Dominant Elongase Activity of Elovl5a but Higher Expression of Elovl5b in Common Carp (Cyprinus carpio)
Abstract
:1. Introduction
2. Results
2.1. The Structure Difference between Common Carp Elovl5a and Elovl5b
2.2. Dominant Elongase Activities of Elovl5a Compared with Elovle5b
2.3. In Silico Molecular Docking Difference between Elovl5a and Elovl5b
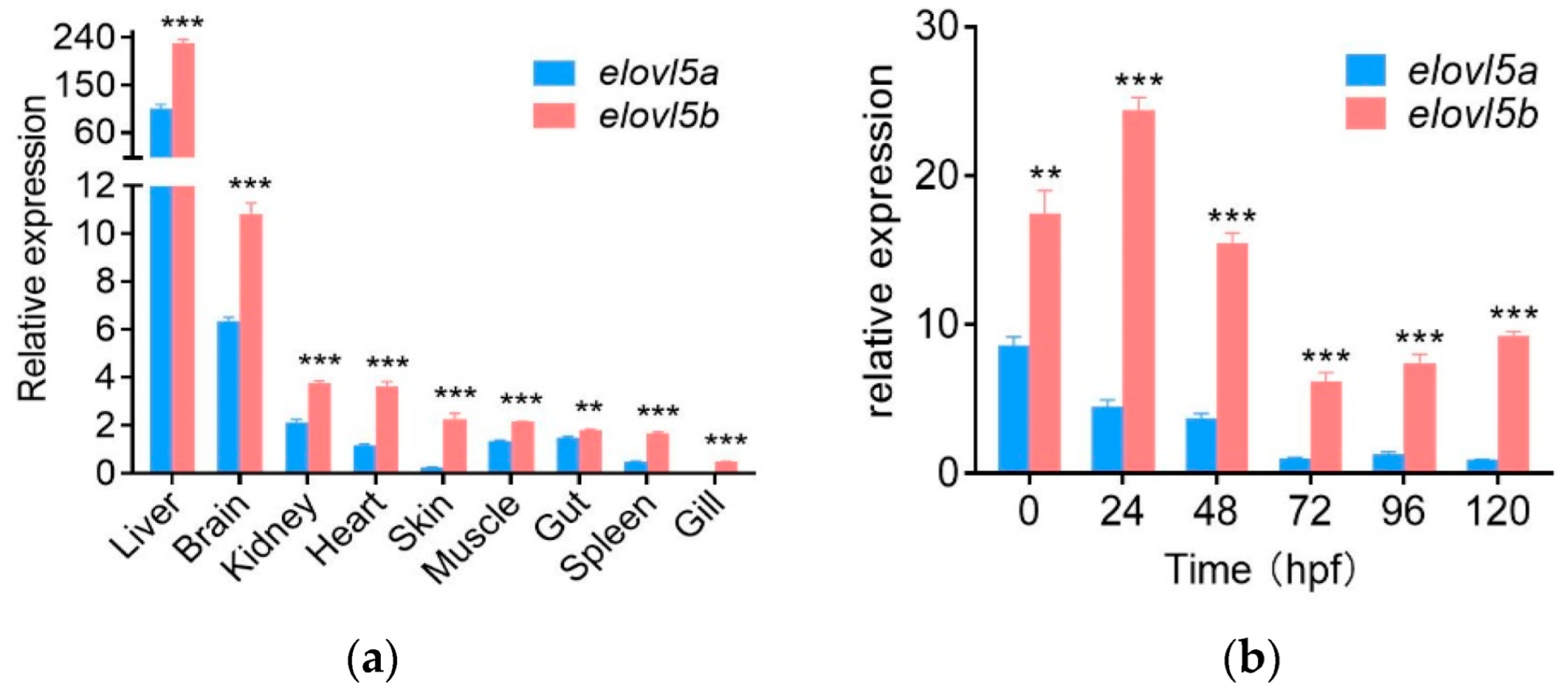
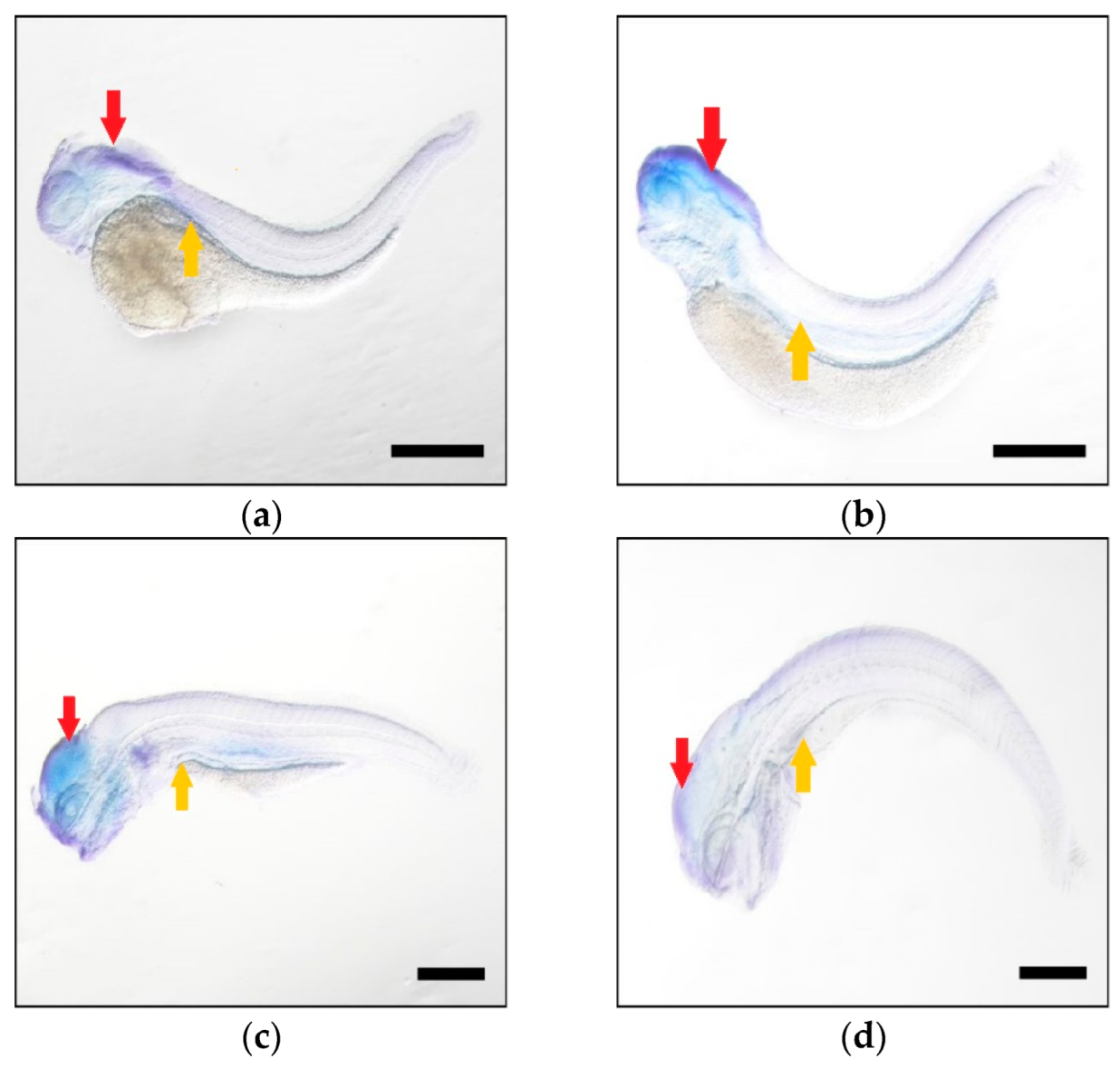
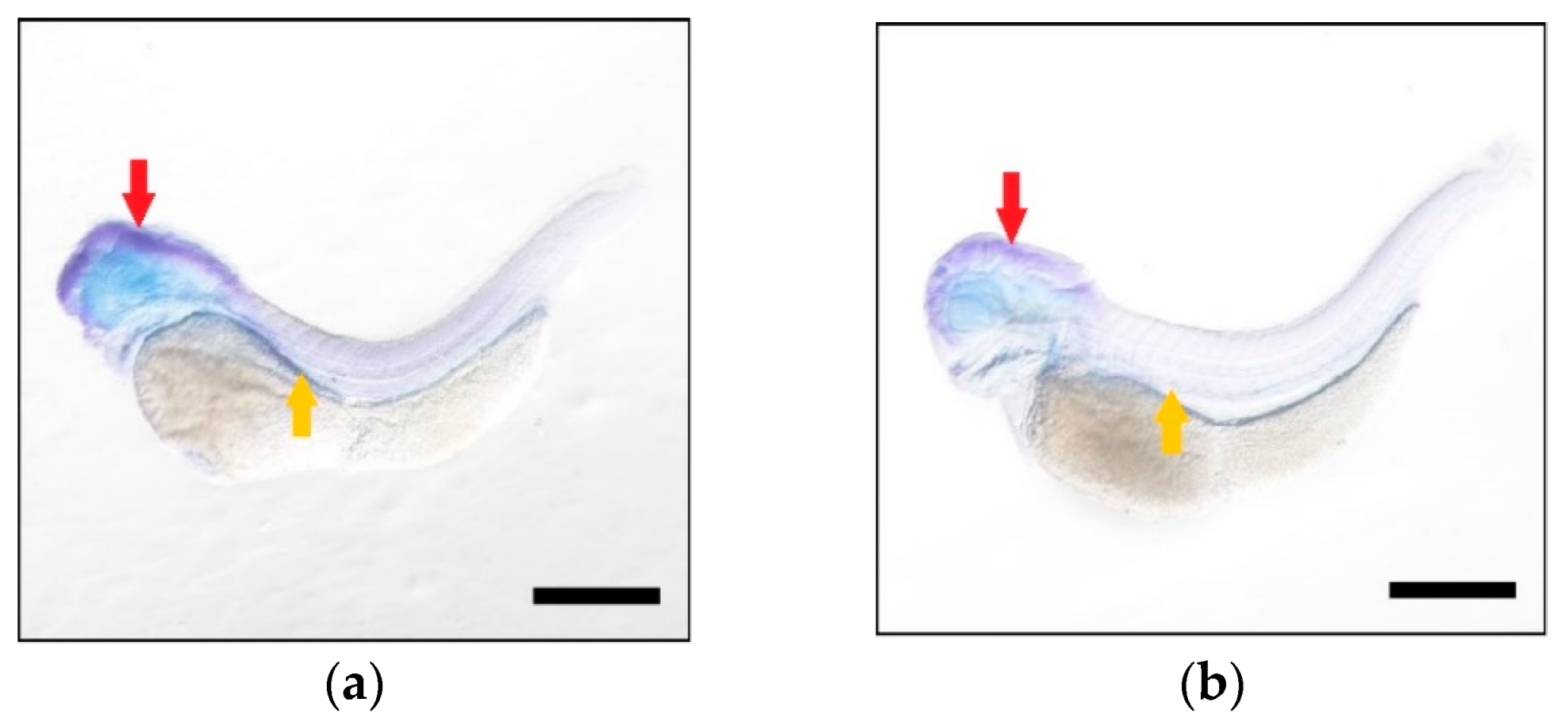
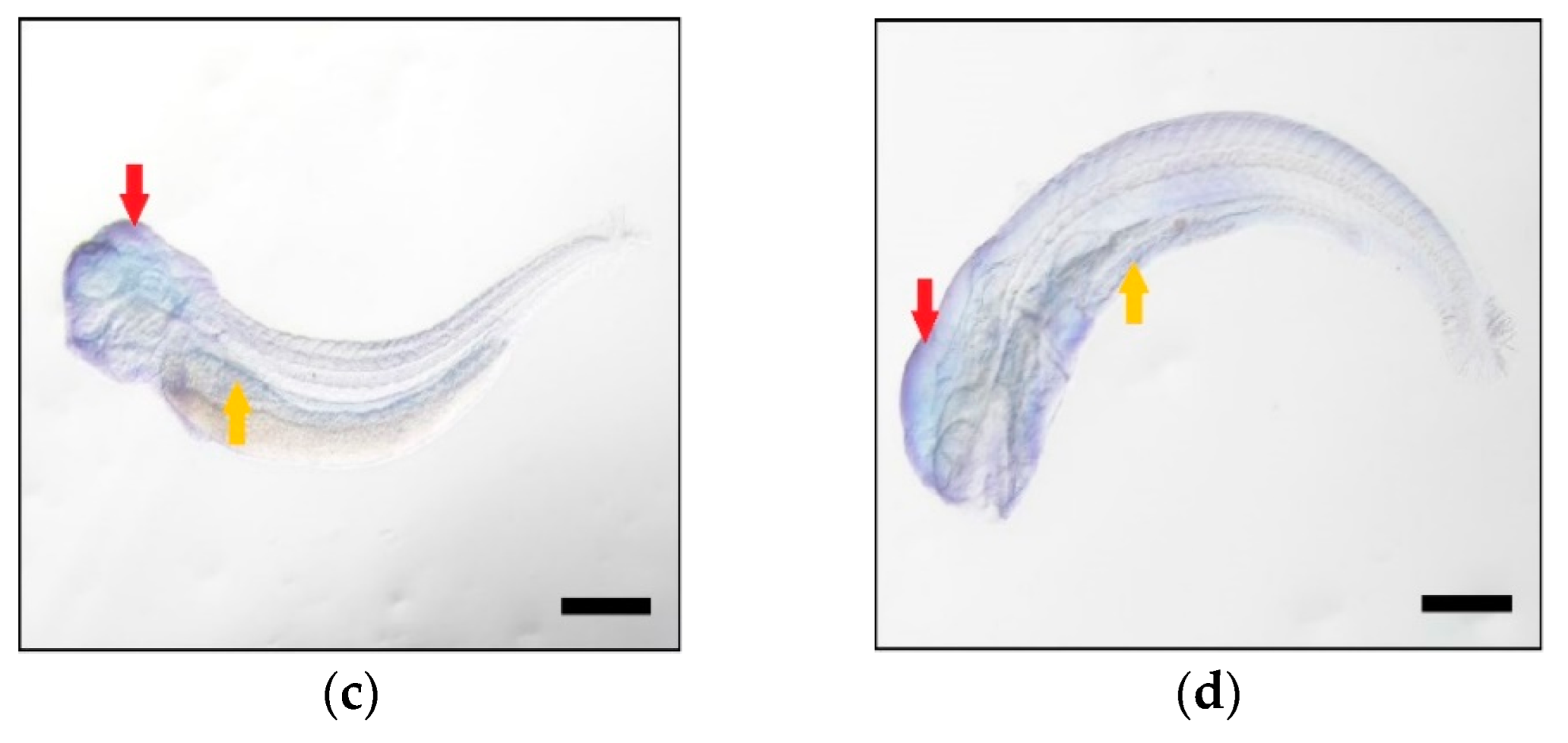
2.4. Higher Expression Levels of Elovl5b Than Elovl5a
2.5. Correlation between the Larvae PUFA Contents and the Expressions of Two Genes
2.6. Promoter Activity Analysis of Elovl5a and Elovl5b
3. Discussion
4. Materials and Methods
4.1. Animals and Tissue Collection
4.2. Cloning of Elovl5a and Elovl5b cDNAs
4.3. Sequence and Phylogenetic Analysis
4.4. Functional Characterization of the Common Carp Elovl5a and Elovl5b in Yeast
4.5. Structure Modelling and Docking of Elovl5a and Elovl5b
4.6. Spatial-Temporal Expression of Elovl5a and Elovl5b
4.7. Fatty Acid Analysis of Common Carp Embryos at Different Developmental Stages
4.8. Identifying the Core Promoters of Common Carp Elovl5a and Elovl5b
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gow, R.V.; Hibbeln, J.R. Omega-3 fatty acid and nutrient deficits in adverse neurodevelopment and childhood behaviors. Child Adolesc. Psychiatr. Clin. N. Am. 2014, 23, 555–590. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lands, B. Highly unsaturated fatty acids (HUFA) mediate and monitor food’s impact on health. Prostaglandins Other Lipid Mediat. 2017, 133, 4–10. [Google Scholar] [CrossRef] [PubMed]
- Rennison, J.H.; Van Wagoner, D.R. Impact of dietary fatty acids on cardiac arrhythmogenesis. Circ. Arrhythm. Electrophysiol. 2009, 2, 460–469. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Monroig, O.; Zhang, L.; Wang, S.; Zheng, X.; Dick, J.R.; You, C.; Tocher, D.R. Vertebrate fatty acyl desaturase with Δ4 activity. Proc. Natl. Acad. Sci. USA 2010, 107, 16840–16845. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davis-Bruno, K.; Tassinari, M.S. Essential fatty acid supplementation of DHA and ARA and effects on neurodevelopment across animal species: A review of the literature. Birth Defects Res. Part B Dev. Reprod. Toxicol. 2011, 92, 240–250. [Google Scholar] [CrossRef]
- Roussel, C.; Anunciação Braga Guebara, S.; Plante, P.L.; Desjardins, Y.; Di Marzo, V.; Silvestri, C. Short-term supplementation with ω-3 polyunsaturated fatty acids modulates primarily mucolytic species from the gut luminal mucin niche in a human fermentation system. Gut Microbes 2022, 14, 2120344. [Google Scholar] [CrossRef]
- Sun, G.Y.; Simonyi, A.; Fritsche, K.L.; Chuang, D.Y.; Hannink, M.; Gu, Z.; Greenlief, C.M.; Yao, J.K.; Lee, J.C.; Beversdorf, D.Q. Docosahexaenoic acid (DHA): An essential nutrient and a nutraceutical for brain health and diseases. Prostaglandins Leukot. Essent. Fat. Acids 2018, 136, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Jakobsson, A.; Westerberg, R.; Jacobsson, A. Fatty acid elongases in mammals: Their regulation and roles in metabolism. Prog. Lipid Res. 2006, 45, 237–249. [Google Scholar] [CrossRef]
- Guo, F.; Bunn, S.E.; Brett, M.T.; Kainz, M.J. Polyunsaturated fatty acids in stream food webs—High dissimilarity among producers and consumers. Freshw. Biol. 2017, 62, 1325–1334. [Google Scholar] [CrossRef] [Green Version]
- Carmona-Antoñanzas, G.; Tocher, D.R.; Taggart, J.B.; Leaver, M.J. An evolutionary perspective on Elovl5 fatty acid elongase: Comparison of Northern pike and duplicated paralogs from Atlantic salmon. BMC Evol. Biol. 2013, 13, 85. [Google Scholar] [CrossRef] [PubMed]
- Gregory, M.K.; James, M.J. Rainbow trout (Oncorhynchus mykiss) Elovl5 and Elovl2 differ in selectivity for elongation of omega-3 docosapentaenoic acid. Biochim. Biophys. Acta (BBA)-Mol. Cell Biol. Lipids 2014, 1841, 1656–1660. [Google Scholar] [CrossRef]
- Li, J.T.; Wang, Q.; Huang Yang, M.D.; Li, Q.S.; Cui, M.S.; Dong, Z.J.; Wang, H.W.; Yu, J.H.; Zhao, Y.J.; Yang, C.R.; et al. Parallel subgenome structure and divergent expression evolution of allo-tetraploid common carp and goldfish. Nat. Genet. 2021, 53, 1493–1503. [Google Scholar] [CrossRef]
- Castro, L.F.; Tocher, D.R.; Monroig, O. Long-chain polyunsaturated fatty acid biosynthesis in chordates: Insights into the evolution of Fads and Elovl gene repertoire. Prog. Lipid Res. 2016, 62, 25–40. [Google Scholar] [CrossRef]
- Ma, W.; Zhu, Z.H.; Bi, X.Y.; Murphy, R.W.; Wang, S.Y.; Gao, Y.; Xiao, H.; Zhang, Y.P.; Luo, J. Allopolyploidization is not so simple: Evidence from the origin of the tribe Cyprinini (Teleostei: Cypriniformes). Curr. Mol. Med. 2014, 14, 1331–1338. [Google Scholar] [CrossRef] [PubMed]
- Agaba, M.; Tocher, D.R.; Dickson, C.A.; Dick, J.R.; Teale, A.J. Zebrafish cDNA encoding multifunctional Fatty Acid elongase involved in production of eicosapentaenoic (20:5n-3) and docosahexaenoic (22:6n-3) acids. Mar. Biotechnol. 2004, 6, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Roy, A.; Kucukural, A.; Zhang, Y. I-TASSER: A unified platform for automated protein structure and function prediction. Nat. Protoc. 2010, 5, 725–738. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stank, A.; Kokh, D.B.; Fuller, J.C.; Wade, R.C. Protein Binding Pocket Dynamics. Acc. Chem. Res. 2016, 49, 809–815. [Google Scholar] [CrossRef] [Green Version]
- Dworkin, M.B.; Dworkin-Rastl, E. Functions of maternal mRNA in early development. Mol. Reprod. Dev. 1990, 26, 261–297. [Google Scholar] [CrossRef]
- Wolf, S.S.; Roder, K.H.; Schweizer, M. The general transcription factor Sp1 plays an important role in the regulation of fatty acid synthase. Biochem. Soc. Trans. 1998, 26, S95. [Google Scholar] [CrossRef]
- Wang, X.; Hassan, W.; Zhao, J.; Bakht, S.; Nie, Y.; Wang, Y.; Pang, Q.; Huang, Z. The impact of hepatocyte nuclear factor-1α on liver malignancies and cell stemness with metabolic consequences. Stem Cell Res. Ther. 2019, 10, 315. [Google Scholar] [CrossRef]
- Sabatier, M.; Birsen, R.; Lauture, L.; Dehairs, J.; Angelino, P.; Mouche, S.; Heiblig, M.; Boet, E.; Sahal, A.; Saland, E.; et al. C/EBPα confers dependence to fatty acid anabolic pathways and vulnerability to lipid oxidative stress in FLT3-mutant leukemia. bioRxiv 2022. [Google Scholar] [CrossRef]
- Dong, Y.; Wang, S.; Chen, J.; Zhang, Q.; Liu, Y.; You, C.; Monroig, Ó.; Tocher, D.R.; Li, Y. Hepatocyte Nuclear Factor 4α (HNF4α) Is a Transcription Factor of Vertebrate Fatty Acyl Desaturase Gene as Identified in Marine Teleost Siganus canaliculatus. PLoS ONE 2016, 11, e0160361. [Google Scholar] [CrossRef] [Green Version]
- Agaba, M.K.; Tocher, D.R.; Zheng, X.; Dickson, C.A.; Dick, J.R.; Teale, A.J. Cloning and functional characterisation of polyunsaturated fatty acid elongases of marine and freshwater teleost fish. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2005, 142, 342–352. [Google Scholar] [CrossRef]
- Marrero, M.; Monroig, Ó.; Navarro, J.C.; Ribes-Navarro, A.; Pérez, J.A.; Galindo, A.; Rodríguez, C. Metabolic and molecular evidence for long-chain PUFA biosynthesis capacity in the grass carp Ctenopharyngodon idella. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2022, 270, 111232. [Google Scholar] [CrossRef] [PubMed]
- Morais, S.; Monroig, O.; Zheng, X.; Leaver, M.J.; Tocher, D.R. Highly unsaturated fatty acid synthesis in Atlantic salmon: Characterization of ELOVL5- and ELOVL2-like elongases. Mar. Biotechnol. 2009, 11, 627–639. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Ye, D.; Wang, H.; He, M.; Sun, Y. Elovl2 But Not Elovl5 Is Essential for the Biosynthesis of Docosahexaenoic Acid (DHA) in Zebrafish: Insight from a Comparative Gene Knockout Study. Mar. Biotechnol. 2020, 22, 613–619. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, Q.-S.; Ye, Y.-Q.; Wang, Q.; Sun, X.-Q.; Zhao, R.; Li, J.-T. Association Analysis between Genetic Variants of elovl5a and elovl5b and Poly-Unsaturated Fatty Acids in Common Carp (Cyprinus carpio). Biology 2022, 11, 466. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Madan, A. CAP3: A DNA sequence assembly program. Genome Res. 1999, 9, 868–877. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wheeler, D.L.; Chappey, C.; Lash, A.E.; Leipe, D.D.; Madden, T.L.; Schuler, G.D.; Tatusova, T.A.; Rapp, B.A. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2000, 28, 10–14. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [Green Version]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, B.; Gao, S.; Lercher, M.J.; Hu, S.; Chen, W.H. Evolview v3: A webserver for visualization, annotation, and management of phylogenetic trees. Nucleic Acids Res. 2019, 47, W270–W275. [Google Scholar] [CrossRef] [PubMed]
- Zhu, K.C.; Song, L.; Guo, H.Y.; Guo, L.; Zhang, N.; Liu, B.S.; Jiang, S.G.; Zhang, D.C. Identification of Fatty Acid Desaturase 6 in Golden Pompano Trachinotus Ovatus (Linnaeus 1758) and Its Regulation by the PPARαb Transcription Factor. Int. J. Mol. Sci. 2018, 20, 23. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Sun, X.-Q.; Ye, Y.-Q.; Wang, Q.; Li, Q.-S.; Zhao, R.; Wang, H.-W.; Li, J.-T. Association between the Polymorphisms of fads2a and fads2b and Poly-Unsaturated Fatty Acids in Common Carp (Cyprinus carpio). Animals 2021, 11, 1780. [Google Scholar] [CrossRef] [PubMed]
- Burley, S.K.; Bhikadiya, C.; Bi, C.; Bittrich, S.; Chen, L.; Crichlow, G.V.; Christie, C.H.; Dalenberg, K.; Di Costanzo, L.; Duarte, J.M.; et al. RCSB Protein Data Bank: Powerful new tools for exploring 3D structures of biological macromolecules for basic and applied research and education in fundamental biology, biomedicine, biotechnology, bioengineering and energy sciences. Nucleic Acids Res. 2021, 49, D437–D451. [Google Scholar] [CrossRef]
- Case, D.; Betz, R.; Cerutti, D.S.; Cheatham, T.; Darden, T.; Duke, R.; Giese, T.J.; Gohlke, H.; Götz, A.; Homeyer, N.; et al. Amber 16, University of California, San Francisco. 2016. Available online: https://doi.org/10.13140/RG.2.2.27958.70729 (accessed on 2 October 2022).
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A.; et al. PubChem Substance and Compound databases. Nucleic Acids Res. 2016, 44, D1202–D1213. [Google Scholar] [CrossRef] [PubMed]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rao, X.; Huang, X.; Zhou, Z.; Lin, X. An improvement of the 2ˆ(-delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat. Bioinform. Biomath. 2013, 3, 71–85. [Google Scholar]
- Cunningham, R.L.; Monk, K.R. Whole Mount In Situ Hybridization and Immunohistochemistry for Zebrafish Larvae. Methods Mol. Biol. 2018, 1739, 371–384. [Google Scholar] [CrossRef] [PubMed]
- Thisse, B.; Thisse, C. In situ hybridization on whole-mount zebrafish embryos and young larvae. Methods Mol. Biol. 2014, 1211, 53–67. [Google Scholar] [CrossRef] [PubMed]
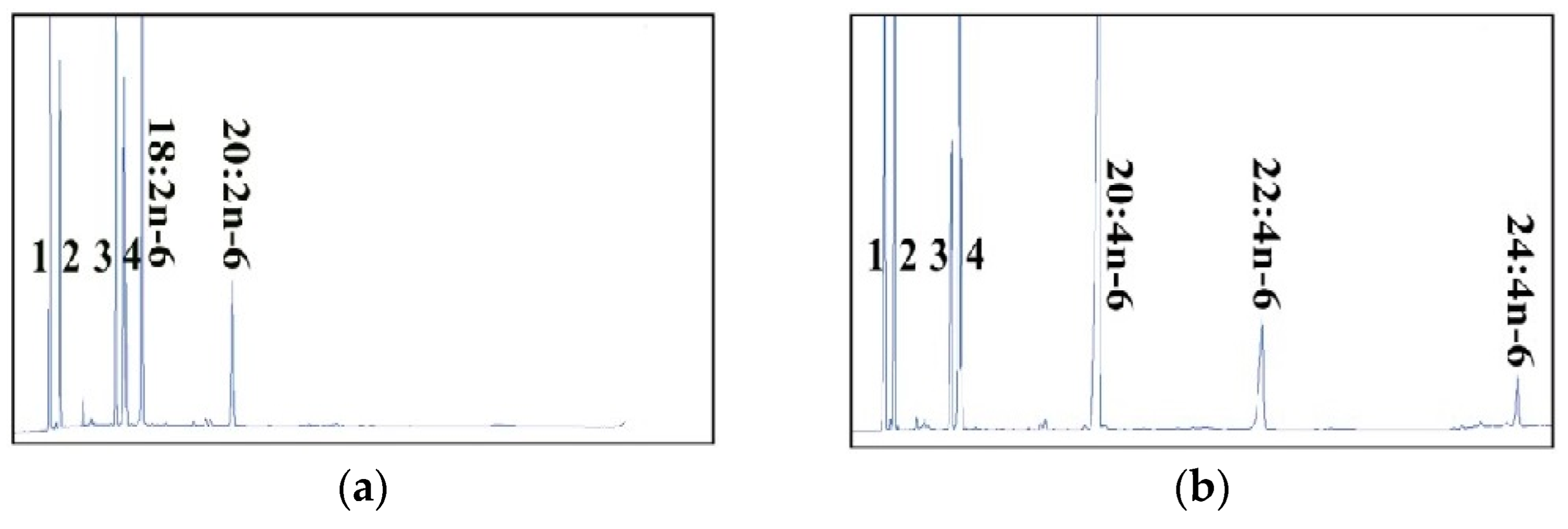
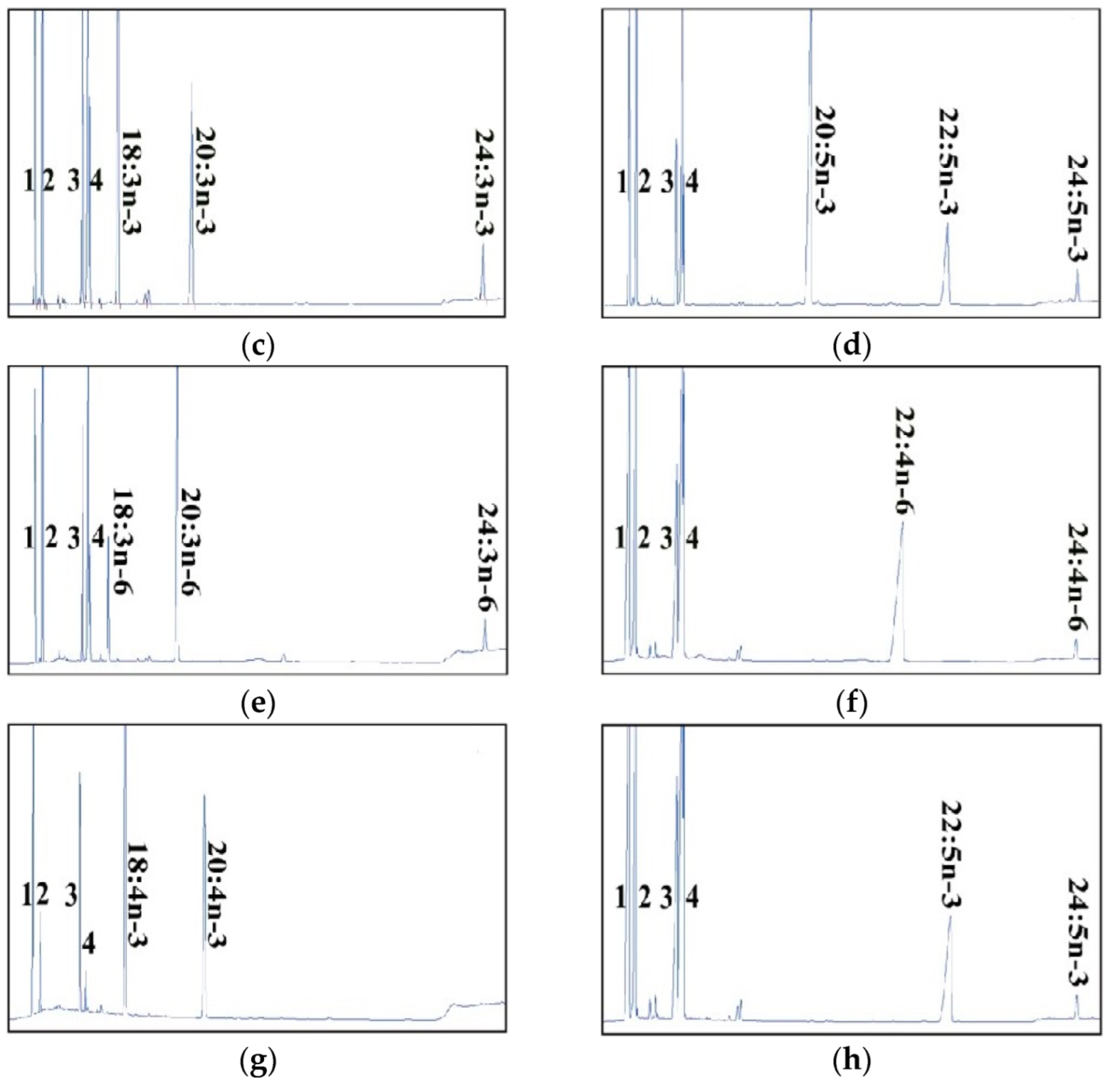
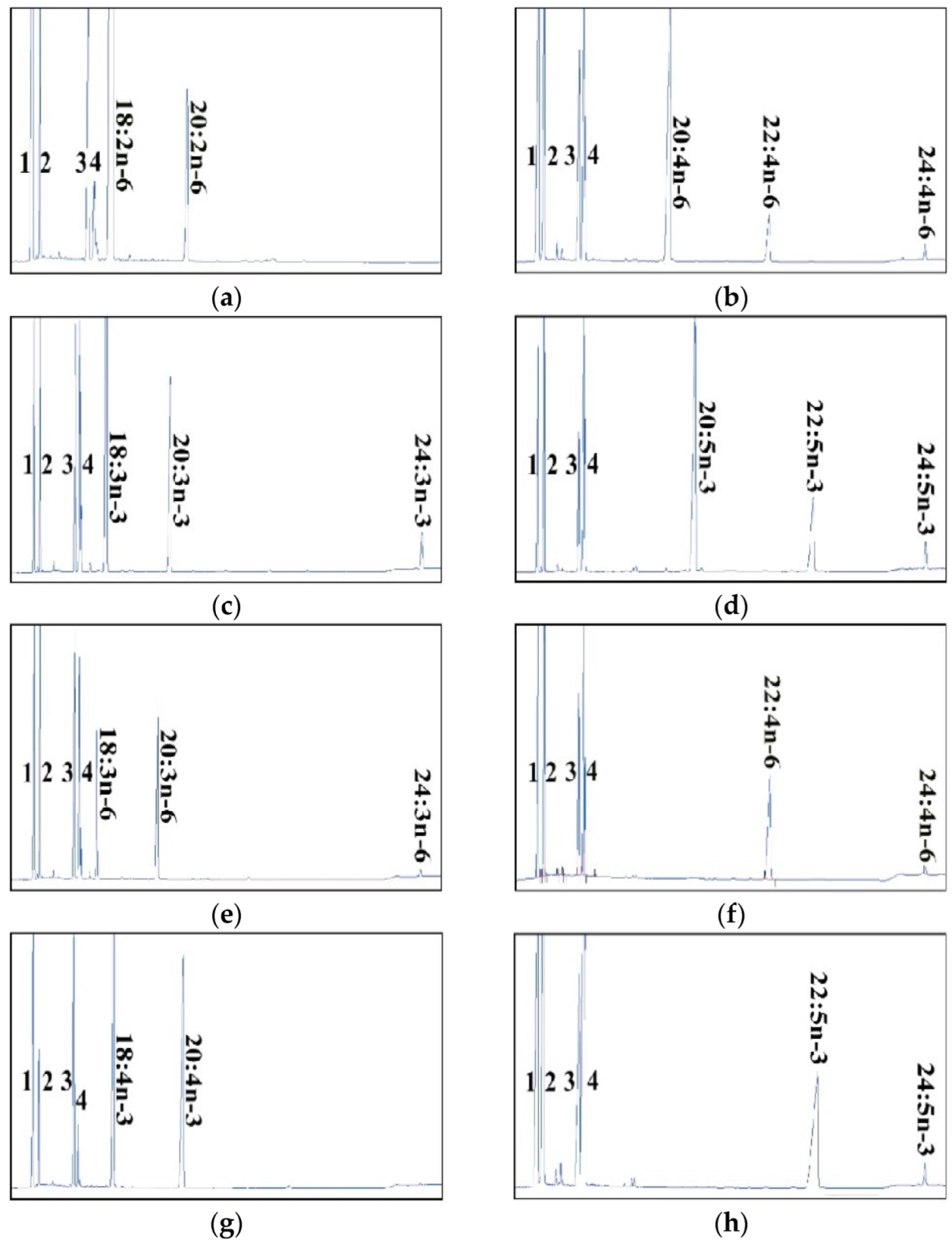

| FA Substrate | Product | Elovl5a Conversion (%) | Elovl5b Conversion (%) | Activity |
|---|---|---|---|---|
| C18:2n-6 | C20:2n-6 | 11.63 | 8.42 | C18→C20 |
| C18:3n-6 | C20:3n-6 | 74.24 | 69.63 | C18→C20 |
| C24:3n-6 | 8.45 | 1.14 | C22→C24 | |
| Total | 82.69 | 70.74 | ||
| C18:4n-3 | C20:4n-3 | 44.91 | 43.91 | C18→C20 |
| C18:3n-3 | C20:3n-3 | 36 | 22.97 | C18→C20 |
| C24:3n-3 | 9.2 | 3.9 | C22→C24 | |
| Total | 45.2 | 26.87 | ||
| C20:5n-3 | C22:5n-3 | 25.12 | 26.52 | C20→C22 |
| C24:5n-3 | 4.83 | 3.68 | C22→C24 | |
| Total | 29.95 | 30.2 | ||
| C20:4n-6 | C22:4n-6 | 17.90 | 13.29 | C20→C22 |
| C24:4n-6 | 5.02 | 2.92 | C22→C24 | |
| Total | 22.92 | 16.21 | ||
| C22:4n-6 | C24:4n-6 | 4.66 | 3.87 | C22→C24 |
| C22:5n-3 | C24:5n-3 | 8.24 | 6.00 | C22→C24 |
| FA Substrates | 0 hpf | 24 hpf | 48 hpf | 72 hpf | 96 hpf | 120 hpf |
|---|---|---|---|---|---|---|
| C18:2n-6 | 121.22 | 108.9 | 104.32 | 85.53 | 68.28 | 37.92 |
| C20:2n-6 | 13.26 | 11.67 | 11.9 | 10.99 | 10.39 | 7.91 |
| C20:3n-6 | 75.96 | 19.89 | 19.49 | 17.58 | 16.17 | 11.6 |
| C22:4n-6 | 12.3 | 11.26 | 10.87 | 10.18 | 10.42 | 8.12 |
| elovl5-related n-6 PUFAs | 222.74 | 151.72 | 146.58 | 124.28 | 105.26 | 65.55 |
| total n-6 PUFA products | 101.52 | 42.82 | 42.26 | 38.75 | 36.98 | 27.63 |
| n-6 products/substrate ratio | 0.84 | 0.39 | 0.405 | 0.453 | 0.54 | 0.73 |
| C18:3n-3 | 5.71 | 5.06 | 4.56 | 3.06 | 2.08 | 1.06 |
| C20:3n-3 | 0.94 | 1.08 | 1.02 | 0.85 | 0.83 | 0.71 |
| C20:4n-3 | 0.59 | 0.47 | 0.48 | 0.4 | 0.37 | 0.17 |
| C22:5n-3 | 1.47 | 3.7 | 3.54 | 3.86 | 3.65 | 3.04 |
| elovl5-related n-3 PUFAs | 8.71 | 10.31 | 9.6 | 8.17 | 6.93 | 4.98 |
| total n-3 PUFA products | 3 | 5.25 | 5.04 | 5.11 | 4.85 | 3.92 |
| n-3 products/substrate ratio | 0.525 | 1.04 | 1.11 | 1.67 | 2.33 | 3.70 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, R.; Wang, Y.-X.; Yang, C.-R.; Li, S.-Q.; Li, J.-C.; Sun, X.-Q.; Wang, H.-W.; Wang, Q.; Zhang, Y.; Li, J.-T. Dominant Elongase Activity of Elovl5a but Higher Expression of Elovl5b in Common Carp (Cyprinus carpio). Int. J. Mol. Sci. 2022, 23, 14666. https://doi.org/10.3390/ijms232314666
Zhao R, Wang Y-X, Yang C-R, Li S-Q, Li J-C, Sun X-Q, Wang H-W, Wang Q, Zhang Y, Li J-T. Dominant Elongase Activity of Elovl5a but Higher Expression of Elovl5b in Common Carp (Cyprinus carpio). International Journal of Molecular Sciences. 2022; 23(23):14666. https://doi.org/10.3390/ijms232314666
Chicago/Turabian StyleZhao, Ran, Ya-Xin Wang, Chen-Ru Yang, Shang-Qi Li, Jin-Cheng Li, Xiao-Qing Sun, Hong-Wei Wang, Qi Wang, Yan Zhang, and Jiong-Tang Li. 2022. "Dominant Elongase Activity of Elovl5a but Higher Expression of Elovl5b in Common Carp (Cyprinus carpio)" International Journal of Molecular Sciences 23, no. 23: 14666. https://doi.org/10.3390/ijms232314666





