1. Introduction
Colorectal cancer is one of the leading cancers globally and is particularly challenging in Western countries. In high/very high human development index (HDI) countries, the age-standardised incidence of colorectal cancer came second at 20 per 100,000 females population after breast cancer in females and third at 29.0 per 100,000 male population after prostate and lung cancers [
1]. The death rate of colorectal cancer in HDI countries ranked third in males and females alike [
1]. Collectively, this represents 1.9 milion new colorectal cases and more than 930,000 death from colorectal cancer globally. In the UK where the present study is conducted, colorectal cancer ranked the 4th most common cancer type and 2nd most common cause of cancer death [
2]. Thus, colorectal cancer (CRC) has become a heavy burden to public health. Despite a stable overall incidence rate and steadily improving survival rates in recent years, there is an increasing trend of early-onset CRC cases and challenging 5-year survival rates in more aggressive stages [
2]. As one of the major treatments of CRC apart from surgery, chemotherapy remains important in treating CRC patients and has been delivered as single drug treatment such as 5-Fluorouracil (5-FU), Oxaliplatin, Docetaxel, Irinotecan, etc. and in combinational regimes such as OLFOX (5-FU, leucovorin (LV) and Oxaliplatin), Capeox (Capecitabine with Oxaliplatin), FOLFIRI (5-FU, LV with Irinotecan), CAPIRI (Capecitabine with Irinotecan), etc. [
3]. However, therapeutic resistances to such agents are also frequently reported and are linked with worsened prognosis of the patients. Aberration in genes/translation factors and related signalling events may contribute to such resistances [
4,
5,
6] as well as the development and progression of CRC [
7,
8]. Therefore, research focused on identifying novel markers to shed light on therapeutic strategies have become more and more urgent.
Encoded by the
LIMA1 gene, Epithelial Protein Lost In Neoplasm (EPLIN) was initially discovered to be downregulated in oral cancer [
9,
10]. EPLIN has been reported to be downregulated in multiple cancer types including breast cancer [
11], prostate cancer [
12,
13,
14], lung cancer [
15] and ovarian cancer [
16]. The downregulation of EPLIN links to poor prognosis of such cancer patients [
11,
13,
15,
16]. EPLIN has also been demonstrated to regulates cellular functions such as adhesion, growth, migration and invasion, essential to the aggressive phenotype of cancer cells [
11,
12,
13,
14,
15,
17]. To achieve its cellular impacts, EPLIN can be phosphorylated by extracellular signal-regulated kinase 1/2 (ERK1/2) upon activation of platelet derived growth factor (PDGF), subsequently leading to upregulation of β-catenin and Zinc Finger E-Box Binding Homeobox 1 (ZEB1) and promote the epithelial mesenchymal transition (EMT) process [
12,
18]. Meanwhile, EPLIN can be regulated by p53 family members to lessen the invasiveness of cancer cells [
19,
20]. Moreover, EPLIN is involved in a carcinogenetic progress namely apical elimination by interacting with plectin and paxillin [
21].
Hence, EPLIN has been demonstrated to act as a potential novel tumour suppressor. In the present study, we explored the role of EPLIN in CRC in clinical cohorts and in vitro assays. Novel interacting partners of EPLIN were also investigated for their influence on clinical outcomes of CRC patients and impact on drug resistance to chemotherapeutic and targeted therapeutic agents.
3. Discussion
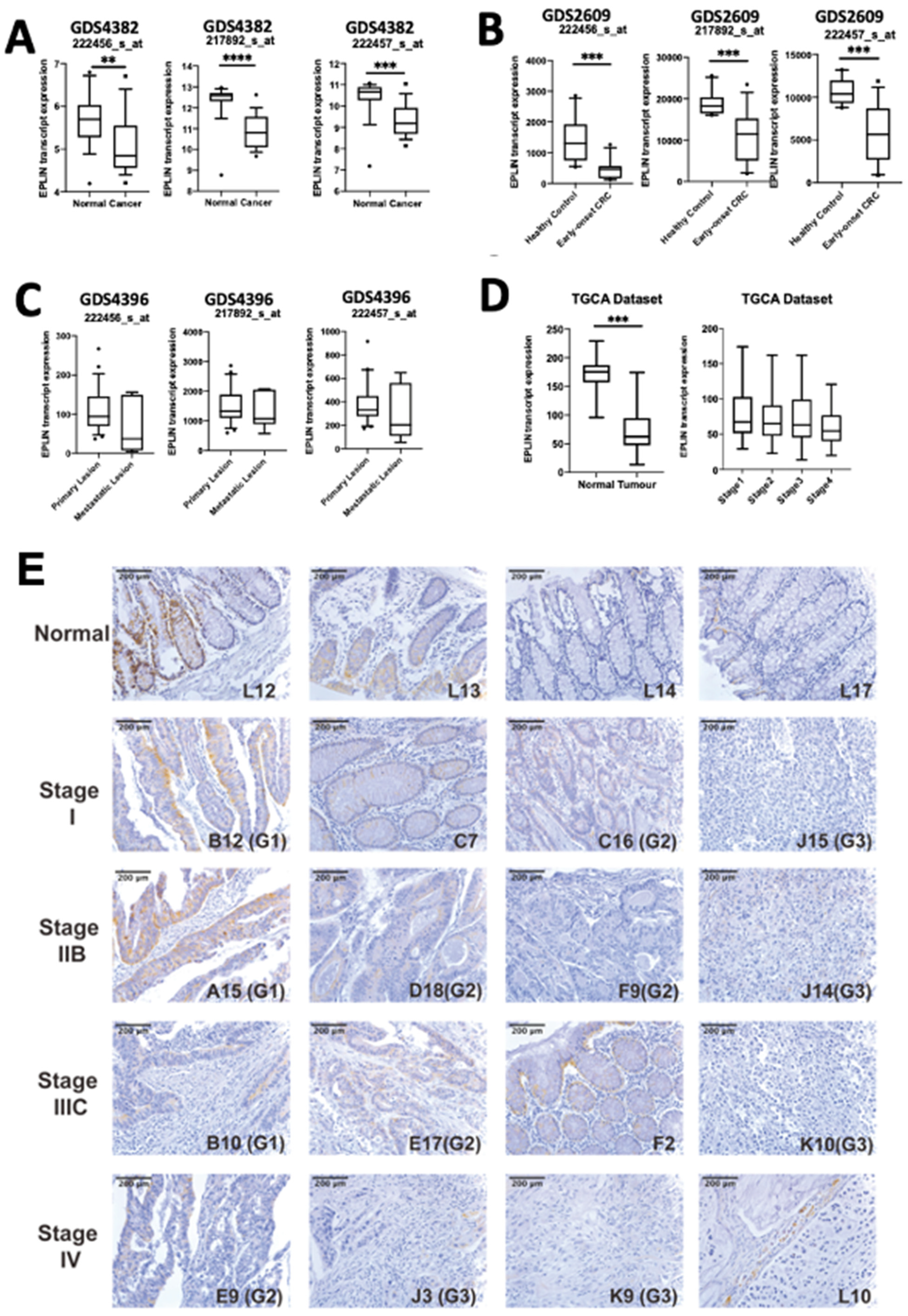
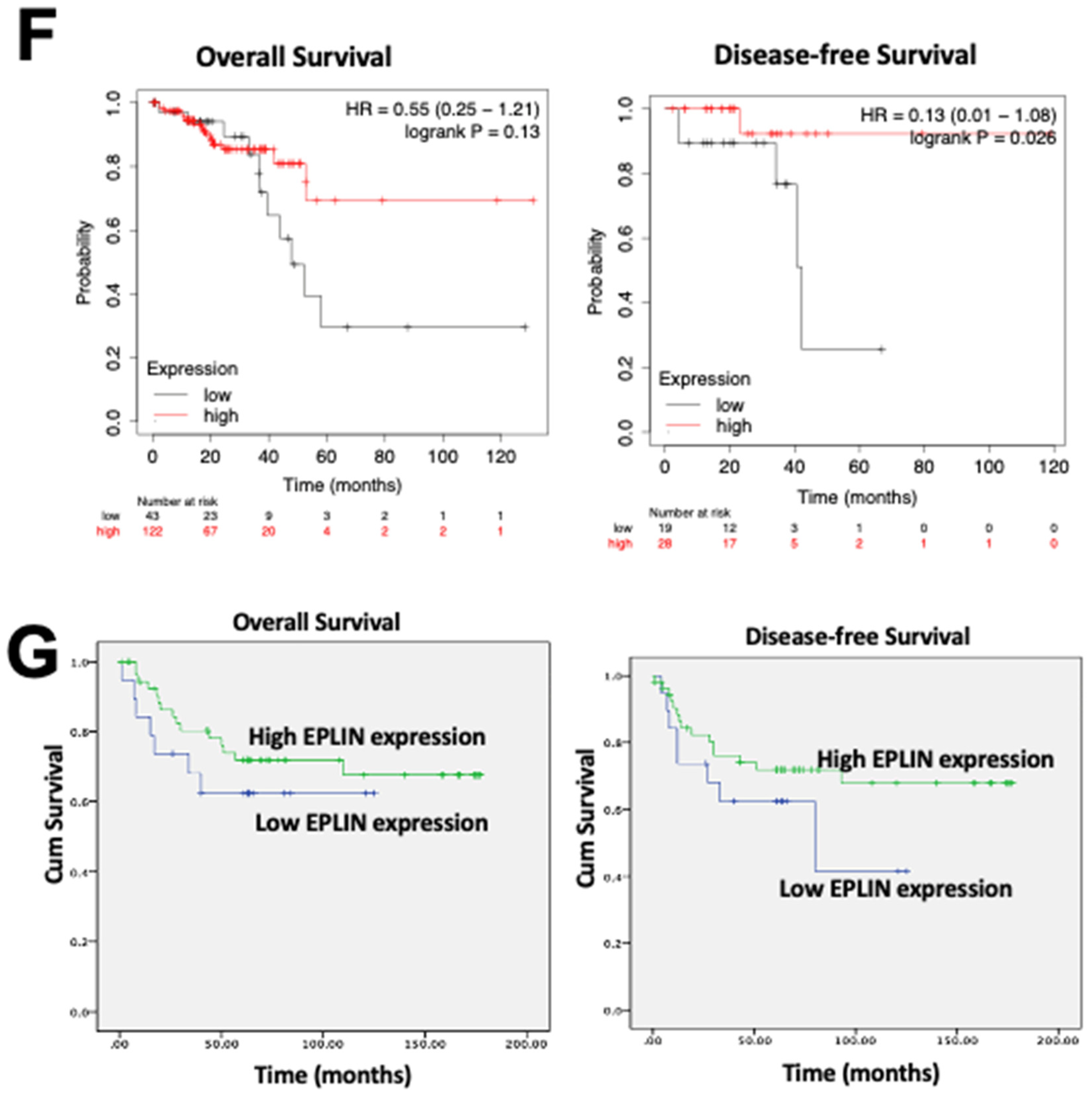
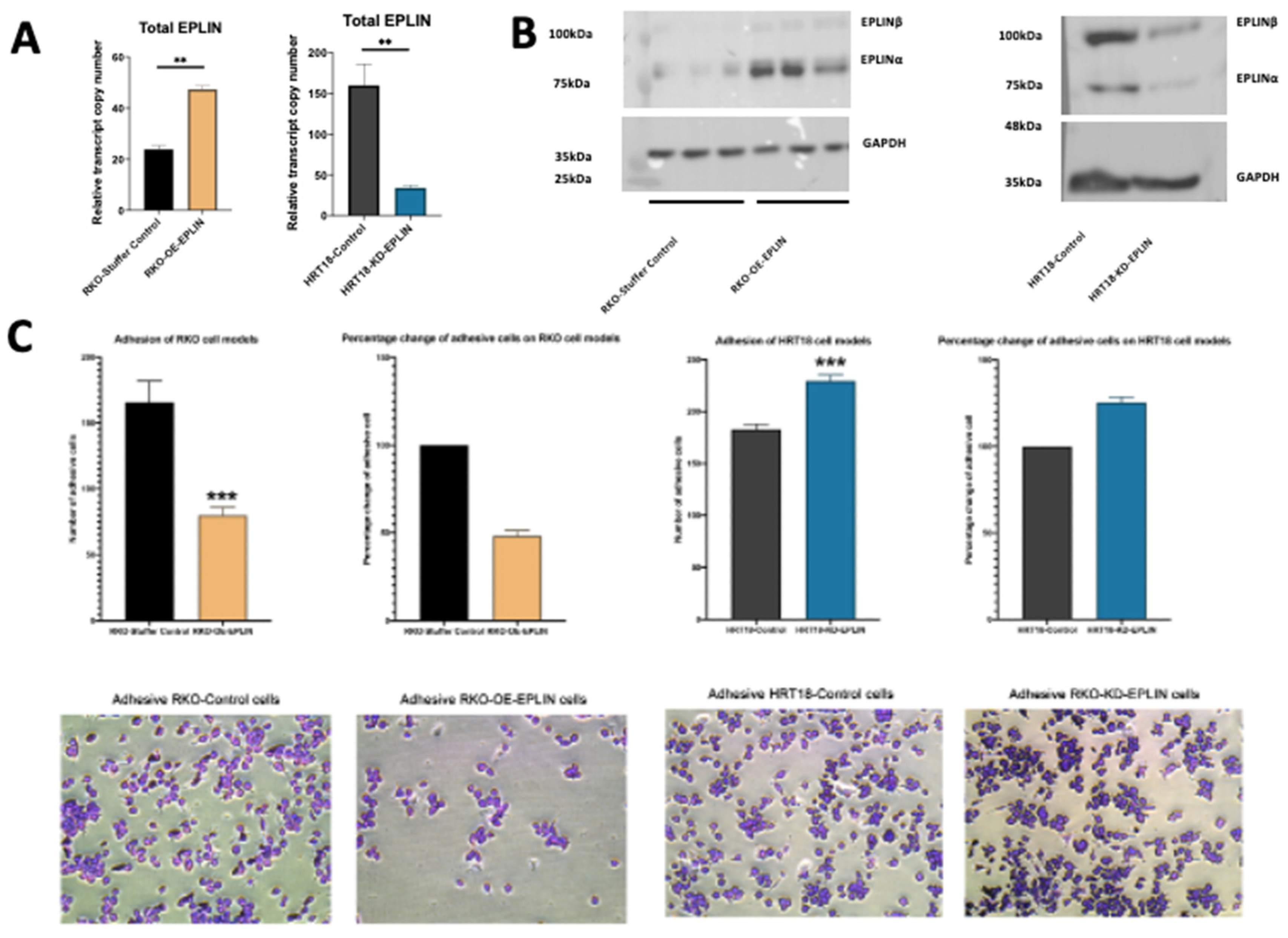
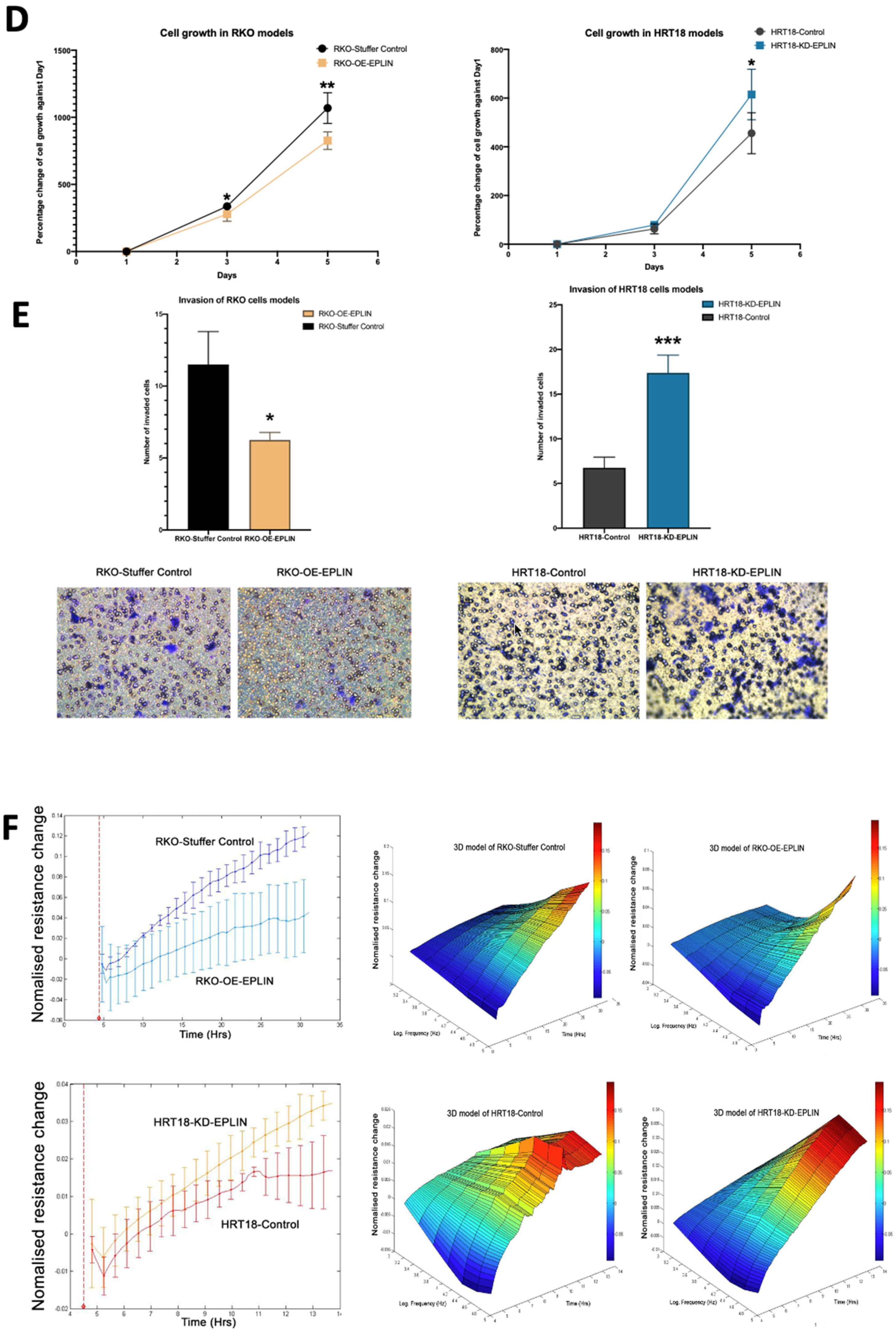
EPLIN has been implicated as a tumour suppressor and is associated with patient’s clinical outcomes as well as drug resistances in some cancer types. In the present study, we have shown that EPLIN is also an important regulator in colorectal cancer. Downregulation of EPLIN was found in colorectal cancer at transcript and protein levels and this downregulation was also related to poor OS and RFS in colorectal cancer patients. In vitro, we demonstrated that genetic modification of EPLIN expression rendered cells with less aggressive phenotypes, impacting growth, adhesion, migration and invasion negatively. These properties of EPLIN on colorectal cancer cells thus indicates itself as an important player in influencing some of the hallmarks of cancer, namely invasion and metastasis [
32,
33]. These findings collectively suggest that in colorectal cancer, EPLIN behaves as a potential tumour suppressive molecule, as suggested in a number of other cancer types [
11,
12,
13,
14,
15,
17,
18,
34].
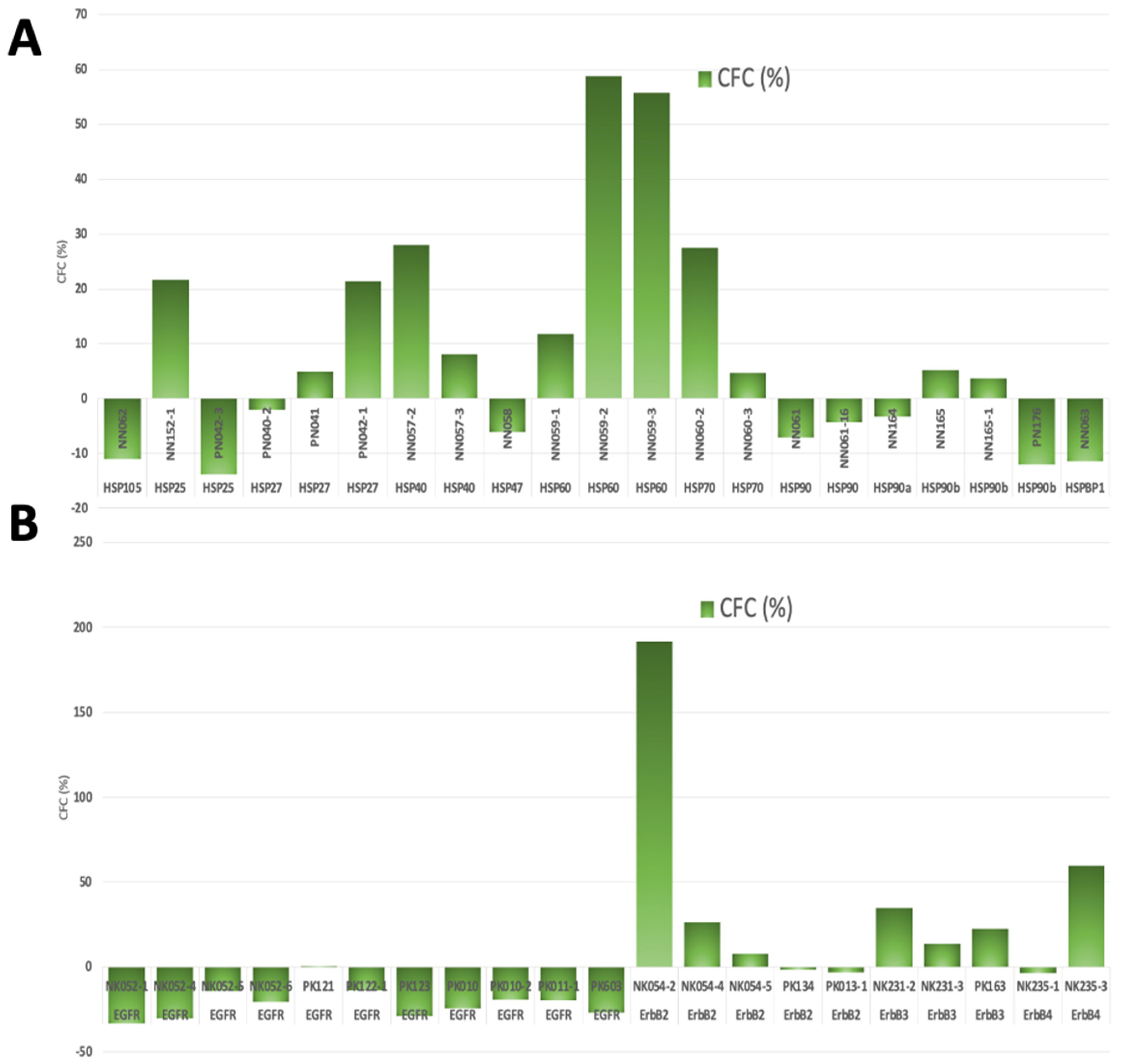
Apart from emphasizing EPLIN’s impact on clinical outcomes and cellular functions, we also search for potential mechanistic or interacting partners. Our search for potential EPLIN partners in colon cancer cells utilising Kinexus
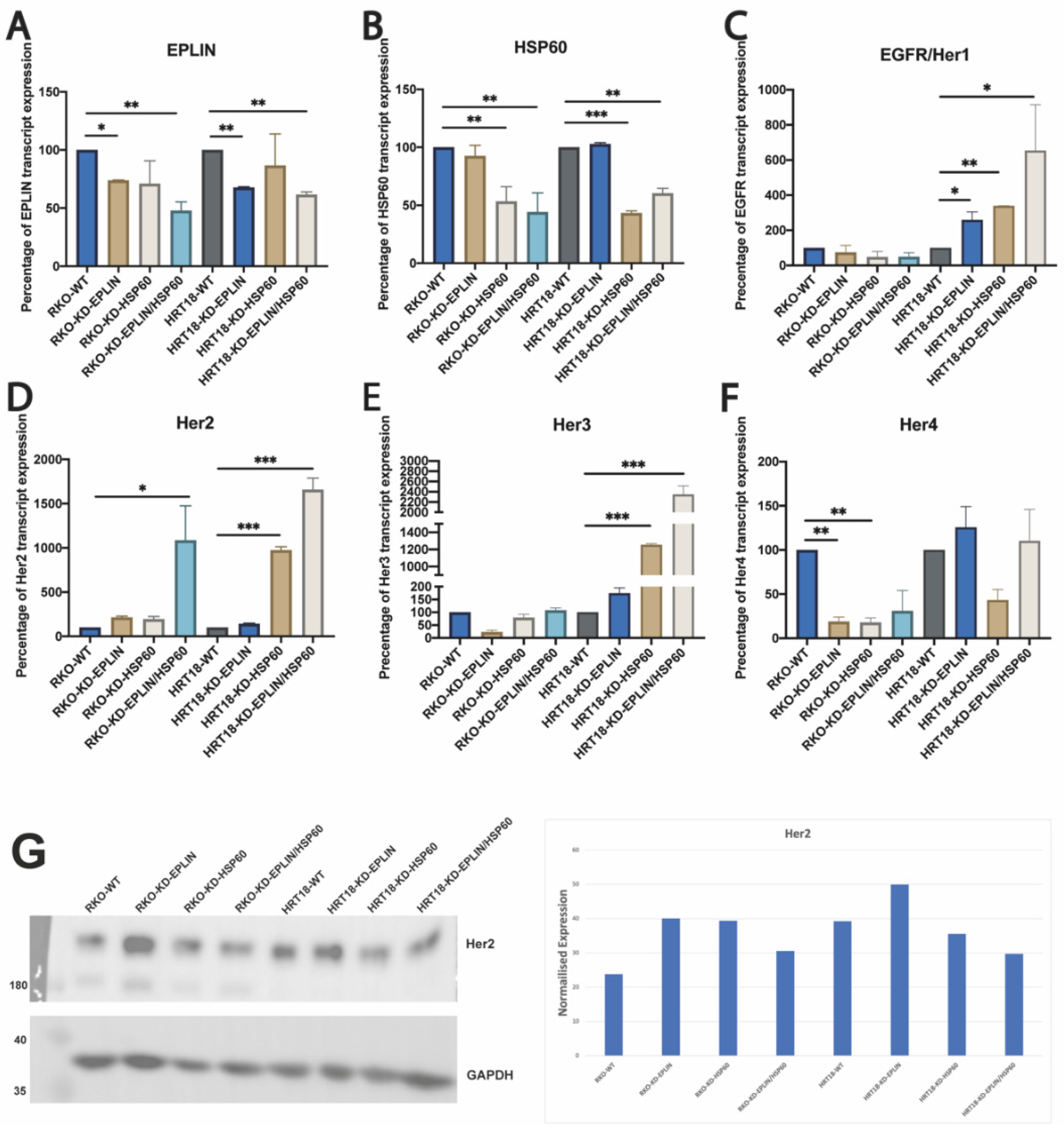
TM antibody-based protein microarray identified HSP60 and Her2 are as potential interacting partners, together with a few others related to MAPK signalling pathway. By following this path, we did not observe direct protein–protein interaction between EPLIN, HSP60 and Her2. This could be due to the limitation of in vitro study based on epithelial colorectal cancer cell lines. Proteins that were applied to the protein microarray contains not only epithelial cells, but also elements from microenvironment, such as stomal cells, stem cells, etc. Thus, this close interaction might exist in other elements but not in epithelial cells. After manipulating the expressions of EPLIN and HSP60 in RKO and HRT18 cell lines, the regulatory relationship between EPLIN, HSP60 and EGFR family members were investigated. Interestingly, in both cell lines, inhibition of EPLIN and HSP60 together resulted in a marked upregulation of transcript level of Her2. This interesting finding was supported by our analysis on clinical cohort, in that HSP60 expression was found to be upregulated in Her2 positive breast cancer and such regulation was predicted to be affected by MAPK signalling [
35]. EGFR and Her3 were only found to be regulated in HRT18 cellular models, while Her4 was found to be regulated in RKO cellular models. Such results might be due to the different genetic profiles between RKO and HRT18. For example, Her4 was mutated in HRT18 but not in RKO.
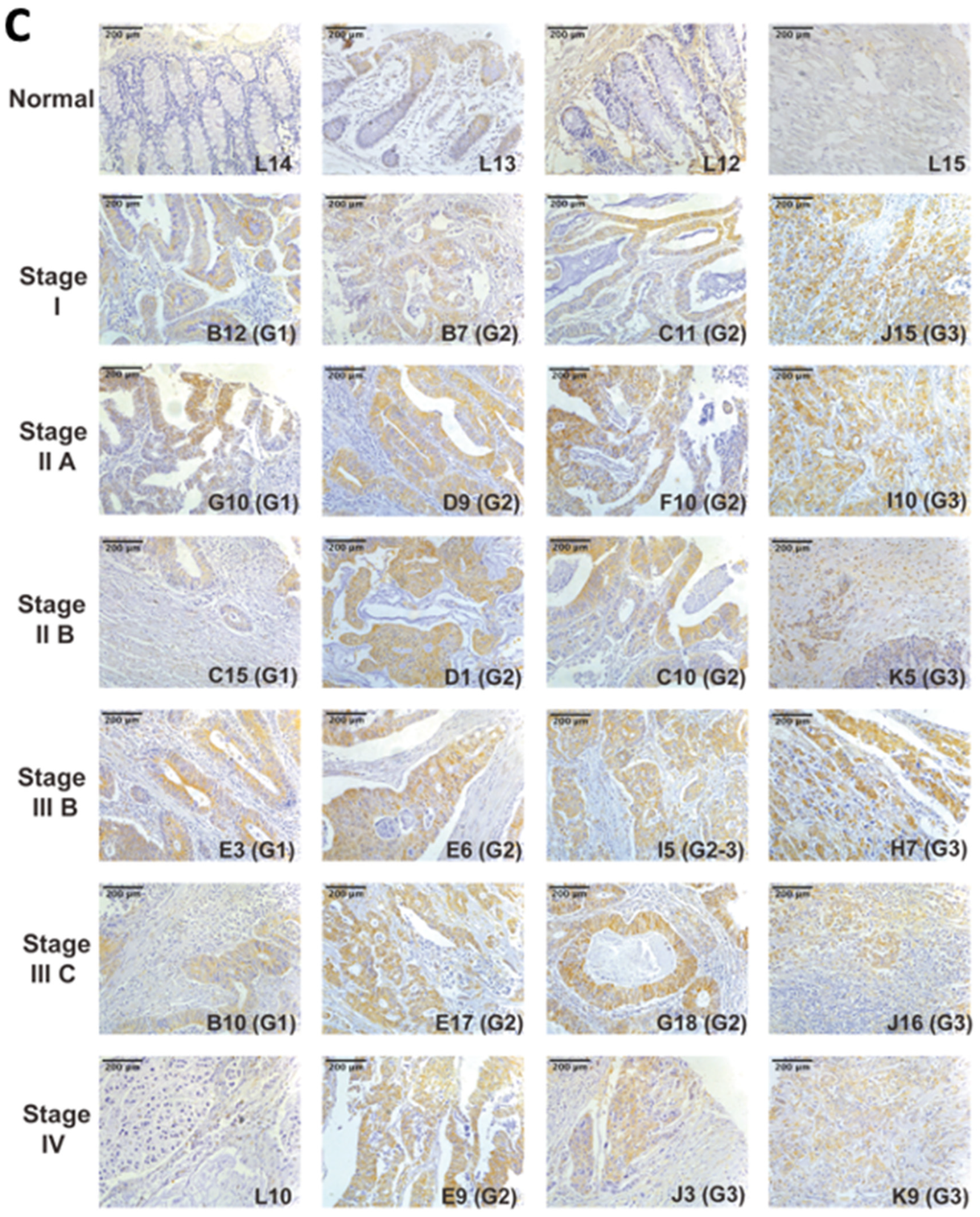
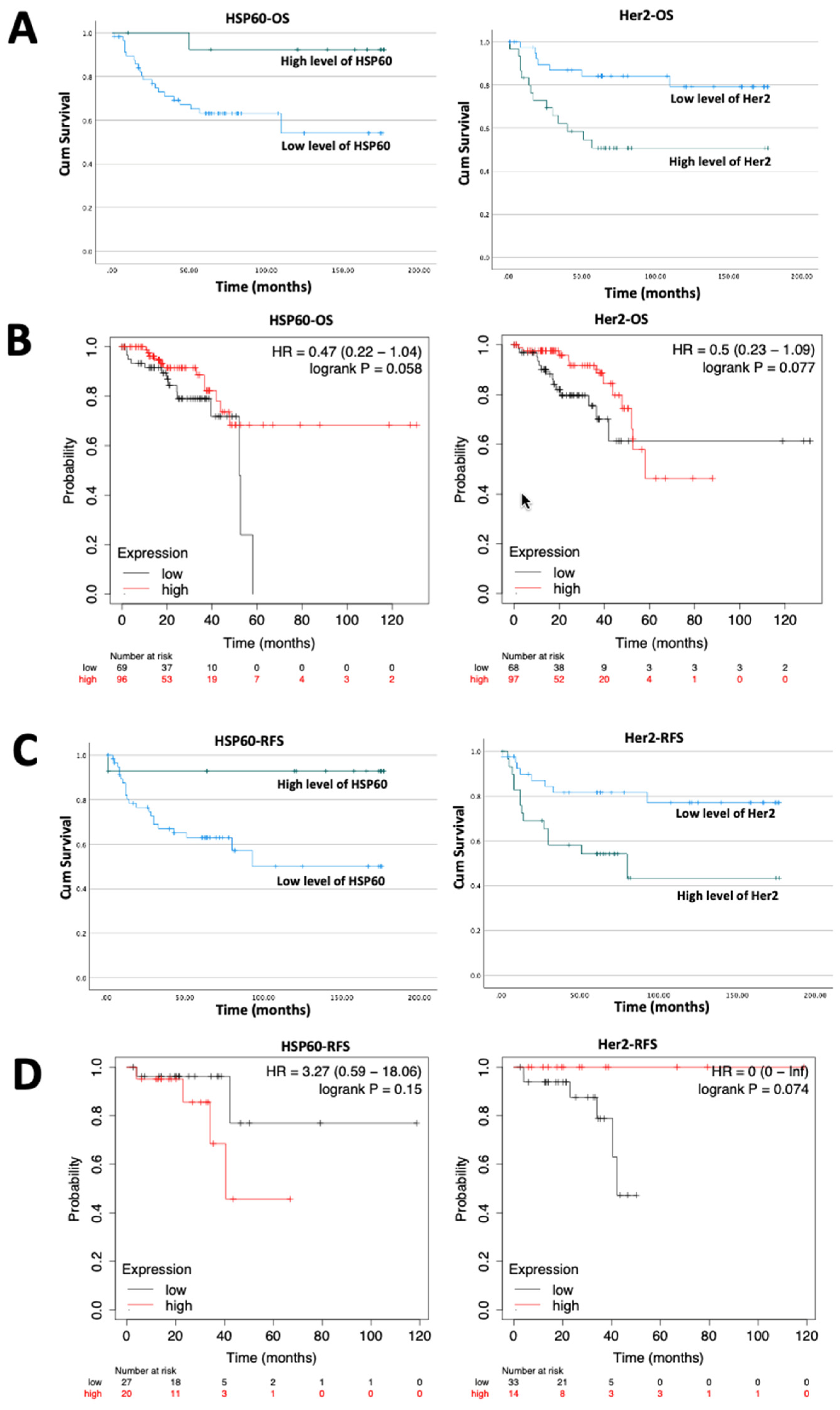
We also assessed transcript profiles of HSP60 and Her2 in clinical colorectal cancer cohorts and highlighted that high levels of HSP60 and Her2 were observed in tumour tissues compared to normal ones. However, based on our analysis of a clinical cohort and KM plotter, patients with high level of HSP60 was found to have better OS and RFS. These findings seem in conflict to our observation on transcript profile. Vocka et al. reported that high serum level of HSP60 was related to worse OS in colorectal cancer patients [
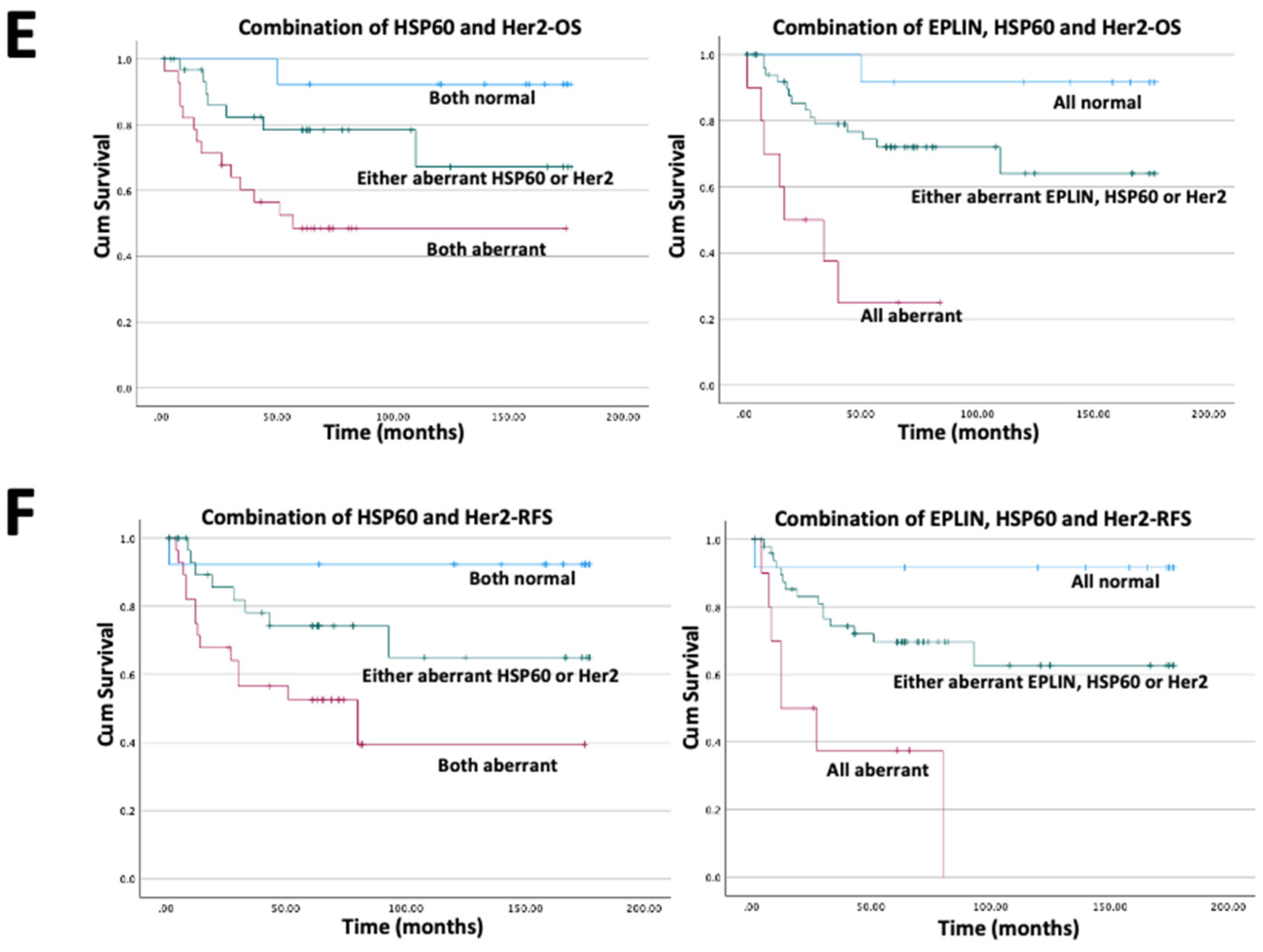
36]. Hence, a larger cohort is needed for better understanding of the clinical impact of HSP60. However, the significant finding that the combination of EPLIN, HSP60 and Her2 expression presents a significant independent prognostic indicator for the clinical outcome of the patients strongly argues that this interacting partner group has a value that need to be further explored in this cancer type and potentially in other types of cancers.
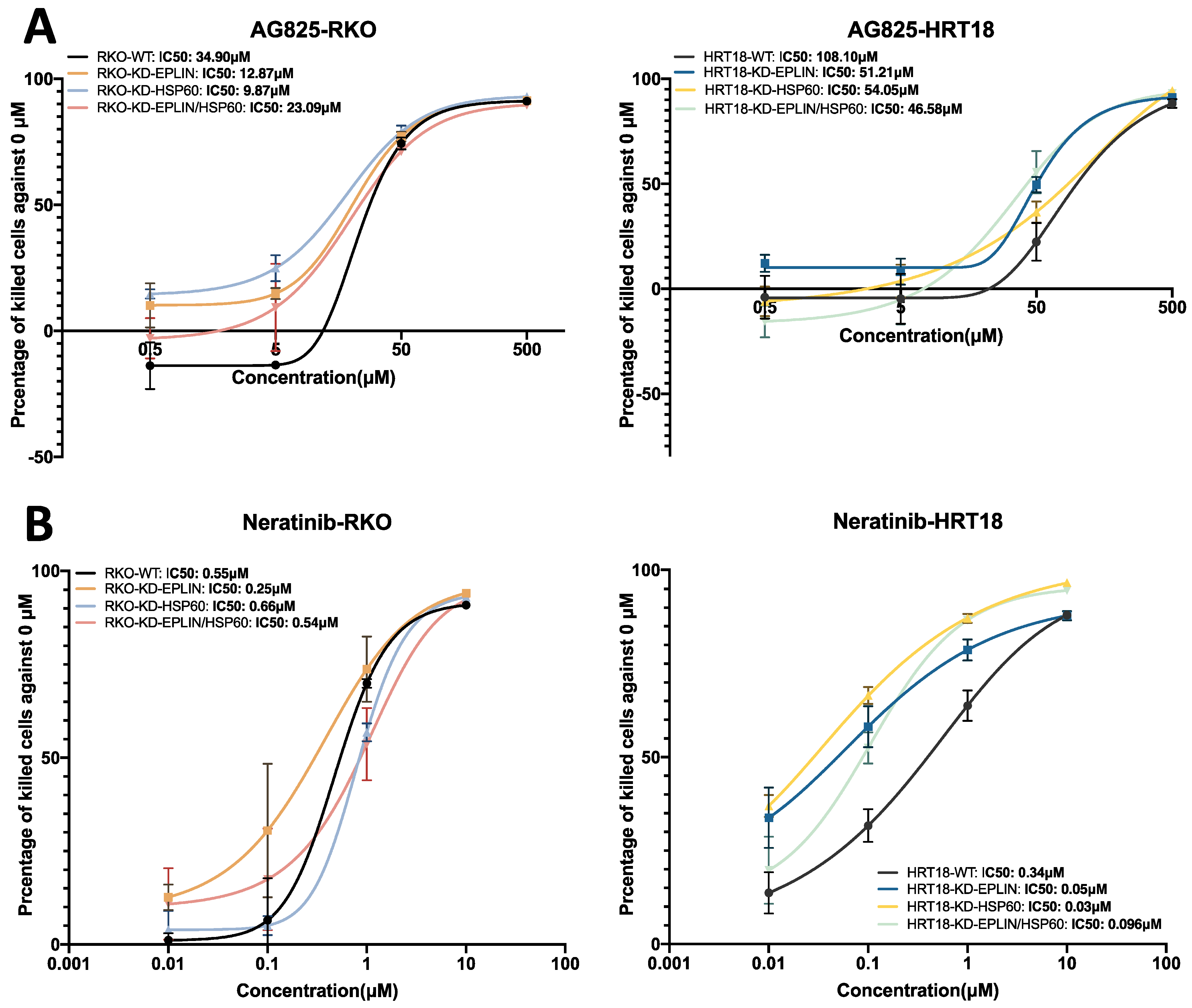
Cytotoxicity assays revealed EPLIN and HSP60 might have the potential to affect chemotherapeutic and targeted therapeutic resistances. Inhibition of both molecules in RKO and HRT18 resulted in less sensitive response to 5-FU. While more sensitive cell’s response to Docetaxel was found in HRT18 when both molecules were suppressed. Interestingly, inhibition of either of the molecules alone in HRT18 increase cell’s response to Oxaliplatin, but an opposite effect was observed when both were inhibited together. Therefore, we showed the potential of EPLIN and HSP60 in regulating cell’s response to chemotherapeutic agents. Inhibition of EPLIN and HSP60 alone or together in our cellular models resulted in more sensitive response to an HER2 inhibitor and Neratinib. This might be related to the upregulation of Her2 caused by inhibiting EPLIN and HSP60 and argues a novel direction for identifying patients with colorectal cancer who have a favourable pattern of expression of the molecular pair and may be more sensitive to target therapies to Her family.
Since its first discovery more than two decades ago [
10], there have been sustained efforts in not only determining its role in cancer progression and metastasis, but also the potential mechanism(s) played by EPLIN in cancer cells. As it has been recently summarised [
37], a small number of potential leads have been reported to be potential partners of EPLIN including p53, caveolin-1, plectin, paxillin and FAK (focal adhesion kinase). These potential partners are known players in cell adhesion and migration and cell growth. These early findings together with the discovery of the present study contribute to the unveiling of how EPLIN interplays in the complex networking within the ‘blackbox’ of the cells and its networking. However, it is also clear that there is a long way to go to fully clarify the signalling events up and down stream of EPLIN, an exciting prospect to further expand this area, together with the understanding of its role of a broad sense of clinical cancers.
In conclusion, EPLIN appears to act as a tumour suppressor in clinical colorectal cancer and regulates cellular growth, invasion, migration and adhesion negatively in colorectal cancer cells. Her2, HSP60 and their related signalling events (i.e.,MAPK) are potential novel interacting partners of EPLIN. Aberrant expression of EPLIN, HSP60 and Her2 is identified as an independent predictor for OS and RFS. EPLIN and HSP60 have the potential to regulate cell’s response to chemotherapeutic agents. While inhibition of both molecules leads to upregulation of Her2 and more sensitive cell’s response to Her2 inhibitor and Neratinib. These findings warrant more intensive studies on EPLIN both in clinical and therapeutics in colorectal cancer.
4. Materials and Methods
4.1. Collection and Processing of Colorectal Cancer Tissues
A clinical cohort of colorectal cancer containing 94 colorectal cancer tissues from the patients. Of these 94 samples, we were able to obtain 80 normal matched colorectal cancer samples from the same patients. The normal colon tissues were obtained from the same patients with at least 10 cm away from tumour margins. Both tumour and normal tissues were used in the present study. Tissues were collected in the University Hospital of Wales after surgery and consenting the patients. Tissue samples were examined by a consultant pathologist and stored at −80 °C until use. Pathological information was obtained via hospital clinical portal system and CANISC data. Tissues were used under ethical approval by East Wales Local Research Ethics Committee (reference number SJT/C617/08).
4.2. Cell Culture
Two colorectal cancer cell lines were used for this study, RKO (CRL-2577, poorly differentiated colorectal carcinoma cell line derived from colon and with no known P53 mutation) and HRT18 (ECACC 86040306, an epithelial adherent cell line derived from large intestine. Both cell lines were purchased from American Type Culture Collection (ATCC, Rockville, MD, USA) (LGC Standard, ATCC UK agent). Cells were cultured at low passages at 37 °C with 95% humidity and 5% CO2 in either Dulbecco’s Modified Eagle’s medium (DMEM) or RPMI-1640 medium with L-glutamine and sodium, which were supplement with 10% heat inactivated foetal calf serum (FCS) and 1% antibiotics mixture (Sigma-Aldrich, Pooled, Dorset, UK).
4.3. RNA Extraction and Reverse Transcription
TRI Reagent Kit (Sigma-Aldrich, Poole, Dorset, UK) was used to extract RNA from tissues samples and cells from colorectal cancer cell lines following manufacture’s instruction. After isolation, RNA samples were quantified to 500 ng/μL and used to performed reverse transcription in a Simpliamp thermocycler (Fisher Scientific UK Leicestershire, UK), using a GoScriptTM Reverse Transcription System Kit (Promega Corporation, Madison, WI, USA). cDNA samples were then stored at −20 °C until required.
4.4. Real Time Quantitative PCR (qPCR)
qPCR was carried out to assess transcript expression of genes of interest. Amplifilour Uniprimer
TM Universal system (Intergen company, New York, USA) was utilised for qPCR [
29]. In brief, each reaction was made up with 5 μL precision FAST2x qPCR Master Mix (PrimerDesign, Southampton, UK), 0.3 μL forward primers (EPLIN: AAGCAAAAATGAAAACGAAG; GAPDH: AAGGTCATCCATGACAACTT; Her1: GACCTCCATGCCTTTGAGAA; Her2: CCTCCTCGCCCTCTTG; Her3: CCCCACACCAAGTATCAGTA; Her4: CTGCTGAGTTTTCAAGGATG; HSP60: TGTAGACCTTTTAGCCGATG), 0.3 μL reverse primer with z sequence whose concentration was 1/10 of forward primer (EPLIN: ACTGAACCTGACCGTACAGACACCCACCTTAGCAATAG; GAPDH: ACTGAACCTGACCGTACAGCCATCCACAGTCTTCTG; Her1: ACTGAACCTGACCGTACAGCACAAATTTTTGTTTCCTGA; Her2: ACTGAACCTGACCGTACACATGTCCAGGTGGGTCT; Her3: ACTGAACCTGACCGTACAACACAGGATGTTTGATCCAC; Her4: ACTGAACCTGACCGTACAAACTTGCTGTCATTTGGACT; HSP60: ACTGAACCTGACCGTACAACAGTCACACCATCTTTTGT) (Sigma-Aldrich Co, Poole, Dorset, UK), 0.3 μL uniprimer and 4.1 μL cDNA samples. In addition to the test cDNA samples, a set of known transcript copy number samples (ranging from 10
1 to 10
8) was also run as standard relative copy number of cDNA samples were calculated based on this standard and normalised to the housekeeping control, GAPDH.
4.5. Preparation of Protein Samples
Protein samples were collected from colorectal cancer patients or from cell lines. In brief, colorectal cancer cell lines were harvested with PBS once they reached desired confluences (<90%). After centrifuging, supernatant was discarded and pellet was resuspended with home-made modified RIPA lysis buffer. Samples were put on a rotational platform and incubated overnight at 4 °C before being centrifuged at 4 °C to form pellets. Pellets was then discarded and samples processed and quantified. The concentration of samples was tested using a BioRadDC Protein Assay (BioRad Laboratories, Hertfordshire, UK) in accordance with the manufacturer’s instructions. Samples were quantified to at least 2 μg/μL with lysis buffer and were stored at −20 °C before use.
4.6. Western Blotting
Western blotting was performed to detect expression of protein samples. Briefly, protein samples were loaded on a Sodium dodecyl sulfate-polyacrylamide (SDS-PAGE) gel. The gel was subject to electrophoresis to separate proteins based on mass at 120 V, 50 W and 50 mA until sufficient separation was obtained. The gel was then transferred to an Immobilon-P PVDF membrane (Merck Millipore, Hertfordshire, UK) to perform semi-dry transfer via a semi-dry blotter at 15 V, 500 mA, 20 W for 50 min. Immunoblotting was then carried out as follows: The membrane was blocked with 5% milk mixture which was diluted in tris buffered saline (TBS) (Sigma-Aldrich Co, Poole, Dorset, UK) with 0.1% tween-20 (Melford Laboratories Ltd., Suffolk, UK) for an hour before incubating with desired primary antibody (anti-EPLIN: monoclonal, mouse, dilution 1:500; sc-136339; anti-GAPDH: monoclonal, mouse, dilution 1:1000, sc-32233; anti-Her2, mouse monoclonal, SC-33684 dilution 1:500; purchased from Santa Cruz Biotechnology, Inc. Dallas, Texas, USA) (diluted with 2.5% mixture) overnight at 4 °C. The membrane was washed in 2.5% milk mixture three times for 15 min at room temperature then was incubated with secondary antibody, mouse (whole molecule) IgG peroxidise conjugate (Sigma-Aldrich Co, Poole, Dorset, UK), which was diluted 1000 times in 2.5 milk mixture, for an hour at room temperature. After incubation, membrane was washed with TBS-T and TBS twice for 10 min. EZ-ECL solution (equal parts of solution A mix with solution B) (Geneflow Ltd., Litchfield, UK) was used to incubate with membrane in the dark before capturing images on a G-BOX (Syngene, Cambridge, UK) detection system.
4.7. Immunohistochemical Staining and Analysis
Immunohistochemical staining was performed on the tissue microarray (TMA) slides from colorectal cancer (
https://www.biomax.us/tissue-arrays/Colon/CO2161a, accessed on 16 July 2022; code: CO2161a; US Biomax, Inc., Derwood, MD, USA) to detect the intensity and distribution of EPLIN and HSP60. In brief, IHC assay was carried out as follows. After rehydration and washing, slides were blocked with 10% horse serum for an hour, then were incubated with primary antibodies (EPLIN: monoclonal, mouse, 1:500, sc-136339; HSP60: monoclonal, mouse, 1:500, Santa Cruz Biotechnology, Inc. Dallas, TX, USA) overnight. Secondary antibody was applied after stringent washing followed by incubating with avidin-biotin complex (ABC) reagent (Vector Laboratories, Inc., Newark, CA, USA) and 3′3 diaminobenzidine (DAB) substrate (5 mg/mL) to develop staining. Counterstaining was performed with Gill’s haematoxylin (Vector Laboratories Inc., CA, USA) and rehydrated through a series of graded alcohols and cleared in xylene (Fisher Scientific, Leicestershire, UK). Analysis of IHC staining was performed by scoring intensity and distribution of targeted proteins by two researchers.
4.8. Exploration of Rectum Adenocarcinoma Datasets in Kaplan–Meier Plotter
An online platform, Kaplan–Meier Plotter was carried out to investigate implication of EPLIN, Her2 and HSP60 on OS and RFS of rectum adenocarcinoma patients. Data and graphs were assessed from
https://kmplot.com/analysis/index.php?p = service&cancer = pancancer_rnaseq (accessed on 1 October 2022).
4.9. Generation of EPLINα Overexpression Cell Lines Using Electroporation-Based Transfection
Overexpressed EPLIN models were generated by using a plasmid DNA which contains the open reading frame of a sequence of EPLINα or Stuffer300 control and a puromycin resistant sequence (VectorBuilder Inc., Chicago, IL, USA) via electroporation-based transfection. In brief, 5 μL of plasmid DNA was mixed with 1 mL of cell mixture containing 1 × 106 of cells with antibiotic-free medium in an electroporation cuvette (Geneflow Ltd., Litchfield, UK). After incubating for 5 min, the solution was applied to the BioRad Cell Pulser Xcell electroporation system (BioRad Laboratories, Hertfordshire, UK) and was pulsed at 290 V and 1000 μF. Cells went through selection with culture medium that contains 2 μg/mL puromycin for 72 h and were subsequently kept culturing in medium that contains 0.2 μg/mL puromycin.
4.10. Generation EPLIN Knockdown Cell Lines Using shRNA-Based Transfection
EPLIN shRNA plasmid (sc-60593-SH) was used to knock down expression of EPLIN in cell lines from colorectal cancer with shRNA transfection reagent (Santa Cruz Biotechnology, Inc., Dallas, TX, USA), following instruction from the manufacturer. After selecting with medium that contains 2 μg/mL puromycin, cells were cultured in medium that contains 0.2 μg/mL puromycin to maintain the effect of transfection.
4.11. Generation HSP60 Knockdown Cell Lines Using siRNA-Based Transfection
HSP60 siRNA (sc-29351) was utilised to silence expression of HSP60 in cell lines from colorectal cancer using siRNA transfection reagent (Santa Cruz Biotechnology, Inc., Dallas, TX, USA). The transfection was performed following instruction from the manufacturer.
4.12. Thiazolyl Blue Tetrazolium Bromide (MTT) Based Cellular Growth Assay
MTT based cellular growth assays were performed to investigate EPLIN’s impact on cellular growth. In brief, 3 × 103 cells from each cell models were seeded onto three 96-well plates at triplicate, then were incubated at 37 °C with 5% CO2. At Day 1, Day 3 and Day 5, each well of the 96-well plate was supplemented with 22 μL of 5 mg/mL MTT solution (Sigma- Aldrich Co., Poole, Dorset, UK) and was incubated for 4 h at 37 °C with 5% CO2. After incubation, medium was aspirated and 100 μL of Dimethyl sulfoxide (DMSO) (Sigma-Aldrich Co., Poole, Dorset, UK) was added into each well. The plate was allowed to be incubated for 10 min at 37 °C with 5% CO2. Absorbance was detected in a LT4500 plate reader (Wolf Laboratories, York, UK) at 540 nm.
4.13. Matrigel Adhesion Assay
Matrigel adhesion assay was performed to investigate cellular adhesive function. In brief, Matrigel (BD Biosciences, Oxford, UK) was diluted in serum-free medium (SFM) to the concentration of 0.05 mg/mL. One well of 96-well was coated with 5 μg of Matrigel solution and was allowed to dry in oven at 55 °C before use. After rehydration for 30 min with SFM, each precoated well was seeded by 4 × 104 cells from each cell models at 6 repeats, then was incubated for 40 min at 37 °C with 5% CO2. Medium was discarded and each well of the 96-well plate was fixed with 100 μL of 4% formalin solution for 10 min followed by staining with 100 μL of 0.5% crystal violet solution for 10 min. Once the plate was dry. Photos of 4 random areas of each well were taken under a Leica DM IRB microscope (X20) with the Leica LAS EZ system (Leica Microsystems (UK) Ltd., England, UK). Image J (National Institute of Mental Health, Bethesda, Maryland, USA) was utilised to count cells and statistical analysis was performed using GraphPad.
4.14. Matrigel Invasion Assay
Matrigel invasion assay was carried out to examine cellular invasion. In brief, Matrigel was diluted with SFM to the concentration of 0.5 mg/mL. Then, 50 μg was used to coat a trans-well insert which was placed in a 24-well plate and contains 8.0μm pores. After the inserts were dry and rehydrated with SFM for 30 min. 2.5 × 104 cells from each model were seed into the upper chamber and were incubated for 72 h at 37 °C with 5% CO2. Medium was then discarded carefully before invaded cells at the underside of the insert was fixed with 4% formalin solution for 10 min. Cells were then stained with 0.5% crystal violet solution for 10 min and were allowed to dry at room temperature. 4 random fields were chosen under a Leica DM IRB microscope (X20) and photos of such areas were taken by the Leica LAS EZ system. ImageJ was applied to count invaded stained cells and statistical analysis was performed using GraphPad.
4.15. Electric Cell-Substrate Impedance Sensing (ECIS) Based Cell Migration Assay
ECIS assay was used to measure impedance after wounding to investigate cell’s ability to migrate across the wound. 20 × 103 cells from each model were seeded into the 96-well ECIS W961E electrode arrays in 5–6 repeats. The arrays were then placed on the placed on the ECIS Ztheta instrument (Applied Biophysics Ltd., Troy, NJ, USA). Wounding (2000 mA for 20 s) was initiated after incubating for 5 h at 37 °C. Impedance was recorded from 1000 to 64 × 103 Hz by the ECIS system for 10–24 h. Data was analysed using the ECIS software.
4.16. KinexusTM Antibody-Based Protein Microarray
KinexTM KAM-880 antibody-based protein microarray (Kinexus Bioinformatics Ltd., Vancouver, BC, Canada) was performed on two pairs of patient’s protein (normal: ID126 & 128; tumour: ID127 & 129) to investigate potential interacting partners. In brief, protein samples were extracted and quantified before they were precipitated with EPLIN antibody (immunoglobulin G (IgG) monoclonal, Sc- 136399, Santa Cruz Biotechnology, Inc. Dallas, TX, USA). Samples were then sent to Kinexus Bioinformatics Ltd., and applied on the arrays to test with 877 antibodies.
4.17. Cytotoxicity Assays
Cytotoxicity assays were utilised to investigate implication on drug resistance. In brief, 96-well plates were precoated with 100 μL of serial diluted chemotherapeutic or EGFR/Her2 targeted therapeutic agents. 100 μL cell solution which contained 5 × 103 cells from each cell models were seeded into the 96-well plate in 3 repeats for each concentration. After incubating for 72 h at 37 °C with 5% CO2, medium was discarded and the plates were fixed with 100 μL of 4% formalin solution for 10 min. Cells were then stained with 100 μL of 0.5% crystal violet solution, washed and were allowed to dry at room temperature. After the plates were dry, 100 μL of 10% acetic acid was added into each well and incubated for 10 min to extract crystal violet stain. Absorbance was read in a LT4500 plate reader at 595 nm.
4.18. Statistical Analysis
GraphPad (Prism 8) (GraphPad Software, San Diego, CA, USA), Minitab (Minitab Ltd. Coventry, UK) and SPSS version 26 (IBM, Armonk, New York, NY, USA) were used for statistical analysis in this study.