oncol. 2015, 7, 422. [Google Scholar] [CrossRef] [Green Version]
- Chen, B.; Xia, Z.; Deng, Y.N.; Yang, Y.; Zhang, P.; Zhu, H.; Xu, N.; Liang, S. Emerging microRNA biomarkers for colorectal cancer diagnosis and prognosis. Open Biol. 2019, 9, 180212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hassan, C.; Rossi, P.G.; Camilloni, L.; Rex, D.; Jimenez-Cendales, B.; Ferroni, E.; Borgia, P.; Zullo, A.; Guasticchi, G.; Group, H. Meta-analysis: Adherence to colorectal cancer screening and the detection rate for advanced neoplasia, according to the type of screening test. Aliment. Pharmacol. Ther. 2012, 36, 929–940. [Google Scholar] [CrossRef]
- Levin, B.; Lieberman, D.A.; McFarland, B.; Smith, R.A.; Brooks, D.; Andrews, K.S.; Dash, C.; Giardiello, F.M.; Glick, S.; Levin, T.R. Screening and surveillance for the early detection of colorectal cancer and adenomatous polyps, 2008: A joint guideline from the American Cancer Society, the US Multi-Society Task Force on Colorectal Cancer, and the American College of Radiology. CA Cancer J. Clin. 2008, 58, 130–160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mousavinezhad, M.; Majdzadeh, R.; Sari, A.A.; Delavari, A.; Mohtasham, F. The effectiveness of FOBT vs. FIT: A meta-analysis on colorectal cancer screening test. Med. J. Islamic Repub. Iran 2016, 30, 366. [Google Scholar]
- Obuch, J.C.; Ahnen, D.J. Colorectal cancer: Genetics is changing everything. Gastroenterol. Clin. 2016, 45, 459–476. [Google Scholar] [CrossRef] [PubMed]
- LeGolvan, M.P.; Taliano, R.J.; Resnick, M.B. Application of molecular techniques in the diagnosis, prognosis and management of patients with colorectal cancer: A practical approach. Hum. Pathol. 2012, 43, 1157–1168. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Peng, R.; Wang, J.; Qin, Z.; Xue, L. Circulating microRNAs as potential cancer biomarkers: The advantage and disadvantage. Clin. Epigenet. 2018, 10, 59. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zahran, R.; Ghozy, A.; Elkholy, S.S.; El-Taweel, F.; El-Magd, M.A. Combination therapy with melatonin, stem cells and extracellular vesicles is effective in limiting renal ischemia–reperfusion injury in a rat model. Int. J. Urol. 2020, 27, 1039–1049. [Google Scholar] [CrossRef]
- Su, C.; Zhang, J.; Yarden, Y.; Fu, L. The key roles of cancer stem cell-derived extracellular vesicles. Signal Transduct. Target. Ther. 2021, 6, 109. [Google Scholar] [CrossRef]
- López de Andrés, J.; Griñán-Lisón, C.; Jiménez, G.; Marchal, J.A. Cancer stem cell secretome in the tumor microenvironment: A key point for an effective personalized cancer treatment. J. Hematol. Oncol. 2020, 13, 136. [Google Scholar] [CrossRef] [PubMed]
- Martellucci, S.; Orefice, N.S.; Angelucci, A.; Luce, A.; Caraglia, M.; Zappavigna, S. Extracellular Vesicles: New Endogenous Shuttles for miRNAs in Cancer Diagnosis and Therapy? Int. J. Mol. Sci. 2020, 21, 6486. [Google Scholar] [CrossRef]
- Doyle, L.M.; Wang, M.Z. Overview of extracellular vesicles, their origin, composition, purpose, and methods for exosome isolation and analysis. Cells 2019, 8, 727. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hessvik, N.P.; Llorente, A. Current knowledge on exosome biogenesis and release. Cell. Mol. Life Sci. 2018, 75, 193–208. [Google Scholar] [CrossRef] [Green Version]
- Vautrot, V.; Chanteloup, G.; Elmallah, M.; Cordonnier, M.; Aubin, F.; Garrido, C.; Gobbo, J. Exosomal miRNA: Small molecules, big impact in colorectal cancer. J. Oncol. 2019, 2019, 8585276. [Google Scholar] [CrossRef] [PubMed]
- Lv, L.-H.; Wan, Y.-L.; Lin, Y.; Zhang, W.; Yang, M.; Li, G.-L.; Lin, H.-M.; Shang, C.-Z.; Chen, Y.-J.; Min, J. Anticancer drugs cause release of exosomes with heat shock proteins from human hepatocellular carcinoma cells that elicit effective natural killer cell antitumor responses in vitro. J. Biol. Chem. 2012, 287, 15874–15885. [Google Scholar] [CrossRef] [Green Version]
- Kalra, H.; Drummen, G.P.; Mathivanan, S. Focus on extracellular vesicles: Introducing the next small big thing. Int. J. Mol. Sci. 2016, 17, 170. [Google Scholar] [CrossRef] [Green Version]
- Janas, T.; Janas, P.; Sapoń, K.; Janas, T. Binding of RNA Aptamers to Membrane Lipid Rafts: Implications for Exosomal miRNAs Transfer from Cancer to Immune Cells. Int. J. Mol. Sci. 2020, 21, 8503. [Google Scholar] [CrossRef]
- Badawy, A.A.; Othman, R.Q.A.; El-Magd, M.A. Effect of combined therapy with camel milk-derived exosomes, tamoxifen, and hesperidin on breast cancer. Mol. Cell. Toxicol. 2021. [Google Scholar] [CrossRef]
- Ibrahim, H.M.; Mohammed-Geba, K.; Tawfic, A.A.; El-Magd, M.A. Camel milk exosomes modulate cyclophosphamide-induced oxidative stress and immuno-toxicity in rats. Food Funct. 2019, 10, 7523–7532. [Google Scholar] [CrossRef]
- Xiao, Y.; Zhong, J.; Zhong, B.; Huang, J.; Jiang, L.; Jiang, Y.; Yuan, J.; Sun, J.; Dai, L.; Yang, C. Exosomes as potential sources of biomarkers in colorectal cancer. Cancer Lett. 2020, 476, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Gerloff, D.; Lützkendorf, J.; Moritz, R.K.; Wersig, T.; Mäder, K.; Müller, L.P.; Sunderkötter, C. Melanoma-derived exosomal miR-125b-5p educates tumor associated macrophages (TAMs) by targeting lysosomal acid lipase A (LIPA). Cancers 2020, 12, 464. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Daassi, D.; Mahoney, K.M.; Freeman, G.J. The importance of exosomal PDL1 in tumour immune evasion. Nat. Rev. Immunol. 2020, 20, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Matsumura, T.; Sugimachi, K.; Iinuma, H.; Takahashi, Y.; Kurashige, J.; Sawada, G.; Ueda, M.; Uchi, R.; Ueo, H.; Takano, Y. Exosomal microRNA in serum is a novel biomarker of recurrence in human colorectal cancer. Br. J. Cancer 2015, 113, 275–281. [Google Scholar] [CrossRef] [PubMed]
- Ogata-Kawata, H.; Izumiya, M.; Kurioka, D.; Honma, Y.; Yamada, Y.; Furuta, K.; Gunji, T.; Ohta, H.; Okamoto, H.; Sonoda, H. Circulating exosomal microRNAs as biomarkers of colon cancer. PLoS ONE 2014, 9, e92921. [Google Scholar]
- Francavilla, A.; Turoczi, S.; Tarallo, S.; Vodicka, P.; Pardini, B.; Naccarati, A. Exosomal microRNAs and other non-coding RNAs as colorectal cancer biomarkers: A review. Mutagenesis 2020, 35, 243–260. [Google Scholar] [CrossRef]
- Plotnikova, O.; Baranova, A.; Skoblov, M. Comprehensive analysis of human microRNA–mRNA interactome. Front. Genet. 2019, 10, 933. [Google Scholar] [CrossRef]
- Badawy, A.A.; El-Magd, M.A.; AlSadrah, S.A.; Alruwaili, M.M. Altered expression of some miRNAs and their target genes following mesenchymal stem cell treatment in busulfan-induced azoospermic rats. Gene 2020, 737, 144481. [Google Scholar] [CrossRef]
- Hunter, S.; Nault, B.; Ugwuagbo, K.C.; Maiti, S.; Majumder, M. Mir526b and Mir655 promote tumour associated angiogenesis and lymphangiogenesis in breast cancer. Cancers 2019, 11, 938. [Google Scholar] [CrossRef] [Green Version]
- Baroni, S.; Romero-Cordoba, S.; Plantamura, I.; Dugo, M.; D’ippolito, E.; Cataldo, A.; Cosentino, G.; Angeloni, V.; Rossini, A.; Daidone, M. Exosome-mediated delivery of miR-9 induces cancer-associated fibroblast-like properties in human breast fibroblasts. Cell Death Dis. 2016, 7, e2312. [Google Scholar] [CrossRef]
- Dilsiz, N. Role of exosomes and exosomal microRNAs in cancer. Future Sci. OA 2020, 6, FSO465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ostenfeld, M.S.; Jeppesen, D.K.; Laurberg, J.R.; Boysen, A.T.; Bramsen, J.B.; Primdal-Bengtson, B.; Hendrix, A.; Lamy, P.; Dagnaes-Hansen, F.; Rasmussen, M.H. Cellular disposal of miR23b by RAB27-dependent exosome release is linked to acquisition of metastatic properties. Cancer Res. 2014, 74, 5758–5771. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Q.; Huang, Z.; Ni, S.; Xiao, X.; Xu, Q.; Wang, L.; Huang, D.; Tan, C.; Sheng, W.; Du, X. Plasma miR-601 and miR-760 are novel biomarkers for the early detection of colorectal cancer. PLoS ONE 2012, 7, e44398. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Formentini, A.; Chien, M.; Weir, D.B.; Russo, J.J.; Ju, J.; Kornmann, M.; Ju, J. Prognostic values of microRNAs in colorectal cancer. Biomark. Insights 2006, 1, 117727190600100009. [Google Scholar] [CrossRef] [Green Version]
- Sohel, M.H. Extracellular/circulating microRNAs: Release mechanisms, functions and challenges. Achiev. Life Sci. 2016, 10, 175–186. [Google Scholar] [CrossRef] [Green Version]
- Nedaeinia, R.; Manian, M.; Jazayeri, M.; Ranjbar, M.; Salehi, R.; Sharifi, M.; Mohaghegh, F.; Goli, M.; Jahednia, S.; Avan, A. Circulating exosomes and exosomal microRNAs as biomarkers in gastrointestinal cancer. Cancer Gene Ther. 2017, 24, 48–56. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Vijayan, M.; Bhatti, J.; Reddy, P.H. MicroRNAs as peripheral biomarkers in aging and age-related diseases. Prog. Mol. Biol. Transl. Sci. 2017, 146, 47–94. [Google Scholar]
- Igaz, P. Circulating Micrornas in Disease Diagnostics and Their Potential Biological Relevance; Springer: Basel, Switzerland, 2015; Volume 106. [Google Scholar]
- Koga, Y.; Yasunaga, M.; Moriya, Y.; Akasu, T.; Fujita, S.; Yamamoto, S.; Matsumura, Y. Exosome can prevent RNase from degrading microRNA in feces. J. Gastrointest. Oncol. 2011, 2, 215. [Google Scholar]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [Green Version]
- Shigeyasu, K.; Toden, S.; Zumwalt, T.J.; Okugawa, Y.; Goel, A. Emerging role of microRNAs as liquid biopsy biomarkers in gastrointestinal cancers. Clin. Cancer Res. 2017, 23, 2391–2399. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, H.; Wang, N.; Yi, X.; Tang, C.; Wang, D. Low-level serum miR-24-2 is associated with the progression of colorectal cancer. Cancer Biomark. 2018, 21, 261–267. [Google Scholar] [CrossRef] [PubMed]
- Ya, G.; Wang, H.; Ma, Y.; Hu, A.; Ma, Y.; Hu, J.; Yu, Y. Serum miR-129 functions as a biomarker for colorectal cancer by targeting estrogen receptor (ER) β. Die Pharm. Int. J. Pharm. Sci. 2017, 72, 107–112. [Google Scholar]
- Hur, K.; Toiyama, Y.; Okugawa, Y.; Ide, S.; Imaoka, H.; Boland, C.R.; Goel, A. Circulating microRNA-203 predicts prognosis and metastasis in human colorectal cancer. Gut 2017, 66, 654–665. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ng, L.; Wan, T.M.-H.; Man, J.H.-W.; Chow, A.K.-M.; Iyer, D.; Chen, G.; Yau, T.C.-C.; Lo, O.S.-H.; Foo, D.C.-C.; Poon, J.T.-C. Identification of serum miR-139-3p as a non-invasive biomarker for colorectal cancer. Oncotarget 2017, 8, 27393. [Google Scholar] [CrossRef]
- Huang, X.; Yuan, T.; Tschannen, M.; Sun, Z.; Jacob, H.; Du, M.; Liang, M.; Dittmar, R.L.; Liu, Y.; Liang, M. Characterization of human plasma-derived exosomal RNAs by deep sequencing. BMC Genom. 2013, 14, 319. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Temoche-Diaz, M.M.; Shurtleff, M.J.; Nottingham, R.M.; Yao, J.; Fadadu, R.P.; Lambowitz, A.M.; Schekman, R. Distinct mechanisms of microRNA sorting into cancer cell-derived extracellular vesicle subtypes. eLife 2019, 8, e47544. [Google Scholar] [CrossRef]
- Wei, H.; Chen, Q.; Lin, L.; Sha, C.; Li, T.; Liu, Y.; Yin, X.; Xu, Y.; Chen, L.; Gao, W. Regulation of exosome production and cargo sorting. Int. J. Biol. Sci. 2021, 17, 163. [Google Scholar] [CrossRef]
- McKenzie, A.J.; Hoshino, D.; Hong, N.H.; Cha, D.J.; Franklin, J.L.; Coffey, R.J.; Patton, J.G.; Weaver, A.M. KRAS-MEK signaling controls Ago2 sorting into exosomes. Cell Rep. 2016, 15, 978–987. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guduric-Fuchs, J.; O’Connor, A.; Camp, B.; O’Neill, C.L.; Medina, R.J.; Simpson, D.A. Selective extracellular vesicle-mediated export of an overlapping set of microRNAs from multiple cell types. BMC Genom. 2012, 13, 357. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Deng, T.; Liu, R.; Ning, T.; Yang, H.; Liu, D.; Zhang, Q.; Lin, D.; Ge, S.; Bai, M. CAF secreted miR-522 suppresses ferroptosis and promotes acquired chemo-resistance in gastric cancer. Mol. Cancer 2020, 19, 43. [Google Scholar] [CrossRef] [Green Version]
- Gao, T.; Shu, J.; Cui, J. A systematic approach to RNA-associated motif discovery. BMC Genom. 2018, 19, 146. [Google Scholar] [CrossRef] [Green Version]
- Xu, Y.-F.; Hannafon, B.N.; Khatri, U.; Gin, A.; Ding, W.-Q. The origin of exosomal miR-1246 in human cancer cells. RNA Biol. 2019, 16, 770–784. [Google Scholar] [CrossRef]
- Villarroya-Beltri, C.; Gutiérrez-Vázquez, C.; Sánchez-Cabo, F.; Pérez-Hernández, D.; Vázquez, J.; Martin-Cofreces, N.; Martinez-Herrera, D.J.; Pascual-Montano, A.; Mittelbrunn, M.; Sánchez-Madrid, F. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat. Commun. 2013, 4, 2980. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tietje, A.; Maron, K.N.; Wei, Y.; Feliciano, D.M. Cerebrospinal fluid extracellular vesicles undergo age dependent declines and contain known and novel non-coding RNAs. PLoS ONE 2014, 9, e113116. [Google Scholar] [CrossRef] [PubMed]
- Santangelo, L.; Giurato, G.; Cicchini, C.; Montaldo, C.; Mancone, C.; Tarallo, R.; Battistelli, C.; Alonzi, T.; Weisz, A.; Tripodi, M. The RNA-binding protein SYNCRIP is a component of the hepatocyte exosomal machinery controlling microRNA sorting. Cell Rep. 2016, 17, 799–808. [Google Scholar] [CrossRef] [Green Version]
- Hobor, F.; Dallmann, A.; Ball, N.J.; Cicchini, C.; Battistelli, C.; Ogrodowicz, R.W.; Christodoulou, E.; Martin, S.R.; Castello, A.; Tripodi, M. A cryptic RNA-binding domain mediates Syncrip recognition and exosomal partitioning of miRNA targets. Nat. Commun. 2018, 9, 831. [Google Scholar] [CrossRef]
- Cooks, T.; Pateras, I.S.; Jenkins, L.M.; Patel, K.M.; Robles, A.I.; Morris, J.; Forshew, T.; Appella, E.; Gorgoulis, V.G.; Harris, C.C. Mutant p53 cancers reprogram macrophages to tumor supporting macrophages via exosomal miR-1246. Nat. Commun. 2018, 9, 771. [Google Scholar] [CrossRef] [Green Version]
- Mittelbrunn, M.; Gutiérrez-Vázquez, C.; Villarroya-Beltri, C.; González, S.; Sánchez-Cabo, F.; González, M.Á.; Bernad, A.; Sánchez-Madrid, F. Unidirectional transfer of microRNA-loaded exosomes from T cells to antigen-presenting cells. Nat. Commun. 2011, 2, 282. [Google Scholar] [CrossRef] [Green Version]
- Kosaka, N.; Iguchi, H.; Hagiwara, K.; Yoshioka, Y.; Takeshita, F.; Ochiya, T. Neutral sphingomyelinase 2 (nSMase2)-dependent exosomal transfer of angiogenic microRNAs regulate cancer cell metastasis. J. Biol. Chem. 2013, 288, 10849–10859. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kosaka, N.; Iguchi, H.; Yoshioka, Y.; Takeshita, F.; Matsuki, Y.; Ochiya, T. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J. Biol. Chem. 2010, 285, 17442–17452. [Google Scholar] [CrossRef] [Green Version]
- Wei, J.x.; Lv, L.h.; Wan, Y.l.; Cao, Y.; Li, G.l.; Lin, H.m.; Zhou, R.; Shang, C.z.; Cao, J.; He, H. Vps4A functions as a tumor suppressor by regulating the secretion and uptake of exosomal microRNAs in human hepatoma cells. Hepatology 2015, 61, 1284–1294. [Google Scholar] [CrossRef] [PubMed]
- Lin, F.; Zeng, Z.; Song, Y.; Li, L.; Wu, Z.; Zhang, X.; Li, Z.; Ke, X.; Hu, X. YBX-1 mediated sorting of miR-133 into hypoxia/reoxygenation-induced EPC-derived exosomes to increase fibroblast angiogenesis and MEndoT. Stem Cell Res. Ther. 2019, 10, 263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shurtleff, M.J.; Temoche-Diaz, M.M.; Karfilis, K.V.; Ri, S.; Schekman, R. Y-box protein 1 is required to sort microRNAs into exosomes in cells and in a cell-free reaction. eLife 2016, 5, e19276. [Google Scholar] [CrossRef] [PubMed]
- Lu, P.; Li, H.; Li, N.; Singh, R.N.; Bishop, C.E.; Chen, X.; Lu, B. MEX3C interacts with adaptor-related protein complex 2 and involves in miR-451a exosomal sorting. PLoS ONE 2017, 12, e0185992. [Google Scholar] [CrossRef] [Green Version]
- Dickman, C.T.; Lawson, J.; Jabalee, J.; MacLellan, S.A.; LePard, N.E.; Bennewith, K.L.; Garnis, C. Selective extracellular vesicle exclusion of miR-142-3p by oral cancer cells promotes both internal and extracellular malignant phenotypes. Oncotarget 2017, 8, 15252. [Google Scholar] [CrossRef] [Green Version]
- Teng, Y.; Ren, Y.; Hu, X.; Mu, J.; Samykutty, A.; Zhuang, X.; Deng, Z.; Kumar, A.; Zhang, L.; Merchant, M.L. MVP-mediated exosomal sorting of miR-193a promotes colon cancer progression. Nat. Commun. 2017, 8, 14448. [Google Scholar] [CrossRef]
- Janas, T.; Janas, M.M.; Sapoń, K.; Janas, T. Mechanisms of RNA loading into exosomes. FEBS Lett. 2015, 589, 1391–1398. [Google Scholar] [CrossRef] [Green Version]
- Momose, F.; Seo, N.; Akahori, Y.; Sawada, S.-I.; Harada, N.; Ogura, T.; Akiyoshi, K.; Shiku, H. Guanine-rich sequences are a dominant feature of exosomal microRNAs across the mammalian species and cell types. PLoS ONE 2016, 11, e0154134. [Google Scholar] [CrossRef]
- Bolukbasi, M.F.; Mizrak, A.; Ozdener, G.B.; Madlener, S.; Ströbel, T.; Erkan, E.P.; Fan, J.-B.; Breakefield, X.O.; Saydam, O. miR-1289 and “Zipcode”-like sequence enrich mRNAs in microvesicles. Mol. Ther.-Nucleic Acids 2012, 1, e10. [Google Scholar] [CrossRef]
- Squadrito, M.L.; Baer, C.; Burdet, F.; Maderna, C.; Gilfillan, G.D.; Lyle, R.; Ibberson, M.; De Palma, M. Endogenous RNAs modulate microRNA sorting to exosomes and transfer to acceptor cells. Cell Rep. 2014, 8, 1432–1446. [Google Scholar] [CrossRef] [Green Version]
- Cha, D.J.; Franklin, J.L.; Dou, Y.; Liu, Q.; Higginbotham, J.N.; Beckler, M.D.; Weaver, A.M.; Vickers, K.; Prasad, N.; Levy, S. KRAS-dependent sorting of miRNA to exosomes. eLife 2015, 4, e07197. [Google Scholar] [CrossRef] [PubMed]
- Jin, W.; Shi, J.; Liu, M. Overexpression of miR-671-5p indicates a poor prognosis in colon cancer and accelerates proliferation, migration, and invasion of colon cancer cells. Onco Targets 2019, 12, 6865–6873. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, J.; Zhao, J.; Zhang, R. Prognostic significance of serum miR-193b in colorectal cancer. Int. J. Clin. Exp. Pathol. 2017, 10, 9509–9514. [Google Scholar]
- Li, J.; Peng, W.; Yang, P.; Chen, R.; Gu, Q.; Qian, W.; Ji, D.; Wang, Q.; Zhang, Z.; Tang, J.; et al. MicroRNA-1224-5p Inhibits Metastasis and Epithelial-Mesenchymal Transition in Colorectal Cancer by Targeting SP1-Mediated NF-κB Signaling Pathways. Front. Oncol. 2020, 10, 294. [Google Scholar] [CrossRef] [Green Version]
- Fujino, Y.; Takeishi, S.; Nishida, K.; Okamoto, K.; Muguruma, N.; Kimura, T.; Kitamura, S.; Miyamoto, H.; Fujimoto, A.; Higashijima, J.; et al. Downregulation of microRNA-100/microRNA-125b is associated with lymph node metastasis in early colorectal cancer with submucosal invasion. Cancer Sci. 2017, 108, 390–397. [Google Scholar] [CrossRef] [PubMed]
- Danac, J.M.C.; Uy, A.G.G.; Garcia, R.L. Exosomal microRNAs in colorectal cancer: Overcoming barriers of the metastatic cascade (Review). Int. J. Mol. Med. 2021, 47, 112. [Google Scholar] [CrossRef]
- Niu, Z.-Y.; Li, W.-L.; Jiang, D.-L.; Li, Y.-S.; Xie, X.-J. Mir-483 inhibits colon cancer cell proliferation and migration by targeting TRAF1. Kaohsiung J. Med. Sci. 2018, 34, 479–486. [Google Scholar] [CrossRef]
- Zhu, X.; Luo, X.; Song, Z.; Jiang, S.; Long, X.; Gao, X.; Xie, X.; Zheng, L.; Wang, H. miR-188-5p promotes oxaliplatin resistance by targeting RASA1 in colon cancer cells. Oncol. Lett. 2021, 21, 481. [Google Scholar] [CrossRef]
- Yu, Y.; Lu, X.; Yang, C.; Yin, F. Long Noncoding RNA LINC00173 Contributes to the Growth, Invasiveness and Chemo-Resistance of Colorectal Cancer Through Regulating miR-765/PLP2 Axis. Cancer Manag. Res. 2020, 12, 3363–3369. [Google Scholar] [CrossRef]
- Zhang, J.; Fei, B.; Wang, Q.; Song, M.; Yin, Y.; Zhang, B.; Ni, S.; Guo, W.; Bian, Z.; Quan, C.; et al. MicroRNA-638 inhibits cell proliferation, invasion and regulates cell cycle by targeting tetraspanin 1 in human colorectal carcinoma. Oncotarget 2014, 5, 12083–12096. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, P.; Cai, J.-X.; Han, F.; Wang, J.; Zhou, J.-J.; Shen, K.-W.; Wang, L.-H. Expression and significance of miR-654-5p and miR-376b-3p in patients with colon cancer. World J. Gastrointest. Oncol. 2020, 12, 492–502. [Google Scholar] [CrossRef]
- Lai, H.; Zhang, J.; Zuo, H.; Liu, H.; Xu, J.; Feng, Y.; Lin, Y.; Mo, X. Overexpression of miR-17 is correlated with liver metastasis in colorectal cancer. Medicine 2020, 99, e19265. [Google Scholar] [CrossRef]
- Yang, J.; Zhao, H.; Xin, Y.; Fan, L. MicroRNA-198 Inhibits Proliferation and Induces Apoptosis of Lung Cancer Cells Via Targeting FGFR1. J. Cell. Biochem. 2014, 115, 987–995. [Google Scholar] [CrossRef]
- Cui, F.C.; Chen, Y.; Wu, X.Y.; Hu, M.; Qin, W.S. MicroRNA-493-5p suppresses colorectal cancer progression via the PI3K-Akt-FoxO3a signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 4212–4223. [Google Scholar] [CrossRef]
- Chai, B.; Guo, Y.; Cui, X.; Liu, J.; Suo, Y.; Dou, Z.; Li, N. MiR-223-3p promotes the proliferation, invasion and migration of colon cancer cells by negative regulating PRDM1. Am. J. Transl. Res. 2019, 11, 4516–4523. [Google Scholar] [PubMed]
- Vishnubalaji, R.; Hamam, R.; Yue, S.; Al-Obeed, O.; Kassem, M.; Liu, F.-F.; Aldahmash, A.; Alajez, N.M. MicroRNA-320 suppresses colorectal cancer by targeting SOX4, FOXM1, and FOXQ1. Oncotarget 2016, 7, 35789–35802. [Google Scholar] [CrossRef]
- Liu, X.; Chen, X.; Zeng, K.; Xu, M.; He, B.; Pan, Y.; Sun, H.; Pan, B.; Xu, X.; Xu, T.; et al. DNA-methylation-mediated silencing of miR-486-5p promotes colorectal cancer proliferation and migration through activation of PLAGL2/IGF2/β-catenin signal pathways. Cell Death Dis. 2018, 9, 1037. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baassiri, A.; Nassar, F.; Mukherji, D.; Shamseddine, A.; Nasr, R.; Temraz, S. Exosomal Non Coding RNA in LIQUID Biopsies as a Promising Biomarker for Colorectal Cancer. Int. J. Mol. Sci. 2020, 21, 1398. [Google Scholar] [CrossRef] [Green Version]
- Bach, D.H.; Hong, J.Y.; Park, H.J.; Lee, S.K. The role of exosomes and miRNAs in drug-resistance of cancer cells. Int. J. Cancer 2017, 141, 220–230. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, Q.-R.; Wang, H.; Yan, Y.-D.; Liu, Y.; Su, C.-Y.; Chen, H.-B.; Yan, Y.-Y.; Adhikari, R.; Wu, Q.; Zhang, J.-Y. The role of exosomal microRNA in cancer drug resistance. Front. Oncol. 2020, 10, 472. [Google Scholar] [CrossRef] [PubMed]
- Lopes-Rodrigues, V.; Seca, H.; Sousa, D.; Sousa, E.; Lima, R.T.; Vasconcelos, M.H. The network of P-glycoprotein and microRNAs interactions. Int. J. Cancer 2014, 135, 253–263. [Google Scholar] [CrossRef]
- Liu, T.; Zhang, X.; Du, L.; Wang, Y.; Liu, X.; Tian, H.; Wang, L.; Li, P.; Zhao, Y.; Duan, W. Exosome-transmitted miR-128-3p increase chemosensitivity of oxaliplatin-resistant colorectal cancer. Mol. Cancer 2019, 18, 43. [Google Scholar] [CrossRef] [Green Version]
- Holohan, C.; Van Schaeybroeck, S.; Longley, D.B.; Johnston, P.G. Cancer drug resistance: An evolving paradigm. Nat. Rev. Cancer 2013, 13, 714–726. [Google Scholar] [CrossRef]
- Goldstein, M.; Kastan, M.B. The DNA damage response: Implications for tumor responses to radiation and chemotherapy. Annu. Rev. Med. 2015, 66, 129–143. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jackson, S.P.; Bartek, J. The DNA-damage response in human biology and disease. Nature 2009, 461, 1071–1078. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakajima, G.; Hayashi, K.; Xi, Y.; Kudo, K.; Uchida, K.; Takasaki, K.; Yamamoto, M.; Ju, J. Non-coding microRNAs hsa-let-7g and hsa-miR-181b are associated with chemoresponse to S-1 in colon cancer. Cancer Genom.-Proteom. 2006, 3, 317–324. [Google Scholar]
- Wang, S.; Zeng, Y.; Zhou, J.M.; Nie, S.L.; Peng, Q.; Gong, J.; Huo, J.R. MicroRNA-1246 promotes growth and metastasis of colorectal cancer cells involving CCNG2 reduction. Mol. Med. Rep. 2016, 13, 273–280. [Google Scholar] [CrossRef] [Green Version]
- Hasegawa, S.; Eguchi, H.; Nagano, H.; Konno, M.; Tomimaru, Y.; Wada, H.; Hama, N.; Kawamoto, K.; Kobayashi, S.; Nishida, N. MicroRNA-1246 expression associated with CCNG2-mediated chemoresistance and stemness in pancreatic cancer. Br. J. Cancer 2014, 111, 1572–1580. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.J.; Ren, Z.J.; Tang, J.H.; Yu, Q. Exosomal MicroRNA MiR-1246 promotes cell proliferation, invasion and drug resistance by targeting CCNG2 in breast cancer. Cell. Physiol. Biochem. 2017, 44, 1741–1748. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, G.J. Metabolic reprogramming: The emerging concept and associated therapeutic strategies. J. Exp. Clin. Cancer Res. 2015, 34, 111. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gatenby, R.A.; Gillies, R.J. Why do cancers have high aerobic glycolysis? Nat. Rev. Cancer 2004, 4, 891–899. [Google Scholar] [CrossRef]
- Hirschhaeuser, F.; Sattler, U.G.; Mueller-Klieser, W. Lactate: A metabolic key player in cancer. Cancer Res. 2011, 71, 6921–6925. [Google Scholar] [CrossRef] [Green Version]
- Song, B.; Wang, Y.; Xi, Y.; Kudo, K.; Bruheim, S.; Botchkina, G.I.; Gavin, E.; Wan, Y.; Formentini, A.; Kornmann, M. Mechanism of chemoresistance mediated by miR-140 in human osteosarcoma and colon cancer cells. Oncogene 2009, 28, 4065–4074. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| miRNA | Function |
|---|---|
| miR-671-5p [94] | Oncogenic miR that is overexpressed in the large intestine of CRC patients and colorectal cancer cell lines. Its expression is associated with metastasis, proliferation, migration, and invasion of CRC cells. |
| miR-193b [95] | Tumor-suppressive miR that is downregulated in the serum of CRC patients. Its low levels are correlated with TNM stage and metastasis in CRC patients. |
| miR-1224-5p [96] | Tumor-suppressive miR that is reduced in CRC tissues and cell lines mainly due to the hypoxic microenvironment. It prevents the epithelial–mesenchymal transition (EMT), invasion, and migration of CRC cells by directly interfering with the SP1-mediated NF-κB pathway. |
| miR-125b-1 [97] | Tumor-suppressive miR that is downregulated in early CRC cell lines. Its low levels induce metastasis by increasing the expression of the XIAP gene. |
| miR-125a-3p [98] | Tumor-suppressive miR suppressing fucosyltransferase (FUT)5 and FUT6 to regulate the PI3K/Akt signaling pathway, subsequently inhibiting the proliferation, migration, invasion, and angiogenesis of CRC cells. It also inhibits CRC development by directly targeting the angiogenesis-related gene VEGFA and the antiapoptotic gene Bcl-2. |
| miR-483-5p [99] | Tumor-suppressive miR that inhibits CRC cell proliferation and metastasis, possibly through inhibiting tumor necrosis factor-receptor associated factor (TRAF), which plays critical roles in immune cell signaling). |
| miR-188-5p [100] | Oncogenic miR that is overexpressed in CRC tissue and cell lines. Its higher expression is accompanied by tumor cell proliferation, invasion, and metastasis through inhibition of FOXL1/Wnt signaling. |
| miR-765 [101] | Tumor-suppressive miR that inhibits proliferation, migration, and invasion of CRC cells by targeting patatin-like protein 2 (PLP2). |
| miR-638 [102] | Tumor-suppressive miR that is downregulated in the serum and within exosomes of CRC patients and cell lines. It represses CRC cell viability and migration and modulates the cell cycle by inhibiting tetraspanin 1 (TSPAN1). A reduction in miR-638 is associated with poor overall survival. |
| miR-1246 [79] | Oncogenic miR that is involved in tumor progression and metastasis. miR-1246 targets normal p53; however, miR-1246 was found to be overexpressed in mutant p53 tumor-derived exosomes. |
| miR-654-5p [103] | The abundant expression of miR-654-5p is correlated with colon cancer development, metastasis, and a low survival rate of CRC patients. |
| miR-17 [104] | Overexpression of miR-17 can increase cell proliferation and liver metastases in CRC. It is also associated with the progression of colorectal adenoma to adenocarcinoma in CRC patients. |
| miR-198 [105] | Tumor-suppressive miR that hinders CRC cell viability, triggers death, and inhibits metastasis. |
| miR-601 and miR-760 [54] | Their expression was reduced in the serum of CRC patients, and they could be used as predictors of advanced CRC. |
| miR-493-5p [106] | Tumor-suppressive miR that inhibits CRC progression by targeting the PI3K–Akt–FoxO3a signaling pathway. |
| miR-223-3p [107] | Oncogenic miR that is upregulated in CRC tissues and associated with the proliferation and metastasis of CRC cells. |
| miR-320 [108] | Tumor-suppressive miR that is downregulated in CRC tissues and cell lines. Its upregulation is associated with the inhibition of CRC cell proliferation and metastasis. Its downstream targets are FOXQ1 and SOX4 genes. |
| miR-486-5p [109] | Tumor-suppressive miR that is downregulated in CRC tissues. It is a negative regulator of pleiomorphic adenoma gene-like 2 (PLAGL2), a transcription factor for β-catenin and insulin-like growth factor 2 (IGF2) with roles in promoting proliferation, cell survival, and metastasis, as well as decreasing E-cadherin and increasing N-cadherin expression. |
| miR-150 [110] | Tumor-suppressive miR that is downregulated in serum exosomes of CRC patients; however, this expression was increased in postoperative samples. The downregulated expression was associated with higher tumor progression, metastasis, and poor survival rate. |
| miR-100 [97] | When the expression of this tumor-suppressive miR decreased, CRC growth and metastasis increased. The mechanism involves the induction of downstream targets mTOR, IGF1R, Fas, and XIAP. |
| miR-92a-3p [98] | Exosomal miR-92a-3p facilitates tumor angiogenesis by inducing partial EMT in endothelial cells and through the downregulation of Dkk-3 and claudin-11. Exosomes derived from colon cancer cells and plasma derived from murine xenograft models were enriched with miR-92a-3p, and it has been found to stimulate tube formation in human umbilical vein endothelial cells upon transfer. |
| miR-193a [88] | The expression of this tumor-suppressive miR is abundantly increased in the exosomes of metastatic CRC cell lines and plasma of CRC patients with liver metastasis. Its upregulated expression is associated with a cell cycle arrest in the G1 phase followed by the hindering of CRC cell proliferation through the inhibition of Caprin1, followed by CCND2 and c-MYC. The loss of major vault protein (MVP) caused the upregulation of miR-193a in cells rather than exosomes. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bakhsh, T.; Alhazmi, S.; Alburae, N.A.; Farsi, A.; Alzahrani, F.; Choudhry, H.; Bahieldin, A. Exosomal miRNAs as a Promising Source of Biomarkers in Colorectal Cancer Progression. Int. J. Mol. Sci. 2022, 23, 4855. https://doi.org/10.3390/ijms23094855
Bakhsh T, Alhazmi S, Alburae NA, Farsi A, Alzahrani F, Choudhry H, Bahieldin A. Exosomal miRNAs as a Promising Source of Biomarkers in Colorectal Cancer Progression. International Journal of Molecular Sciences. 2022; 23(9):4855. https://doi.org/10.3390/ijms23094855
Chicago/Turabian StyleBakhsh, Tahani, Safiah Alhazmi, Najla Ali Alburae, Ali Farsi, Faisal Alzahrani, Hani Choudhry, and Ahmed Bahieldin. 2022. "Exosomal miRNAs as a Promising Source of Biomarkers in Colorectal Cancer Progression" International Journal of Molecular Sciences 23, no. 9: 4855. https://doi.org/10.3390/ijms23094855
APA StyleBakhsh, T., Alhazmi, S., Alburae, N. A., Farsi, A., Alzahrani, F., Choudhry, H., & Bahieldin, A. (2022). Exosomal miRNAs as a Promising Source of Biomarkers in Colorectal Cancer Progression. International Journal of Molecular Sciences, 23(9), 4855. https://doi.org/10.3390/ijms23094855







