Nitric Oxide Implication in Potato Immunity to Phytophthora infestans via Modifications of Histone H3/H4 Methylation Patterns on Defense Genes
Abstract
:1. Introduction
2. Results
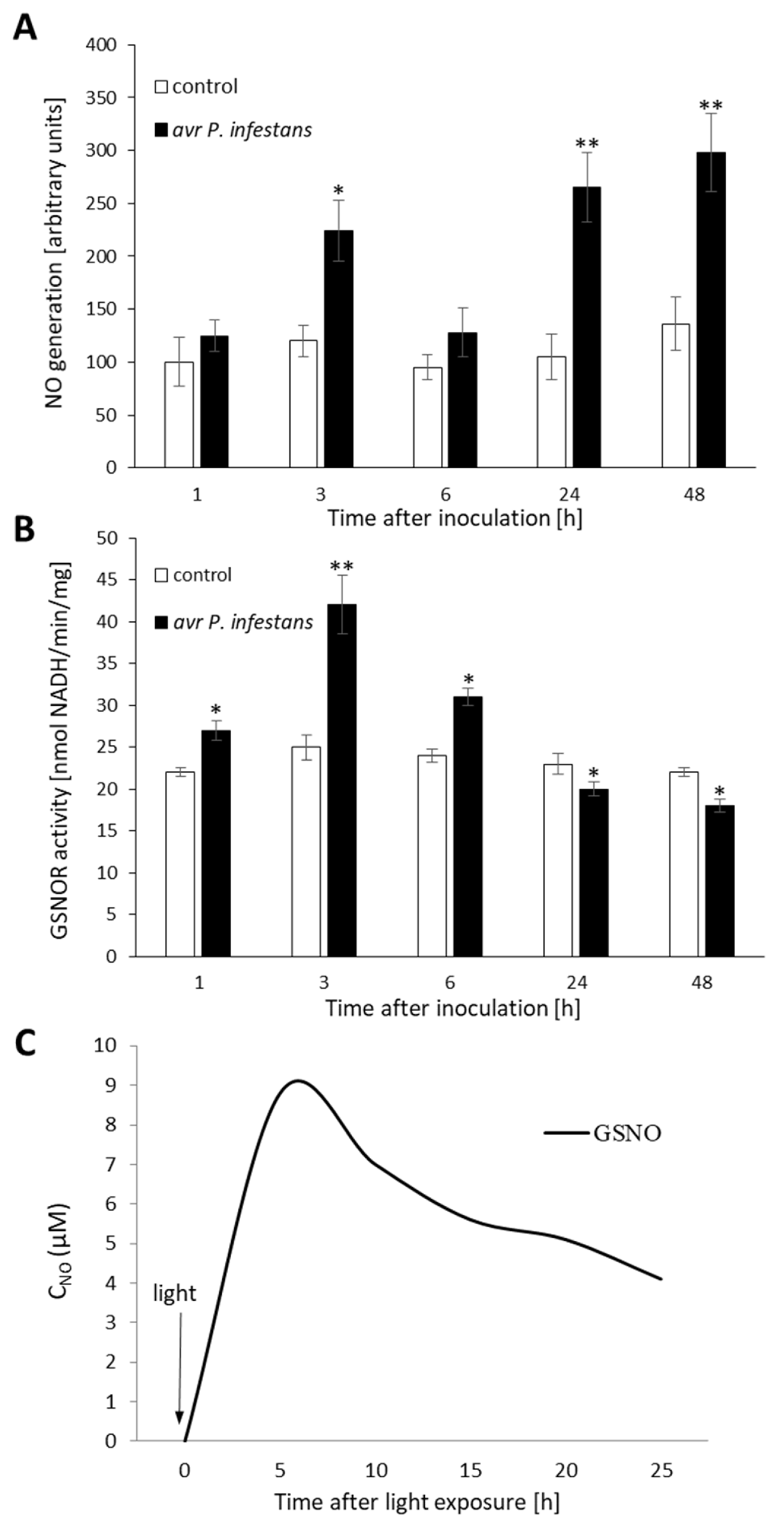
2.1. Biphasic NO Production under GSNOR Controlling
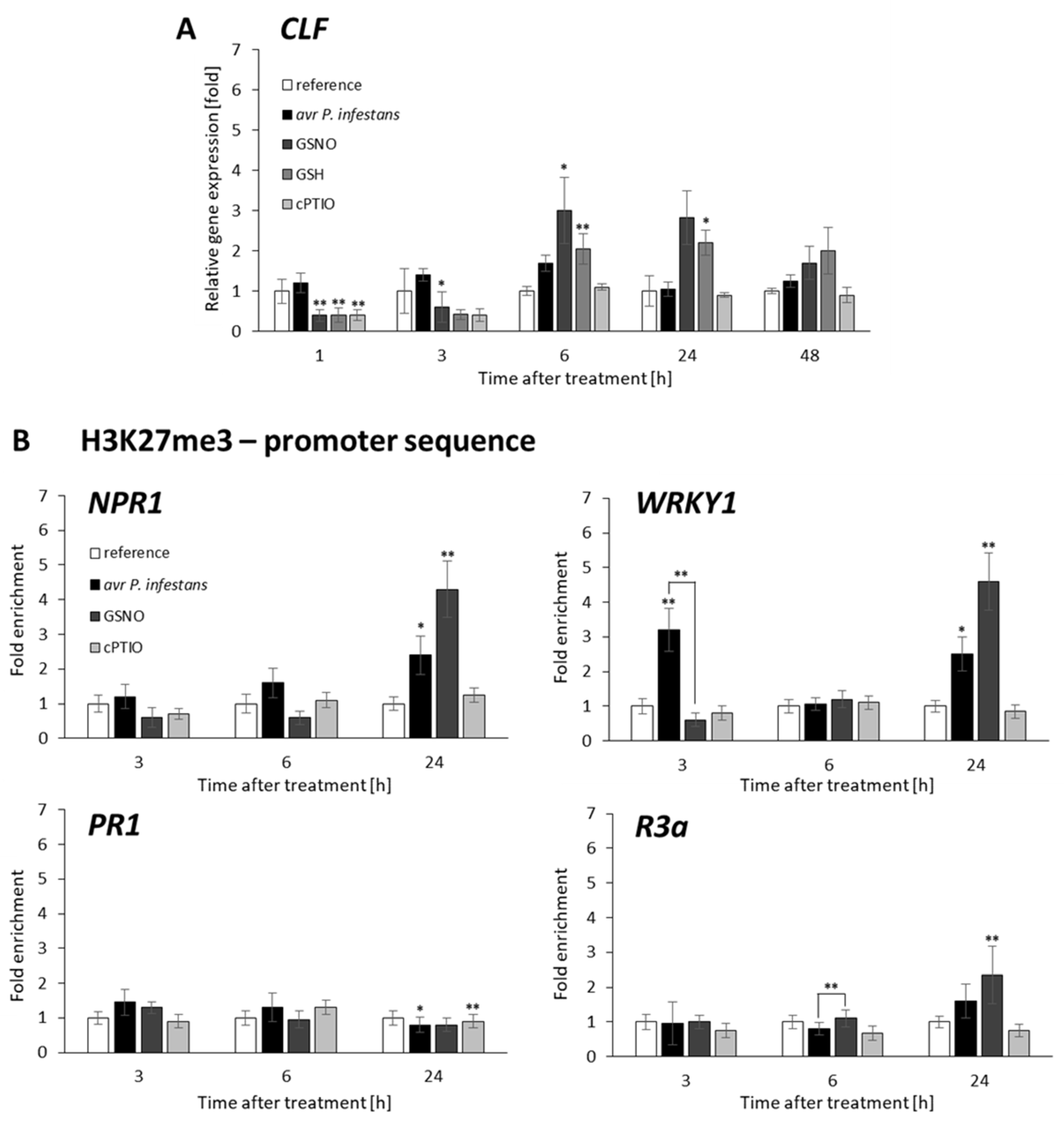
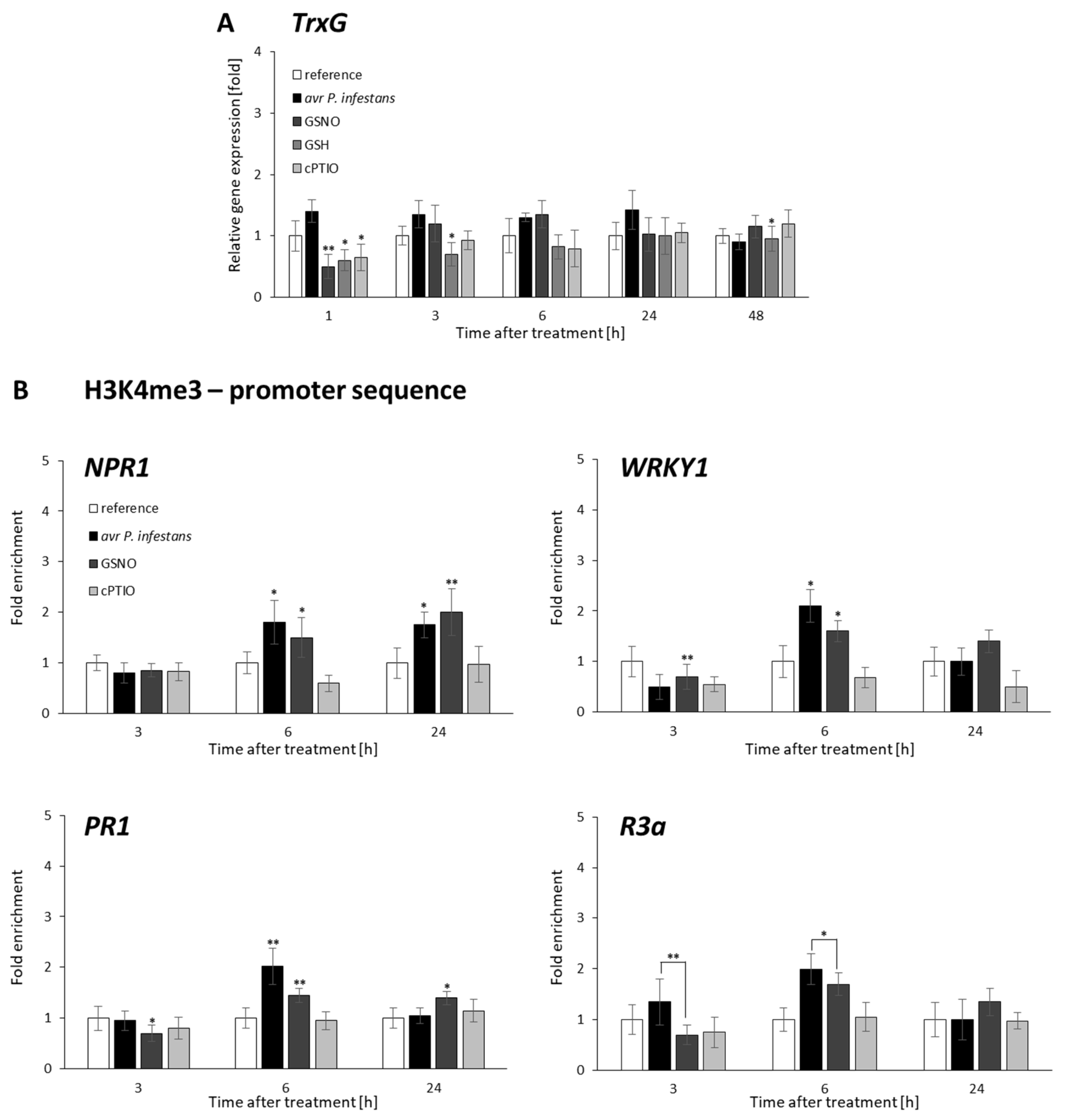
2.2. CLF and TrxG Gene Expression under Redox-Dependent Changes
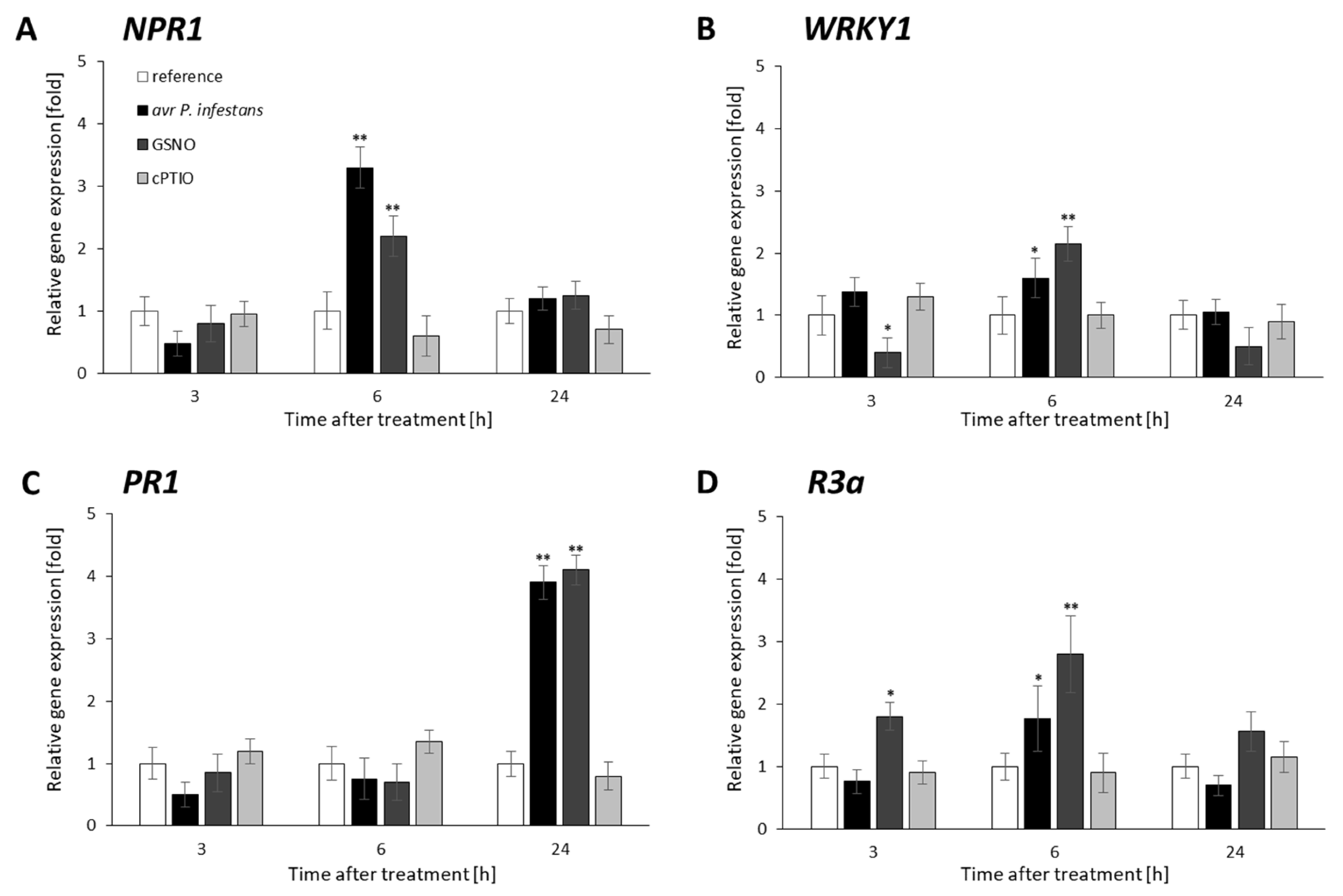
2.3. Distribution Status of CLF-Mediated H3K27me3 and TrxG-Mediated H3K4me3 Marks on Stress-Responsive Genes Shows Some Similarities in the Response to Pathogens and GSNO
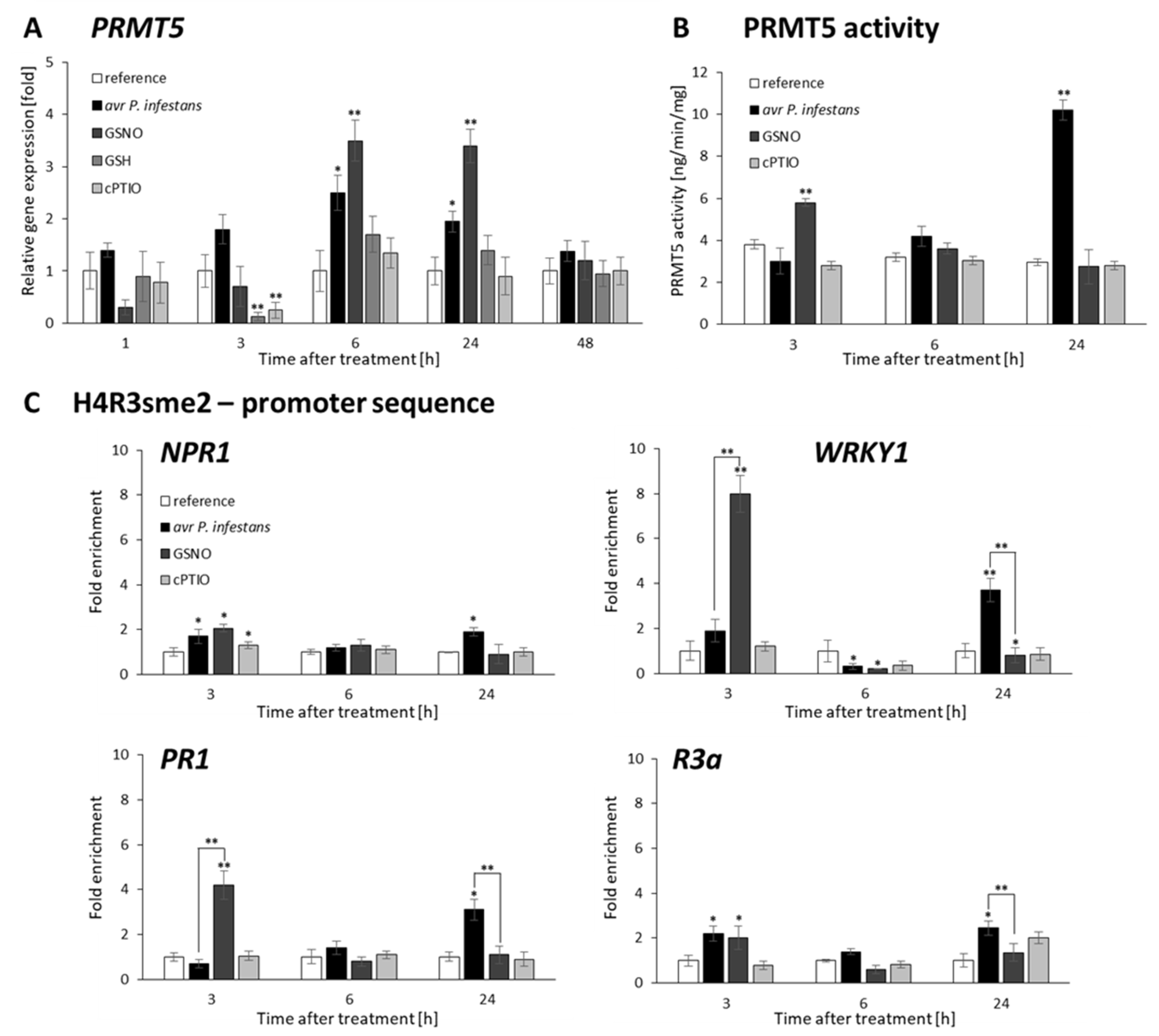
2.4. P. infestans and GSNO Modify PRMT5 Activity and Expression
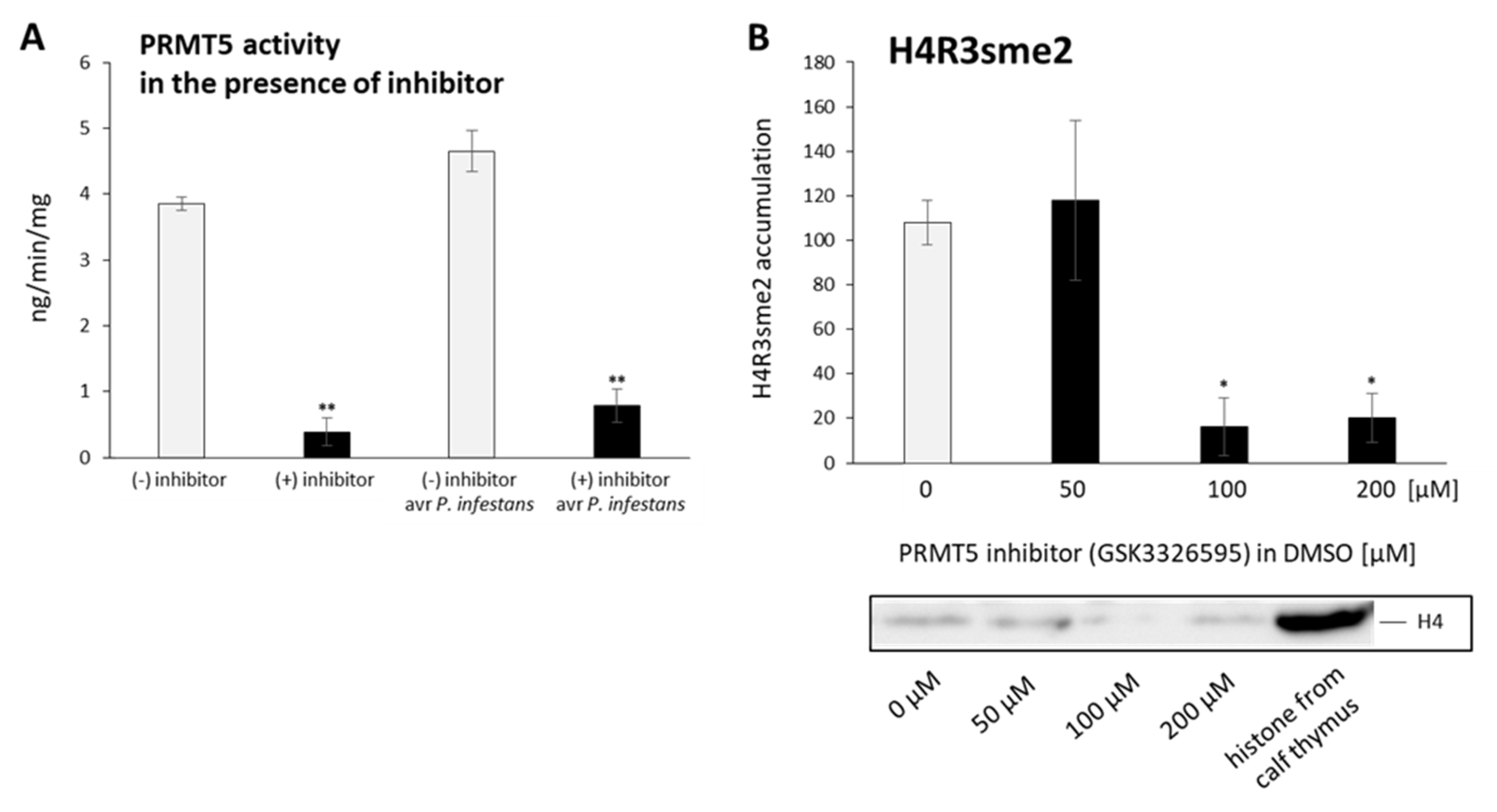
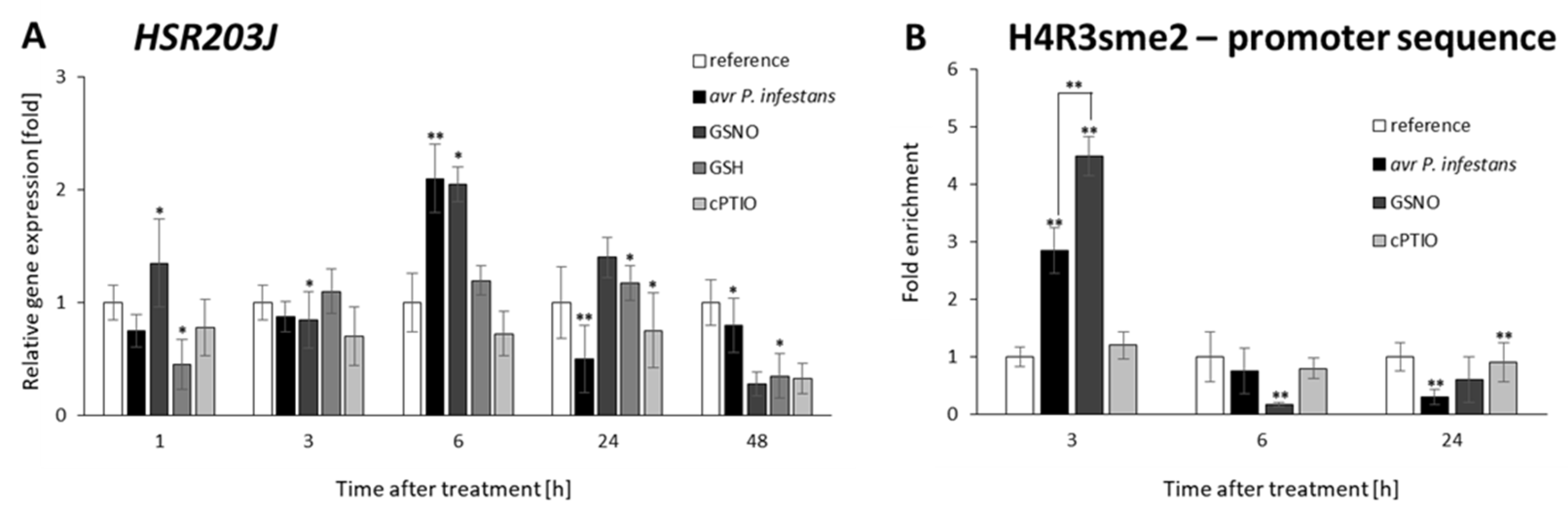
2.5. PRMT5 Affects Defense Genes Expression by Transient Deposition of the H4R3sme2 Mark
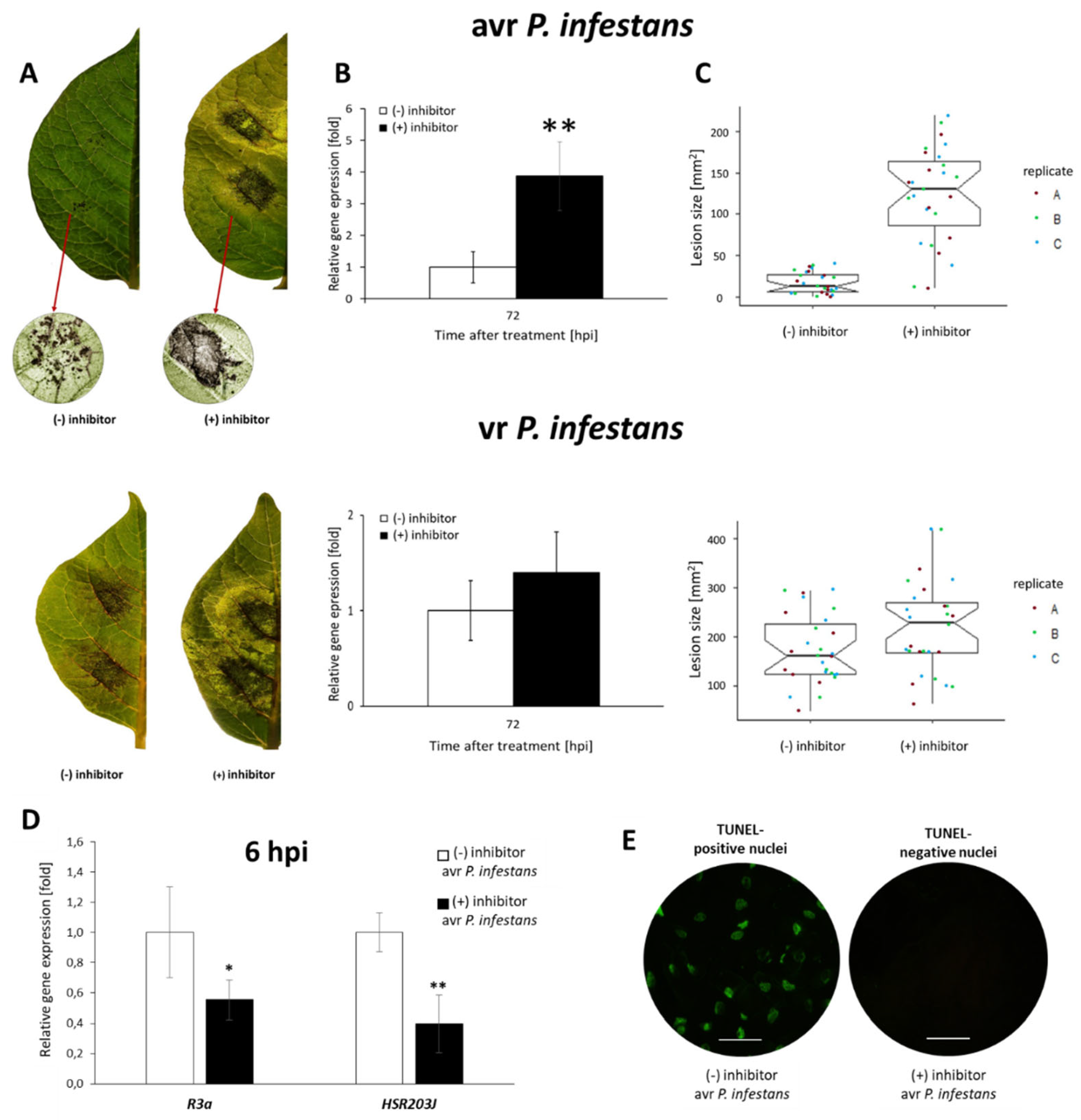
2.6. PRMT5 Contributes to the Hypersensitive Response of Potato to Avr P. infestans
3. Discussion
3.1. Biphasic NO Generation Indirect Reprograms Defense Gene Expression
3.1.1. NO and PRMT5 Activity Are Required to Integrate the Transcription of Defense Genes
3.1.2. NO Cooperates with PRMT5 in the Regulation of Hypersensitive Cell Death and Potato Resistance to Late Blight
4. Materials and Methods
4.1. Plant Material
4.2. Pathogen Culture and Inoculation
4.3. Molecular Quantification of Pathogen
4.4. Assessment of Disease Development
4.5. NO Donor and Scavenger Treatment
4.6. PRMT5 Inhibitor Treatment
4.7. NO Detection and Quantification by the Electrochemical Method
4.8. Measurement of Nitric Oxide Generation
4.9. S-Nitrosoglutathione Reductase [EC 1.2.1.46]
4.10. TUNEL Assay
4.11. Gene Expression Analysis
4.12. Chromatin Immunoprecipitation Assay
4.13. ELISA Test for PRMT5 Activity
4.14. Histone-Enriched Protein Isolation
4.15. Immunoblot Analysis
4.16. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| CLF | curly leaf |
| cPTIO | carboxy-PTIO |
| DNICs | dinitrosyl-iron complexes |
| ETI | effector-triggered immunity |
| GSH | glutathione |
| GSK3326595 | inhibitor of protein arginine N-methyltransferase |
| GSNO | S-nitrosoglutathione |
| GSNOR | S-nitrosoglutathione reductase |
| H4R3sme2 | symmetricdi methylation atthearginine residue Ron histoneH4 |
| H3K27me3 | trimethylation of histone H3lysine27 |
| H3K4me3 | trimethylation of histone H3lysine4 |
| HDACs | histone deacetylases |
| HMTs | methyltransferases |
| HR | hypersensitive response |
| KDMs | lysine demethylases |
| NR | nitrate reductase |
| NO | nitricoxide |
| ONOO¯ | peroxynitrite |
| PAMP | pathogen-associated molecular pattern |
| PR | pathogenesis-related proteins |
| PRC | polycomb repressor complex |
| PRMT5 | protein arginine symmetric methyl transferase5 |
| PTI | PAMP-triggered immunity |
| PTMs | post-translational protein modifications |
| RdDM | RNA-directed DNA methylation |
| ROS | reactive oxygen species |
| siRNA | small non-coding interfering RNA |
| SKB1 | kinasebindingprotein1 |
| TrxG | trithorax |
References
- Mur, L.A.J.; Carver, T.L.W.; Prats, E. NO Way to Live; the Various Roles of Nitric Oxide in Plant–Pathogen Interactions. J. Exp. Bot. 2006, 57, 489–505. [Google Scholar] [CrossRef] [PubMed]
- Trapet, P.; Kulik, A.; Lamotte, O.; Jeandroz, S.; Bourque, S.; Nicolas-Francès, V.; Rosnoblet, C.; Besson-Bard, A.; Wendehenne, D. NO Signaling in Plant Immunity: A Tale of Messengers. Phytochemistry 2015, 112, 72–79. [Google Scholar] [CrossRef] [PubMed]
- Delledonne, M.; Xia, Y.; Dixon, R.A.; Lamb, C. Nitric Oxide Functions as a Signal in Plant Disease Resistance. Nature 1998, 394, 585–588. [Google Scholar] [CrossRef]
- Foissner, I.; Wendehenne, D.; Langebartels, C.; Durner, J. In Vivo Imaging of an Elicitor-Induced Nitric Oxide Burst in Tobacco. Plant J. 2000, 23, 817–824. [Google Scholar] [CrossRef]
- Tada, Y.; Mori, T.; Shinogi, T.; Yao, N.; Takahashi, S.; Betsuyaku, S.; Sakamoto, M.; Park, P.; Nakayashiki, H.; Tosa, Y.; et al. Nitric Oxide and Reactive Oxygen Species Do Not Elicit Hypersensitive Cell Death but Induce Apoptosis in the Adjacent Cells During the Defense Response of Oat. Mol. Plant. Microbe Interact. 2004, 17, 245–253. [Google Scholar] [CrossRef] [Green Version]
- Floryszak-Wieczorek, J.; Arasimowicz, M.; Milczarek, G.; Jelen, H.; Jackowiak, H. Only an Early Nitric Oxide Burst and the Following Wave of Secondary Nitric Oxide Generation Enhanced Effective Defence Responses of Pelargonium to a Necrotrophic Pathogen. New Phytol. 2007, 175, 718–730. [Google Scholar] [CrossRef]
- Asai, S.; Ohta, K.; Yoshioka, H. MAPK Signaling Regulates Nitric Oxide and NADPH Oxidase-Dependent Oxidative Bursts in Nicotiana Benthamiana. Plant Cell 2008, 20, 1390–1406. [Google Scholar] [CrossRef] [Green Version]
- Wendehenne, D.; Gao, Q.-M.; Kachroo, A.; Kachroo, P. Free Radical-Mediated Systemic Immunity in Plants. Curr. Opin. Plant Biol. 2014, 20, 127–134. [Google Scholar] [CrossRef]
- Arasimowicz-Jelonek, M.; Floryszak-Wieczorek, J. Understanding the Fate of Peroxynitrite in Plant Cells—From Physiology to Pathophysiology. Phytochemistry 2011, 72, 681–688. [Google Scholar] [CrossRef]
- Chaki, M.; Valderrama, R.; Fernández-Ocaña, A.M.; Carreras, A.; Gómez-Rodríguez, M.V.; López-Jaramillo, J.; Begara-Morales, J.C.; Sánchez-Calvo, B.; Luque, F.; Leterrier, M.; et al. High Temperature Triggers the Metabolism of S-Nitrosothiols in Sunflower Mediating a Process of Nitrosative Stress Which Provokes the Inhibition of Ferredoxin–NADP Reductase by Tyrosine Nitration. Plant Cell Environ. 2011, 34, 1803–1818. [Google Scholar] [CrossRef]
- Corpas, F.; Palma, J.; del Río, L.; Barroso, J. Protein Tyrosine Nitration in Higher Plants Grown under Natural and Stress Conditions. Front. Plant Sci. 2013, 4, 29. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mata-Pérez, C.; Begara-Morales, J.C.; Chaki, M.; Sánchez-Calvo, B.; Valderrama, R.; Padilla, M.N.; Corpas, F.J.; Barroso, J.B. Protein Tyrosine Nitration during Development and Abiotic Stress Response in Plants. Front. Plant Sci. 2016, 7, 1699. [Google Scholar] [CrossRef] [PubMed]
- Izbiańska, K.; Floryszak-Wieczorek, J.; Gajewska, J.; Meller, B.; Kuźnicki, D.; Arasimowicz-Jelonek, M. RNA and MRNA Nitration as a Novel Metabolic Link in Potato Immune Response to Phytophthora Infestans. Front. Plant Sci. 2018, 9, 672. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Romero-Puertas, M.C.; Laxa, M.; Mattè, A.; Zaninotto, F.; Finkemeier, I.; Jones, A.M.E.; Perazzolli, M.; Vandelle, E.; Dietz, K.-J.; Delledonne, M. S-Nitrosylation of Peroxiredoxin II E Promotes Peroxynitrite-Mediated Tyrosine Nitration. Plant Cell 2007, 19, 4120–4130. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhan, N.; Wang, C.; Chen, L.; Yang, H.; Feng, J.; Gong, X.; Ren, B.; Wu, R.; Mu, J.; Li, Y.; et al. S-Nitrosylation Targets GSNO Reductase for Selective Autophagy during Hypoxia Responses in Plants. Mol. Cell 2018, 71, 142–154. [Google Scholar] [CrossRef] [Green Version]
- Feng, J.; Chen, L.; Zuo, J. Protein S-Nitrosylation in Plants: Current Progresses and Challenges. J. Integr. Plant Biol. 2019, 61, 1206–1223. [Google Scholar] [CrossRef]
- Falak, N.; Imran, Q.M.; Hussain, A.; Yun, B.-W. Transcription Factors as the “Blitzkrieg” of Plant Defense: A Pragmatic View of Nitric Oxide’s Role in Gene Regulation. Int. J. Mol. Sci. 2021, 22, 522. [Google Scholar] [CrossRef]
- Shumaev, K.B.; Kosmachevskaya, O.V.; Chumikina, L.V.; Topunov, A.F. Dinitrosyl Iron Complexes and Other Physiological Metabolites of Nitric Oxide: Multifarious Role in Plants. Nat. Prod. Commun. 2016, 11, 1189–1192. [Google Scholar] [CrossRef] [Green Version]
- Srikant, T.; Drost, H.-G. How Stress Facilitates Phenotypic Innovation Through Epigenetic Diversity. Front. Plant Sci. 2021, 11, 606800. [Google Scholar] [CrossRef]
- Pikaard, C.S.; Scheid, O.M. Epigenetic Regulation in Plants. Cold Spring Harb. Perspect. Biol. 2014, 6, a019315. [Google Scholar] [CrossRef]
- Santos, A.P.; Ferreira, L.J.; Oliveira, M.M. Concerted Flexibility of Chromatin Structure, Methylome, and Histone Modifications along with Plant Stress Responses. Biology 2017, 6, 3. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alvarez, M.E.; Nota, F.; Cambiagno, D.A. Epigenetic Control of Plant Immunity. Mol. Plant Pathol. 2010, 11, 563–576. [Google Scholar] [CrossRef] [PubMed]
- Lafos, M.; Kroll, P.; Hohenstatt, M.L.; Thorpe, F.L.; Clarenz, O.; Schubert, D. Dynamic Regulation of H3K27 Trimethylation during Arabidopsis Differentiation. PLOS Genet. 2011, 7, e1002040. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pu, L.; Sung, Z.R. PcG and TrxG in Plants—Friends or Foes. Trends Genet. 2015, 31, 252–262. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Liu, Y.; Liang, Y.; Zhou, D.; Li, S.; Lin, S.; Dong, H.; Huang, L. The Function of Histone Lysine Methylation Related SET Domain Group Proteins in Plants. Protein Sci. 2020, 29, 1120–1137. [Google Scholar] [CrossRef] [PubMed]
- Bedford, M.T.; Richard, S. Arginine Methylation: An Emerging Regulatorof Protein Function. Mol. Cell 2005, 18, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Wysocka, J.; Allis, C.D.; Coonrod, S. Histone Arginine Methylation and Its Dynamic Regulation. Front. Biosci. 2006, 11, 344–355. [Google Scholar] [CrossRef] [Green Version]
- Pei, Y.; Niu, L.; Lu, F.; Liu, C.; Zhai, J.; Kong, X.; Cao, X. Mutations in the Type II Protein Arginine Methyltransferase AtPRMT5 Result in Pleiotropic Developmental Defects in Arabidopsis. Plant Physiol. 2007, 144, 1913–1923. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Zhang, Y.; Ma, Q.; Zhang, Z.; Xue, Y.; Bao, S.; Chong, K. SKB1-Mediated Symmetric Dimethylation of Histone H4R3 Controls Flowering Time in Arabidopsis. EMBO J. 2007, 26, 1934–1941. [Google Scholar] [CrossRef] [Green Version]
- Schmitz, R.J.; Sung, S.; Amasino, R.M. Histone Arginine Methylation Is Required for Vernalization-Induced Epigenetic Silencing of FLC in Winter-Annual Arabidopsis Thaliana. Proc. Natl. Acad. Sci. USA 2008, 105, 411–416. [Google Scholar] [CrossRef] [Green Version]
- Deng, X.; Gu, L.; Liu, C.; Lu, T.; Lu, F.; Lu, Z.; Cui, P.; Pei, Y.; Wang, B.; Hu, S.; et al. Arginine Methylation Mediated by the Arabidopsis Homolog of PRMT5 Is Essential for Proper Pre-MRNA Splicing. Proc. Natl. Acad. Sci. USA 2010, 107, 19114–19119. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Z.; Zhang, S.; Zhang, Y.; Wang, X.; Li, D.; Li, Q.; Yue, M.; Li, Q.; Zhang, Y.; Xu, Y.; et al. Arabidopsis Floral Initiator SKB1 Confers High Salt Tolerance by Regulating Transcription and Pre-MRNA Splicing through Altering Histone H4R3 and Small Nuclear Ribonucleoprotein LSM4 Methylation. Plant Cell 2011, 23, 396–411. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fan, H.; Zhang, Z.; Wang, N.; Cui, Y.; Sun, H.; Liu, Y.; Wu, H.; Zheng, S.; Bao, S.; Ling, H.-Q. SKB1/PRMT5-Mediated Histone H4R3 Dimethylation of Ib Subgroup BHLH Genes Negatively Regulates Iron Homeostasis in Arabidopsis Thaliana. Plant J. 2014, 77, 209–221. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Zhao, Y.; Yue, M.; Xue, Y.; Bao, S. The Protein Arginine Methylase 5 (PRMT5/SKB1) Gene Is Required for the Maintenance of Root Stem Cells in Response to DNA Damage. J. Genet. Genom. 2016, 43, 187–197. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Ma, H.; Chen, S.; Gu, T.; Gong, J. Control of Proline Accumulation under Drought via a Novel Pathway Comprising the Histone Methylase CAU1 and the Transcription Factor ANAC055. J. Exp. Bot. 2018, 69, 579–588. [Google Scholar] [CrossRef] [Green Version]
- Espejo, A.B.; Gao, G.; Black, K.; Gayatri, S.; Veland, N.; Kim, J.; Chen, T.; Sudol, M.; Walker, C.; Bedford, M.T. PRMT5 C-Terminal Phosphorylation Modulates a 14-3-3/PDZ Interaction Switch. J. Biol. Chem. 2017, 292, 2255–2265. [Google Scholar] [CrossRef] [Green Version]
- Hu, J.; Yang, H.; Mu, J.; Lu, T.; Peng, J.; Deng, X.; Kong, Z.; Bao, S.; Cao, X.; Zuo, J. Nitric Oxide Regulates Protein Methylation during Stress Responses in Plants. Mol. Cell 2017, 67, 702–710. [Google Scholar] [CrossRef]
- Kovacs, I.; Ageeva, A.; König, E.-E.; Lindermayr, C. S-Nitrosylation of Nuclear Proteins: New Pathways in Regulation of Gene Expression. Adv. Bot. Res. 2016, 77, 15–39. [Google Scholar] [CrossRef]
- Ageeva-Kieferle, A.; Rudolf, E.E.; Lindermayr, C. Redox-Dependent Chromatin Remodeling: A New Function of Nitric Oxide as Architect of Chromatin Structure in Plants. Front. Plant Sci. 2019, 10, 625. [Google Scholar] [CrossRef] [Green Version]
- Wurm, C.J.; Lindermayr, C. Nitric Oxide Signaling in the Plant Nucleus: The Function of Nitric Oxide in Chromatin Modulation and Transcription. J. Exp. Bot. 2020, 72, 808–818. [Google Scholar] [CrossRef]
- Lindermayr, C.; Rudolf, E.E.; Durner, J.; Groth, M. Interactions between Metabolism and Chromatin in Plant Models. Mol. Metab. 2020, 38, 100951. [Google Scholar] [CrossRef] [PubMed]
- Saravana Kumar, R.M.; Wang, Y.; Zhang, X.; Cheng, H.; Sun, L.; He, S.; Hao, F. Redox Components: Key Regulators of Epigenetic Modifications in Plants. Int. J. Mol. Sci. 2020, 21, 1419. [Google Scholar] [CrossRef] [Green Version]
- Samo, N.; Ebert, A.; Kopka, J.; Mozgová, I. Plant Chromatin, Metabolism and Development—An Intricate Crosstalk. Curr. Opin. Plant Biol. 2021, 61, 102002. [Google Scholar] [CrossRef] [PubMed]
- Chaki, M.; Valderrama, R.; Fernández-Ocaña, A.M.; Carreras, A.; López-Jaramillo, J.; Luque, F.; Palma, J.M.; Pedrajas, J.R.; Begara-Morales, J.C.; Sánchez-Calvo, B.; et al. Protein Targets of Tyrosine Nitration in Sunflower (Helianthus Annuus L.) Hypocotyls. J. Exp. Bot. 2009, 60, 4221–4234. [Google Scholar] [CrossRef] [PubMed]
- Mengel, A.; Ageeva, A.; Georgii, E.; Bernhardt, J.; Wu, K.; Durner, J.; Lindermayr, C. Nitric Oxide Modulates Histone Acetylation at Stress Genes by Inhibition of Histone Deacetylases. Plant Physiol. 2017, 173, 1434–1452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chaki, M.; Shekariesfahlan, A.; Ageeva, A.; Mengel, A.; von Toerne, C.; Durner, J.; Lindermayr, C. Identification of Nuclear Target Proteins for S-Nitrosylation in Pathogen-Treated Arabidopsis Thaliana Cell Cultures. Plant Sci. 2015, 238, 115–126. [Google Scholar] [CrossRef]
- Ageeva-Kieferle, A.; Georgii, E.; Winkler, B.; Ghirardo, A.; Albert, A.; Hüther, P.; Mengel, A.; Becker, C.; Schnitzler, J.-P.; Durner, J.; et al. Nitric Oxide Coordinates Growth, Development, and Stress Response via Histone Modification and Gene Expression. Plant Physiol. 2021, 187, 336–360. [Google Scholar] [CrossRef]
- Rudolf, E.E.; Hüther, P.; Forné, I.; Georgii, E.; Han, Y.; Hell, R.; Wirtz, M.; Imhof, A.; Becker, C.; Durner, J.; et al. GSNOR Contributes to Demethylation and Expression of Transposable Elements and Stress-Responsive Genes. Antioxidants 2021, 10, 1128. [Google Scholar] [CrossRef]
- Varotto, S.; Tani, E.; Abraham, E.; Krugman, T.; Kapazoglou, A.; Melzer, R.; Radanović, A.; Miladinović, D. Epigenetics: Possible Applications in Climate-Smart Crop Breeding. J. Exp. Bot. 2020, 71, 5223–5236. [Google Scholar] [CrossRef]
- Kakoulidou, I.; Avramidou, E.V.; Baránek, M.; Brunel-Muguet, S.; Farrona, S.; Johannes, F.; Kaiserli, E.; Lieberman-Lazarovich, M.; Martinelli, F.; Mladenov, V.; et al. Epigenetics for Crop Improvement in Times of Global Change. Biology 2021, 10, 766. [Google Scholar] [CrossRef]
- Zhi, P.; Chang, C. Exploiting Epigenetic Variations for Crop Disease Resistance Improvement. Front. Plant Sci. 2021, 12, 692328. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Andrews, H.; Eglitis-Sexton, J.; Godwin, I.; Tanurdžić, M.; Crisp, P.A. Epigenome Guided Crop Improvement: Current Progress and Future Opportunities. Emerg. Top. Life Sci. 2022, ETLS20210258. [Google Scholar] [CrossRef] [PubMed]
- Haverkort, A.J.; Struik, P.C.; Visser, R.G.F.; Jacobsen, E. Applied Biotechnology to Combat Late Blight in Potato Caused by Phytophthora Infestans. Potato Res. 2009, 52, 249–264. [Google Scholar] [CrossRef]
- Floryszak-Wieczorek, J.; Milczarek, G.; Arasimowicz, M.; Ciszewski, A. Do Nitric Oxide Donors Mimic Endogenous NO-Related Response in Plants? Planta 2006, 224, 1363–1372. [Google Scholar] [CrossRef] [PubMed]
- Komar, D.N.; Mouriz, A.; Jarillo, J.A.; Piñeiro, M. Chromatin Immunoprecipitation Assay for the Identification of Arabidopsis Protein-DNA Interactions In Vivo. J. Vis. Exp. 2016, e53422. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tao, H.; Yan, X.; Zhu, K.; Zhang, H. Discovery of Novel PRMT5 Inhibitors by Virtual Screening and Biological Evaluations. Chem. Pharm. Bull. 2019, 67, 382–388. [Google Scholar] [CrossRef] [Green Version]
- Pontier, D.; Tronchet, M.; Rogowsky, P.; Lam, E.; Roby, D. Activation of Hsr203, a Plant Gene Expressed during Incompatible Plant-Pathogen Interactions, Is Correlated with Programmed Cell Death. Mol. Plant Microbe Interact. 1998, 11, 544–554. [Google Scholar] [CrossRef]
- Pontier, D.; Balagué, C.; Bezombes-Marion, I.; Tronchet, M.; Deslandes, L.; Roby, D. Identification of a Novel Pathogen-Responsive Element in the Promoter of the Tobacco Gene HSR203J, a Molecular Marker of the Hypersensitive Response. Plant J. 2001, 26, 495–507. [Google Scholar] [CrossRef] [Green Version]
- Lamb, C.; Dixon, R.A. The Oxidative Burst in Plant Disease Resistance. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1997, 48, 251–275. [Google Scholar] [CrossRef]
- Mur, L.A.J.; Brown, I.R.; Darby, R.M.; Bestwick, C.S.; Bi, Y.-M.; Mansfield, J.W.; Draper, J. A Loss of Resistance to Avirulent Bacterial Pathogens in Tobacco Is Associated with the Attenuation of a Salicylic Acid-Potentiated Oxidative Burst. Plant J. 2000, 23, 609–621. [Google Scholar] [CrossRef] [Green Version]
- Floryszak-Wieczorek, J.; Arasimowicz-Jelonek, M.; Izbiańska, K. The Combined Nitrate Reductase and Nitrite-Dependent Route of NO Synthesis in Potato Immunity to Phytophthora Infestans. Plant Physiol. Biochem. 2016, 108, 468–477. [Google Scholar] [CrossRef] [PubMed]
- Feechan, A.; Kwon, E.; Yun, B.-W.; Wang, Y.; Pallas, J.A.; Loake, G.J. A Central Role for S-Nitrosothiols in Plant Disease Resistance. Proc. Natl. Acad. Sci. USA 2005, 102, 8054–8059. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Treffon, P.; Rossi, J.; Gabellini, G.; Trost, P.; Zaffagnini, M.; Vierling, E. Quantitative Proteome Profiling of a S-Nitrosoglutathione Reductase (GSNOR) Null Mutant Reveals a New Class of Enzymes Involved in Nitric Oxide Homeostasis in Plants. Front. Plant Sci. 2021, 12, 2869. [Google Scholar] [CrossRef] [PubMed]
- Delorme-Hinoux, V.; Bangash, S.A.K.; Meyer, A.J.; Reichheld, J.-P. Nuclear Thiol Redox Systems in Plants. Plant Sci. 2016, 243, 84–95. [Google Scholar] [CrossRef] [PubMed]
- Sani, E.; Herzyk, P.; Perrella, G.; Colot, V.; Amtmann, A. Hyperosmotic Priming of Arabidopsis Seedlings Establishes a Long-Term Somatic Memory Accompanied by Specific Changes of the Epigenome. Genome Biol. 2013, 14, R59. [Google Scholar] [CrossRef] [Green Version]
- Sneppen, K.; Ringrose, L. Theoretical Analysis of Polycomb-Trithorax Systems Predicts That Poised Chromatin Is Bistable and Not Bivalent. Nat. Commun. 2019, 10, 2133. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.-H. Multifaceted Chromatin Structure and Transcription Changes in Plant Stress Response. Int. J. Mol. Sci. 2021, 22, 2013. [Google Scholar] [CrossRef]
- Liu, N.; Fromm, M.; Avramova, Z. H3K27me3 and H3K4me3 Chromatin Environment at Super-Induced Dehydration Stress Memory Genes of Arabidopsis Thaliana. Mol. Plant 2014, 7, 502–513. [Google Scholar] [CrossRef] [Green Version]
- Vasudevan, D.; Hickok, J.R.; Bovee, R.C.; Pham, V.; Mantell, L.L.; Bahroos, N.; Kanabar, P.; Cao, X.-J.; Maienschein-Cline, M.; Garcia, B.A.; et al. Nitric Oxide Regulates Gene Expression in Cancers by Controlling Histone Posttranslational Modifications. Cancer Res. 2015, 75, 5299–5308. [Google Scholar] [CrossRef] [Green Version]
- Socco, S.; Bovee, R.C.; Palczewski, M.B.; Hickok, J.R.; Thomas, D.D. Epigenetics: The Third Pillar of Nitric Oxide Signaling. Pharmacol. Res. 2017, 121, 52–58. [Google Scholar] [CrossRef]
- Vasudevan, D.; Bovee, R.C.; Thomas, D.D. Nitric Oxide, the New Architect of Epigenetic Landscapes. Nitric Oxide 2016, 59, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Palczewski, M.B.; Petraitis, H.; Thomas, D.D. Nitric Oxide Is an Epigenetic Regulator of Histone Post-Translational Modifications in Cancer. Curr. Opin. Physiol. 2019, 9, 94–99. [Google Scholar] [CrossRef]
- Nott, A.; Watson, P.M.; Robinson, J.D.; Crepaldi, L.; Riccio, A. S-Nitrosylation of Histone Deacetylase 2 Induces Chromatin Remodelling in Neurons. Nature 2008, 455, 411–415. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.H.; Jing, F.B.; Fang, H.; Gu, L.C.; Xu, W.F. Expression, Purification, and S-Nitrosylation of Recombinant Histone Deacetylase 8 in Escherichia Coli. Biosci. Trends 2011, 5, 17–22. [Google Scholar] [CrossRef] [Green Version]
- Okuda, K.; Ito, A.; Uehara, T. Regulation of Histone Deacetylase 6 Activity via S-Nitrosylation. Biol. Pharm. Bull. 2015, 38, 1434–1437. [Google Scholar] [CrossRef] [Green Version]
- Hernando, C.E.; Sanchez, S.E.; Mancini, E.; Yanovsky, M.J. Genome Wide Comparative Analysis of the Effects of PRMT5 and PRMT4/CARM1 Arginine Methyltransferases on the Arabidopsis Thaliana Transcriptome. BMC Genom. 2015, 16, 192. [Google Scholar] [CrossRef] [Green Version]
- Hickok, J.R.; Vasudevan, D.; Antholine, W.E.; Thomas, D.D. Nitric Oxide Modifies Global Histone Methylation by Inhibiting Jumonji C Domain-Containing Demethylases. J. Biol. Chem. 2013, 288, 16004–16015. [Google Scholar] [CrossRef] [Green Version]
- Sanchez, S.E.; Petrillo, E.; Beckwith, E.J.; Zhang, X.; Rugnone, M.L.; Hernando, C.E.; Cuevas, J.C.; Godoy Herz, M.A.; Depetris-Chauvin, A.; Simpson, C.G.; et al. A Methyl Transferase Links the Circadian Clock to the Regulation of Alternative Splicing. Nature 2010, 468, 112–116. [Google Scholar] [CrossRef] [Green Version]
- Yue, M.; Li, Q.; Zhang, Y.; Zhao, Y.; Zhang, Z.; Bao, S. Histone H4R3 Methylation Catalyzed by SKB1/PRMT5 Is Required for Maintaining Shoot Apical Meristem. PLoS ONE 2013, 8, e83258. [Google Scholar] [CrossRef] [Green Version]
- Balint-Kurti, P. The Plant Hypersensitive Response: Concepts, Control and Consequences. Mol. Plant Pathol. 2019, 20, 1163–1178. [Google Scholar] [CrossRef] [Green Version]
- Yogendra, K.N.; Kumar, A.; Sarkar, K.; Li, Y.; Pushpa, D.; Mosa, K.A.; Duggavathi, R.; Kushalappa, A.C. Transcription Factor StWRKY1 Regulates Phenylpropanoid Metabolites Conferring Late Blight Resistance in Potato. J. Exp. Bot. 2015, 66, 7377–7389. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chacón-Cerdas, R.; Barboza-Barquero, L.; Albertazzi, F.J.; Rivera-Méndez, W. Transcription Factors Controlling Biotic Stress Response in Potato Plants. Physiol. Mol. Plant Pathol. 2020, 112, 101527. [Google Scholar] [CrossRef]
- Naveed, Z.A.; Wei, X.; Chen, J.; Mubeen, H.; Ali, G.S. The PTI to ETI Continuum in Phytophthora-Plant Interactions. Front. Plant Sci. 2020, 11, 2030. [Google Scholar] [CrossRef] [PubMed]
- Sachamitr, P.; Ho, J.C.; Ciamponi, F.E.; Ba-Alawi, W.; Coutinho, F.J.; Guilhamon, P.; Kushida, M.M.; Cavalli, F.M.G.; Lee, L.; Rastegar, N.; et al. PRMT5 Inhibition Disrupts Splicing and Stemness in Glioblastoma. Nat. Commun. 2021, 12, 979. [Google Scholar] [CrossRef]
- Metz, P.J.; Ching, K.A.; Xie, T.; Delgado Cuenca, P.; Niessen, S.; Tatlock, J.H.; Jensen-Pergakes, K.; Murray, B.W. Symmetric Arginine Dimethylation Is Selectively Required for MRNA Splicing and the Initiation of Type I and Type III Interferon Signaling. Cell Rep. 2020, 30, 1935–1950.e8. [Google Scholar] [CrossRef] [Green Version]
- Pal, S.; Sif, S. Interplay between Chromatin Remodelers and Protein Arginine Methyltransferases. J. Cell. Physiol. 2007, 213, 306–315. [Google Scholar] [CrossRef]
- Ahmad, A.; Cao, X. Plant PRMTs Broaden the Scope of Arginine Methylation. J. Genet. Genom. 2012, 39, 195–208. [Google Scholar] [CrossRef]
- Karkhanis, V.; Hu, Y.-J.; Baiocchi, R.A.; Imbalzano, A.N.; Sif, S. Versatility of PRMT5-Induced Methylation in Growth Control and Development. Trends Biochem. Sci. 2011, 36, 633–641. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Jing, L.; Li, M.; He, L.; Guo, Z. Regulation of Histone Arginine Methylation/Demethylation by Methylase and Demethylase (Review). Mol. Med. Rep. 2019, 19, 3963–3971. [Google Scholar] [CrossRef] [Green Version]
- Liu, F.; Xu, Y.; Lu, X.; Hamard, P.-J.; Karl, D.L.; Man, N.; Mookhtiar, A.K.; Martinez, C.; Lossos, I.S.; Sun, J.; et al. PRMT5-Mediated Histone Arginine Methylation Antagonizes Transcriptional Repression by Polycomb Complex PRC2. Nucleic Acids Res. 2020, 48, 2956–2968. [Google Scholar] [CrossRef]
- van’t Klooster, J.W.; van den Berg-Velthuis, G.; van West, P.; Govers, F. Tef1, a Phytophthora Infestans Gene Encoding Translation Elongation Factor 1α. Gene 2000, 249, 145–151. [Google Scholar] [CrossRef]
- Barroso, J.B.; Corpas, F.J.; Carreras, A.; Rodríguez-Serrano, M.; Esteban, F.J.; Fernández-Ocaña, A.; Chaki, M.; Romero-Puertas, M.C.; Valderrama, R.; Sandalio, L.M.; et al. Localization of S-Nitrosoglutathione and Expression of S-Nitrosoglutathione Reductase in Pea Plants under Cadmium Stress. J. Exp. Bot. 2006, 57, 1785–1793. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Janus, Ł.; Milczarek, G.; Arasimowicz-Jelonek, M.; Abramowski, D.; Billert, H.; Floryszak-Wieczorek, J. Normoergic NO-Dependent Changes, Triggered by a SAR Inducer in Potato, Create More Potent Defense Responses to Phytophthora Infestans. Plant Sci. 2013, 211, 23–34. [Google Scholar] [CrossRef] [PubMed]
- Floryszak-Wieczorek, J.; Arasimowicz-Jelonek, M. Contrasting Regulation of NO and ROS in Potato Defense-Associated Metabolism in Response to Pathogens of Different Lifestyles. PLoS ONE 2016, 11, e0163546. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Fernald, R.D. Comprehensive Algorithm for Quantitative Real-Time Polymerase Chain Reaction. J. Comput. Biol. 2005, 12, 1047–1064. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A New Mathematical Model for Relative Quantification in Real-Time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- Tichopad, A.; Didier, A.; Pfaffl, M.W. Inhibition of Real-Time RT-PCR Quantification Due to Tissue-Specific Contaminants. Mol. Cell Probes 2004, 18, 45–50. [Google Scholar] [CrossRef]
- Haring, M.; Offermann, S.; Danker, T.; Horst, I.; Peterhansel, C.; Stam, M. Chromatin Immunoprecipitation: Optimization, Quantitative Analysis and Data Normalization. Plant Methods 2007, 3, 11. [Google Scholar] [CrossRef] [Green Version]
- Jarillo, J.A.; Komar, D.N.; Piñeiro, M. The Use of the Chromatin Immunoprecipitation Technique for In Vivo Identification of Plant Protein-DNA Interactions. Methods Mol Biol. 2018, 1794, 323–334. [Google Scholar] [CrossRef]
- Moehs, C.P.; McElwain, E.F.; Spiker, S. Chromosomal Proteins of Arabidopsis Thaliana. Plant Mol. Biol. 1988, 11, 507–515. [Google Scholar] [CrossRef]
- Bradford, M.M. A Rapid and Sensitive Method for the Quantitation of Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Drozda, A.; Kurpisz, B.; Arasimowicz-Jelonek, M.; Kuźnicki, D.; Jagodzik, P.; Guan, Y.; Floryszak-Wieczorek, J. Nitric Oxide Implication in Potato Immunity to Phytophthora infestans via Modifications of Histone H3/H4 Methylation Patterns on Defense Genes. Int. J. Mol. Sci. 2022, 23, 4051. https://doi.org/10.3390/ijms23074051
Drozda A, Kurpisz B, Arasimowicz-Jelonek M, Kuźnicki D, Jagodzik P, Guan Y, Floryszak-Wieczorek J. Nitric Oxide Implication in Potato Immunity to Phytophthora infestans via Modifications of Histone H3/H4 Methylation Patterns on Defense Genes. International Journal of Molecular Sciences. 2022; 23(7):4051. https://doi.org/10.3390/ijms23074051
Chicago/Turabian StyleDrozda, Andżelika, Barbara Kurpisz, Magdalena Arasimowicz-Jelonek, Daniel Kuźnicki, Przemysław Jagodzik, Yufeng Guan, and Jolanta Floryszak-Wieczorek. 2022. "Nitric Oxide Implication in Potato Immunity to Phytophthora infestans via Modifications of Histone H3/H4 Methylation Patterns on Defense Genes" International Journal of Molecular Sciences 23, no. 7: 4051. https://doi.org/10.3390/ijms23074051
APA StyleDrozda, A., Kurpisz, B., Arasimowicz-Jelonek, M., Kuźnicki, D., Jagodzik, P., Guan, Y., & Floryszak-Wieczorek, J. (2022). Nitric Oxide Implication in Potato Immunity to Phytophthora infestans via Modifications of Histone H3/H4 Methylation Patterns on Defense Genes. International Journal of Molecular Sciences, 23(7), 4051. https://doi.org/10.3390/ijms23074051






