Comprehensive Transcriptome Analysis Reveals Genome-Wide Changes Associated with Endoplasmic Reticulum (ER) Stress in Potato (Solanum tuberosum L.)
Abstract
1. Introduction
2. Results
2.1. Expression Profiles of Genes Involved in the Early ER Stress Response
2.2. Chromatin Modifications and Altered Gene Regulation Are Early Stress Responses to TM Treatment
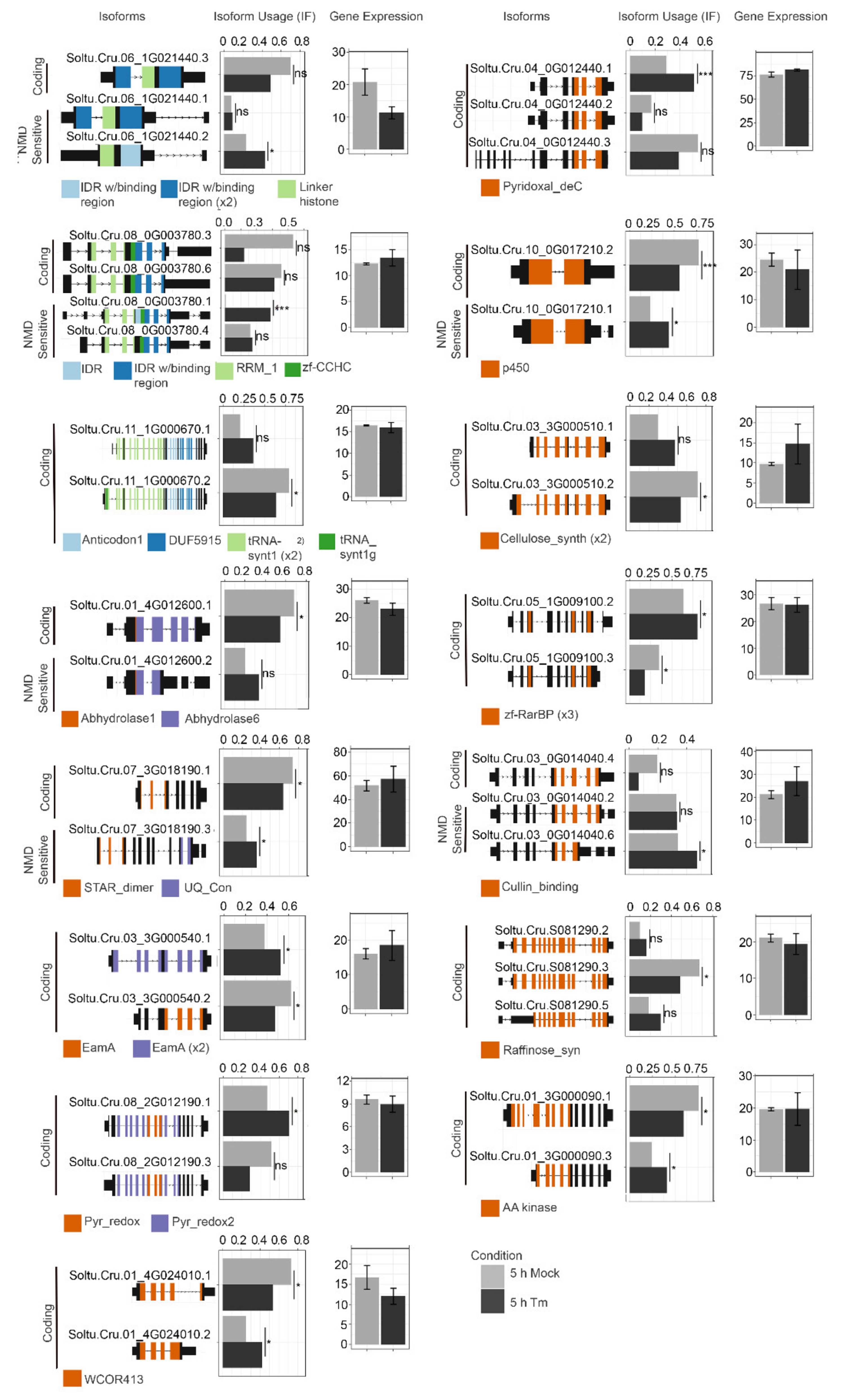
2.3. Changes in the Accumulation of Transcript Isoforms and Genes Involved in RNA Metabolism Are Seen in TM-Treated Leaves
2.4. Gene Expression Involving Protein Maturation and Degradation Pathways
2.5. Changes in Oxygen Metabolic Enzymes and Protein Kinases
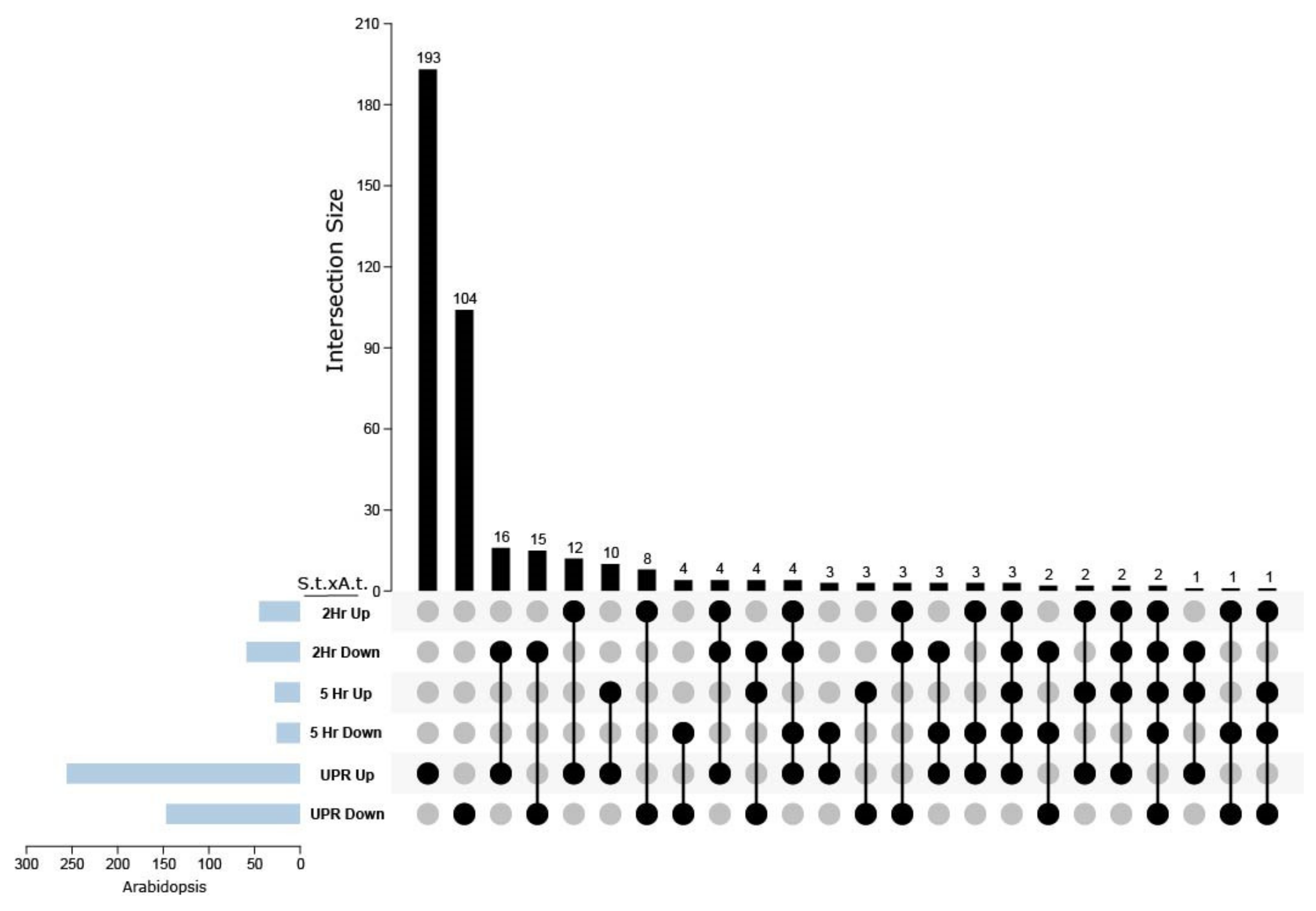
2.6. DEGs That Are Common between Potato and Arabidopsis Treated with TM
3. Discussion
4. Materials and Methods
4.1. Plant Material and TM Treatment
4.2. Transcriptomic Analysis
4.3. Gene Annotation and Characterization
4.4. Isoform Analysis
4.5. Comparative Analysis of TM-Induced DEG in Potato and Arabidopsis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gülow, K.; Bienert, D.; Haas, I.G. BiP Is Feed-Back Regulated by Control of Protein Translation Efficiency. J. Cell Sci. 2002, 115, 2443–2452. [Google Scholar] [CrossRef] [PubMed]
- Ellgaard, L.; McCaul, N.; Chatsisvili, A.; Braakman, I. Co- and Post-Translational Protein Folding in the ER. Traffic 2016, 17, 615–638. [Google Scholar] [CrossRef]
- Nagashima, Y.; Mishiba, K.; Suzuki, E.; Shimado, Y.; Iwata, Y.; Koizumi, N. Arabidopsis IRE1 catalyzes unconventional splicing of bZIP60 mRNA to produce the active transcription factor. Sci. Reports 2011, 1, 1–29. [Google Scholar] [CrossRef]
- Urquidi Camacho, R.A.; Lokdarshi, A.; von Arnim, A.G. Translational Gene Regulation in Plants: A Green New Deal. Wiley Interdiscip. Rev. RNA 2020, 11, e1597. [Google Scholar] [CrossRef]
- Afrin, T.; Diwan, D.; Sahawneh, K.; Pajerowska-Mukhtar, K. Multilevel Regulation of Endoplasmic Reticulum Stress Re-sponses in Plants: Where Old Roads and New Paths Meet. J. Exp. Bot. 2020, 71, 1659–1667. [Google Scholar] [CrossRef] [PubMed]
- Iwawaki, T.; Hosoda, A.; Okuda, T.; Kamigori, Y.; Nomura-Furuwatari, C.; Kimata, Y.; Tsuru, A.; Kohno, K. Translational Control by the ER Transmembrane Kinase/Ribonuclease IRE1 under ER Stress. Nat. Cell Biol. 2001, 3, 158–164. [Google Scholar] [CrossRef] [PubMed]
- Mishiba, K.I.; Nagashima, Y.; Suzukia, E.; Hayashi, N.; Ogata, Y.; Shimada, Y.; Koizumi, N. Defects in IRE1 Enhance Cell Death and Fail to Degrade MRNAs Encoding Secretory Pathway Proteins in the Arabidopsis Unfolded Protein Response. Proc. Natl. Acad. Sci. USA 2013, 110, 5713–5718. [Google Scholar] [CrossRef] [PubMed]
- Tajima, H.; Iwata, Y.; Iwano, M.; Takayama, S.; Koizumi, N. Identification of an Arabidopsis Transmembrane BZIP Tran-scription Factor Involved in the Endoplasmic Reticulum Stress Response. Biochem. Biophys. Res. Commun. 2008, 374, 242–247. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.-S.; Yamaguchi-Shinozaki, K.; Shinozaki, K. ER-Anchored Transcription Factors BZIP17 and BZIP28 Regulate Root Elongation. Plant Physiol. 2018, 176, 2221–2230. [Google Scholar] [CrossRef]
- Srivastava, R.; Deng, Y.; Shah, S.; Rao, A.G.; Howell, S.H. BINDING PROTEIN Is a Master Regulator of the Endoplasmic Re-ticulum Stress Sensor/Transducer BZIP28 in Arabidopsis. Plant Cell 2013, 25, 1416–1429. [Google Scholar] [CrossRef]
- Ruberti, C.; Lai, Y.S.; Brandizzi, F. Recovery from Temporary Endoplasmic Reticulum Stress in Plants Relies on the Tis-sue-Specific and Largely Independent Roles of BZIP28 and BZIP60, as Well as an Antagonizing Function of BAX-Inhibitor 1 upon the pro-Adaptive Signaling Mediated by BZIP28. Plant J. 2018, 93, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Iwata, Y.; Ashida, M.; Hasegawa, C.; Tabara, K.; Mishiba, K.; Koizumi, N. Activation of the Arabidopsis Membrane-Bound Transcription Factor BZIP28 Is Mediated by Site-2 Protease, but Not Site-1 Protease. Plant J. 2017, 91, 408–415. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Lu, S.J.; Zhang, S.S.; Zhou, S.F.; Sun, L.; Liu, J.X. The Lumen-Facing Domain Is Important for the Biological Function and Organelle-to-Organelle Movement of BZIP28 during ER Stress in Arabidopsis. Mol. Plant 2013, 6, 1605–1615. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.X.; Howell, S.H. BZIP28 and NF-Y Transcription Factors Are Activated by ER Stress and Assemble into a Transcrip-tional Complex to Regulate Stress Response Genes in Arabidopsis. Plant Cell 2010, 22, 782–796. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, R.; Deng, Y.; Howell, S.H. Stress Sensing in Plants by an ER Stress Sensor/Transducer, BZIP28. Front Plant Sci. 2014, 5, 59. [Google Scholar] [CrossRef]
- Ishikawa, T.; Watanabe, N.; Nagano, M.; Kawai-Yamada, M.; Lam, E. Bax Inhibitor-1: A Highly Conserved Endoplasmic Reticulum-Resident Cell Death Suppressor. Cell Death Differ. 2011, 18, 1271–1278. [Google Scholar] [CrossRef]
- Liu, J.X.; Howell, S.H. Managing the Protein Folding Demands in the Endoplasmic Reticulum of Plants. New Phytol. 2016, 211, 418–428. [Google Scholar] [CrossRef]
- Chen, Y.; Brandizzi, F. IRE1: ER Stress Sensor and Cell Fate Executor. Trends Cell Biol. 2013, 23, 547–555. [Google Scholar] [CrossRef]
- Srivastava, R.; Li, Z.; Russo, G.; Tang, J.; Bi, R.; Muppirala, U.; Chudalayandi, S.; Severin, A.; He, M.; Vaitkevicius, S.I.; et al. Response to Persistent ER Stress in Plants: A Multiphasic Process That Transitions Cells from Prosurvival Activities to Cell Death. Plant Cell 2018, 30, 1220–1242. [Google Scholar] [CrossRef]
- Iwata, Y.; Koizumi, N. Unfolded Protein Response Followed by Induction of Cell Death in Cultured Tobacco Cells Treated with Tunicamycin. Planta 2005, 220, 804–807. [Google Scholar] [CrossRef]
- Kamauchi, S.; Nakatani, H.; Nakano, C.; Urade, R. Gene Expression in Response to Endoplasmic Reticulum Stress in Arabidopsis Thaliana. FEBS J. 2005, 272, 3461–3476. [Google Scholar] [CrossRef] [PubMed]
- Chawla, A.; Chakrabarti, S.; Ghosh, G.; Niwa, M. Attenuation of Yeast UPR Is Essential for Survival and Is Mediated by IRE1 Kinase. J. Cell Biol. 2011, 193, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Brandizzi, F. AtIRE1A/AtIRE1B and AGB1 Independently Control Two Essential Unfolded Protein Response Pathways in Arabidopsis. Plant J. 2012, 69, 266–277. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Pu, Y.; Yu, X.; Gregory, B.D.; Srivastava, R.; Howell, S.H.; Bassham, D.C. IRE1B Degrades RNAs Encoding Proteins That Interfere with the Induction of Autophagy by ER Stress in Arabidopsis Thaliana. Autophagy 2018, 14, 1562–1573. [Google Scholar] [CrossRef]
- Bao, Y.; Howell, S.H. The Unfolded Protein Response Supports Plant Development and Defense as Well as Responses to Abi-otic Stress. Front Plant Sci. 2017, 8, 344. [Google Scholar] [CrossRef]
- Iwata, Y.; Nishino, T.; Takayama, S.; Koizumi, N. Characterization of a Plant-Specific Gene Induced by Endoplasmic Reticu-lum Stress in Arabidopsis Thaliana. Biosci. Biotechnol. Biochem. 2010, 74, 2087–2091. [Google Scholar] [CrossRef]
- Iwata, Y.; Sakiyama, M.; Lee, M.H.; Koizumi, N. Transcriptomic Response of Arabidopsis Thaliana to Tunicamycin Induced Endoplasmic Reticulum Stress. Plant Biotechnol. 2010, 27, 161–171. [Google Scholar] [CrossRef]
- Travers, K.J.; Patil, C.K.; Wodicka, L.; Lockhart, D.J.; Weissman, J.S.; Walter, P. Functional and Genomic Analyses Reveal an Essential Coordination between the Unfolded Protein Response and ER-Associated Degradation. Cell 2000, 101, 249–258. [Google Scholar] [CrossRef]
- Melo, B.P.; Fraga, O.T.; Silva, J.C.F.; Ferreira, D.O.; Brustolini, O.J.B.; Carpinetti, P.A.; Machado, J.P.B.; Reis, P.A.B.; Fontes, E.P.B. Revisiting the Soybean GmNAC Superfamily. Front Plant Sci. 2018, 871, 1–22. [Google Scholar] [CrossRef]
- Wakasa, Y.; Oono, Y.; Yazawa, T.; Hayashi, S.; Ozawa, K.; Handa, H.; Matsumoto, T.; Takaiwa, F. RNA Sequenc-ing-Mediated Transcriptome Analysis of Rice Plants in Endoplasmic Reticulum Stress Conditions. BMC Plant Biol. 2014, 14, 101. [Google Scholar] [CrossRef]
- Ruberti, C.; Kim, S.-J.; Stefano, G.; Brandizzi, F. Unfolded Protein Response in Plants: One Master, Many Questions. Curr. Opin. Plant Biol. 2015, 27, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Kyriakidou, M.; Anglin, N.L.; Ellis, D.; Tai, H.H.; Stromvik, M.V. Genome Assembly of Six Polyploid Potato genomes. Sci. Data 2020, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Pan, S.; Cheng, S.; Zhang, B.; Mu, D.; Ni, P.; Zhang, G.; Yang, S.; Li, R.; Wang, J.; et al. Genome Sequence and Analysis of the Tuber Crop Potato. Nature 2011, 475, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Zhang, Y.; Cai, G.; Qing, Y.; Song, J.; Wang, H.; Tan, X.; Liu, C.; Yang, M.; Fang, Z.; et al. Genome Assembly of Primitive Cultivated Potato Solanum Stenotomum Provides Insights into Potato Evolution. G3 Genes Genomes Genet. 2021, 11, jkab262. [Google Scholar] [CrossRef]
- Lihodeevskiy, G.A.; Shanina, E.P. Structural Variations in the Genome of Potato Varieties of the Ural Selection. Agronomy 2021, 111, 1703. [Google Scholar] [CrossRef]
- Petek, M.; Zagorščak, M.; Ramšak, Ž.; Sanders, S.; Tomaž, Š.; Tseng, E.; Zouine, M.; Coll, A.; Gruden, K. Cultivar-Specific Transcriptome and Pan-Transcriptome Reconstruction of Tetraploid Potato. Sci. Data 2020, 7, 249. [Google Scholar] [CrossRef]
- Herath, V.; Gayral, M.; Adhikari, N.; Miller, R.; Verchot, J. Genome-Wide Identification and Characterization of Solanum Tuberosum BiP Genes Reveal the Role of the Promoter Architecture in BiP Gene Diversity. Sci. Rep. 2020, 10, 11327. [Google Scholar] [CrossRef]
- Herath, V.; Verchot, J. Insight into the BZIP Gene Family in Solanum Tuberosum: Genome and Transcriptome Analysis to Understand the Roles of Gene Diversification in Spatiotemporal Gene Expression and Function. Int. J. Mol. Sci. 2020, 22, 253. [Google Scholar] [CrossRef]
- Jakoby, M.; Weisshaar, B.; Dröge-Laser, W.; Vicente-Carbajosa, J.; Tiedemann, J.; Kroj, T.; Parcy, F. BZIP Transcription Factors in Arabidopsis. Trends Plant Sci. 2002, 7, 106–111. [Google Scholar] [CrossRef]
- Dröge-Laser, W.; Snoek, B.L.; Snel, B.; Weiste, C. The Arabidopsis BZIP Transcription Factor Family—An Update. Curr. Opin. Plant Biol. 2018, 45, 36–49. [Google Scholar] [CrossRef]
- Jindrich, K.; Degnan, B.M. The Diversification of the Basic Leucine Zipper Family in Eukaryotes Correlates with the Evolu-tion of Multicellularity Genome Evolution and Evolutionary Systems Biology. BMC Evol. Biol. 2016, 16, 28. [Google Scholar] [CrossRef] [PubMed]
- Gayral, M.; Arias Gaguancela, O.; Vasquez, E.; Herath, V.; Flores, F.J.; Dickman, M.B.; Verchot, J. Multiple ER-to-nucleus Stress Signaling Pathways Are Activated during Plantago Asiatica Mosaic Virus and Turnip Mosaic Virus Infection in Arabidopsis Thaliana. Plant J. 2020, 103, 1233–1245. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.-T.; Sun, L.; Lu, S.-J.; Tian, Y.; Ding, Y.; Liu, J.-X. Transcription Factor Interaction with COMPASS-like Complex Regu-lates Histone H3K4 Trimethylation for Specific Gene Expression in Plants. Proc. Natl. Acad. Sci. USA 2015, 112, 2900–2905. [Google Scholar] [CrossRef] [PubMed]
- Ye, C.-M.; Kelly, V.; Payton, M.; Dickman, M.B.; Verchot, J. SGT1 Is Induced by the Potato Virus X TGBp3 and Enhances Virus Accumulation in Nicotiana Benthamiana. Mol. Plant 2012, 5, 1151–1153. [Google Scholar] [CrossRef] [PubMed]
- Götz, S.; García-Gómez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talón, M.; Dopazo, J.; Conesa, A. High-Throughput Functional Annotation and Data Mining with the Blast2GO Suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef]
- Rashid, H.O.; Yadav, R.K.; Kim, H.R.; Chae, H.J. ER Stress: Autophagy Induction, Inhibition and Selection. Autophagy 2015, 11, 1956–1977. [Google Scholar] [CrossRef]
- Sharma, M.; Bhattacharyya, S.; Sharma, K.B.; Chauhan, S.; Asthana, S.; Abdin, M.Z.; Vrati, S.; Kalia, M. Japanese Encephalitis Virus Activates Autophagy through XBP1 and ATF6 ER Stress Sensors in Neuronal Cells. J. Gen. Virol. 2017, 98, 1027–1039. [Google Scholar] [CrossRef]
- Fung, T.S.; Liu, D.X. The ER Stress Sensor IRE1 and MAP Kinase ERK Modulate Autophagy Induction in Cells Infected with Coronavirus Infectious Bronchitis Virus. Virology 2019, 533, 34–44. [Google Scholar] [CrossRef]
- Li, Y.; Dickman, M. Processing of AtBAG6 Triggers Autophagy and Fungal Resistance. Plant Signal Behav. 2016, 11, 233–247. [Google Scholar] [CrossRef]
- Bernales, S.; McDonald, K.L.; Walter, P. Autophagy Counterbalances Endoplasmic Reticulum Expansion during the Unfold-ed Protein Response. PLoS Biol. 2006, 4, e423. [Google Scholar] [CrossRef]
- Yang, X.; Srivastava, R.; Howell, S.H.; Bassham, D.C. Activation of Autophagy by Unfolded Proteins during Endoplasmic Reticulum Stress. Plant J. 2016, 85, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Jenuwein, T.; Allis, C.D. Translating the Histone Code. Science 2001, 293, 1074–1080. [Google Scholar] [CrossRef] [PubMed]
- Rea, S.; Eisenhaber, F.; O’Carroll, D.; Strahl, B.D.; Sun, Z.W.; Schmid, M.; Opravil, S.; Mechtier, K.; Ponting, C.P.; Allis, C.D.; et al. Regulation of Chromatin Structure by Site-Specific Histone H3 Methyltransferases. Nature 2000, 406, 593–599. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Zhu, Y.; Gao, J.; Yu, F.; Dong, A.; Shen, W.H. Molecular and Reverse Genetic Characterization of NUCLEOSOME AS-SEMBLY PROTEIN1 (NAP1) Genes Unravels Their Function in Transcription and Nucleotide Excision Repair in Arabidopsis Thaliana. Plant J. 2009, 59, 27–38. [Google Scholar] [CrossRef]
- Liu, Z.Q.; Gao, J.; Dong, A.W.; Shen, W.H. A Truncated Arabidopsis NUCLEOSOME ASSEMBLY PROTEIN 1, ATNAP1;3T, Alters Plant Growth Responses to Abscisic Acid and Salt in the ATNAP1;3-2 Mutant. Mol. Plant 2009, 2, 688–699. [Google Scholar] [CrossRef]
- Gao, J.; Zhu, Y.; Zhou, W.; Molinier, J.; Dong, A.; Shen, W.H. NAP1 Family Histone Chaperones Are Required for Somatic Homologous Recombination in Arabidopsis. Plant Cell 2012, 24, 1437–1447. [Google Scholar] [CrossRef]
- Pajoro, A.; Severing, E.; Angenent, G.C.; Immink, R.G.H. Histone H3 Lysine 36 Methylation Affects Temperature-Induced Al-ternative Splicing and Flowering in Plants. Genome Biol. 2017, 18, 102. [Google Scholar] [CrossRef]
- Bhadouriya, S.L.; Mehrotra, S.; Basantani, M.K.; Loake, G.J.; Mehrotra, R. Role of Chromatin Architecture in Plant Stress Re-sponses: An Update. Front Plant Sci. 2021, 11, 603380. [Google Scholar] [CrossRef]
- Jaskiewicz, M.; Conrath, U.; Peterhälnsel, C. Chromatin Modification Acts as a Memory for Systemic Acquired Resistance in the Plant Stress Response. EMBO Rep. 2011, 12, 50–55. [Google Scholar] [CrossRef]
- Feng, B.; Liu, C.; de Oliveira, M.V.V.; Intorne, A.C.; Li, B.; Babilonia, K.; de Souza Filho, G.A.; Shan, L.; He, P. Protein Poly(ADP-Ribosyl)Ation Regulates Arabidopsis Immune Gene Expression and Defense Responses. PLoS Genet. 2015, 11, e1004936. [Google Scholar] [CrossRef]
- Rissel, D.; Peiter, E. Poly(ADP-Ribose) Polymerases in Plants and Their Human Counterparts: Parallels and Peculiarities. Int. J. Mol. Sci. 2019, 20, 1638. [Google Scholar] [CrossRef] [PubMed]
- Elfving, N.; Davoine, C.; Benlloch, R.; Blomberg, J.; Brännström, K.; Müller, D.; Nilsson, A.; Ulfstedt, M.; Ronne, H.; Wingsle, G.; et al. The Arabidopsis Thaliana Med25 Mediator Subunit Integrates Environmental Cues to Control Plant Development. Proc. Natl. Acad. Sci. USA 2011, 108, 8245–8250. [Google Scholar] [CrossRef] [PubMed]
- Zhai, Q.; Deng, L.; Li, C. Mediator Subunit MED25: At the Nexus of Jasmonate Signaling. Curr. Opin. Plant Biol. 2020, 57, 78–86. [Google Scholar] [CrossRef] [PubMed]
- An, C.; Mou, Z. The Function of the Mediator Complex in Plant Immunity. Plant Signal Behav. 2013, 8, e23182. [Google Scholar] [CrossRef]
- Kim, Y.S.; An, C.; Park, S.; Gilmour, S.J.; Wang, L.; Renna, L.; Brandizzi, F.; Grumet, R.; Thomashow, M.F. CAMTA-Mediated Regulation of Salicylic Acid Immunity Pathway Genes in Arabidopsis Exposed to Low Temperature and Pathogen Infection. Plant Cell 2017, 29, 2465–2477. [Google Scholar] [CrossRef]
- Pandey, N.; Ranjan, A.; Pant, P.; Tripathi, R.K.; Ateek, F.; Pandey, H.P.; Patre, U.V.; Sawant, S.V. CAMTA1 Regulates Drought Responses in Arabidopsis Thaliana. BMC Genom. 2013, 14, 216. [Google Scholar] [CrossRef]
- Qiao, L.; Zhang, W.; Li, X.; Zhang, L.; Zhang, X.; Li, X.; Guo, H.; Ren, Y.; Zheng, J.; Chang, Z. Characterization and Expression Eatterns of Auxin Response Factors in Wheat. Front Plant Sci. 2018, 9, 1395. [Google Scholar] [CrossRef]
- Thomas, Q.A.; Ard, R.; Liu, J.; Li, B.; Wang, J.; Pelechano, V.; Marquardt, S. Transcript Isoform Sequencing Reveals Wide-spread Promoter-Proximal Transcriptional Termination in Arabidopsis. Nat. Commun. 2020, 11, 2589. [Google Scholar] [CrossRef]
- Martinez-Seidel, F.; Beine-Golovchuk, O.; Hsieh, Y.C.; Kopka, J. Systematic Review of Plant Ribosome Heterogeneity and Spe-cialization. Front Plant Sci. 2020, 11, 948. [Google Scholar] [CrossRef]
- Sáez-Vásquez, J.; Delseny, M. Ribosome Biogenesis in Plants: From Functional 45S Ribosomal DNA Organization to Ribo-some Assembly Factors. Plant Cell 2019, 31, 1945–1967. [Google Scholar] [CrossRef]
- Kanodia, P.; Vijayapalani, P.; Srivastava, R.; Bi, R.; Liu, P.; Miller, W.A.; Howell, S.H. Control of Translation During the Un-folded Protein Response in Maize Seedlings: Life without PERKs. Plant Direct 2020, 4, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Duc, K.D.; Batra, S.S.; Bhattacharya, N.; Cate, J.H.D.; Song, Y.S. Differences in the Path to Exit the Ribosome across the Three Domains of Life. Nucleic Acids Res. 2019, 47, 4198–4210. [Google Scholar] [CrossRef]
- Trivedi, D.K.; Ansari, M.W.; Tuteja, N. Multiple Abiotic Stress Responsive Rice Cyclophilin: (OsCYP-25) Mediates a Wide Range of Cellular Responses. Commun. Integr. Biol. 2013, 6. [Google Scholar] [CrossRef] [PubMed]
- Amin-Wetzel, N.; Saunders, R.A.; Kamphuis, M.J.; Rato, C.; Preissler, S.; Harding, H.P.; Ron, D. A J-Protein Co-chaperone Re-cruits BiP to Monomerize IRE1 and Repress the Unfolded Protein Response. Cell 2017, 171, 1625–1637. [Google Scholar] [CrossRef] [PubMed]
- Upton, J.P.; Wang, L.; Han, D.; Wang, E.S.; Huskey, N.E.; Lim, L.; Truitt, M.; McManus, M.T.; Ruggero, D.; Goga, A.; et al. Ire1alpha cleaves select microRNAs during ER stress to derepress translation of proapoptotic Caspase-2. Science 2012, 338, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.L.; Li, J.Y.; Wang, M.J.; Song, Z.T.; Liu, J.X. Protein Quality Control in Plant Organelles: Current Progress and Future Perspectives. Mol. Plant 2021, 14, 95–114. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Peng, D.; Zhou, J.; Lin, S.; Wang, C.; Ning, W.; Xu, H.; Deng, W.; Xue, Y. IEKPD 2.0: An Update with Rich Annotations for Eukaryotic Protein Kinases, Protein Phosphatases and Proteins Containing Phosphoprotein-Binding Domains. Nucleic Acids Res. 2019, 47, D344–D350. [Google Scholar] [CrossRef]
- Bonturi, C.R.; Teixeira, A.B.S.; Rocha, V.M.; Valente, P.F.; Oliveira, J.R.; Filho, C.M.B.; Batista, I.F.C.; Oliva, M.L.V. Plant Ku-nitz Inhibitors and Their Interaction with Proteases: Current and Potential Pharmacological Targets. Int. J. Mol. Sci. 2022, 23, 4742. [Google Scholar] [CrossRef]
- Nishimura, K.; Kato, Y.; Sakamoto, W. Essentials of Proteolytic Machineries in Chloroplasts. Mol. Plant 2017, 10, 4–19. [Google Scholar] [CrossRef]
- Deng, Y.; Srivastava, R.; Quilichini, T.D.; Dong, H.; Bao, Y.; Horner, H.T.; Howell, S.H. IRE1, a Component of the Unfolded Protein Response Signaling Pathway, Protects Pollen Development in Arabidopsis from Heat Stress. Plant J. 2016, 88, 193–204. [Google Scholar] [CrossRef]
- Xu, J.; Wang, X.Y.; Guo, W.Z. The Cytochrome P450 Superfamily: Key Players in Plant Development and Defense. J. Integr. Agric. 2015, 14, 1673–1686. [Google Scholar] [CrossRef]
- Fischer, M.; Kuckenberg, M.; Kastilan, R.; Muth, J.; Gebhardt, C. Novel in Vitro Inhibitory Functions of Potato Tuber Protein-aceous Inhibitors. Mol. Genet. Genom. 2014, 290, 387–398. [Google Scholar] [CrossRef] [PubMed]
- Haak, D.C.; Fukao, T.; Grene, R.; Hua, Z.; Ivanov, R.; Perrella, G.; Li, S. Multilevel Regulation of Abiotic Stress Responses in Plants. Front Plant Sci. 2017, 8, 1564. [Google Scholar] [CrossRef] [PubMed]
- Aydemir, B.Ç.; Ergül, A. Transcriptomic Analysis of Endoplasmic Reticulum Stress in Roots of Grapevine Rootstock. Plant Biotechnol. Rep. 2021, 15, 683–706. [Google Scholar] [CrossRef]
- Sun, L.; Yang, Z.-T.; Song, Z.-T.; Wang, M.-J.; Sun, L.; Lu, S.-J.; Liu, J.-X. The Plant-Specific Transcription Factor Gene NAC103 Is Induced by BZIP60 through a New Cis-Regulatory Element to Modulate the Unfolded Protein Response in Arabidopsis. Plant J. 2013, 76, 274–286. [Google Scholar] [CrossRef]
- Herath, V.; Verchot, J. Transcriptional Regulatory Networks Associate with Early Stages of Potato Virus X Infection of So-lanum Tuberosum. Int. J. Mol. Sci. 2021, 22, 2837. [Google Scholar] [CrossRef]
- Mészáros, B.; Erdös, G.; Dosztányi, Z. IUPred2A: Context-Dependent Prediction of Protein Disorder as a Function of Redox State and Protein Binding. Nucleic Acids Res. 2018, 46, W329–W337. [Google Scholar] [CrossRef]
- Chen, Y.; Chen, Y.; Shi, C.; Huang, Z.; Zhang, Y.; Li, S.; Li, Y.; Yu, C.; Li, Z.; Ahang, X.; et al. SOAPnuke: A MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. GigaScience 2018, 7, 1–6. [Google Scholar] [CrossRef]
- Hoopes, G.; Meng, X.; Hamilton, J.P.; Achakkagari, S.R.; de Alves Freitas Guesdes, F.; Bolger, M.E.; Coombs, J.J.; Esselink, D.; Kaiser, N.R.; Kodde, L.; et al. Phased, Chromosome-Scale Genome Assemblies of Tetraploid Potato Reveal a Complex Ge-nome, Transcriptome, and Predicted Proteome Landscape Underpinning Genetic Diversity. Mol. Plant 2022, 15, 520–536. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A Fast Spliced Aligner with Low Memory Requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Etherington, G.J.; Ramirez-Gonzalez, R.H.; MacLean, D. Bio-Samtools 2: A Package for Analysis and Visualization of Se-quence and Alignment Data with SAMtools in Ruby. Bioinformatics 2015, 31, 2565–2567. [Google Scholar] [CrossRef] [PubMed]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve Years of SAMtools and BCFtools. GigaScience 2021, 10, giab008. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie Enables Improved Reconstruc-tion of a Transcriptome from RNA-Seq Reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Putri, G.H.; Anders, S.; Pyl, P.T.; Pimanda, J.E.; Zanini, F. Analysing High-Throughput Sequencing Data in Python with HTSeq 2.0. Bioinformatics 2022, 38, 2943–2945. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. EdgeR: A Bioconductor Package for Differential Expression Analysis of Digital Gene Expression Data. Bioinformatics 2009, 26, 139–140. [Google Scholar] [CrossRef]
- Blighe, K.; Rana, S.; Lewis, M. EnhancedVolcano: Publication-Ready Volcano Plots with Enhanced Colouring and Labeling. 2018. Available online: https://github.com/kevinblighe/EnhancedVolcano (accessed on 27 May 2022).
- Agarwala, R.; Barrett, T.; Beck, J.; Benson, D.A.; Bollin, C.; Bolton, E.; Bourexis, D.; Brister, J.R.; Bryant, S.H.; Canese, K.; et al. Database Resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2016, 44, D7–D19. [Google Scholar] [CrossRef]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. In-terProScan 5: Genome-Scale Protein Function Classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef]
- Blum, M.; Chang, H.-Y.; Chuguransky, S.; Grego, T.; Kandasaamy, S.; Mitchell, A.; Nuka, G.; Paysan-Lafosse, T.; Qureshi, M.; Raj, S.; et al. The InterPro Protein Families and Domains Database: 20 Years On. Nucleic Acids Res. 2020, 49, D344–D354. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools—An Integrative Toolkit Developed for Inter-active Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Bray, N.L.; Pimentel, H.; Melsted, P.; Pachter, L. Near-Optimal Probabilistic RNA-Seq Quantification. Nat. Biotechnol. 2016, 34, 525–527. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Vitting-Seerup, K.; Sandelin, A.; Berger, B. IsoformSwitchAnalyzeR: Analysis of Changes in Genome-Wide Patterns of Alter-native Splicing and Its Functional Consequences. Bioinformatics 2019, 35, 4469–4471. [Google Scholar] [CrossRef] [PubMed]
- Soneson, C.; Love, M.I.; Robinson, M.D. Differential Analyses for RNA-Seq: Transcript-Level Estimates Improve Gene-Level Inferences. F1000Research 2015, 4, 1521. [Google Scholar] [CrossRef] [PubMed]
- Vitting-Seerup, K.; Sandelin, A. The Landscape of Isoform Switches in Human Cancers. Mol. Cancer Res. 2017, 15, 1206–1220. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Reyes, A.; Huber, W. Detecting differential usage of exons from RNA-seq data. Genome Res. 2012, 22, 2008–2017. [Google Scholar] [CrossRef] [PubMed]
- Huber, W.; Carey, V.J.; Gentleman, R.; Anders, S.; Carlson, M.; Carvalho, B.S.; Bravo, H.C.; Davis, S.; Gatto, L.; Girke, T.; et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 2015, 12, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.J.; Yang, D.C.; Kong, L.; Hou, M.; Meng, Y.Q.; Wei, L.; Gao, G. CPC2: A Fast and Accurate Coding Potential Calculator Based on Sequence Intrinsic Features. Nucleic Acids Res. 2017, 45, W12–W16. [Google Scholar] [CrossRef]
- El-Gebali, S.; Mistry, J.; Bateman, A.; Eddy, S.R.; Luciani, A.; Potter, S.C.; Qureshi, M.; Richardson, L.J.; Salazar, G.A.; Smart, A.; et al. The Pfam Protein Families Database in 2019. Nucleic Acids Res. 2019, 47, D427–D432. [Google Scholar] [CrossRef]
- Choo, K.H.; Tan, T.W.; Ranganathan, S. A Comprehensive assessment of N-terminal signal peptides prediction methods. BMC Bioinform. 2009, 10, 1–12. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Herath, V.; Verchot, J. Comprehensive Transcriptome Analysis Reveals Genome-Wide Changes Associated with Endoplasmic Reticulum (ER) Stress in Potato (Solanum tuberosum L.). Int. J. Mol. Sci. 2022, 23, 13795. https://doi.org/10.3390/ijms232213795
Herath V, Verchot J. Comprehensive Transcriptome Analysis Reveals Genome-Wide Changes Associated with Endoplasmic Reticulum (ER) Stress in Potato (Solanum tuberosum L.). International Journal of Molecular Sciences. 2022; 23(22):13795. https://doi.org/10.3390/ijms232213795
Chicago/Turabian StyleHerath, Venura, and Jeanmarie Verchot. 2022. "Comprehensive Transcriptome Analysis Reveals Genome-Wide Changes Associated with Endoplasmic Reticulum (ER) Stress in Potato (Solanum tuberosum L.)" International Journal of Molecular Sciences 23, no. 22: 13795. https://doi.org/10.3390/ijms232213795
APA StyleHerath, V., & Verchot, J. (2022). Comprehensive Transcriptome Analysis Reveals Genome-Wide Changes Associated with Endoplasmic Reticulum (ER) Stress in Potato (Solanum tuberosum L.). International Journal of Molecular Sciences, 23(22), 13795. https://doi.org/10.3390/ijms232213795







