Combined Model-Based Prediction for Non-Invasive Prenatal Screening
Abstract
:1. Introduction
2. Results
2.1. Basic Information on the Study Samples
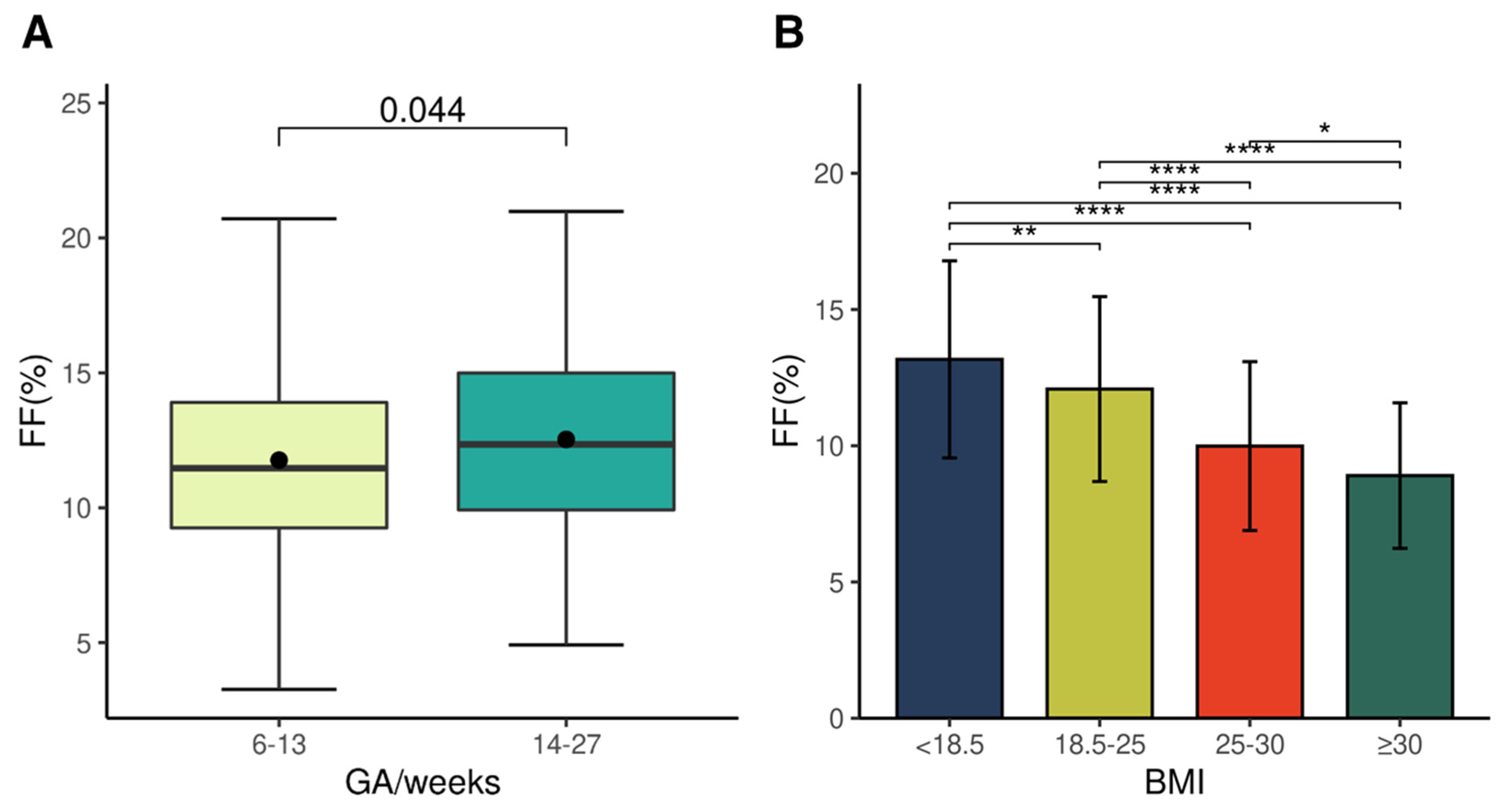
2.2. Features of a Pregnanncy-Affecting Fetal Fraction
2.3. Factors Affecting the Z-Score of NIPT
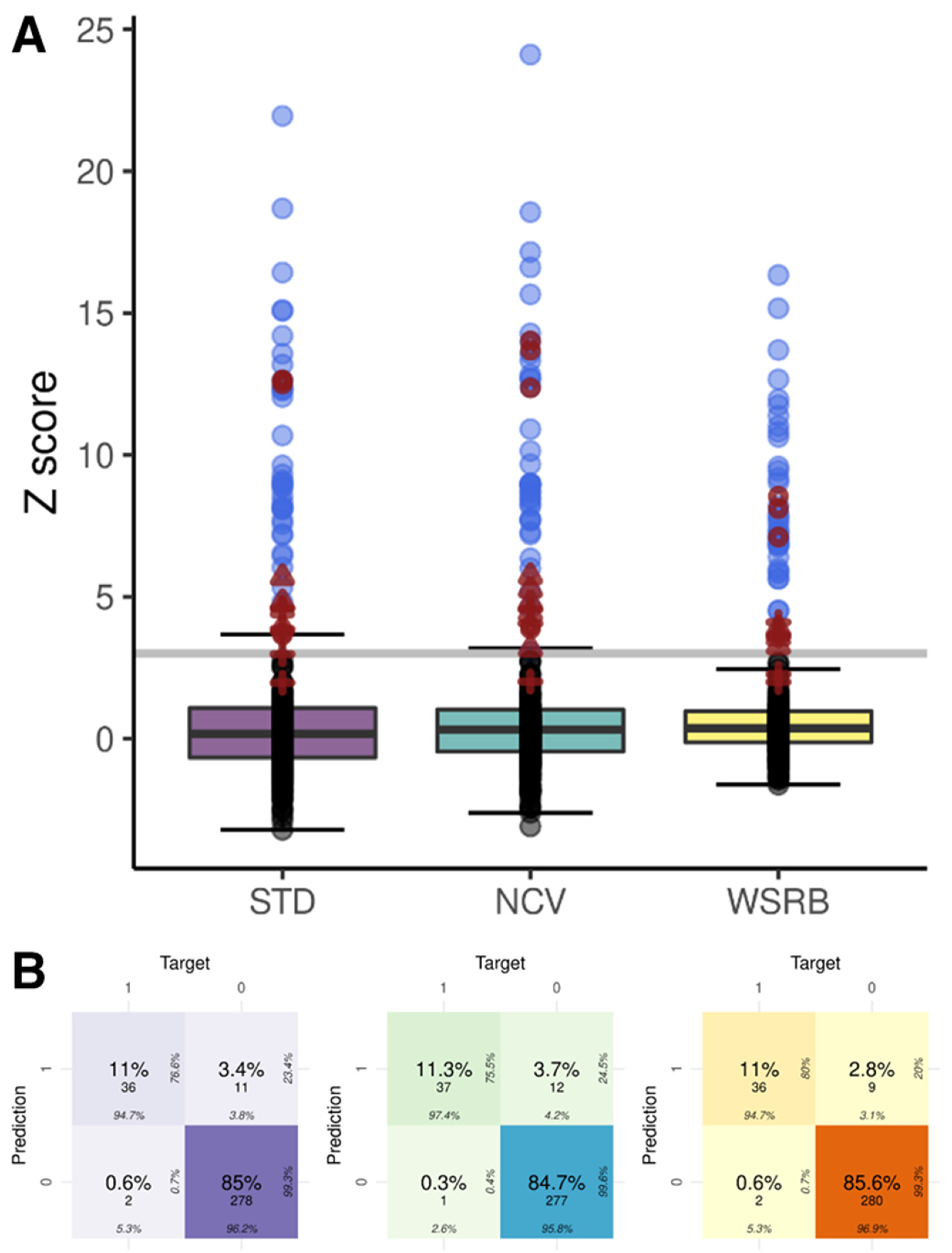
2.4. NIPT Results of the Three Methods
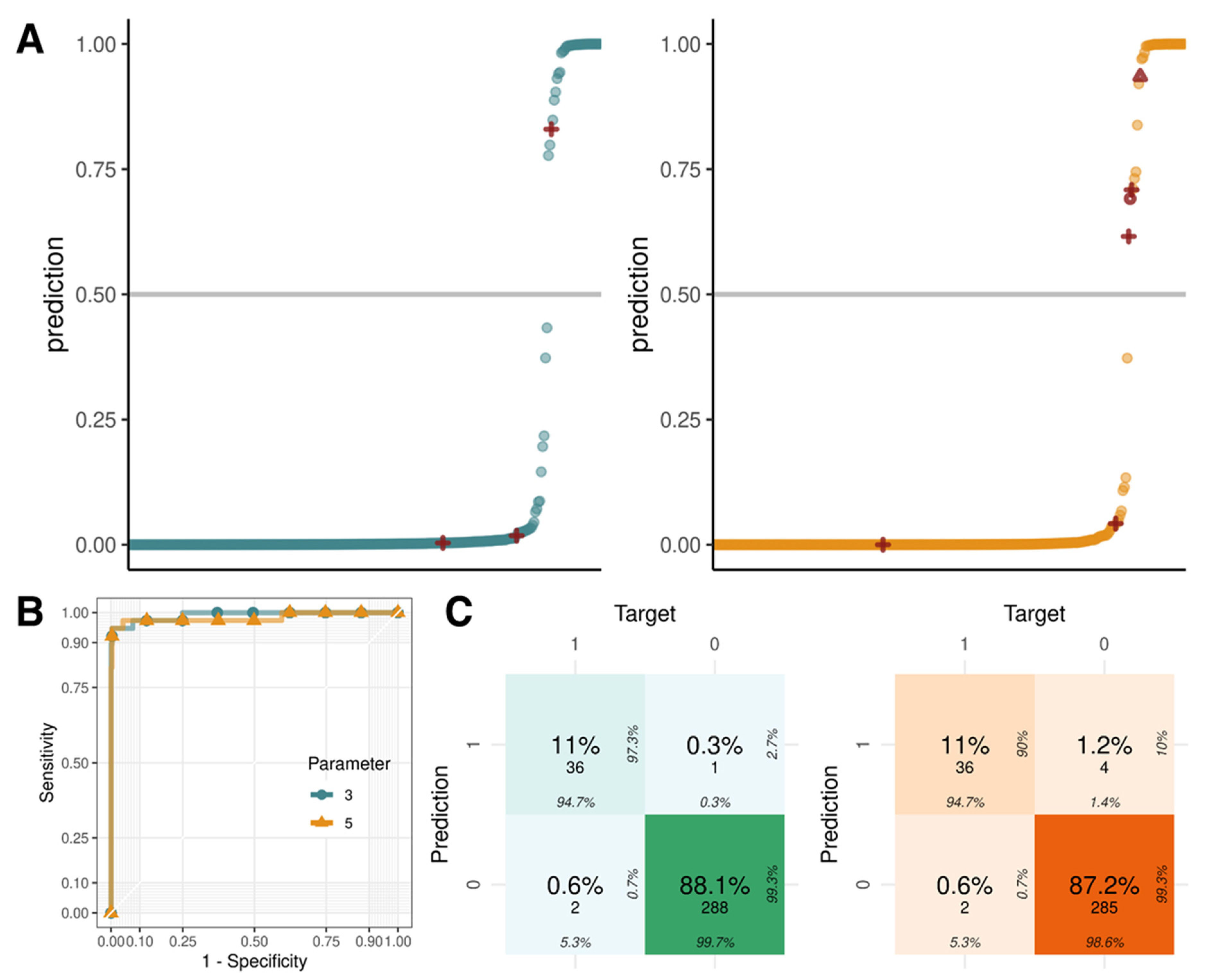
2.5. Logistic Regression with Parameters Using Z-Scores, GC Content, and Fetal Fraction
3. Discussion
4. Materials and Methods
4.1. Sample Preparation and Sequencing
4.2. Shallow NGS Data Analysis
4.3. NIPT Analysis
4.4. Logistic Regression
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- OECD Family Database: SF2.3: Age of Mothers at Childbirth and Age-Specific Fertility. Available online: https://www.oecd.org/els/family/database.htm (accessed on 30 October 2022).
- Norton, M.E.; Jacobsson, B.; Swamy, G.K.; Laurent, L.C.; Ranzini, A.C.; Brar, H.; Tomlinson, M.W.; Pereira, L.; Spitz, J.L.; Hollemon, D.; et al. Cell-free DNA analysis for noninvasive examination of trisomy. N. Engl. J. Med. 2015, 372, 1589–1597. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tabor, A.; Philip, J.; Madsen, M.; Bang, J.; Obel, E.B.; Norgaard-Pedersen, B. Randomised controlled trial of genetic amniocentesis in 4606 low-risk women. Lancet 1986, 1, 1287–1293. [Google Scholar] [CrossRef]
- Benn, P.; Rebarber, A. Non-invasive prenatal testing in the management of twin pregnancies. Prenat. Diagn. 2021, 41, 1233–1240. [Google Scholar] [CrossRef] [PubMed]
- Errol, R.; Norwitz, M.; Brynn, L. Noninvasive Prenatal Testing_The Future is Now. Rev. Obstet. Gynecol. 2013, 6, 48. [Google Scholar] [CrossRef]
- Mayo, S.; Gomez-Manjon, I.; Atencia, G.; Moreno-Izquierdo, A.; Escribano, D.; Fernandez-Martinez, F.J. Noninvasive prenatal testing: How far can we reach detecting fetal copy number variations. Eur. J. Obstet. Gynecol. Reprod. Biol. 2022, 272, 150–155. [Google Scholar] [CrossRef] [PubMed]
- Junhui, W.; Ru, L.; Qiuxia, Y.; Dan, W.; Xiuhong, S.; Yongling, Z.; Xiangyi, J.; Fatao, L.; Xuewei, T.; Guilan, C.; et al. Evaluation of the Z-score accuracy of noninvasive prenatal testing for fetal trisomies 13, 18 and 21 at a single center. Prenat. Diagn. 2021, 41, 690–696. [Google Scholar] [CrossRef]
- Zhu, H.; Jin, X.; Xu, Y.; Zhang, W.; Liu, X.; Jin, J.; Qian, Y.; Dong, M. Efficiency of non-invasive prenatal screening in pregnant women at advanced maternal age. BMC Pregnancy Childbirth 2021, 21, 86. [Google Scholar] [CrossRef]
- Wang, E.; Batey, A.; Struble, C.; Musci, T.; Song, K.; Oliphant, A. Gestational age and maternal weight effects on fetal cell-free DNA in maternal plasma. Prenat. Diagn. 2013, 33, 662–666. [Google Scholar] [CrossRef]
- Chan, K.C.; Zhang, J.; Hui, A.B.; Wong, N.; Lau, T.K.; Leung, T.N.; Lo, K.W.; Huang, D.W.; Lo, Y.M. Size distributions of maternal and fetal DNA in maternal plasma. Clin. Chem. 2004, 50, 88–92. [Google Scholar] [CrossRef] [Green Version]
- Lo, Y.M.; Zhang, J.; Leung, T.N.; Lau, T.K.; Chang, A.M.; Hjelm, N.M. Rapid clearance of fetal DNA from maternal plasma. Am. J. Hum. Genet. 1999, 64, 218–224. [Google Scholar] [CrossRef]
- Smid, M.; Galbiati, S.; Vassallo, A.; Gambini, D.; Ferrari, A.; Viora, E.; Pagliano, M.; Restagno, G.; Ferrari, M.; Cremonesi, L. No evidence of fetal DNA persistence in maternal plasma after pregnancy. Hum. Genet. 2003, 112, 617–618. [Google Scholar] [CrossRef] [PubMed]
- Chiu, R.W.K.; Chan, K.C.A.; Gao, Y.; Lau, V.Y.M.; Zheng, W.; Leung, T.Y.; Foo, C.H.F.; Xie, B.; Tsui, N.B.Y.; Lun, F.M.F.; et al. Noninvasive prenatal diagnosis of fetal chromosomal aneuploidy by massively parallel genomic sequencing of DNA in maternal plasma. Proc. Natl. Acad. Sci. USA 2008, 105, 20458–20463. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sehnert, A.J.; Rhees, B.; Comstock, D.; de Feo, E.; Heilek, G.; Burke, J.; Rava, R.P. Optimal detection of fetal chromosomal abnormalities by massively parallel DNA sequencing of cell-free fetal DNA from maternal blood. Clin. Chem. 2011, 57, 1042–1049. [Google Scholar] [CrossRef] [PubMed]
- Johansson, L.F.; de Boer, E.N.; de Weerd, H.A.; van Dijk, F.; Elferink, M.G.; Schuring-Blom, G.H.; Suijkerbuijk, R.F.; Sinke, R.J.; Te Meerman, G.J.; Sijmons, R.H.; et al. Novel Algorithms for Improved Sensitivity in Non-Invasive Prenatal Testing. Sci. Rep. 2017, 7, 1838. [Google Scholar] [CrossRef] [Green Version]
- Liu, B.; Tang, X.; Qiu, F.; Tao, C.; Gao, J.; Ma, M.; Zhong, T.; Cai, J.; Li, Y.; Ding, G. DASAF: An R Package for Deep Sequencing-Based Detection of Fetal Autosomal Abnormalities from Maternal Cell-Free DNA. Biomed Res. Int. 2016, 2016, 2714341. [Google Scholar] [CrossRef] [Green Version]
- Lo, K.K.; Boustred, C.; Chitty, L.S.; Plagnol, V. RAPIDR: An analysis package for non-invasive prenatal testing of aneuploidy. Bioinformatics 2014, 30, 2965–2967. [Google Scholar] [CrossRef] [Green Version]
- Raman, L.; Dheedene, A.; De Smet, M.; Van Dorpe, J.; Menten, B. WisecondorX: Improved copy number detection for routine shallow whole-genome sequencing. Nucleic Acids Res. 2019, 47, 1605–1614. [Google Scholar] [CrossRef]
- Johansson, L.F.; de Weerd, H.A.; de Boer, E.N.; van Dijk, F.; Te Meerman, G.J.; Sijmons, R.H.; Sikkema-Raddatz, B.; Swertz, M.A. NIPTeR: An R package for fast and accurate trisomy prediction in non-invasive prenatal testing. BMC Bioinform. 2018, 19, 531. [Google Scholar] [CrossRef] [Green Version]
- Mehari, M.A.; Maeruf, H.; Robles, C.C.; Woldemariam, S.; Adhena, T.; Mulugeta, M.; Haftu, A.; Hagose, H.; Kumsa, H. Advanced maternal age pregnancy and its adverse obstetrical and perinatal outcomes in Ayder comprehensive specialized hospital, Northern Ethiopia, 2017: A comparative cross-sectional study. BMC Pregnancy Childbirth 2020, 20, 60. [Google Scholar] [CrossRef] [Green Version]
- Hou, Y.; Yang, J.; Qi, Y.; Guo, F.; Peng, H.; Wang, D.; Wang, Y.; Luo, X.; Li, Y.; Yin, A. Factors affecting cell-free DNA fetal fraction: Statistical analysis of 13,661 maternal plasmas for non-invasive prenatal screening. Hum. Genom. 2019, 13, 62. [Google Scholar] [CrossRef]
- Deng, C.; Liu, S. Factors Affecting the Fetal Fraction in Noninvasive Prenatal Screening: A Review. Front. Pediatr. 2022, 10, 812781. [Google Scholar] [CrossRef] [PubMed]
- Zaninović, L.; Bašković, M.; Ježek, D.; Katušić Bojanac, A. Validity and Utility of Non-Invasive Prenatal Testing for Copy Number Variations and Microdeletions: A Systematic Review. J. Clin. Med. 2022, 11, 3350. [Google Scholar] [CrossRef] [PubMed]
- Mardy, A.; Wapner, R.J. Confined placental mosaicism and its impact on confirmation of NIPT results. Am. J. Med. Genet. Part C Semin. Med. Genet. 2016, 172, 118–122. [Google Scholar] [CrossRef]
- Hui, L.; Bianchi, D.W. Cell-free fetal nucleic acids in amniotic fluid. Hum. Reprod. Update 2010, 17, 362–371. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tjoa, M.L.; Cindrova-Davies, T.; Spasic-Boskovic, O.; Bianchi, D.W.; Burton, G.J. Trophoblastic oxidative stress and the release of cell-free feto-placental DNA. Am. J. Pathol. 2006, 169, 400–404. [Google Scholar] [CrossRef] [Green Version]
- Salomon, L.J.; Sotiriadis, A.; Wulff, C.B.; Odibo, A.; Akolekar, R. Risk of miscarriage following amniocentesis or chorionic villus sampling: Systematic review of literature and updated meta-analysis. Ultrasound Obstet. Gynecol. 2019, 54, 442–451. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [Green Version]
- Kim, S.K.; Hannum, G.; Geis, J.; Tynan, J.; Hogg, G.; Zhao, C.; Jensen, T.J.; Mazloom, A.R.; Oeth, P.; Ehrich, M.; et al. Determination of fetal DNA fraction from the plasma of pregnant women using sequence read counts. Prenat. Diagn. 2015, 35, 810–815. [Google Scholar] [CrossRef]

| Characteristic | Reference Set (n = 1698) | Test Set (n = 109) |
|---|---|---|
| GA at NIPT (weeks) | ||
| First trimester (6–13) | 1594 | 89 |
| Second trimester (14–27) | 104 | 20 |
| Maternal age (years) | ||
| 20–29 | 54 | 3 |
| 30–39 | 1314 | 79 |
| ≥40 | 330 | 27 |
| Advanced maternal age (≥35 years) | 1291 | 85 |
| BMI | ||
| <18.5 | 140 | 12 |
| 18.5–25 | 1293 | 81 |
| 25–30 | 213 | 9 |
| ≥30 | 52 | 7 |
| Pregnancy | ||
| Singleton | 1623 | 105 |
| Twin | 75 | 4 |
| Secondary outcomes | ||
| Trisomy 18 | 5 | |
| Trisomy 21 | 33 |
| Accuracy | Sensitivity | Specificity | PPV | NPV | |
|---|---|---|---|---|---|
| STD | 0.960 | 0.947 | 0.962 | 0.766 | 0.993 |
| NCV | 0.960 | 0.974 | 0.958 | 0.755 | 0.996 |
| WSRB | 0.966 | 0.947 | 0.969 | 0.800 | 0.993 |
| Accuracy | Sensitivity | Specificity | PPV | NPV | |
|---|---|---|---|---|---|
| 3 parameters | 0.991 | 0.947 | 0.997 | 0.973 | 0.993 |
| 5 parameters | 0.982 | 0.947 | 0.986 | 0.900 | 0.993 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, S.-Y.; Kang, K.M.; Kim, S.-Y.; Lim, S.Y.; Jang, H.Y.; Hong, K.; Cha, D.H.; Shim, S.H.; Joung, J.-G. Combined Model-Based Prediction for Non-Invasive Prenatal Screening. Int. J. Mol. Sci. 2022, 23, 14990. https://doi.org/10.3390/ijms232314990
Yang S-Y, Kang KM, Kim S-Y, Lim SY, Jang HY, Hong K, Cha DH, Shim SH, Joung J-G. Combined Model-Based Prediction for Non-Invasive Prenatal Screening. International Journal of Molecular Sciences. 2022; 23(23):14990. https://doi.org/10.3390/ijms232314990
Chicago/Turabian StyleYang, So-Yun, Kyung Min Kang, Sook-Young Kim, Seo Young Lim, Hee Yeon Jang, Kirim Hong, Dong Hyun Cha, Sung Han Shim, and Je-Gun Joung. 2022. "Combined Model-Based Prediction for Non-Invasive Prenatal Screening" International Journal of Molecular Sciences 23, no. 23: 14990. https://doi.org/10.3390/ijms232314990
APA StyleYang, S.-Y., Kang, K. M., Kim, S.-Y., Lim, S. Y., Jang, H. Y., Hong, K., Cha, D. H., Shim, S. H., & Joung, J.-G. (2022). Combined Model-Based Prediction for Non-Invasive Prenatal Screening. International Journal of Molecular Sciences, 23(23), 14990. https://doi.org/10.3390/ijms232314990






