Key Genetic Components of Fibrosis in Diabetic Nephropathy: An Updated Systematic Review and Meta-Analysis
Abstract
1. Introduction
2. Methods
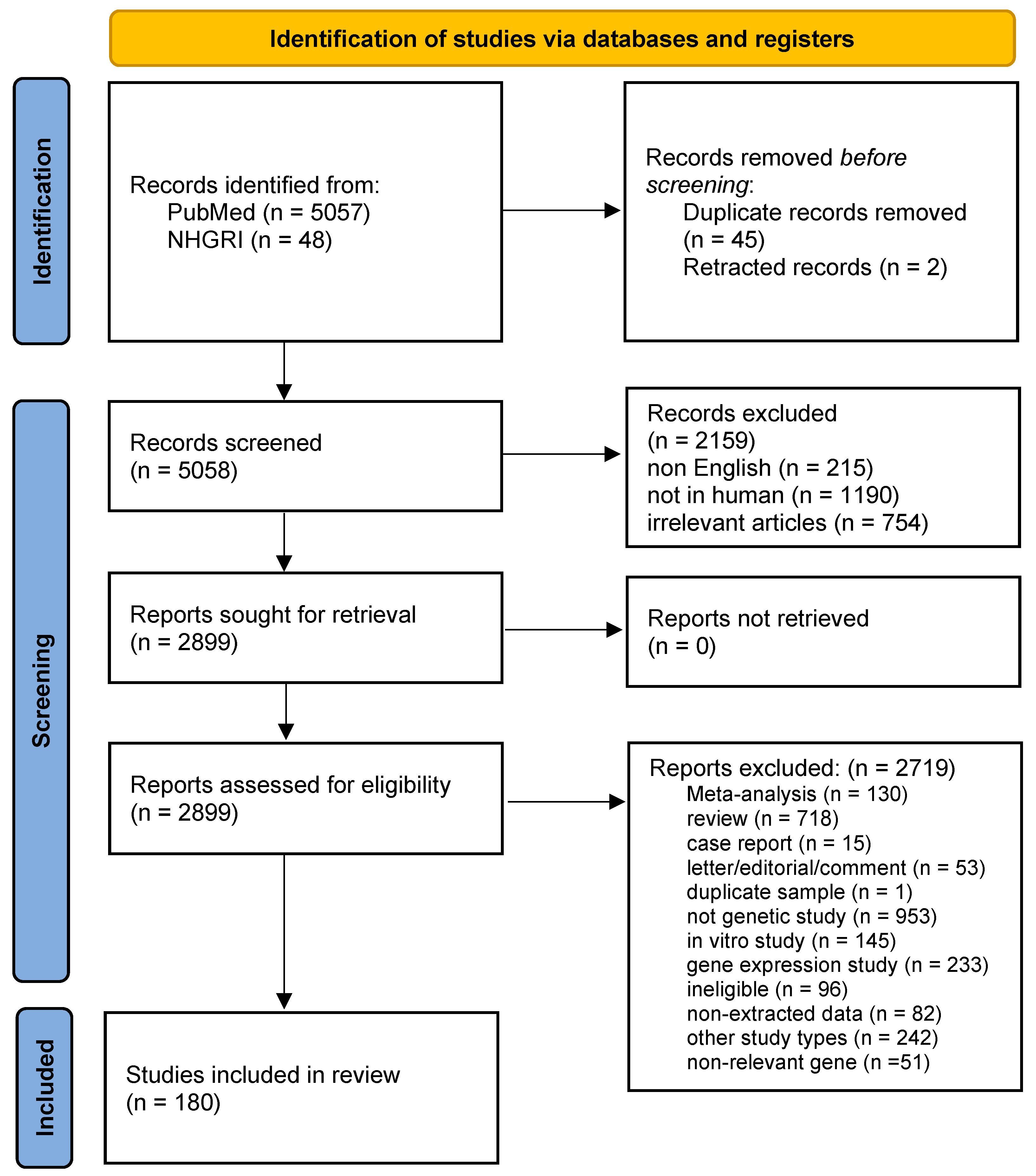
2.1. Identification and Eligibility of Relevant Studies
2.2. Data Synthesis and Analysis
3. Results and Discussion
3.1. Study Characteristics
3.2. Meta-Analysis Results
3.3. Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Rich, S.S. Genetics of diabetes and its complications. J. Am. Soc. Nephrol. 2006, 17, 353–360. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cowie, C.C.; Port, F.K.; Wolfe, R.A.; Savage, P.J.; Moll, P.P.; Hawthorne, V.M. Disparities in incidence of diabetic end-stage renal disease according to race and type of diabetes. N. Engl. J. Med. 1989, 321, 1074–1079. [Google Scholar] [CrossRef] [PubMed]
- Seaquist, E.R.; Goetz, F.C.; Rich, S.; Barbosa, J. Familial clustering of diabetic kidney disease. Evidence for genetic susceptibility to diabetic nephropathy. N. Engl. J. Med. 1989, 320, 1161–1165. [Google Scholar] [CrossRef] [PubMed]
- Thomas, M.C.; Groop, P.H.; Tryggvason, K. Towards understanding the inherited susceptibility for nephropathy in diabetes. Curr. Opin. Nephrol. Hypertens. 2012, 21, 195–202. [Google Scholar] [CrossRef]
- Rutledge, J.C.; Ng, K.F.; Aung, H.H.; Wilson, D.W. Role of triglyceride-rich lipoproteins in diabetic nephropathy. Nat. Rev. Nephrol. 2010, 6, 361–370. [Google Scholar] [CrossRef]
- Zhang, Y.; Jin, D.; Kang, X.; Zhou, R.; Sun, Y.; Lian, F.; Tong, X. Signaling Pathways Involved in Diabetic Renal Fibrosis. Front. Cell Dev. Biol. 2021, 9, 696542. [Google Scholar] [CrossRef]
- Li, L.; Fu, H.; Liu, Y. The fibrogenic niche in kidney fibrosis: Components and mechanisms. Nat. Rev. Nephrol. 2022, 18, 545–557. [Google Scholar] [CrossRef]
- Hu, L.; Ding, M.; He, W. Emerging Therapeutic Strategies for Attenuating Tubular EMT and Kidney Fibrosis by Targeting Wnt/β -Catenin Signaling. Front. Pharmacol. 2022, 12, 830340. [Google Scholar] [CrossRef]
- Edeling, M.; Ragi, G.; Huang, S.; Pavenstädt, H.; Susztak, K. Developmental Signalling Pathways in Renal Fibrosis: The Roles of Notch, Wnt and Hedgehog; Nature Publishing Group: Berlin/Heidelberg, Germany, 2016. [Google Scholar]
- Arai, H.; Yanagita, M. Janus-Faced: Molecular Mechanisms and Versatile Nature of Renal Fibrosis. Kidney360 2020, 1, 697–704. [Google Scholar] [CrossRef]
- Fragiadaki, M.; Macleod, F.M.; Ong, A.C.M. The controversial role of fibrosis in autosomal dominant polycystic kidney disease. Int. J. Mol. Sci. 2020, 21, 8936. [Google Scholar] [CrossRef]
- Tziastoudi, M.; Stefanidis, I.; Stravodimos, K.; Zintzaras, E. Identification of Chromosomal Regions Linked to Diabetic Nephropathy: A Meta-Analysis of Genome-Wide Linkage Scans. Genet. Test. Mol. Biomark. 2019, 23, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Tziastoudi, M.; Tziastoudi, M. The genetic map of diabetic nephropathy: Evidence from a systematic review and meta-analysis of genetic association studies. Clin. Kidney J. 2020, 13, 768–781. [Google Scholar] [CrossRef] [PubMed]
- Tziastoudi, M.; Stefanidis, I.; Hadjigeorgiou, G.M.; Stravodimos, K.; Zintzaras, E. A systematic review and meta-analysis of genetic association studies for the role of inflammation and the immune system in diabetic nephropathy. Clin. Kidney J. 2017, 10, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Tziastoudi, M.; Dardiotis, E.; Pissas, G.; Filippidis, G.; Golfinopoulos, S.; Siokas, V.; Tachmitzi, S.V.; Eleftheriadis, T.; Hadjigeorgiou, G.M.; Tsironi, E.; et al. Serpin Family E Member 1 Tag Single-Nucleotide Polymorphisms in Patients with Diabetic Nephropathy: An Association Study and Meta-Analysis Using a Genetic Model-Free Approach. Genes 2021, 12, 1887. [Google Scholar] [CrossRef] [PubMed]
- Stefanidis, I.; Tziastoudi, M.; Tsironi, E.E.; Dardiotis, E.; Tachmitzi, S.V.; Fotiadou, A.; Pissas, G.; Kytoudis, K.; Sounidaki, M.; Ampatzis, G.; et al. The contribution of genetic variants of SLC2A1 gene in T2DM and T2DM-nephropathy: Association study and meta-analysis. Ren. Fail. 2018, 40, 561–576. [Google Scholar] [CrossRef] [PubMed]
- Maeda, S.; Osawa, N.; Hayashi, T.; Tsukada, S.; Kobayashi, M.; Kikkawa, R. Genetic variations associated with diabetic nephropathy and type II diabetes in a Japanese population. Kidney Int. Suppl. 2007, 72, S43–S48. [Google Scholar] [CrossRef] [PubMed]
- Germain, M.; Pezzolesi, M.G.; Sandholm, N.; McKnight, A.J.; Susztak, K.; Lajer, M.; Forsblom, C.; Marre, M.; Parving, H.H.; Rossing, P.; et al. SORBS1 gene, a new candidate for diabetic nephropathy: Results from a multi-stage genome-wide association study in patients with type 1 diabetes. Diabetologia 2015, 58, 543–548. [Google Scholar] [CrossRef]
- Taira, M.; Imamura, M.; Takahashi, A.; Kamatani, Y.; Yamauchi, T.; Araki, S.I.; Tanaka, N.; Van Zuydam, N.R.; Ahlqvist, E.; Toyoda, M.; et al. A variant within the FTO confers susceptibility to diabetic nephropathy in Japanese patients with type 2 diabetes. PLoS ONE 2018, 13, e0208654. [Google Scholar] [CrossRef]
- Pezzolesi, M.G.; Poznik, G.D.; Mychaleckyj, J.C.; Paterson, A.D.; Barati, M.T.; Klein, J.B.; Ng, D.P.; Placha, G.; Canani, L.H.; Bochenski, J.; et al. Genome-Wide Association Scan for Diabetic Nephropathy Susceptibility Genes in Type 1 Diabetes. Diabetes 2009, 58, 1403–1410. [Google Scholar] [CrossRef]
- McDonough, C.W.; Palmer, N.D.; Hicks, P.J.; Roh, B.H.; An, S.S.; Cooke, J.N.; Hester, J.M.; Wing, M.R.; Bostrom, M.A.; Rudock, M.E.; et al. A genome-wide association study for diabetic nephropathy genes in African Americans. Kidney Int. 2011, 79, 563–572. [Google Scholar] [CrossRef]
- Hsieh, A.R.; Huang, Y.C.; Yang, Y.F.; Lin, H.J.; Lin, J.M.; Chang, Y.W.; Wu, C.M.; Liao, W.L.; Tsai, F.J. Lack of association of genetic variants for diabetic retinopathy in Taiwanese patients with diabetic nephropathy. BMJ Open Diabetes Res. Care 2020, 8, e000727. [Google Scholar] [CrossRef] [PubMed]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. BMJ 2021, 372, 89. Available online: https://www.bmj.com/content/372/bmj.n71 (accessed on 14 August 2022).
- Zintzaras, E. The power of generalized odds ratio in assessing association in genetic studies. J. Appl. Stat. 2012, 39, 2569–2581. [Google Scholar] [CrossRef]
- Zintzaras, E. The generalized odds ratio as a measure of genetic risk effect in the analysis and meta-analysis of association studies. Stat. Appl. Genet. Mol. Biol. 2010, 9, 21. [Google Scholar] [CrossRef] [PubMed]
- DerSimonian, R.; Laird, N. Meta-analysis in clinical trials. Control Clin. Trials 1986, 7, 177–188. [Google Scholar] [CrossRef]
- Higgins, J.P.T.; Thompson, S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 2002, 21, 1539–1558. [Google Scholar] [CrossRef]
- Cochran, W. The combination of estimates from different experiments. Biometrics 1954, 10, 101–129. [Google Scholar] [CrossRef]
- Egger, M.; Davey Smith, G.; Schneider, M.; Minder, C. Bias in meta-analysis detected by a simple, graphical test. BMJ 1997, 315, 629–634. [Google Scholar] [CrossRef]
- Karihaloo, A. Anti-fibrosis therapy and diabetic nephropathy. Curr. Diabetes Rep. 2012, 12, 414–422. [Google Scholar] [CrossRef]
- Mezzano, S.A.; Ruiz-Ortega, M.; Egido, J. Angiotensin II and Renal Fibrosis [Internet]. 2001. Available online: http://www.hypertensionaha.org (accessed on 11 June 2022).
- Meng, X.M.; Nikolic-Paterson, D.J.; Lan, H.Y. TGF-β: The master regulator of fibrosis. Nat. Rev. Nephrol. 2016, 12, 325–338. [Google Scholar] [CrossRef]
- van Meeteren, L.A.; ten Dijke, P. Regulation of endothelial cell plasticity by TGF-β. Cell Tissue Res. 2012, 347, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Lemos, D.R.; McMurdo, M.; Karaca, G.; Wilflingseder, J.; Leaf, I.A.; Gupta, N.; Miyoshi, T.; Susa, K.; Johnson, B.G.; Soliman, K.; et al. Interleukin-1b activates a MYC-dependent metabolic switch in kidney stromal cells necessary for progressive tubulointerstitial fibrosis. J. Am. Soc. Nephrol. 2018, 29, 1690–1705. [Google Scholar] [CrossRef] [PubMed]
- Ranganathan, P.; Jayakumar, C.; Ramesh, G. Proximal tubule-specific overexpression of netrin-1 suppresses acute kidney injury-induced interstitial fibrosis and glomerulosclerosis through suppression of IL-6/STAT3 signaling. Am. J. Physiol. Ren. Physiol. 2013, 304, 1054–1065. [Google Scholar]
- Steen, E.H.; Wang, X.; Balaji, S.; Butte, M.J.; Bollyky, P.L.; Keswani, S.G. The Role of the Anti-Inflammatory Cytokine Interleukin-10 in Tissue Fibrosis. Adv. Wound Care 2020, 9, 184–198. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, K.; Taguchi, K.; Kishi, S.; Brooks, C.R.; Ochi, A.; Kadoya, H.; Ikeda, Y.; Miyoshi, M.; Tamaki, M.; Abe, H.; et al. Dual disruption of eNOS and ApoE gene accelerates kidney fibrosis and senescence after injury. Biochem. Biophys. Res. Commun. 2021, 556, 142–148. [Google Scholar] [CrossRef]
- Sun, X.; Gan, H.; Xia, Y. A meta-analysis of the effects of endothelial nitric oxide synthase 4ba polymorphism on renal interstitial fibrosis in diabetic nephropathy. Ann. Palliat. Med. 2021, 10, 633–645. [Google Scholar] [CrossRef] [PubMed]
- Neder, T.H.; Schrankl, J.; Fuchs, M.A.A.; Broeker, K.A.E.; Wagner, C. Endothelin receptors in renal interstitial cells do not contribute to the development of fibrosis during experimental kidney disease. Pflug. Arch. 2021, 473, 1667–1683. [Google Scholar] [CrossRef] [PubMed]
- Heerspink, H.J.; Parving, H.H.; Andress, D.L.; Bakris, G.; Correa-Rotter, R.; Hou, F.F.; Kitzman, D.W.; Kohan, D.; Makino, H.; McMurray, J.J.; et al. Atrasentan and renal events in patients with type 2 diabetes and chronic kidney disease (SONAR): A double-blind, randomised, placebo-controlled trial. Lancet 2019, 393, 1937–1947. [Google Scholar]
- Geng, X.C.; Hu, Z.P.; Lian, G.Y. Erythropoietin ameliorates renal interstitial fibrosis via the inhibition of fibrocyte accumulation. Mol. Med. Rep. 2015, 11, 3860–3865. [Google Scholar] [CrossRef]
- Church, R.H.; Ali, I.; Tate, M.; Lavin, D.; Krishnakumar, A.; Kok, H.M.; Hombrebueno, J.R.; Dunne, P.D.; Bingham, V.; Goldschmeding, R.; et al. Gremlin1 plays a key role in kidney development and renal fibrosis. Am. J. Physiol. Ren. Physiol. 2017, 312, 1141–1157. [Google Scholar] [CrossRef]
- Kinashi, H.; Ito, Y.; Sun, T.; Katsuno, T.; Takei, Y. Roles of the TGF-β–VEGF-C pathway in fibrosis-related lymphangiogenesis. Int. J. Mol. Sci. 2018, 19, 2487. [Google Scholar] [CrossRef] [PubMed]
- Grotendorst, G.R.; Rahmanie, H.; Duncan, M.R. Combinatorial signaling pathways determine fibroblast proliferation and myofibroblast differentiation. FASEB J. 2004, 18, 469–479. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Liu, H.; Takagi, S.; Nitta, K.; Kitada, M.; Srivastava, S.P.; Takagaki, Y.; Kanasaki, K.; Koya, D. Renal protective effects of empagliflozin via inhibition of EMT and aberrant glycolysis in proximal tubules. JCI Insight 2020, 5, e129034. [Google Scholar] [CrossRef] [PubMed]
- Miao, C.; Zhu, X.; Wei, X.; Long, M.; Jiang, L.; Li, C.; Jin, D.; Du, Y. Pro- and anti-fibrotic effects of vascular endothelial growth factor in chronic kidney diseases. Ren. Fail. 2022, 44, 881–892. [Google Scholar] [CrossRef] [PubMed]
- Moretti, L.; Stalfort, J.; Barker, T.H.; Abebayehu, D. The interplay of fibroblasts, the extracellular matrix, and inflammation in scar formation. J. Biol. Chem. 2022, 298, 101530. [Google Scholar] [CrossRef]
- Owens, E.P.; Vesey, D.A.; Kassianos, A.J.; Healy, H.; Hoy, W.E.; Gobe, G.C. Biomarkers and the role of mast cells as facilitators of inflammation and fibrosis in chronic kidney disease. Transl. Androl. Urol. 2019, 8, S175–S183. [Google Scholar] [CrossRef]
- Strattan, E.; Hildebrandt, G.C. Mast cell involvement in fibrosis in chronic graft-versus-host disease. Int. J. Mol. Sci. 2021, 22, 2385. [Google Scholar] [CrossRef]
- Conti, P.; Caraffa, A.; Mastrangelo, F.; Tettamanti, L.; Ronconi, G.; Frydas, I.; Kritas, S.K.; Theoharides, T.C. Critical role of inflammatory mast cell in fibrosis: Potential therapeutic effect of IL-37. Cell Prolif. 2018, 51, e12475. [Google Scholar] [CrossRef]
- Bradding, P.; Pejler, G. The controversial role of mast cells in fibrosis. Immunol. Rev. 2018, 282, 198–231. [Google Scholar]
- El-Koraie, A.F.; Baddour, N.M.; Adam, A.G.; el Kashef, E.H.; el Nahas, A.M. Role of stem cell factor and mast cells in the progression of chronic glomerulonephritides. Kidney Int. 2001, 60, 167–172. [Google Scholar] [CrossRef]
- Ehara, T.; Shigematsu, H. Contribution of mast cells to the tubulointerstitial lesions in IgA nephritis. Kidney Int. 1998, 54, 1675–1683. [Google Scholar] [CrossRef]
- Kurusu, A.; Suzuki, Y.; Horikoshi, S.; Shirato, I.; Tomino, Y. Relationship between Mast Cells in the Tubulointerstitium and Prognosis of Patients with IgA Nephropathy. Nephron 2001, 89, 391–397. [Google Scholar] [CrossRef] [PubMed]
- Kondo, S.; Kagami, S.; Kido, H.; Strutz, F.; Müller, G.A.; Kuroda, Y. Role of Mast Cell Tryptase in Renal Interstitial Fibrosis. J. Am. Soc. Nephrol. 2001, 12, 1668–1676. [Google Scholar] [CrossRef] [PubMed]
- Roberts, I.S.D.; Brenchley, P.E.C. Mast cells: The forgotten cells of renal fibrosis. J. Clin. Pathol. 2000, 53, 858–862. [Google Scholar] [CrossRef] [PubMed]
- Gieseck, R.L.; Wilson, M.S.; Wynn, T.A. Type 2 immunity in tissue repair and fibrosis. Nat. Rev. Immunol. 2018, 18, 62–76. [Google Scholar] [PubMed]
- Mukai, K.; Tsai, M.; Saito, H.; Galli, S.J. Mast cells as sources of cytokines, chemokines, and growth factors. Immunol. Rev. 2018, 282, 121–150. [Google Scholar]
- Theoharides, T.C.; Alysandratos, K.D.; Angelidou, A.; Delivanis, D.A.; Sismanopoulos, N.; Zhang, B.; Asadi, S.; Vasiadi, M.; Weng, Z.; Miniati, A.; et al. Mast cells and inflammation. Biochim. Et Biophys. Acta Mol. Basis Dis. 2012, 1822, 21–33. [Google Scholar]
- Hügle, T. Beyond allergy: The role of mast cells in fibrosis. Swiss Med. Wkly. 2014, 144, w13999. [Google Scholar] [CrossRef]
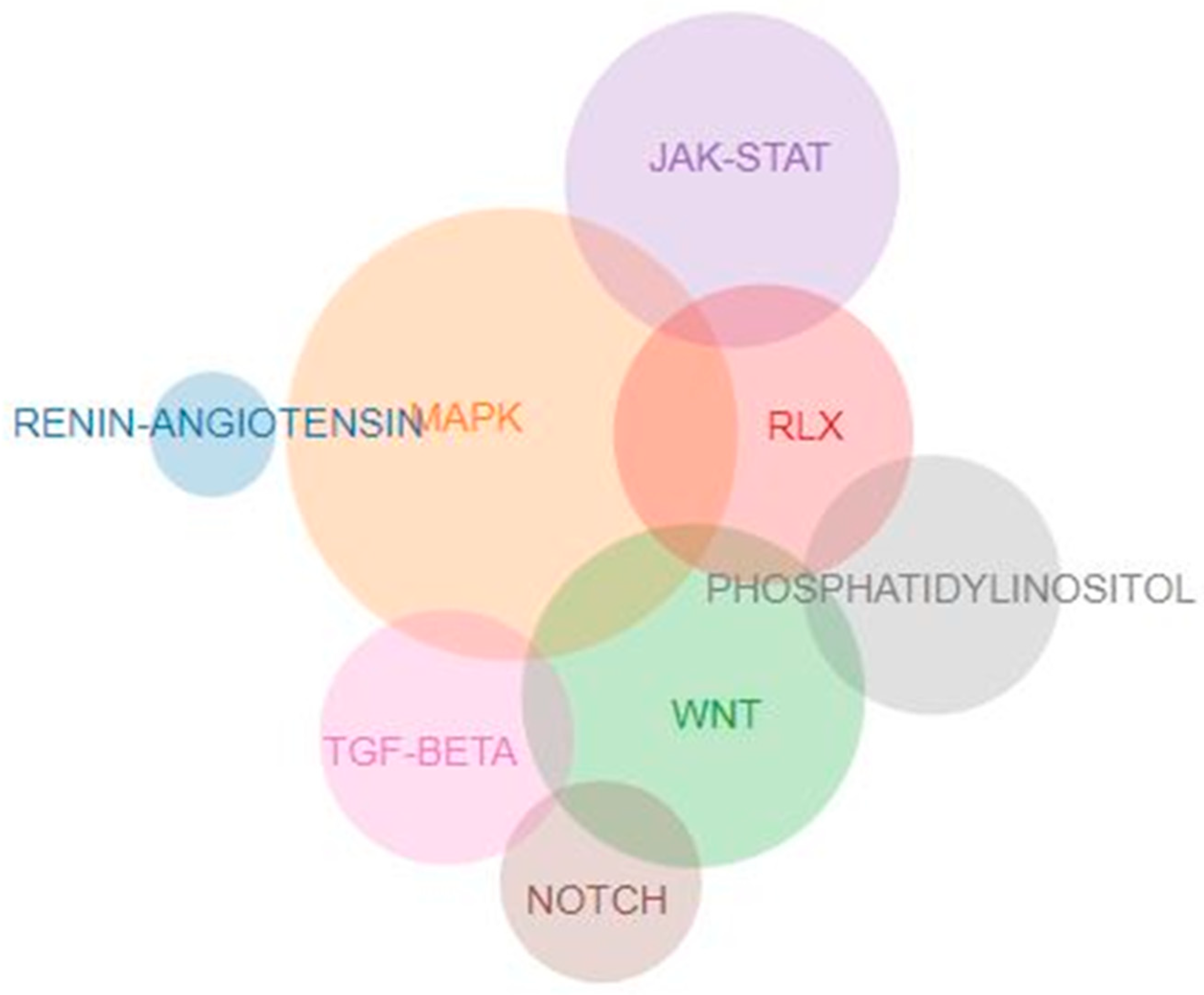


| ACE pathway ACE, ACE2, AGT, AGTR1, AGTR2, ANPEP, ATP6AP2, CMA1, CPA3, CTSA, CTSG, ENPEP, KLK1, KLK2, LNPEP, MAS1, MME, MRGPRD, NLN, PRCP, PREP, REN, THOP1 |
| Relaxin pathway ACTA2, ADCY1, ADCY2, ADCY3, ADCY4, ADCY5, ADCY6, ADCY7, ADCY8, ADCY9, AKT1, AKT2, AKT3, ARRB1, ARRB2, ATF2, ATF4, ATF6B, COL1A1, COL1A2, COL3A1, COL4A1, COL4A2, COL4A3, COL4A4, COL4A5, COL4A6, CREB1, CREB3, CREB3L1, CREB3L2, CREB3L3, CREB3L4, CREB5, EDN1, EDNRB, EGFR, FOS, GNA15, GNAI1, GNAI2, GNAI3, GNAO1, GNAS, GNB1, GNB2, GNB3, GNB4, GNB5, GNG10, GNG11, GNG12, GNG13, GNG2, GNG3, GNG4, GNG5, GNG7, GNG8, GNGT1, GNGT2, GRB2, HRAS, INSL3, INSL5, JUN, KRAS, MAP2K1, MAP2K2, MAP2K4, MAP2K7, MAPK1, MAPK10, MAPK11, MAPK12, MAPK13, MAPK14, MAPK3, MAPK8, MAPK9, MMP1, MMP13, MMP2, MMP9, NFKB1, NFKBIA, NOS1, NOS2, NOS3, NRAS, PIK3CA, PIK3CB, PIK3CD, PIK3R1, PIK3R2, PIK3R3, PLCB1, PLCB2, PLCB3, PLCB4, PRKACA, PRKACB, PRKACG, PRKCA, PRKCZ, RAF1, RELA, RLN1, RLN2, RLN3, RXFP1, RXFP2, RXFP3, RXFP4, SHC1, SHC2, SHC3, SHC4, SMAD2, SOS1, SOS2, SRC, TGFB1, TGFBR1, TGFBR2, VEGFA, VEGFB, VEGFC, VEGFD |
| Wnt signaling pathway APC, APC2, APCDD1, APCDD1L, AXIN1, AXIN2, BAMBI, BTRC, CACYBP, CAMK2A, CAMK2B, CAMK2D, CAMK2G, CBY1, CCDC88C, CCN4, CCND1, CCND2, CCND3, CER1, CHD8, CREBBP, CSNK1A1, CSNK1A1L, CSNK1E, CSNK2A1, CSNK2A2, CSNK2A3, CSNK2B, CTBP1, CTBP2, CTNNB1, CTNNBIP1, CTNND2, CUL1, CXXC4, DAAM1, DAAM2, DKK1, DKK2, DKK4, DVL1, DVL2, DVL3, EP300, FBXW11, FOSL1, FRAT1, FRAT2, FRZB, FZD1, FZD10, FZD2, FZD3, FZD4, FZD5, FZD6, FZD7, FZD8, FZD9, GPC4, GSK3B, INVS, JUN, LEF1, LGR4, LGR5, LGR6, LRP5, LRP6, MAP3K7, MAPK10, MAPK8, MAPK9, MMP7, MYC, NFATC1, NFATC2, NFATC3, NFATC4, NKD1, NKD2, NLK, NOTUM, PLCB1, PLCB2, PLCB3, PLCB4, PORCN, PPARD, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PRICKLE1, PRICKLE2, PRICKLE3, PRICKLE4, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB, PRKCG, PSEN1, RAC1, RAC2, RAC3, RBX1, RHOA, RNF43, ROCK2, ROR1, ROR2, RSPO1, RSPO2, RSPO3, RSPO4, RUVBL1, RYK, SENP2, SERPINF1, SFRP1, SFRP2, SFRP4, SFRP5, SIAH1, SKP1, SMAD3, SMAD4, SOST, SOX17, TBL1X, TBL1XR1, TBL1Y, TCF7, TCF7L1, TCF7L2, TLE1, TLE2, TLE3, TLE4, TLE6, TLE7, TP53, TPTEP2-CSNK1E, VANGL1, VANGL2, WIF1, WNT1, WNT10A, WNT10B, WNT11, WNT16, WNT2, WNT2B, WNT3, WNT3A, WNT4, WNT5A, WNT5B, WNT6, WNT7A, WNT7B, WNT8A, WNT8B, WNT9A, WNT9B, ZNRF3 |
| MAPK signaling pathway AKT1, AKT2, AKT3, ANGPT1, ANGPT2, ANGPT4, ARAF, AREG, ARRB1, ARRB2, ATF2, ATF4, BDNF, BRAF, CACNA1A, CACNA1B, CACNA1C, CACNA1D, CACNA1E, CACNA1F, CACNA1G, CACNA1H, CACNA1I, CACNA1S, CACNA2D1, CACNA2D2, CACNA2D3, CACNA2D4, CACNB1, CACNB2, CACNB3, CACNB4, CACNG1, CACNG2, CACNG3, CACNG4, CACNG5, CACNG6, CACNG7, CACNG8, CASP3, CD14, CDC25B, CDC42, CHUK, CRK, CRKL, CSF1, CSF1R, DAXX, DDIT3, DUSP1, DUSP10, DUSP16, DUSP2, DUSP3, DUSP4, DUSP5, DUSP6, DUSP7, DUSP8, DUSP9, ECSIT, EFNA1, EFNA2, EFNA3, EFNA4, EFNA5, EGF, EGFR, ELK1, ELK4, EPHA2, ERBB2, ERBB3, ERBB4, EREG, FAS, FASLG, FGF1, FGF10, FGF16, FGF17, FGF18, FGF19, FGF2, FGF20, FGF21, FGF22, FGF23, FGF3, FGF4, FGF5, FGF6, FGF7, FGF8, FGF9, FGFR1, FGFR2, FGFR3, FGFR4, FLNA, FLNB, FLNC, FLT1, FLT3, FLT3LG, FLT4, FOS, GADD45A, GADD45B, GADD45G, GNA12, GNG12, GRB2, HGF, HRAS, HSPA1A, HSPA1B, HSPA1L, HSPA2, HSPA6, HSPA8, HSPB1, IGF1, IGF1R, IGF2, IKBKB, IKBKG, IL1A, IL1B, IL1R1, IL1RAP, INS, INSR, IRAK1, IRAK4, JMJD7-PLA2G4B, JUN, JUND, KDR, KIT, KITLG, KRAS, LAMTOR3, MAP2K1, MAP2K2, MAP2K3, MAP2K4, MAP2K5, MAP2K6, MAP2K7, MAP3K1, MAP3K11, MAP3K12, MAP3K13, MAP3K14, MAP3K2, MAP3K20, MAP3K3, MAP3K4, MAP3K5, MAP3K6, MAP3K7, MAP3K8, MAP4K1, MAP4K2, MAP4K3, MAP4K4, MAPK1, MAPK10, MAPK11, MAPK12, MAPK13, MAPK14, MAPK3, MAPK7, MAPK8, MAPK8IP1, MAPK8IP2, MAPK8IP3, MAPK9, MAPKAPK2, MAPKAPK3, MAPKAPK5, MAPT, MAX, MECOM, MEF2C, MET, MKNK1, MKNK2, MRAS, MYC, MYD88, NF1, NFATC1, NFATC3, NFKB1, NFKB2, NGF, NGFR, NLK, NR4A1, NRAS, NTF3, NTF4, NTRK1, NTRK2, PAK1, PAK2, PDGFA, PDGFB, PDGFC, PDGFD, PDGFRA, PDGFRB, PGF, PLA2G4A, PLA2G4B, PLA2G4C, PLA2G4D, PLA2G4E, PLA2G4F, PPM1A, PPM1B, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PPP5C, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB, PRKCG, PTPN5, PTPN7, PTPRR, RAC1, RAC2, RAC3, RAF1, RAP1A, RAP1B, RAPGEF2, RASA1, RASA2, RASGRF1, RASGRF2, RASGRP1, RASGRP2, RASGRP3, RASGRP4, RELA, RELB, RPS6KA1, RPS6KA2, RPS6KA3, RPS6KA4, RPS6KA5, RPS6KA6, RRAS, RRAS2, SOS1, SOS2, SRF, STK3, STK4, STMN1, TAB1, TAB2, TAOK1, TAOK2, TAOK3, TEK, TGFA, TGFB1, TGFB2, TGFB3, TGFBR1, TGFBR2, TNF, TNFRSF1A, TP53, TRADD, TRAF2, TRAF6, VEGFA, VEGFB, VEGFC, VEGFD |
| PI3KC signaling pathway BPNT2, CALM1, CALM2, CALM3, CALML3, CALML4, CALML5, CALML6, CDIPT, CDS1, CDS2, DGKA, DGKB, DGKD, DGKE, DGKG, DGKH, DGKI, DGKK, DGKQ, DGKZ, IMPA1, IMPA2, INPP1, INPP4A, INPP4B, INPP5A, INPP5B, INPP5D, INPP5E, INPP5F, INPPL1, IP6K1, IP6K2, IP6K3, IPMK, IPPK, ITPK1, ITPKA, ITPKB, ITPKC, ITPR1, ITPR2, ITPR3, MTM1, MTMR1, MTMR14, MTMR2, MTMR3, MTMR4, MTMR6, MTMR7, MTMR8, OCRL, PI4K2A, PI4K2B, PI4KA, PI4KB, PIK3C2A, PIK3C2B, PIK3C2G, PIK3C3, PIK3CA, PIK3CB, PIK3CD, PIK3R1, PIK3R2, PIK3R3, PIKFYVE, PIP4K2A, PIP4K2B, PIP4K2C, PIP4P1, PIP4P2, PIP5K1A, PIP5K1B, PIP5K1C, PLCB1, PLCB2, PLCB3, PLCB4, PLCD1, PLCD3, PLCD4, PLCE1, PLCG1, PLCG2, PLCZ1, PPIP5K1, PPIP5K2, PRKCA, PRKCB, PRKCG, PTEN, SACM1L, SYNJ1, SYNJ2 |
| TGFB1 signaling pathway ACVR1, ACVR1B, ACVR1C, ACVR2A, ACVR2B, AMH, AMHR2, BAMBI, BMP2, BMP4, BMP5, BMP6, BMP7, BMP8A, BMP8B, BMPR1A, BMPR1B, BMPR2, CDKN2B, CHRD, CREBBP, CUL1, DCN, E2F4, E2F5, EP300, FBN1, FMOD, FST, GDF5, GDF6, GDF7, GREM1, GREM2, HAMP, HJV, ID1, ID2, ID3, ID4, IFNG, INHBA, INHBB, INHBC, INHBE, LEFTY1, LEFTY2, LTBP1, MAPK1, MAPK3, MICOS10-NBL1, MYC, NBL1, NEO1, NODAL, NOG, PITX2, PPP2CA, PPP2CB, PPP2R1A, PPP2R1B, RBL1, RBX1, RGMA, RGMB, RHOA, ROCK1, RPS6KB1, RPS6KB2, SKP1, SMAD1, SMAD2, SMAD3, SMAD4, SMAD5, SMAD6, SMAD7, SMAD9, SMURF1, SMURF2, SP1, TFDP1, TGFB1, TGFB2, TGFB3, TGFBR1, TGFBR2, TGIF1, TGIF2, THBS1, THSD4, TNF, ZFYVE16, ZFYVE9 |
| NOTCH signaling pathway ADAM17, APH1A, APH1B, ATXN1, ATXN1L, CIR1, CREBBP, CTBP1, CTBP2, DLL1, DLL3, DLL4, DTX1, DTX2, DTX3, DTX3L, DTX4, DVL1, DVL2, DVL3, EP300, HDAC1, HDAC2, HES1, HES5, HEY1, HEY2, HEYL, JAG1, JAG2, KAT2A, KAT2B, LFNG, MAML1, MAML2, MAML3, MFNG, NCOR2, NCSTN, NOTCH1, NOTCH2, NOTCH3, NOTCH4, NUMB, NUMBL, PSEN1, PSEN2, PSENEN, PTCRA, RBPJ, RBPJL, RFNG, SNW1, TLE1, TLE2, TLE3, TLE4, TLE6, TLE7 |
| JAK signaling pathway AKT1, AKT2, AKT3, AOX1, BCL2, BCL2L1, CCND1, CCND2, CCND3, CDKN1A, CISH, CNTF, CNTFR, CREBBP, CRLF2, CSF2, CSF2RA, CSF2RB, CSF3, CSF3R, CSH1, CSH2, CTF1, EGF, EGFR, EP300, EPO, EPOR, FHL1, GFAP, GH1, GH2, GHR, GRB2, HRAS, IFNA1, IFNA10, IFNA13, IFNA14, IFNA16, IFNA17, IFNA2, IFNA21, IFNA4, IFNA5, IFNA6, IFNA7, IFNA8, IFNAR1, IFNAR2, IFNB1, IFNE, IFNG, IFNGR1, IFNGR2, IFNK, IFNL1, IFNL2, IFNL3, IFNLR1, IFNW1, IL10, IL10RA, IL10RB, IL11, IL11RA, IL12A, IL12B, IL12RB1, IL12RB2, IL13, IL13RA1, IL13RA2, IL15, IL15RA, IL17D, IL19, IL2, IL20, IL20RA, IL20RB, IL21, IL21R, IL22, IL22RA1, IL22RA2, IL23A, IL23R, IL24, IL27RA, IL2RA, IL2RB, IL2RG, IL3, IL3RA, IL4, IL4R, IL5, IL5RA, IL6, IL6R, IL6ST, IL7, IL7R, IL9, IL9R, IRF9, JAK1, JAK2, JAK3, LEP, LEPR, LIF, LIFR, MCL1, MPL, MTOR, MYC, OSM, OSMR, PDGFA, PDGFB, PDGFRA, PDGFRB, PIAS1, PIAS2, PIAS3, PIAS4, PIK3CA, PIK3CB, PIK3CD, PIK3R1, PIK3R2, PIK3R3, PIM1, PRL, PRLR, PTPN11, PTPN2, PTPN6, RAF1, SOCS1, SOCS2, SOCS3, SOCS4, SOCS5, SOCS6, SOCS7, SOS1, SOS2, STAM, STAM2, STAT1, STAT2, STAT3, STAT4, STAT5A, STAT5B, STAT6, THPO, TSLP, TYK2 |
| Gene | Variant | RS | Studies (n) | Cases/Controls (n) | RE ORG | 95% LL | 95% UL | I2(%) | PQ | PE | Current Status |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Diseased Controls versus Cases | |||||||||||
| ACE | I > D | rs4646994 | 66 | 11437/10984 | 1.22 | 1.10 | 1.35 | 76.34 | 0.00 | 0.70 | updated |
| All in HWE | I > D | 56 | 9383/8847 | 1.28 | 1.16 | 1.41 | 67.29 | 0.00 | 0.65 | ||
| AGT | M235T | rs699 | 26 | 5015/5253 | 1.21 | 1.01 | 1.45 | 82,45 | 0,00 | 0.84 | [13] |
| All in HWE | 19 | 3181/3655 | 1.09 | 0.92 | 1.31 | 72.76 | 0.00 | 0.95 | |||
| EPO | G > T | rs1617640 | 3 | 1618/954 | 1.64 | 1.43 | 1.89 | 0.00 | 0.78 | 0.03 | [13] |
| GREM1 | rs1129456 (A/T) | 2 | 859/940 | 1.55 | 1.23 | 1.94 | 1.02 | 0.31 | na | new | |
| IL1B | −511C > T | rs16944 | 3 | 774/667 | 1.66 | 1.38 | 2.01 | 0.00 | 0.86 | 0.28 | [13] |
| All in HWE | 3 | ||||||||||
| IL10 | −1082 A > G | rs1800896 | 4 | 677/761 | 1.23 | 1.01 | 1.49 | 0 | 0.56 | 0.63 | [13] |
| All in HWE | 2 | 610/690 | 1.25 | 1.02 | 1.53 | 0 | 0.62 | na | |||
| NOS3 | T-786C | rs2070744 | 9 | 2288/2154 | 1.21 | 1.08 | 1.36 | 0.00 | 0.62 | 0.40 | updated |
| All in HWE | 7 | 2026/1862 | 1.21 | 1.08 | 1.37 | 3.08 | 0.40 | 0.29 | |||
| NOS3 | G894T | rs1799983 | 21 | 4538/3774 | 1.19 | 0.98 | 1.44 | 77.97 | <0.001 | 0.36 | updated |
| All in HWE | 19 | 4306/3564 | 1.24 | 1.02 | 1.51 | 77.68 | 0.00 | 0.20 | |||
| NOS3 | rs869109213 | 2 | 354/444 | 1.47 | 1.11 | 1.95 | 0.00 | 0.95 | na | updated | |
| All in HWE | 2 | na | |||||||||
| Healthy Controls versus Cases | |||||||||||
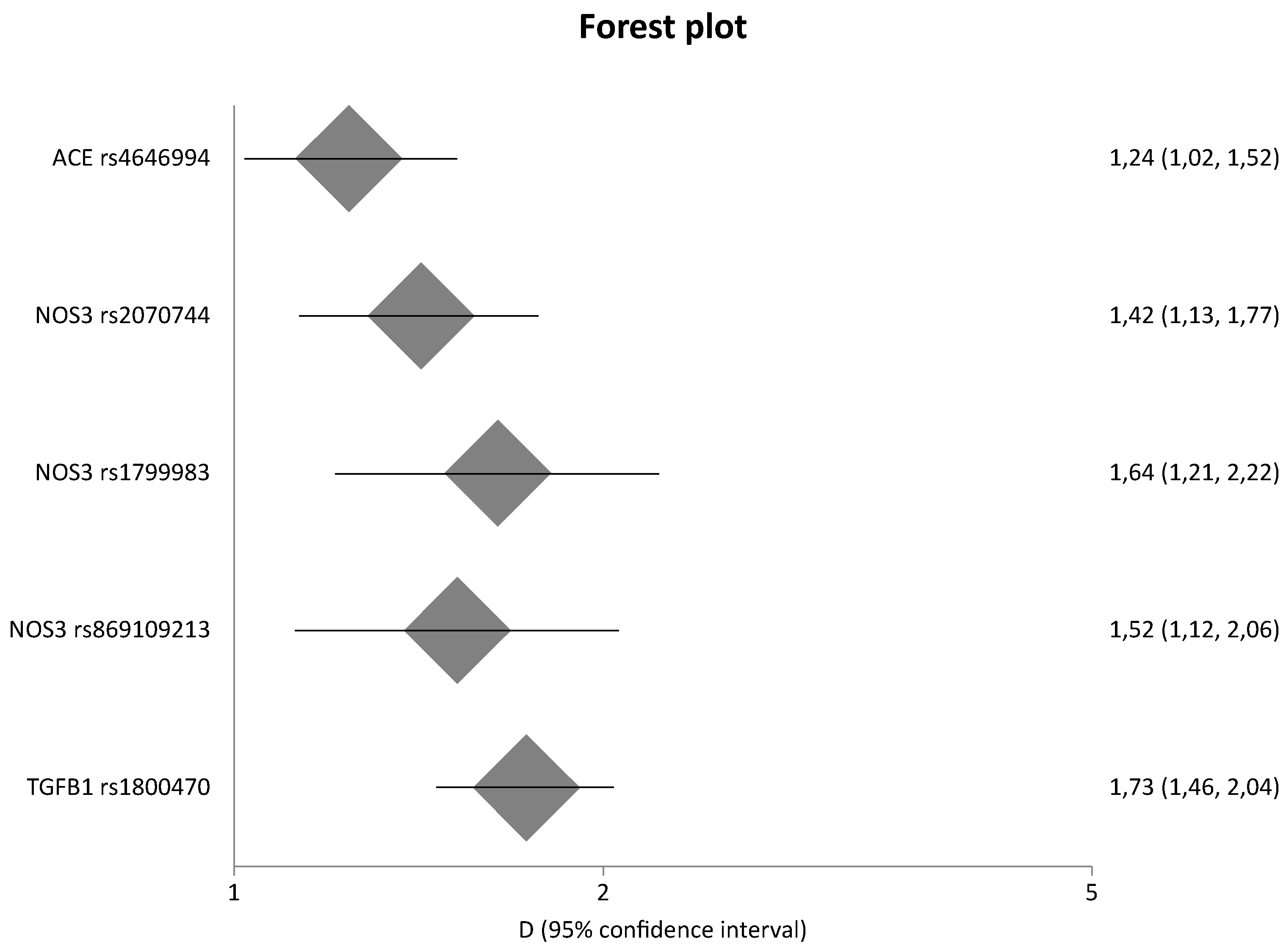
| ACE | I > D | rs4646994 | 30 | 3690/4927 | 1.24 | 1.02 | 1.52 | 83.20 | 0.00 | 0.03 | [13] |
| All in HWE | I > D | 29 | 3283/4695 | 1.26 | 1.02 | 1.55 | 82.87 | 0.00 | 0.01 | ||
| NOS3 | T-786C | rs2070744 | 9 | 1583/2142 | 1.42 | 1.13 | 1.77 | 58.00 | 0.01 | 0.84 | updated |
| All in HWE | 8 | 1516/2042 | 1.41 | 1.11 | 1.79 | 63.04 | 0.01 | 0.80 | |||
| NOS3 | G894T | rs1799983 | 11 | 2295/2737 | 1.64 | 1.21 | 2.22 | 82.07 | <0.001 | 0.11 | updated |
| All in HWE | 10 | 2247/2467 | 1.55 | 1.14 | 2.11 | 82.40 | <0.001 | 0.21 | |||
| NOS3 | rs869109213 | 2 | 354/444 | 1.52 | 1.12 | 2.06 | 17.59 | 0.27 | na | updated | |
| All in HWE | 2 | ||||||||||
| TGFB1 | T869C | rs1800470 | 6 | 814/1450 | 1.30 | 0.86 | 1.96 | 83.64 | 0 | 0.18 | |
| All in HWE | 4 | 706/1103 | 1.73 | 1.46 | 2.04 | 0 | 0.41 | 0.21 | |||
| Healthy Controls versus Diseased Controls versus Cases | |||||||||||
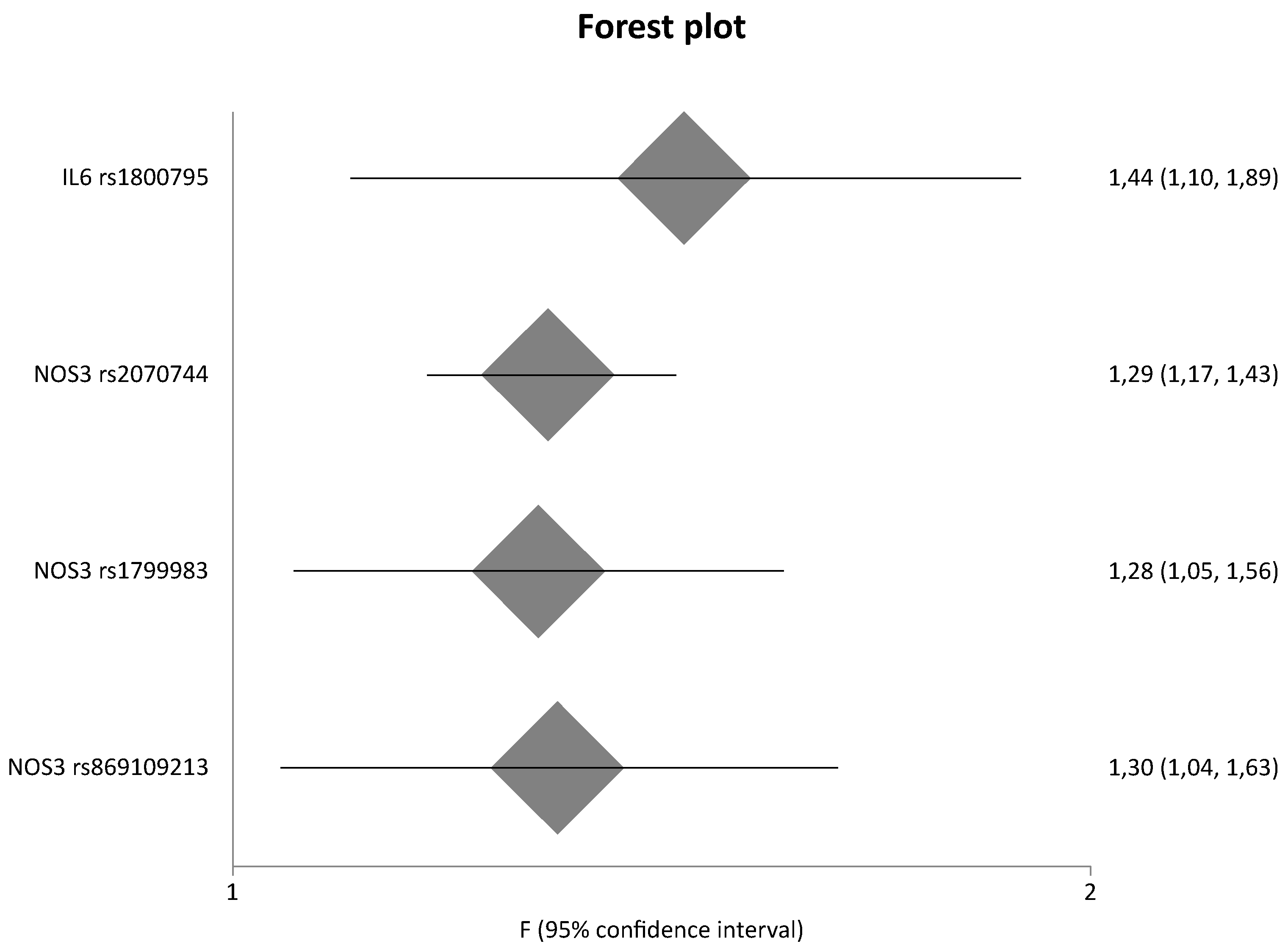
| IL6 | G(−174)C | rs1800795 | 2 | 90/234/212 | 1.44 | 1.10 | 1.89 | 0.00 | 0.42 | na | updated |
| All in HWE | 1 | ||||||||||
| NOS3 | T-786C | rs2070744 | 5 | 1307/1117/1451 | 1.29 | 1.17 | 1.43 | 0.00 | 0.53 | 0.46 | updated |
| All in HWE | 4 | 1240/1080/1351 | 1.29 | 1.16 | 1.43 | 3.59 | 0.37 | 0.51 | |||
| NOS3 | G894T | rs1799983 | 8 | 1506/1255/1642 | 1.28 | 1.05 | 1.56 | 70.00 | 0.01 | 0.32 | updated |
| All in HWE | 7 | 1.35 | 1.17 | 1.56 | 40.52 | 0.12 | 0.41 | ||||
| NOS3 | rs869109213 | 2 | 354/444/515 | 1.30 | 1.04 | 1.63 | 30.45 | 0.23 | na | updated | |
| All in HWE | 2 | ||||||||||
| GENE | Variant | RS | Studies (n) | Cases/ Controls (n) | RE OR | 95% LL | 95% UL | I2 (%) | PQ | PE | Current Status |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Diseased Controls versus Cases | |||||||||||
| EDN1 | rs1794849 | 3 | 1176/1323 | 1.16 | 1.02 | 1.31 | 0 | 0.62 | 0.08 | [13] | |
| FLT4 | rs2242221 | 3 | 1176/1323 | 1.14 | 1.01 | 1.29 | 0 | 0.38 | 0.43 | [13] | |
| IGF2/INS/TH cluster | rs1004446 | 3 | 1176/1323 | 1.16 | 1.03 | 1.31 | 0 | 0.49 | 0.22 | [13] | |
| IGF2/INS/TH cluster | rs4320932 | 3 | 1176/1323 | 0.84 | 0.73 | 0.96 | 0 | 0.43 | 0.06 | [13] | |
| VEGFA | C > A | rs2146323 | 3 | 1176/1323 | 0.85 | 0.76 | 0.95 | 0.2 | 0.2 | [13] | |
| Healthy Controls versus Cases | |||||||||||
| IL12RB1 | rs372889 | 2 | 1674/1719 | 1.243 | 1.130 | 1.367 | 0 | 0.567 | - | new | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tziastoudi, M.; Theoharides, T.C.; Nikolaou, E.; Efthymiadi, M.; Eleftheriadis, T.; Stefanidis, I. Key Genetic Components of Fibrosis in Diabetic Nephropathy: An Updated Systematic Review and Meta-Analysis. Int. J. Mol. Sci. 2022, 23, 15331. https://doi.org/10.3390/ijms232315331
Tziastoudi M, Theoharides TC, Nikolaou E, Efthymiadi M, Eleftheriadis T, Stefanidis I. Key Genetic Components of Fibrosis in Diabetic Nephropathy: An Updated Systematic Review and Meta-Analysis. International Journal of Molecular Sciences. 2022; 23(23):15331. https://doi.org/10.3390/ijms232315331
Chicago/Turabian StyleTziastoudi, Maria, Theoharis C. Theoharides, Evdokia Nikolaou, Maria Efthymiadi, Theodoros Eleftheriadis, and Ioannis Stefanidis. 2022. "Key Genetic Components of Fibrosis in Diabetic Nephropathy: An Updated Systematic Review and Meta-Analysis" International Journal of Molecular Sciences 23, no. 23: 15331. https://doi.org/10.3390/ijms232315331
APA StyleTziastoudi, M., Theoharides, T. C., Nikolaou, E., Efthymiadi, M., Eleftheriadis, T., & Stefanidis, I. (2022). Key Genetic Components of Fibrosis in Diabetic Nephropathy: An Updated Systematic Review and Meta-Analysis. International Journal of Molecular Sciences, 23(23), 15331. https://doi.org/10.3390/ijms232315331








