Genome-Wide Identification of AP2/ERF Superfamily Genes in Juglans mandshurica and Expression Analysis under Cold Stress
Abstract
:1. Introduction
2. Results
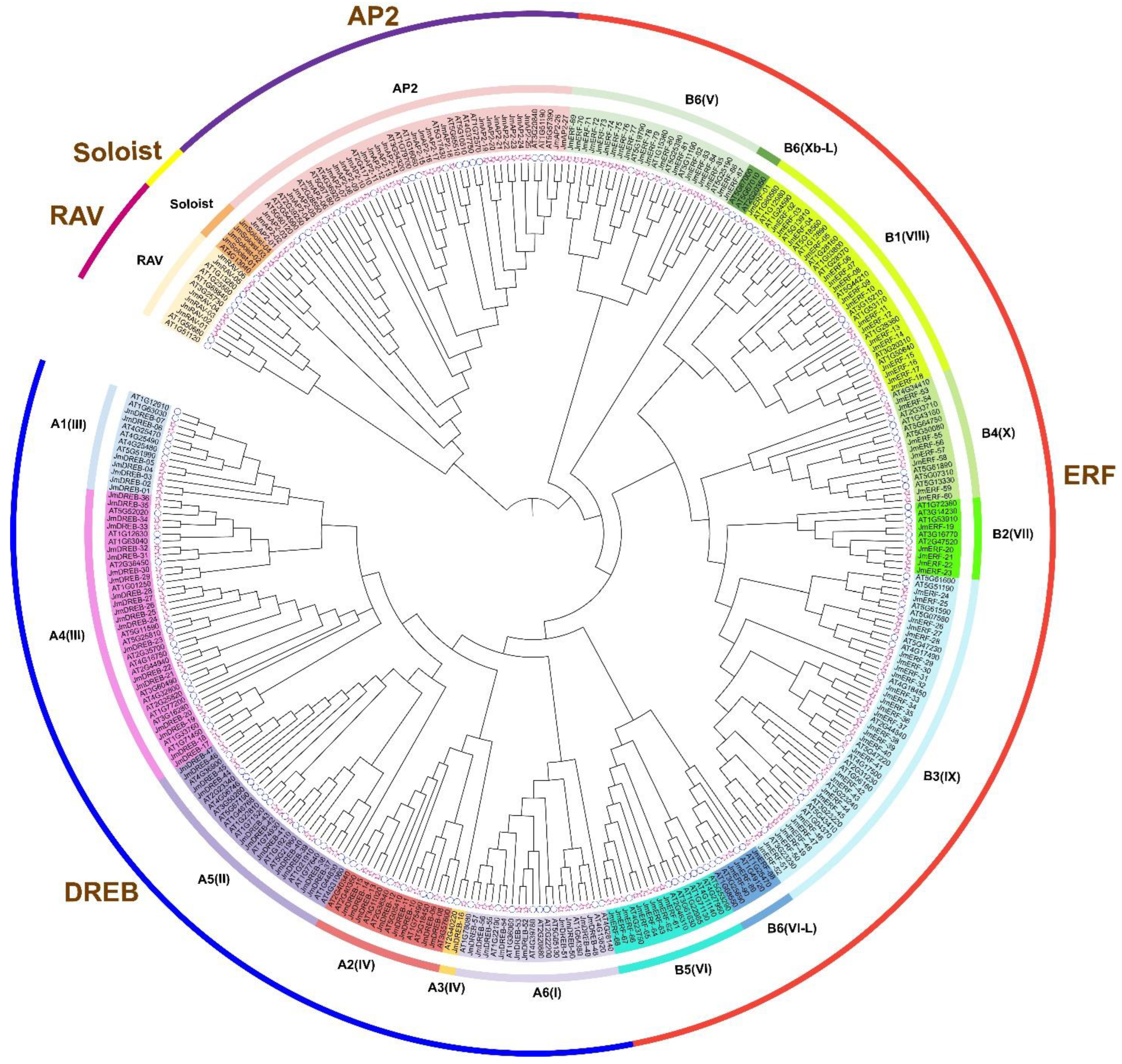
2.1. Identification and Phylogenetic Analysis of JmAP2/ERF Genes
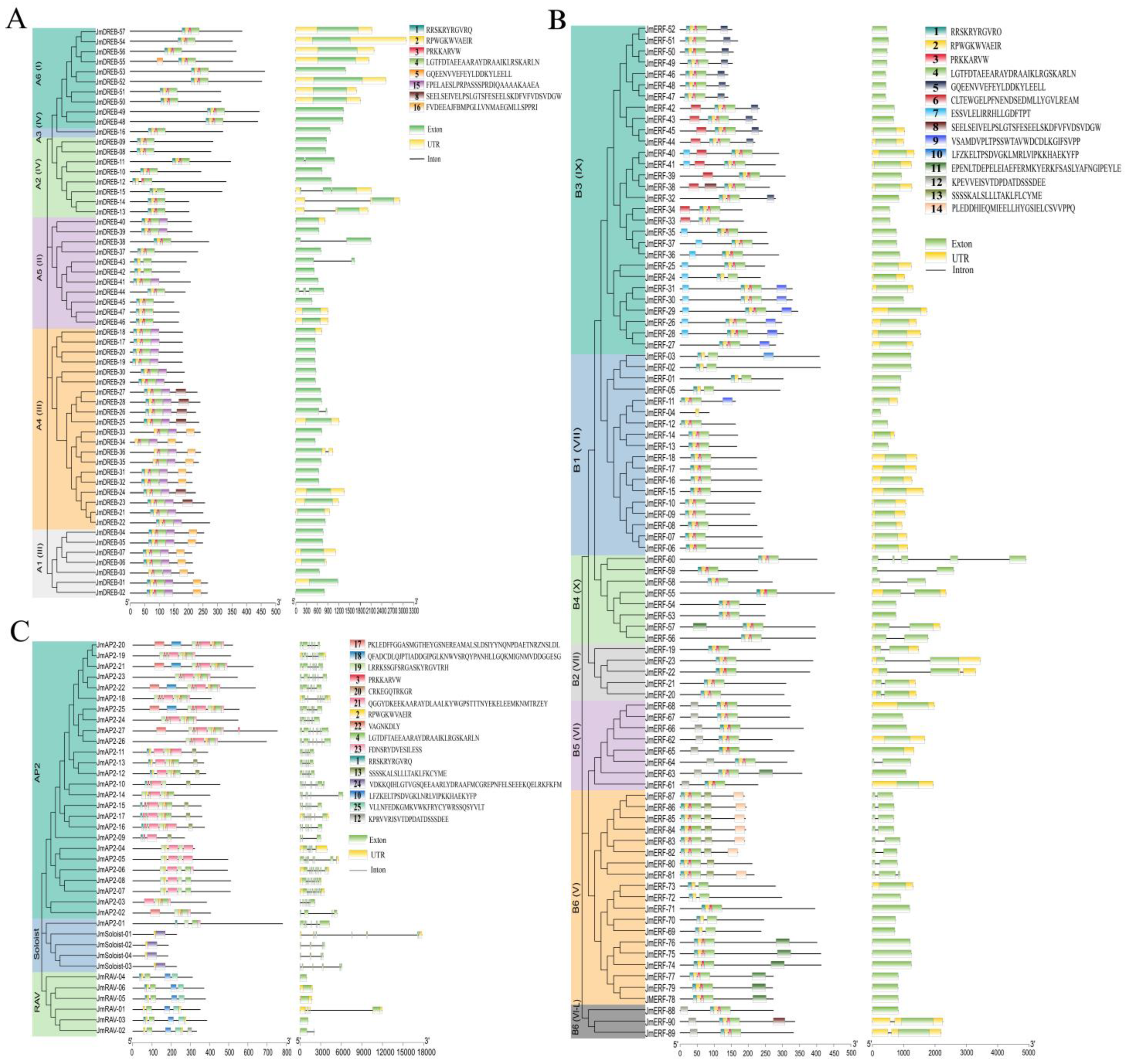
2.2. Gene Structure and Conserved Motif Analysis of JmAP2/ERFs
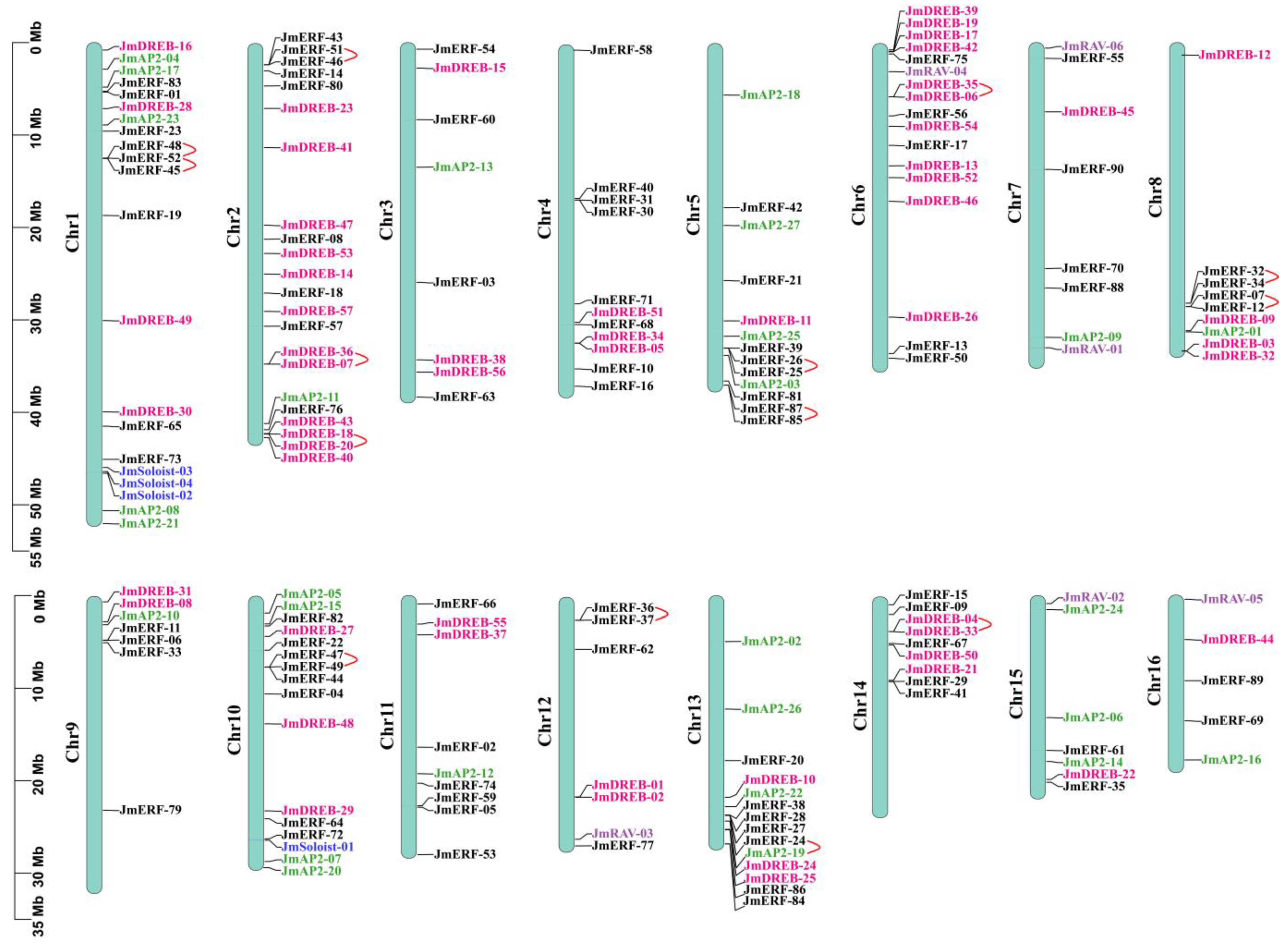
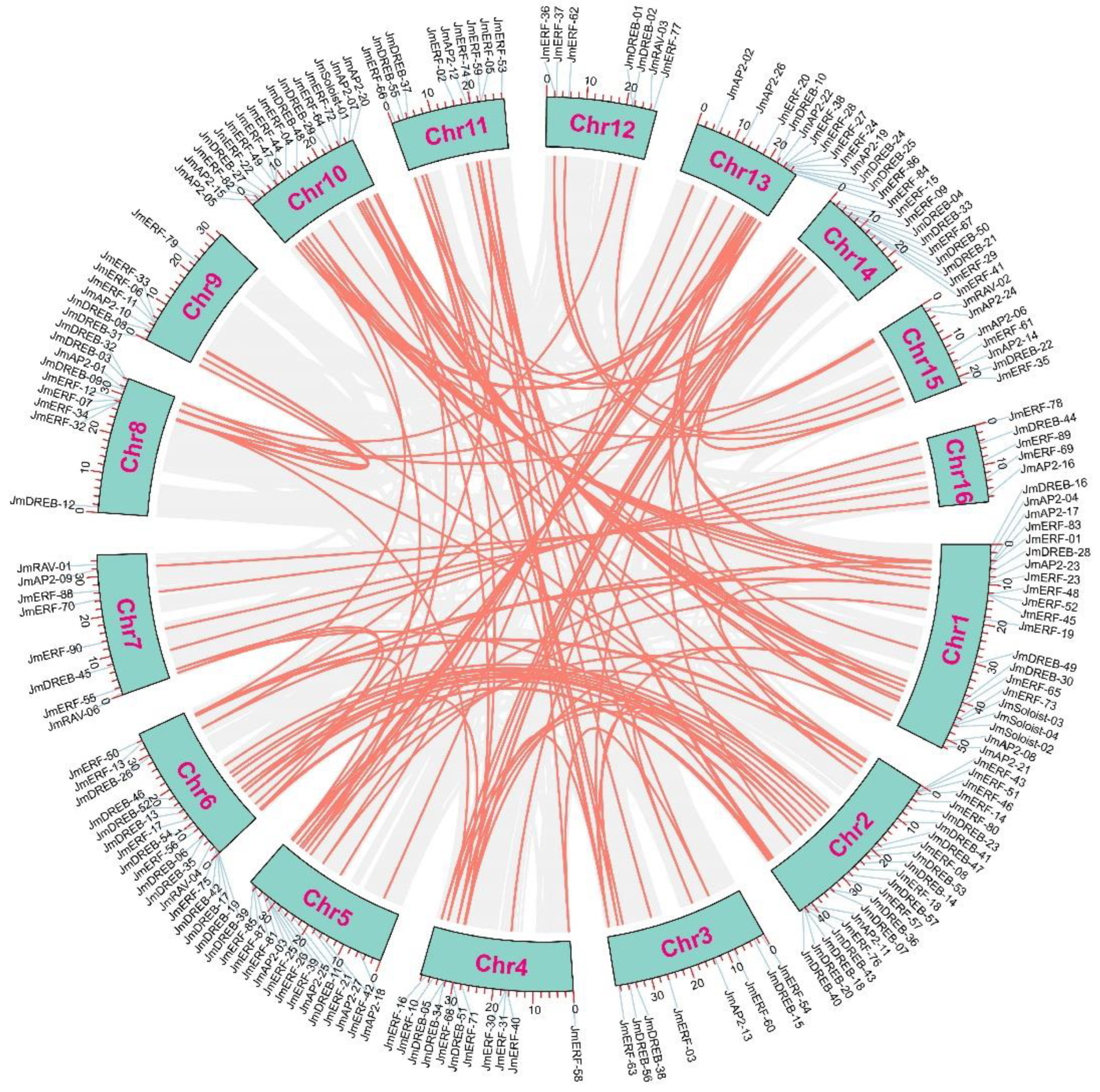
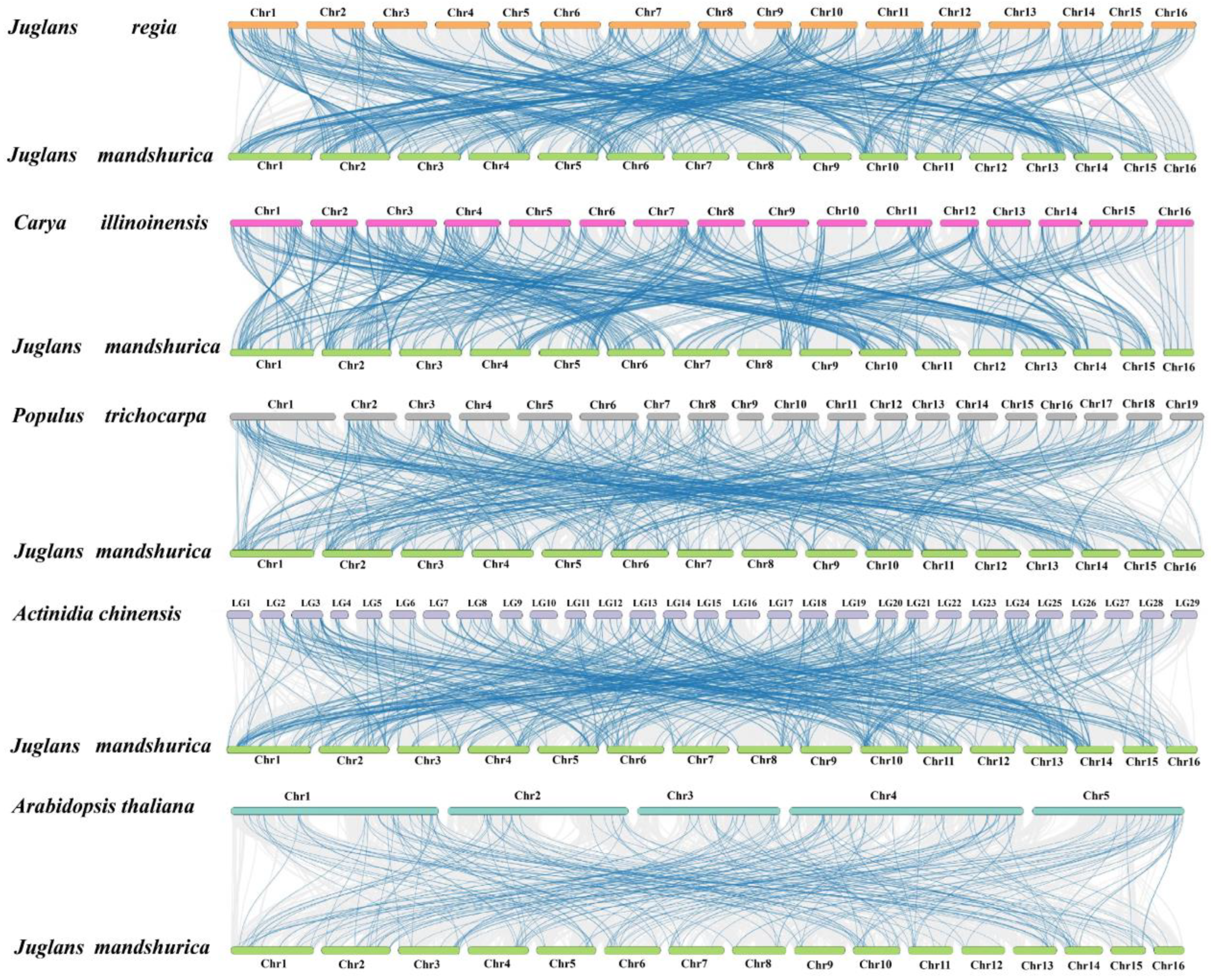
2.3. Chromosome Distribution, Duplication Events and Synteny Analysis of JmAP2/ERFs
2.4. Analysis of Cis-Acting Elements in JmAP2/ERF Promoters
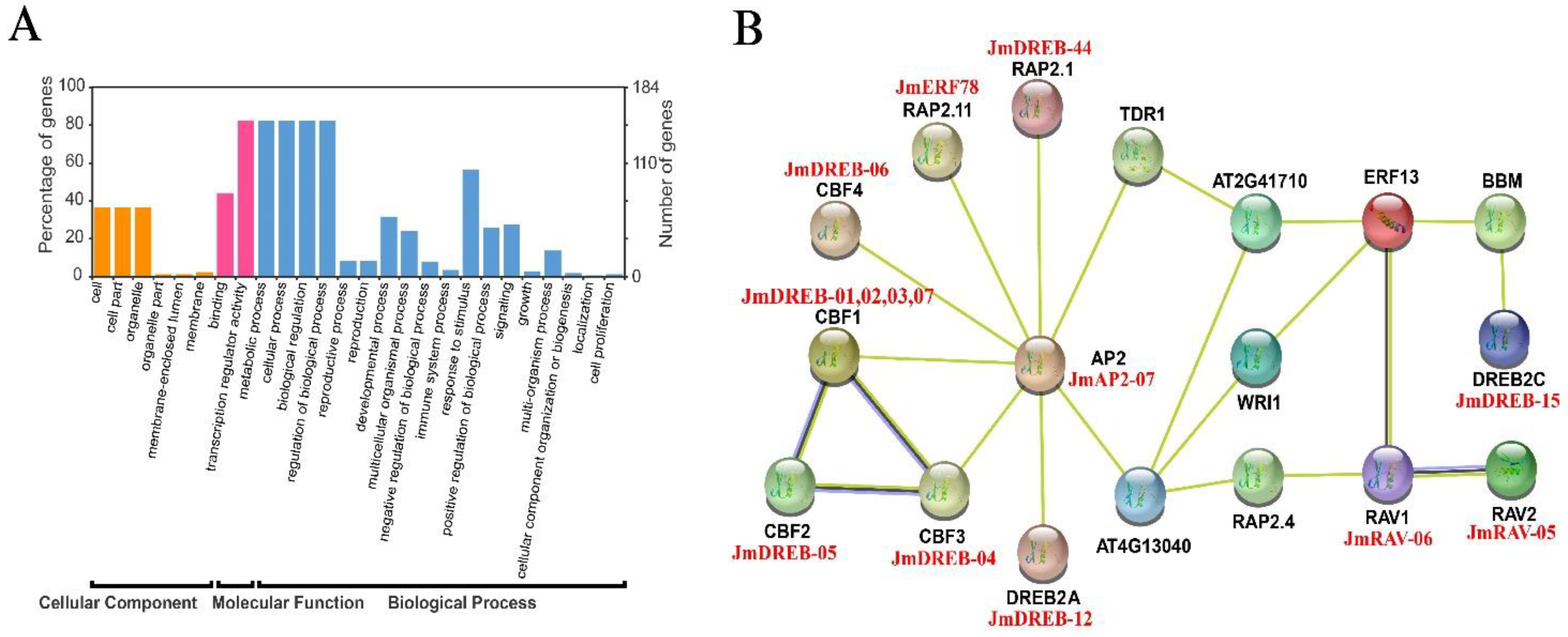
2.5. Interaction Network Analysis of JmAP2/ERF Proteins
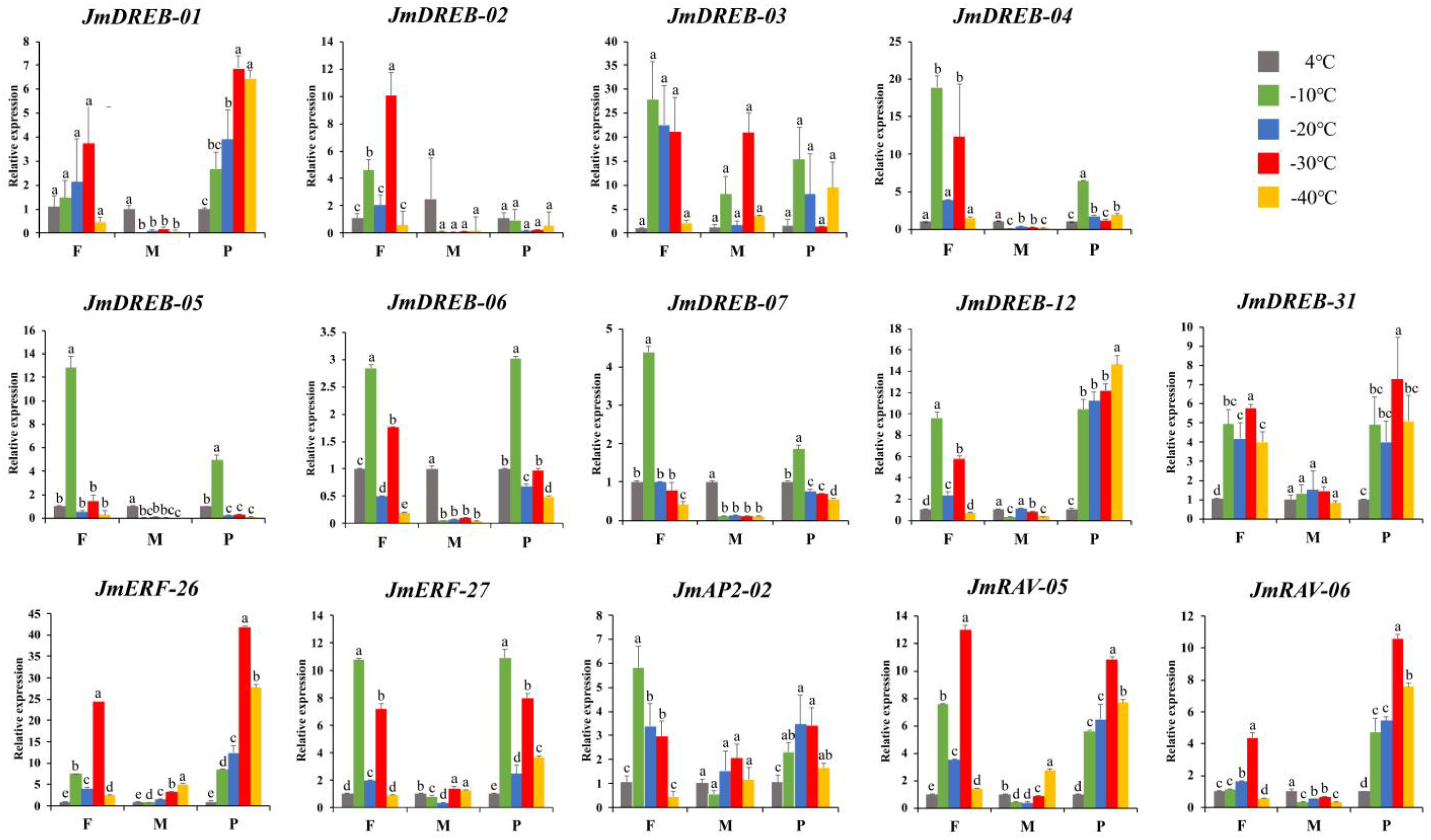
2.6. Expression Profiles of JmAP2/ERF Genes under Cold Stress Using qRT-PCR
3. Discussion
3.1. Identification of the AP2/ERF Superfamily
3.2. The Characteristics Information of AP2/ERF Superfamily in J. mandshurica
3.3. Gene Duplication Events Are Vital Factors in the JmAP2/ERF Superfamily Evolution
3.4. Exploring AP2/ERF Genes Function and Expression Profile in J. mandshurica
4. Materials and Methods
4.1. Identification and Sequence Analysis of JmAP2/ERF Genes
4.2. Phylogenetic Analysis, Gene Structure Identification and Conserved Motif Distribution
4.3. Chromosomal Distribution, Gene Duplications, and Evolutionary Analysis
4.4. Cis-Acting Element Analysis
4.5. Prediction of the Protein Interaction Network
4.6. Plant Materials and Cold Stress Treatments
4.7. RNA Extraction and qRT-PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Singh, K.; Foley, R.C.; Onate-Sanchez, L. Transcription factors in plant defense and stress responses. Curr. Opin. Plant Biol. 2002, 5, 430–436. [Google Scholar] [CrossRef]
- Feng, K.; Hou, X.L.; Xing, G.M.; Liu, J.X.; Duan, A.Q.; Xu, Z.S.; Li, M.Y.; Zhuang, J.; Xiong, A.S. Advances in AP2/ERF super-family transcription factors in plant. Crit. Rev. Biotechnol. 2020, 40, 750–776. [Google Scholar] [CrossRef] [PubMed]
- Licausi, F.; Ohme-Takagi, M.; Perata, P. APETALA/Ethylene Responsive Factor (AP2/ERF) transcription factors: Mediators of stress responses and developmental programs. New Phytol. 2013, 199, 639–649. [Google Scholar] [CrossRef] [PubMed]
- Andriankaja, A.; Boisson-Demier, A.; Frances, L.; Sauviac, L.; Jauneau, A.; Barker, D.G.; de Carvalho-Niebel, F. AP2-ERF transcription factors mediate nod factor-dependent mt ENOD11 activation in root hairs via a novel cis-regulatory motif. Plant Cell 2007, 19, 2866–2885. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakano, T.; Suzuki, K.; Fujimura, T.; Shinshi, H. Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 2006, 140, 411–432. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ohme-Takagi, M.; Shinshi, H. Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 1995, 7, 173–182. [Google Scholar] [CrossRef] [Green Version]
- Sakuma, Y.; Liu, Q.; Dubouzet, J.G.; Abe, H.; Shinozaki, K.; Yamaguchi-Shinozaki, K. DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem. Biophys. Res. Commun. 2002, 290, 998–1009. [Google Scholar] [CrossRef]
- Faraji, S.; Filiz, E.; Kazemitabar, S.K.; Vannozzi, A.; Palumbo, F.; Barcaccia, G.; Heidari, P. The AP2/ERF Gene Family in Triticum durum: Genome-Wide Identification and Expression Analysis under Drought and Salinity Stresses. Genes 2020, 11, 1464. [Google Scholar] [CrossRef]
- Cui, M.J.; Haider, M.S.; Chai, P.P.; Guo, J.J.; Du, P.; Li, H.Y.; Dong, W.Z.; Huang, B.Y.; Zheng, Z.; Shi, L.; et al. Genome-wide identification and expression analysis of AP2/ERF transcription factor related to drought stress in cultivated peanut (Arachis hypogaea L.). Front. Genet. 2021, 12, 750761. [Google Scholar] [CrossRef]
- Zhang, L.; Chen, L.J.; Pang, S.Q.; Zheng, Q.; Quan, S.W.; Liu, Y.F.; Xu, T.; Liu, Y.D.; Qi, M.F. Function analysis of the ERF and DREB subfamilies in tomato fruit development and ripening. Front. Plant Sci. 2022, 13, 849048. [Google Scholar] [CrossRef]
- Yao, W.; An, T.Y.; Xu, Z.C.; Zhang, L.B.; Gao, H.; Sun, W.; Liao, B.S.; Jiang, C.H.; Liu, Z.Q.; Duan, L.X.; et al. Genomic-wide identification and expression analysis of AP2/ERF transcription factors related to andrographolide biosynthesis in Andrographis paniculata. Ind. Crops Prod. 2020, 157, 112878. [Google Scholar] [CrossRef]
- Kabir, S.M.T.; Hossain, M.S.; Bashar, K.K.; Honi, U.; Ahmed, B.; Emdad, E.M.; Alam, M.M.; Haque, M.S.; Islam, M.S. Genome-wide identification and expression profiling of AP2/ERF superfamily genes under stress conditions in dark jute (Corchorus olitorius L.). Ind. Crops Prod. 2021, 166, 113469. [Google Scholar] [CrossRef]
- Zhu, Y.Y.; Liu, X.L.; Gao, Y.D.; Li, K.; Guo, W.D. Transcriptome-based identification of AP2/ERF family genes and their cold-regulated expression during the dormancy phase transition of Chinese cherry flower buds. Sci. Hortic. 2021, 275, 109666. [Google Scholar] [CrossRef]
- Nakano, T.; Fujisawa, M.; Shima, Y.; Ito, Y. The AP2/ERF transcription factor SlERF52 functions in flower pedicel abscission in tomato. J. Exp. Bot. 2014, 65, 3111–3119. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shigyo, M.; Ito, M. Analysis of gymnosperm two-AP2-domain-containing genes. Dev. Genes Evol. 2004, 214, 105–114. [Google Scholar] [CrossRef]
- Licausi, F.; Giorgi, F.M.; Zenoni, S.; Osti, F.; Pezzotti, M.; Perata, P. Genomic and transcriptomic analysis of the AP2/ERF superfamily in Vitis vinifera. BMC Genom. 2010, 11, 719. [Google Scholar] [CrossRef] [Green Version]
- Xie, Z.; Nolan, T.M.; Jiang, H.; Yin, Y. AP2/ERF Transcription Factor Regulatory Networks in Hormone and Abiotic Stress Responses in Arabidopsis. Front. Plant Sci. 2019, 10, 228. [Google Scholar] [CrossRef] [Green Version]
- Xie, Z.L.; Nolan, R.O.R.; Jiang, H.; Tang, B.Y.; Zhang, M.C.; Li, Z.H.; Yin, Y.H. The AP2/ERF transcription factor TINY Modulates Brassinosteroid-Regulated plant growth and drought responses in Arabidopsis. Plant Cell 2019, 31, 1788–1806. [Google Scholar] [CrossRef] [Green Version]
- Lee, D.K.; Jung, H.; Jang, G.; Jeong, J.S.; Kim, Y.S.; Ha, S.H.; Do Choi, Y.; Kim, J.K. Overexpression of the OsERF71 Transcription Factor Alters Rice Root Structure and Drought Resistance. Plant Physiol. 2016, 172, 575–588. [Google Scholar] [CrossRef] [Green Version]
- Stockinger, E.J.; Gilmour, S.J.; Thomashow, M.F. Arabidopsis thaliana CBF1 encodes an AP2 domain-containing transcriptional activator that binds to the C-repeat/DRE, a cis-acting DNA regulatory element that stimulates transcription in response to low temperature and water deficit. Proc. Natl. Acad. Sci. USA 1997, 94, 1035–1040. [Google Scholar] [CrossRef]
- Hu, Z.; Ban, Q.; Hao, J.; Zhu, X.; Cheng, Y.; Mao, J.; Lin, M.; Xia, E.; Li, Y. Genome-Wide Characterization of the C-repeat Binding Factor (CBF) Gene Family Involved in the Response to Abiotic Stresses in Tea Plant (Camellia sinensis). Front. Plant Sci. 2020, 11, 921. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.L.; Shi, Y.T.; Yang, S.H. Advances and challenges in uncovering cold tolerance regulatory mechanisms in plants. New Phytol. 2019, 222, 1690–1704. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, S.; Tang, X.F.; Ma, N.N.; Wang, L.Y.; Meng, Q.W. Heterology expression of the sweet pepper CBF3 gene confers elevated tolerance to chilling stress in transgenic tobacco. J. Plant Physiol. 2011, 168, 1804–1812. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.X.; Wang, W.H.; Zhang, H.X.; Liu, J.H.; Shi, B.Y.; Dai, W.Z.; Liu, K.W.; Zhang, H.G. Diversity in fruit morphology and nutritional composition of Juglans mandshurica Maxim in northeast China. Front. Plant Sci. 2022, 13, 820457. [Google Scholar] [CrossRef]
- Zhang, L.J.; Guo, C.; Lu, X.J.; Sun, X.M.; Liu, C.P.; Zhou, Q.; Deng, J.F. Flower development of heterodichogamous Juglans mandshurica (Juglandaceae). Front. Plant Sci. 2021, 12, 541163. [Google Scholar] [CrossRef]
- Zhang, S.K.; Zhan, W.; Zhang, H.; Song, S.L.; Qu, X.B.; Wang, J.Y.; Tigabu, M.; Li, X.; Zhang, Q.H.; Zhao, M.H.; et al. Genetic improvement in Juglans mandshurica and its uses in China: Current status and future prospects. Phyton-Int. J. Exp. Bot. 2022, 91, 489–505. [Google Scholar] [CrossRef]
- Zhang, Q.H.; Yu, S.H.; Pei, X.N.; Wang, Q.C.; Lu, A.J.; Cao, Y.; Tigabu, M.; Feng, J.; Zhao, X.Y. Within- and between-population variations in seed and seedling traits of Juglans mandshurica (Aug, 10.1007/s11676-021-01381-1, 2021). J. For. Res. 2022, 33, 1711. [Google Scholar] [CrossRef]
- Luan, F.; Wang, Z.Y.; Yang, Y.; Ji, Y.F.; Lv, H.Z.; Han, K.Q.; Liu, D.H.; Shang, X.F.; He, X.R.; Zeng, N. Juglans mandshurica Maxim.: A review of its traditional usages, phytochemical constituents, and pharmacological properties. Front. Pharmacol. 2021, 11, 569800. [Google Scholar] [CrossRef]
- Islam, M.S.; Saito, J.A.; Emdad, E.M.; Ahmed, B.; Islam, M.M.; Halim, A.; Hossen, Q.M.M.; Hossain, M.Z.; Ahmed, R.; Hossain, M.S.; et al. Comparative genomics of two jute species and insight into fibre biogenesis. Nat. Plants 2017, 3, 16223. [Google Scholar] [CrossRef] [Green Version]
- Doudna, J.A.; Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 2014, 346, 1258096. [Google Scholar] [CrossRef]
- Yan, F.; Xi, R.M.; She, R.X.; Chen, P.P.; Yan, Y.J.; Yang, G.; Dang, M.; Yue, M.; Pei, D.; Woeste, K.; et al. Improved de novo chromosome-level genome assembly of the vulnerable walnut tree Juglans mandshurica reveals gene family evolution and possible genome basis of resistance to lesion nematode. Mol. Ecol. Resour. 2021, 21, 2063–2076. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Bian, J.; Guan, Y.; Xu, F.; Han, X.; Deng, X.; Liu, X. Genome-Wide Analysis and Expression Profiles of Ethylene Signal Genes and Apetala2/Ethylene-Responsive Factors in Peanut (Arachis hypogaea L.). Front. Plant Sci. 2022, 13, 828482. [Google Scholar] [CrossRef]
- Yang, B.B.; Yao, X.H.; Zeng, Y.R.; Zhang, C.C. Genome-wide identification, characterization, and expression profiling of AP2/ERF superfamily genes under different development and abiotic stress conditions in pecan (Carya illinoinensis). Int. J. Mol. Sci. 2022, 23, 2920. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Wang, J.; Ma, X.; Li, Y.; Lei, T.; Wang, L.; Ge, W.; Guo, D.; Wang, Z.; Li, C.; et al. Origination, Expansion, Evolutionary Trajectory, and Expression Bias of AP2/ERF Superfamily in Brassica napus. Front. Plant Sci. 2016, 7, 1186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.; Cai, K.W.; Pei, X.N.; Li, Y.; Hu, Y.B.; Meng, F.J.; Song, X.S.; Tigabu, M.; Ding, C.J.; Zhao, X.Y. Genome-wide identification of NAC transcription factor family in Juglans mandshurica and their expression analysis during the fruit development and ripening. Int. J. Mol. Sci. 2021, 22, 2414. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Zhang, L.; Chen, P.; Wu, C.; Zhang, H.; Yuan, J.; Zhou, J.; Li, X. Genome-wide identification of APETALA2/ETHYLENE RESPONSIVE FACTOR transcription factors in Cucurbita moschata and their involvement in ethylene response. Front. Plant Sci. 2022, 13, 847754. [Google Scholar] [CrossRef] [PubMed]
- Karniely, S.; Pines, O. Single translation-dual destination: Mechanisms of dual protein targeting in eukaryotes. EMBO Rep. 2005, 6, 420–425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, S.; Zhu, C.; Lyu, Y.; Chen, Y.; Zhang, Z.; Lai, Z.; Lin, Y. Genome-wide identification, molecular evolution, and expression analysis provide new insights into the APETALA2/ethylene responsive factor (AP2/ERF) superfamily in Dimocarpus longan Lour. BMC Genom. 2020, 21, 62. [Google Scholar] [CrossRef] [Green Version]
- Liu, M.; Sun, W.; Ma, Z.; Zheng, T.; Huang, L.; Wu, Q.; Zhao, G.; Tang, Z.; Bu, T.; Li, C.; et al. Genome-wide investigation of the AP2/ERF gene family in tartary buckwheat (Fagopyum Tataricum). BMC Plant Biol. 2019, 19, 84. [Google Scholar] [CrossRef] [Green Version]
- Donde, R.; Gupta, M.K.; Gouda, G.; Kumar, J.; Vadde, R.; Sahoo, K.K.; Dash, S.K.; Behera, L. Computational characterization of structural and functional roles of DREB1A, DREB1B and DREB1C in enhancing cold tolerance in rice plant. Amino Acids 2019, 51, 839–853. [Google Scholar] [CrossRef]
- Zhang, Y.J.; Yang, J.S.; Guo, S.J.; Meng, J.J.; Zhang, Y.L.; Wan, S.B.; He, Q.W.; Li, X.G. Over-expression of the Arabidopsis CBF1 gene improves resistance of tomato leaves to low temperature under low irradiance. Plant Biol. 2011, 13, 362–367. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.J.; Kwon, C.W.; Choi, D.W.; Song, S.I.; Kim, J.K. Expression of barley HvCBF4 enhances tolerance to abiotic stress in transgenic rice. Plant Biotechnol. J. 2007, 5, 646–656. [Google Scholar] [CrossRef] [PubMed]
- Yun, K.Y.; Park, M.R.; Mohanty, B.; Herath, V.; Xu, F.Y.; Mauleon, R.; Wijaya, E.; Bajic, V.B.; Bruskiewich, R.; de los Reyes, B.G. Transcriptional regulatory network triggered by oxidative signals configures the early response mechanisms of japonica rice to chilling stress. BMC Plant Biol. 2010, 10, 16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kong, H.; Landherr, L.L.; Frohlich, M.W.; Leebens-Mack, J.; Ma, H.; de Pamphilis, C.W. Patterns of gene duplication in the plant SKP1 gene family in angiosperms: Evidence for multiple mechanisms of rapid gene birth. Plant J. 2007, 50, 873–885. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.Y.; He, Y.; Zhu, Y.S.; Li, M.Y.; Song, S.; Bo, W.H.; Li, Y.Y.; Pang, X.M. Comparative transcriptome profiling reveals cold stress responsiveness in two contrasting Chinese jujube cultivars. BMC Plant Biol. 2020, 20, 240. [Google Scholar] [CrossRef] [PubMed]
- Tian, J.; Wang, H.; Gao, Y.; Zhang, Z. Assessment of Freezing Tolerance of Juglans Germplasms by Using Annual Dormant Branches. Acta Hortic. Sin. 2013, 40, 1051–1060. [Google Scholar] [CrossRef]
- Villouta, C.; Workmaster, B.A.; Atucha, A. Freezing stress damage and growth viability in Vaccinium macrocarpon Ait. bud structures. Physiol. Plant. 2021, 172, 2238–2250. [Google Scholar] [CrossRef]
- Dubouzet, J.G.; Sakuma, Y.; Ito, Y.; Kasuga, M.; Dubouzet, E.G.; Miura, S.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J. Cell Mol. Biol. 2003, 33, 751–763. [Google Scholar] [CrossRef]
- Gilmour, S.J.; Sebolt, A.M.; Salazar, M.P.; Everard, J.D.; Thomashow, M.F. Overexpression of the Arabidopsis CBF3 transcriptional activator mimics multiple biochemical changes associated with cold acclimation. Plant Physiol. 2000, 124, 1854–1865. [Google Scholar] [CrossRef] [Green Version]
- Lee, B.H.; Henderson, D.A.; Zhu, J.K. The Arabidopsis cold-responsive transcriptome and its regulation by ICE1. Plant Cell 2005, 17, 3155–3175. [Google Scholar] [CrossRef]
- Illgen, S.; Zintl, S.; Zuther, E.; Hincha, D.K.; Schmuelling, T. Characterisation of the ERF102 to ERF105 genes of Arabidopsis thaliana and their role in the response to cold stress. Plant Mol. Biol. 2020, 103, 303–320. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, Y.T.; Ding, Y.L.; Yang, S.H. Molecular reculation of CBF sicnalinc in cold acclimation. Trends Plant Sci. 2018, 23, 623–637. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Ma, J.; Liu, H.; Ou, M.; Ye, H.; Zhao, P. Identification and Characterization of Wall-Associated Kinase (WAK) and WAK-like (WAKL) Gene Family in Juglans regia and Its Wild Related Species Juglans mandshurica. Genes 2022, 13, 134. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Cai, K.W.; Zhang, Q.H.; Pei, X.N.; Chen, S.; Jiang, L.P.; Han, Z.M.; Zhao, M.H.; Li, Y.; Zhang, X.X.; et al. The Manchurian Walnut genome: Insights into juglone and lipid biosynthesis. Gigascience 2022, 11, giac057. [Google Scholar] [CrossRef] [PubMed]
- Gasteiger, E.; Gattiker, A.; Hoogland, C.; Ivanyi, I.; Appel, R.D.; Bairoch, A. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 2003, 31, 3784–3788. [Google Scholar] [CrossRef] [Green Version]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [Green Version]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Cantalapiedra, C.P.; Hernandez-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. EggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef]
- Ye, J.; Zhang, Y.; Cui, H.H.; Liu, J.W.; Wu, Y.Q.; Cheng, Y.; Xu, H.X.; Huang, X.X.; Li, S.T.; Zhou, A.; et al. WEGO 2.0: A web tool for analyzing and plotting GO annotations, 2018 update. Nucleic Acids Res. 2018, 46, W71–W75. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING database in 2021: Customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef] [PubMed]

| Classification | Domains | Group | Number of Genes |
|---|---|---|---|
| RAV subfamily | AP2-B3 | 6 | |
| Soloist subfamily | AP2 | 4 | |
| AP2 subfamily | AP2 | ANT | 3 |
| AP2-AP2 | AP2 | 24 | |
| Total JmAP2 genes | 27 | ||
| DREB subfamily | AP2 | I | 10 |
| AP2 | II | 11 | |
| AP2 | III | 27 | |
| AP2 | IV | 9 | |
| Total JmDREB genes | 57 | ||
| ERF subfamily | AP2 | V | 19 |
| AP2 | VI | 8 | |
| AP2 | VII | 5 | |
| AP2 | VIII | 18 | |
| AP2 | IX | 29 | |
| AP2 | X | 8 | |
| AP2 | VI-L | 3 | |
| Total JmERF genes | 90 | ||
| Total JmAP2/ERF genes | 184 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, M.; Li, Y.; Zhang, X.; You, X.; Yu, H.; Guo, R.; Zhao, X. Genome-Wide Identification of AP2/ERF Superfamily Genes in Juglans mandshurica and Expression Analysis under Cold Stress. Int. J. Mol. Sci. 2022, 23, 15225. https://doi.org/10.3390/ijms232315225
Zhao M, Li Y, Zhang X, You X, Yu H, Guo R, Zhao X. Genome-Wide Identification of AP2/ERF Superfamily Genes in Juglans mandshurica and Expression Analysis under Cold Stress. International Journal of Molecular Sciences. 2022; 23(23):15225. https://doi.org/10.3390/ijms232315225
Chicago/Turabian StyleZhao, Minghui, Yan Li, Xinxin Zhang, Xiangling You, Haiyang Yu, Ruixue Guo, and Xiyang Zhao. 2022. "Genome-Wide Identification of AP2/ERF Superfamily Genes in Juglans mandshurica and Expression Analysis under Cold Stress" International Journal of Molecular Sciences 23, no. 23: 15225. https://doi.org/10.3390/ijms232315225





