Circular RNA Expression and Interaction Patterns Are Perturbed in Amyotrophic Lateral Sclerosis
Abstract
1. Introduction
2. Results
2.1. Differentially Expressed circRNAs
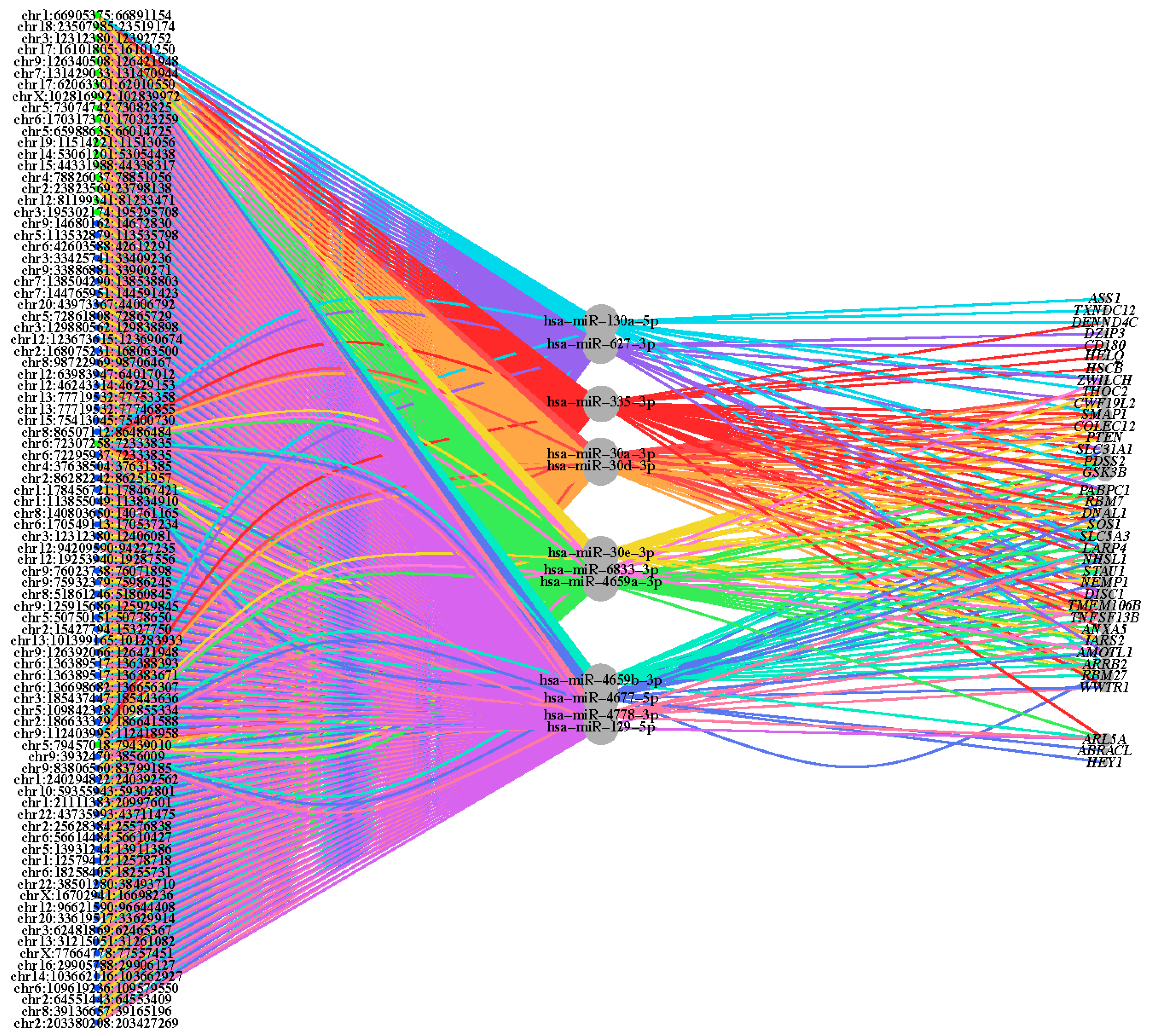
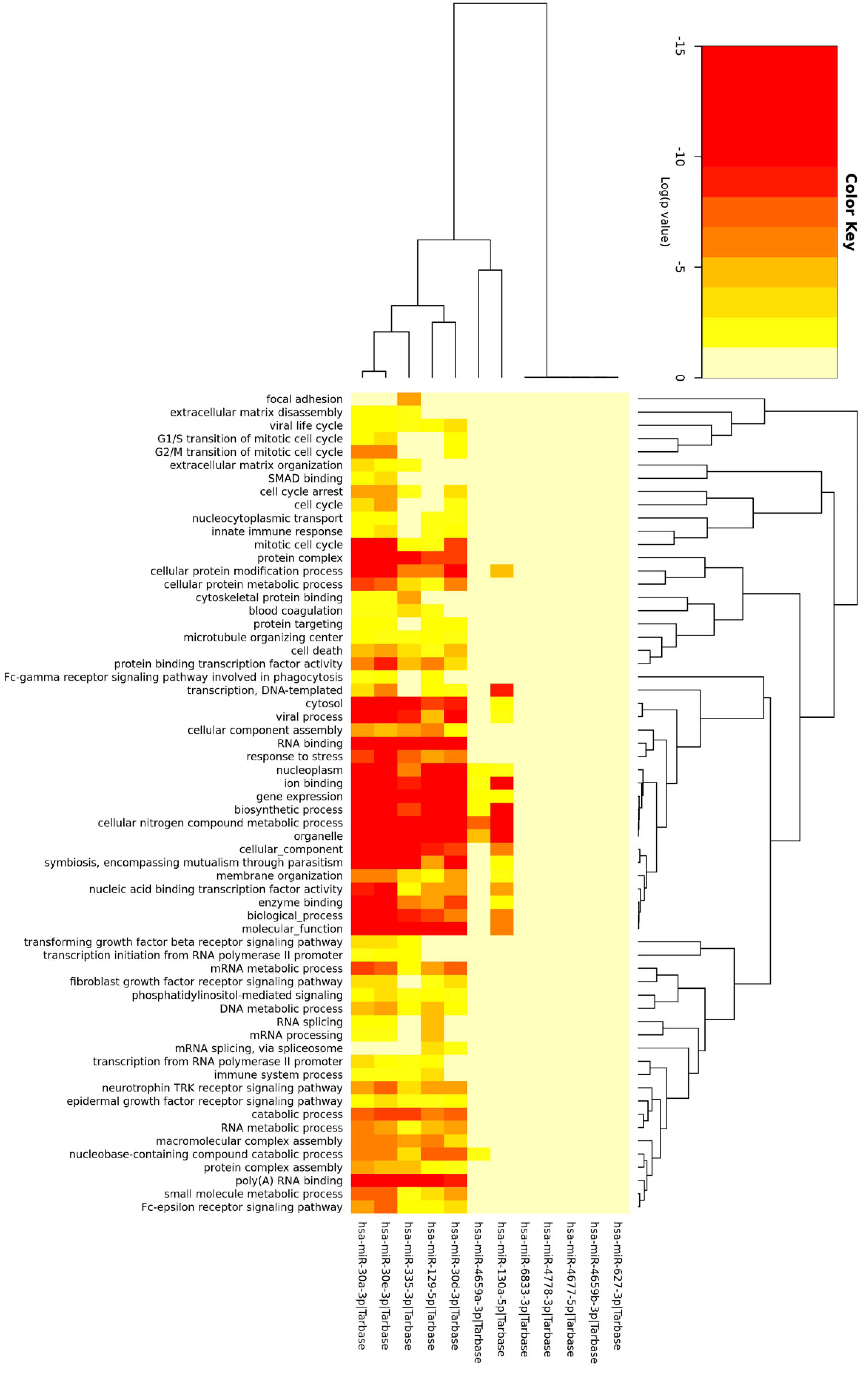
2.2. Binding Capacity of miRNAs to Differentially Expressed circRNAs
2.3. Comparison of Differentially Expressed circRNAs and mRNAs
3. Discussion
3.1. ALS-Specific Circular Transcriptome
3.2. circRNA‒miRNA‒mRNA Interactions
4. Materials and Methods
4.1. Dataset Structure and Quality Control
4.2. circRNA Detection and Differential Expression Analysis
4.3. Linear RNA Detection and Differential Expression Analysis
4.4. circRNA‒miRNA‒mRNA Interaction Analysis
4.5. Data Visualization
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Hardiman, O.; Al-Chalabi, A.; Brayne, C.; Beghi, E.; van den Berg, L.H.; Chio, A.; Martin, S.; Logroscino, G.; Rooney, J. The changing picture of amyotrophic lateral sclerosis: Lessons from European registers. J. Neurol. Neurosurg. Psychiatry 2017, 88, 557–563. [Google Scholar] [CrossRef]
- Chio, A.; Logroscino, G.; Traynor, B.J.; Collins, J.; Simeone, J.C.; Goldstein, L.A.; White, L.A. Global epidemiology of amyotrophic lateral sclerosis: A systematic review of the published literature. Neuroepidemiology 2013, 41, 118–130. [Google Scholar] [CrossRef]
- Vijayakumar, U.G.; Milla, V.; Stafford, M.Y.C.; Bjourson, A.J.; Duddy, W.; Duguez, S.M. A systematic review of suggested molecular strata, biomarkers and their tissue sources in ALS. Front. Neurol. 2019, 10, 400. [Google Scholar] [CrossRef]
- Brown, R.H.; Al-Chalabi, A. Amyotrophic lateral sclerosis. N. Engl. J. Med. 2017, 377, 162–172. [Google Scholar] [CrossRef]
- Jeck, W.R.; Sharpless, N.E. Detecting and characterizing circular RNAs. Nat. Biotechnol. 2014, 32, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Yang, E.; van Nimwegen, E.; Zavolan, M.; Rajewsky, N.; Schroeder, M.; Magnasco, M.; Darnell, J.E., Jr. Decay rates of human mRNAs: Correlation with functional characteristics and sequence attributes. Genome Res. 2003, 13, 1863–1872. [Google Scholar] [CrossRef]
- Zhang, Z.; Yang, T.; Xiao, J. Circular RNAs: Promising biomarkers for human diseases. eBioMedicine 2018, 34, 267–274. [Google Scholar] [CrossRef]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef]
- Xu, H.; Zhang, Y.; Qi, L.; Ding, L.; Jiang, H.; Yu, H. NFIX circular RNA promotes glioma progression by regulating miR-34a-5p via notch signaling pathway. Front. Mol. Neurosci. 2018, 11, 225. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Alexandrov, P.N.; Jaber, V.; Lukiw, W.J. Deficiency in the ubiquitin conjugating enzyme UBE2A in Alzheimer’s disease (AD) is linked to deficits in a natural circular miRNA-7 sponge (circRNA; ciRS-7). Genes 2016, 7, 116. [Google Scholar] [CrossRef]
- Sang, Q.; Liu, X.; Wang, L.; Qi, L.; Sun, W.; Wang, W.; Sun, Y.; Zhang, H. CircSNCA downregulation by pramipexole treatment mediates cell apoptosis and autophagy in Parkinson’s disease by targeting miR-7. Aging 2018, 10, 1281–1293. [Google Scholar] [CrossRef]
- Dolinar, A.; Koritnik, B.; Glavac, D.; Ravnik-Glavac, M. Circular RNAs as potential blood biomarkers in amyotrophic lateral sclerosis. Mol. Neurobiol. 2019, 56, 8052–8062. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Friedman, R.C.; Farh, K.K.; Burge, C.B.; Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009, 19, 92–105. [Google Scholar] [CrossRef]
- Grimson, A.; Farh, K.K.; Johnston, W.K.; Garrett-Engele, P.; Lim, L.P.; Bartel, D.P. MicroRNA targeting specificity in mammals: Determinants beyond seed pairing. Mol. Cell 2007, 27, 91–105. [Google Scholar] [CrossRef]
- Garcia, D.M.; Baek, D.; Shin, C.; Bell, G.W.; Grimson, A.; Bartel, D.P. Weak seed-pairing stability and high target-site abundance decrease the proficiency of lsy-6 and other microRNAs. Nat. Struct. Mol. Biol. 2011, 18, 1139–1146. [Google Scholar] [CrossRef]
- Zhao, M.; Kim, J.R.; van Bruggen, R.; Park, J. RNA-binding proteins in amyotrophic lateral sclerosis. Mol. Cells 2018, 41, 818–829. [Google Scholar] [CrossRef]
- Kim, H.J.; Taylor, J.P. Lost in transportation: Nucleocytoplasmic transport defects in ALS and other neurodegenerative diseases. Neuron 2017, 96, 285–297. [Google Scholar] [CrossRef]
- Tyzack, G.E.; Luisier, R.; Taha, D.M.; Neeves, J.; Modic, M.; Mitchell, J.S.; Meyer, I.; Greensmith, L.; Newcombe, J.; Ule, J.; et al. Widespread FUS mislocalization is a molecular hallmark of amyotrophic lateral sclerosis. Brain 2019, 142, 2572–2580. [Google Scholar] [CrossRef]
- Jain, M.R.; Ge, W.W.; Elkabes, S.; Li, H. Amyotrophic lateral sclerosis: Protein chaperone dysfunction revealed by proteomic studies of animal models. Proteom. Clin. Appl. 2008, 2, 670–684. [Google Scholar] [CrossRef] [PubMed]
- Ranganathan, S.; Bowser, R. Alterations in G1 to S phase cell-cycle regulators during amyotrophic lateral sclerosis. Am. J. Pathol. 2003, 162, 823–835. [Google Scholar] [CrossRef] [PubMed]
- Pradhan, J.; Noakes, P.G.; Bellingham, M.C. The role of altered BDNF/TrkB signaling in amyotrophic lateral sclerosis. Front. Cell. Neurosci. 2019, 13, 368. [Google Scholar] [CrossRef]
- Karagkouni, D.; Paraskevopoulou, M.D.; Chatzopoulos, S.; Vlachos, I.S.; Tastsoglou, S.; Kanellos, I.; Papadimitriou, D.; Kavakiotis, I.; Maniou, S.; Skoufos, G.; et al. DIANA-TarBase v8: A decade-long collection of experimentally supported miRNA-gene interactions. Nucleic Acids Res. 2018, 46, D239–D245. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Guan, Y.; Zhang, Z.; Liu, H.; Wang, S.; Yu, L.; Wu, X.; Wang, X. Wnt signaling pathway is involved in the pathogenesis of amyotrophic lateral sclerosis in adult transgenic mice. Neurol. Res. 2012, 34, 390–399. [Google Scholar] [CrossRef]
- Momtaz, S.; Memariani, Z.; El-Senduny, F.F.; Sanadgol, N.; Golab, F.; Katebi, M.; Abdolghaffari, A.H.; Farzaei, M.H.; Abdollahi, M. Targeting ubiquitin-proteasome pathway by natural products: Novel therapeutic strategy for treatment of neurodegenerative diseases. Front. Physiol. 2020, 11, 361. [Google Scholar] [CrossRef]
- Aliabadi, F.; Sohrabi, B.; Mostafavi, E.; Pazoki-Toroudi, H.; Webster, T.J. Ubiquitin-proteasome system and the role of its inhibitors in cancer therapy. Open Biol. 2021, 11, 200390. [Google Scholar] [CrossRef]
- Gong, B.; Radulovic, M.; Figueiredo-Pereira, M.E.; Cardozo, C. The ubiquitin-proteasome system: Potential therapeutic targets for Alzheimer’s disease and spinal cord injury. Front. Mol. Neurosci. 2016, 9, 4. [Google Scholar] [CrossRef]
- Frankish, A.; Diekhans, M.; Ferreira, A.-M.; Johnson, R.; Jungreis, I.; Loveland, J.; Mudge, J.M.; Sisu, C.; Wright, J.; Armstrong, J.; et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 2019, 47, D766–D773. [Google Scholar] [CrossRef]
- PVC (Research Infrastructure). Katana; UNSW: Sydney, Australia, 2010. [Google Scholar]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Hoffman, G.E.; Schadt, E.E. variancePartition: Interpreting drivers of variation in complex gene expression studies. BMC Bioinform. 2016, 17, 483. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data; Babraham Institute: Cambridge, UK, 2010. [Google Scholar]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Zhang, J.; Zhao, F. Circular RNA identification based on multiple seed matching. Brief. Bioinform. 2017, 19, 803–810. [Google Scholar] [CrossRef]
- Zhang, X.-O.; Dong, R.; Zhang, Y.; Zhang, J.-L.; Luo, Z.; Zhang, J.; Chen, L.-L.; Yang, L. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 2016, 26, 1277–1287. [Google Scholar] [CrossRef] [PubMed]
- R-Core-Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Pertea, M.; Kim, D.; Pertea, G.M.; Leek, J.T.; Salzberg, S.L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and ballgown. Nat. Protoc. 2016, 11, 1650–1667. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Aufiero, S.; Reckman, Y.J.; Tijsen, A.J.; Pinto, Y.M.; Creemers, E.E. circRNAprofiler: An R-based computational framework for the downstream analysis of circular RNAs. BMC Bioinform. 2020, 21, 164. [Google Scholar] [CrossRef] [PubMed]
- Acton, R.M. scrapeR: Tools for Scraping Data from HTML and XML Documents; University of California: Irvine, CA, USA, 2010. [Google Scholar]
- Xu, T.; Su, N.; Liu, L.; Zhang, J.; Wang, H.; Zhang, W.; Gui, J.; Yu, K.; Li, J.; Le, T.D. miRBaseConverter: An R/Bioconductor package for converting and retrieving miRNA name, accession, sequence and family information in different versions of miRBase. BMC Bioinform. 2018, 19, 514. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. Stringr: Simple, Consistent Wrappers for Common String Operations; CRAN: Vienna, Austria, 2019. [Google Scholar]
- Vlachos, I.S.; Zagganas, K.; Paraskevopoulou, M.D.; Georgakilas, G.; Karagkouni, D.; Vergoulis, T.; Dalamagas, T.; Hatzigeorgiou, A.G. DIANA-miRPath v3.0: Deciphering microRNA function with experimental support. Nucleic Acids Res. 2015, 43, W460–W466. [Google Scholar] [CrossRef] [PubMed]
- Durinck, S.; Moreau, Y.; Kasprzyk, A.; Davis, S.; De Moor, B.; Brazma, A.; Huber, W. BioMart and bioconductor: A powerful link between biological databases and microarray data analysis. Bioinformatics 2005, 21, 3439–3440. [Google Scholar] [CrossRef]
- Durinck, S.; Spellman, P.T.; Birney, E.; Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 2009, 4, 1184–1191. [Google Scholar] [CrossRef]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Liberzon, A.; Subramanian, A.; Pinchback, R.; Thorvaldsdottir, H.; Tamayo, P.; Mesirov, J.P. Molecular signatures database (MSigDB) 3.0. Bioinformatics 2011, 27, 1739–1740. [Google Scholar] [CrossRef]
- Liberzon, A.; Birger, C.; Thorvaldsdottir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 2015, 1, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Krijthe, J.H. Rtsne: T-Distributed Stochastic Neighbor Embedding Using a Barnes-Hut Implementation; TU Delft: Delft, The Netherlands, 2015. [Google Scholar]

| Tissue | Condition | Number of Samples | Number of circRNAs Detected | Number of Detected circRNAs Common to ALS and Control Samples | Number of circRNAs Unique to the Condition |
|---|---|---|---|---|---|
| Spinal Cord Cervical | ALS | 18 | 24,259 | 12,557 | 11,702 |
| Control | 7 | 15,210 | 2653 | ||
| Spinal Cord Lumbar | ALS | 16 | 24,020 | 14,044 | 9976 |
| Control | 6 | 18,262 | 4218 | ||
| Spinal Cord Thoracic | ALS | 16 | 20,581 | 12,100 | 8481 |
| Control | 6 | 16,209 | 4109 |
| Spinal Cord Cervical | Spinal Cord Lumbar | Spinal Cord Thoracic | ||
|---|---|---|---|---|
| Number of DE genes (adjusted p value < 0.05) | Upregulated | 1944 | 524 | 242 |
| Downregulated | 1052 | 460 | 90 | |
| Total | 2996 | 984 | 332 | |
| Total genes tested | 60,662 | 60,662 | 60,662 | |
| GO Accession and Name | Number of Genes in Overlap | p Value |
|---|---|---|
| GO:1901699: Cellular response to nitrogen compound | 7 | 2.97 × 10−6 |
| GO:0198738: Cell‒cell signaling by wnt | 6 | 6.94 × 10−6 |
| GO:0050806: Positive regulation of synaptic transmission | 4 | 9.41 × 10−6 |
| GO:1901701: Cellular response to oxygen-containing compound | 8 | 9.67 × 10−6 |
| GO:0071375: Cellular response to peptide hormone stimulus | 5 | 1.16 × 10−5 |
| GO:0001837: Epithelial to mesenchymal transition | 4 | 1.20 × 10−5 |
| GO:0051100: Negative regulation of binding | 4 | 1.39 × 10−5 |
| GO:1905114: cell surface receptor signaling pathway involved in cell‒cell signaling | 6 | 1.88 × 10−5 |
| GO:0032870: Cellular response to hormone stimulus | 6 | 1.98 × 10−5 |
| GO:1901653: Cellular response to peptide | 5 | 2.83 × 10−5 |
| Tissue | No. of Samples in ALS Cohort | No. of Samples in Control Cohort |
|---|---|---|
| Spinal Cord Cervical | 18 | 7 |
| Spinal Cord Thoracic | 16 | 6 |
| Spinal Cord Lumbar | 16 | 6 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aquilina-Reid, C.; Brennan, S.; Curry-Hyde, A.; Teunisse, G.M.; The NYGC ALS Consortium; Janitz, M. Circular RNA Expression and Interaction Patterns Are Perturbed in Amyotrophic Lateral Sclerosis. Int. J. Mol. Sci. 2022, 23, 14665. https://doi.org/10.3390/ijms232314665
Aquilina-Reid C, Brennan S, Curry-Hyde A, Teunisse GM, The NYGC ALS Consortium, Janitz M. Circular RNA Expression and Interaction Patterns Are Perturbed in Amyotrophic Lateral Sclerosis. International Journal of Molecular Sciences. 2022; 23(23):14665. https://doi.org/10.3390/ijms232314665
Chicago/Turabian StyleAquilina-Reid, Chiara, Samuel Brennan, Ashton Curry-Hyde, Guus M. Teunisse, The NYGC ALS Consortium, and Michael Janitz. 2022. "Circular RNA Expression and Interaction Patterns Are Perturbed in Amyotrophic Lateral Sclerosis" International Journal of Molecular Sciences 23, no. 23: 14665. https://doi.org/10.3390/ijms232314665
APA StyleAquilina-Reid, C., Brennan, S., Curry-Hyde, A., Teunisse, G. M., The NYGC ALS Consortium, & Janitz, M. (2022). Circular RNA Expression and Interaction Patterns Are Perturbed in Amyotrophic Lateral Sclerosis. International Journal of Molecular Sciences, 23(23), 14665. https://doi.org/10.3390/ijms232314665







