Identification of Iguania Ancestral Syntenic Blocks and Putative Sex Chromosomes in the Veiled Chameleon (Chamaeleo calyptratus, Chamaeleonidae, Iguania)
Abstract
1. Introduction
2. Results
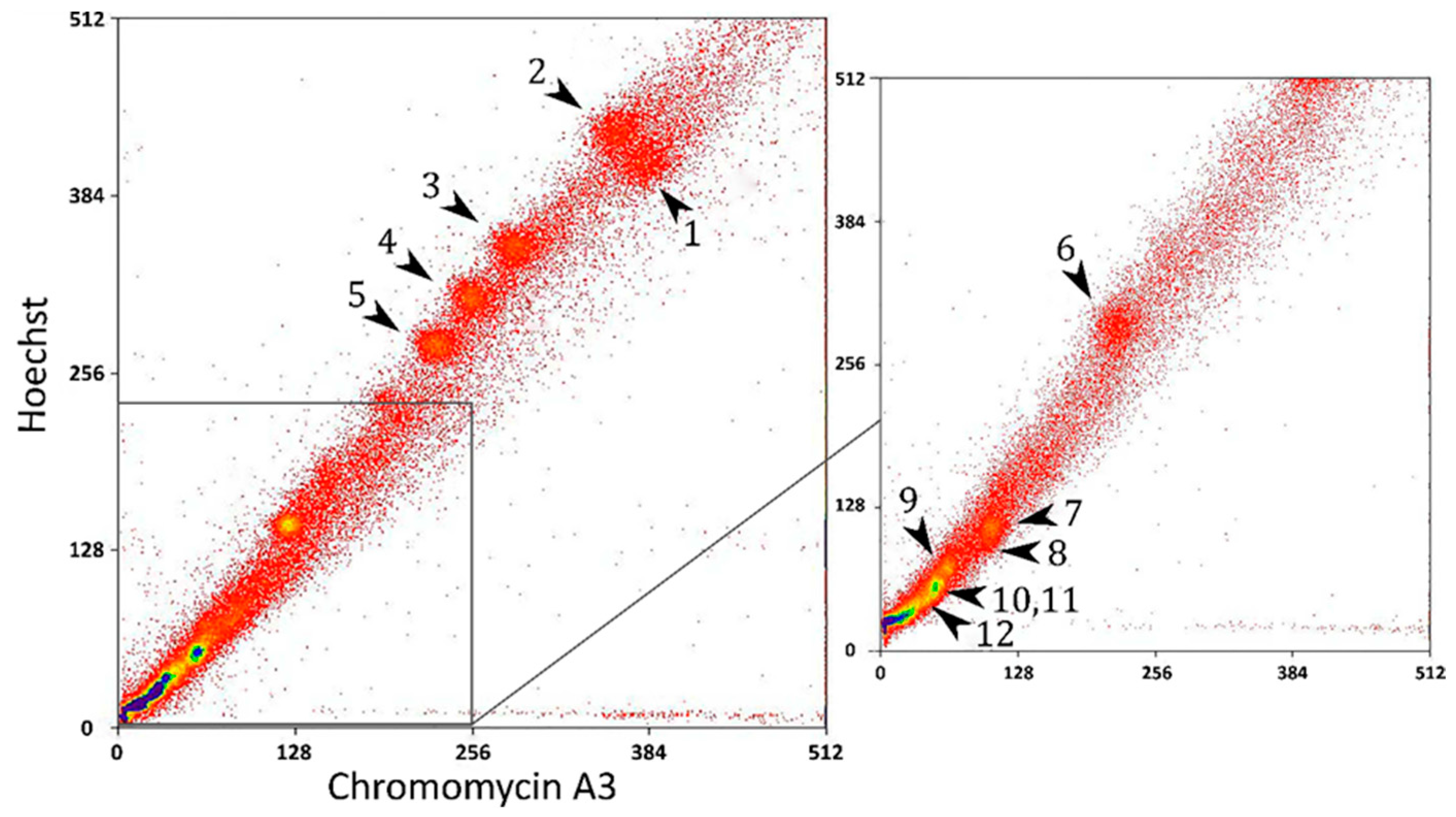
2.1. FISH with Flow-Sorted Chromosome-Specific Probes of Chamaeleo calyptratus
2.2. Sequencing of the Flow-Sorted Chromosome-Specific Probes
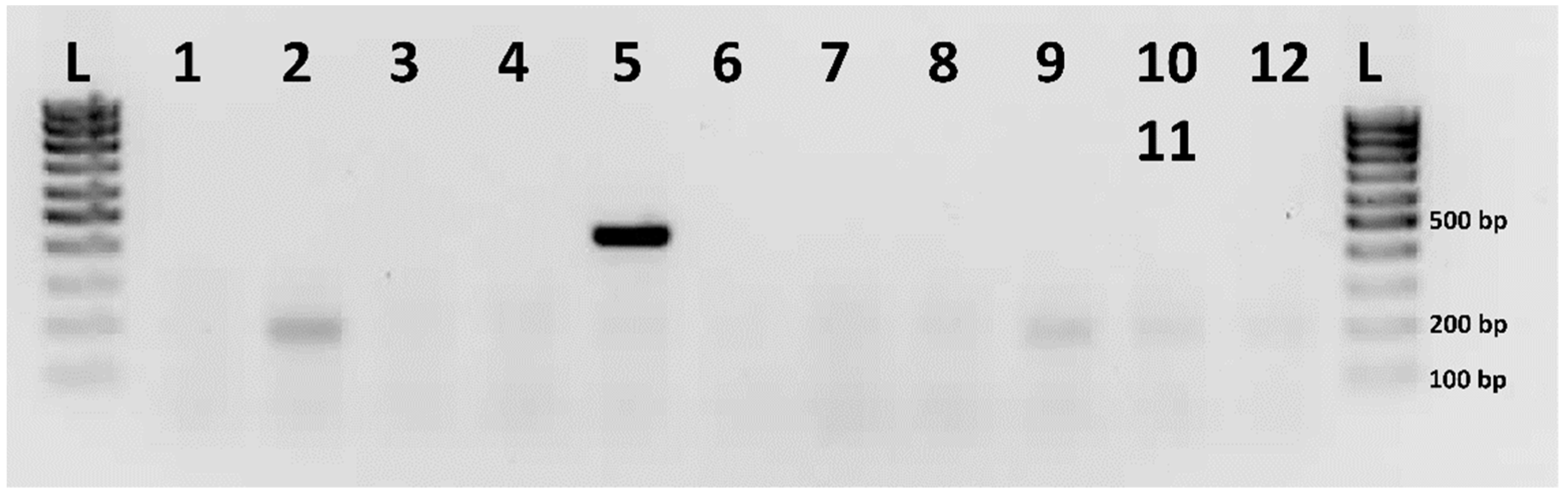
2.3. Identification of the Putative Sex Chromosome Pair
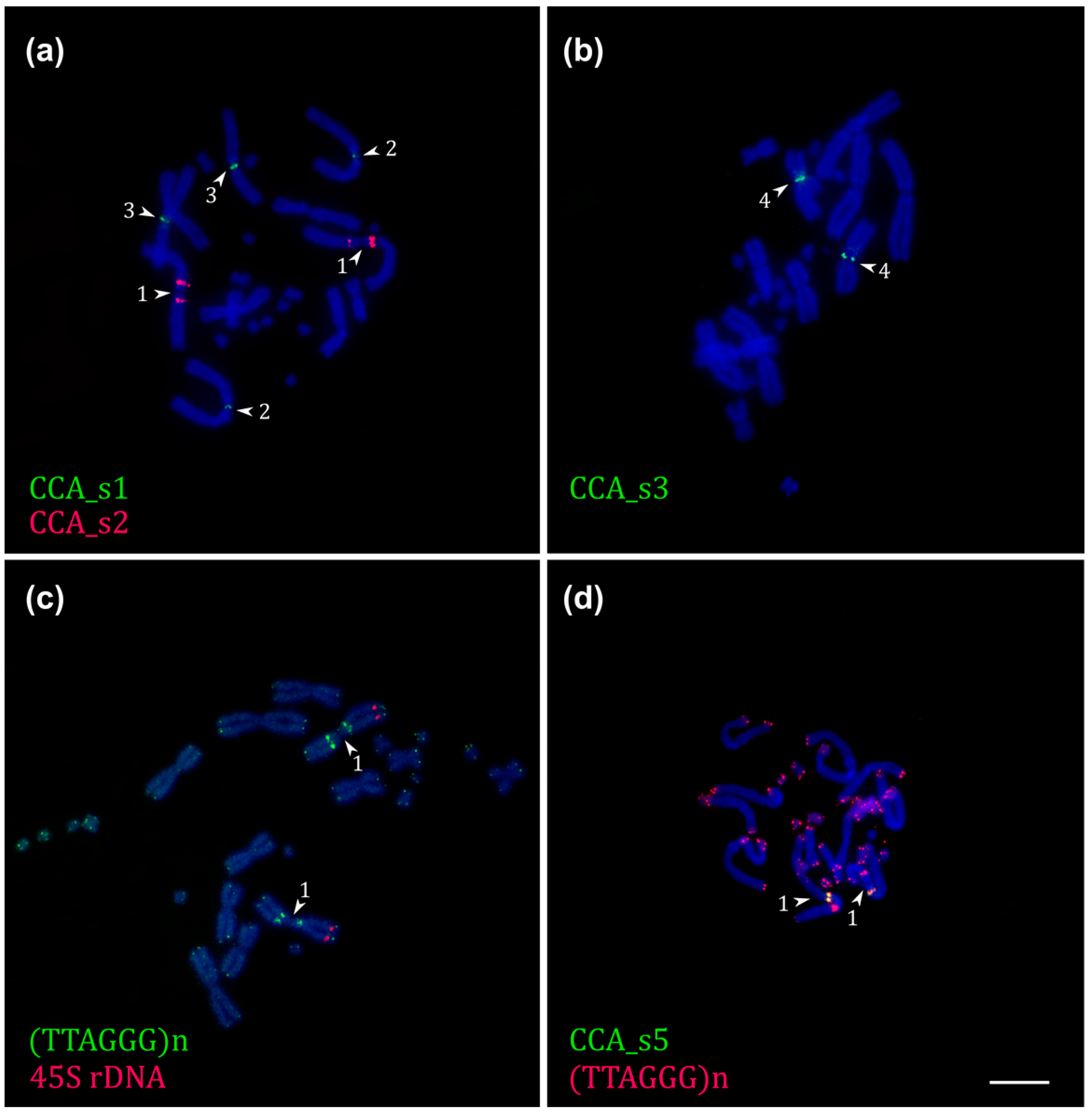
2.4. Repetitive DNA Bioinformatic Analysis and FISH Localization
2.5. FISH with Telomeric and rDNA Probes of Chamaeleo calyptratus
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. Cell Culture and Chromosome Suspension
4.3. Chromosome Sorting and Probe Preparation
4.4. DNA Extraction
4.5. Telomeric and rDNA Probes
4.6. Probes of Sex-Specific RAD-Seq Markers
4.7. Male-Specific RAD-Seq Markers Mapping
4.8. DNA Sequencing
4.9. Bioinformatic Analysis
4.10. Probes for Repetitive DNA Monomers
4.11. Fluorescent in Situ Hybridization
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tonini, J.F.R.; Beard, K.H.; Ferreira, R.B.; Jetz, W.; Pyron, R.A. Fully-Sampled Phylogenies of Squamates Reveal Evolutionary Patterns in Threat Status. Biol. Conserv. 2016, 204, 23–31. [Google Scholar] [CrossRef]
- Hedges, S.B.; Marin, J.; Suleski, M.; Kumar, S. Tree of Life Reveals Clock-Like Speciation and Diversification. Mol. Biol. Evol. 2015, 32, 835–845. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Wiens, J.J. Combining Phylogenomic and Supermatrix Approaches, and a Time-Calibrated Phylogeny for Squamate Reptiles (Lizards and Snakes) Based on 52 Genes and 4162 Species. Mol. Phylogenet. Evol. 2016, 94, 537–547. [Google Scholar] [CrossRef] [PubMed]
- Vitt, L.J.; Caldwell, J.P. Herpetology: An Introductory Biology of Amphibians and Reptiles, 4th ed.; Gomez, K., Gonzalez, P., Eds.; Academic Press: Cambridge, MA, USA, 2014. [Google Scholar]
- Uetz, P.; Hosek, J. The Reptile Database. Available online: http://www.reptile-database.org/ (accessed on 5 November 2022).
- Bickel, R.; Losos, J.B. Patterns of Morphological Variation and Correlates of Habitat Use in Chameleons. Biol. J. Linn. Soc. 2002, 76, 91–103. [Google Scholar] [CrossRef]
- Stuart-Fox, D.; Moussalli, A. Selection for Social Signalling Drives the Evolution of Chameleon Colour Change. PLoS Biol. 2008, 6, e25. [Google Scholar] [CrossRef] [PubMed]
- Rovatsos, M.; Altmanová, M.; Pokorná, M.J.; Velenský, P.; Baca, A.S.; Kratochvíl, L. Evolution of Karyotypes in Chameleons. Genes 2017, 8, 382. [Google Scholar] [CrossRef]
- Olmo, E.; Signorino, G. Chromorep: A Reptile Chromosomes Dabase; ScienceOpen: Burlington, MA, USA, 2005. [Google Scholar]
- Rovatsos, M.; Pokorna, M.J.; Altmanova, M.; Kratochvil, L. Female Heterogamety in Madagascar Chameleons (Squamata: Chamaeleonidae: Furcifer): Differentiation of Sex and Neo-Sex Chromosomes. Sci. Rep. 2015, 5, 13196. [Google Scholar] [CrossRef]
- Rovatsos, M.; Altmanová, M.; Augstenová, B.; Mazzoleni, S.; Velenský, P.; Kratochvíl, L. ZZ/ZW Sex Determination with Multiple Neo-Sex Chromosomes Is Common in Madagascan Chameleons of the Genus Furcifer (Reptilia: Chamaeleonidae). Genes 2019, 10, 1020. [Google Scholar] [CrossRef]
- Andrews, R.M. Incubation Temperature and Sex Ratio of the Veiled Chameleon (Chamaeleo calyptratus). J. Herptol. 2005, 39, 515–518. [Google Scholar] [CrossRef]
- Nielsen, S.V.; Banks, J.L.; Diaz, R.E.; Trainor, P.A.; Gamble, T. Dynamic Sex Chromosomes in Old World Chameleons (Squamata: Chamaeleonidae). J. Evol. Biol. 2018, 31, 484–490. [Google Scholar] [CrossRef]
- Sidhom, M.; Said, K.; Chatti, N.; Guarino, F.M.; Odierna, G.; Petraccioli, A.; Picariello, O.; Mezzasalma, M. Karyological Characterization of the Common Chameleon (Chamaeleo chamaeleon) Provides Insights on the Evolution and Diversification of Sex Chromosomes in Chamaeleonidae. Zoology 2020, 141, 125738. [Google Scholar] [CrossRef] [PubMed]
- Diaz, R.E.; Anderson, C.V.; Baumann, D.P.; Kupronis, R.; Jewell, D.; Piraquive, C.; Kupronis, J.; Winter, K.; Bertocchini, F.; Trainor, P.A. The Veiled Chameleon (Chamaeleo calyptratus Duméril and Duméril 1851): A Model for Studying Reptile Body Plan Development and Evolution. Cold Spring Harb. Protoc. 2015, 2015, pdb-emo087700. [Google Scholar] [CrossRef]
- Ballen, C.; Shine, R.; Olsson, M. Effects of Early Social Isolation on the Behaviour and Performance of Juvenile Lizards, Chamaeleo Calyptratus. Anim. Behav. 2014, 88, 1–6. [Google Scholar] [CrossRef][Green Version]
- Pokorná, M.; Giovannotti, M.; Kratochvíl, L.; Kasai, F.; Trifonov, V.A.; O’Brien, P.C.M.; Caputo, V.; Olmo, E.; Ferguson-Smith, M.A.; Rens, W. Strong Conservation of the Bird Z Chromosome in Reptilian Genomes Is Revealed by Comparative Painting despite 275 Million Years Divergence. Chromosoma 2011, 120, 455–468. [Google Scholar] [CrossRef] [PubMed]
- Pinto, B.J.; Card, D.C.; Castoe, T.A.; Diaz, R.E.; Nielsen, S.V.; Trainor, P.A.; Gamble, T. The Transcriptome of the Veiled Chameleon (Chamaeleo calyptratus): A Resource for Studying the Evolution and Development of Vertebrates. Dev. Dyn. 2019, 248, 702–708. [Google Scholar] [CrossRef]
- Makunin, A.I.; Kichigin, I.G.; Larkin, D.M.; O’brien, P.C.M.; Ferguson-Smith, M.A.; Yang, F.; Proskuryakova, A.A.; Vorobieva, N.V.; Chernyaeva, E.N.; O’brien, S.J.; et al. Contrasting Origin of B Chromosomes in Two Cervids (Siberian Roe Deer and Grey Brocket Deer) Unravelled by Chromosome-Specific DNA Sequencing. BMC Genom. 2016, 17, 618. [Google Scholar] [CrossRef]
- Lisachov, A.P.; Tishakova, K.V.; Romanenko, S.A.; Molodtseva, A.S.; Prokopov, D.Y.; Pereira, J.C.; Ferguson-Smith, M.A.; Borodin, P.M.; Trifonov, V.A. Whole-Chromosome Fusions in the Karyotype Evolution of Sceloporus (Iguania, Reptilia) Are More Frequent in Sex Chromosomes than Autosomes. Philos. Trans. R. Soc. B Biol. Sci. 2021, 376, 20200099. [Google Scholar] [CrossRef]
- Giovannotti, M.; Trifonov, V.A.; Paoletti, A.; Kichigin, I.G.; O’brien, P.C.M.; Kasai, F.; Giovagnoli, G.; Ng, B.L.; Ruggeri, P.; Cerioni, P.N.; et al. New Insights into Sex Chromosome Evolution in Anole Lizards (Reptilia, Dactyloidae). Chromosoma 2017, 126, 245–260. [Google Scholar] [CrossRef]
- Lisachov, A.P.; Makunin, A.I.; Giovannotti, M.; Pereira, J.C.; Druzhkova, A.S.; Caputo Barucchi, V.; Ferguson-Smith, M.A.; Trifonov, V.A. Genetic Content of the Neo-Sex Chromosomes in Ctenonotus and Norops (Squamata, Dactyloidae) and Degeneration of the Y Chromosome as Revealed by High-Throughput Sequencing of Individual Chromosomes. Cytogenet. Genome Res. 2019, 157, 115–122. [Google Scholar] [CrossRef]
- Kichigin, I.G.; Giovannotti, M.; Makunin, A.I.; Ng, B.L.; Kabilov, M.R.; Tupikin, A.E.; Caputo Barucchi, V.; Splendiani, A.; Ruggeri, P.; Rens, W.; et al. Evolutionary Dynamics of Anolis Sex Chromosomes Revealed by Sequencing of Flow Sorting-Derived Microchromosome-Specific DNA. Mol. Genet. Genom. 2016, 291, 1955–1966. [Google Scholar] [CrossRef]
- McQueen, H.A.; Siriaco, G.; Bird, A.P. Chicken Microchromosomes Are Hyperacetylated, Early Replicating, and Gene Rich. Genome Res. 1998, 8, 621–630. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.; Bruley, C.K.; Paton, I.R.; Dunn, I.; Jones, C.T.; Windsor, D.; Morrice, D.R.; Law, A.S.; Masabanda, J.; Sazanov, A.; et al. Differences in Gene Density on Chicken Macrochromosomes and Microchromosomes. Anim. Genet. 2000, 31, 96–103. [Google Scholar] [CrossRef] [PubMed]
- Burt, D.W. Origin and Evolution of Avian Microchromosomes. Cytogenet. Genome Res. 2002, 96, 97–112. [Google Scholar] [CrossRef] [PubMed]
- International Chicken Genome Sequencing Consortium. Sequence and Comparative Analysis of the Chicken Genome Provide Unique Perspectives on Vertebrate Evolution. Nature 2004, 432, 695–716. [Google Scholar] [CrossRef] [PubMed]
- Perry, B.W.; Schield, D.R.; Adams, R.H.; Castoe, T.A. Microchromosomes Exhibit Distinct Features of Vertebrate Chromosome Structure and Function with Underappreciated Ramifications for Genome Evolution. Mol. Biol. Evol. 2020, 38, 904–910. [Google Scholar] [CrossRef] [PubMed]
- Waters, P.D.; Patel, H.R.; Ruiz-Herrera, A.; Álvarez-González, L.; Lister, N.C.; Simakov, O.; Ezaz, T.; Kaur, P.; Frere, C.; Grützner, F.; et al. Microchromosomes Are Building Blocks of Bird, Reptile, and Mammal Chromosomes. Proc. Natl. Acad. Sci. USA 2021, 118, e2112494118. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, E.H.C.; Habermann, F.A.; Lacerda, O.; Sbalqueiro, I.J.; Wienberg, J.; Müller, S. Chromosome Reshuffling in Birds of Prey: The Karyotype of the World’s Largest Eagle (Harpy Eagle, Harpia harpyja) Compared to That of the Chicken (Gallus gallus). Chromosoma 2005, 114, 338–343. [Google Scholar] [CrossRef]
- Koochekian, N.; Ascanio, A.; Farleigh, K.; Card, D.C.; Schield, D.R.; Castoe, T.A.; Jezkova, T. A Chromosome-Level Genome Assembly and Annotation of the Desert Horned Lizard, Phrynosoma Platyrhinos, Provides Insight into Chromosomal Rearrangements among Reptiles. Gigascience 2022, 11, giab098. [Google Scholar] [CrossRef]
- Ruiz-Herrera, A.; Nergadze, S.G.; Santagostino, M.; Giulotto, E. Telomeric Repeats Far from the Ends: Mechanisms of Origin and Role in Evolution. Cytogenet. Genome Res. 2008, 122, 219–228. [Google Scholar] [CrossRef]
- Srikulnath, K.; Azad, B.; Singchat, W.; Ezaz, T. Distribution and Amplification of Interstitial Telomeric Sequences (ITSs) in Australian Dragon Lizards Support Frequent Chromosome Fusions in Iguania. PLoS ONE 2019, 14, e0212683. [Google Scholar] [CrossRef]
- Altmanová, M.; Rovatsos, M.; Johnson Pokorná, M.; Veselý, M.; Wagner, F.; Kratochvíl, L. All Iguana Families with the Exception of Basilisks Share Sex Chromosomes. Zoology 2018, 126, 98–102. [Google Scholar] [CrossRef] [PubMed]
- Rovatsos, M.; Pokorná, M.; Altmanová, M.; Kratochvíl, L. Cretaceous Park of Sex Determination: Sex Chromosomes Are Conserved across Iguanas. Biol. Lett. 2014, 10, 20131093. [Google Scholar] [CrossRef] [PubMed]
- Rovatsos, M.; Praschag, P.; Fritz, U.; Kratochvíl, L. Stable Cretaceous Sex Chromosomes Enable Molecular Sexing in Softshell Turtles (Testudines: Trionychidae). Sci. Rep. 2017, 7, 42150. [Google Scholar] [CrossRef] [PubMed]
- Kawagoshi, T.; Uno, Y.; Matsubara, K.; Matsuda, Y.; Nishida, C. The ZW Micro-Sex Chromosomes of the Chinese Soft-Shelled Turtle (Pelodiscus Sinensis, Trionychidae, Testudines) Have the Same Origin as Chicken Chromosome 15. Cytogenet. Genome Res. 2009, 125, 125–131. [Google Scholar] [CrossRef] [PubMed]
- Rovatsos, M.; Farkačová, K.; Altmanová, M.; Johnson Pokorná, M.; Kratochvíl, L. The Rise and Fall of Differentiated Sex Chromosomes in Geckos. Mol. Ecol. 2019, 28, 3042–3052. [Google Scholar] [CrossRef]
- Rovatsos, M.; Gamble, T.; Nielsen, S.; Georges, A.; Ezaz, T.; Kratochvíl, L. Do Male and Female Heterogamety Really Differ in Expression Regulation? Lack of Global Dosage Balance in Pygopodid Geckos. Philos. Trans. R. Soc. B 2021, 376, 20200102. [Google Scholar] [CrossRef]
- Zhu, Z.; Matsubara, K.; Shams, F.; Dobry, J.; Wapstra, E.; Gamble, T.; Sarre, S.D.; Georges, A.; Graves, J.A.M.; Zhou, Q.; et al. Diversity of Reptile Sex Chromosome Evolution Revealed by Cytogenetic and Linked-Read Sequencing. Zool. Res. 2022, 43, 719–733. [Google Scholar] [CrossRef]
- Kostmann, A.; Kratochvíl, L.; Rovatsos, M. Poorly Differentiated XX/XY Sex Chromosomes Are Widely Shared across Skink Radiation. Proc. R. Soc. B Biol. Sci. 2021, 288, 20202139. [Google Scholar] [CrossRef]
- Pensabene, E.; Kratochvíl, L.; Rovatsos, M. Independent Evolution of Sex Chromosomes in Eublepharid Geckos, a Lineage with Environmental and Genotypic Sex Determination. Life 2020, 10, 342. [Google Scholar] [CrossRef]
- Nandi, S.S.; Ghosh, P.; Roy, S.S. Expression of PITX2 Homeodomain Transcription Factor during Rat Gonadal Development in a Sexually Dimorphic Manner. Cell. Physiol. Biochem. 2011, 27, 159–170. [Google Scholar] [CrossRef]
- Guioli, S.; Lovell-Badge, R. PITX2 Controls Asymmetric Gonadal Development in Both Sexes of the Chick and Can Rescue the Degeneration of the Right Ovary. Development 2007, 134, 4199–4208. [Google Scholar] [CrossRef] [PubMed]
- Schartl, M.; Schories, S.; Wakamatsu, Y.; Nagao, Y.; Hashimoto, H.; Bertin, C.; Mourot, B.; Schmidt, C.; Wilhelm, D.; Centanin, L.; et al. Sox5 Is Involved in Germ-Cell Regulation and Sex Determination in Medaka Following Co-Option of Nested Transposable Elements. BMC Biol. 2018, 16, 16. [Google Scholar] [CrossRef] [PubMed]
- Polanco, J.C.; Wilhelm, D.; Davidson, T.-L.; Knight, D.; Koopman, P. Sox10 Gain-of-Function Causes XX Sex Reversal in Mice: Implications for Human 22q-Linked Disorders of Sex Development. Hum. Mol. Genet. 2010, 19, 506–516. [Google Scholar] [CrossRef] [PubMed]
- Deakin, J.E.; Edwards, M.J.; Patel, H.; O’meally, D.; Lian, J.; Stenhouse, R.; Ryan, S.; Livernois, A.M.; Azad, B.; Holleley, C.E.; et al. Anchoring Genome Sequence to Chromosomes of the Central Bearded Dragon (Pogona Vitticeps) Enables Reconstruction of Ancestral Squamate Macrochromosomes and Identifies Sequence Content of the Z Chromosome. BMC Genom. 2016, 17, 447. [Google Scholar] [CrossRef] [PubMed]
- Kratochvíl, L.; Gamble, T.; Rovatsos, M. Sex Chromosome Evolution among Amniotes: Is the Origin of Sex Chromosomes Non-Random? Philos. Trans. R. Soc. B 2021, 376, 20200108. [Google Scholar] [CrossRef]
- Maomao, Z.; Yuezhao, W.; Zhijun, L.; Zili, F.; Guanfu, W.; Papenfuss, T.J.; Mace, R.J. Karyotypes of Nine Species in the Genus Phrynocephalus, with Discussion of Karyotypic Evolution of Chinese Phrynocephalus. Acta Zool. Sin. 1997, 43, 399–410. [Google Scholar]
- Stanyon, R.; Galleni, L. A Rapid Fibroblast Culture Technique for High Resolution Karyotypes. Ital. J. Zool. 1991, 58, 81–83. [Google Scholar] [CrossRef]
- Romanenko, S.A.; Biltueva, L.S.; Serdyukova, N.A.; Kulemzina, A.I.; Beklemisheva, V.R.; Gladkikh, O.L.; Lemskaya, N.A.; Interesova, E.A.; Korentovich, M.A.; Vorobieva, N.V.; et al. Segmental Paleotetraploidy Revealed in Sterlet (Acipenser ruthenus) Genome by Chromosome Painting. Mol. Cytogenet. 2015, 8, 90. [Google Scholar] [CrossRef]
- Yang, F.; Carter, N.P.; Shiu, L.; Ferguson-Smith, M.A. A Comparative Study of Karyotypes of Muntjacs by Chromosome Painting. Chromosoma 1995, 103, 642–652. [Google Scholar] [CrossRef]
- Telenius, H.; Pelmear, A.H.; Tunnacliffe, A.; Carter, N.P.; Behmel, A.; Ferguson-Smith, M.A.; Nordenrkjold, M.; Pfragner, R.; Ponder, B.A.J. Cytogenetic Analysis by Chromosome Painting Using DOP-PCR Amplified Flow-Sorted Chromosomes. Genes Chromosom. Cancer 1992, 4, 251–263. [Google Scholar] [CrossRef]
- Maden, B.E.H.; Dent, C.L.; Farrell, T.E.; Garde, J.; Mccallumt, F.S.; Wakeman, J.A. Clones of Human Ribosomal DNA Containing the Complete 18 S-RRNA and 28 S-RRNA Genes Characterization, a Detailed Map of the Human Ribosomal Transcription Unit and Diversity among Clones. Biochem. J. 1987, 246, 519–527. [Google Scholar] [CrossRef] [PubMed]
- Ijdo, J.W.; Wells, R.A.; Baldini, A.; Reeders, S.T. Improved Telomere Detection Using a Telomere Repeat Probe (TTAGGG)n Generated by PCR. Nucleic Acids Res. 1991, 19, 4780. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.-J.; Yu, W.-B.; Yang, J.-B.; Song, Y.; DePamphilis, C.W.; Yi, T.-S.; Li, D.-Z. GetOrganelle: A Fast and Versatile Toolkit for Accurate de Novo Assembly of Organelle Genomes Germplasm Bank of Wild Species. Genome Biol. 2020, 21, 241. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: A Multiple Sequence Alignment Method with Reduced Time and Space Complexity. BMC Bioinform. 2004, 5, 113. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- Novák, P.; Neumann, P.; Pech, J.; Steinhaisl, J.; MacAs, J. RepeatExplorer: A Galaxy-Based Web Server for Genome-Wide Characterization of Eukaryotic Repetitive Elements from next-Generation Sequence Reads. Bioinformatics 2013, 29, 792–793. [Google Scholar] [CrossRef]
- Novák, P.; Robledillo, L.Á.; Koblížková, A.; Vrbová, I.; Neumann, P.; Macas, J. TAREAN: A Computational Tool for Identification and Characterization of Satellite DNA from Unassembled Short Reads. Nucleic Acids Res. 2017, 45, e111. [Google Scholar] [CrossRef]
- Bishani, A.; Prokopov, D.Y.; Romanenko, S.A.; Molodtseva, A.S.; Perelman, P.L.; Interesova, E.A.; Beklemisheva, V.R.; Graphodatsky, A.S.; Trifonov, V.A. Evolution of Tandemly Arranged Repetitive DNAs in Three Species of Cyprinoidei with Different Ploidy Levels. Cytogenet. Genome Res. 2021, 161, 32–42. [Google Scholar] [CrossRef]
- Johnson, M.; Zaretskaya, I.; Raytselis, Y.; Merezhuk, Y.; McGinnis, S.; Madden, T.L. NCBI BLAST: A Better Web Interface. Nucleic Acids Res. 2008, 36, W5–W9. [Google Scholar] [CrossRef]
- Liehr, T.; Kreskowski, K.; Ziegler, M.; Piaszinski, K.; Rittscher, K. The Standard FISH Procedure. In Fluorescence In Situ Hybridisation (FISH); Springer Nature: Berlin, Germany, 2016; pp. 109–118. [Google Scholar]

| C. calyptratus (CCA) Flow Sorting Peak | Anolis carolinensis (ACA) | Phrynosoma platyrhinos (PPL) | Sceloporus tristichus (STR) |
|---|---|---|---|
| CCA1 | ACA2 | PPLma2 | STR2 |
| ACA15 | PPLmi6 * (part’) ** | STR11 | |
| ACA16 | PPLmi8 | ||
| CCA2 | ACA1 | PPLma1 | STR1 |
| CCA3 | ACA3 | PPLma3 | STR3 |
| ACA18 | PPLmi11 | ||
| ACA6 (small fragment) | |||
| CCA4 | ACA4 | PPLma4 | STR4 |
| STR8 (part’’) | |||
| CCA5 | ACA5 ACAX | PPLma5 PPLmi9 (X) | STR5 STR10 |
| CCA6 | ACA6 | PPLma6 | STR6 (part’) |
| CCA7 | ACA7 ACA14 | PPLmi1 PPLmi6 (part’’) | STR6 (part’’) STR9 (part’’) |
| CCA8 | ACA9 ACA17 | PPLmi4 PPLmi10 | STR7 STR9 (part’) STR9 (part’’’) |
| CCA9 | ACA8 | PPLmi2 | STR8 |
| CCA10,11 | ACA10 ACA11 | PPLmi3 PPLmi5 | STR6 (part’’’) STR7 (part’) |
| CCA12 | ACA12 | PPLmi7 | STR8 |
| Repetitive Element Name | Monomer Length, bp | Genome Fraction | GC-Content, % | GenBank Accession Number |
|---|---|---|---|---|
| CCA_s1 | 462 | 0.077 | 35.4 | OP297933 |
| CCA_s2 | 98 | 0.045 | 38.8 | OP297934 |
| CCA_s3 | 1228 | 0.024 | 39.1 | OP297935 |
| CCA_s4 | 51 | 0.13 | 54.9 | OP297936 |
| CCA_s5 | 880 | 0.021 | 49 | OP297937 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tishakova, K.V.; Prokopov, D.Y.; Davletshina, G.I.; Rumyantsev, A.V.; O’Brien, P.C.M.; Ferguson-Smith, M.A.; Giovannotti, M.; Lisachov, A.P.; Trifonov, V.A. Identification of Iguania Ancestral Syntenic Blocks and Putative Sex Chromosomes in the Veiled Chameleon (Chamaeleo calyptratus, Chamaeleonidae, Iguania). Int. J. Mol. Sci. 2022, 23, 15838. https://doi.org/10.3390/ijms232415838
Tishakova KV, Prokopov DY, Davletshina GI, Rumyantsev AV, O’Brien PCM, Ferguson-Smith MA, Giovannotti M, Lisachov AP, Trifonov VA. Identification of Iguania Ancestral Syntenic Blocks and Putative Sex Chromosomes in the Veiled Chameleon (Chamaeleo calyptratus, Chamaeleonidae, Iguania). International Journal of Molecular Sciences. 2022; 23(24):15838. https://doi.org/10.3390/ijms232415838
Chicago/Turabian StyleTishakova, Katerina V., Dmitry Yu. Prokopov, Guzel I. Davletshina, Alexander V. Rumyantsev, Patricia C. M. O’Brien, Malcolm A. Ferguson-Smith, Massimo Giovannotti, Artem P. Lisachov, and Vladimir A. Trifonov. 2022. "Identification of Iguania Ancestral Syntenic Blocks and Putative Sex Chromosomes in the Veiled Chameleon (Chamaeleo calyptratus, Chamaeleonidae, Iguania)" International Journal of Molecular Sciences 23, no. 24: 15838. https://doi.org/10.3390/ijms232415838
APA StyleTishakova, K. V., Prokopov, D. Y., Davletshina, G. I., Rumyantsev, A. V., O’Brien, P. C. M., Ferguson-Smith, M. A., Giovannotti, M., Lisachov, A. P., & Trifonov, V. A. (2022). Identification of Iguania Ancestral Syntenic Blocks and Putative Sex Chromosomes in the Veiled Chameleon (Chamaeleo calyptratus, Chamaeleonidae, Iguania). International Journal of Molecular Sciences, 23(24), 15838. https://doi.org/10.3390/ijms232415838









