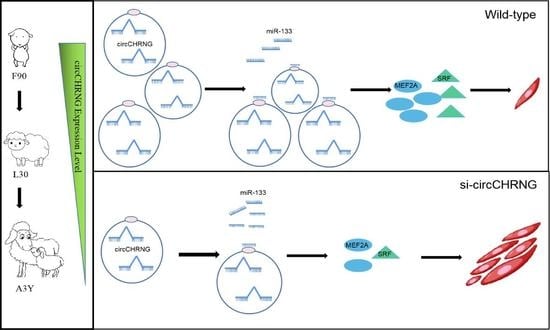
Genome-Wide Analysis of Circular RNAs Reveals circCHRNG Regulates Sheep Myoblast Proliferation via miR-133/SRF and MEF2A Axis
Abstract
1. Introduction
2. Results
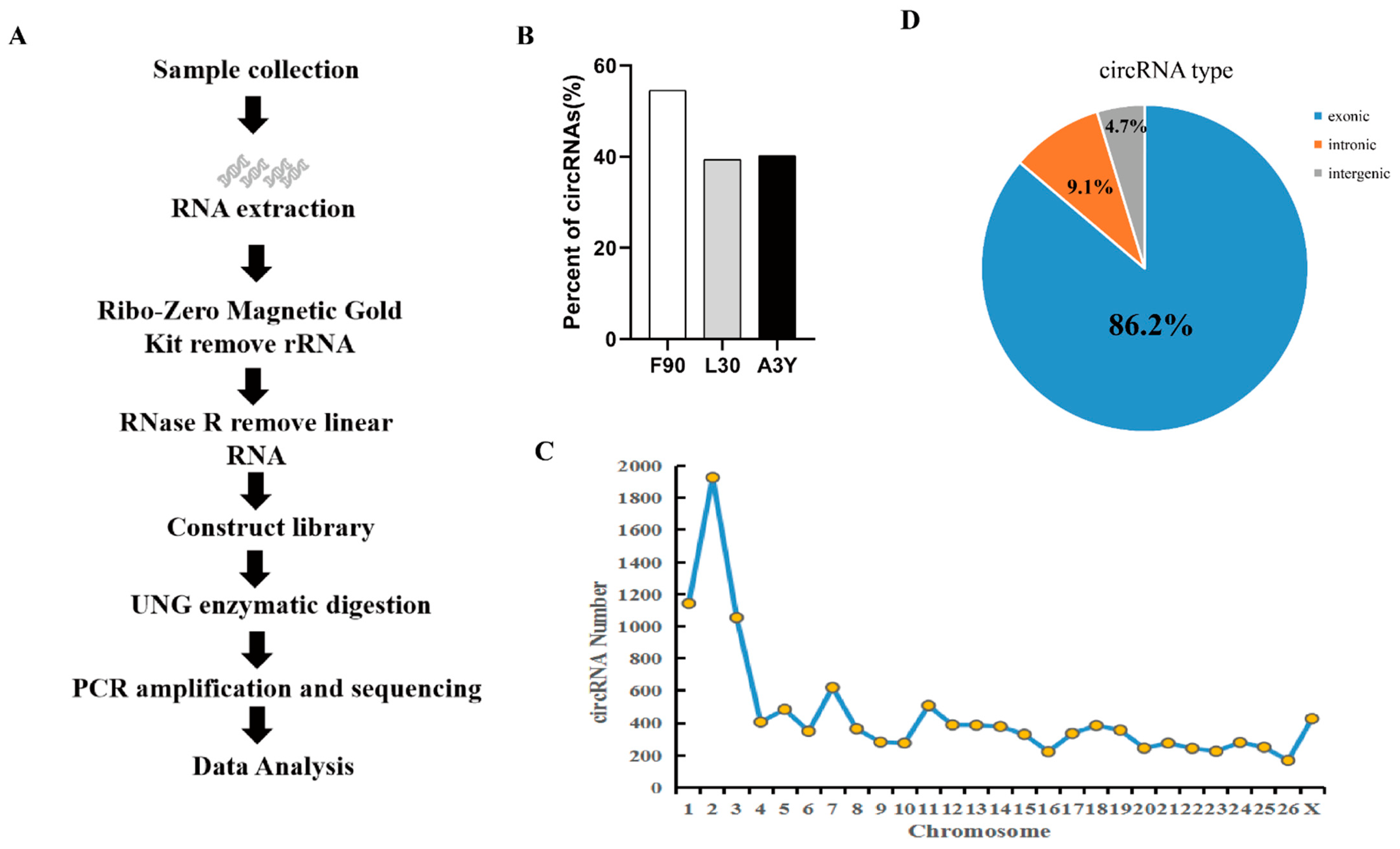
2.1. Identification of circRNAs during Sheep Skeletal Muscle Development
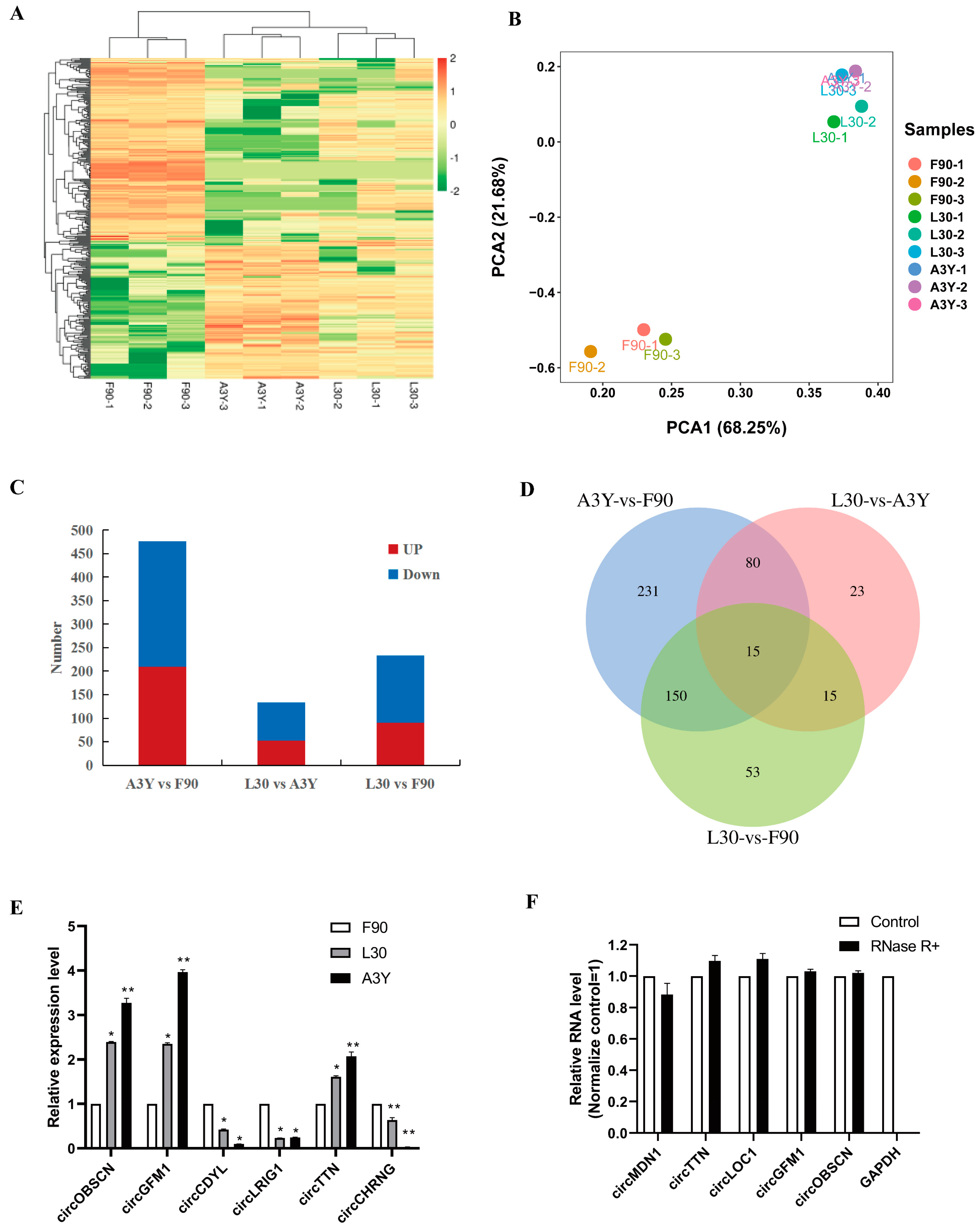
2.2. Identification of DEcircRNAs
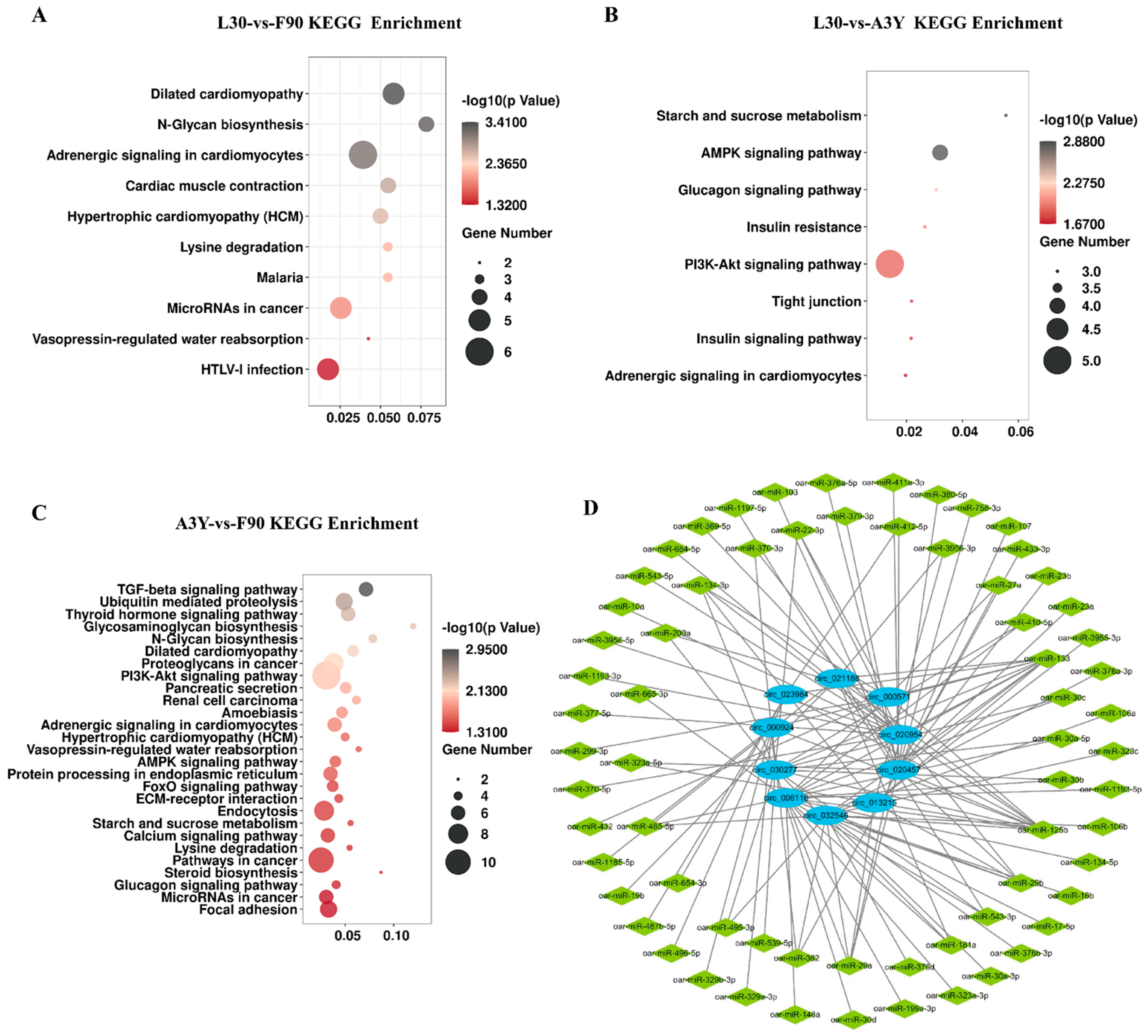
2.3. Functional Enrichment Analysis of DEcircRNAs
2.4. Trend Analysis of circRNAs during Muscle Development
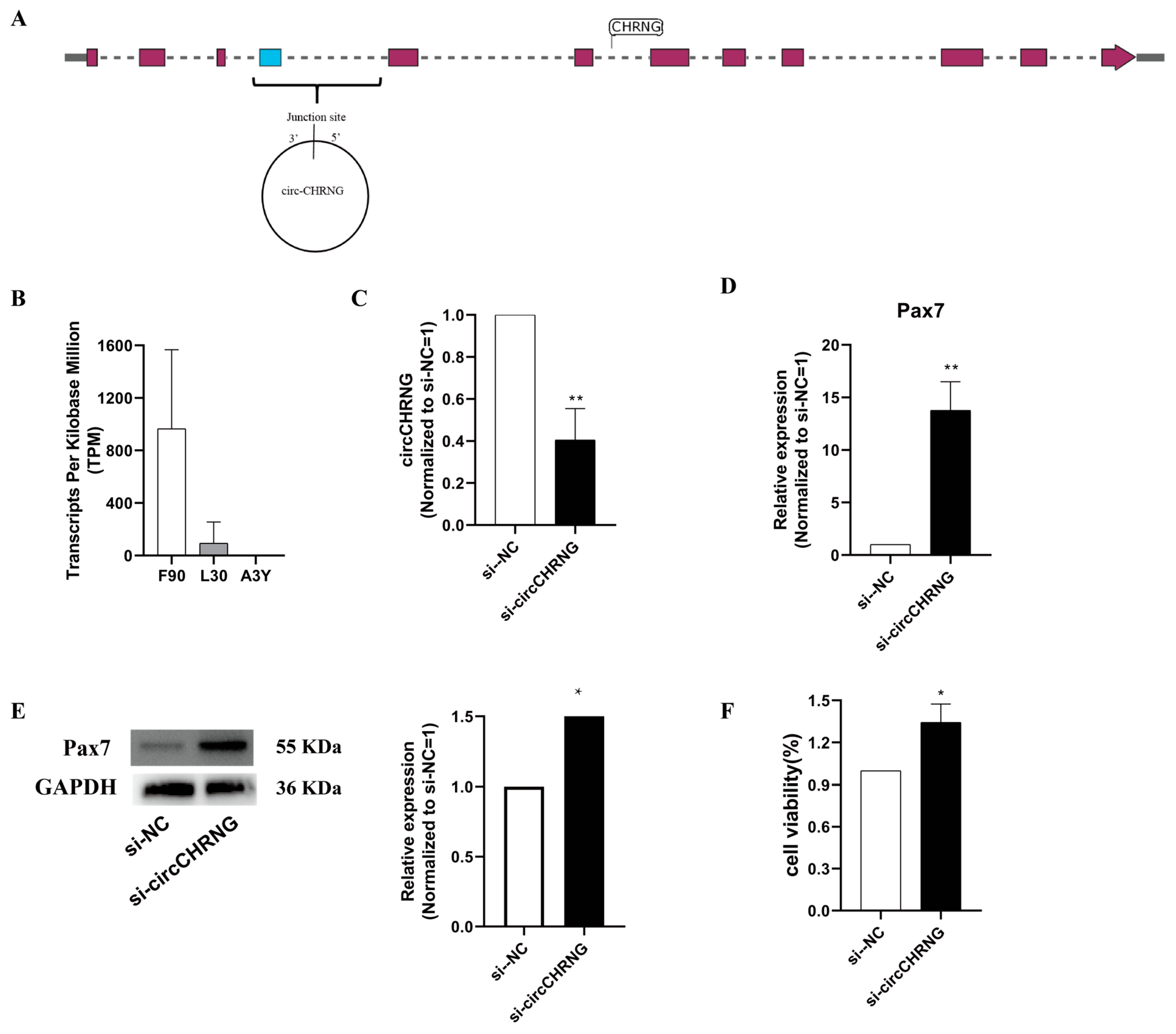
2.5. CircCHRNG Promote Myoblast Proliferation
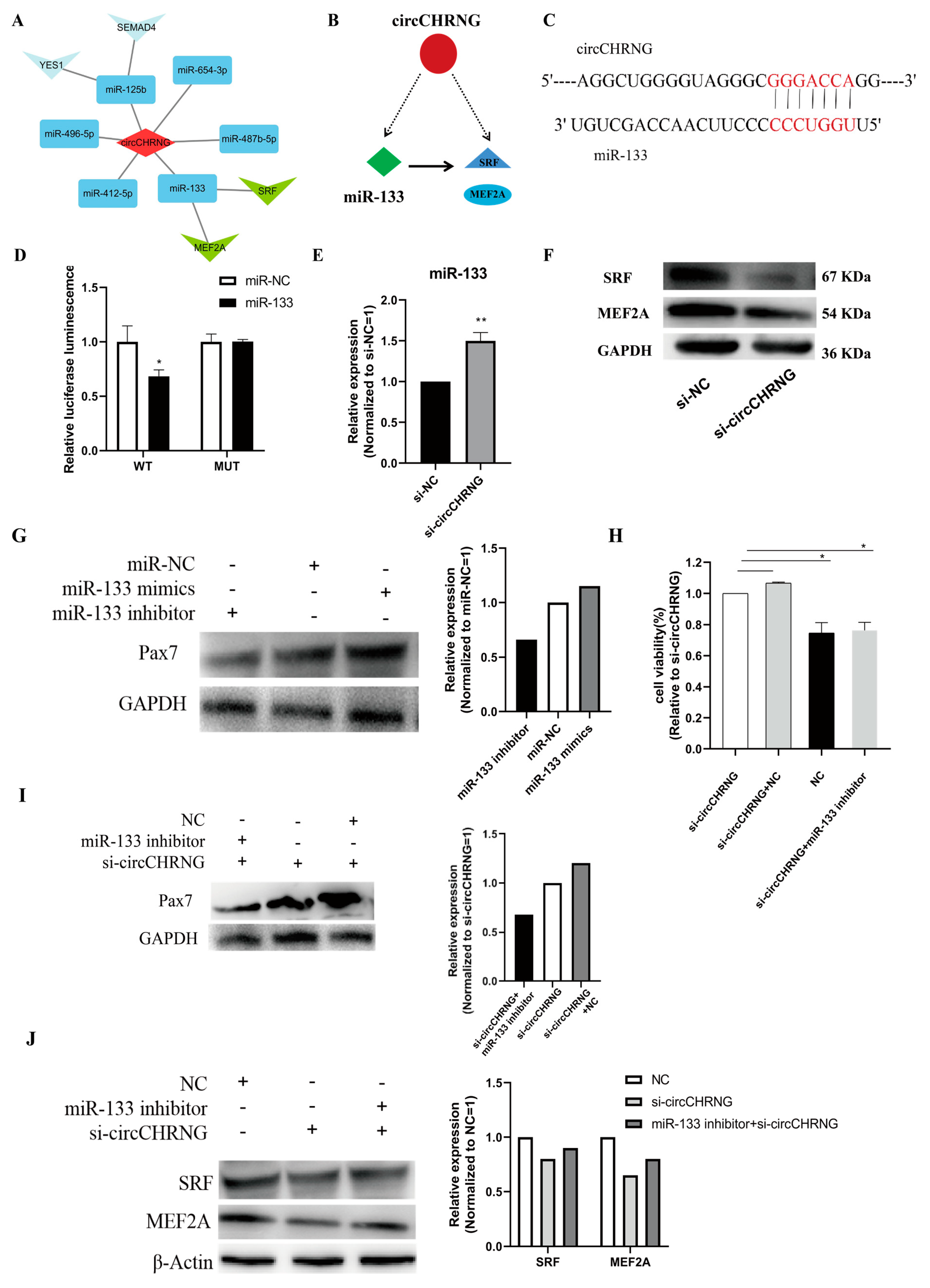
2.6. CircCHRNG interacts with miR-133 and Regulates the Expression of SRF and MEF2A in Myoblasts
2.7. CircCHRNG Functions as a Sponge of miR-133 to Promote Myoblast Proliferation
3. Discussion
4. Materials and Methods
4.1. Experimental Samples and Ethics Statement
4.2. Library Construction and Sequencing
4.3. Identification of Circular RNAs in Sheep Muscle
4.4. Differential Expression Analysis
4.5. Target Site Prediction and Functional Enrichment Analysis
4.6. Data Validation
4.7. Cell Culture and Transfection
4.8. Dual-Luciferase Reporter Assay
4.9. Cell Proliferation Assays
4.10. Western Blot Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Guttman, M.; Rinn, J.L. Modular regulatory principles of large non-coding RNAs. Nature 2012, 482, 339–346. [Google Scholar] [CrossRef]
- Sanger, H.L.; Klotz, G.; Riesner, D.; Gross, H.J.; Kleinschmidt, A.K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc. Natl. Acad. Sci. USA 1976, 73, 3852–3856. [Google Scholar] [CrossRef] [PubMed]
- Kai, D.; Yannian, L.; Yitian, C.; Dinghao, G.; Xin, Z.; Wu, J. Circular RNA HIPK3 promotes gallbladder cancer cell growth by sponging microRNA-124. Biochem. Biopyhs. Res. Commun. 2018, 503, 863–869. [Google Scholar] [CrossRef]
- Veno, M.T.; Hansen, T.B.; Venø, S.T.; Clausen, B.H.; Grebing, M.; Finsen, B.; Holm, I.E.; Kjems, J. Spatio-temporal regulation of circular RNA expression during porcine embryonic brain development. Genome Biol. 2015, 16, 245. [Google Scholar] [CrossRef]
- Chen, W.; Schuman, E. Circular RNAs in brain and other tissues: A functional enigma. Trends Neurosci. 2016, 39, 597–604. [Google Scholar] [CrossRef]
- Meng, S.; Zhou, H.; Feng, Z.; Xu, Z.; Tang, Y.; Li, P.; Wu, M. CircRNA: Functions and properties of a novel potential biomarker for cancer. Mol. Cancer 2017, 16, 94. [Google Scholar] [CrossRef]
- Chen, B.; Huang, S. Circular RNA: An emerging non-coding RNA as a regulator and biomarker in cancer. Cancer Lett. 2018, 418, 41–50. [Google Scholar] [CrossRef]
- Das, A.; Sinha, T.; Shyamal, S.; Panda, A. Emerging role of circular RNA–protein interactions. Noncoding RNA 2021, 7, 48. [Google Scholar] [CrossRef]
- Sinha, T.; Panigrahi, C.; Das, D.; Panda, A.C. Circular RNA translation, a path to hidden proteome. Wiley Interdiscip. Rev.-RNA 2022, 13, e1685. [Google Scholar] [CrossRef]
- Ke, S.A.; Zhao, S.; Liu, Y.; Zhuo, Q.; Tong, X.; Xu, Y. Circular RNA-encoded peptides and proteins: Implications to cancer. Sheng Wu Gong Xue Bao Chin. J. Biotechnol. 2022, 38, 3131–3140. [Google Scholar]
- Zhang, Y.; Zhang, X.-O.; Chen, T.; Xiang, J.-F.; Yin, Q.-F.; Xing, Y.-H.; Zhu, S.; Yang, L.; Chen, L.-L. Circular intronic long noncoding RNAs. Mol. Cell 2013, 51, 792–806. [Google Scholar] [CrossRef]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef]
- Nie, M.; Deng, Z.-L.; Liu, J.; Wang, D.-Z. Noncoding RNAs, emerging regulators of skeletal muscle development and diseases. BioMed Res. Int. 2015, 2015, 676575. [Google Scholar] [CrossRef]
- Wang, J.; Ren, Q.; Hua, L.; Chen, J.; Zhang, J.; Bai, H.; Li, H.; Xu, B.; Shi, Z.; Cao, H.; et al. Comprehensive analysis of differentially expressed mRNA, lncRNA and circRNA and their ceRNA networks in the longissimus dorsi muscle of two different pig breeds. Int. J. Mol. Sci. 2019, 20, 1107. [Google Scholar] [CrossRef]
- Liang, G.; Yang, Y.; Niu, G.; Tang, Z.; Li, K. Genome-wide profiling of Sus scrofa circular RNAs across nine organs and three developmental stages. DNA Res. 2017, 24, 523–535. [Google Scholar] [CrossRef]
- Ouyang, H.; Chen, X.; Wang, Z.; Yu, J.; Jia, X.; Li, Z.; Luo, W.; Abdalla, B.A.; Jebessa, E.; Nie, Q.; et al. Circular RNAs are abundant and dynamically expressed during embryonic muscle development in chickens. DNA Res. 2018, 25, 71–86. [Google Scholar] [CrossRef]
- Legnini, I.; Di Timoteo, G.; Rossi, F.; Morlando, M.; Briganti, F.; Sthandier, O.; Fatica, A.; Santini, T.; Andronache, A.; Wade, M.; et al. Circ-ZNF609 is a circular RNA that can be translated and functions in myogenesis. Mol. Cell 2017, 66, 22–37.e29. [Google Scholar] [CrossRef]
- Zhang, P.; Xu, H.; Li, R.; Wu, W.; Chao, Z.; Li, C.; Xia, W.; Wang, L.; Yang, J.; Xu, Y. Assessment of myoblast circular RNA dynamics and its correlation with miRNA during myogenic differentiation. Int. J. Biochem. Cell Biol. 2018, 99, 211–218. [Google Scholar] [CrossRef]
- Du, J.; Zhao, H.; Song, G.; Pang, Y.; Jiang, L.; Zan, L.; Wang, H. Overexpression of cholinergic receptor nicotinic gamma subunit inhibits proliferation and differentiation of bovine preadipocytes. Anim. Biosci. 2022, 00, 1–9. [Google Scholar] [CrossRef]
- Koenen, M.; Peter, C.; Villarroel, A.; Witzemann, V.; Sakmann, B. Acetylcholine receptor channel subtype directs the innervation pattern of skeletal muscle. EMBO Rep. 2005, 6, 570–576. [Google Scholar] [CrossRef][Green Version]
- Chen, J.-F.; Mandel, E.M.; Thomson, J.M.; Wu, Q.; Callis, T.E.; Hammond, S.M.; Conlon, F.L.; Wang, D.-Z. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat. Genet. 2006, 38, 228–233. [Google Scholar] [CrossRef]
- Horak, M.; Novak, J.; Bienertova-Vasku, J. Muscle-specific microRNAs in skeletal muscle development. Dev. Biol. 2016, 410, 1–13. [Google Scholar] [CrossRef]
- Liu, N.; Williams, A.H.; Kim, Y.; McAnally, J.; Bezprozvannaya, S.; Sutherland, L.B.; Richardson, J.A.; Bassel-Duby, R.; Olson, E.N. An intragenic MEF2-dependent enhancer directs muscle-specific expression of microRNAs 1 and 133. Proc. Natl. Acad. Sci. USA 2007, 104, 20844–20849. [Google Scholar] [CrossRef]
- Zhao, Y.; Samal, E.; Srivastava, D. Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis. Nature 2005, 436, 214–220. [Google Scholar] [CrossRef]
- Fathi, M.; Gharakhanlou, R.; Rezaei, R. The Changes of Heart miR-1 and miR-133 Expressions following Physiological Hypertrophy Due to Endurance Training. Cell J. 2020, 22, 133. [Google Scholar]
- Jin, L.; Tang, Q.; Hu, S.; Chen, Z.; Zhou, X.; Zeng, B.; Wang, Y.; He, M.; Li, Y.; Gui, L.; et al. A pig BodyMap transcriptome reveals diverse tissue physiologies and evolutionary dynamics of transcription. Nat. Commun. 2021, 12, 3715. [Google Scholar] [CrossRef]
- Huang, M.; Shen, Y.; Mao, H.; Chen, L.; Chen, J.; Guo, X.; Xu, N. Circular RNA expression profiles in the porcine liver of two distinct phenotype pig breeds. Asian Austalas. J. Anim. Sci. 2018, 31, 812. [Google Scholar] [CrossRef]
- Yang, Z.; He, T.; Chen, Q. The roles of circRNAs in regulating muscle development of livestock animals. Front. Cell Dev. Biol. 2021, 9, 619329. [Google Scholar] [CrossRef]
- Hong, L.; Gu, T.; He, Y.; Zhou, C.; Hu, Q.; Wang, X.; Zheng, E.; Huang, S.; Xu, Z.; Yang, J.; et al. Genome-wide analysis of circular RNAs mediated ceRNA regulation in porcine embryonic muscle development. Front. Cell Dev. Biol. 2019, 7, 289. [Google Scholar] [CrossRef]
- Wei, X.; Li, H.; Yang, J.; Hao, D.; Dong, D.; Huang, Y.; Lan, X.; Plath, M.; Lei, C.; Lin, F.; et al. Circular RNA profiling reveals an abundant circLMO7 that regulates myoblasts differentiation and survival by sponging miR-378a-3p. Cell Death Dis. 2017, 8, e3153. [Google Scholar] [CrossRef]
- Lai, K.-M.V.; Gonzalez, M.; Poueymirou, W.T.; Kline, W.O.; Na, E.; Zlotchenko, E.; Stitt, T.N.; Economides, A.; Yancopoulos, G.D.; Glass, D.J. Conditional activation of akt in adult skeletal muscle induces rapid hypertrophy. Mol. Cell. Biol. 2004, 24, 9295–9304. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, J.; Vernus, B.; Chelh, I.; Cassar-Malek, I.; Gabillard, J.C.; Sassi, A.H.; Seiliez, I.; Picard, B.; Bonnieu, A. Myostatin and the skeletal muscle atrophy and hypertrophy signaling pathways. Cell. Mol. Life Sci. 2014, 71, 4361–4371. [Google Scholar] [CrossRef]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA hypothesis: The Rosetta Stone of a hidden RNA language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef]
- Ni, H.; Li, W.; Zhuge, Y.; Xu, S.; Wang, Y.; Chen, Y.; Shen, G.; Wang, F. Inhibition of circHIPK3 prevents angiotensin II-induced cardiac fibrosis by sponging miR-29b-3p. Int. J. Cardiol. 2019, 292, 188–196. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.-X.; Lu, J.; Xie, H.; Wang, D.-P.; Ni, H.; Zhu, Y.; Ren, L.-H.; Meng, X.-X.; Wang, R.-L. circHIPK3 regulates lung fibroblast-to-myofibroblast transition by functioning as a competing endogenous RNA. Cell Death Dis. 2019, 10, 182. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Yang, F.; Cao, H.; Liang, Z. Malat1 regulates serum response factor through miR-133 as a competing endogenous RNA in myogenesis. FASEB J. 2015, 29, 3054–3064. [Google Scholar] [CrossRef] [PubMed]
- Velasco, E.L.; Galiano-Torres, J.; Jodar-Garcia, A.; Aranega, A.E.; Franco, D. miR-27 and miR-125 distinctly regulate muscle-enriched transcription factors in cardiac and skeletal myocytes. BioMed Res. Int. 2015, 2015, 391306. [Google Scholar]
- Wang, X.; Chen, S.; Gao, Y.; Yu, C.; Nie, Z.; Lu, R.; Sun, Y.; Guan, Z. MicroRNA-125b inhibits the proliferation of vascular smooth muscle cells induced by platelet-derived growth factor BB. Exp. Ther. Med. 2021, 22, 791. [Google Scholar] [CrossRef] [PubMed]
- Naguibneva, I.; Ameyar-Zazoua, M.; Polesskaya, A.; Ait-Si-Ali, S.; Groisman, R.; Souidi, M.; Cuvellier, S.; Harel-Bellan, A. The microRNA miR-181 targets the homeobox protein Hox-A11 during mammalian myoblast differentiation. Nat. Cell Biol. 2006, 8, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Wu, N.; Gu, T.; Lu, L.; Cao, Z.; Song, Q.; Wang, Z.; Zhang, Y.; Chang, G.; Xu, Q.; Chen, G. Roles of miRNA-1 and miRNA-133 in the Proliferation and Differentiation of Myoblasts in Duck Skeletal Muscle. J. Cell. Physiol. 2019, 234, 3490–3499. [Google Scholar] [CrossRef]
- Huang, M.; Xu, H.; Xie, S.-J.; Zhou, H.; Qu, L.-H. Insulin-like growth factor-1 receptor is regulated by microRNA-133 during skeletal myogenesis. PLoS ONE 2011, 6, e29173. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Yang, S.; Kang, Z.; Ru, W.; Shen, X.; Li, M.; Lan, X.; Chen, H. circMEF2D Negatively Regulated by HNRNPA1 Inhibits Proliferation and Differentiation of Myoblasts via miR-486-PI3K/AKT Axis. J. Agric. Food Chem. 2022, 70, 8145–8163. [Google Scholar] [CrossRef]
- Guo, Z.; Zhao, L.; Ji, S.; Long, T.; Huang, Y.; Ju, R.; Tang, W.; Tian, W.; Long, J. CircRNA-23525 regulates osteogenic differentiation of adipose-derived mesenchymal stem cells via miR-30a-3p. Cell Tissue Res. 2021, 383, 795–807. [Google Scholar] [CrossRef]
- Kariminejad, A.; Almadani, N.; Khoshaeen, A.; Olsson, B.; Moslemi, A.-R.; Tajsharghi, H. Truncating CHRNG mutations associated with interfamilial variability of the severity of the Escobar variant of multiple pterygium syndrome. BMC Genet. 2016, 17, 71. [Google Scholar] [CrossRef]
- Wang, Y.; Pei, W.; Lu, P. Circ_ARHGAP32 acts as miR-665 sponge to upregulate FGF2 to promote ox-LDL induced vascular smooth muscle cells proliferation and migration. Clin. Hemorheol. Microcirc. 2022, 82, 169–182. [Google Scholar] [CrossRef]
- Feng, Y.; Niu, L.; Wei, W.; Zhang, W.; Li, X.; Cao, J.; Zhao, S. A feedback circuit between miR-133 and the ERK1/2 pathway involving an exquisite mechanism for regulating myoblast proliferation and differentiation. Cell Death Dis. 2013, 4, e934. [Google Scholar] [CrossRef]
- Wang, Y.-N.; Yang, W.-C.; Li, P.-W.; Wang, H.-B.; Zhang, Y.-Y.; Zan, L.-S. Myocyte enhancer factor 2A promotes proliferation and its inhibition attenuates myogenic differentiation via myozenin 2 in bovine skeletal muscle myoblast. PLoS ONE 2018, 13, e0196255. [Google Scholar] [CrossRef]
- Snyder, C.M.; Rice, A.L.; Estrella, N.L.; Held, A.; Kandarian, S.C.; Naya, F.J. MEF2A regulates the Gtl2-Dio3 microRNA mega-cluster to modulate WNT signaling in skeletal muscle regeneration. Development 2013, 140, 31–42. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Chen, Q.; Bao, J.; Pu, Y.; Han, J.; Zhao, H.; Ma, Y.; Zhao, Q. Genome-Wide Analysis of Circular RNAs Reveals circCHRNG Regulates Sheep Myoblast Proliferation via miR-133/SRF and MEF2A Axis. Int. J. Mol. Sci. 2022, 23, 16065. https://doi.org/10.3390/ijms232416065
Liu Y, Chen Q, Bao J, Pu Y, Han J, Zhao H, Ma Y, Zhao Q. Genome-Wide Analysis of Circular RNAs Reveals circCHRNG Regulates Sheep Myoblast Proliferation via miR-133/SRF and MEF2A Axis. International Journal of Molecular Sciences. 2022; 23(24):16065. https://doi.org/10.3390/ijms232416065
Chicago/Turabian StyleLiu, Yue, Qian Chen, Jingjing Bao, Yabin Pu, Jianlin Han, Huijing Zhao, Yuehui Ma, and Qianjun Zhao. 2022. "Genome-Wide Analysis of Circular RNAs Reveals circCHRNG Regulates Sheep Myoblast Proliferation via miR-133/SRF and MEF2A Axis" International Journal of Molecular Sciences 23, no. 24: 16065. https://doi.org/10.3390/ijms232416065
APA StyleLiu, Y., Chen, Q., Bao, J., Pu, Y., Han, J., Zhao, H., Ma, Y., & Zhao, Q. (2022). Genome-Wide Analysis of Circular RNAs Reveals circCHRNG Regulates Sheep Myoblast Proliferation via miR-133/SRF and MEF2A Axis. International Journal of Molecular Sciences, 23(24), 16065. https://doi.org/10.3390/ijms232416065