miR−122−5p Regulates Renal Fibrosis In Vivo
Abstract
:1. Introduction
2. Results
2.1. Profiling of miRNAs in Renal Fibrosis
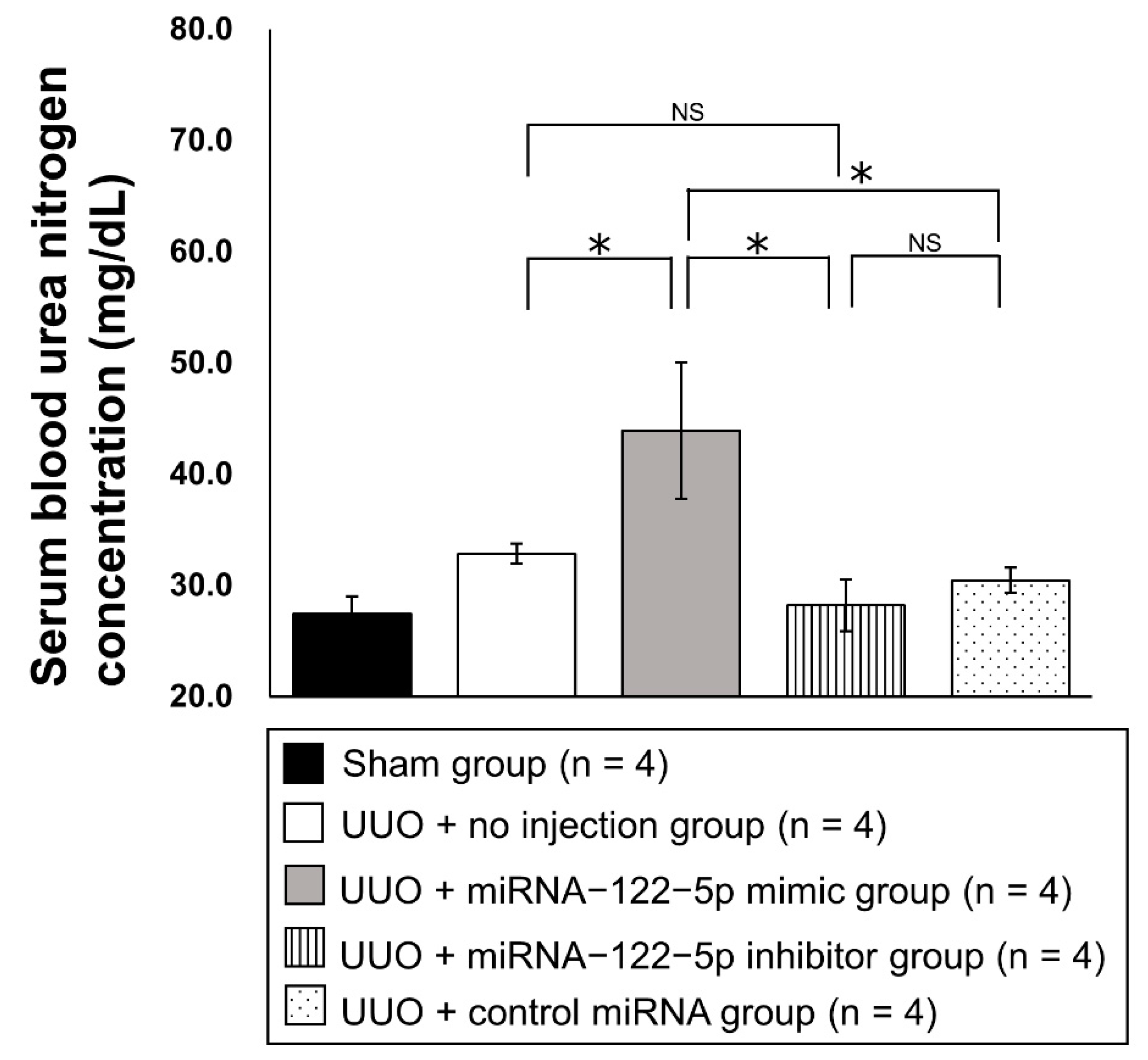
2.2. Regulation of Renal Fibrosis by a Mimic and Inhibitor of miR−122−5p
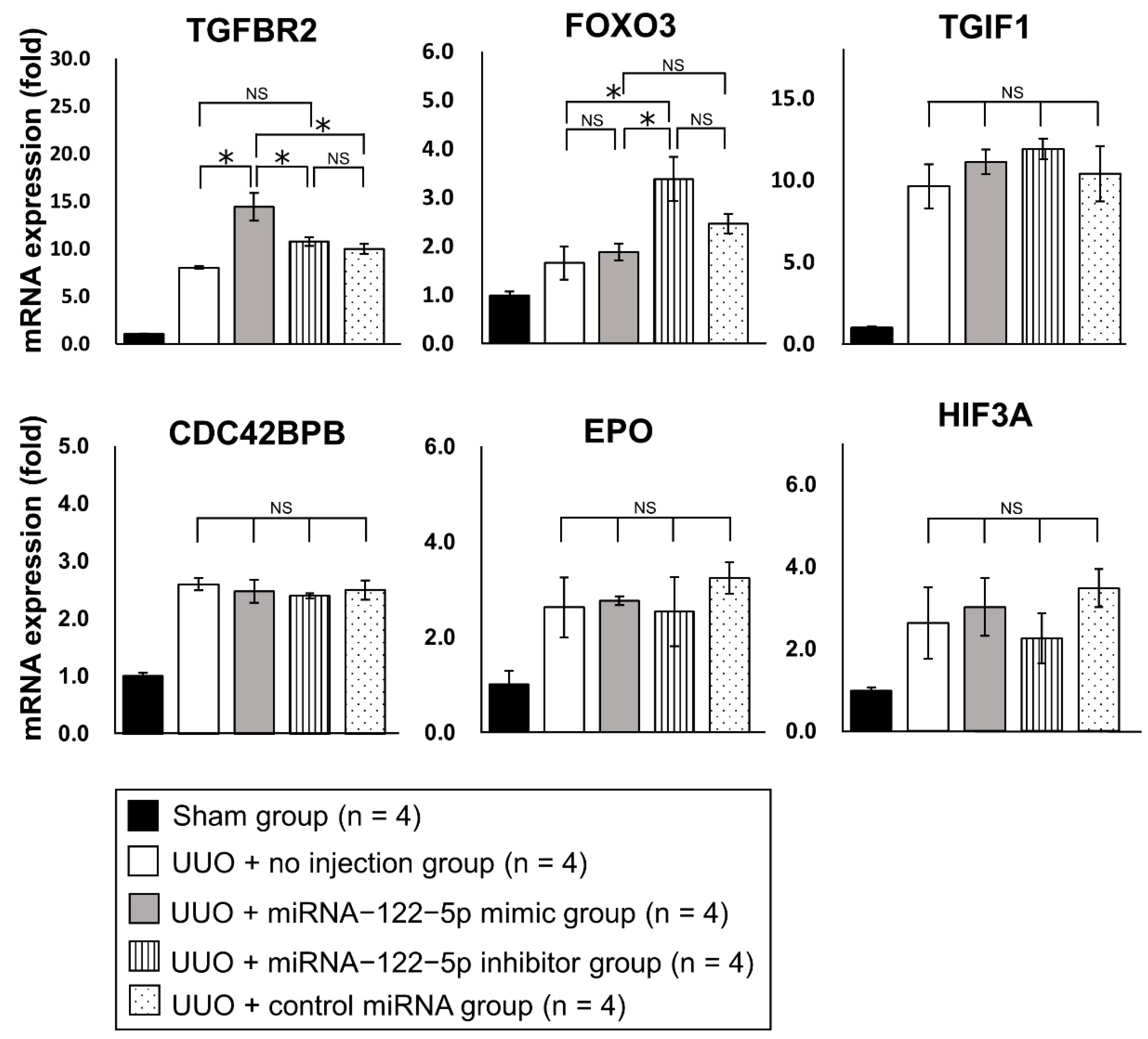
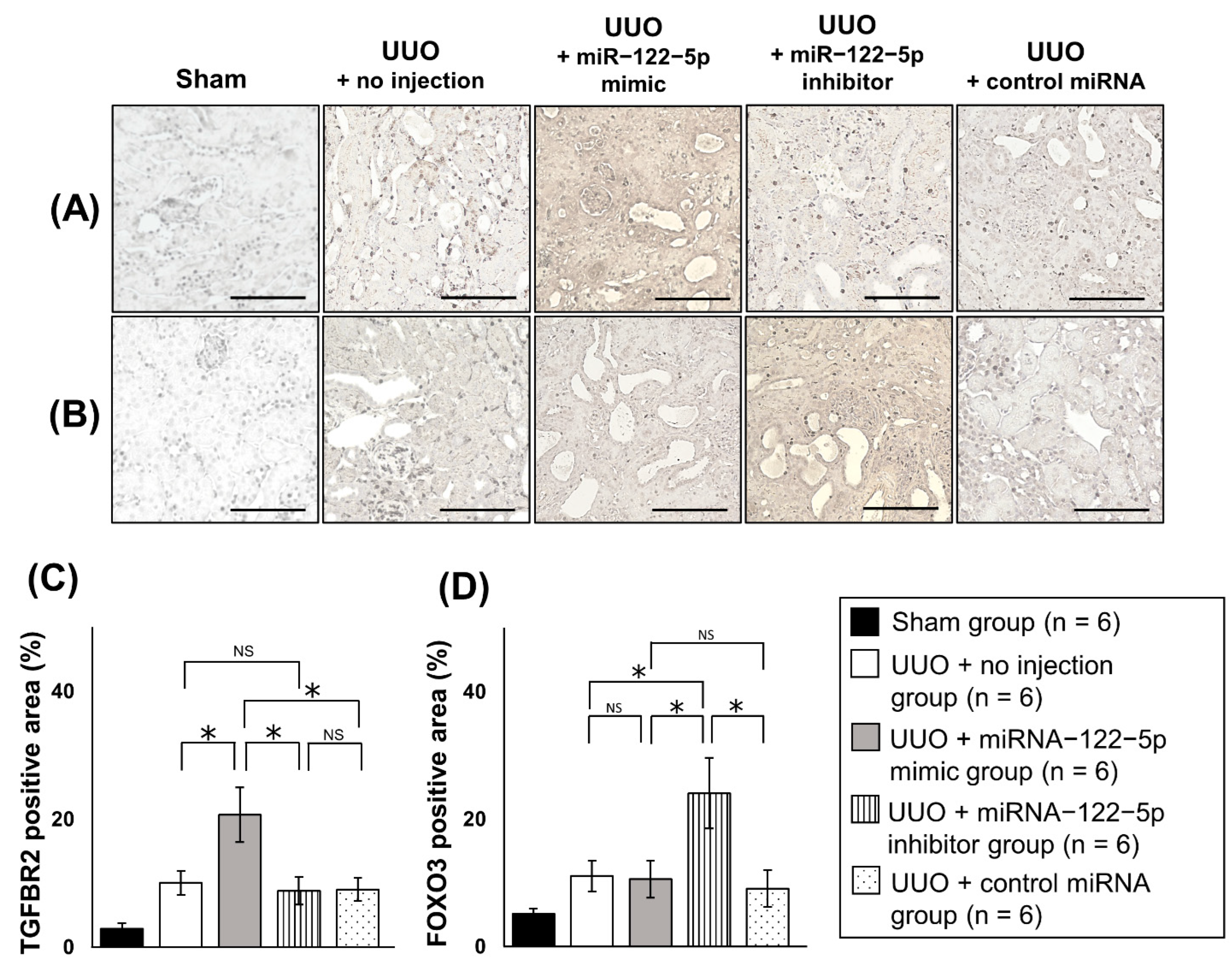
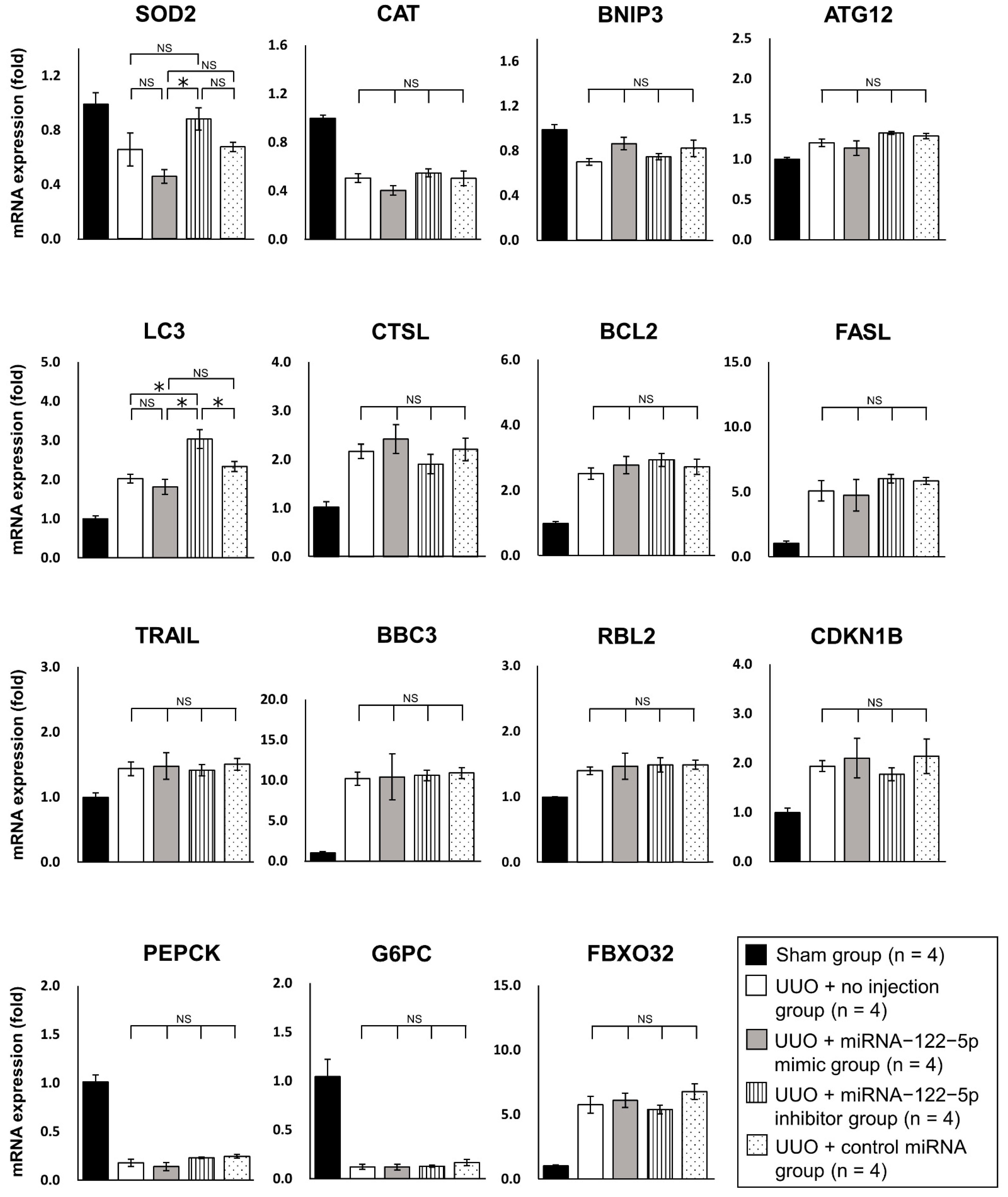
2.3. Mechanism of the Modulation of Renal Fibrosis by miR−122−5p
3. Discussion
4. Materials and Methods
4.1. Ethical Approval
4.2. Renal Fibrotic Mouse Model
4.3. miRNA Microarray Analysis, Data Processing, and Statistical Analysis
4.4. qRT-PCR
4.5. miR−122−5p Mimic and miR−122−5p Inhibitor
4.6. Delivery of the miRNA Mimic and miRNA Inhibitor to Kidneys
4.7. Kidney Histology (AZAN Staining and Sirius Red Staining)
4.8. IHC
4.9. Measurement of Serum BUN
4.10. Nomenclature System for miRNAs
4.11. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Nogueira, A.; Pires, M.J.; Oliveira, P.A. Pathophysiological Mechanisms of Renal Fibrosis: A Review of Animal Models and Therapeutic Strategies. In Vivo 2017, 31, 1–22. [Google Scholar] [CrossRef] [Green Version]
- Bülow, R.D.; Boor, P. Extracellular Matrix in Kidney Fibrosis: More Than Just a Scaffold. J. Histochem. Cytochem. Off. J. Histochem. Soc. 2019, 67, 643–661. [Google Scholar] [CrossRef] [Green Version]
- Lv, W.; Fan, F.; Wang, Y.; Gonzalez-Fernandez, E.; Wang, C.; Yang, L.; Booz, G.W.; Roman, R.J. Therapeutic potential of microRNAs for the treatment of renal fibrosis and CKD. Physiol. Genom. 2018, 50, 20–34. [Google Scholar] [CrossRef]
- Schefold, J.C.; Filippatos, G.; Hasenfuss, G.; Anker, S.D.; von Haehling, S. Heart failure and kidney dysfunction: Epidemiology, mechanisms and management. Nat. Rev. Nephrol. 2016, 12, 610–623. [Google Scholar] [CrossRef]
- Wang, H.; Wang, B.; Zhang, A.; Hassounah, F.; Seow, Y.; Wood, M.; Ma, F.; Klein, J.D.; Price, S.R.; Wang, X.H. Exosome-Mediated miR-29 Transfer Reduces Muscle Atrophy and Kidney Fibrosis in Mice. Mol. Ther. 2019, 27, 571–583. [Google Scholar] [CrossRef] [Green Version]
- Campanholle, G.; Ligresti, G.; Gharib, S.A.; Duffield, J.S. Cellular mechanisms of tissue fibrosis. 3. Novel mechanisms of kidney fibrosis. Am. J. Physiol. Cell Physiol. 2013, 304, C591–C603. [Google Scholar] [CrossRef] [Green Version]
- Cho, M.H. Renal fibrosis. Korean J. Pediatr. 2010, 53, 735–740. [Google Scholar] [CrossRef] [Green Version]
- Humphreys, B.D. Mechanisms of Renal Fibrosis. Annu. Rev. Physiol. 2018, 80, 309–326. [Google Scholar] [CrossRef]
- Morishita, Y.; Imai, T.; Yoshizawa, H.; Watanabe, M.; Ishibashi, K.; Muto, S.; Nagata, D. Delivery of microRNA-146a with polyethylenimine nanoparticles inhibits renal fibrosis in vivo. Int. J. Nanomed. 2015, 10, 3475–3488. [Google Scholar] [CrossRef] [Green Version]
- Fu, Y.; Tang, C.; Cai, J.; Chen, G.; Zhang, D.; Dong, Z. Rodent models of AKI-CKD transition. Am. J. Physiol. Ren. Physiol. 2018, 315, F1098–F1106. [Google Scholar] [CrossRef]
- Vishnoi, A.; Rani, S. MiRNA Biogenesis and Regulation of Diseases: An Overview. Methods Mol. Biol. 2017, 1509, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Rupaimoole, R.; Slack, F.J. MicroRNA therapeutics: Towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 2017, 16, 203–222. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.M.; Tang, P.M.; Li, J.; Lan, H.Y. TGF-β/Smad signaling in renal fibrosis. Front. Physiol. 2015, 6, 82. [Google Scholar] [CrossRef] [Green Version]
- Gomez, I.G.; Nakagawa, N.; Duffield, J.S. MicroRNAs as novel therapeutic targets to treat kidney injury and fibrosis. Am. J. Physiol. Ren. Physiol. 2016, 310, F931–F944. [Google Scholar] [CrossRef] [Green Version]
- Patel, V.; Noureddine, L. MicroRNAs and fibrosis. Curr. Opin. Nephrol. Hypertens. 2012, 21, 410–416. [Google Scholar] [CrossRef]
- Wang, B.; Ricardo, S. Role of microRNA machinery in kidney fibrosis. Clin. Exp. Pharmacol. Physiol. 2014, 41, 543–550. [Google Scholar] [CrossRef] [Green Version]
- Meng, X.M.; Nikolic-Paterson, D.J.; Lan, H.Y. TGF-β: The master regulator of fibrosis. Nat. Rev. Nephrol. 2016, 12, 325–338. [Google Scholar] [CrossRef]
- Van der Hauwaert, C.; Glowacki, F.; Pottier, N.; Cauffiez, C. Non-Coding RNAs as New Therapeutic Targets in the Context of Renal Fibrosis. Int. J. Mol. Sci. 2019, 20, 1977. [Google Scholar] [CrossRef] [Green Version]
- Fierro-Fernández, M.; Miguel, V.; Márquez-Expósito, L.; Nuevo-Tapioles, C.; Herrero, J.I.; Blanco-Ruiz, E.; Tituaña, J.; Castillo, C.; Cannata, P.; Monsalve, M.; et al. MiR-9-5p protects from kidney fibrosis by metabolic reprogramming. FASEB J. 2020, 34, 410–431. [Google Scholar] [CrossRef] [Green Version]
- Glowacki, F.; Savary, G.; Gnemmi, V.; Buob, D.; Van der Hauwaert, C.; Lo-Guidice, J.M.; Bouyé, S.; Hazzan, M.; Pottier, N.; Perrais, M.; et al. Increased circulating miR-21 levels are associated with kidney fibrosis. PLoS ONE 2013, 8, e58014. [Google Scholar] [CrossRef]
- Morizane, R.; Fujii, S.; Monkawa, T.; Hiratsuka, K.; Yamaguchi, S.; Homma, K.; Itoh, H. miR-34c attenuates epithelial-mesenchymal transition and kidney fibrosis with ureteral obstruction. Sci. Rep. 2014, 4, 4578. [Google Scholar] [CrossRef] [Green Version]
- Zeisberg, M.; Kalluri, R. The role of epithelial-to-mesenchymal transition in renal fibrosis. J. Mol. Med. 2004, 82, 175–181. [Google Scholar] [CrossRef]
- Zhong, X.; Chung, A.C.; Chen, H.Y.; Meng, X.M.; Lan, H.Y. Smad3-mediated upregulation of miR-21 promotes renal fibrosis. J. Am. Soc. Nephrol. JASN 2011, 22, 1668–1681. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, Y.; Wang, H.; Li, Y.; Liu, S.; Chen, J.; Ying, H. miR-24 and miR-122 Negatively Regulate the Transforming Growth Factor-β/Smad Signaling Pathway in Skeletal Muscle Fibrosis. Mol. Ther. Nucleic Acids 2018, 11, 528–537. [Google Scholar] [CrossRef] [Green Version]
- Lin, F. Molecular regulation and function of FoxO3 in chronic kidney disease. Curr. Opin. Nephrol. Hypertens. 2020, 29, 439–445. [Google Scholar] [CrossRef]
- Yoon, H.E.; Kim, S.J.; Kim, S.J.; Chung, S.; Shin, S.J. Tempol attenuates renal fibrosis in mice with unilateral ureteral obstruction: The role of PI3K-Akt-FoxO3a signaling. J. Korean Med. Sci. 2014, 29, 230–237. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Kang, H.; Zhang, Q.; D’Agati, V.D.; Al-Awqati, Q.; Lin, F. FoxO3 activation in hypoxic tubules prevents chronic kidney disease. J. Clin. Investig. 2019, 129, 2374–2389. [Google Scholar] [CrossRef] [Green Version]
- Bijkerk, R.; de Bruin, R.G.; van Solingen, C.; van Gils, J.M.; Duijs, J.M.; van der Veer, E.P.; Rabelink, T.J.; Humphreys, B.D.; van Zonneveld, A.J. Silencing of microRNA-132 reduces renal fibrosis by selectively inhibiting myofibroblast proliferation. Kidney Int. 2016, 89, 1268–1280. [Google Scholar] [CrossRef]
- Zhou, H.; Hasni, S.A.; Perez, P.; Tandon, M.; Jang, S.I.; Zheng, C.; Kopp, J.B.; Austin, H., 3rd; Balow, J.E.; Alevizos, I.; et al. miR-150 promotes renal fibrosis in lupus nephritis by downregulating SOCS1. J. Am. Soc. Nephrol. JASN 2013, 24, 1073–1087. [Google Scholar] [CrossRef] [Green Version]
- He, F.; Peng, F.; Xia, X.; Zhao, C.; Luo, Q.; Guan, W.; Li, Z.; Yu, X.; Huang, F. MiR-135a promotes renal fibrosis in diabetic nephropathy by regulating TRPC1. Diabetologia 2014, 57, 1726–1736. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhao, S.; Wu, D.; Liu, X.; Shi, M.; Wang, Y.; Zhang, F.; Ding, J.; Xiao, Y.; Guo, B. MicroRNA-22 Promotes Renal Tubulointerstitial Fibrosis by Targeting PTEN and Suppressing Autophagy in Diabetic Nephropathy. J. Diabetes Res. 2018, 2018, 4728645. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, M.L.; Khosroheidari, M.; Eddy, E.; Kiefer, J. Role of microRNA 1207-5P and its host gene, the long non-coding RNA Pvt1, as mediators of extracellular matrix accumulation in the kidney: Implications for diabetic nephropathy. PLoS ONE 2013, 8, e77468. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, Y.; Zhao, Y.; Wang, L.; Zhang, W.; Liu, C.; Yin, A. MicroRNA-194 overexpression protects against hypoxia/reperfusion-induced HK-2 cell injury through direct targeting Rheb. J. Cell Biochem. 2018. Online ahead of print. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Yang, Z.; Heng, Y.; Miao, C. MicroRNA-181 exerts an inhibitory role during renal fibrosis by targeting early growth response factor-1 and attenuating the expression of profibrotic markers. Mol. Med. Rep. 2019, 19, 3305–3313. [Google Scholar] [CrossRef] [Green Version]
- Ning, Y.X.; Wang, X.Y.; Wang, J.Q.; Zeng, R.; Wang, G.Q. miR-152 regulates TGF-β1-induced epithelial-mesenchymal transition by targeting HPIP in tubular epithelial cells. Mol. Med. Rep. 2018, 17, 7973–7979. [Google Scholar] [CrossRef] [PubMed]
- Halász, T.; Horváth, G.; Pár, G.; Werling, K.; Kiss, A.; Schaff, Z.; Lendvai, G. miR-122 negatively correlates with liver fibrosis as detected by histology and FibroScan. World J. Gastroenterol. 2015, 21, 7814–7823. [Google Scholar] [CrossRef] [Green Version]
- Zhuang, Y.; Yang, D.; Shi, S.; Wang, L.; Yu, M.; Meng, X.; Fan, Y.; Zhou, R.; Wang, F. MiR-375-3p Promotes Cardiac Fibrosis by Regulating the Ferroptosis Mediated by GPX4. Comput. Intell. Neurosci. 2022, 2022, 9629158. [Google Scholar] [CrossRef]
- Yang, Y.Z.; Zhao, X.J.; Xu, H.J.; Wang, S.C.; Pan, Y.; Wang, S.J.; Xu, Q.; Jiao, R.Q.; Gu, H.M.; Kong, L.D. Author Correction: Magnesium isoglycyrrhizinate ameliorates high fructose-induced liver fibrosis in rat by increasing miR-375-3p to suppress JAK2/STAT3 pathway and TGF-β1/Smad signaling. Acta Pharmacol. Sin. 2022, 43, 1616. [Google Scholar] [CrossRef]
- Conde, E.; Giménez-Moyano, S.; Martín-Gómez, L.; Rodríguez, M.; Ramos, M.E.; Aguado-Fraile, E.; Blanco-Sanchez, I.; Saiz, A.; García-Bermejo, M.L. HIF-1α induction during reperfusion avoids maladaptive repair after renal ischemia/reperfusion involving miR127-3p. Sci. Rep. 2017, 7, 41099. [Google Scholar] [CrossRef] [Green Version]
- Biernacka, A.; Dobaczewski, M.; Frangogiannis, N.G. TGF-β signaling in fibrosis. Growth Factors 2011, 29, 196–202. [Google Scholar] [CrossRef]
- Xin, Z.; Ma, Z.; Hu, W.; Jiang, S.; Yang, Z.; Li, T.; Chen, F.; Jia, G.; Yang, Y. FOXO1/3: Potential suppressors of fibrosis. Ageing Res. Rev. 2018, 41, 42–52. [Google Scholar] [CrossRef] [PubMed]
- Yakymovych, I.; Yakymovych, M.; Heldin, C.H. Intracellular trafficking of transforming growth factor β receptors. Acta Biochim. Et Biophys. Sin. 2018, 50, 3–11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sanese, P.; Forte, G.; Disciglio, V.; Grossi, V.; Simone, C. FOXO3 on the Road to Longevity: Lessons From SNPs and Chromatin Hubs. Comput. Struct. Biotechnol. J. 2019, 17, 737–745. [Google Scholar] [CrossRef] [PubMed]
- Morris, B.J.; Willcox, D.C.; Donlon, T.A.; Willcox, B.J. FOXO3: A Major Gene for Human Longevity—A Mini-Review. Gerontology 2015, 61, 515–525. [Google Scholar] [CrossRef] [Green Version]
- Ding, Y.; Kim, S.; Lee, S.Y.; Koo, J.K.; Wang, Z.; Choi, M.E. Autophagy regulates TGF-β expression and suppresses kidney fibrosis induced by unilateral ureteral obstruction. J. Am. Soc. Nephrol. JASN 2014, 25, 2835–2846. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, Y.A.; Kim, H.J.; Gwon, M.G.; Gu, H.; An, H.J.; Bae, S.; Leem, J.; Jung, H.J.; Park, K.K. Inhibitory Effects of STAT3 Transcription Factor by Synthetic Decoy ODNs on Autophagy in Renal Fibrosis. Biomedicines 2021, 9, 331. [Google Scholar] [CrossRef]
- Chung, S.; Son, M.; Chae, Y.; Oh, S.; Koh, E.S.; Kim, Y.K.; Shin, S.J.; Park, C.W.; Jung, S.C.; Kim, H.S. Fabry disease exacerbates renal interstitial fibrosis after unilateral ureteral obstruction via impaired autophagy and enhanced apoptosis. Kidney Res. Clin. Pract. 2021, 40, 208–219. [Google Scholar] [CrossRef]
- Hartleben, B.; Gödel, M.; Meyer-Schwesinger, C.; Liu, S.; Ulrich, T.; Köbler, S.; Wiech, T.; Grahammer, F.; Arnold, S.J.; Lindenmeyer, M.T.; et al. Autophagy influences glomerular disease susceptibility and maintains podocyte homeostasis in aging mice. J. Clin. Investig. 2010, 120, 1084–1096. [Google Scholar] [CrossRef] [Green Version]
- Yu, H.X.; Lin, W.; Yang, K.; Wei, L.J.; Chen, J.L.; Liu, X.Y.; Zhong, K.; Chen, X.; Pei, M.; Yang, H.T. Transcriptome-Based Network Analysis Reveals Hirudin Potentiates Anti-Renal Fibrosis Efficacy in UUO Rats. Front. Pharmacol. 2021, 12, 741801. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Jin, L.; Zheng, D.; Tang, X.; Yang, J.; Fan, L.; Xie, X. Fucoxanthin Alleviates Oxidative Stress through Akt/Sirt1/FoxO3α Signaling to Inhibit HG-Induced Renal Fibrosis in GMCs. Mar. Drugs 2019, 17, 702. [Google Scholar] [CrossRef] [PubMed]
- Möllsten, A.; Jorsal, A.; Lajer, M.; Vionnet, N.; Tarnow, L. The V16A polymorphism in SOD2 is associated with increased risk of diabetic nephropathy and cardiovascular disease in type 1 diabetes. Diabetologia 2009, 52, 2590–2593. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prunotto, M.; Carnevali, M.L.; Candiano, G.; Murtas, C.; Bruschi, M.; Corradini, E.; Trivelli, A.; Magnasco, A.; Petretto, A.; Santucci, L.; et al. Autoimmunity in membranous nephropathy targets aldose reductase and SOD2. J. Am. Soc. Nephrol. JASN 2010, 21, 507–519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olsson, J.; Jacobson, T.A.; Paulsson, J.M.; Dadfar, E.; Moshfegh, A.; Jacobson, S.H.; Lundahl, J. Expression of neutrophil SOD2 is reduced after lipopolysaccharide stimulation: A potential cause of neutrophil dysfunction in chronic kidney disease. Nephrol. Dial. Transplant. Off. Publ. Eur. Dial. Transpl. Assoc.—Eur. Ren. Assoc. 2011, 26, 2195–2201. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zaza, G.; Granata, S.; Masola, V.; Rugiu, C.; Fantin, F.; Gesualdo, L.; Schena, F.P.; Lupo, A. Downregulation of nuclear-encoded genes of oxidative metabolism in dialyzed chronic kidney disease patients. PLoS ONE 2013, 8, e77847. [Google Scholar] [CrossRef] [Green Version]
- Mohammedi, K.; Bellili-Muñoz, N.; Driss, F.; Roussel, R.; Seta, N.; Fumeron, F.; Hadjadj, S.; Marre, M.; Velho, G. Manganese superoxide dismutase (SOD2) polymorphisms, plasma advanced oxidation protein products (AOPP) concentration and risk of kidney complications in subjects with type 1 diabetes. PLoS ONE 2014, 9, e96916. [Google Scholar] [CrossRef] [Green Version]
- Krueger, K.; Shen, J.; Maier, A.; Tepel, M.; Scholze, A. Lower Superoxide Dismutase 2 (SOD2) Protein Content in Mononuclear Cells Is Associated with Better Survival in Patients with Hemodialysis Therapy. Oxidative Med. Cell. Longev. 2016, 2016, 7423249. [Google Scholar] [CrossRef] [Green Version]
- Jerotic, D.; Matic, M.; Suvakov, S.; Vucicevic, K.; Damjanovic, T.; Savic-Radojevic, A.; Pljesa-Ercegovac, M.; Coric, V.; Stefa novic, A.; Ivanisevic, J.; et al. Association of Nrf2, SOD2 and GPX1 Polymorphisms with Biomarkers of Oxidative Distress and Survival in End-Stage Renal Disease Patients. Toxins 2019, 11, 431. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Corredor, Z.; Filho, M.; Rodríguez-Ribera, L.; Velázquez, A.; Hernández, A.; Catalano, C.; Hemminki, K.; Coll, E.; Silva, I.; Diaz, J.M.; et al. Genetic Variants Associated with Chronic Kidney Disease in a Spanish Population. Sci. Rep. 2020, 10, 144. [Google Scholar] [CrossRef] [Green Version]
- Miyazaki, N.; Murata, I.; Takemura, G.; Okada, H.; Kanamori, H.; Matsumoto-Miyazaki, J.; Yoshida, G.; Izumi, K.; Kashi, H.; Niimi, K.; et al. Expression of prorenin receptor in renal biopsies from patients with IgA nephropathy. Int. J. Clin. Exp. Pathol. 2014, 7, 7485–7496. [Google Scholar]
- Xiong, G.; Tang, W.; Zhang, D.; He, D.; Wei, G.; Atala, A.; Liang, X.J.; Bleyer, A.J.; Bleyer, M.E.; Yu, J.; et al. Impaired Regen eration Potential in Urinary Stem Cells Diagnosed from the Patients with Diabetic Nephropathy. Theranostics 2019, 9, 4221–4232. [Google Scholar] [CrossRef]
- Liu, W.J.; Gan, Y.; Huang, W.F.; Wu, H.L.; Zhang, X.Q.; Zheng, H.J.; Liu, H.F. Lysosome restoration to activate podocyte autophagy: A new therapeutic strategy for diabetic kidney disease. Cell Death Dis. 2019, 10, 806. [Google Scholar] [CrossRef] [PubMed]
- Ogawa-Akiyama, A.; Sugiyama, H.; Kitagawa, M.; Tanaka, K.; Kano, Y.; Mise, K.; Otaka, N.; Tanabe, K.; Morinaga, H.; Ki nomura, M.; et al. Podocyte autophagy is associated with foot process effacement and proteinuria in patients with minimal change nephrotic syndrome. PLoS ONE 2020, 15, e0228337. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- da Silva, C.A.; Monteiro, M.; Araújo, L.S.; Urzedo, M.G.; Rocha, L.B.; Dos Reis, M.A.; Machado, J.R. In situ evaluation of podocytes in patients with focal segmental glomerulosclerosis and minimal change disease. PLoS ONE 2020, 15, e0241745. [Google Scholar] [CrossRef]
- Yanai, K.; Kaneko, S.; Ishii, H.; Aomatsu, A.; Ito, K.; Hirai, K.; Ookawara, S.; Ishibashi, K.; Morishita, Y. Quantitative Real-Time PCR Evaluation of microRNA Expressions in Mouse Kidney with Unilateral Ureteral Obstruction. J. Vis. Exp. 2020, 27. [Google Scholar] [CrossRef] [PubMed]
- Chevalier, R.L.; Forbes, M.S.; Thornhill, B.A. Ureteral obstruction as a model of renal interstitial fibrosis and obstructive nephropathy. Kidney Int. 2009, 75, 1145–1152. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martínez-Klimova, E.; Aparicio-Trejo, O.E.; Tapia, E.; Pedraza-Chaverri, J. Unilateral Ureteral Obstruction as a Model to Investigate Fibrosis-Attenuating Treatments. Biomolecules 2019, 9, 141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morishita, Y.; Yoshizawa, H.; Watanabe, M.; Imai, R.; Imai, T.; Hirahara, I.; Akimoto, T.; Ookawara, S.; Muto, S.; Nagata, D. MicroRNA expression profiling in peritoneal fibrosis. Transl. Res. J. Lab. Clin. Med. 2016, 169, 47–66. [Google Scholar] [CrossRef]
- Godbey, W.T.; Wu, K.K.; Mikos, A.G. Tracking the intracellular path of poly(ethylenimine)/DNA complexes for gene delivery. Proc. Natl. Acad. Sci. USA 1999, 96, 5177–5181. [Google Scholar] [CrossRef] [Green Version]
- Boussif, O.; Lezoualc’h, F.; Zanta, M.A.; Mergny, M.D.; Scherman, D.; Demeneix, B.; Behr, J.P. A versatile vector for gene and oligonucleotide transfer into cells in culture and in vivo: Polyethylenimine. Proc. Natl. Acad. Sci. USA 1995, 92, 7297–7301. [Google Scholar] [CrossRef] [Green Version]
- Akinc, A.; Thomas, M.; Klibanov, A.M.; Langer, R. Exploring polyethylenimine-mediated DNA transfection and the proton sponge hypothesis. J. Gene Med. 2005, 7, 657–663. [Google Scholar] [CrossRef]
- Zhao, Q.Q.; Chen, J.L.; Lv, T.F.; He, C.X.; Tang, G.P.; Liang, W.Q.; Tabata, Y.; Gao, J.Q. N/P ratio significantly influences the transfection efficiency and cytotoxicity of a polyethylenimine/chitosan/DNA complex. Biol. Pharm. Bull. 2009, 32, 706–710. [Google Scholar] [CrossRef] [PubMed]
- Hao, F.; Li, Y.; Zhu, J.; Sun, J.; Marshall, B.; Lee, R.J.; Teng, L.; Yang, Z.; Xie, J. Polyethylenimine-based Formulations for Delivery of Oligonucleotides. Curr. Med. Chem. 2019, 26, 2264–2284. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Hartley, A.V.; Yin, Y.; Herskowitz, J.H.; Lah, J.J.; Ressler, K.J. AAV2 production with optimized N/P ratio and PEI-mediated transfection results in low toxicity and high titer for in vitro and in vivo applications. J. Virol. Methods 2013, 193, 270–277. [Google Scholar] [CrossRef] [Green Version]
- Griffiths-Jones, S.; Saini, H.K.; van Dongen, S.; Enright, A.J. miRBase: Tools for microRNA genomics. Nucleic Acids Res. 2008, 36, D154–D158. [Google Scholar] [CrossRef] [PubMed]
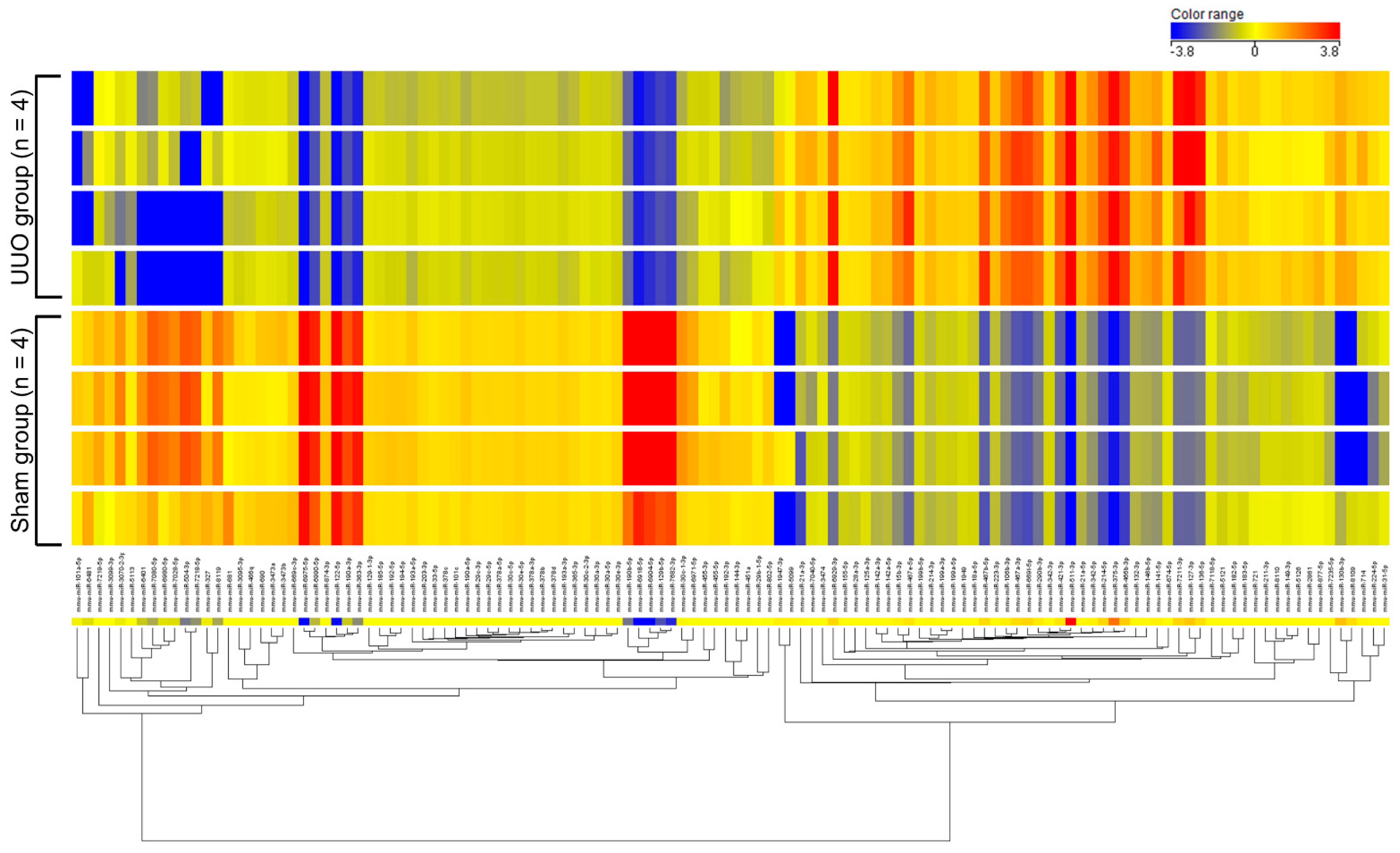

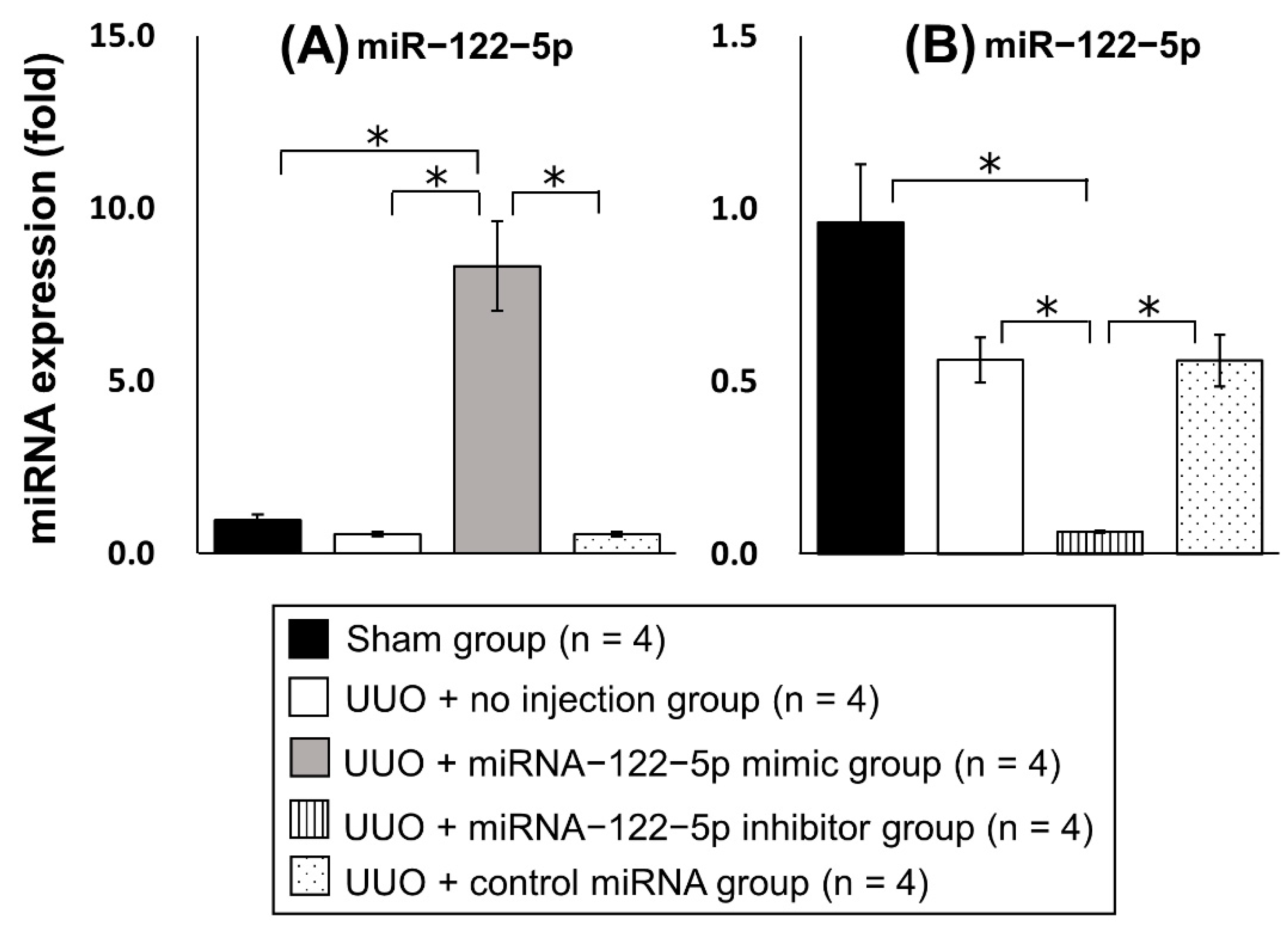
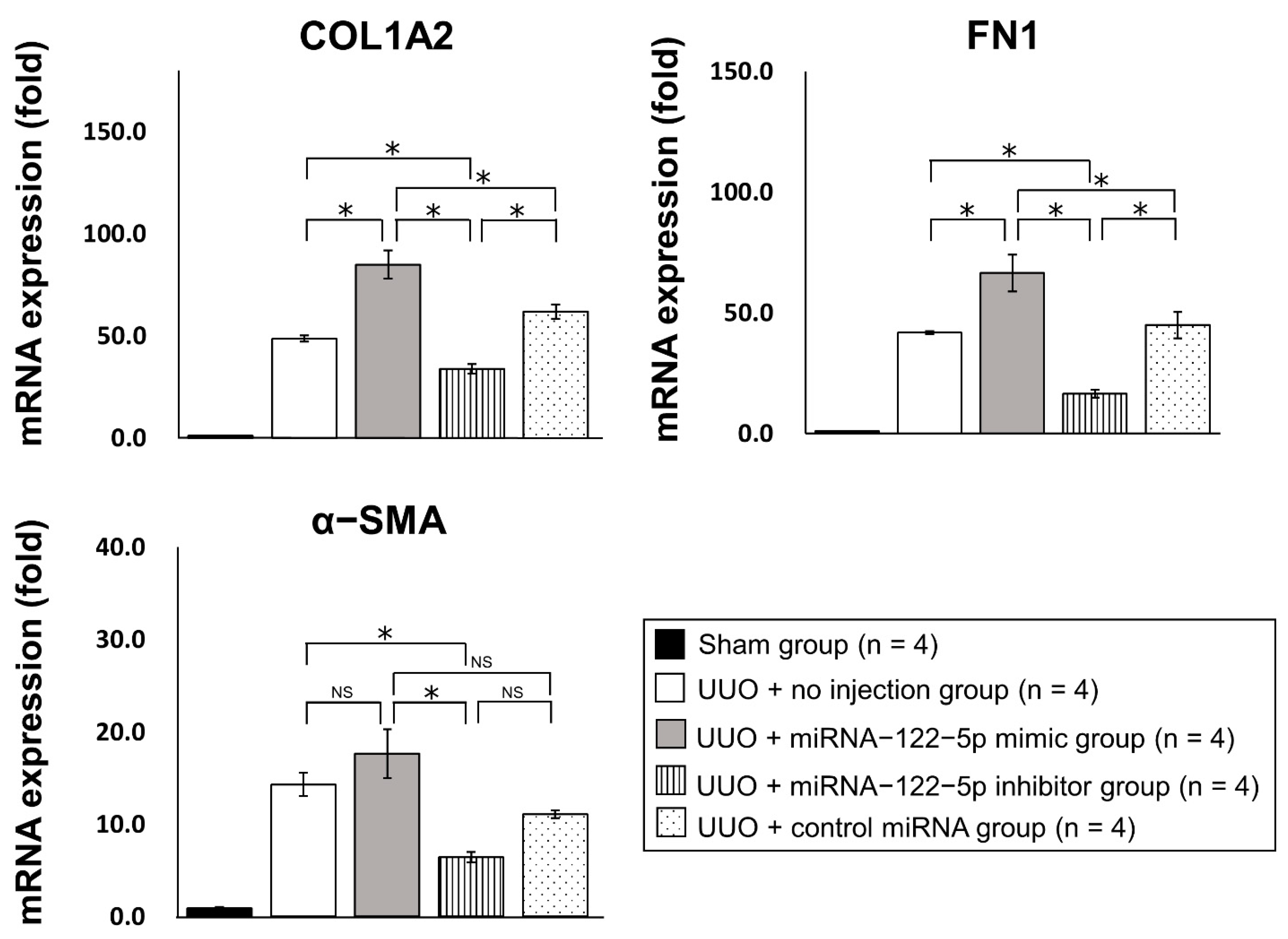
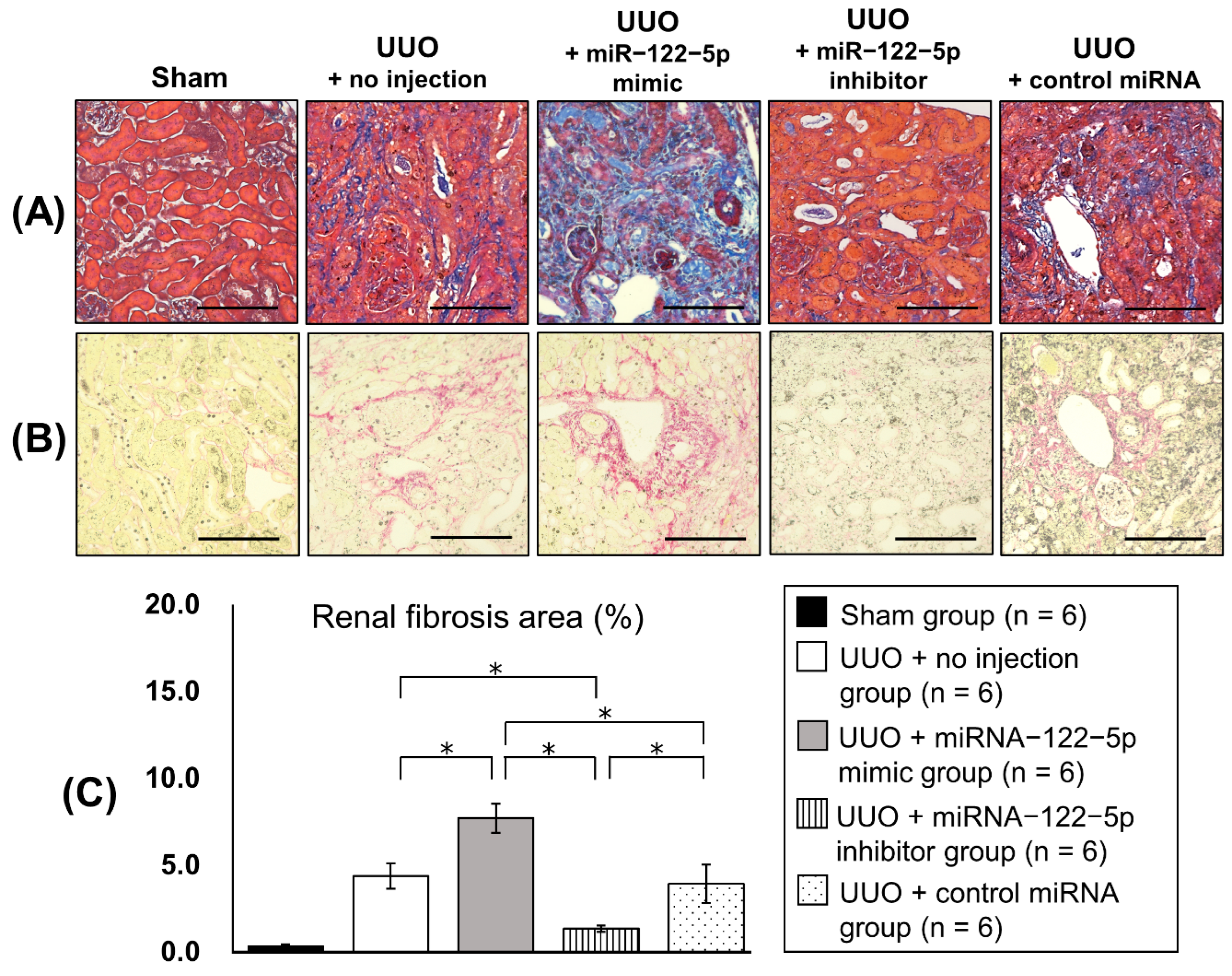
| microRNA | Sequences | Fold Change | p |
|---|---|---|---|
| (UUO Mice/Control Mice) | |||
| miR−511−3p | AAUGUGUAGCAAAAGACAGGAU | 312.3 | <0.01 |
| miR−375−3p | UUUGUUCGUUCGGCUCGCGUGA | 143.0 | <0.01 |
| miR−130b−3p | CAGUGCAAUGAUGAAAGGGCAU | 61.8 | <0.01 |
| miR−127−3p | UCGGAUCCGUCUGAGCUUGGCU | 55.7 | <0.01 |
| miR−466b−3p | AUACAUACACGCACACAUAAGA | 48.8 | <0.01 |
| miR−7211−3p | UGGAGUGACUGUAGGGAGGAUGC | 41.7 | <0.01 |
| miR−669l−5p | AGUUGUGUGUGCAUGUAUAUGU | 40.6 | <0.01 |
| miR−214−5p | UGCCUGUCUACACUUGCUGUGC | 39.1 | <0.01 |
| miR−6929−3p | AGGGAGGAGCAGCAUCUGUGA | 38.9 | <0.01 |
| miR−467b−5p | GUAAGUGCCUGCAUGUAUAUG | 35.5 | <0.01 |
| miR−8109 | CCCGCGGCCG | 33.4 | <0.01 |
| miR−467a−3p | TGTAGGTGTGTGTATGTATA | 33.3 | <0.01 |
| miR−136−5p | CCATCATCAAAACAAATGGAGT | 30.9 | <0.01 |
| miR−421−3p | GCGCCCAATTAATGTCTG | 26.0 | <0.01 |
| miR−467e−5p | ACATATACATGCTCACACT | 25.6 | <0.01 |
| miR−290b−3p | TACTCAAACTATGGGGGC | 19.0 | <0.01 |
| miR−1907 | ACCTCCAGATCCTCTG | 17.7 | <0.01 |
| miR−1967 | GCATCTTCTCCCCAG | 17.7 | <0.01 |
| miR−7687−5p | AGCCTGCGCCTCA | 17.4 | <0.01 |
| miR−106b−3p | GCAGCAAGTACCCAC | 15.1 | <0.01 |
| miR−379−5p | CCTACGTTCCATAGTC | 14.5 | <0.01 |
| miR−3547−5p | TCCCGGGCCCC | 14.1 | <0.01 |
| miR−1947−3p | GGAGGGAGAGCTAGC | 13.6 | <0.01 |
| miR−3091−5p | GCGGGCCCAACC | 13.5 | <0.01 |
| miR−134−5p | CCCCTCTGGTCAA | 13.2 | <0.01 |
| miR−31−3p | GATGGCAATATGTTGGCAT | 13.2 | <0.01 |
| miR−5099 | GGAGCACCACATCG | 11.4 | <0.01 |
| miR−141−5p | TCCAACACTGCACTGGA | 10.9 | <0.01 |
| miR−15b−3p | TAGAGCAGCAAATAATGATTCG | 10.9 | <0.01 |
| miR−300−3p | GAAGAGAGCTTGCCCTTG | 10.8 | <0.01 |
| miR−342−5p | CTCAATCACAGATAGCACC | 10.4 | <0.01 |
| miR−8095 | GGGACAGACAGCAGA | 10.2 | <0.01 |
| miR−6984−3p | AGAAAGACAGGAAAGAAAG | 10.0 | <0.01 |
| miR−714 | GCGACCGACCGGCC | 9.6 | <0.01 |
| miR−3472 | GGTTCCTTCCAGCTT | 9.5 | <0.01 |
| miR−1957a | GTCATATGCTCTACCACT | 9.4 | <0.01 |
| miR−5131 | GCTCGGGGCTCC | 8.8 | <0.01 |
| miR−7047−3p | GGAAGGAGGAAGGGT | 8.6 | <0.01 |
| miR−3473c | CTTATTATGGGGGCTGG | 8.1 | <0.01 |
| miR−21a−3p | GACAGCCCATCGACT | 7.8 | <0.01 |
| miR−6392−3p | AGAGGACCCGGCA | 7.2 | <0.01 |
| miR−710 | CTCAACTCTCCCCA | 7.0 | <0.01 |
| miR−7046−5p | CCTGGCTCCCAGC | 6.9 | <0.01 |
| miR−146b−5p | AGCCTATGGAATTCAGTTC | 6.4 | <0.01 |
| miR−770−3p | CCAGCTCCACGTC | 6.0 | <0.01 |
| miR−6961−5p | GGCCATTTCCTTCCC | 5.6 | <0.01 |
| miR−376a−3p | ACGTGGATTTTCCTCTA | 5.5 | <0.01 |
| miR−6354 | AGAGACCTGATCCCCA | 5.3 | <0.01 |
| miR−1931 | GCCATCGCACCAGC | 5.2 | <0.01 |
| miR−501−3p | CAAATCCTTGCCCGG | 4.9 | <0.01 |
| miR−21a−5p | TCAACATCAGTCTGATAAGC | 4.9 | <0.01 |
| miR−132−3p | CGACCATGGCTGTAGA | 4.9 | <0.01 |
| miR−468−3p | CAGACACACGCACATCA | 4.7 | <0.01 |
| miR−214−3p | ACTGCCTGTCTGT | 4.4 | <0.01 |
| miR−7672−5p | TCGCCCGCTGTCA | 4.4 | <0.01 |
| miR−292b−3p | ATACTCAAACTGGGGGC | 4.0 | <0.01 |
| miR−7063−5p | TGTGCTCAGCCTGC | 4.0 | <0.01 |
| miR−5132−5p | CCTGAGTCCACCACC | 4.0 | <0.01 |
| miR−3093−3p | CCAACCTCCCACGG | 4.0 | <0.01 |
| miR−142a−3p | TCCATAAAGTAGGAAACACTACA | 4.0 | <0.01 |
| miR−5121 | GGAGATGTCTCATCACA | 4.0 | <0.01 |
| miR−223−3p | TGGGGTATTTGACAAACTGAC | 3.9 | <0.01 |
| miR−224−5p | AACGGAACCACTAGTGACTTA | 3.9 | <0.01 |
| miR−3572−5p | GTCCACCTTGCCCT | 3.8 | <0.01 |
| miR−6963−5p | CCAGGTTCTGCCATC | 3.8 | <0.01 |
| miR−6921−5p | GCTTCCTACCTCATGC | 3.7 | <0.01 |
| miR−467a−5p | CGCATATACATGCAGGCA | 3.5 | <0.01 |
| miR−6349 | CGCATGCCCCTCC | 3.5 | <0.01 |
| miR−669b−5p | ACATGCACATGCACACA | 3.5 | <0.01 |
| miR−299b−5p | ATGTATGTGGGACGGTAAAC | 3.5 | <0.01 |
| miR−7080−3p | AGGGAACGGAGGGG | 3.5 | <0.01 |
| miR−142a−5p | AGTAGTGCTTTCTACTTTA | 3.4 | <0.01 |
| miR−199a−5p | GAACAGGTAGTCTGAACAC | 3.2 | <0.01 |
| miR−8093 | CACTCATGCTCTGCTC | 3.1 | <0.01 |
| miR−183−5p | AGTGAATTCTACCAGTGCC | 3.1 | <0.01 |
| miR−7040−5p | CGCCTCCATCTCCC | 3.0 | <0.01 |
| miR−718 | CGACACCCGGCCG | 3.0 | <0.01 |
| miR−7235−5p | GCCCAGACCCCTC | 3.0 | <0.01 |
| miR−762 | GCTCTGTCCCGGC | 3.0 | <0.01 |
| miR−199a−3p | TAACCAATGTGCAGACTACT | 3.0 | <0.01 |
| miR−672−5p | TCACACACAGTACACCA | 2.9 | <0.01 |
| miR−18a−5p | CTATCTGCACTAGATGCAC | 2.8 | <0.01 |
| miR−3067−3p | CCTCTCCCAGGGC | 2.8 | <0.01 |
| miR−292a−5p | CAAAAGAGCCCCCAG | 2.7 | <0.01 |
| miR−342−3p | ACGGGTGCGATTTCTGT | 2.7 | <0.01 |
| miR−20a−3p | CTTTAAGTGCTCGTAATGCA | 2.7 | <0.01 |
| miR−let−7i−3p | AGCAAGGCAGTAGCTT | 2.6 | <0.01 |
| miR−290a−3p | GGGCTTAAAACTAGGCGGC | 2.6 | <0.01 |
| miR−182−5p | CGGTGTGAGTTCTACC | 2.5 | <0.01 |
| miR−149−3p | GCACCGCCCCC | 2.5 | <0.01 |
| miR−877−5p | CCCTGCGCCATCT | 2.5 | <0.01 |
| miR−1949 | AACTATGCTGACATCCTG | 2.5 | <0.01 |
| miR−674−5p | TACACCACTCCCAT | 2.5 | <0.01 |
| miR−199b−5p | GAACAGGTAGTCTAAACACTGG | 2.5 | <0.01 |
| miR−31−5p | CAGCTATGCCAGCATCT | 2.4 | <0.01 |
| miR−125a−3p | GGCTCCCAAGAACCTC | 2.4 | <0.01 |
| miR−721 | TTCCCCCTTTTAATT | 2.4 | <0.01 |
| miR−7115−3p | CTGTGGGGGCAGG | 2.3 | <0.01 |
| miR−3474 | GAATCCACGTCTCCTC | 2.3 | <0.01 |
| miR−28a−3p | TCCAGCAGCTCACA | 2.2 | <0.01 |
| miR−3075−5p | GTCCTTGGCTGCTC | 2.2 | <0.01 |
| miR−5126 | CCCCGCCCCCG | 2.2 | <0.01 |
| miR−155−5p | ACCCCTATCACAATTAGC | 2.1 | <0.01 |
| miR−2861 | CCGCCCGCCG | 2.1 | <0.01 |
| miR−5128 | AGCCATCTCGCCAGC | 2.1 | <0.01 |
| miR−135a−1−3p | CGCCACGGCTCCA | 2.1 | <0.01 |
| miR−8110 | CCCCCCCCCCA | 2.1 | <0.01 |
| miR−211−3p | GCCCCCCTTTGCT | 2.0 | <0.01 |
| miR−7118−5p | GTTCCCTCTCCCGC | 2.0 | <0.01 |
| miR−6975−5p | GCUGGGGAGAAAGGGGUUUGGCA | −253.2 | <0.01 |
| miR−6918−5p | UGCUGAGGACGGGAUUAGGUUCU | −252.2 | <0.01 |
| miR−6904−5p | UCCUGGGGUUAGAGUUGAGUGG | −194.5 | <0.01 |
| miR−122−5p | UGGAGUGUGACAAUGGUGUUUG | −183.2 | <0.01 |
| miR−7682−3p | CCUGUGGGUUGGGUUGGCUUU | −164.8 | <0.01 |
| miR−129b−5p | GCUUUUUGGGGUAAGGGCUUCC | −134.9 | <0.01 |
| miR−190b−5p | UGAUAUGUUUGAUAUUGGGUUG | −107.9 | <0.01 |
| miR−504−3p | AGGGAGAGCAGGGCAGGGUUUC | −104.0 | <0.01 |
| miR−363−3p | AAUUGCACGGUAUCCAUCUGUA | −89.6 | <0.01 |
| miR−7218−5p | UGCAGGGUUUAGUGUAGAGGG | −85.3 | <0.01 |
| miR−8119 | GACCCTAGCTCCCTC | −70.6 | <0.01 |
| miR−7080−5p | CCAAACCCACCTCC | −67.1 | <0.01 |
| miR−1927 | TCAGTCCCTAACATCCA | −57.8 | <0.01 |
| miR−6990−5p | AGAGCCCTGACTCACC | −56.3 | <0.01 |
| miR−7234−3p | CCTTCTACCCTAGAAAGA | −51.7 | <0.01 |
| miR−6905−5p | TCATTCAACCCAACCTG | −49.6 | <0.01 |
| miR−7055−5p | CCAACTCAGATAACCCA | −46.6 | <0.01 |
| miR−190a−3p | AGGAATATGCTTGATATATAGT | −42.8 | <0.01 |
| miR−6401 | ACCCGACACCACTG | −42.2 | <0.01 |
| miR−7233−5p | CATCTATCTGTCCCTAACT | −41.4 | <0.01 |
| miR−7657−5p | TTACCTAACTATCCAACTATT | −36.4 | <0.01 |
| miR−3078−3p | CCTAAAGACTACCCCAG | −35.4 | <0.01 |
| miR−7664−3p | ATTAGTTAACCCAGCCTAA | −33.4 | <0.01 |
| miR−7086−5p | TGCCCAAACCTTTCTC | −33.1 | <0.01 |
| miR−6902−3p | CTGAACCCACACATCA | −32.7 | <0.01 |
| miR−188−3p | TGCAAACCCTGCATGTG | −31.2 | <0.01 |
| miR−7028−5p | CTCCTGACCCAAGC | −27.7 | <0.01 |
| miR−6481 | CATCTAAGCATTTTCAGTG | −25.8 | <0.01 |
| miR−302c−5p | GCAGGTAACCCCAT | −24.6 | <0.01 |
| miR−6980−5p | CTAACCTAGCCTCCCC | −24.2 | <0.01 |
| miR−467b−3p | GTGTTGGTGTGTGTAT | −23.8 | <0.01 |
| miR−7219−3p | AGTGTGTTAGAAACCCG | −21.1 | <0.01 |
| miR−194−2−3p | CAGATAACAGCAGCCC | −19.6 | <0.01 |
| miR−7074−5p | ACTGGAGCCCTAGCC | −19.0 | <0.01 |
| miR−6998−5p | AGTCACTTTGCCCTCT | −18.2 | <0.01 |
| miR−874−5p | CTTACCCTGGTGCG | −17.4 | <0.01 |
| miR−6923−5p | ACACCCCAATCCTCC | −16.4 | <0.01 |
| miR−327 | ATCCTCATGCCCCT | −15.7 | <0.01 |
| miR−1188−3p | GCAGGGTGTGGTGG | −15.7 | <0.01 |
| miR−3070−2−3p | TCTACCCCTGACCATAG | −14.9 | <0.01 |
| miR−101a−5p | GCATCAGCACTGTGAT | −10.9 | <0.01 |
| miR−3070−3p | TCTACCCCTGACGGT | −10.5 | <0.01 |
| miR−6926−5p | TCACCATCCCTCACC | −8.8 | <0.01 |
| miR−6981−5p | GCCTTCAGCCTCTTC | −8.6 | <0.01 |
| miR−3063−3p | GGCGAGAGATCAGGA | −8.6 | <0.01 |
| miR−6236 | CCTGACTGCCGGC | −7.2 | <0.01 |
| miR−30c−1−3p | GGAGTAAACAACCCTCTCC | −6.0 | <0.01 |
| miR−883b−5p | TGACTGCTACCCATT | −5.6 | <0.01 |
| miR−26b−3p | GAGCCAAGTAATGGAGAACA | −5.3 | <0.01 |
| miR−744−3p | AGGTTGAGGTTAGTGGCA | −5.1 | <0.01 |
| miR−6988−5p | TGGGCCTCAGCTCT | −4.9 | <0.01 |
| miR−376b−3p | AAGTGGATGTTCCTCTAT | −4.3 | <0.01 |
| miR−6393 | ACTCAGTGTGCTTCGT | −4.1 | <0.01 |
| miR−7094b−2−5p | TCAGACCCTGTATCCTC | −3.9 | <0.01 |
| miR−7056−5p | AACCTCTCTGTCCTCC | −3.9 | <0.01 |
| miR−6971−5p | AGCCTCTACACCCTCC | −3.9 | <0.01 |
| miR−497b | CCACGTCCAAACCA | −3.9 | <0.01 |
| miR−874−3p | TCGGTCCCTCGGG | −3.6 | <0.01 |
| miR−192−3p | CTGTGACCTATGGAATTG | −3.5 | <0.01 |
| miR−873b | GTGTGCATTTGCAGGA | −3.5 | <0.01 |
| miR−7075−5p | AAAACCATGTCCTCCTC | −3.4 | <0.01 |
| miR−703 | TTCTTTCCTTCTGAAGGTT | −3.3 | <0.01 |
| miR−193a−5p | TCATCTTGCCCGCA | −3.3 | <0.01 |
| miR−7670−5p | TTCCCAATCTGCCCA | −3.2 | <0.01 |
| miR−8094 | TCTTCTCGTTGTCCTTC | −3.2 | <0.01 |
| miR−144−5p | ACTTACAGTATATGATGATATCC | −3.2 | <0.01 |
| miR−669c−3p | TTTACTTGTGTGTGTGTG | −3.0 | <0.01 |
| miR−193a−3p | ACTGGGACTTTGTAGGC | −3.0 | <0.01 |
| miR−190a−5p | ACCTAATATATCAAACATATCA | −3.0 | <0.01 |
| miR−30e−5p | CTTCCAGTCAAGGATGT | −2.9 | <0.01 |
| miR−302c−3p | CCACTGAAACATGGAAGCAC | −2.8 | <0.01 |
| miR−192−5p | GGCTGTCAATTCATAGGTC | −2.7 | <0.01 |
| miR−30e−3p | GCTGTAAACATCCGACTG | −2.7 | <0.01 |
| miR−681 | AGCTGCCTGCCAG | −2.6 | <0.01 |
| miR−376c−3p | ACGTGAAATTTCCTCTATGTT | −2.6 | <0.01 |
| miR−466g | TGTGTGTGCATGTGTC | −2.6 | <0.01 |
| miR−30c−5p | GCTGAGAGTGTAGGATGT | −2.6 | <0.01 |
| miR−194−5p | TCCACATGGAGTTGCT | −2.6 | <0.01 |
| miR−802−5p | AAGGATGAATCTTTGTTACTGA | −2.6 | <0.01 |
| miR−365−3p | ATAAGGATTTTTAGGGGCATTA | −2.5 | <0.01 |
| miR−378a−5p | ACACAGGACCTGGAGTCA | −2.5 | <0.01 |
| miR−7211−5p | GGTGGAGTGGCAGA | −2.5 | <0.01 |
| miR−5113 | ACAGGATCTCTCTCCTC | −2.5 | <0.01 |
| miR−378b | TCTTCTGACTCCAAGTC | −2.5 | <0.01 |
| miR−29c−5p | GAACACCAGGAGAAATCGGTC | −2.4 | <0.01 |
| miR−378d | ACCTTCTGACTCCAAGG | −2.4 | <0.01 |
| miR−185−5p | TCAGGAACTGCCTTTCT | −2.4 | <0.01 |
| miR−378a−3p | CCTTCTGACTCCAA | −2.4 | <0.01 |
| miR−7219−5p | TCTCAACCCTGAGCTC | −2.3 | <0.01 |
| miR−29c−3p | TAACCGATTTCAAATGGTGCTA | −2.3 | <0.01 |
| miR−378c | GCTTCTGACTCCAAGT | −2.3 | <0.01 |
| miR−3473b | CTGAGCCATCTCTCCA | −2.3 | <0.01 |
| miR−30a−3p | GCTGCAAACATCCGACT | −2.3 | <0.01 |
| miR−let−7f−1−3p | GGGAAGGCAATAGATTGTAT | −2.3 | <0.01 |
| miR−203−3p | CTAGTGGTCCTAAACATT | −2.3 | <0.01 |
| miR−101c | TCAGTTATCACAGTACTGT | −2.2 | <0.01 |
| miR−129−1−3p | ATACTTTTTGGGGTAAGGG | −2.2 | <0.01 |
| miR−144−3p | AGTACATCATCTATACTGTA | −2.2 | <0.01 |
| miR−455−5p | CGATGTAGTCCAAAGGCA | −2.2 | <0.01 |
| miR−451a | AACTCAGTAATGGTAACGGTTT | −2.2 | <0.01 |
| miR−3095−3p | AAAAGCTCTCTCTCCAGT | −2.2 | <0.01 |
| miR−690 | TTTGGTTGTGAGCCTA | −2.2 | <0.01 |
| miR−455−3p | GTGTATATGCCCGTGG | −2.1 | <0.01 |
| miR−466q | ACGTATGTGTGTGTGTG | −2.1 | <0.01 |
| miR−3099−3p | TCCCCAACCTCTCTC | −2.1 | <0.01 |
| miR−760−3p | TCCCCACAGACCCA | −2.1 | <0.01 |
| miR−3473a | TGCTGAGCCATCTCTC | −2.1 | <0.01 |
| miR−30a−5p | CTTCCAGTCGAGGATGT | −2.1 | <0.01 |
| miR−29b−1−5p | TAAACCACCATATGAAACCAGC | −2.1 | <0.01 |
| miR−664−5p | CCAGTCATTTTCCCCA | −2.0 | <0.01 |
| miR−30c−2−3p | AGAGTAAACAGCCTTCTCC | −2.0 | <0.01 |
| miR−33−5p | TGCAATGCAACTACAATGCAC | −2.0 | <0.01 |
| miR−295−5p | GAAGTGTGCCCCAC | −2.0 | <0.01 |
| Target | Authors | Country | Year | Patients and Number | Results and Findings | References |
|---|---|---|---|---|---|---|
| SOD2 | Möllsten et al. | Sweden | 2009 | Type I DM patients (n = 411) | Genetic SOD2 polymorphisms were associated with the development of DM nephropathy caused by type Ⅰ DM. | [51] |
| SOD2 | Prunotto et al. | Italy | 2010 | MN patient (n = 24) | Anti-SOD2 antibodies were specifically detected in the serum of primary MN patients. Anti-SOD2 antibodies were deposited in the glomerular podocytes of MN patients. | [52] |
| SOD2 | Olsson et al. | Sweden | 2011 | CKD patients (n = 30) | Neutrophils from CKD patients were stimulated with lipopolysaccharide. SOD2 gene expression was decreased in the neutrophils of CKD patients but not controls. | [53] |
| SOD2 | Zaza et al. | Italy | 2013 | PD patients (n = 15) | Plasma malondialdehyde (an oxidative marker) levels were higher in PD patients compared with controls, leading to increased SOD2 gene expression. | [54] |
| SOD2 | Mohammedi et al | France | 2014 | Type Ⅰ DM patients (n = 1285) | SOD2 gene mutations were associated with the onset and progression of DM nephropathy, plasma advanced oxidation protein product (an oxidative marker) concentrations, and antioxidant activity in type Ⅰ DM patients. | [55] |
| SOD2 | Krueger et al. | Denmark Germany | 2016 | CKD patients (n = 120) HD patients (n = 81) | SOD2 protein levels in monocytes decreased as the CKD stage progressed. After HD induction, SOD2 protein levels began to increase. | [56] |
| SOD2 | Jerotic et al. | Serbia. | 2019 | HD patients (n = 256) | Genetic SOD2 polymorphisms were associated with risk of end-stage renal disease. | [57] |
| SOD2 | Corredor et al. | Spain | 2020 | CKD patients (n = 548) | Genetic SOD2 polymorphisms were associated with the erythropoietin resistance of renal anemia. | [58] |
| LC3 | Miyazaki et al. | Japan | 2014 | IgAN patients (n = 48) | In IgAN patients, prorenin receptor expression was an autophagy (LC3)-mediated compensatory response to IgAN progression. | [59] |
| LC3 | Xiong et al. | USA | 2019 | DM nephropathy patients (n = 12) | LC3 protein obtained from urinary stem cells in DM nephropathy patients was decreased compared with the control group. | [60] |
| LC3 | Liu et al. | China | 2019 | DM nephropathy patients (n = 11) | Podocytes of DM nephropathy patients had more LC3 positive puncta compared with the control group. | [61] |
| LC3 | Ogawa-Akiyama et al. | Japan | 2020 | MCNS patients (n = 41) MN patients (n = 37) | LC3 was localized to glomerular podocytes, suggesting autophagy mainly occurred in the glomerular podocytes of MCNS patients. | [62] |
| LC3 | da Silva et al. | Brazil | 2020 | FSGS patients (n = 22) MCNS patients (n = 27) | LC3-positive glomerular podocytes were denser in MCNS patients than in FSGS patients. | [63] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaneko, S.; Yanai, K.; Ishii, H.; Aomatsu, A.; Hirai, K.; Ookawara, S.; Ishibashi, K.; Morishita, Y. miR−122−5p Regulates Renal Fibrosis In Vivo. Int. J. Mol. Sci. 2022, 23, 15423. https://doi.org/10.3390/ijms232315423
Kaneko S, Yanai K, Ishii H, Aomatsu A, Hirai K, Ookawara S, Ishibashi K, Morishita Y. miR−122−5p Regulates Renal Fibrosis In Vivo. International Journal of Molecular Sciences. 2022; 23(23):15423. https://doi.org/10.3390/ijms232315423
Chicago/Turabian StyleKaneko, Shohei, Katsunori Yanai, Hiroki Ishii, Akinori Aomatsu, Keiji Hirai, Susumu Ookawara, Kenichi Ishibashi, and Yoshiyuki Morishita. 2022. "miR−122−5p Regulates Renal Fibrosis In Vivo" International Journal of Molecular Sciences 23, no. 23: 15423. https://doi.org/10.3390/ijms232315423
APA StyleKaneko, S., Yanai, K., Ishii, H., Aomatsu, A., Hirai, K., Ookawara, S., Ishibashi, K., & Morishita, Y. (2022). miR−122−5p Regulates Renal Fibrosis In Vivo. International Journal of Molecular Sciences, 23(23), 15423. https://doi.org/10.3390/ijms232315423








