Genome-Wide Identification and Expression Analysis of the 14-3-3 Gene Family in Mango (Mangifera indica L.)
Abstract
:1. Introduction
2. Results
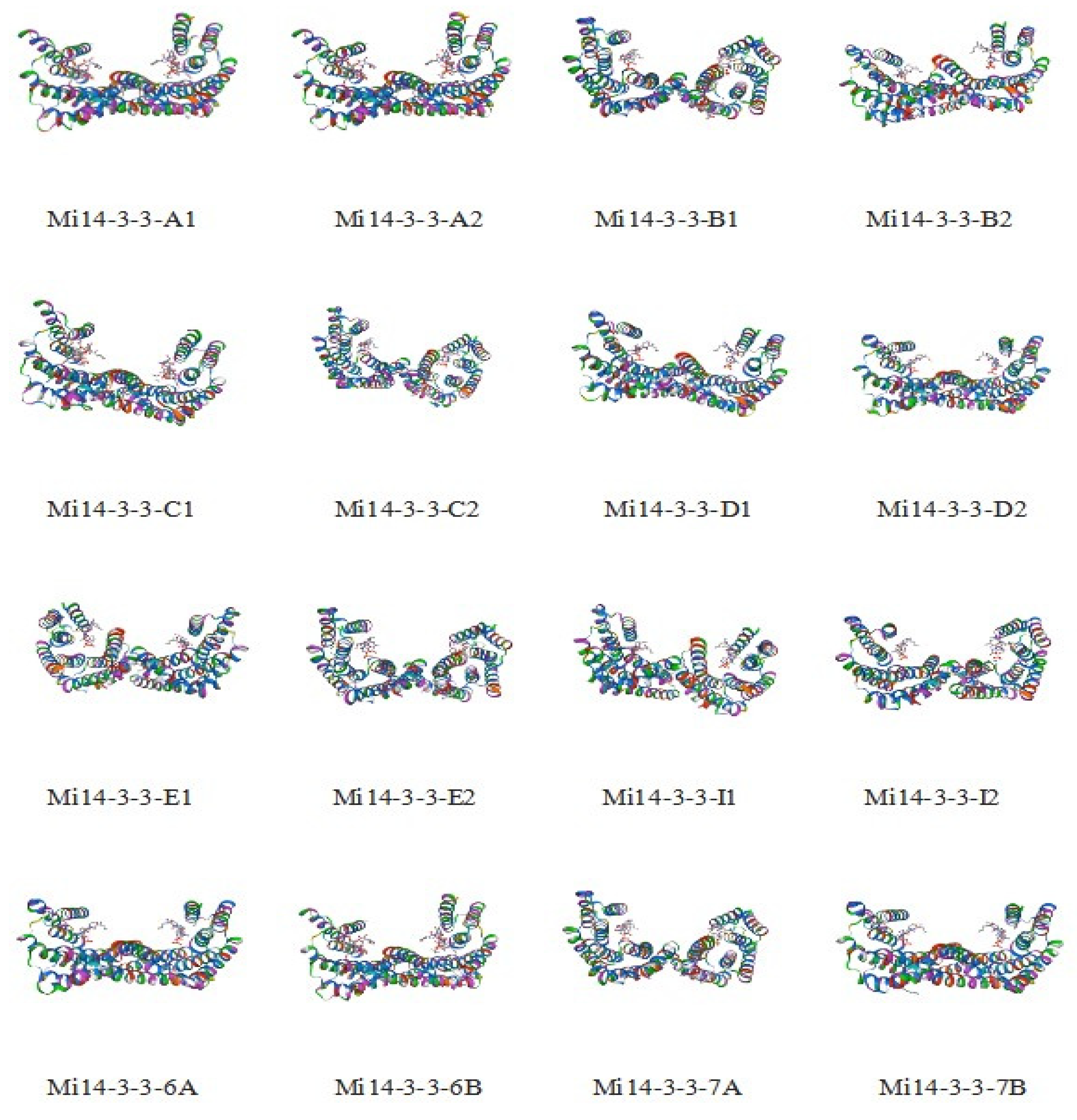
2.1. Genome-Wide Identification of Mi14-3-3 Gene Family Members and Characterization of Their Proteins
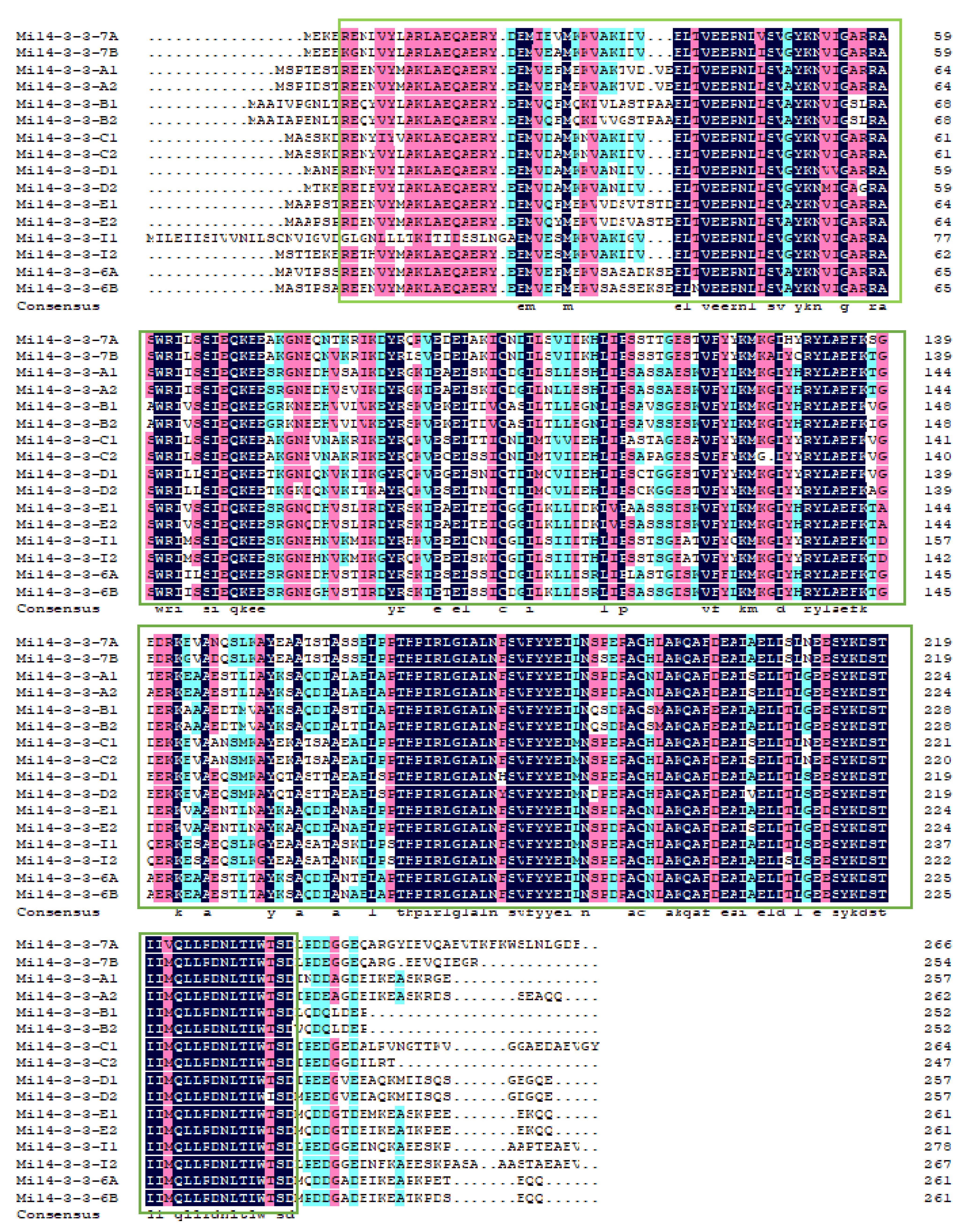
2.2. Multiple Sequence Alignment of Mi14-3-3 Proteins
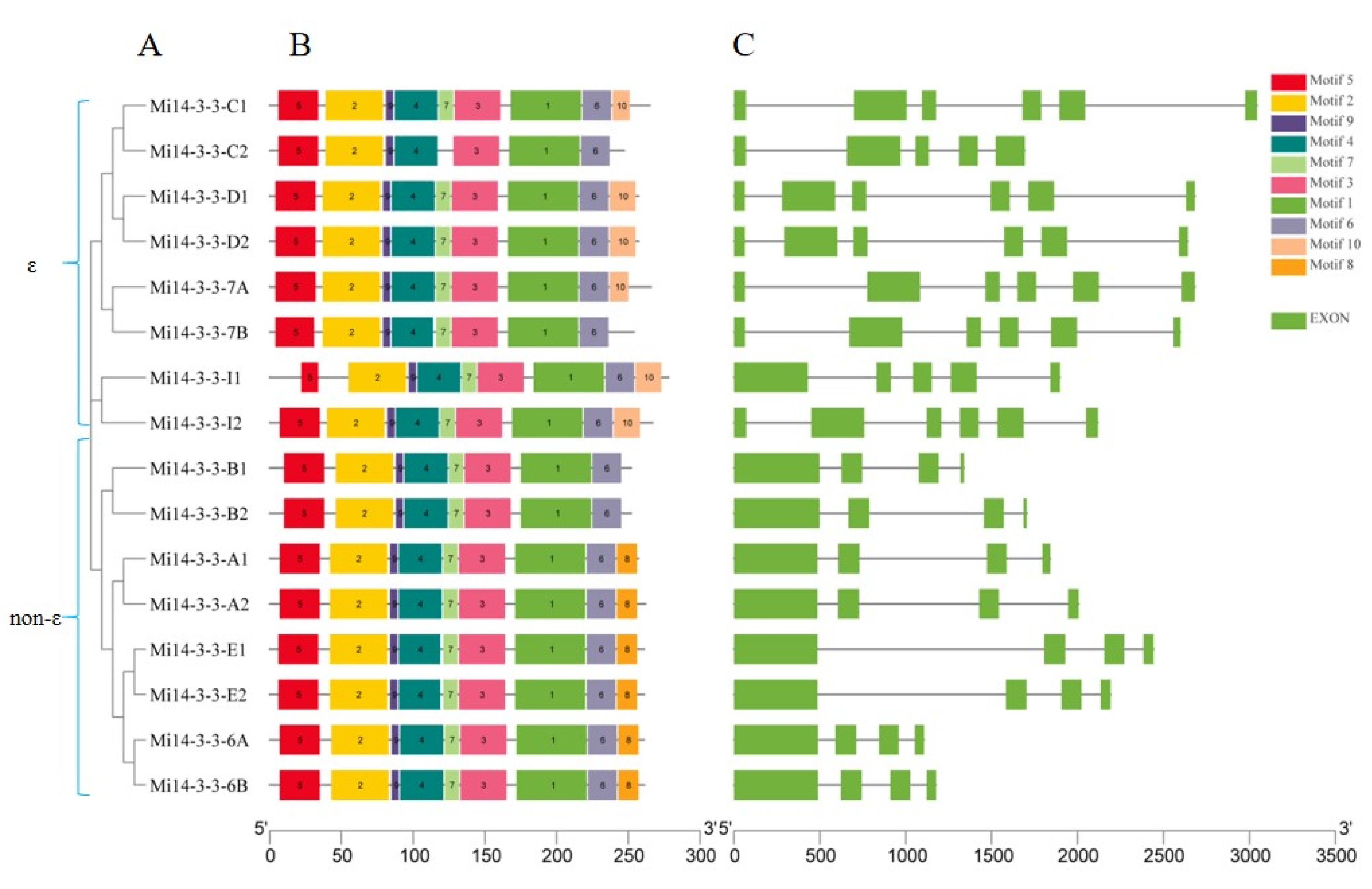
2.3. Gene structure and Motif Analysis
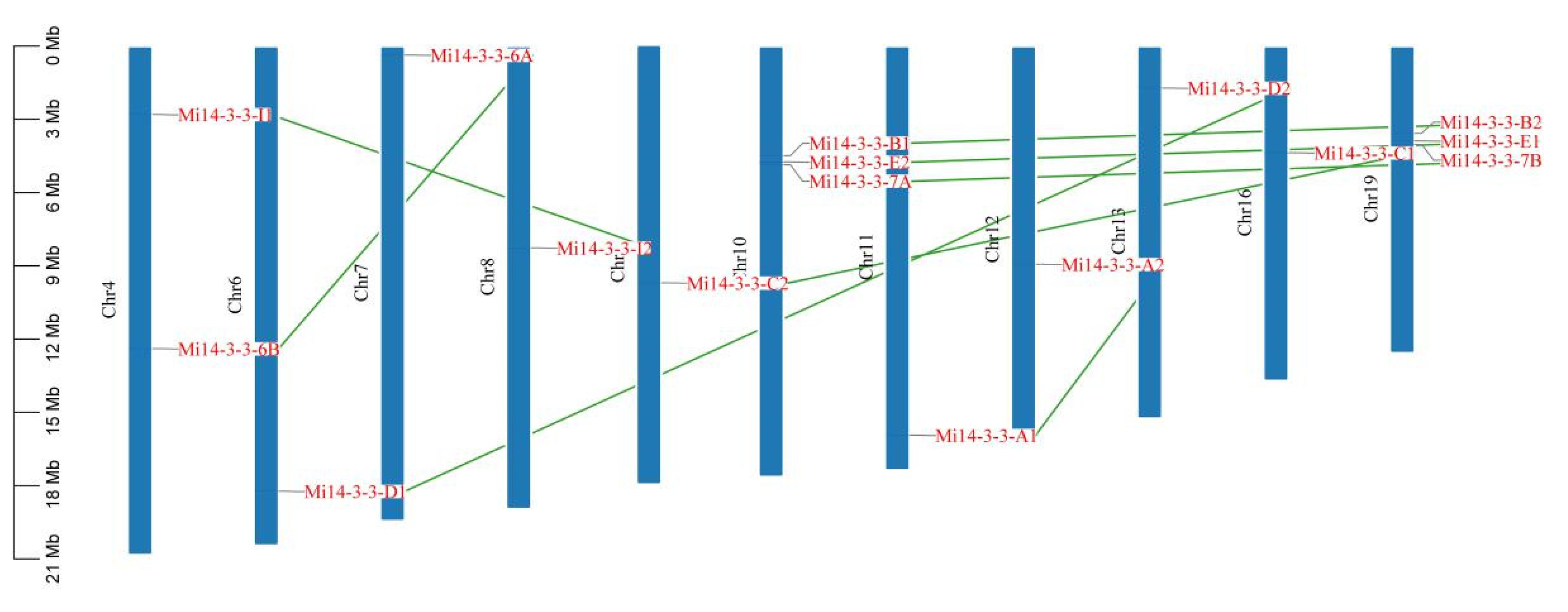
2.4. Chromosome Distribution of 14-3-3 Gene Family Members in Mango
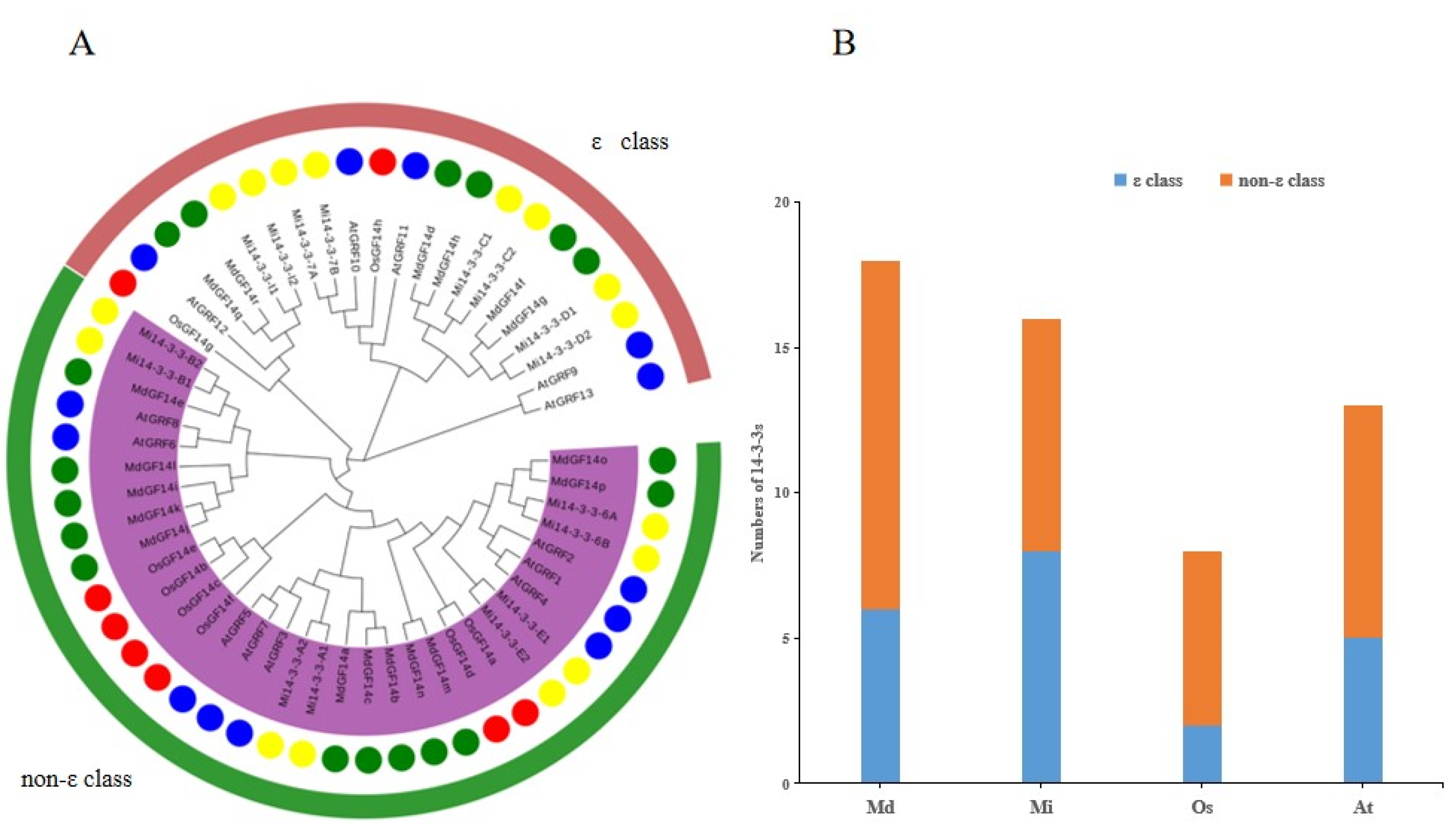
2.5. Phylogenetic Analysis of the Mi14-3-3 Genes
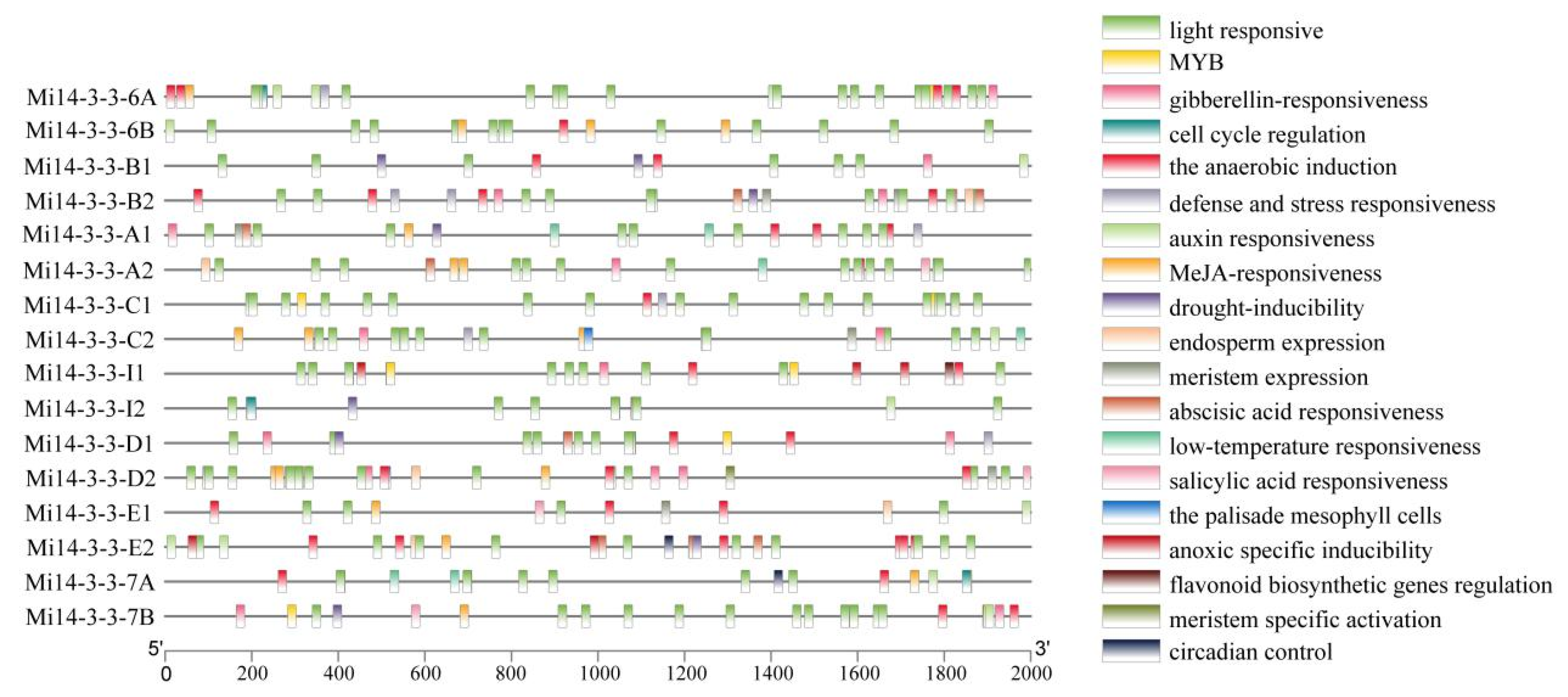
2.6. Analysis of the Cis-Regulatory Elements of Mi14-3-3 Gene Family Member Promoters
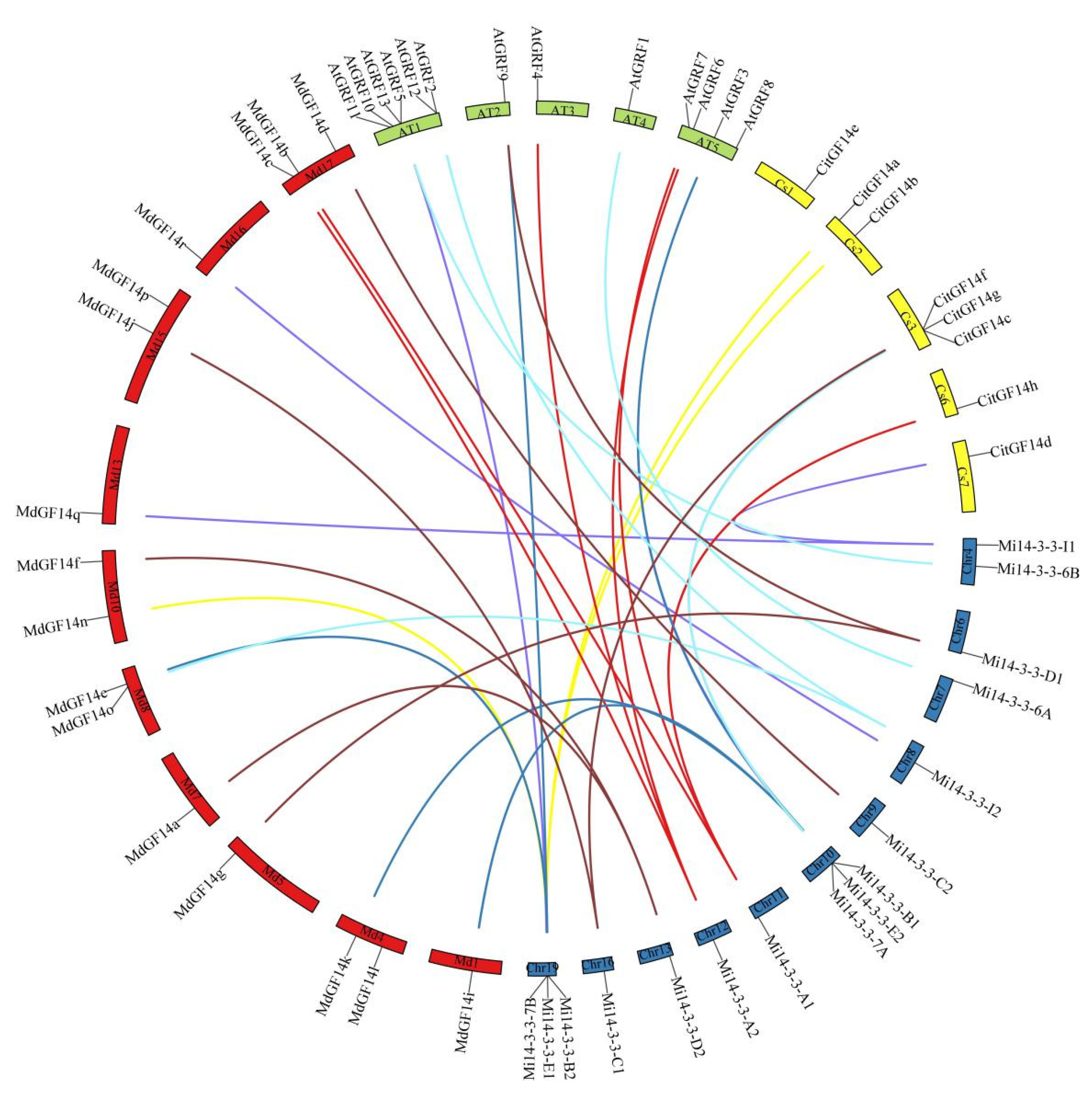
2.7. Synteny Analysis of Mi14-3-3 Genes
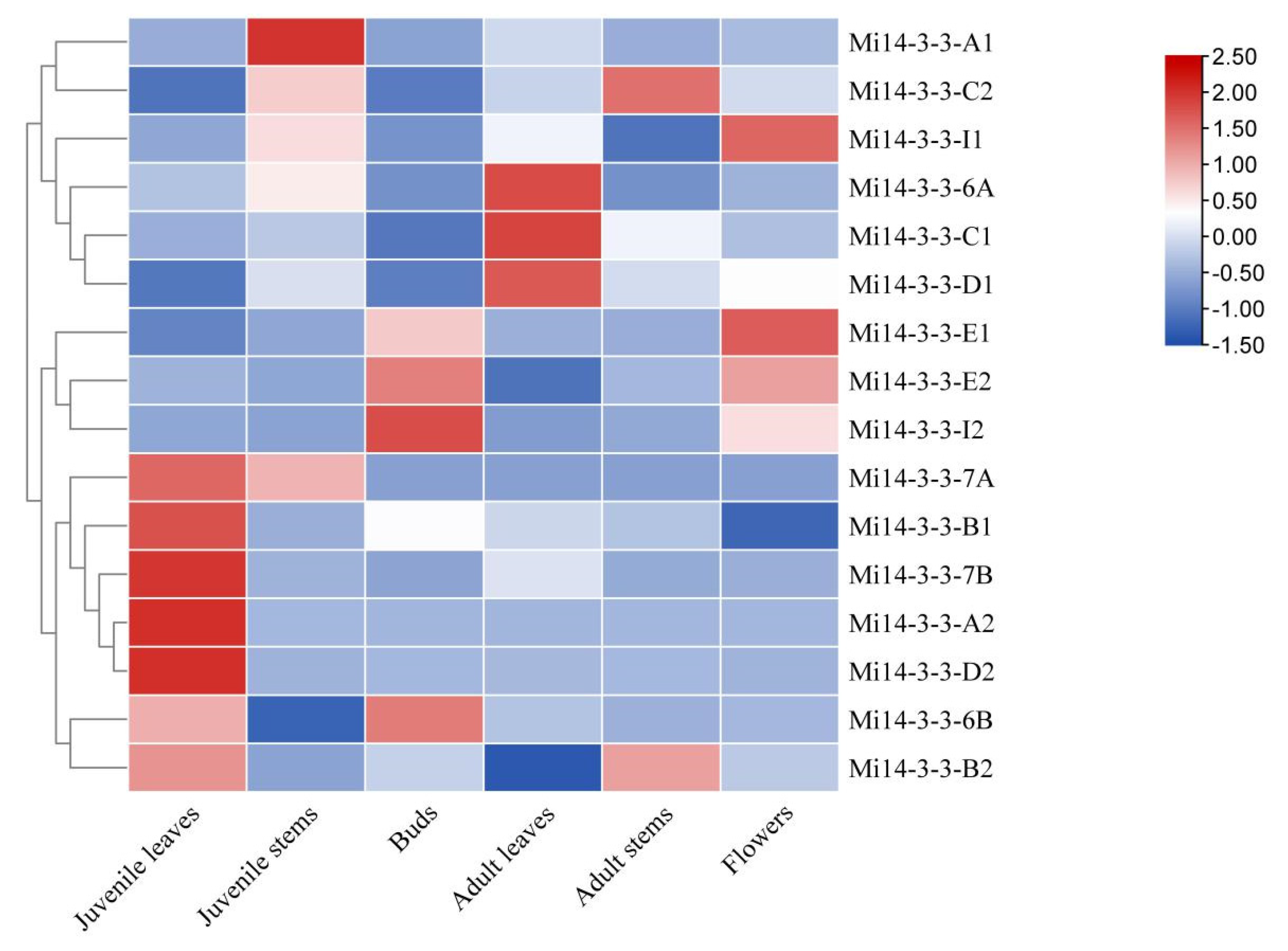
2.8. Expression Profiles of Mi14-3-3 Genes in Different Tissues
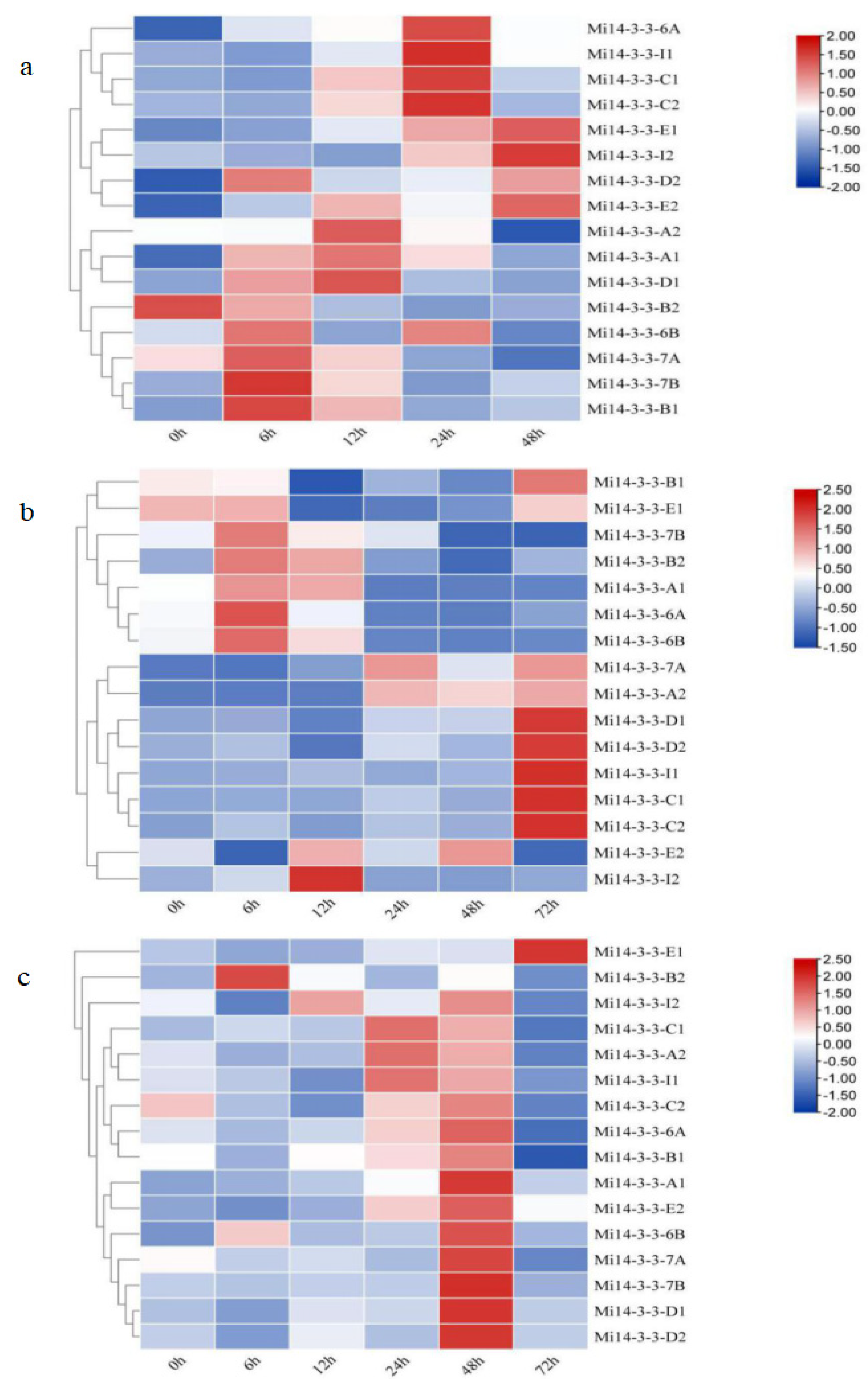
2.9. Expression Patterns of Mi14-3-3s under Abiotic Stress
3. Discussion
4. Materials and Methods
4.1. Materials and Treatments
4.2. Identification and Sequence Analysis of 14-3-3 Gene Family Members in Mango
4.3. Chromosomal Location and Gene Structure Analyses
4.4. Sequence Alignment and Phylogenetic Analyses
4.5. Prediction of Cis-Regulatory Elements
4.6. Tandem Duplication and Synteny Analysis
4.7. RNA Extraction and qRT–PCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board statement
Informed consent statement
Data availability statement
Conflicts of Interest
References
- Camoni, L.; Visconti, S.; Aducci, P.; Marra, M. 14-3-3 proteins in Plant Hormone Signaling: Doing Several Things at Once. Front. Plant Sci. 2018, 9, 297. [Google Scholar] [CrossRef] [PubMed]
- Pnueli, L.; Gutfinger, T.; Hareven, D.; Ben-Naim, O.; Ron, N.; Adir, N.; Lifschitz, E. Tomato SP-interacting proteins define a conserved signaling system that regulates shoot architecture and flowering. Plant Cell 2001, 13, 2687–2702. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, B.; Xiao, G.; Luo, K.; Wang, Z.; Mao, B.; Lin, X.; Guo, X. Overexpression of PvGF14c from Phyllostachys violascens Delays Flowering Time in Transgenic Arabidopsis. Front. Plant Sci. 2018, 9, 105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mayfield, J.D.; Folta, K.M.; Pau, A.L.; Ferl, R.J. The 14-3-3 proteins mu and upsilon influence transition to flowering and early phytochrome response. Plant Physiol. 2007, 145, 1692–1702. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaneko-Suzuki, M.; Kurihara-Ishikawa, R.; Okushita-Terakawa, C.; Kojima, C.; Nagano-Fujiwara, M.; Ohki, I.; Tsuji, H.; Shimamoto, K.; Taoka, K.I. TFL1-Like Proteins in Rice Antagonize Rice FT-Like Protein in Inflorescence Development by Competition for Complex Formation with 14-3-3 and FD. Plant Cell Physiol. 2018, 59, 458–468. [Google Scholar] [CrossRef] [Green Version]
- Schoonheim, P.J.; Costa Pereira, D.D.; De Boer, A.H. Dual role for 14-3-3 proteins and ABF transcription factors in gibberellic acid and abscisic acid signalling in barley (Hordeum vulgare) aleurone cells. Plant Cell Environ. 2009, 32, 439–447. [Google Scholar] [CrossRef]
- Xu, W.; Jia, L.; Shi, W.; Baluska, F.; Kronzucker, H.J.; Liang, J.; Zhang, J. The Tomato 14-3-3 protein TFT4 modulates H+ efflux, basipetal auxin transport, and the PKS5-J3 pathway in the root growth response to alkaline stress. Plant Physiol. 2013, 163, 1817–1828. [Google Scholar] [CrossRef] [Green Version]
- Xu, W.F.; Shi, W.M. Expression profiling of the 14-3-3 gene family in response to salt stress and potassium and iron deficiencies in young tomato (Solanum lycopersicum) roots: Analysis by real-time RT-PCR. Ann. Bot. 2006, 98, 965–974. [Google Scholar] [CrossRef] [Green Version]
- Catalá, R.; López-Cobollo, R.; Mar-Castellano, M.; Angosto, T.; Alonso, J.M.; Ecker, J.R.; Salinas, J. The Arabidopsis 14-3-3 protein RARE COLD INDUCIBLE 1A links low-temperature response and ethylene biosynthesis to regulate freezing tolerance and cold acclimation. Plant Cell 2014, 26, 3326–3342. [Google Scholar] [CrossRef] [Green Version]
- Sun, X.; Luo, X.; Sun, M.; Chen, C.; Ding, X.; Wang, X.; Yang, S.; Yu, Q.; Jia, B.; Ji, W.; et al. A Glycine soja 14-3-3 protein GsGF14o participates in stomatal and root hair development and drought tolerance in Arabidopsis thaliana. Plant Cell Physiol. 2014, 55, 99–118. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Zhou, X.; Chang, S.; Chu, Z.; Wang, H.; Han, S.; Wang, Y. Calcium-dependent protein kinase 21 phosphorylates 14-3-3 proteins in response to ABA signaling and salt stress in rice. Biochem. Biophys. Res. Commun. 2017, 493, 1450–1456. [Google Scholar] [CrossRef] [PubMed]
- Mayfield, J.D.; Paul, A.L.; Ferl, R.J. The 14-3-3 proteins of Arabidopsis regulate root growth and chloroplast development as components of the photosensory system. J. Exp. Bot. 2012, 63, 3061–3070. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.; David, H.; Tuan, H.; Liu, L.H.; Lee, D.H.; Lee, C.H.; Chen, Y.R.; Lin, S.Y.; Lu, C.A.; Yu, S.M. Sugar starvation-regulated MYBS2 and 14-3-3 protein interactions enhance plant growth, stress tolerance, and grain weight in rice. Proc. Natl. Acad. Sci. USA 2019, 116, 21925–21935. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, L.; You, J.; Wang, Y.; Li, J.; Quan, W.; Yin, M.; Wang, Q.; Chan, Z. Systematic analysis of the G-box Factor 14-3-3 gene family and functional characterization of GF14a in Brachypodium distachyon. Plant Physiol. Biochem. 2017, 117, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.P.; Li, H.L.; Guo, D.; Tang, X.; Peng, S.Q. Identification and characterization of the 14-3-3 gene family in Hevea brasiliensis. Plant Physiol. Biochem. 2014, 80, 121–127. [Google Scholar] [CrossRef]
- Purwestri, Y.A.; Ogaki, Y.; Tamaki, S.; Tsuji, H.; Shimamoto, K. The 14-3-3 protein GF14c acts as a negative regulator of flowering in rice by interacting with the florigen Hd3a. Plant Cell Physiol. 2009, 50, 429–438. [Google Scholar] [CrossRef] [Green Version]
- Taoka, K.; Ohki, I.; Tsuji, H.; Furuita, K.; Hayashi, K.; Yanase, T.; Yamaguchi, M.; Nakashima, C.; Purwestri, Y.A.; Tamaki, S.; et al. 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature 2011, 476, 332–335. [Google Scholar] [CrossRef]
- Cao, J.; Cao, J.F.; Huang, J.Q.; Liu, X.; Huang, C.C.; Zheng, Z.S.; Zhang, X.F.; Shangguan, X.X.; Wang, L.J.; Zhang, Y.G.; et al. Genome-wide characterization of the GRF family and their roles in response to salt stress in Gossypium. BMC Genom. 2020, 21, 575. [Google Scholar] [CrossRef]
- Rosenquist, M.; Alsterfjord, M.; Larsson, C.; Sommarin, M. Data Mining the Arabidopsis genome reveals fifteen 14-3-3 genes, expression is demonstrated for two out of five novel genes1. Plant Physiol. 2001, 127, 142–149. [Google Scholar] [CrossRef] [Green Version]
- Chen, F.; Li, Q.; Sun, L.; He, Z. The rice 14-3-3 gene family and its involvement in responses to biotic and abiotic stress. DNA Res. 2006, 13, 53–63. [Google Scholar] [CrossRef] [Green Version]
- Li, X.; Dhaubhadel, S. Soybean 14-3-3 gene family: Identification and molecular characterization. Planta 2011, 233, 569–582. [Google Scholar] [CrossRef] [PubMed]
- Konagaya, K.; Matsushita, Y.; Kasahara, M.; Nyunoya, H. Members of 14-3-3 protein isoforms interacting with the resistance gene product N and the elicitor of Tobacco mosaic virus. J. Gen. Plant Pathol. 2004, 70, 221–231. [Google Scholar] [CrossRef]
- Tian, F.; Wang, T.; Xie, Y.; Zhang, J.; Hu, J. Genome-wide identification, classification, and expression analysis of 14-3-3 gene family in Populus. PLoS ONE. 2015, 10, e0123225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zuo, X.; Wang, S.; Xiang, W.; Yang, H.; Tahir, M.M.; Zheng, S.; An, N.; Han, M.; Zhao, C.; Zhang, D. Genome-wide identification of the 14-3-3 gene family and its participation in floral transition by interacting with TFL1/FT in apple. BMC Genom. 2021, 22, 41. [Google Scholar] [CrossRef] [PubMed]
- Lyu, S.; Chen, G.; Pan, D.; Chen, J.; She, W. Molecular Analysis of 14-3-3 Genes in Citrus sinensis and Their Responses to Different Stresses. Int. J Mol Sci. 2021, 22, 568. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, A.B.; Anamika, K.; Jha, V.; Chidley, H.G.; Oak, P.S.; Kadoo, N.Y.; Pujari, K.H.; Giri, A.P.; Gupta, V.S. Transcriptional transitions in Alphonso mango (Mangifera indica L.) during fruit development and ripening explain its distinct aroma and shelf life characteristics. Sci. Rep. 2017, 7, 8711. [Google Scholar] [CrossRef] [PubMed]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef] [Green Version]
- Yang, X.W.; Lee, W.H.; Sobott, F.; Papagrigoriou, E.; Robinson, C.V.; Grossmann, J.G.; Sundström, M.; Doyle, D.A.; Elkins, J.M. Structural basis for protein-protein interactions in the 14-3-3 protein family. Proc. Natl. Acad. Sci. USA 2006, 103, 17237–17242. [Google Scholar] [CrossRef] [Green Version]
- Aducci, P.; Camoni, L.; Marra, M.; Visconti, S. From cytosol to organelles: 14-3-3 proteins as multifunctional regulators of plant cell. IUBMB Life 2002, 53, 49–55. [Google Scholar] [CrossRef]
- Sullivan, S.; Thomson, C.E.; Kaiserli, E.; Christie, J.M. Interaction specificity of Arabidopsis 14-3-3 proteins with phototropin receptor kinases. FEBS Lett. 2009, 583, 2187–2193. [Google Scholar] [CrossRef] [Green Version]
- Folta, K.M.; Paul, A.; Mayfield, J.D.; Ferl, R.J. 14-3-3 isoforms participate in red light signaling and photoperiodic flowering. Plant Signal. Behav. 2008, 3, 304–306. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schoonheim, P.J.; Sinnige, M.P.; Casaretto, J.A.; Veiga, H.; Bunney, T.D.; Quatrano, R.S.; De Boer, A.H. 14-3-3 adaptor proteins are intermediates in ABA signal transduction during barley seed germination. Plant J. 2007, 49, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Ishida, S.; Fukazawa, J.; Yuasa, T.; Takahashi, Y. Involvement of 14-3-3 signaling protein binding in the functional regulation of the transcriptional activator REPRESSION OF SHOOT GROWTH by gibberellins. Plant Cell 2004, 16, 2641–2651. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoon, G.M.; Kieber, J.J. 14-3-3 regulates 1-aminocyclopropane-1-carboxylate synthase Protein turnover in Arabidopsis. Plant Cell 2013, 25, 1016–1028. [Google Scholar] [CrossRef] [Green Version]
- Keicher, J.; Jaspert, N.; Weckermann, K.; Moller, C.; Throm, C.; Kintzi, A.; Oecking, C. Arabidopsis 14-3-3 epsilon members contribute to polarity of PIN auxin carrier and auxin transport-related development. eLife 2017, 6, 605. [Google Scholar] [CrossRef]
- Liang, M.; Ow, D.W. Nucleocytoplasmic OXIDATIVE STRESS 2 can relocate FLOWERING LOCUS T. Biochem. Biophys. Res. Commun. 2019, 517, 735–740. [Google Scholar] [CrossRef]
- Ishida, S.; Yuasa, T.; Nakata, M.; Takahashi, Y. A tobacco calcium-dependent protein kinase, CDPK1, regulates the transcription factor REPRESSION OF SHOOT GROWTH in response to gibberellins. Plant Cell 2008, 20, 3273–3288. [Google Scholar] [CrossRef] [Green Version]
- Ormancey, M.; Thuleau, P.; Mazars, C.; Cotelle, V. CDPKs and 14-3-3 proteins: Emerging Duo in Signaling. Trends Plant Sci. 2017, 22, 263–272. [Google Scholar] [CrossRef]
- Liu, J.; Sun, X.; Liao, W.; Zhang, J.; Liang, J.; Xu, W. Involvement of OsGF14b Adaptation in the Drought Resistance of Rice Plants. Rice 2019, 12, 82. [Google Scholar] [CrossRef] [Green Version]
- Chen, B.Z.; Wang, X.Y.; Lv, J.Y.; Ge, M.J.; Qiao, K.K.; Chen, Q.J.; Zhang, K.P.; Wang, J.S.; Fan, S.L.; Ma, Q.F. GhN/AINV13 positively regulates cotton stress tolerance by interacting with the 14-3-3 protein. Genomics. 2021, 113, 44–56. [Google Scholar] [CrossRef]
- He, Y.; Wu, J.; Lv, B.; Li, J.; Gao, Z.; Xu, W.; Baluška, F.; Shi, W.; Shaw, P.C.; Zhang, J. Involvement of 14-3-3 protein GRF9 in root growth and response under polyethylene glycol-induced water stress. J. Exp. Bot. 2015, 66, 2271–2281. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Campo, S.; Peris-Peris, C.; Montesinos, L.; Peñas, G.; Messeguer, J.; San, S.B. Expression of the maize ZmGF14-6 gene in rice confers tolerance to drought stress while enhancing susceptibility to pathogen infection. J Exp Bot. 2012, 63, 983–999. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, J.M.; Jiang, C.Y.; Kang, L.; Zhang, H.C.; Song, Y.; Zou, Z.R.; Zheng, W.J. Over-Expression of a 14-3-3 protein From Foxtail Millet Improves Plant Tolerance to Salinity Stress in Arabidopsis thaliana. Front. Plant Sci. 2020, 11, 449. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ren, Y.; Yang, Y.Y.; Zhang, R.; You, C.X.; Zhao, Q.; Hao, Y.J. MdGRF11, an apple 14-3-3 protein, acts as a positive regulator of drought and salt tolerance. Plant Sci. 2019, 288, 110219. [Google Scholar] [CrossRef] [PubMed]
- Poole, R.L. The TAIR database. Methods Mol. Biol. 2007, 406, 179–212. [Google Scholar] [PubMed]
- Letunic, I.; Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 2018, 46, D493–D496. [Google Scholar] [CrossRef]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server; Walker, J.M., Ed.; The Proteomics Protocols Handbook University of Hertfordshire: Hatfield, UK, 2005; pp. 571–607. [Google Scholar]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol Biol. 1999, 112, 531–552. [Google Scholar]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef] [Green Version]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2014, 31, 1296–1297. [Google Scholar] [CrossRef] [Green Version]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gu, Z.; Cavalcanti, A.; Chen, F.C.; Bouman, P.; Li, W.H. Extent of gene duplication in the genomes of Drosophila, nematode, and yeast. Mol. Biol. Evol. 2002, 19, 256–262. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Yue, H.; Feng, K.; Deng, P.; Song, W.; Nie, X. Genome-wide identification, phylogeny and expressional profiles of mitogen activated protein kinase kinase kinase (MAPKKK) gene family in bread wheat (Triticum aestivum L.). BMC Genom. 2016, 17, 668. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
| Gene Name | Gene Locus | Chr | aa | pI | II | MW (Da) | Gene Entry Number |
|---|---|---|---|---|---|---|---|
| Mi14-3-3-A1 | 15934509- 15936351 | Chr11 | 257 | 4.78 | 47.14 | 29,051.64 | OK491862 |
| Mi14-3-3-A2 | 8935256- 8937263 | Chr12 | 262 | 4.74 | 48.11 | 29,631.29 | OK491863 |
| Mi14-3-3-B1 | 4489787- 4491126 | Chr10 | 252 | 4.83 | 37.22 | 28,431.38 | OK491864 |
| Mi14-3-3-B2 | 3572447- 3574125 | Chr19 | 252 | 4.79 | 39.12 | 28,503.45 | OK491865 |
| Mi14-3-3-C1 | 4366341- 4369386 | Chr16 | 265 | 4.80 | 35.13 | 29,899.64 | OK491866 |
| Mi14-3-3-C2 | 9746038- 9747732 | Chr9 | 247 | 4.85 | 41.86 | 28,078.73 | OK491867 |
| Mi14-3-3-D1 | 18226835- 18229518 | Chr6 | 257 | 4.77 | 46.99 | 29,284.95 | OK491868 |
| Mi14-3-3-D2 | 1716202- 1718844 | Chr13 | 257 | 4.68 | 45.77 | 29,422.24 | OK491869 |
| Mi14-3-3-E1 | 3882198- 3883306 | Chr19 | 261 | 4.68 | 44.49 | 29,583.16 | OK491870 |
| Mi14-3-3-E2 | 4744386- 4745564 | Chr10 | 261 | 4.69 | 51.96 | 29,607.16 | OK491871 |
| Mi14-3-3-I1 | 2799060- 2800971 | Chr4 | 278 | 4.83 | 49.60 | 30,968.14 | OK491872 |
| Mi14-3-3-I2 | 8264404- 8266523 | Chr8 | 267 | 4.93 | 51.64 | 30,230.90 | OK491873 |
| Mi14-3-3-6A | 364937- 367380 | Chr7 | 261 | 4.72 | 48.41 | 29,541.14 | OK203791 |
| Mi14-3-3-6B | 12389595- 12391788 | Chr4 | 261 | 4.76 | 50.54 | 29,376.81 | OK203792 |
| Mi14-3-3-7A | 4859531- 4862214 | Chr10 | 266 | 5.01 | 42.85 | 30,506.27 | OK491860 |
| Mi14-3-3-7B | 4027530- 4010130 | Chr19 | 254 | 4.85 | 45.50 | 28,807.40 | OK491861 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xia, L.; He, X.; Huang, X.; Yu, H.; Lu, T.; Xie, X.; Zeng, X.; Zhu, J.; Luo, C. Genome-Wide Identification and Expression Analysis of the 14-3-3 Gene Family in Mango (Mangifera indica L.). Int. J. Mol. Sci. 2022, 23, 1593. https://doi.org/10.3390/ijms23031593
Xia L, He X, Huang X, Yu H, Lu T, Xie X, Zeng X, Zhu J, Luo C. Genome-Wide Identification and Expression Analysis of the 14-3-3 Gene Family in Mango (Mangifera indica L.). International Journal of Molecular Sciences. 2022; 23(3):1593. https://doi.org/10.3390/ijms23031593
Chicago/Turabian StyleXia, Liming, Xinhua He, Xing Huang, Haixia Yu, Tingting Lu, Xiaojie Xie, Xuemei Zeng, Jiawei Zhu, and Cong Luo. 2022. "Genome-Wide Identification and Expression Analysis of the 14-3-3 Gene Family in Mango (Mangifera indica L.)" International Journal of Molecular Sciences 23, no. 3: 1593. https://doi.org/10.3390/ijms23031593
APA StyleXia, L., He, X., Huang, X., Yu, H., Lu, T., Xie, X., Zeng, X., Zhu, J., & Luo, C. (2022). Genome-Wide Identification and Expression Analysis of the 14-3-3 Gene Family in Mango (Mangifera indica L.). International Journal of Molecular Sciences, 23(3), 1593. https://doi.org/10.3390/ijms23031593






