Transcriptomic and Physiological Responses of Chlorella pyrenoidosa during Exposure to 17α-Ethinylestradiol
Abstract
1. Introduction
2. Results
2.1. The Growth of C. pyrenoidosa after Exposure to 17α-EE2
2.2. Characterization by SEM and FTIR Spectroscopy
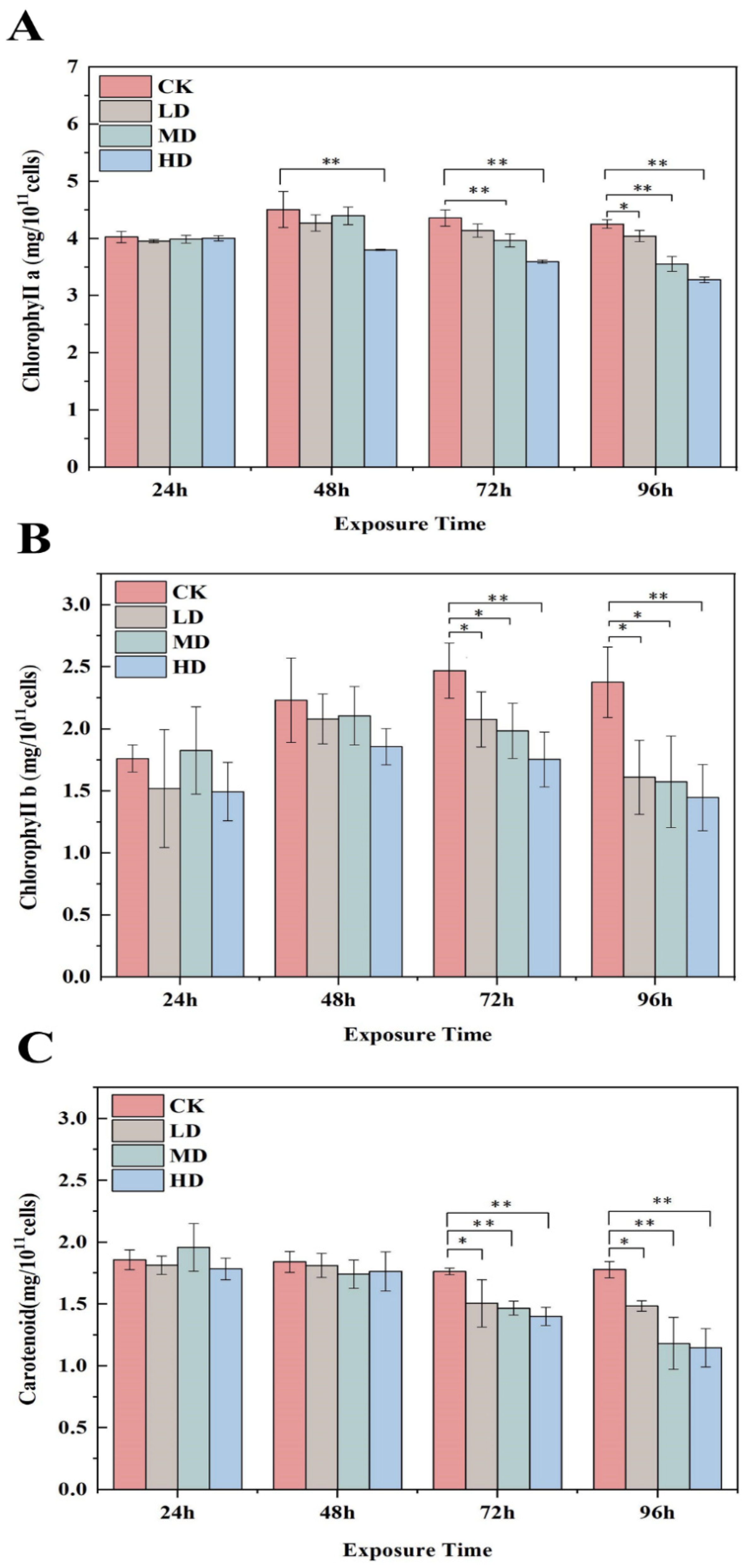
2.3. Effects on Photosynthesis
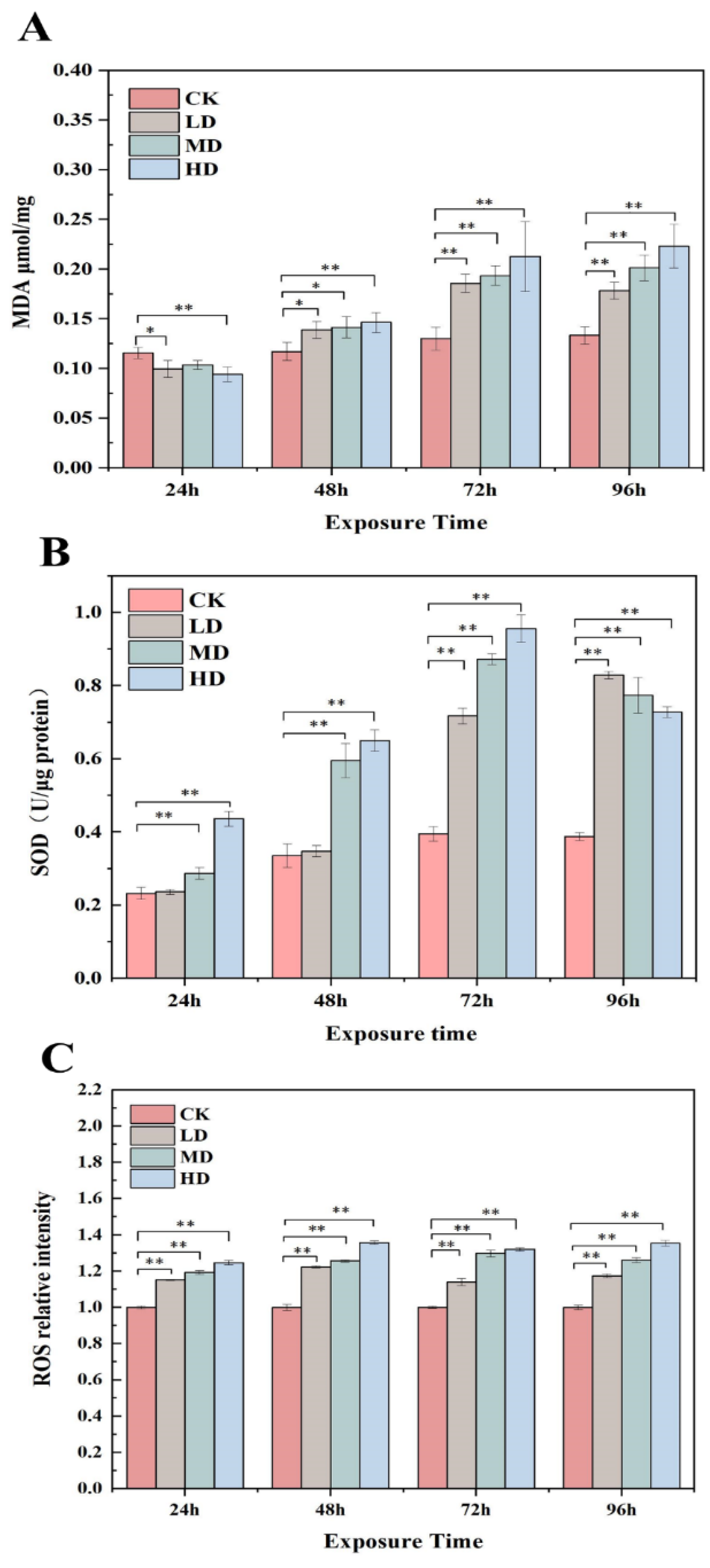
2.4. Variation in the MDA Levels, SOD Activity, and ROS Generation in Response to 17α-EE2
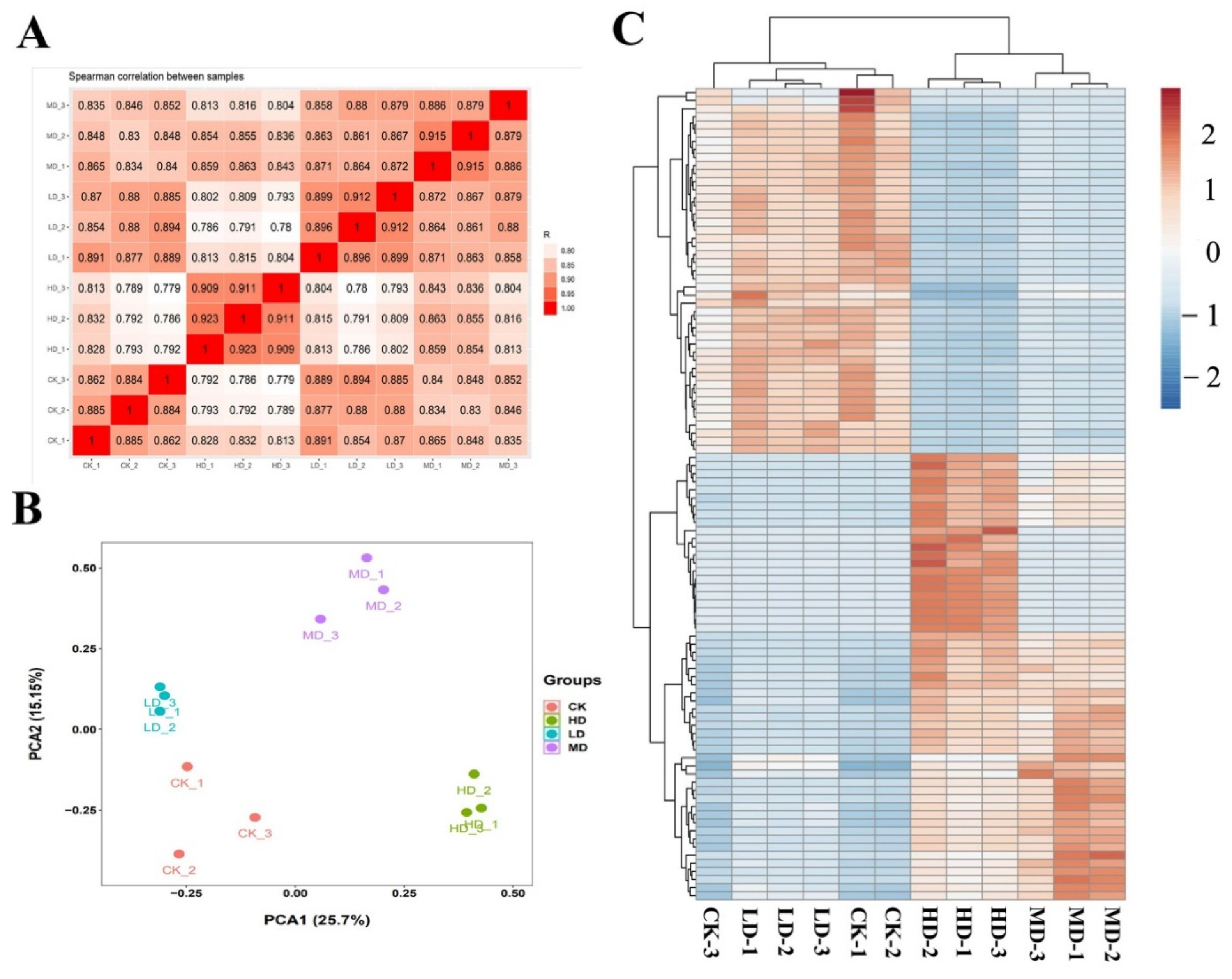
2.5. Differentially Expressed Genes
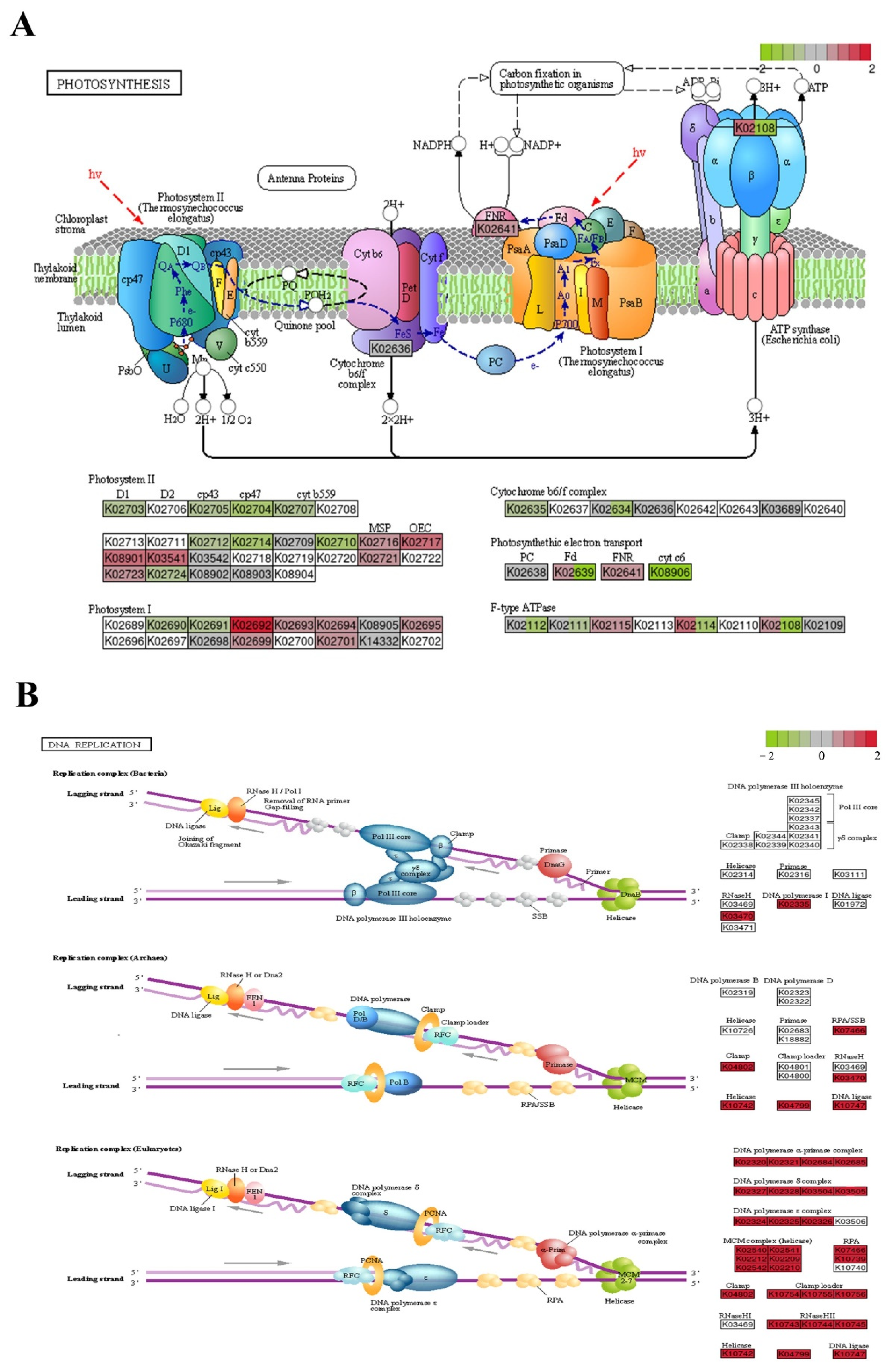
2.6. Analysis of Transcriptional Changes
3. Discussion
4. Materials and Methods
4.1. Algal Cultures
4.2. 17α-EE2 Exposure
4.3. Determination of Cell Density and Photosynthetic Pigment Content
4.4. Measurement of Chlorophyll Fluorescence
4.5. Analysis of Contents of MDA, SOD, and ROS
4.6. Morphological Examination
4.7. Determination of Surface Characteristics
4.8. Transcriptome Sequencing Analysis
4.9. Quantitative Real-Time PCR
4.10. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pocock, T.; Falk, S. Negative impact on growth and photosynthesis in the green alga Chlamydomonas reinhardtii in the presence of the estrogen 17α-ethynylestradiol. PLoS ONE 2014, 9, e109289. [Google Scholar] [CrossRef] [PubMed]
- Segner, H.; Caroll, K.; Fenske, M.; Janssen, C.R.; Maack, G.; Pascoe, D.; Schäfers, C.; Vandenbergh, G.F.; Watts, M.; Wenzel, A. Identification of endocrine-disrupting effects in aquatic vertebrates and invertebrates: Report from the European IDEA project. Ecotoxicol. Environ. Saf. 2003, 54, 302–314. [Google Scholar] [CrossRef]
- Adeel, M.; Song, X.; Wang, Y.; Francis, D.; Yang, Y. Environmental impact of estrogens on human, animal and plant life: A critical review. Environ. Int. 2017, 99, 107–119. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.M.; Lee, M.C.; Kang, H.M.; Rhee, J.S.; Lee, J.S. Genomic organization and transcriptional modulation in response to endocrine disrupting chemicals of three vitellogenin genes in the self-fertilizing fish Kryptolebias marmoratus. J. Environ. Sci. 2016, 42, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.X.; Wu, J.L.; Lv, H.B.; Zhang, H.Y.; Zhang, J.; Limbu, S.M.; Qiao, F.; Chen, L.Q.; Yang, Y.; Zhang, M.L.; et al. Environmental estrogen exposure converts lipid metabolism in male fish to a female pattern mediated by AMPK and mTOR signaling pathways. J. Hazard. Mater. 2020, 394, 122537. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Liu, Z.-H.; Wang, H.; Dang, Z.; Liu, Y. A review of 17α-ethynylestradiol (EE2) in surface water across 32 countries: Sources, concentrations, and potential estrogenic effects. J. Environ. Manag. 2021, 292, 112804. [Google Scholar] [CrossRef]
- De Mes, T.; Zeeman, G.; Lettinga, G. Occurrence and fate of estrone, 17β-estradiol and 17α-ethynylestradiol in STPs for domestic wastewater. Rev. Environ. Sci. Biotechnol. 2005, 4, 275–311. [Google Scholar] [CrossRef]
- Campbell, C.G.; Borglin, S.E.; Green, F.B.; Grayson, A.; Wozei, E.; Stringfellow, W.T. Biologically directed environmental monitoring, fate, and transport of estrogenic endocrine disrupting compounds in water: A review. Chemosphere 2006, 65, 1265–1280. [Google Scholar] [CrossRef] [PubMed]
- Sornalingam, K.; McDonagh, A.; Zhou, J.L. Photodegradation of estrogenic endocrine disrupting steroidal hormones in aqueous systems: Progress and future challenges. Sci. Total Environ. 2016, 550, 209–224. [Google Scholar] [CrossRef]
- Vilela, C.L.S.; Bassin, J.P.; Peixoto, R.S. Water contamination by endocrine disruptors: Impacts, microbiological aspects and trends for environmental protection. Environ. Pollut. 2018, 235, 546–559. [Google Scholar] [CrossRef] [PubMed]
- Pawlowski, S.; Ternes, T.; Bonerz, M.; Kluczka, T.; van der Burg, B.; Nau, H.; Erdinger, L.; Braunbeck, T. Combined in situ and in vitro assessment of the estrogenic activity of sewage and surface water samples. Toxicol. Sci. 2003, 75, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, P.Y.; Carvalho, G.; Reis, M.A.M.; Oehmen, A. A review of the biotransformations of priority pharmaceuticals in biological wastewater treatment processes. Water Res. 2021, 188, 116446. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Tang, J.; He, H.; Gu, L.; Pan, X. Ecotoxicological effects and removal of 17β-estradiol in chlorella algae. Ecotoxicol. Environ. Saf. 2019, 174, 377–383. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.Q.; Kurade, M.B.; Abou-Shanab, R.A.I.; Ji, M.K.; Choi, J.; Kim, J.O.; Jeon, B.H. Biodegradation of carbamazepine using freshwater microalgae Chlamydomonas mexicana and Scenedesmus obliquus and the determination of its metabolic fate. Bioresour. Technol. 2016, 205, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Joo, S.H. The ecotoxicological impact of metal oxide Nanoparticles on pool algae in the presence and absence of disinfection byproducts: A new research direction for the public health and safety of engineered Nanoparticles used in consumer products. Water. Air. Soil Pollut. 2013, 224, 1681. [Google Scholar] [CrossRef]
- Fastelli, P.; Renzi, M. Exposure of key marine species to sunscreens: Changing ecotoxicity as a possible indirect effect of global warming. Mar. Pollut. Bull. 2019, 149, 110517. [Google Scholar] [CrossRef] [PubMed]
- Maes, H.M.; Maletz, S.X.; Ratte, H.T.; Hollender, J.; Schaeffer, A. Uptake, elimination, and biotransformation of 17α-ethinylestradiol by the freshwater alga Desmodesmus subspicatus. Environ. Sci. Technol. 2014, 48, 12354–12361. [Google Scholar] [CrossRef]
- Ding, T.; Lin, K.; Yang, B.; Yang, M.; Li, J.; Li, W.; Gan, J. Biodegradation of naproxen by freshwater algae Cymbella sp. and Scenedesmus quadricauda and the comparative toxicity. Bioresour. Technol. 2017, 238, 164–173. [Google Scholar] [CrossRef]
- Ma, Y.; Shen, W.; Tang, T.; Li, Z.; Dai, R. Environmental estrogens in surface water and their interaction with microalgae: A review. Sci. Total Environ. 2022, 807, 150637. [Google Scholar] [CrossRef]
- Czarny, K.; Szczukocki, D.; Krawczyk, B.; Skrzypek, S.; Zieliński, M.; Gadzała-Kopciuch, R. Toxic effects of single animal hormones and their mixtures on the growth of Chlorella vulgaris and Scenedesmus armatus. Chemosphere 2019, 224, 93–102. [Google Scholar] [CrossRef]
- Balina, K.; Balode, M.; Muzikante, L.; Blumberga, D. Impact of synthetic hormone 17α-ethinylestradiol on growth of microalgae Desmodesmus communis. Agron. Res. 2015, 13, 445–454. [Google Scholar]
- Duan, L.; Chen, Q.; Duan, S. Transcriptional analysis of chlorella pyrenoidosa exposed to bisphenol A. Int. J. Environ. Res. Public Health 2019, 16, 1374. [Google Scholar] [CrossRef]
- Zhu, Q.; Zhang, M.; Bao, J.; Liu, J. Physiological, metabolomic, and transcriptomic analyses reveal the dynamic redox homeostasis upon extended exposure of Dunaliella salina GY-H13 cells to Cd. Ecotoxicol. Environ. Saf. 2021, 223, 112593. [Google Scholar] [CrossRef] [PubMed]
- Kang, W.; Li, X.; Sun, A.; Yu, F.; Hu, X. Study of the Persistence of the Phytotoxicity Induced by Graphene Oxide Quantum Dots and of the Specific Molecular Mechanisms by Integrating Omics and Regular Analyses. Environ. Sci. Technol. 2019, 53, 3791–3801. [Google Scholar] [CrossRef]
- Tripathi, S.; Poluri, K.M. Heavy metal detoxification mechanisms by microalgae: Insights from transcriptomics analysis. Environ. Pollut. 2021, 285, 117443. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Jiang, X.; Xiao, Y.; Li, M. Exposure of microalgae Euglena gracilis to polystyrene microbeads and cadmium: Perspective from the physiological and transcriptional responses. Aquat. Toxicol. 2020, 228, 105650. [Google Scholar] [CrossRef]
- Xiang, R.; Shi, J.; Zhang, H.; Dong, C.; Liu, L.; Fu, J.K.; He, X.; Yan, Y.; Wu, Z. Chlorophyll a fluorescence and transcriptome reveal the toxicological effects of bisphenol A on an invasive cyanobacterium, Cylindrospermopsis raciborskii. Aquat. Toxicol. 2018, 200, 188–196. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Wang, A.; Fu, W.; Song, D. Growth performance, antioxidant response, biodegradation and transcriptome analysis of Chlorella pyrenoidosa after nonylphenol exposure. Sci. Total Environ. 2022, 806, 150507. [Google Scholar] [CrossRef]
- Yang, M.; Fan, Z.; Xie, Y.; Fang, L.; Wang, X.; Yuan, Y.; Li, R. Transcriptome analysis of the effect of bisphenol A exposure on the growth, photosynthetic activity and risk of microcystin-LR release by Microcystis aeruginosa. J. Hazard. Mater. 2020, 397, 122746. [Google Scholar] [CrossRef]
- Li, Z.; Yi, X.; Zhou, H.; Chi, T.; Li, W.; Yang, K. Combined effect of polystyrene microplastics and dibutyl phthalate on the microalgae Chlorella pyrenoidosa. Environ. Pollut. 2019, 257, 113604. [Google Scholar] [CrossRef]
- Xu, D.; Li, C.; Chen, H.; Shao, B. Cellular response of freshwater green algae to perfluorooctanoic acid toxicity. Ecotoxicol. Environ. Saf. 2013, 88, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Dong, J.; Li, B.; Xue, C.; Tetteh, P.A.; Li, D.; Gao, K.; Deng, X. Using a freshwater green alga Chlorella pyrenoidosa to evaluate the biotoxicity of ionic liquids with different cations and anions. Ecotoxicol. Environ. Saf. 2020, 198, 110604. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Peñalver, A.; Reigosa, M.J.; Sánchez-Moreiras, A.M. Imaging chlorophyll a fluorescence reveals specific spatial distributions under different stress conditions. Flora Morphol. Distrib. Funct. Ecol. Plants 2011, 206, 836–844. [Google Scholar] [CrossRef]
- Xiong, J.Q.; Kurade, M.B.; Kim, J.R.; Roh, H.S.; Jeon, B.H. Ciprofloxacin toxicity and its co-metabolic removal by a freshwater microalga Chlamydomonas mexicana. J. Hazard. Mater. 2017, 323, 212–219. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Wang, X.; Yang, L.; Sun, Y.; Guo, H. Bioaccumulation and ROS generation in coontail Ceratophyllum demersum L. exposed to phenanthrene. Ecotoxicology 2010, 19, 1102–1110. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, G.; Dineshkumar, R.; Sen, R. Modelling of oxygen-evolving-complex ionization dynamics for energy-efficient production of microalgal biomass, pigment and lipid with carbon capture: An engineering vision for a biorefinery. RSC Adv. 2016, 6, 51941–51956. [Google Scholar] [CrossRef]
- Chen, S.; Yang, J.; Zhang, M.; Strasser, R.J.; Qiang, S. Classification and characteristics of heat tolerance in Ageratina adenophora populations using fast chlorophyll a fluorescence rise O-J-I-P. Environ. Exp. Bot. 2016, 122, 126–140. [Google Scholar] [CrossRef]
- Zaman, N.K.; Abdullah, M.Y.; Othman, S.; Zaman, N.K. Growth and Physiological Performance of Aerobic and Lowland Rice as Affected by Water Stress at Selected Growth Stages. Rice Sci. 2018, 25, 82–93. [Google Scholar] [CrossRef]
- Zhang, F.; Yao, X.; Sun, S.; Wang, L.; Liu, W.; Jiang, X.; Wang, J. Effects of mesotrione on oxidative stress, subcellular structure, and membrane integrity in Chlorella vulgaris. Chemosphere 2020, 247, 125668. [Google Scholar] [CrossRef]
- Ma, Y.; Lu, M.; Li, J.Y.; Qin, Y.; Gong, X.G. Evolutive and structural characterization of Nostoc commune iron-superoxide dismutase that is fit for modification. Genet. Mol. Res. 2012, 11, 3607–3617. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Guan, Y.; Gao, B.; Yue, Q. Antioxidant responses and degradation of two antibiotic contaminants in Microcystis aeruginosa. Ecotoxicol. Environ. Saf. 2012, 86, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Liu, Y.; Zhang, J. Mechanisms for the stimulatory effects of a five-component mixture of antibiotics in Microcystis aeruginosa at transcriptomic and proteomic levels. J. Hazard. Mater. 2021, 406, 124722. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Cheng, J.; Wang, Z.; Zhang, X.; Chen, S.; Zhou, J. Enhancing Photosynthetic Characterization and Biomass Productivity of Nannochloropsis Oceanica by Nuclear Radiation. Front. Energy Res. 2020, 8, 143. [Google Scholar] [CrossRef]
- Zhang, Z.; Sun, D.; Zhang, Y.; Chen, F. Chloroplast morphogenesis in Chromochloris zofingiensis in the dark. Algal Res. 2020, 45, 101742. [Google Scholar] [CrossRef]
- Yang, W.; Gao, P.; Li, H.; Huang, J.; Zhang, Y.; Ding, H.; Zhang, W. Mechanism of the inhibition and detoxification effects of the interaction between nanoplastics and microalgae Chlorella pyrenoidosa. Sci. Total Environ. 2021, 783, 146919. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Ma, Z.; Peng, J.; Mo, J.; Li, Q.; Guo, J.; Yang, F. Transcriptomic analysis of Raphidocelis subcapitata exposed to erythromycin: The role of DNA replication in hormesis and growth inhibition. J. Hazard. Mater. 2021, 402, 123512. [Google Scholar] [CrossRef] [PubMed]
- Luo, S.W.; Alimujiang, A.; Balamurugan, S.; Zheng, J.W.; Wang, X.; Yang, W.D.; Cui, J.; Li, H.Y. Physiological and molecular responses in halotolerant Dunaliella salina exposed to molybdenum disulfide nanoparticles. J. Hazard. Mater. 2021, 404, 124014. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.; Guissé, B.; Greppin, H.; Strasser, R.J. Regulation of antenna structure and electron transport in Photosystem II of Pisum sativum under elevated temperature probed by the fast polyphasic chlorophyll a fluorescence transient: OKJIP. Biochim. Biophys. Acta—Bioenerg. 1997, 1320, 95–106. [Google Scholar] [CrossRef]
- Zhong, X.; Li, Y.; Che, X.; Zhang, Z.; Li, Y.; Liu, B.; Li, Q.; Gao, H. Significant inhibition of photosynthesis and respiration in leaves of Cucumis sativus L. by oxybenzone, an active ingredient in sunscreen. Chemosphere 2019, 219, 456–462. [Google Scholar] [CrossRef]
- Sabatini, S.E.; Juárez, Á.B.; Eppis, M.R.; Bianchi, L.; Luquet, C.M.; del Carmen Ríos de Molina, M. Oxidative stress and antioxidant defenses in two green microalgae exposed to copper. Ecotoxicol. Environ. Saf. 2009, 72, 1200–1206. [Google Scholar] [CrossRef]
- He, Y.Y.; Häder, D.P. UV-B-induced formation of reactive oxygen species and oxidative damage of the cyanobacterium Anabaena sp.: Protective effects of ascorbic acid and N-acetyl-L-cysteine. J. Photochem. Photobiol. B Biol. 2002, 66, 115–124. [Google Scholar] [CrossRef]
- Mao, Y.; Ai, H.; Chen, Y.; Zhang, Z.; Zeng, P.; Kang, L.; Li, W.; Gu, W.; He, Q.; Li, H. Phytoplankton response to polystyrene microplastics: Perspective from an entire growth period. Chemosphere 2018, 208, 59–68. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Q.; Zhou, Q.; Jin, Z.; Jiang, Y.; Xu, L.; Jiang, H.; Zhao, Y. Bioaccumulation, growth performance, and transcriptomic response of Dictyosphaerium sp. after exposure to nonylphenol. Sci. Total Environ. 2019, 687, 416–422. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. J. Embnet J. 2011, 17, 1. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2014, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Chen, Z.; Tao, Y.; Wu, W.; Zeng, Y.; Liao, K.; Li, X.; Chen, L. Transcriptomic and Physiological Responses of Chlorella pyrenoidosa during Exposure to 17α-Ethinylestradiol. Int. J. Mol. Sci. 2022, 23, 3583. https://doi.org/10.3390/ijms23073583
Zhang Y, Chen Z, Tao Y, Wu W, Zeng Y, Liao K, Li X, Chen L. Transcriptomic and Physiological Responses of Chlorella pyrenoidosa during Exposure to 17α-Ethinylestradiol. International Journal of Molecular Sciences. 2022; 23(7):3583. https://doi.org/10.3390/ijms23073583
Chicago/Turabian StyleZhang, Yurui, Zixu Chen, Yue Tao, Wanyin Wu, Yuyang Zeng, Kejun Liao, Xinyue Li, and Lanzhou Chen. 2022. "Transcriptomic and Physiological Responses of Chlorella pyrenoidosa during Exposure to 17α-Ethinylestradiol" International Journal of Molecular Sciences 23, no. 7: 3583. https://doi.org/10.3390/ijms23073583
APA StyleZhang, Y., Chen, Z., Tao, Y., Wu, W., Zeng, Y., Liao, K., Li, X., & Chen, L. (2022). Transcriptomic and Physiological Responses of Chlorella pyrenoidosa during Exposure to 17α-Ethinylestradiol. International Journal of Molecular Sciences, 23(7), 3583. https://doi.org/10.3390/ijms23073583




