Investigation of MicroRNA Biomarkers in Equine Distal Interphalangeal Joint Osteoarthritis
Abstract
1. Introduction
2. Results and Discussion
2.1. Samples
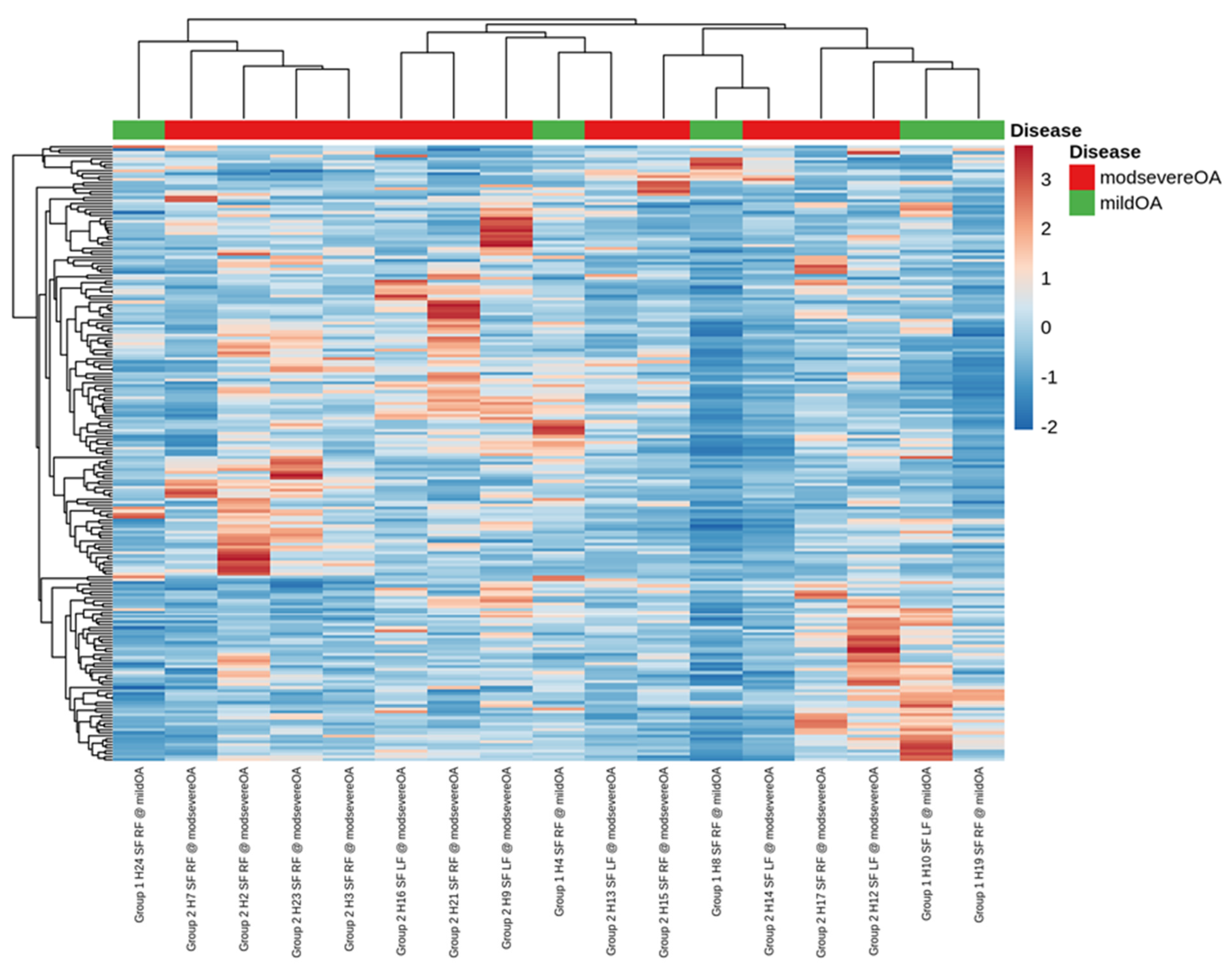
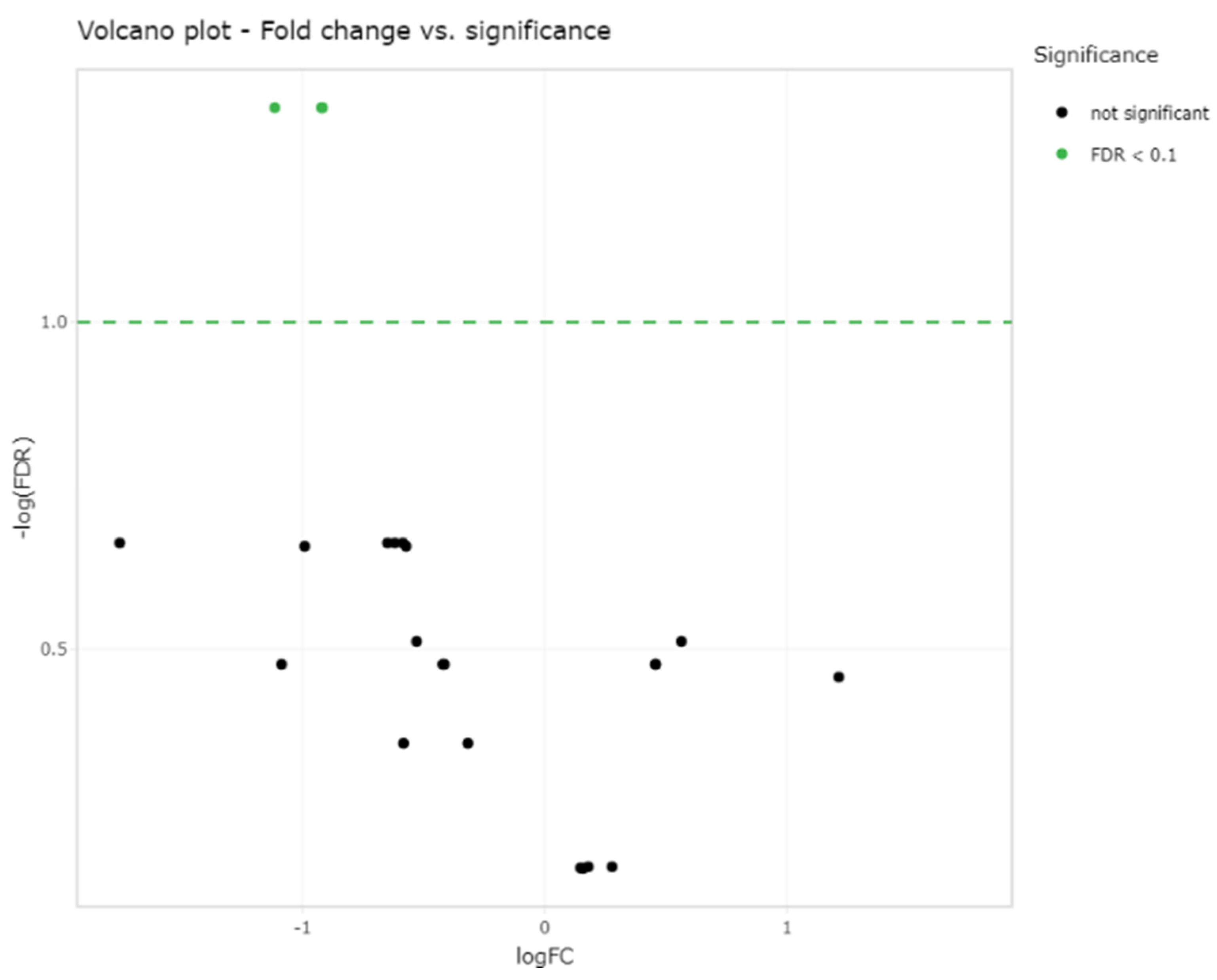
2.2. Small RNA Sequencing Analysis
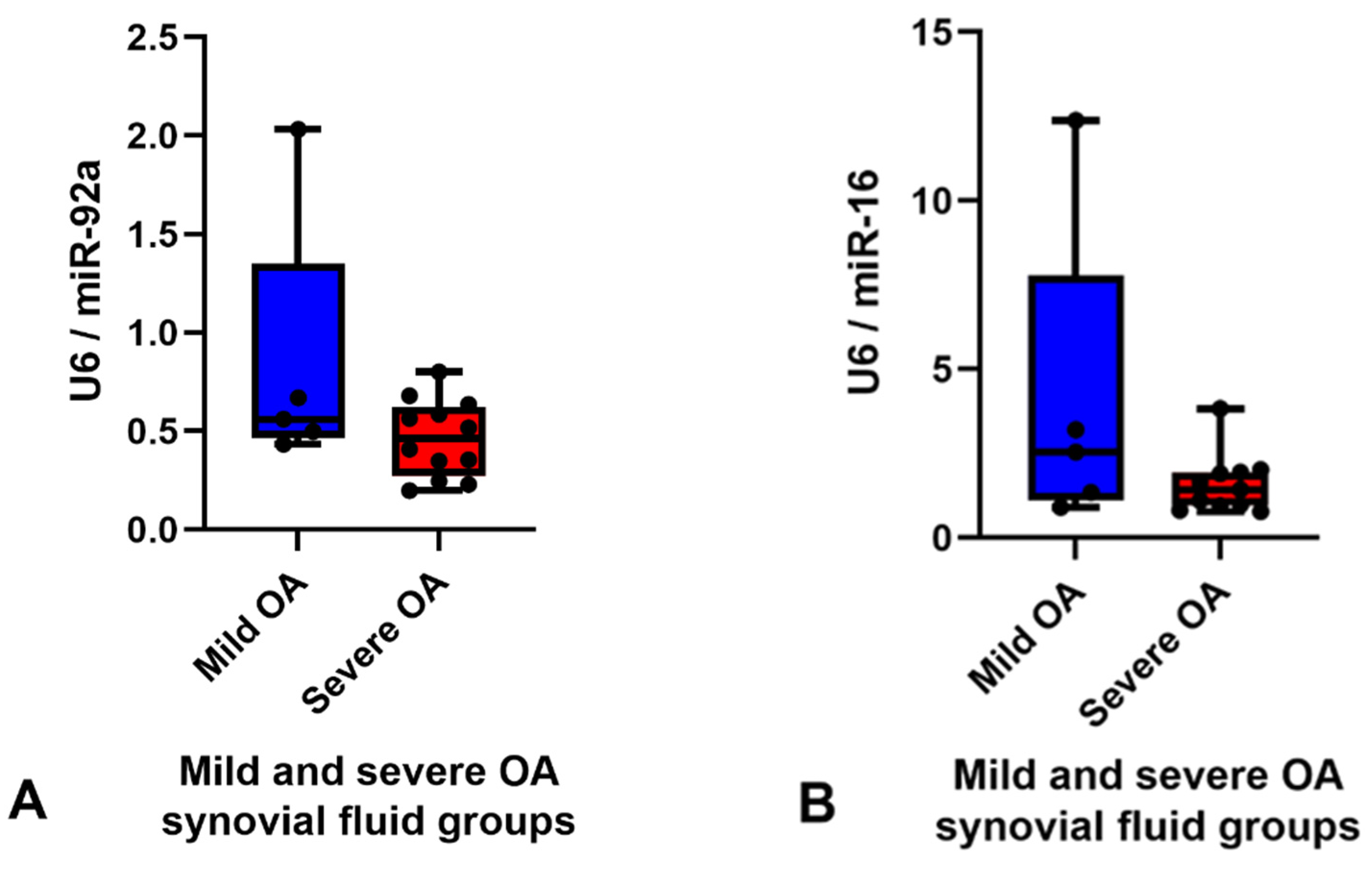
2.3. Validation of Small RNA Sequencing by RT-qPCR
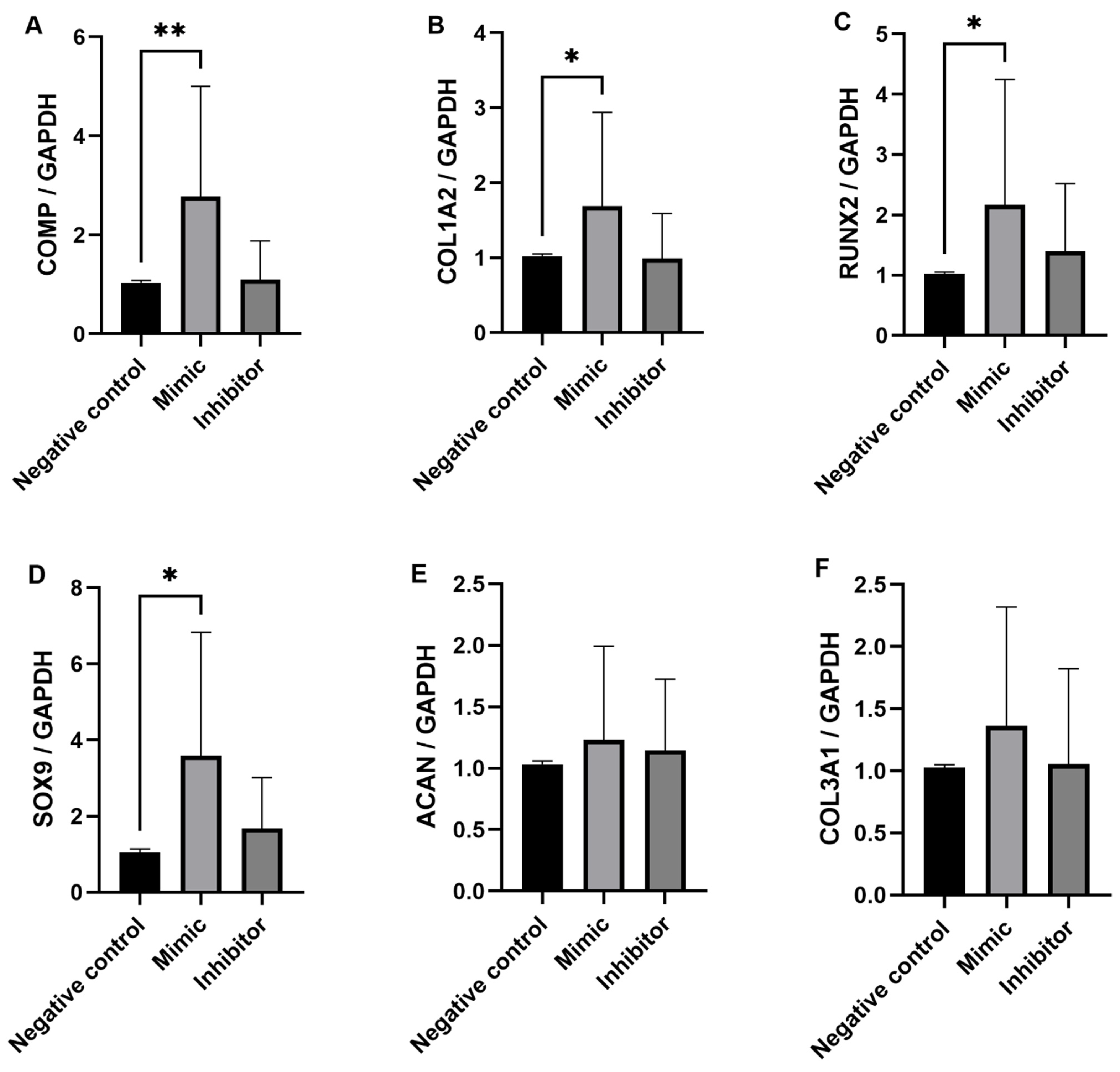
2.4. Gain and Loss Function Studies for miR-92a
Discussion
3. Methods and Materials
3.1. Sample Collection
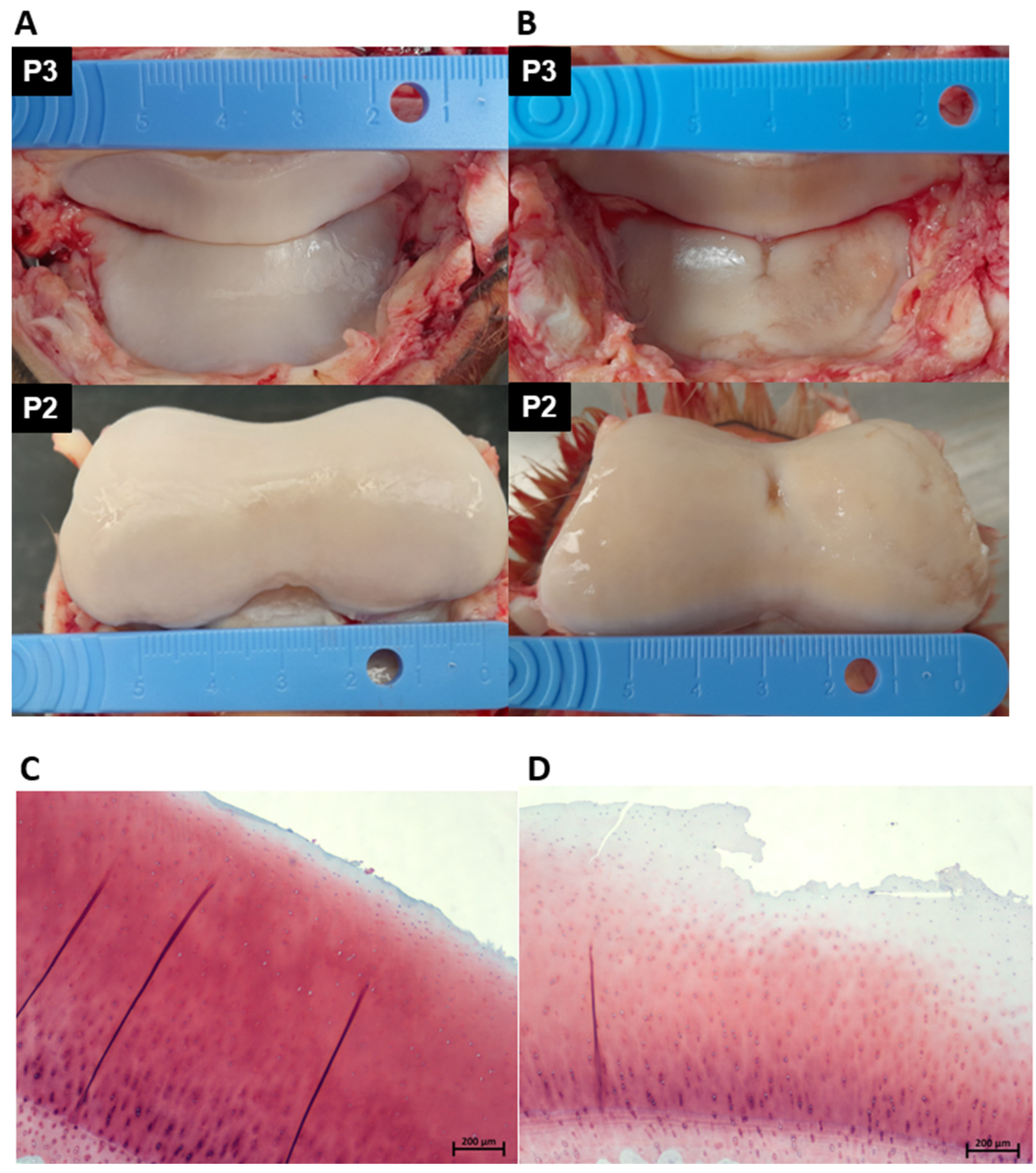
3.2. Macroscopic and Microscopic Cartilage Assessment
3.3. Total RNA Extraction, Library Preparation and Small RNA Sequencing
3.4. Small RNA Sequencing Data Analysis
3.5. Small RNA Sequencing Validation
3.6. Primary Chondrocyte Collection, Isolation and Cell Culture
3.7. miRNA Cell Transfection
3.8. RNA Extraction, Reverse Transcription and RT-qPCR of Equine Chondrocytes
3.9. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ireland, J.L.; Clegg, P.D.; McGowan, C.M.; Platt, L.; Pinchbeck, G.L. Factors associated with mortality of geriatric horses in the United Kingdom. Prev. Vet. Med. 2011, 101, 204–218. [Google Scholar] [CrossRef] [PubMed]
- Ireland, J.L.; Clegg, P.D.; McGowan, C.M.; McKane, S.A.; Chandler, K.J.; Pinchbeck, G.L. Disease prevalence in geriatric horses in the United Kingdom: Veterinary clinical assessment of 200 cases. Equine Vet. J. 2012, 44, 101–106. [Google Scholar] [CrossRef] [PubMed]
- McIlwraith, C.W. 3-Traumatic Arthritis and Posttraumatic Osteoarthritis in the Horse. In Joint Disease in the Horse, 2nd ed.; McIlwraith, C.W., Frisbie, D.D., Kawcak, C.E., van Weeren, P.R., Eds.; W.B. Saunders: Edinburgh, Scotland, 2016; pp. 33–48. [Google Scholar] [CrossRef]
- Vincent, T.L.; Watt, F.E. Osteoarthritis. Medicine 2014, 42, 213–219. [Google Scholar] [CrossRef]
- Brommer, H.; van Weeren, P.R.; Brama, P.A.; Barneveld, A. Quantification and age-related distribution of articular cartilage degeneration in the equine fetlock joint. Equine Vet. J. 2003, 35, 697–701. [Google Scholar] [CrossRef]
- Wilsher, S.; Allen, W.R.; Wood, J.L. Factors associated with failure of thoroughbred horses to train and race. Equine Vet. J. 2006, 38, 113–118. [Google Scholar] [CrossRef]
- van Zadelhoff, C.; Schwarz, T.; Smith, S.; Engerand, A.; Taylor, S. Identification of Naturally Occurring Cartilage Damage in the Equine Distal Interphalangeal Joint Using Low-Field Magnetic Resonance Imaging and Magnetic Resonance Arthrography. Front. Vet. Sci. 2020, 6, 508. [Google Scholar] [CrossRef]
- Global Burden of Disease Study 2013 Collaborators. Global, regional, and national incidence, prevalence, and years lived with disability for 301 acute and chronic diseases and injuries in 188 countries, 1990–2013: A systematic analysis for the Global Burden of Disease Study 2013. Lancet 2015, 386, 743–800. [Google Scholar] [CrossRef]
- Reginster, J.Y. The prevalence and burden of arthritis. Rheumatology 2002, 41, 3–6. [Google Scholar] [CrossRef]
- Peterfy, C.G.; Guermazi, A.; Zaim, S.; Tirman, P.F.; Miaux, Y.; White, D.; Kothari, M.; Lu, Y.; Fye, K.; Zhao, S.; et al. Whole-Organ Magnetic Resonance Imaging Score (WORMS) of the knee in osteoarthritis. Osteoarthr. Cartil. 2004, 12, 177–190. [Google Scholar] [CrossRef]
- Peterfy, C.G.; Schneider, E.; Nevitt, M. The osteoarthritis initiative: Report on the design rationale for the magnetic resonance imaging protocol for the knee. Osteoarthr. Cartil. 2008, 16, 1433–1441. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Ramchandran, R.; Chaluvally-Raghavan, P. miRNA-Mediated RNA Activation in Mammalian Cells. Adv. Exp. Med. Biol. 2017, 983, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Zhang, W.; Li, F.; Guo, F.; Chen, A. miR-139 is up-regulated in osteoarthritis and inhibits chondrocyte proliferation and migration possibly via suppressing EIF4G2 and IGF1R. Biochem. Biophys. Res. Commun. 2016, 474, 296–302. [Google Scholar] [CrossRef]
- Tao, S.C.; Yuan, T.; Zhang, Y.L.; Yin, W.J.; Guo, S.C.; Zhang, C.Q. Exosomes derived from miR-140-5p-overexpressing human synovial mesenchymal stem cells enhance cartilage tissue regeneration and prevent osteoarthritis of the knee in a rat model. Theranostics 2017, 7, 180–195. [Google Scholar] [CrossRef] [PubMed]
- Ntoumou, E.; Tzetis, M.; Braoudaki, M.; Lambrou, G.; Poulou, M.; Malizos, K.; Stefanou, N.; Anastasopoulou, L.; Tsezou, A. Serum microRNA array analysis identifies miR-140-3p, miR-33b-3p and miR-671-3p as potential osteoarthritis biomarkers involved in metabolic processes. Clin. Epigenetics 2017, 9, 127. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- Laterza, O.F.; Lim, L.; Garrett-Engele, P.W.; Vlasakova, K.; Muniappa, N.; Tanaka, W.K.; Johnson, J.M.; Sina, J.F.; Fare, T.L.; Sistare, F.D.; et al. Plasma MicroRNAs as sensitive and specific biomarkers of tissue injury. Clin. Chem. 2009, 55, 1977–1983. [Google Scholar] [CrossRef]
- Park, N.J.; Zhou, H.; Elashoff, D.; Henson, B.S.; Kastratovic, D.A.; Abemayor, E.; Wong, D.T. Salivary microRNA: Discovery, characterization, and clinical utility for oral cancer detection. Clin. Cancer Res. 2009, 15, 5473–5477. [Google Scholar] [CrossRef]
- Sapre, N.; Macintyre, G.; Clarkson, M.; Naeem, H.; Cmero, M.; Kowalczyk, A.; Anderson, P.D.; Costello, A.J.; Corcoran, N.M.; Hovens, C.M. A urinary microRNA signature can predict the presence of bladder urothelial carcinoma in patients undergoing surveillance. Br. J. Cancer 2016, 114, 454–462. [Google Scholar] [CrossRef]
- Kosaka, N.; Izumi, H.; Sekine, K.; Ochiya, T. microRNA as a new immune-regulatory agent in breast milk. Silence 2010, 1, 7. [Google Scholar] [CrossRef]
- Murata, K.; Yoshitomi, H.; Tanida, S.; Ishikawa, M.; Nishitani, K.; Ito, H.; Nakamura, T. Plasma and synovial fluid microRNAs as potential biomarkers of rheumatoid arthritis and osteoarthritis. Arthritis. Res. Ther. 2010, 12, R86. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Chen, W.; Zhao, M.; Xu, Y.; Yu, H.; Qin, J.; Li, H. Exploration of Exosomal miRNAs from Serum and Synovial Fluid in Arthritis Patients. Diagnostics 2022, 12, 239. [Google Scholar] [CrossRef] [PubMed]
- Antunes, J.; Koch, T.G.; Koenig, J.; Cote, N.; Dubois, M.S. On the road to biomarkers: Developing a robust system for miRNA evaluation in equine blood and synovial fluid. Osteoarthr. Cartil. 2019, 27, S110–S111. [Google Scholar] [CrossRef]
- Castanheira, C.; Balaskas, P.; Falls, C.; Ashraf-Kharaz, Y.; Clegg, P.; Burke, K.; Fang, Y.; Dyer, P.; Welting, T.J.M.; Peffers, M.J. Equine synovial fluid small non-coding RNA signatures in early osteoarthritis. BMC Vet. Res. 2021, 17, 26. [Google Scholar] [CrossRef]
- Anderson, J.R.; Jacobsen, S.; Walters, M.; Bundgaard, L.; Diendorfer, A.; Hackl, M.; Clarke, E.J.; James, V.; Peffers, M.J. Small non-coding RNA landscape of extracellular vesicles from a post-traumatic model of equine osteoarthritis. Front. Vet. Sci. 2022, 9, 901269. [Google Scholar] [CrossRef]
- Yang, Q.; Zhou, Y.; Cai, P.; Fu, W.; Wang, J.; Wei, Q.; Li, X. Downregulation of microRNA-23b-3p alleviates IL-1beta-induced injury in chondrogenic CHON-001 cells. Drug Des. Devel. Ther. 2019, 13, 2503–2512. [Google Scholar] [CrossRef]
- Guo, Y.; Min, Z.; Jiang, C.; Wang, W.; Yan, J.; Xu, P.; Xu, K.; Xu, J.; Sun, M.; Zhao, Y.; et al. Downregulation of HS6ST2 by miR-23b-3p enhances matrix degradation through p38 MAPK pathway in osteoarthritis. Cell Death Dis. 2018, 9, 699. [Google Scholar] [CrossRef]
- Lu, X.; Weng, X.; Li, Z.; Yang, B.; Qian, J. Suppression of microRNA-23a/b-3p delays the development degenerative joint disease by activating Grem1 and TGF-beta/Smad signaling pathway. Am. J. Physiol. Cell Physiol. 2020, 319, C456. [Google Scholar] [CrossRef]
- Diendorfer, A.; Khamina, K.; Pultar, M.; Hackl, M. miND (miRNA NGS Discovery pipeline): A small RNA-seq analysis pipeline and report generator for microRNA biomarker discovery studies [version 1; peer review: 1 approved with reservations]. F1000Research 2022, 11, 233. [Google Scholar] [CrossRef]
- Björnsdóttir, S.; Axelsson, M.; Eksell, P.; Sigurdsson, H.; Carlsten, J. Radiographic and clinical survey of degenerative joint disease in the distal tarsal joints in Icelandic horses. Equine Vet. J. 2000, 32, 268–272. [Google Scholar] [CrossRef]
- Byam-Cook, K.L.; Singer, E.R. Is there a relationship between clinical presentation, diagnostic and radiographic findings and outcome in horses with osteoarthritis of the small tarsal joints? Equine Vet. J. 2009, 41, 118–123. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.X.; He, S.H.; Liang, X.; Li, W.; Li, T.F.; Li, D.F. Aging, Cell Senescence, the Pathogenesis and Targeted Therapies of Osteoarthritis. Front. Pharmacol. 2021, 12, 728100. [Google Scholar] [CrossRef] [PubMed]
- Childs, B.G.; Gluscevic, M.; Baker, D.J.; Laberge, R.M.; Marquess, D.; Dananberg, J.; van Deursen, J.M. Senescent cells: An emerging target for diseases of ageing. Nat. Rev. Drug Discov. 2017, 16, 718–735. [Google Scholar] [CrossRef] [PubMed]
- Yin, C.M.; Suen, W.C.; Lin, S.; Wu, X.M.; Li, G.; Pan, X.H. Dysregulation of both miR-140-3p and miR-140-5p in synovial fluid correlate with osteoarthritis severity. Bone Jt. Res. 2017, 6, 612–618. [Google Scholar] [CrossRef]
- Xu, J.F.; Zhang, S.J.; Zhao, C.; Qiu, B.S.; Gu, H.F.; Hong, J.F.; Cao, L.; Chen, Y.; Xia, B.; Bi, Q.; et al. Altered microRNA expression profile in synovial fluid from patients with knee osteoarthritis with treatment of hyaluronic acid. Mol. Diagn. Ther. 2015, 19, 299–308. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.; Su, W.; Xia, H.; Wang, Z.; Su, C.; Su, B. Synovial Fluid MicroRNA-210 as a Potential Biomarker for Early Prediction of Osteoarthritis. Biomed. Res. Int. 2019, 2019, 7165406. [Google Scholar] [CrossRef]
- Li, H.Y.; Tavallaee, G.; Tokar, T.; Nakamura, A.; Sundararajan, K.; Weston, A.; Sharma, A.; Mahomed, N.N.; Gandhi, R.; Jurisica, I.; et al. Identification of synovial fluid microRNA signature in knee osteoarthritis: Differentiating early- and late-stage knee osteoarthritis. Osteoarthr. Cartil. 2016, 24, 1577–1586. [Google Scholar] [CrossRef]
- Hamati, M.C.; Maynard, R.; Bartolomei, J.; Zuscik, M.J.; Ackert-Bicknell, C.L.; Hunt, K.J. Toward Preventing Arthritis: A Comparison of Synovial RNA in Osteoarthritic Ankles and Knees. Foot Ankle Orthop. 2022, 7, 2473011421S2473000224. [Google Scholar] [CrossRef]
- Arroyo, J.D.; Chevillet, J.R.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.S.; Bennett, C.F.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef]
- Patton, J.G.; Franklin, J.L.; Weaver, A.M.; Vickers, K.; Zhang, B.; Coffey, R.J.; Mark Ansel, K.; Blelloch, R.; Goga, A.; Huang, B.; et al. Biogenesis, delivery, and function of extracellular RNA. J. Extracell. Vesicles 2015, 4, 27494. [Google Scholar] [CrossRef]
- Everaert, C.; Luypaert, M.; Maag, J.L.V.; Cheng, Q.X.; Dinger, M.E.; Hellemans, J.; Mestdagh, P. Benchmarking of RNA-sequencing analysis workflows using whole-transcriptome RT-qPCR expression data. Sci. Rep. 2017, 7, 1559. [Google Scholar] [CrossRef] [PubMed]
- Xia, B.; Di, C.; Zhang, J.; Hu, S.; Jin, H.; Tong, P. Osteoarthritis pathogenesis: A review of molecular mechanisms. Calcif. Tissue Int. 2014, 95, 495–505. [Google Scholar] [CrossRef]
- Martel-Pelletier, J.; Barr, A.J.; Cicuttini, F.M.; Conaghan, P.G.; Cooper, C.; Goldring, M.B.; Goldring, S.R.; Jones, G.; Teichtahl, A.J.; Pelletier, J.P. Osteoarthritis. Nat. Rev. Dis. Prim. 2016, 2, 16072. [Google Scholar] [CrossRef]
- Mao, G.; Wu, P.; Zhang, Z.; Zhang, Z.; Liao, W.; Li, Y.; Kang, Y. MicroRNA-92a-3p Regulates Aggrecanase-1 and Aggrecanase-2 Expression in Chondrogenesis and IL-1β-Induced Catabolism in Human Articular Chondrocytes. Cell. Physiol. Biochem. 2017, 44, 38–52. [Google Scholar] [CrossRef]
- Mao, G.; Zhang, Z.; Huang, Z.; Chen, W.; Huang, G.; Meng, F.; Zhang, Z.; Kang, Y. MicroRNA-92a-3p regulates the expression of cartilage-specific genes by directly targeting histone deacetylase 2 in chondrogenesis and degradation. Osteoarthr. Cartil. 2017, 25, 521–532. [Google Scholar] [CrossRef]
- Mao, G.; Zhang, Z.; Hu, S.; Zhang, Z.; Chang, Z.; Huang, Z.; Liao, W.; Kang, Y. Exosomes derived from miR-92a-3p-overexpressing human mesenchymal stem cells enhance chondrogenesis and suppress cartilage degradation via targeting WNT5A. Stem Cell Res. Ther. 2018, 9, 247. [Google Scholar] [CrossRef]
- De Angelis, E.; Cacchioli, A.; Ravanetti, F.; Bileti, R.; Cavalli, V.; Martelli, P.; Borghetti, P. Gene expression markers in horse articular chondrocytes: Chondrogenic differentiaton IN VITRO depends on the proliferative potential and ageing. Implication for tissue engineering of cartilage. Res. Vet. Sci. 2020, 128, 107–117. [Google Scholar] [CrossRef]
- Semevolos, S.A.; Nixon, A.J.; Brower-Toland, B.D. Changes in molecular expression of aggrecan and collagen types I, II, and X, insulin-like growth factor-I, and transforming growth factor-beta1 in articular cartilage obtained from horses with naturally acquired osteochondrosis. Am. J. Vet. Res. 2001, 62, 1088–1094. [Google Scholar] [CrossRef] [PubMed]
- Benya, P.D.; Padilla, S.R.; Nimni, M.E. Independent regulation of collagen types by chondrocytes during the loss of differentiated function in culture. Cell 1978, 15, 1313–1321. [Google Scholar] [CrossRef]
- Von der Mark, K.; Gauss, V.; von der Mark, H.; Müller, P. Relationship between cell shape and type of collagen synthesised as chondrocytes lose their cartilage phenotype in culture. Nature 1977, 267, 531–532. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.-C.; Lee, S.-W.; Ryu, D.-Y.; Cui, F.-J.; Bhak, J.; Kim, Y. Identification and Characterization of MicroRNAs in Normal Equine Tissues by Next Generation Sequencing. PLoS ONE 2014, 9, e93662. [Google Scholar] [CrossRef]
- Sebastian, A.; McCool, J.L.; Hum, N.R.; Murugesh, D.K.; Wilson, S.P.; Christiansen, B.A.; Loots, G.G. Single-Cell RNA-Seq Reveals Transcriptomic Heterogeneity and Post-Traumatic Osteoarthritis-Associated Early Molecular Changes in Mouse Articular Chondrocytes. Cells 2021, 10, 1462. [Google Scholar] [CrossRef]
- Lecchi, C.; Dalla Costa, E.; Lebelt, D.; Ferrante, V.; Canali, E.; Ceciliani, F.; Stucke, D.; Minero, M. Circulating miR-23b-3p, miR-145-5p and miR-200b-3p are potential biomarkers to monitor acute pain associated with laminitis in horses. Animal 2018, 12, 366–375. [Google Scholar] [CrossRef] [PubMed]
- Caron, M.M.J.; Emans, P.J.; Coolsen, M.M.E.; Voss, L.; Surtel, D.A.M.; Cremers, A.; van Rhijn, L.W.; Welting, T.J.M. Redifferentiation of dedifferentiated human articular chondrocytes: Comparison of 2D and 3D cultures. Osteoarthr. Cartil. 2012, 20, 1170–1178. [Google Scholar] [CrossRef]
- Bernstein, P.; Dong, M.; Corbeil, D.; Gelinsky, M.; Günther, K.P.; Fickert, S. Pellet culture elicits superior chondrogenic redifferentiation than alginate-based systems. Biotechnol. Prog. 2009, 25, 1146–1152. [Google Scholar] [CrossRef] [PubMed]
- Ortved, K.F.; Nixon, A.J. Cell-based cartilage repair strategies in the horse. Vet. J. 2016, 208, 1–12. [Google Scholar] [CrossRef] [PubMed]
- McIlwraith, C.W.; Frisbie, D.D.; Kawcak, C.E.; Fuller, C.J.; Hurtig, M.; Cruz, A. The OARSI histopathology initiative-recommendations for histological assessments of osteoarthritis in the horse. Osteoarthr. Cartil. 2010, 18 (Suppl. 3), S93–S105. [Google Scholar] [CrossRef] [PubMed]
- Pritzker, K.P.H.; Gay, S.; Jimenez, S.A.; Ostergaard, K.; Pelletier, J.P.; Revell, P.A.; Salter, D.; van den Berg, W.B. Osteoarthritis cartilage histopathology: Grading and staging. Osteoarthr. Cartil. 2006, 14, 13–29. [Google Scholar] [CrossRef] [PubMed]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 11 April 2022).
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: Summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 2011, 17, 3. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome. Biol. 2009, 10, R25. [Google Scholar] [CrossRef]
- Friedländer, M.R.; Mackowiak, S.D.; Li, N.; Chen, W.; Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2011, 40, 37–52. [Google Scholar] [CrossRef]
- Zerbino, D.R.; Achuthan, P.; Akanni, W.; Amode, M.R.; Barrell, D.; Bhai, J.; Billis, K.; Cummins, C.; Gall, A.; Girón, C.G.; et al. Ensembl 2018. Nucleic Acids Res. 2017, 46, D754–D761. [Google Scholar] [CrossRef]
- Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Res. 2004, 32, D109–D111. [Google Scholar] [CrossRef]
- The RNAcentral Consortium. RNAcentral: A hub of information for non-coding RNA sequences. Nucleic Acids Res. 2018, 47, D221–D229. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2009, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Andersen, C.L.; Jensen, J.L.; Ørntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]

| Mild OA Group | Severe OA Group | |
|---|---|---|
| Sex | Three geldings Two mares | Six geldings Six mares |
| Mean weight | 550 ± 70 kg | 440 ± 70 kg |
| Mean age | 9 ± 3 years | 18 ± 9 years |
| Breeds | Highland Pony, Dutch Warmblood, Belgian Warmblood and Thoroughbred | Irish Draught, Welsh Pony, Icelandic Pony, Irish Sports Horse, Cob, Cross-breed pony, Thoroughbred, Sports Horse and Connemara Pony |
| Reasons for euthanasia | Colic, lameness, cervical vertebral malformation and one horse had a complete open third metacarpal bone fracture | Colic, lameness, incisional hernia, congestive heart failure, severe behavioural problems and advanced dental disease |
| Horse Number | Breed | Age (Years) | Sex | Weight (kg) | Reason for Euthanasia | Group |
|---|---|---|---|---|---|---|
| H3 | Irish Draught | 28 | G | 500 kg | Lameness | Severe OA |
| H16 | Cross-breed horse | 22 | G | 400 kg | Age-related concerns | Severe OA |
| H17 | Cob-cross | 26 | G | 400 kg | Age-related concerns | Severe OA |
| H26 | Irish Sports Horse | 5 | M | 500 kg | Multi-limb lameness | Mild OA |
| H27 | Cob | 7 | M | 500 kg | Grass sickness | Mild OA |
| H28 | Irish Draught | 12 | G | 600 kg | Lameness | Severe OA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Baker, M.E.; Lee, S.; Clinton, M.; Hackl, M.; Castanheira, C.; Peffers, M.J.; Taylor, S.E. Investigation of MicroRNA Biomarkers in Equine Distal Interphalangeal Joint Osteoarthritis. Int. J. Mol. Sci. 2022, 23, 15526. https://doi.org/10.3390/ijms232415526
Baker ME, Lee S, Clinton M, Hackl M, Castanheira C, Peffers MJ, Taylor SE. Investigation of MicroRNA Biomarkers in Equine Distal Interphalangeal Joint Osteoarthritis. International Journal of Molecular Sciences. 2022; 23(24):15526. https://doi.org/10.3390/ijms232415526
Chicago/Turabian StyleBaker, Melissa E., Seungmee Lee, Michael Clinton, Matthias Hackl, Catarina Castanheira, Mandy J. Peffers, and Sarah E. Taylor. 2022. "Investigation of MicroRNA Biomarkers in Equine Distal Interphalangeal Joint Osteoarthritis" International Journal of Molecular Sciences 23, no. 24: 15526. https://doi.org/10.3390/ijms232415526
APA StyleBaker, M. E., Lee, S., Clinton, M., Hackl, M., Castanheira, C., Peffers, M. J., & Taylor, S. E. (2022). Investigation of MicroRNA Biomarkers in Equine Distal Interphalangeal Joint Osteoarthritis. International Journal of Molecular Sciences, 23(24), 15526. https://doi.org/10.3390/ijms232415526





