Molecular Management of High-Grade Serous Ovarian Carcinoma
Abstract
1. Introduction
2. Current Molecular Approaches for the Diagnosis and Prognosis of HGSOC
2.1. Molecular Markers and Algorithm Decisions for the Diagnosis of HGSOC: Carbohydrate Antigen 125 (CA125), Alone or in Combination with Other Imaging Techniques or Biomarkers
2.2. Gene Expression Profiling and Gene Panels
3. Novel Trends in the Understanding of the Origin of HGSOC
3.1. Cell Origin
3.2. Hormones
- (a) Gonadotropin hypothesis: The risk of serous EOC increases due to the excessive ovarian tissue uptake of FSH and LH [136,137]. The FSH-mediated proliferation and migration of EOC cells via SphK [138] and the FSH-R/LH-R-mediated cell migration and invasiveness via COX2 [139,140] or ERBB-2 [141] support this theory.
- (b) Androgen/progestin hypothesis: This dual hypothesis acknowledges high androgen levels (usually linked to polycystic ovarian syndrome (PCOS) or obesity) as an EOC risk factor [142,143,144] and P4 as a protective factor [132,133]. High expression of AR [145,146] in HGSOC [147,148] and FTE is associated with the onset of serous EOC [149]. Conversely, PR expression is associated with a favorable prognosis and a reduced metastatic risk [134,150].
3.3. Genetic Traits
3.3.1. Inherited Mutations
3.3.2. Somatic Mutations
3.3.3. Chromosomal Aberrations
3.4. Epigenetics
3.5. Target Signaling Pathways in HGSOC
4. Ongoing Clinical Trials
5. Liquid Biopsy as a Non-Invasive Tool for the Early Diagnosis of HGSOC
5.1. Circulating Tumor Cells
5.2. Cell-Free DNA and Circulating DNA
5.3. Cell-Free RNA and Circulating RNA
5.4. Exosomes
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Abbreviation | Whole Name |
| 5-hmc | 5-Hydroxymethylcytosine |
| 5-mc | 5-Methylcytosine |
| ADNEX | Assessment of Different Neoplasias in the Adnexa |
| AUC | Area under the curve |
| CA125 | Carbohydrate antigen 125 |
| CICs | Cortical inclusion cysts |
| CIN | Chromosomal instability |
| CLP | Chromothripsis-like pattern |
| CTCs | Circulating tumor cells |
| ctDNA | Circulating tumor DNA |
| ctRNA | Circulating tumor RNA |
| EOC | Epithelial ovarian carcinoma |
| eQTL | Expression quantitative trait locus |
| EVs | Extracellular vesicles |
| FDA | Food and Drug Administration |
| FIGO | International Federation of Gynecology and Obstetrics |
| FTE | Fallopian tube epithelium |
| FSH | Follicle-stimulating hormone |
| GWASs | Genome-wide association studies |
| HDACs | Deacetylases |
| HGSOC | High-grade serous ovarian carcinoma |
| HMTs | Histone methyl transferases |
| HPO | Hypothalamic–pituitary–ovarian |
| HRD | HRR pathway deficiency |
| HRR | Homologous recombination repair |
| IHC | Immunohistochemistry |
| LGSOC | Low-grade serous ovarian carcinoma |
| LH | Luteinizing hormone |
| lncRNAs | Long non-coding RNAs |
| MIA | Multivariate index assay |
| miRNAs | MicroRNAs |
| NACT | Neoadjuvant chemotherapy |
| ncRNAs | Non-coding RNAs |
| NGS | Next-generation sequencing |
| NIH | National Institutes of Health |
| NROSS | Normal Risk Ovarian Screening Study |
| OC | Ovarian cancer |
| OSE | Ovarian surface epithelium |
| P4 | Progesterone |
| PCOS | Polycystic ovarian syndrome |
| PLCO | Prostate, Lung, Ovarian, Colorectal |
| PPE | Pelvic palpation examination |
| PRMT | Protein arginine methyltransferase |
| ROCA | Risk of Ovarian Cancer Algorithm |
| ROMA | Risk of Ovarian Malignancy Algorithm |
| ROS | Reactive oxygen species |
| RMI | Risk of Malignancy Index |
| SNPs | Single-nucleotide polymorphisms |
| STICs | Serous tubular intra-epithelial carcinomas |
| TCGA | The Cancer Genome Atlas |
| TEPs | Tumor-educated platelets |
| TVS | Transvaginal ultrasound/transvaginal sonography |
| TWAS | Transcriptome-wide association studies |
| UKCTOCS | United Kingdom Collaborative Trial of Ovarian Cancer Screening |
| WGD | Whole-genome duplication |
| Gene Symbol | Gene Name |
| ANGPTL1/2 | Angiopoietin Like 1/2 |
| APC | APC regulator of WNT signaling pathway |
| ARID1A | AT-rich interaction domain 1A |
| ATM | ATM serine/threonine kinase |
| ATR | ATR serine/threonine kinase |
| BARD1 | BRCA1-associated RING domain 1 |
| BMP7 | Bone morphogenetic protein 7 |
| BRAF | B-Raf proto-oncogene, serine/threonine kinase |
| BRCA1/2 | BRCA1/2 DNA-repair-associated |
| BRIP1 | BRCA1-interacting helicase 1 |
| C2CD4D | C2 calcium-dependent domain containing 4D |
| CASC11 | Cancer susceptibility 11 |
| CCNE1 | Cyclin E1 |
| CDK12 | Cyclin-dependent kinase 12 |
| CHEK2 | Checkpoint kinase 2 |
| CKB | Creatine kinase B |
| COL23A1 | Collagen type XXIII alpha 1 chain |
| CPNE1 | Copine 1 |
| CSMD3 | CUB and Sushi multiple domains 3 |
| CTBP1-DT | C-terminal binding protein 1-divergent transcript |
| CTNNBL1 | Catenin beta-like 1 |
| CXCL10 | C-X-C motif chemokine ligand 10 |
| CXCL11 | C-X-C motif chemokine ligand 11 |
| CXCR3 | C-X-C motif chemokine receptor 3 |
| CCND | Cyclin D |
| FANCD2 | FA complementation group D2 |
| FAP | Fibroblast activation protein alpha |
| FAT3 | FAT atypical cadherin 3 |
| FOXC2 | Forkhead box C2 |
| FOXL2 | Forkhead box L2 |
| FOXM1 | Forkhead box M1 |
| GATA4 | GATA-binding protein 4 |
| GATA6 | GATA-binding protein 6 |
| GNG11 | G protein subunit gamma 11 |
| HEATR3 | HEAT repeat-containing 3 |
| HIC1 | HIC ZBTB transcriptional repressor 1 |
| HIF1α | Hypoxia-inducible factor 1 subunit alpha |
| HMGA2 | High mobility group AT-hook 2 |
| HOX | Homeobox family |
| HOXA9 | Homeobox A9 |
| IGFBP7 | Insulin-like growth-factor-binding protein 7 |
| KLF6 | Kruppel-like factor 6 |
| KRAS | KRAS proto-oncogene, GTPase |
| MCM2 | Minichromosome maintenance complex component 2 |
| MECOM | MDS1 and EVI1 complex locus |
| miR-1246 | MicroRNA 1246 |
| miR-1290 | MicroRNA 1290 |
| miR-141 | MicroRNA 141 |
| miR-145 | MicroRNA 145 |
| miR-200 | MicroRNA 200 |
| miR-200a/b/c | MicroRNA 200a/b/c |
| miR-203 | MicroRNA 203 |
| miR-205 | MicroRNA 205 |
| miR-21 | MicroRNA 21 |
| miR-214 | MicroRNA 214 |
| miR-2278 | MicroRNA 2278 |
| miR-23a | MicroRNA 23a |
| miR-27-a-3p | MicroRNA 27-a-3p |
| miR-34-a | MicroRNA 34-a |
| MIR4435-2HG | MIR4435-2 Host Gene |
| miR-595 | MicroRNA 595 |
| miR-93 | MicroRNA 93 |
| MLF1 | Myeloid leukemia factor 1 |
| MLH3 | MutL homolog 3 |
| MRE11A | MRE11 homolog, double-strand break repair nuclease |
| MUC16 | Mucin 16, cell-surface-associated |
| MYC | MYC proto-oncogene, bHLH transcription factor |
| NF1 | Neurofibromin 1 |
| NF-κβ | Nuclear factor kappa B |
| NOXA | Phorbol-12-myristate-13-acetate-induced protein 1 |
| NR5A1 | Nuclear receptor subfamily 5 group A member 1 |
| OPCML | Opioid-binding protein/cell adhesion molecule-like |
| P16 (CCDKN2A) | Cyclin-dependent kinase inhibitor 2A |
| P21cip1 (CCDKN1A) | Cyclin-dependent kinase inhibitor 1A |
| PALB2 | Partner and localizer of BRCA2 |
| PCNA | Proliferating cell nuclear antigen |
| PIK3CA | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha |
| PLCG2 | Phospholipase C gamma 2 |
| PTEN | Phosphatase and tensin homolog |
| RAD50 | RAD50 double-strand break repair protein |
| RAD51 | RAD51 recombinase |
| RAD51D | RAD51 paralog D |
| RASSF1A | Ras association domain family member 1 |
| RB1 | RB transcriptional corepressor 1 |
| RUNX3 | RUNX family transcription factor 3 |
| SOX11 | SRY-box transcription factor 11 |
| STRCP1 | Stereocilin pseudogene 1 |
| TP53 | Tumor protein p53 |
| TWIST2 | Twist family bHLH transcription factor 2 |
| UBE2Q1 | Ubiquitin-conjugating enzyme E2 Q1 |
| WNT6 | Wnt family member 6 |
| WT1 | WT1 transcription factor |
| Protein Symbol | Protein Name |
| ADAM8 | ADAM metallopeptidase domain 8 |
| anti-HSF1 | Anti-heat-shock transcription factor 1 |
| AR | Androgen receptor |
| AURKA | Aurora kinase A |
| CA125 | Carbohydrate antigen 125 |
| CA72-4 | Cancer antigen 72-4 |
| CARM1 | Coactivator-associated arginine methyltransferase 1 |
| CCDC155 | KASH domain-containing 5 (KASH5) |
| CDKs | Cyclin-dependent kinases |
| CEA | Carcinoembryonic antigen |
| CNNE1 | G1/S-specific cyclin-E1 |
| COX2 | Cytochrome c oxidase subunit 2 |
| CXCL13 | C-X-C motif chemokine ligand 13 |
| E2F | E2F transcription factor family |
| ER | Estrogen receptor |
| ERBB2 | Receptor tyrosine-protein kinase erbB-2 |
| EZH2 | Enhancer of zeste 2 polycomb repressive complex 2 subunit |
| FGG | Fibrinogen gamma chain |
| FOLR1 | Folate receptor alpha |
| FSH-R | Follicle-stimulating hormone receptor |
| GFA | Glutamine–fructose-6-phosphate transaminase 1 (GFPT1) |
| GPCR | G-protein-coupled receptor |
| GSN | Gelsolin |
| HE4 | Human epididymis 4 |
| IL-6 | Interleukin-6 |
| KLK11 | Kallikrein-related peptidase 11 |
| LBP | Lipopolysaccharide-binding protein |
| LCAT | Lecithin cholesterol acyltransferase |
| LH-R | Luteinizing hormone receptor |
| MAPKs | Mitogen-activated protein kinases |
| MDK | Midkine |
| MMP-7 | Matrilysin |
| MSLN | Mesothelin |
| P53 | Cellular tumor antigen p53 |
| PARP | Poly [ADP-ribose] polymerase |
| PR | Progesterone receptor |
| RAF | RAF proto-oncogene serine/threonine-protein kinase |
| RAS | RAS protein |
| RB | Retinoblastoma-associated protein |
| RTKs | Receptor tyrosine kinases |
| SphK | Sphingosine kinase |
| TKR | Tyrosine kinase receptor |
| VCAM-1 | Vascular cell adhesion protein 1 |
| VEGF | Vascular endothelial growth factor |
| WISP1 | Cellular communication network factor 4 (CCN4) |
| γ-H2AX | γ-Histone H2AX |
References
- Common Cancer Sites—Cancer Stat Facts. Available online: https://seer.cancer.gov/statfacts/html/common.html (accessed on 10 September 2022).
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.; Sun, H.; Li, X.; Lin, F.; Wang, Z.; Wang, X. Ovarian Cancer: Epigenetics, Drug Resistance, and Progression. Cancer Cell Int. 2021, 21, 434. [Google Scholar] [CrossRef]
- Köbel, M.; Kang, E.Y. The Evolution of Ovarian Carcinoma Subclassification. Cancers 2022, 14, 416. [Google Scholar] [CrossRef]
- Lisio, M.A.; Fu, L.; Goyeneche, A.; Gao, Z.H.; Telleria, C. High-Grade Serous Ovarian Cancer: Basic Sciences, Clinical and Therapeutic Standpoints. Int. J. Mol. Sci. 2019, 20, 952. [Google Scholar] [CrossRef]
- Lheureux, S.; Braunstein, M.; Oza, A.M. Epithelial Ovarian Cancer: Evolution of Management in the Era of Precision Medicine. CA Cancer J. Clin. 2019, 69, 280–304. [Google Scholar] [CrossRef] [PubMed]
- Torre, L.A.; Trabert, B.; DeSantis, C.E.; Miller, K.D.; Samimi, G.; Runowicz, C.D.; Gaudet, M.M.; Jemal, A.; Siegel, R.L. Ovarian Cancer Statistics, 2018. CA Cancer J. Clin. 2018, 68, 284–296. [Google Scholar] [CrossRef] [PubMed]
- Hirst, J.; Crow, J.; Godwin, A. Ovarian Cancer Genetics: Subtypes and Risk Factors. In Ovarian Cancer—From Pathogenesis to Treatment; IntechOpen: London, UK, 2018. [Google Scholar]
- Prat, J. Ovarian Carcinomas: Five Distinct Diseases with Different Origins, Genetic Alterations, and Clinicopathological Features. Virchows Archiv 2012, 460, 237–249. [Google Scholar] [CrossRef]
- Berek, J.S.; Renz, M.; Kehoe, S.; Kumar, L.; Friedlander, M. Cancer of the Ovary, Fallopian Tube, and Peritoneum: 2021 Update. Int. J. Gynaecol. Obstet. 2021, 155, 61–85. [Google Scholar] [CrossRef]
- Peres, L.C.; Cushing-Haugen, K.L.; Köbel, M.; Harris, H.R.; Berchuck, A.; Rossing, M.A.; Schildkraut, J.M.; Doherty, J.A. Invasive Epithelial Ovarian Cancer Survival by Histotype and Disease Stage. J. Natl. Cancer Inst. 2019, 111, 60–68. [Google Scholar] [CrossRef]
- Bowtell, D.D.; Böhm, S.; Ahmed, A.A.; Aspuria, P.J.; Bast, R.C.; Beral, V.; Berek, J.S.; Birrer, M.J.; Blagden, S.; Bookman, M.A.; et al. Rethinking Ovarian Cancer II: Reducing Mortality from High-Grade Serous Ovarian Cancer. Nat. Rev. Cancer 2015, 15, 668–679. [Google Scholar] [CrossRef] [PubMed]
- Bast, R.C.; Lu, Z.; Han, C.Y.; Lu, K.H.; Anderson, K.S.; Drescher, C.W.; Skates, S.J. Biomarkers and Strategies for Early Detection of Ovarian Cancer. Cancer Epidemiol. Biomark. Prev. 2020, 29, 2504–2512. [Google Scholar] [CrossRef] [PubMed]
- Chacón, E.; Dasí, J.; Caballero, C.; Alcázar, J.L. Risk of Ovarian Malignancy Algorithm versus Risk Malignancy Index-I for Preoperative Assessment of Adnexal Masses: A Systematic Review and Meta-Analysis. Gynecol. Obstet. Investig. 2019, 84, 591–598. [Google Scholar] [CrossRef]
- Ratnavelu, N.D.; Brown, A.P.; Mallett, S.; Scholten, R.J.; Patel, A.; Founta, C.; Galaal, K.; Cross, P.; Naik, R. Intraoperative Frozen Section Analysis for the Diagnosis of Early Stage Ovarian Cancer in Suspicious Pelvic Masses. Cochrane Database Syst. Rev. CDS 2016, 2016, CD010360. [Google Scholar] [CrossRef]
- Querleu, D.; Planchamp, F.; Chiva, L.; Fotopoulou, C.; Barton, D.; Cibula, D.; Aletti, G.; Carinelli, S.; Creutzberg, C.; Davidson, B.; et al. European Society of Gynaecological Oncology (ESGO) Guidelines for Ovarian Cancer Surgery. Int. J. Gynecol. Cancer 2017, 27, 1534–1542. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Cheng, S.; Jin, Y.; Zhao, Y.; Wang, Y. Roles of CA125 in Diagnosis, Prediction, and Oncogenesis of Ovarian Cancer. Biochim. Biophys. Acta Rev. Cancer 2021, 1875, 188503. [Google Scholar] [CrossRef] [PubMed]
- Charkhchi, P.; Cybulski, C.; Gronwald, J.; Wong, F.O.; Narod, S.A.; Akbari, M.R. CA125 and Ovarian Cancer: A Comprehensive Review. Cancers 2020, 12, 3730. [Google Scholar] [CrossRef]
- Srivastava, A.; Gupta, A.; Patidar, S. Review of Biomarker Systems as an Alternative for Early Diagnosis of Ovarian Carcinoma. Clin. Transl. Oncol. 2021, 23, 1967–1978. [Google Scholar] [CrossRef]
- Matulonis, U.A.; Sood, A.K.; Fallowfield, L.; Howitt, B.E.; Sehouli, J.; Karlan, B.Y. Ovarian Cancer. Nat. Rev. Dis. Primers 2016, 2, 16061. [Google Scholar] [CrossRef]
- Shinagare, A.B.; Sadowski, E.A.; Park, H.; Brook, O.R.; Forstner, R.; Wallace, S.K.; Horowitz, J.M.; Horowitz, N.; Javitt, M.; Jha, P.; et al. Ovarian Cancer Reporting Lexicon for Computed Tomography (CT) and Magnetic Resonance (MR) Imaging Developed by the SAR Uterine and Ovarian Cancer Disease-Focused Panel and the ESUR Female Pelvic Imaging Working Group. Eur. Radiol. 2022, 32, 3220–3235. [Google Scholar] [CrossRef]
- Sokalska, A.; Timmerman, D.; Testa, A.C.; van Holsbeke, C.; Lissoni, A.A.; Leone, F.P.G.; Jurkovic, D.; Valentin, L. Diagnostic Accuracy of Transvaginal Ultrasound Examination for Assigning a Specific Diagnosis to Adnexal Masses. Ultrasound Obstet. Gynecol. 2009, 34, 462–470. [Google Scholar] [CrossRef] [PubMed]
- Jung, S.E.; Lee, J.M.; Rha, S.E.; Byun, J.Y.; Jung, J.I.; Hahn, S.T. CT and MR Imaging of Ovarian Tumors with Emphasis on Differential Diagnosis. Radiographics 2002, 22, 1305–1325. [Google Scholar] [CrossRef] [PubMed]
- Liberto, J.M.; Chen, S.-Y.; Shih, I.-M.; Wang, T.-H.; Wang, T.-L.; Pisanic, T.R. Current and Emerging Methods for Ovarian Cancer Screening and Diagnostics: A Comprehensive Review. Cancers 2022, 14, 2885. [Google Scholar] [CrossRef]
- Bast, R.C.; Feeney, M.; Lazarus, H.; Nadler, L.M.; Colvin, R.B.; Knapp, R.C. Reactivity of a Monoclonal Antibody with Human Ovarian Carcinoma. J. Clin. Investig. 1981, 68, 1331–1337. [Google Scholar] [CrossRef] [PubMed]
- Köbel, M.; Kalloger, S.E.; Boyd, N.; McKinney, S.; Mehl, E.; Palmer, C.; Leung, S.; Bowen, N.J.; Ionescu, D.N.; Rajput, A.; et al. Ovarian Carcinoma Subtypes Are Different Diseases: Implications for Biomarker Studies. PLoS Med. 2008, 5, 1749–1760. [Google Scholar] [CrossRef]
- Van Haaften-day, C.; Shen, Y.; Xu, F.; Yu, Y.; Berchuck, A.; Havrilesky, L.J.; De Bruijn, H.W.A.; Hacker, N.F. OVX1, Macrophague-Colony Stimulating Factor, and CA-125-II as Tumor Markers for Epithelial. A Critical Appraisal. Cancer 2001, 92, 2837–2844. [Google Scholar] [CrossRef]
- Urban, N.; McIntosh, M.W.; Andersen, M.; Karlan, B.Y. Ovarian Cancer Screening. Hematol. Oncol. Clin. N. Am. 2003, 17, 989–1005. [Google Scholar] [CrossRef]
- Kobayashi, H.; Yamada, Y.; Sado, T.; Sakata, M.; Yoshida, S.; Kawaguchi, R.; Kanayama, S.; Shigetomi, H.; Haruta, S.; Tsuji, Y.; et al. A Randomized Study of Screening for Ovarian Cancer: A Multicenter Study in Japan. Int. J. Gynecol. Cancer 2008, 18, 414–420. [Google Scholar] [CrossRef]
- Jacobs, I.J.; Skates, S.J.; MacDonald, N.; Menon, U.; Rosenthal, A.N.; Davies, A.P.; Woolas, R.; Jeyarajah, A.R.; Sibley, K.; Lowe, D.G.; et al. Screening for Ovarian Cancer: A Pilot Randomised Controlled Trial. Lancet 1999, 353, 1207–1210. [Google Scholar] [CrossRef]
- Dochez, V.; Caillon, H.; Vaucel, E.; Dimet, J.; Winer, N.; Ducarme, G. Biomarkers and Algorithms for Diagnosis of Ovarian Cancer: CA125, HE4, RMI and ROMA, a Review. J. Ovarian Res. 2019, 12, 28. [Google Scholar] [CrossRef]
- Kafali, H.; Artuc, H.; Demir, N. Use of CA125 Fluctuation during the Menstrual Cycle as a Tool in the Clinical Diagnosis of Endometriosis; a Preliminary Report. Eur. J. Obstet. Gynecol. Reprod. Biol. 2004, 116, 85–88. [Google Scholar] [CrossRef] [PubMed]
- Kokot, I.; Piwowar, A.; Jędryka, M.; Sołkiewicz, K.; Kratz, E.M. Diagnostic Significance of Selected Serum Inflammatory Markers in Women with Advanced Endometriosis. Int. J. Mol. Sci. 2021, 22, 2295. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhou, F.; Xiao, X.; Ying, C. Serum Levels of Human Epididymis Protein 4 Are More Stable than Cancer Antigen 125 in Early and Mid-Term Pregnancy. J. Obstet. Gynaecol. Res. 2018, 44, 2053–2058. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, M.; Duffy, J.M.N.; Davis, C.J.; Nieves Plana, M.; Khan, K.S. Diagnostic Accuracy of Cancer Antigen 125 for Endometriosis: A Systematic Review and Meta-Analysis. BJOG 2016, 123, 1761–1768. [Google Scholar] [CrossRef]
- Szecsi, P.B.; Andersen, M.R.; Bjørngaard, B.; Hedengran, K.K.; Stender, S. Cancer Antigen 125 after Delivery in Women with a Normal Pregnancy: A Prospective Cohort Study. Acta Obstet. Gynecol. Scand. 2014, 93, 1295–1301. [Google Scholar] [CrossRef]
- Machado-Lopez, A.; Simón, C.; Mas, A. Molecular and Cellular Insights into the Development of Uterine Fibroids. Int. J. Mol. Sci. 2021, 22, 8483. [Google Scholar] [CrossRef]
- Hu, X.; Zhang, J.; Cao, Y. Factors Associated with Serum CA125 Level in Women without Ovarian Cancer in the United States: A Population-Based Study. BMC Cancer 2022, 22, 544. [Google Scholar] [CrossRef]
- Ataseven, H.; Öztürk, Z.A.; Arhan, M.; Yüksel, O.; Köklü, S.; Ibiş, M.; Başar, Ö.; Yilmaz, F.M.; Yüksel, I. Cancer Antigen 125 Levels in Inflammatory Bowel Diseases. J. Clin. Lab. Anal. 2009, 23, 244–248. [Google Scholar] [CrossRef]
- Johnson, C.C.; Kessel, B.; Riley, T.L.; Ragard, L.R.; Williams, C.R.; Xu, J.L.; Buys, S.S. The Epidemiology of CA-125 in Women without Evidence of Ovarian Cancer in the Prostate, Lung, Colorectal and Ovarian Cancer (PLCO) Screening Trial. Gynecol. Oncol. 2008, 110, 383–389. [Google Scholar] [CrossRef]
- Fortner, R.T.; Vitonis, A.F.; Schock, H.; Hüsing, A.; Johnson, T.; Fichorova, R.N.; Fashemi, T.; Yamamoto, H.S.; Tjønneland, A.; Hansen, L.; et al. Correlates of Circulating Ovarian Cancer Early Detection Markers and Their Contribution to Discrimination of Early Detection Models: Results from the EPIC Cohort. J. Ovarian Res. 2017, 10, 20. [Google Scholar] [CrossRef]
- Sasamoto, N.; Babic, A.; Rosner, B.A.; Fortner, R.T.; Vitonis, A.F.; Yamamoto, H.; Fichorova, R.N.; Titus, L.J.; Tjønneland, A.; Hansen, L.; et al. Development and Validation of Circulating CA125 Prediction Models in Postmenopausal Women. J. Ovarian Res. 2019, 12, 116. [Google Scholar] [CrossRef] [PubMed]
- Pauler, D.K.; Menon, U.; McIntosh, M.; Symecko, H.L.; Skates, S.J.; Jacobs, I.J. Factors Influencing Serum CA125II Levels in Healthy Postmenopausal Women. Cancer Epidemiol. Biomark. Prev. 2001, 10, 489–493. [Google Scholar]
- Lycke, M.; Kristjansdottir, B.; Sundfeldt, K. A Multicenter Clinical Trial Validating the Performance of HE4, CA125, Risk of Ovarian Malignancy Algorithm and Risk of Malignancy Index. Gynecol. Oncol. 2018, 151, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, I.; Oram, D.; Fairbanks, J.; Turner, J.; Frost, C.; Grudzinskas, J.G. A Risk of Malignancy Index Incorporating CA 125, Ultrasound and Menopausal Status for the Accurate Preoperative Diagnosis of Ovarian Cancer. Maturitas 1991, 13, 177. [Google Scholar] [CrossRef]
- Campos, C.; Sarian, L.O.; Jales, R.M.; Hartman, C.; Araújo, K.G.; Pitta, D.; Yoshida, A.; Andrade, L.; Derchain, S. Performance of the Risk of Malignancy Index for Discriminating Malignant Tumors in Women with Adnexal Masses. J. Med. Ultrasound 2016, 35, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Meys, E.M.J.; Kaijser, J.; Kruitwagen, R.F.P.M.; Slangen, B.F.M.; van Calster, B.; Aertgeerts, B.; Verbakel, J.Y.; Timmerman, D.; van Gorp, T. Subjective Assessment versus Ultrasound Models to Diagnose Ovarian Cancer: A Systematic Review and Meta-Analysis. Eur. J. Cancer 2016, 58, 17–29. [Google Scholar] [CrossRef]
- Buys, S.S.; Partridge, E.; Black, A.; Johnson, C.C.; Lamerato, L.; Isaacs, C.; Reding, D.J.; Greenlee, R.T.; Yokochi, L.A.; Kessel, B.; et al. Effect of Screening on Ovarian Cancer Mortality: The Prostate, Lung, Colorectal and Ovarian (PLCO) Cancer Screening Randomized Controlled Trial. JAMA 2011, 305, 2295–2302. [Google Scholar] [CrossRef]
- Pinsky, P.F.; Yu, K.; Kramer, B.S.; Black, A.; Buys, S.S.; Partridge, E.; Gohagan, J.; Berg, C.D.; Prorok, P.C. Extended Mortality Results for Ovarian Cancer Screening in the PLCO Trial with Median 15 Years Follow-Up. Gynecol. Oncol. 2016, 143, 270–275. [Google Scholar] [CrossRef] [PubMed]
- Skates, S.J. OCS: Development of the Risk of Ovarian Cancer Algorithm (ROCA) and ROCA Screening Trials. Int. J. Gynecol. Cancer 2012, 22, S24–S26. [Google Scholar] [CrossRef]
- Lu, K.H.; Skates, S.; Hernandez, M.A.; Bedi, D.; Bevers, T.; Leeds, L.; Moore, R.; Granai, C.; Harris, S.; Newland, W.; et al. A 2-Stage Ovarian Cancer Screening Strategy Using the Risk of Ovarian Cancer Algorithm (ROCA) Identifies Early-Stage Incident Cancers and Demonstrates High Positive Predictive Value. Cancer 2013, 119, 3454–3461. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, I.J.; Menon, U.; Ryan, A.; Gentry-Maharaj, A.; Burnell, M.; Kalsi, J.K.; Amso, N.N.; Apostolidou, S.; Benjamin, E.; Cruickshank, D.; et al. Ovarian Cancer Screening and Mortality in the UK Collaborative Trial of Ovarian Cancer Screening (UKCTOCS): A Randomised Controlled Trial. Lancet 2016, 387, 945–956. [Google Scholar] [CrossRef]
- Henderson, J.T.; Webber, E.M.; Sawaya, G.F. Screening for Ovarian Cancer Updated Evidence Report and Systematic Review for the US Preventive Services Task Force. JAMA 2018, 319, 595–606. [Google Scholar] [CrossRef]
- Blackman, A.; Mitchell, J.; Rowswell-Turner, R.; Singh, R.; Kim, K.K.; Eklund, E.; Skates, S.; Bast, R.C.; Messerlian, G.; Miller, M.C.; et al. Analysis of Serum HE4 Levels in Various Histologic Subtypes of Epithelial Ovarian Cancer and Other Malignant Tumors. Tumour Biol. 2021, 43, 355–365. [Google Scholar] [CrossRef]
- Drapkin, R.; von Horsten, H.H.; Lin, Y.; Mok, S.C.; Crum, C.P.; Welch, W.R.; Hecht, J.L. Human Epididymis Protein 4 (HE4) Is a Secreted Glycoprotein That Is Overexpressed by Serous and Endometrioid Ovarian Carcinomas. Cancer Res. 2005, 65, 2162–2169. [Google Scholar] [CrossRef] [PubMed]
- Anderson, G.L.; McIntosh, M.; Wu, L.; Barnett, M.; Goodman, G.; Thorpe, J.D.; Bergan, L.; Thornquist, M.D.; Scholler, N.; Kim, N.; et al. Assessing Lead Time of Selected Ovarian Cancer Biomarkers: A Nested Case-Control Study. J. Natl. Cancer Inst. 2010, 102, 26–38. [Google Scholar] [CrossRef]
- Yurkovetsky, Z.; Skates, S.; Lomakin, A.; Nolen, B.; Pulsipher, T.; Modugno, F.; Marks, J.; Godwin, A.; Gorelik, E.; Jacobs, I.; et al. Development of a Multimarker Assay for Early Detection of Ovarian Cancer. J. Clin. Oncol. 2010, 28, 2159–2166. [Google Scholar] [CrossRef] [PubMed]
- Blyuss, O.; Gentry-Maharaj, A.; Fourkala, E.O.; Ryan, A.; Zaikin, A.; Menon, U.; Jacobs, I.; Timms, J.F. Serial Patterns of Ovarian Cancer Biomarkers in a Prediagnosis Longitudinal Dataset. Biomed. Res. Int. 2015, 2015, 681416. [Google Scholar] [CrossRef] [PubMed]
- Chanhee, H.; Bellone, S.; Siegel, E.R.; Altwerger, G.; Menderes, G.; Bonazzoli, E.; Takata, T.; Petinella, F.; Bianchi, A.; Riccio, F.; et al. A Novel Multiple Biomarker Panel for the Early Detection of High-Grade Serous Ovarian Carcinoma. Gynecol. Oncol. 2018, 149, 585–591. [Google Scholar] [CrossRef]
- Zheng, X.; Chen, S.; Li, L.; Liu, X.; Liu, X.; Dai, S.; Zhang, P.; Lu, H.; Lin, Z.; Yu, Y.; et al. Evaluation of HE4 and TTR for Diagnosis of Ovarian Cancer: Comparison with CA-125. J. Gynecol. Obstet. Hum. Reprod. 2018, 47, 227–230. [Google Scholar] [CrossRef]
- van Calster, B.; van Hoorde, K.; Valentin, L.; Testa, A.C.; Fischerova, D.; van Holsbeke, C.; Savelli, L.; Franchi, D.; Epstein, E.; Kaijser, J.; et al. Evaluating the Risk of Ovarian Cancer before Surgery Using the ADNEX Model to Differentiate between Benign, Borderline, Early and Advanced Stage Invasive, and Secondary Metastatic Tumours: Prospective Multicentre Diagnostic Study. BMJ 2014, 349, g5920. [Google Scholar] [CrossRef]
- Moore, R.G.; McMeekin, D.S.; Brown, A.K.; DiSilvestro, P.; Miller, M.C.; Allard, W.J.; Gajewski, W.; Kurman, R.; Bast, R.C.; Skates, S.J. A Novel Multiple Marker Bioassay Utilizing HE4 and CA125 for the Prediction of Ovarian Cancer in Patients with a Pelvic Mass. Gynecol. Oncol. 2009, 112, 40–46. [Google Scholar] [CrossRef]
- Cui, R.; Wang, Y.; Li, Y.; Li, Y. Clinical Value of ROMA Index in Diagnosis of Ovarian Cancer: Meta-Analysis. Cancer Manag. Res. 2019, 11, 2545–2551. [Google Scholar] [CrossRef]
- Elorriaga, M.Á.; Neyro, J.L.; Mieza, J.; Cristóbal, I.; Llueca, A. Biomarkers in Ovarian Pathology: From Screening to Diagnosis. Review of the Literature. J. Pers. Med. 2021, 11, 1115. [Google Scholar] [CrossRef] [PubMed]
- Goff, B.A.; Agnew, K.; Neradilek, M.B.; Gray, H.J.; Liao, J.B.; Urban, R.R. Combining a Symptom Index, CA125 and HE4 (Triple Screen) to Detect Ovarian Cancer in Women with a Pelvic Mass. Gynecol. Oncol. 2017, 147, 291–295. [Google Scholar] [CrossRef]
- Furrer, D.; Grégoire, J.; Turcotte, S.; Plante, M.; Bachvarov, D.; Trudel, D.; Têtu, B.; Douville, P.; Bairati, I. Performance of Preoperative Plasma HE4 and CA-125 Levels in Predicting Ovarian Cancer Mortality in Women with Epithelial Ovarian Cancer (EOC). PLoS ONE 2019, 14, e0218621. [Google Scholar] [CrossRef]
- Qing, X.; Liu, L.; Mao, X. A Clinical Diagnostic Value Analysis of Serum CA125, CA199, and HE4 in Women with Early Ovarian Cancer: Systematic Review and Meta-Analysis. Comput. Math. Methods Med. 2022, 2022, 9339325. [Google Scholar] [CrossRef] [PubMed]
- Bristow, R.E.; Smith, A.; Zhang, Z.; Chan, D.W.; Crutcher, G.; Fung, E.T.; Munroe, D.G. Ovarian Malignancy Risk Stratification of the Adnexal Mass Using a Multivariate Index Assay. Gynecol. Oncol. 2013, 128, 252–259. [Google Scholar] [CrossRef] [PubMed]
- Fung, E.T. A Recipe for Proteomics Diagnostic Test Development: The OVA1 Test, from Biomarker Discovery to FDA Clearance. Clin. Chem. 2010, 56, 327–329. [Google Scholar] [CrossRef]
- Ueland, F.R.; Desimone, C.P.; Seamon, L.G.; Miller, R.A.; Goodrich, S.; Podzielinski, I.; Sokoll, L.; Smith, A.; van Nagell, J.R.; Zhang, Z. Effectiveness of a Multivariate Index Assay in the Preoperative Assessment of Ovarian Tumors. Obstet. Gynecol. 2011, 117, 1289–1297. [Google Scholar] [CrossRef]
- Coleman, R.L.; Herzog, T.J.; Chan, D.W.; Munroe, D.G.; Pappas, T.C.; Smith, A.; Zhang, Z.; Wolf, J. Validation of a Second-Generation Multivariate Index Assay for Malignancy Risk of Adnexal Masses. Am. J. Obstet. Gynecol. 2016, 215, 82.e1–82.e11. [Google Scholar] [CrossRef]
- Russell, M.R.; Graham, C.; D’Amato, A.; Gentry-Maharaj, A.; Ryan, A.; Kalsi, J.K.; Whetton, A.D.; Menon, U.; Jacobs, I.; Graham, R.L.J. Diagnosis of Epithelial Ovarian Cancer Using a Combined Protein Biomarker Panel. Br. J. Cancer 2019, 121, 483–489. [Google Scholar] [CrossRef]
- Simmons, A.R.; Fourkala, E.O.; Gentry-Maharaj, A.; Ryan, A.; Sutton, M.N.; Baggerly, K.; Zheng, H.; Lu, K.H.; Jacobs, I.; Skates, S.; et al. Complementary Longitudinal Serum Biomarkers to CA125 for Early Detection of Ovarian Cancer. Cancer Prev. Res. 2019, 12, 391–399. [Google Scholar] [CrossRef]
- Mukama, T.; Fortner, R.T.; Katzke, V.; Hynes, L.C.; Petrera, A.; Hauck, S.M.; Johnson, T.; Schulze, M.; Schiborn, C.; Rostgaard-Hansen, A.L.; et al. Prospective Evaluation of 92 Serum Protein Biomarkers for Early Detection of Ovarian Cancer. Br. J. Cancer 2022, 126, 1301–1309. [Google Scholar] [CrossRef]
- Ma, Y.; Wang, X.; Qiu, C.; Qin, J.; Wang, K.; Sun, G.; Jiang, D.; Li, J.; Wang, L.; Shi, J.; et al. Using Protein Microarray to Identify and Evaluate Autoantibodies to Tumor-Associated Antigens in Ovarian Cancer. Cancer Sci. 2021, 112, 537–549. [Google Scholar] [CrossRef]
- Nebgen, D.R.; Lu, K.H.; Bast, R.C. Novel Approaches to Ovarian Cancer Screening. Curr. Oncol. Rep. 2019, 21, 75. [Google Scholar] [CrossRef]
- Yang, W.L.; Gentry-Maharaj, A.; Simmons, A.; Ryan, A.; Fourkala, E.O.; Lu, Z.; Baggerly, K.A.; Zhao, Y.; Lu, K.H.; Bowtell, D.; et al. Elevation of TP53 Autoantibody before CA125 in Preclinical Invasive Epithelial Ovarian Cancer. Clin. Cancer Res. 2017, 23, 5912–5922. [Google Scholar] [CrossRef]
- Wilson, A.L.; Moffitt, L.R.; Duffield, N.; Rainczuk, A.; Jobling, T.W.; Plebanski, M.; Stephens, A.N. Autoantibodies against HSF1 and CCDC155 as Biomarkers of Early-Stage, High-Grade Serous Ovarian Cancer. Cancer Epidemiol. Biomark. Prev. 2018, 27, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Sallum, L.F.; Andrade, L.; Ramalho, S.; Ferracini, A.C.; de Andrade Natal, R.; Borsarelli, A.; Brito, C.; Sarian, L.O.; Derchain, S. WT1, P53 and P16 Expression in the Diagnosis of Low-and High-Grade Serous Ovarian Carcinomas and Their Relation to Prognosis. Oncotarget 2018, 9, 15818–15827. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Jaiswal, S.K.; Kaur, R.; Alsaadi, D.; Liang, X.; Drews, F.; DeLoia, J.A.; Krivak, T.; Petrykowska, H.M.; Gotea, V.; et al. Differential Gene Expression Identifies a Transcriptional Regulatory Network Involving ER-Alpha and PITX1 in Invasive Epithelial Ovarian Cancer. BMC Cancer 2021, 21, 768. [Google Scholar] [CrossRef] [PubMed]
- Bell, D.; Berchuck, A.; Birrer, M.; Chien, J.; Cramer, D.W.; Dao, F.; Dhir, R.; Disaia, P.; Gabra, H.; Glenn, P.; et al. Integrated Genomic Analyses of Ovarian Carcinoma. Nature 2011, 474, 609–615. [Google Scholar] [CrossRef]
- Tothill, R.W.; Tinker, A.V.; George, J.; Brown, R.; Fox, S.B.; Lade, S.; Johnson, D.S.; Trivett, M.K.; Etemadmoghadam, D.; Locandro, B.; et al. Novel Molecular Subtypes of Serous and Endometrioid Ovarian Cancer Linked to Clinical Outcome. Clin. Cancer Res. 2008, 14, 5198–5208. [Google Scholar] [CrossRef] [PubMed]
- Testa, U.; Petrucci, E.; Pasquini, L.; Castelli, G.; Pelosi, E. Ovarian Cancers: Genetic Abnormalities, Tumor Heterogeneity and Progression, Clonal Evolution and Cancer Stem Cells. Medicines 2018, 5, 16. [Google Scholar] [CrossRef] [PubMed]
- Verhaak, R.G.W.; Tamayo, P.; Yang, J.Y.; Hubbard, D.; Zhang, H.; Creighton, C.J.; Fereday, S.; Lawrence, M.; Carter, S.L.; Mermel, C.H.; et al. Prognostically Relevant Gene Signatures of High-Grade Serous Ovarian Carcinoma. J. Clin. Investig. 2013, 123, 517–525. [Google Scholar] [CrossRef] [PubMed]
- Konecny, G.E.; Wang, C.; Hamidi, H.; Winterhoff, B.; Kalli, K.R.; Dering, J.; Ginther, C.; Chen, H.W.; Dowdy, S.; Cliby, W.; et al. Prognostic and Therapeutic Relevance of Molecular Subtypes in High-Grade Serous Ovarian Cancer. J. Natl. Cancer Inst. 2014, 106, dju249. [Google Scholar] [CrossRef]
- Shilpi, A.; Kandpal, M.; Ji, Y.; Seagle, B.L.; Shahabi, S.; Davuluri, R.V. Platform-Independent Classification System to Predict Molecular Subtypes of High-Grade Serous Ovarian Carcinoma. JCO Clin. Cancer Inform. 2019, 3, 1–9. [Google Scholar] [CrossRef]
- Sallinen, H.; Janhonen, S.; Pölönen, P.; Niskanen, H.; Liu, O.H.; Kivelä, A.; Hartikainen, J.M.; Anttila, M.; Heinäniemi, M.; Ylä-Herttuala, S.; et al. Comparative Transcriptome Analysis of Matched Primary and Distant Metastatic Ovarian Carcinoma. BMC Cancer 2019, 19, 1121. [Google Scholar] [CrossRef]
- Sohn, M.H.; Kim, S.I.; Shin, J.Y.; Kim, H.S.; Chung, H.H.; Kim, J.W.; Lee, M.; Seo, J.S. Classification of High-Grade Serous Ovarian Carcinoma by Epithelial-to-Mesenchymal Transition Signature and Homologous Recombination Repair Genes. Genes 2021, 12, 1103. [Google Scholar] [CrossRef]
- Matondo, A.; Jo, Y.H.; Shahid, M.; Choi, T.G.; Nguyen, M.N.; Nguyen, N.N.Y.; Akter, S.; Kang, I.; Ha, J.; Maeng, C.H.; et al. The Prognostic 97 Chemoresponse Gene Signature in Ovarian Cancer. Sci. Rep. 2017, 7, 9689. [Google Scholar] [CrossRef]
- Lee, S.; Zhao, L.; Rojas, C.; Bateman, N.W.; Yao, H.; Lara, O.D.; Celestino, J.; Morgan, M.B.; Nguyen, T.V.; Conrads, K.A.; et al. Molecular Analysis of Clinically Defined Subsets of High-Grade Serous Ovarian Cancer. Cell Rep. 2020, 31, 107502. [Google Scholar] [CrossRef]
- Buttarelli, M.; Ciucci, A.; Palluzzi, F.; Raspaglio, G.; Marchetti, C.; Perrone, E.; Minucci, A.; Giacò, L.; Fagotti, A.; Scambia, G.; et al. Identification of a Novel Gene Signature Predicting Response to First-Line Chemotherapy in BRCA Wild-Type High-Grade Serous Ovarian Cancer Patients. J. Exp. Clin. Cancer Res. 2022, 41, 50. [Google Scholar] [CrossRef]
- McGrail, D.J.; Lin, C.C.J.; Garnett, J.; Liu, Q.; Mo, W.; Dai, H.; Lu, Y.; Yu, Q.; Ju, Z.; Yin, J.; et al. Improved Prediction of PARP Inhibitor Response and Identification of Synergizing Agents through Use of a Novel Gene Expression Signature Generation Algorithm. NPJ Syst. Biol. Appl. 2017, 3, 8. [Google Scholar] [CrossRef]
- Talhouk, A.; George, J.; Wang, C.; Budden, T.; Tan, T.Z.; Chiu, D.S.; Kommoss, S.; Leong, H.S.; Chen, S.; Intermaggio, M.P.; et al. Development and Validation of the Gene Expression Predictor of High-Grade Serous Ovarian Carcinoma Molecular SubTYPE (PrOTYPE). Clin. Cancer Res. 2020, 26, 5411–5423. [Google Scholar] [CrossRef] [PubMed]
- Topno, R.; Singh, I.; Kumar, M.; Agarwal, P. Integrated Bioinformatic Analysis Identifies UBE2Q1 as a Potential Prognostic Marker for High Grade Serous Ovarian Cancer. BMC Cancer 2021, 21, 220. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Park, E.Y.; Kim, O.; Schilder, J.M.; Coffey, D.M.; Cho, C.H.; Bast, R.C. Cell Origins of High-Grade Serous Ovarian Cancer. Cancers 2018, 10, 433. [Google Scholar] [CrossRef]
- Labidi-Galy, S.I.; Papp, E.; Hallberg, D.; Niknafs, N.; Adleff, V.; Noe, M.; Bhattacharya, R.; Novak, M.; Jones, S.; Phallen, J.; et al. High Grade Serous Ovarian Carcinomas Originate in the Fallopian Tube. Nat. Commun. 2017, 8, 1093. [Google Scholar] [CrossRef]
- Fathalla, M.F. Incessant Ovulation-a Factor in Ovarian Neoplasia? Lancet 1971, 298, 163. [Google Scholar] [CrossRef]
- Banet, N.; Kurman, R.J. Two Types of Ovarian Cortical Inclusion Cysts: Proposed Origin and Possible Role in Ovarian Serous Carcinogenesis. Int. J. Gynecol. Pathol. 2015, 34, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Choi, P.W.; So, W.W.; Yang, J.; Liu, S.; Tong, K.K.; Kwan, K.M.; Kwok, J.S.L.; Tsui, S.K.W.; Ng, S.K.; Hales, K.H.; et al. MicroRNA-200 Family Governs Ovarian Inclusion Cyst Formation and Mode of Ovarian Cancer Spread. Oncogene 2020, 39, 4045–4060. [Google Scholar] [CrossRef]
- Fleszar, A.J.; Walker, A.; Porubsky, V.; Flanigan, W.; James, D.; Campagnola, P.J.; Weisman, P.S.; Kreeger, P.K. The Extracellular Matrix of Ovarian Cortical Inclusion Cysts Modulates Invasion of Fallopian Tube Epithelial Cells. APL Bioeng. 2018, 2, 031902. [Google Scholar] [CrossRef]
- Ahmed, N.; Abubaker, K.; Findlay, J.; Quinn, M. Cancerous Ovarian Stem Cells: Obscure Targets for Therapy but Relevant to Chemoresistance. J. Cell Biochem. 2013, 114, 21–34. [Google Scholar] [CrossRef]
- Klotz, D.M.; Wimberger, P. Cells of Origin of Ovarian Cancer: Ovarian Surface Epithelium or Fallopian Tube? Arch. Gynecol. Obstet. 2017, 296, 1055–1062. [Google Scholar] [CrossRef] [PubMed]
- Auersperg, N.; Wong, A.S.T.; Choi, K.C.; Kang, S.K.; Leung, P.C.K. Ovarian Surface Epithelium: Biology, Endocrinology, and Pathology. Endocr. Rev. 2001, 22, 255–288. [Google Scholar] [CrossRef] [PubMed]
- King, M.C.; Marks, J.H.; Mandell, J.B. Breast and Ovarian Cancer Risks Due to Inherited Mutations in BRCA1 and BRCA2. Science 2003, 302, 643–646. [Google Scholar] [CrossRef] [PubMed]
- Roy, R.; Chun, J.; Powell, S.N. BRCA1 and BRCA2: Different Roles in a Common Pathway of Genome Protection. Nat. Rev. Cancer 2012, 12, 68–78. [Google Scholar] [CrossRef] [PubMed]
- Piek, J.M.J.; van Diest, P.J.; Zweemer, R.P.; Jansen, J.W.; Poort-Keesom, R.J.J.; Menko, F.H.; Gille, J.J.P.; Jongsma, A.P.M.; Pals, G.; Kenemans, P.; et al. Dysplastic Changes in Prophylactically Removed Fallopian Tubes of Women Predisposed to Developing Ovarian Cancer. J. Pathol. 2001, 195, 451–456. [Google Scholar] [CrossRef]
- Kuhn, E.; Kurman, R.J.; Shih, I.-M. Ovarian Cancer Is an Imported Disease: Fact or Fiction? Curr. Obstet. Gynecol. Rep. 2012, 1, 1–9. [Google Scholar] [CrossRef]
- Finch, A.P.M.; Lubinski, J.; Møller, P.; Singer, C.F.; Karlan, B.; Senter, L.; Rosen, B.; Maehle, L.; Ghadirian, P.; Cybulski, C.; et al. Impact of Oophorectomy on Cancer Incidence and Mortality in Women with a BRCA1 or BRCA2 Mutation. J. Clin. Oncol. 2014, 32, 1547–1553. [Google Scholar] [CrossRef]
- Hartmann, L.C.; Lindor, N.M. The Role of Risk-Reducing Surgery in Hereditary Breast and Ovarian Cancer. N. Engl. J. Med. 2016, 374, 454–468. [Google Scholar] [CrossRef]
- Kurman, R.J.; Shih, I.M. The Dualistic Model of Ovarian Carcinogenesis Revisited, Revised, and Expanded. Am. J. Pathol. 2016, 186, 733–747. [Google Scholar] [CrossRef]
- Jarboe, E.; Folkins, A.; Nucci, M.R.; Kindelberger, D.; Drapkin, R.; Miron, A.; Lee, Y.; Crum, C.P. Serous Carcinogenesis in the Fallopian Tube: A Descriptive Classification. Int. J. Gynecol. Pathol. 2008, 27, 1–9. [Google Scholar] [CrossRef]
- Visvanathan, K.; Vang, R.; Shaw, P.; Gross, A.; Soslow, R.; Parkash, V.; Shih, I.M.; Kurman, R.J. Diagnosis of Serous Tubal Intraepithelial Carcinoma Based on Morphologic and Immunohistochemical Features: A Reproducibility Study. Am. J. Surg. Pathol. 2011, 35, 1766–1775. [Google Scholar] [CrossRef] [PubMed]
- Wethington, S.L.; Park, K.J.; Soslow, R.A.; Kauff, N.D.; Brown, C.L.; Dao, F.; Otegbeye, E.; Sonoda, Y.; Abu-Rustum, N.R.; Barakat, R.R.; et al. Clinical Outcome of Isolated Serous Tubal Intraepithelial Carcinomas (STIC). Int. J. Gynecol. Cancer 2013, 23, 1603–1611. [Google Scholar] [CrossRef] [PubMed]
- Howitt, B.E.; Hanamornroongruang, S.; Lin, D.I.; Conner, J.E.; Schulte, S.; Horowitz, N.; Crum, C.P.; Meserve, E.E. Evidence for a Dualistic Model of High-Grade Serous Carcinoma: BRCA Mutation Status, Histology, and Tubal Intraepithelial Carcinoma. Am. J. Surg. Pathol. 2015, 39, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, E.; Kurman, R.J.; Vang, R.; Sehdev, A.S.; Han, G.; Soslow, R.; Wang, T.L.; Shih, I.M. TP53 Mutations in Serous Tubal Intraepithelial Carcinoma and Concurrent Pelvic High-Grade Serous Carcinoma-Evidence Supporting the Clonal Relationship of the Two Lesions. J. Pathol. 2012, 226, 421–426. [Google Scholar] [CrossRef]
- Kuhn, E.; Wang, T.L.; Doberstein, K.; Bahadirli-Talbott, A.; Ayhan, A.; Sehdev, A.S.; Drapkin, R.; Kurman, R.J.; Shih, I.M. CCNE1 Amplification and Centrosome Number Abnormality in Serous Tubal Intraepithelial Carcinoma: Further Evidence Supporting Its Role as a Precursor of Ovarian High-Grade Serous Carcinoma. Mod. Pathol. 2016, 29, 1254–1261. [Google Scholar] [CrossRef]
- Kindelberger, D.W.; Lee, Y.; Miron, A.; Hirsch, M.S.; Feltmate, C.; Medeiros, F.; Callahan, M.J.; Garner, E.O.; Gordon, R.W.; Birch, C.; et al. Intraepithelial Carcinoma of the Fimbria and Pelvic Serous Carcinoma: Evidence for a Causal Relationship. Am. J. Surg. Pathol. 2007, 31, 161–169. [Google Scholar] [CrossRef]
- Lengyel, E. Ovarian Cancer Development and Metastasis. Am. J. Pathol. 2010, 177, 1053–1064. [Google Scholar] [CrossRef]
- Asaka, S.; Davis, C.; Lin, S.F.; Wang, T.L.; Heaphy, C.M.; Shih, I.M. Analysis of Telomere Lengths in P53 Signatures and Incidental Serous Tubal Intraepithelial Carcinomas Without Concurrent Ovarian Cancer. Am. J. Surg. Pathol. 2019, 43, 1083–1091. [Google Scholar] [CrossRef]
- Kuhn, E.; Meeker, A.; Wang, T.L.; Sehdev, A.S.; Kurman, R.J.; Shih, I.M. Shortened Telomeres in Serous Tubal Intraepithelial Carcinoma: An Early Event in Ovarian High-Grade Serous Carcinogenesis. Am. J. Surg. Pathol. 2010, 34, 829–836. [Google Scholar] [CrossRef]
- Akahane, T.; Masuda, K.; Hirasawa, A.; Kobayashi, Y.; Ueki, A.; Kawaida, M.; Misu, K.; Nakamura, K.; Nagai, S.; Chiyoda, T.; et al. TP53 Variants in P53 Signatures and the Clonality of STICs in RRSO Samples. J. Gynaecol. Oncol. 2022, 33, e50. [Google Scholar] [CrossRef]
- Kyo, S.; Ishikawa, N.; Nakamura, K.; Nakayama, K. The Fallopian Tube as Origin of Ovarian Cancer: Change of Diagnostic and Preventive Strategies. Cancer Med. 2020, 9, 421–431. [Google Scholar] [CrossRef] [PubMed]
- Karnezis, A.N.; Cho, K.R.; Gilks, C.B.; Pearce, C.L.; Huntsman, D.G. The Disparate Origins of Ovarian Cancers: Pathogenesis and Prevention Strategies. Nat. Rev. Cancer 2017, 17, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Lawrenson, K.; Fonseca, M.A.S.; Liu, A.Y.; Segato Dezem, F.; Lee, J.M.; Lin, X.; Corona, R.I.; Abbasi, F.; Vavra, K.C.; Dinh, H.Q.; et al. A Study of High-Grade Serous Ovarian Cancer Origins Implicates the SOX18 Transcription Factor in Tumor Development. Cell Rep. 2019, 29, 3726–3735.e4. [Google Scholar] [CrossRef] [PubMed]
- Ducie, J.; Dao, F.; Considine, M.; Olvera, N.; Shaw, P.A.; Kurman, R.J.; Shih, I.M.; Soslow, R.A.; Cope, L.; Levine, D.A. Molecular Analysis of High-Grade Serous Ovarian Carcinoma with and without Associated Serous Tubal Intra-Epithelial Carcinoma. Nat. Commun. 2017, 8, 990. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Dolgalev, I.; Zhang, T.; Ran, H.; Levine, D.A.; Neel, B.G. Both Fallopian Tube and Ovarian Surface Epithelium Are Cells-of-Origin for High-Grade Serous Ovarian Carcinoma. Nat. Commun. 2019, 10, 5367. [Google Scholar] [CrossRef]
- Lõhmussaar, K.; Kopper, O.; Korving, J.; Begthel, H.; Vreuls, C.P.H.; van Es, J.H.; Clevers, H. Assessing the Origin of High-Grade Serous Ovarian Cancer Using CRISPR-Modification of Mouse Organoids. Nat. Commun. 2020, 11, 2660. [Google Scholar] [CrossRef]
- Chan, K.K.L.; Siu, M.K.Y.; Jiang, Y.X.; Wang, J.J.; Wang, Y.; Leung, T.H.Y.; Liu, S.S.; Cheung, A.N.Y.; Ngan, H.Y.S. Differential Expression of Estrogen Receptor Subtypes and Variants in Ovarian Cancer: Effects on Cell Invasion, Proliferation and Prognosis. BMC Cancer 2017, 17, 606. [Google Scholar] [CrossRef]
- Pavlič, R.; Gjorgoska, M.; Rižner, T.L. Model Cell Lines and Tissues of Different HGSOC Subtypes Differ in Local Estrogen Biosynthesis. Cancers 2022, 14, 2583. [Google Scholar] [CrossRef]
- Li, H.; Liu, Y.; Wang, Y.; Zhao, X.; Qi, X. Hormone Therapy for Ovarian Cancer: Emphasis on Mechanisms and Applications. Oncol. Rep. 2021, 46, 223. [Google Scholar] [CrossRef]
- Gibson, D.A.; Simitsidellis, I.; Collins, F.; Saunders, P.T.K. Evidence of Androgen Action in Endometrial and Ovarian Cancers. Endocr. Relat. Cancer 2014, 21, 203–218. [Google Scholar] [CrossRef]
- Mizushima, T.; Miyamoto, H. The Role of Androgen Receptor Signaling in Ovarian Cancer. Cells 2019, 8, 176. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Li, S.; Zhao, M.; Sheng, B.; Zhu, H.; Zhu, X. Prognostic Value of Progesterone Receptor in Ovarian Cancer: A Meta-Analysis. Chin. J. Evid.-Based Med. 2017, 17, 327–333. [Google Scholar] [CrossRef] [PubMed]
- Schüler, S.; Ponnath, M.; Engel, J.; Ortmann, O. Ovarian Epithelial Tumors and Reproductive Factors: A Systematic Review. Arch. Gynecol. Obstet. 2013, 287, 1187–1204. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, C.; Patel, A.V.; Calle, E.E.; Jacob, E.J.; Thun, M.J. Estrogen Replacement Therapy and Ovarian Cancer Mortality in a Large Prospective Study of US Women. J. Am. Med. Assoc. 2001, 285, 1460–1465. [Google Scholar] [CrossRef] [PubMed]
- Cramer, D.W.; Welch, W.R. Determinants of Ovarian Cancer Risk. II. Inferences Regarding Pathogenesis. J. Natl. Cancer Inst. 1983, 71, 717–721. [Google Scholar]
- Song, K.; Dai, L.; Long, X.; Wang, W.; Di, W. Follicle-Stimulating Hormone Promotes the Proliferation of Epithelial Ovarian Cancer Cells by Activating Sphingosine Kinase. Sci. Rep. 2020, 10, 13834. [Google Scholar] [CrossRef]
- Lau, M.T.; Wong, A.S.T.; Leung, P.C.K. Gonadotropins Induce Tumor Cell Migration and Invasion by Increasing Cyclooxygenases Expression and Prostaglandin E2 Production in Human Ovarian Cancer Cells. Endocrinology 2010, 151, 2985–2993. [Google Scholar] [CrossRef]
- Feng, D.; Zhao, T.; Yan, K.; Liang, H.; Liang, J.; Zhou, Y.; Zhao, W.; Ling, B. Gonadotropins Promote Human Ovarian Cancer Cell Migration and Invasion via a Cyclooxygenase 2-Dependent Pathway. Oncol. Rep. 2017, 38, 1091–1098. [Google Scholar] [CrossRef]
- Warrenfeltz, S.W.; Lott, S.A.; Palmer, T.M.; Gray, J.C.; Puett, D. Luteinizing Hormone-Induced up-Regulation of ErbB-2 Is Insufficient Stimulant of Growth and Invasion in Ovarian Cancer Cells. Mol. Cancer Res. 2008, 6, 1775–1785. [Google Scholar] [CrossRef]
- Zhu, H.; Zhu, X.; Zheng, L.; Hu, X.; Sun, L.Z.; Zhu, X. The Role of the Androgen Receptor in Ovarian Cancer Carcinogenesis and Its Clinical Implications. Oncotarget 2017, 8, 29395–29405. [Google Scholar] [CrossRef]
- Rahim, B.; O’Regan, R. AR Signaling in Breast Cancer. Cancers 2017, 9, 21. [Google Scholar] [CrossRef] [PubMed]
- Calvillo-Robledo, A.; Pedernera, E.; Morales-Vásquez, F.; Pérez-Montiel, D.; Gómora, M.J.; Almaraz, M.Á.; de Alba Graue, P.G.; Rendón, E.; López-Basave, H.N.; Quintanar-Stephano, A.; et al. Simultaneous Expression of Steroid Sulfatase and Androgen Receptor Reduced Overall Survival of Patients with Epithelial Ovarian Tumors. J. Ovarian Res. 2021, 14, 98. [Google Scholar] [CrossRef] [PubMed]
- Elattar, A.; Warburton, K.G.; Mukhopadhyay, A.; Freer, R.M.; Shaheen, F.; Cross, P.; Plummer, E.R.; Robson, C.N.; Edmondson, R.J. Androgen Receptor Expression Is a Biological Marker for Androgen Sensitivity in High Grade Serous Epithelial Ovarian Cancer. Gynecol. Oncol. 2012, 124, 142–147. [Google Scholar] [CrossRef] [PubMed]
- de Toledo, M.C.S.; Sarian, L.O.; Sallum, L.F.; Andrade, L.L.A.; Vassallo, J.; de Paiva Silva, G.R.; Pinto, G.A.; Soares, F.A.; Fonseca, C.D.P.P.; Derchain, S.F.M. Analysis of the Contribution of Immunologically-Detectable HER2, Steroid Receptors and of the “Triple-Negative” Tumor Status to Disease-Free and Overall Survival of Women with Epithelial Ovarian Cancer. Acta Histochem 2014, 116, 440–447. [Google Scholar] [CrossRef]
- Feng, Z.; Wen, H.; Bi, R.; Ju, X.; Chen, X.; Yang, W.; Wu, X. A Clinically Applicable Molecular Classification for High-Grade Serous Ovarian Cancer Based on Hormone Receptor Expression. Sci. Rep. 2016, 6, 25408. [Google Scholar] [CrossRef]
- Feng, Z.; Wen, H.; Ju, X.; Bi, R.; Chen, X.; Yang, W.; Wu, X. Hormone Receptor Expression Profiles Differ between Primary and Recurrent High-Grade Serous Ovarian Cancers. Oncotarget 2017, 8, 32848–32855. [Google Scholar] [CrossRef]
- Maclean, A.; Bunni, E.; Makrydima, S.; Withington, A.; Kamal, A.M.; Valentijn, A.J.; Hapangama, D.K. Fallopian Tube Epithelial Cells Express Androgen Receptor and Have a Distinct Hormonal Responsiveness When Compared with Endometrial Epithelium. Hum. Reprod. 2020, 35, 2097–2106. [Google Scholar] [CrossRef]
- Chen, S.; Dai, X.; Gao, Y.; Shen, F.; Ding, J.; Chen, Q. The Positivity of Estrogen Receptor and Progesterone Receptor May Not Be Associated with Metastasis and Recurrence in Epithelial Ovarian Cancer. Sci. Rep. 2017, 7, 16922. [Google Scholar] [CrossRef]
- Troisi, R.; Bjørge, T.; Gissler, M.; Grotmol, T.; Kitahara, C.M.; Myrtveit Sæther, S.M.; Ording, A.G.; Sköld, C.; Sørensen, H.T.; Trabert, B.; et al. The Role of Pregnancy, Perinatal Factors and Hormones in Maternal Cancer Risk: A Review of the Evidence. J. Intern. Med. 2018, 283, 430–445. [Google Scholar] [CrossRef]
- Fu, Z.; Taylor, S.; Modugno, F. Lifetime Ovulations and Epithelial Ovarian Cancer Risk and Survival: A Systematic Review and Meta-Analysis. Gynecol. Oncol. 2022, 165, 650–663. [Google Scholar] [CrossRef]
- Liu, Y.; Ma, L.; Yang, X.; Bie, J.; Li, D.; Sun, C.; Zhang, J.; Meng, Y.; Lin, J. Menopausal Hormone Replacement Therapy and the Risk of Ovarian Cancer: A Meta-Analysis. Front. Endocrinol. 2019, 10, 16922. [Google Scholar] [CrossRef] [PubMed]
- Trabert, B.; Tworoger, S.S.; O’Brien, K.M.; Townsend, M.K.; Fortner, R.T.; Iversen, E.S.; Hartge, P.; White, E.; Amiano, P.; Arslan, A.A.; et al. The Risk of Ovarian Cancer Increases with an Increase in the Lifetime Number of Ovulatory Cycles: An Analysis from the Ovarian Cancer Cohort Consortium (OC3). Cancer Res. 2020, 80, 1210–1218. [Google Scholar] [CrossRef] [PubMed]
- Toufakis, V.; Katuwal, S.; Pukkala, E.; Tapanainen, J.S. Impact of Parity on the Incidence of Ovarian Cancer Subtypes: A Population-Based Case–Control Study. Acta Oncol. 2021, 60, 850–855. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.; Zhai, Y.; Hu, K.; Liu, C.J.; Udager, A.; Pearce, C.L.; Fearon, E.R.; Cho, K.R. Aging Accelerates While Multiparity Delays Tumorigenesis in Mouse Models of High-Grade Serous Carcinoma. Gynecol. Oncol. 2022, 165, 552–559. [Google Scholar] [CrossRef] [PubMed]
- Kotsopoulos, J.; Lubinski, J.; Gronwald, J.; Cybulski, C.; Demsky, R.; Neuhausen, S.L.; Kim-Sing, C.; Tung, N.; Friedman, S.; Senter, L.; et al. Factors Influencing Ovulation and the Risk of Ovarian Cancer in BRCA1 and BRCA2 Mutation Carriers. Int. J. Cancer 2015, 137, 1136–1146. [Google Scholar] [CrossRef] [PubMed]
- Modugno, F.; Goughnour, S.L.; Wallack, D.; Edwards, R.P.; Odunsi, K.; Kelley, J.L.; Moysich, K.; Ness, R.B.; Brooks, M.M. Breastfeeding Factors and Risk of Epithelial Ovarian Cancer. Gynecol. Oncol. 2019, 153, 116–122. [Google Scholar] [CrossRef]
- Babic, A.; Sasamoto, N.; Rosner, B.A.; Tworoger, S.S.; Jordan, S.J.; Risch, H.A.; Harris, H.R.; Rossing, M.A.; Doherty, J.A.; Fortner, R.T.; et al. Association between Breastfeeding and Ovarian Cancer Risk. JAMA Oncol. 2020, 6, e200421. [Google Scholar] [CrossRef]
- Michels, K.A.; Pfeiffer, R.M.; Brinton, L.A.; Trabert, B. Modification of the Associations between Duration of Oral Contraceptive Use and Ovarian, Endometrial, Breast, and Colorectal Cancers. JAMA Oncol. 2018, 4, 516–521. [Google Scholar] [CrossRef]
- Havrilesky, L.J.; Moorman, P.G.; Lowery, W.J.; Gierisch, J.M.; Coeytaux, R.R.; Urrutia, R.P.; Dinan, M.; McBroom, A.J.; Hasselblad, V.; Sanders, G.D.; et al. Oral Contraceptive Pills as Primary Prevention for Ovarian Cancer: A Systematic Review and Meta-Analysis. Obstet. Gynecol. 2013, 122, 139–147. [Google Scholar] [CrossRef]
- Tan, J.; Song, C.; Wang, D.; Hu, Y.; Liu, D.; Ma, D.; Gao, Q. Expression of Hormone Receptors Predicts Survival and Platinum Sensitivity of High-Grade Serous Ovarian Cancer. BioSci. Rep. 2021, 41, BSR20210478. [Google Scholar] [CrossRef]
- Millstein, J.; Budden, T.; Goode, E.L.; Anglesio, M.S.; Talhouk, A.; Intermaggio, M.P.; Leong, H.S.; Chen, S.; Elatre, W.; Gilks, B.; et al. Prognostic Gene Expression Signature for High-Grade Serous Ovarian Cancer. Ann. Oncol. 2020, 31, 1240–1250. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Lheureux, S.; Karakasis, K.; Burnier, J.V.; Bruce, J.P.; Clouthier, D.L.; Danesh, A.; Quevedo, R.; Dowar, M.; Hanna, Y.; et al. Landscape of Genomic Alterations in High-Grade Serous Ovarian Cancer from Exceptional Long- and Short-Term Survivors. Genome Med. 2018, 10, 81. [Google Scholar] [CrossRef] [PubMed]
- Yamulla, R.J.; Nalubola, S.; Flesken-Nikitin, A.; Nikitin, A.Y.; Schimenti, J.C. Most Commonly Mutated Genes in High-Grade Serous Ovarian Carcinoma Are Nonessential for Ovarian Surface Epithelial Stem Cell Transformation. Cell Rep. 2020, 32, 108086. [Google Scholar] [CrossRef] [PubMed]
- Phelan, C.M.; Kuchenbaecker, K.B.; Tyrer, J.P.; Kar, S.P.; Lawrenson, K.; Winham, S.J.; Dennis, J.; Pirie, A.; Riggan, M.J.; Chornokur, G.; et al. Identification of 12 New Susceptibility Loci for Different Histotypes of Epithelial Ovarian Cancer. Nat. Genet. 2017, 49, 680–691. [Google Scholar] [CrossRef]
- Lawrenson, K.; Li, Q.; Kar, S.; Seo, J.H.; Tyrer, J.; Spindler, T.J.; Lee, J.; Chen, Y.; Karst, A.; Drapkin, R.; et al. Cis-EQTL Analysis and Functional Validation of Candidate Susceptibility Genes for High-Grade Serous Ovarian Cancer. Nat. Commun. 2015, 6, 8234. [Google Scholar] [CrossRef]
- Gusev, A.; Lawrenson, K.; Lin, X.; Lyra, P.C.; Kar, S.; Vavra, K.C.; Segato, F.; Fonseca, M.A.S.; Lee, J.M.; Pejovic, T.; et al. A Transcriptome-Wide Association Study of High-Grade Serous Epithelial Ovarian Cancer Identifies New Susceptibility Genes and Splice Variants. Nat. Genet. 2019, 51, 815–823. [Google Scholar] [CrossRef]
- Kar, S.P.; Considine, D.P.C.; Tyrer, J.P.; Plummer, J.T.; Chen, S.; Dezem, F.S.; Barbeira, A.N.; Rajagopal, P.S.; Rosenow, W.T.; Moreno, F.; et al. Pleiotropy-Guided Transcriptome Imputation from Normal and Tumor Tissues Identifies Candidate Susceptibility Genes for Breast and Ovarian Cancer. Hum. Genet. Genom. Adv. 2021, 2, 100042. [Google Scholar] [CrossRef]
- Cheng, Z.; Mirza, H.; Ennis, D.P.; Smith, P.; Morrill Gavarró, L.; Sokota, C.; Giannone, G.; Goranova, T.; Bradley, T.; Piskorz, A.; et al. The Genomic Landscape of Early-Stage Ovarian High-Grade Serous Carcinoma. Clin. Cancer Res. 2022, 28, 2911–2922. [Google Scholar] [CrossRef]
- Longacre, M.; Snyder, N.A.; Housman, G.; Leary, M.; Lapinska, K.; Heerboth, S.; Willbanks, A.; Sarkar, S. A Comparative Analysis of Genetic and Epigenetic Events of Breast and Ovarian Cancer Related to Tumorigenesis. Int. J. Mol. Sci. 2016, 17, 759. [Google Scholar] [CrossRef]
- Patch, A.M.; Christie, E.L.; Etemadmoghadam, D.; Garsed, D.W.; George, J.; Fereday, S.; Nones, K.; Cowin, P.; Alsop, K.; Bailey, P.J.; et al. Whole-Genome Characterization of Chemoresistant Ovarian Cancer. Nature 2015, 521, 489–494. [Google Scholar] [CrossRef]
- Micci, F.; Haugom, L.; Abeler, V.M.; Davidson, B.; Tropé, C.G.; Heim, S. Genomic Profile of Ovarian Carcinomas. BMC Cancer 2014, 14, 315. [Google Scholar] [CrossRef] [PubMed]
- Engqvist, H.; Parris, T.Z.; Biermann, J.; Rönnerman, E.W.; Larsson, P.; Sundfeldt, K.; Kovács, A.; Karlsson, P.; Helou, K. Integrative Genomics Approach Identifies Molecular Features Associated with Early-Stage Ovarian Carcinoma Histotypes. Sci. Rep. 2020, 10, 7946. [Google Scholar] [CrossRef] [PubMed]
- Evans, D.G.R.; van Veen, E.M.; Byers, H.J.; Wallace, A.J.; Ellingford, J.M.; Beaman, G.; Santoyo-Lopez, J.; Aitman, T.J.; Eccles, D.M.; Lalloo, F.I.; et al. A Dominantly Inherited 5′ UTR Variant Causing Methylation-Associated Silencing of BRCA1 as a Cause of Breast and Ovarian Cancer. Am. J. Hum. Genet. 2018, 103, 213–220. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M. Epigenetics in Cancer. N. Engl. J. Med. 2008, 358, 1148–1159. [Google Scholar] [CrossRef]
- Baylin, S.B.; Jones, P.A. Epigenetic Determinants of Cancer. Cold Spring Harb. Perspect. Biol. 2016, 8, 1–35. [Google Scholar] [CrossRef]
- Karakashev, S.; Zhu, H.; Wu, S.; Yokoyama, Y.; Bitler, B.G.; Park, P.H.; Lee, J.H.; Kossenkov, A.V.; Gaonkar, K.S.; Yan, H.; et al. CARM1-Expressing Ovarian Cancer Depends on the Histone Methyltransferase EZH2 Activity. Nat. Commun. 2018, 9, 631. [Google Scholar] [CrossRef]
- Khabele, D. The Therapeutic Potential of Class I Selective Histone Deacetylase Inhibitors in Ovarian Cancer. Front. Oncol. 2014, 4, 111. [Google Scholar] [CrossRef]
- Kobayashi, M.; Sawada, K.; Nakamura, K.; Yoshimura, A.; Miyamoto, M.; Shimizu, A.; Ishida, K.; Nakatsuka, E.; Kodama, M.; Hashimoto, K.; et al. Exosomal MiR-1290 Is a Potential Biomarker of High-Grade Serous Ovarian Carcinoma and Can Discriminate Patients from Those with Malignancies of Other Histological Types. J. Ovarian Res. 2018, 11, 81. [Google Scholar] [CrossRef]
- Li, R.; Wu, H.; Jiang, H.; Wang, Q.; Dou, Z.; Ma, H.; Yan, S.; Yuan, C.; Yang, N.; Kong, B. FBLN5 Is Targeted by MicroRNA-27a-3p and Suppresses Tumorigenesis and Progression in High-Grade Serous Ovarian Carcinoma. Oncol. Rep. 2020, 44, 2143–2151. [Google Scholar] [CrossRef]
- Mirahmadi, Y.; Nabavi, R.; Taheri, F.; Samadian, M.M.; Ghale-Noie, Z.N.; Farjami, M.; Samadi-Khouzani, A.; Yousefi, M.; Azhdari, S.; Salmaninejad, A.; et al. MicroRNAs as Biomarkers for Early Diagnosis, Prognosis, and Therapeutic Targeting of Ovarian Cancer. J. Oncol. 2021, 2021, 3408937. [Google Scholar] [CrossRef]
- Frugtniet, B.; Morgan, S.; Murray, A.; Palmer-Smith, S.; White, R.; Jones, R.; Hanna, L.; Fuller, C.; Hudson, E.; Mullard, A.; et al. The Detection of Germline and Somatic BRCA1/2 Genetic Variants through Parallel Testing of Patients with High-Grade Serous Ovarian Cancer: A National Retrospective Audit. BJOG 2022, 129, 433–442. [Google Scholar] [CrossRef] [PubMed]
- Walsh, T.; Casadei, S.; Lee, M.K.; Pennil, C.C.; Nord, A.S.; Thornton, A.M.; Roeb, W.; Agnew, K.J.; Stray, S.M.; Wickramanayake, A.; et al. Mutations in 12 Genes for Inherited Ovarian, Fallopian Tube, and Peritoneal Carcinoma Identified by Massively Parallel Sequencing. Proc. Natl. Acad. Sci. USA 2011, 108, 18032–18037. [Google Scholar] [CrossRef] [PubMed]
- Hauke, J.; Hahnen, E.; Schneider, S.; Reuss, A.; Richters, L.; Kommoss, S.; Heimbach, A.; Marmé, F.; Schmidt, S.; Prieske, K.; et al. Deleterious Somatic Variants in 473 Consecutive Individuals with Ovarian Cancer: Results of the Observational AGO-TR1 Study (NCT02222883). J. Med. Genet. 2019, 56, 574–580. [Google Scholar] [CrossRef] [PubMed]
- Risch, H.A.; McLaughlin, J.R.; Cole, D.E.C.; Rosen, B.; Bradley, L.; Fan, I.; Tang, J.; Li, S.; Zhang, S.; Shaw, P.A.; et al. Population BRCA1 and BRCA2 Mutation Frequencies and Cancer Penetrances: A Kin-Cohort Study in Ontario, Canada. J. Natl. Cancer Inst. 2006, 98, 1694–1706. [Google Scholar] [CrossRef] [PubMed]
- Norquist, B.M.; Harrell, M.I.; Brady, M.F.; Walsh, T.; Lee, M.K.; Gulsuner, S.; Bernards, S.S.; Casadei, S.; Yi, Q.; Burger, R.A.; et al. Inherited Mutations in Women with Ovarian Carcinoma. JAMA Oncol. 2016, 2, 482–490. [Google Scholar] [CrossRef]
- Song, H.; Dicks, E.; Ramus, S.J.; Tyrer, J.P.; Intermaggio, M.P.; Hayward, J.; Edlund, C.K.; Conti, D.; Harrington, P.; Fraser, L.; et al. Contribution of Germline Mutations in the RAD51B, RAD51C, and RAD51D Genes to Ovarian Cancer in the Population. J. Clin. Oncol. 2015, 33, 2901–2907. [Google Scholar] [CrossRef]
- Turchiano, A.; Loconte, D.C.; De Nola, R.; Arezzo, F.; Chiarello, G.; Pantaleo, A.; Iacoviello, M.; Bagnulo, R.; De Luisi, A.; Perrelli, S.; et al. Beyond Brca1/2: Homologous Recombination Repair Genetic Profile in A Large Cohort of Apulian Ovarian Cancers. Cancers 2022, 14, 365. [Google Scholar] [CrossRef]
- Weiss, A.S.; Swisher, E.; Pennington, K.P.; Radke, M.; Khasnavis, N.; Garcia, R.L.; Kilgore, M.R.; Lee, M.K.; Norquist, B.M. Inherited Mutations in Fallopian Tube, Ovarian and Primary Peritoneal Carcinoma: Changes in Diagnoses and Mutational Frequency over 20 Years. Gynecol. Oncol. 2020, 159, 214–220. [Google Scholar] [CrossRef]
- Holloman, W.K. Unraveling the Mechanism of BRCA2 in HR. Nat. Struct. Mol. Biol. 2013, 18, 748–754. [Google Scholar] [CrossRef]
- Prakash, R.; Zhang, Y.; Feng, W.; Jasin, M. Homologous Recombination and Human Health: The Roles of BRCA1, BRCA2, and Associated Proteins. Cold Spring Harb. Perspect. Biol. 2015, 7, 1–27. [Google Scholar] [CrossRef]
- Takaya, H.; Nakai, H.; Takamatsu, S.; Mandai, M.; Matsumura, N. Homologous Recombination Deficiency Status-Based Classification of High-Grade Serous Ovarian Carcinoma. Sci. Rep. 2020, 10, 2757. [Google Scholar] [CrossRef]
- Creeden, J.F.; Nanavaty, N.S.; Einloth, K.R.; Gillman, C.E.; Stanbery, L.; Hamouda, D.M.; Dworkin, L.; Nemunaitis, J. Homologous Recombination Proficiency in Ovarian and Breast Cancer Patients. BMC Cancer 2021, 21, 1154. [Google Scholar] [CrossRef] [PubMed]
- Pellegrino, B.; Herencia-Roper, A.; Llop-Guevar, A.; Pedretti, F.; Moles-Fernández, A.; Viaplana, C.; Villacampa, G.; Guzmán, M.; Rodríguez, O.; Grueso, J.; et al. Preclinical In Vivo Validation of the RAD51 Test for Identification of Homologous Recombination-Deficient Tumors and Patient Stratification. Cancer Res. 2022, 82, 1646–1657. [Google Scholar] [CrossRef] [PubMed]
- Liontos, M. Testing of Newly Diagnosed Advanced High Grade Ovarian Cancer (OC) Patients with the Myriad Genetics MyChoice CDx Plus next Generation Sequencing-Based in Vitro Diagnostic Test Emphasizes the Need for Public Insurance Coverage of Genetic Testing: Results of a National Program by the Hellenic Society of Medical Oncology (HeSMO). J. Clin. Oncol. 2022, 40, e18520. [Google Scholar] [CrossRef]
- Uffelmann, E.; Huang, Q.Q.; Munung, N.S.; de Vries, J.; Okada, Y.; Martin, A.R.; Martin, H.C.; Lappalainen, T.; Posthuma, D. Genome-Wide Association Studies. Nat. Rev. Methods Prim. 2021, 1, 59. [Google Scholar] [CrossRef]
- Manichaikul, A.; Peres, L.C.; Wang, X.Q.; Barnard, M.E.; Chyn, D.; Sheng, X.; Du, Z.; Tyrer, J.; Dennis, J.; Schwartz, A.G.; et al. Identification of Novel Epithelial Ovarian Cancer Loci in Women of African Ancestry. Int. J. Cancer 2019, 176, 139–148. [Google Scholar] [CrossRef]
- Flaum, N.; Crosbie, E.J.; Edmondson, R.J.; Smith, M.J.; Evans, D.G. Epithelial Ovarian Cancer Risk: A Review of the Current Genetic Landscape. Clin. Genet. 2020, 97, 54–63. [Google Scholar] [CrossRef] [PubMed]
- Nameki, R.; Shetty, A.; Dareng, E.; Tyrer, J.; Lin, X.; Pharoah, P.; Corona, R.I.; Kar, S.; Lawrenson, K. ChromMAGMA: Regulatory Element-Centric Interrogation of Risk Variants. bioRxiv 2022. [Google Scholar] [CrossRef]
- Schaafsma, E.; Takacs, E.M.; Kaur, S.; Cheng, C.; Kurokawa, M. Predicting Clinical Outcomes of Cancer Patients with a P53 Deficiency Gene Signature. Sci. Rep. 2022, 12, 1317. [Google Scholar] [CrossRef]
- Lacroix, M.; Riscal, R.; Arena, G.; Linares, L.K.; Le Cam, L. Metabolic Functions of the Tumor Suppressor P53: Implications in Normal Physiology, Metabolic Disorders, and Cancer. Mol. Metab. 2020, 33, 2–22. [Google Scholar] [CrossRef]
- Cole, A.J.; Dwight, T.; Gill, A.J.; Dickson, K.A.; Zhu, Y.; Clarkson, A.; Gard, G.B.; Maidens, J.; Valmadre, S.; Clifton-Bligh, R.; et al. Assessing Mutant P53 in Primary High-Grade Serous Ovarian Cancer Using Immunohistochemistry and Massively Parallel Sequencing. Sci. Rep. 2016, 6, 26191. [Google Scholar] [CrossRef] [PubMed]
- Chien, J.; Sicotte, H.; Fan, J.B.; Humphray, S.; Cunningham, J.M.; Kalli, K.R.; Oberg, A.L.; Hart, S.N.; Li, Y.; Davila, J.I.; et al. TP53 Mutations, Tetraploidy and Homologous Recombination Repair Defects in Early Stage High-Grade Serous Ovarian Cancer. Nucleic Acids Res. 2015, 43, 6945–6958. [Google Scholar] [CrossRef] [PubMed]
- Park, E.; Han, H.; Choi, S.E.; Park, H.; Woo, H.Y.; Jang, M.; Shim, H.S.; Hwang, S.; Kang, H.; Cho, N.H. P53 Immunohistochemistry and Mutation Types Mismatching in High-Grade Serous Ovarian Cancer. Diagnostics 2022, 12, 579. [Google Scholar] [CrossRef] [PubMed]
- Duffy, M.J.; Synnott, N.C.; Crown, J. Mutant P53 as a Target for Cancer Treatment. Eur. J. Cancer 2017, 83, 258–265. [Google Scholar] [CrossRef] [PubMed]
- Guo, T.; Dong, X.; Xie, S.; Zhang, L.; Zeng, P.; Zhang, L. Cellular Mechanism of Gene Mutations and Potential Therapeutic Targets in Ovarian Cancer. Cancer Manag. Res. 2021, 13, 3081–3100. [Google Scholar] [CrossRef] [PubMed]
- Chiang, Y.T.; Chien, Y.C.; Lin, Y.H.; Wu, H.H.; Lee, D.F.; Yu, Y.L. The Function of the Mutant P53-R175h in Cancer. Cancers 2021, 13, 4088. [Google Scholar] [CrossRef]
- Klemke, L.; Fehlau, C.F.; Winkler, N.; Toboll, F.; Singh, S.K.; Moll, U.M.; Schulz-Heddergott, R. The Gain-of-Function P53 R248W Mutant Promotes Migration by STAT3 Deregulation in Human Pancreatic Cancer Cells. Front. Oncol. 2021, 11, 642603. [Google Scholar] [CrossRef]
- Li, L.; Li, X.; Tang, Y.; Lao, Z.; Lei, J.; Wei, G. Common Cancer Mutations R175H and R273H Drive the P53 DNA-Binding Domain towards Aggregation-Prone Conformations. Phys. Chem. Chem. Phys. 2020, 22, 9225–9232. [Google Scholar] [CrossRef]
- Mandilaras, V.; Garg, S.; Cabanero, M.; Tan, Q.; Pastrello, C.; Burnier, J.; Karakasis, K.; Wang, L.; Dhani, N.C.; Butler, M.O.; et al. TP53 Mutations in High Grade Serous Ovarian Cancer and Impact on Clinical Outcomes: A Comparison of next Generation Sequencing and Bioinformatics Analyses. Int. J. Gynecol. Cancer 2019, 29, 346–352. [Google Scholar] [CrossRef]
- Tuna, M.; Ju, Z.; Yoshihara, K.; Amos, C.I.; Tanyi, J.L.; Mills, G.B. Clinical Relevance of TP53 Hotspot Mutations in High-Grade Serous Ovarian Cancers. Br. J. Cancer 2020, 122, 405–412. [Google Scholar] [CrossRef]
- Eoh, K.J.; Kim, H.M.; Lee, J.Y.; Kim, S.; Kim, S.W.; Kim, Y.T.; Nam, E.J. Mutation Landscape of Germline and Somatic BRCA1/2 in Patients with High-Grade Serous Ovarian Cancer. BMC Cancer 2020, 20, 204. [Google Scholar] [CrossRef] [PubMed]
- Ji, G.; Yao, Q.; Bao, L.; Zhang, J.; Bai, Q.; Zhu, X.; Tu, X.; Bi, R.; Zhou, X. Germline and Tumor BRCA1/2 Mutations in Chinese High Grade Serous Ovarian Cancer Patients. Ann. Transl. Med. 2021, 9, 453. [Google Scholar] [CrossRef] [PubMed]
- Corona, R.I.; Seo, J.H.; Lin, X.; Hazelett, D.J.; Reddy, J.; Fonseca, M.A.S.; Abassi, F.; Lin, Y.G.; Mhawech-Fauceglia, P.Y.; Shah, S.P.; et al. Non-Coding Somatic Mutations Converge on the PAX8 Pathway in Ovarian Cancer. Nat. Commun. 2020, 11, 2020. [Google Scholar] [CrossRef]
- Ni, J.; Wang, Y.; Cheng, X.; Teng, F.; Wang, C.; Han, S.; Chen, X.; Guo, W. Pathogenic Heteroplasmic Somatic Mitochondrial DNA Mutation Confers Platinum-Resistance and Recurrence of High-Grade Serous Ovarian Cancer. Cancer Manag. Res. 2020, 12, 11085–11093. [Google Scholar] [CrossRef]
- Serio, P.A.D.M.P.; de Lima Pereira, G.F.; Katayama, M.L.H.; Roela, R.A.; Maistro, S.; Folgueira, M.A.A.K. Somatic Mutational Profile of High-Grade Serous Ovarian Carcinoma and Triple-Negative Breast Carcinoma in Young and Elderly Patients: Similarities and Divergences. Cells 2021, 10, 3586. [Google Scholar] [CrossRef]
- Nguyen, B.; Fong, C.; Luthra, A.; Smith, S.A.; DiNatale, R.G.; Nandakumar, S.; Walch, H.; Chatila, W.K.; Madupuri, R.; Kundra, R.; et al. Genomic Characterization of Metastatic Patterns from Prospective Clinical Sequencing of 25,000 Patients. Cell 2022, 185, 563–575.e11. [Google Scholar] [CrossRef] [PubMed]
- Drews, R.M.; Hernando, B.; Tarabichi, M.; Haase, K.; Lesluyes, T.; Philip, S.; Brenton, D.; Van Loo, P.; Macintyre, G.; Markowetz, F. A Pan-Cancer Compendium of Chromosomal Instability. Nature 2022, 606, 976–983. [Google Scholar] [CrossRef]
- Bronder, D.; Tighe, A.; Wangsa, D.; Zong, D.; Meyer, T.J.; Wardenaar, R.; Minshall, P.; Hirsch, D.; Heselmeyer-Haddad, K.; Nelson, L.; et al. TP53 Loss Initiates Chromosomal Instability in Fallopian Tube Epithelial Cells. Dis. Model. Mech. 2021, 14, dmm049001. [Google Scholar] [CrossRef]
- Ertych, N.; Stolz, A.; Valerius, O.; Braus, G.H.; Bastians, O. CHK2-BRCA1 Tumor-Suppressor Axis Restrains Oncogenic Aurora-A Kinase to Ensure Proper Mitotic Microtubule Assembly. Proc. Natl. Acad. Sci. USA 2016, 113, 1817–1822. [Google Scholar] [CrossRef]
- Gallo, D.; Young, J.T.F.; Fourtounis, J.; Martino, G.; Álvarez-Quilón, A.; Bernier, C.; Duffy, N.M.; Papp, R.; Roulston, A.; Stocco, R.; et al. CCNE1 Amplification Is Synthetic Lethal with PKMYT1 Kinase Inhibition. Nature 2022, 604, 749–756. [Google Scholar] [CrossRef]
- Chui, M.H.; Doodnauth, S.A.; Erdmann, N.; Tiedemann, R.E.; Sircoulomb, F.; Drapkin, R.; Shaw, P.; Rottapel, R. Chromosomal Instability and MTORC1 Activation through PTEN Loss Contribute to Proteotoxic Stress in Ovarian Carcinoma. Cancer Res. 2019, 79, 5536–5549. [Google Scholar] [CrossRef] [PubMed]
- Cortés-Ciriano, I.; Lee, J.J.K.; Xi, R.; Jain, D.; Jung, Y.L.; Yang, L.; Gordenin, D.; Klimczak, L.J.; Zhang, C.Z.; Pellman, D.S.; et al. Comprehensive Analysis of Chromothripsis in 2,658 Human Cancers Using Whole-Genome Sequencing. Nat. Genet. 2020, 52, 331–341. [Google Scholar] [CrossRef]
- Bielski, C.M.; Zehir, A.; Penson, A.V.; Donoghue, M.T.A.; Chatila, W.; Armenia, J.; Chang, M.T.; Schram, A.M.; Jonsson, P.; Bandlamudi, C.; et al. Genome Doubling Shapes the Evolution and Prognosis of Advanced Cancers. Nat. Genet. 2018, 50, 1189–1195. [Google Scholar] [CrossRef]
- Martinez-Useros, J.; Martin-Galan, M.; Florez-Cespedes, M.; Garcia-Foncillas, J. Epigenetics of Most Aggressive Solid Tumors: Pathways, Targets and Treatments. Cancers 2021, 13, 3209. [Google Scholar] [CrossRef] [PubMed]
- Matthews, B.G.; Bowden, N.A.; Wong-Brown, M.W. Epigenetic Mechanisms and Therapeutic Targets in Chemoresistant High-Grade Serous Ovarian Cancer. Cancers 2021, 13, 5993. [Google Scholar] [CrossRef] [PubMed]
- Tucker, D.W.; Getchell, C.R.; McCarthy, E.T.; Ohman, A.W.; Sasamoto, N.; Xu, S.; Ko, J.Y.; Gupta, M.; Shafrir, A.; Medina, J.E.; et al. Epigenetic Reprogramming Strategies to Reverse Global Loss of 5-Hydroxymethylcytosine, a Prognostic Factor for Poor Survival in High-Grade Serous Ovarian Cancer. Clin. Cancer Res. 2018, 24, 1389–1401. [Google Scholar] [CrossRef]
- Coughlan, A.Y.; Testa, G. Exploiting Epigenetic Dependencies in Ovarian Cancer Therapy. Int. J. Cancer 2021, 149, 1732–1743. [Google Scholar] [CrossRef]
- Yang, Y.; Bedford, M.T. Protein Arginine Methyltransferases and Cancer. Nat. Rev. Cancer 2013, 13, 37–50. [Google Scholar] [CrossRef]
- Gu, Y.; Yang, P.; Shao, Q.; Liu, X.; Xia, S.; Zhang, M.; Xu, H.; Shao, Q. Investigation of the Expression Patterns and Correlation of DNA Methyltransferases and Class I Histone Deacetylases in Ovarian Cancer Tissues. Oncol. Lett. 2013, 5, 452–458. [Google Scholar] [CrossRef]
- Slack, F.J.; Chinnaiyan, A.M. The Role of Non-Coding RNAs in Oncology. Cell 2019, 179, 1033–1055. [Google Scholar] [CrossRef] [PubMed]
- Bagnoli, M.; Canevari, S.; Califano, D.; Losito, S.; Maio, M.D.; Raspagliesi, F.; Carcangiu, M.L.; Toffoli, G.; Cecchin, E.; Sorio, R.; et al. Development and Validation of a MicroRNA-Based Signature (MiROvaR) to Predict Early Relapse or Progression of Epithelial Ovarian Cancer: A Cohort Study. Lancet Oncol. 2016, 17, 1137–1146. [Google Scholar] [CrossRef]
- Leskelä, S.; Leandro-García, L.J.; Mendiola, M.; Barriuso, J.; Inglada-Pérez, L.; Muñoz, I.; Martínez-Delgado, B.; Redondo, A.; de Santiago, J.; Robledo, M.; et al. The MiR-200 Family Controls β-Tubulin III Expression and Is Associated with Paclitaxel-Based Treatment Response and Progression-Free Survival in Ovarian Cancer Patients. Endocr. Relat. Cancer 2011, 18, 85–95. [Google Scholar] [CrossRef]
- Wang, H.; Fu, Z.; Dai, C.; Cao, J.; Liu, X.; Xu, J.; Lv, M.; Gu, Y.; Zhang, J.; Hua, X.; et al. LncRNAs Expression Profiling in Normal Ovary, Benign Ovarian Cyst and Malignant Epithelial Ovarian Cancer. Sci. Rep. 2016, 6, 38983. [Google Scholar] [CrossRef]
- Liu, P.; Fu, R.; Chen, K.; Zhang, L.; Wang, S.; Liang, W.; Zou, H.; Tao, L.; Jia, W. ETV5-Mediated Upregulation of LncRNA CTBP1-DT as a CeRNA Facilitates HGSOC Progression by Regulating MiR-188-5p/MAP3K3 Axis. Cell Death Dis. 2021, 12, 1146. [Google Scholar] [CrossRef]
- López-Reig, R.; López-Guerrero, J.A. The Hallmarks of Ovarian Cancer: Proliferation and Cell Growth. Eur. J. Cancer Suppl. 2020, 15, 27–37. [Google Scholar] [CrossRef]
- Giacinti, C.; Giordano, A. RB and Cell Cycle Progression. Oncogene 2006, 25, 5220–5227. [Google Scholar] [CrossRef]
- Wiedemeyer, W.R.; Beach, J.A.; Karlan, B.Y. Reversing Platinum Resistance in High-Grade Serous Ovarian Carcinoma: Targeting BRCA and the Homologous Recombination System. Front. Oncol. 2014, 4, 34. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Schneider-Broussard, R.; Kumar, A.P.; MacLeod, M.C.; Johnson, D.G. Regulation of BRCA1 Expression by the Rb-E2F Pathway. J. Biol. Chem. 2000, 275, 4532–4536. [Google Scholar] [CrossRef] [PubMed]
- Garsed, D.W.; Alsop, K.; Fereday, S.; Emmanuel, C.; Kennedy, C.J.; Etemadmoghadam, D.; Gao, B.; Gebski, V.; Gares, V.; Christie, E.L.; et al. Homologous Recombination DNA Repair Pathway Disruption and Retinoblastoma Protein Loss Are Associated with Exceptional Survival in High-Grade Serous Ovarian Cancer. Clin. Cancer Res. 2018, 24, 569–580. [Google Scholar] [CrossRef]
- Ediriweera, M.K.; Tennekoon, K.H.; Samarakoon, S.R. Role of the PI3K/AKT/MTOR Signaling Pathway in Ovarian Cancer: Biological and Therapeutic Significance. Semin. Cancer Biol. 2019, 59, 147–160. [Google Scholar] [CrossRef] [PubMed]
- Rinne, N.; Christie, E.L.; Ardasheva, A.; Kwok, C.H.; Demchenko, N.; Low, C.; Tralau-Stewart, C.; Fotopoulou, C.; Cunnea, P. Targeting the PI3K/AKT/MTOR Pathway in Epithelial Ovarian Cancer, Therapeutic Treatment Options for Platinum-Resistant Ovarian Cancer. Cancer Drug Resist. 2021, 4, 573–595. [Google Scholar] [CrossRef]
- van der Ploeg, P.; Uittenboogaard, A.; Thijs, A.M.J.; Westgeest, H.M.; Boere, I.A.; Lambrechts, S.; van de Stolpe, A.; Bekkers, R.L.M.; Piek, J.M.J. The Effectiveness of Monotherapy with PI3K/AKT/MTOR Pathway Inhibitors in Ovarian Cancer: A Meta-Analysis. Gynecol. Oncol. 2021, 163, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.T.; Lampert, E.J.; Coots, C.; Lee, J.M. Targeting the PI3K Pathway and DNA Damage Response as a Therapeutic Strategy in Ovarian Cancer. Cancer Treat. Rev. 2020, 86, 102021. [Google Scholar] [CrossRef] [PubMed]
- Rendell, A.; Thomas-Bland, I.; McCuish, L.; Taylor, C.; Binju, M.; Yu, Y. Targeting Tyrosine Kinases in Ovarian Cancer: Small Molecule Inhibitor and Monoclonal Antibody, Where Are We Now? Biomedicines 2022, 10, 2113. [Google Scholar] [CrossRef]
- Klempner, S.J.; Myers, A.P.; Mills, G.B.; Westin, S.N. Clinical Investigation of Receptor and Non-Receptor Tyrosine Kinase Inhibitors for the Treatment of Epithelial Ovarian Cancer. Expert Opin. Pharmacother. 2013, 14, 2171–2182. [Google Scholar] [CrossRef]
- Knudsen, E.S.; Wang, J.Y.J. Targeting the RB-Pathway in Cancer Therapy. Clin. Cancer Res. 2010, 16, 1094–1099. [Google Scholar] [CrossRef] [PubMed]
- Nakai, H.; Matsumura, N. The Roles and Limitations of Bevacizumab in the Treatment of Ovarian Cancer. Int. J. Clin. Oncol. 2022, 27, 1120–1126. [Google Scholar] [CrossRef]
- Burger, R.A.; Brady, M.F.; Bookman, M.A.; Fleming, G.F.; Monk, B.J.; Huang, H.; Mannel, R.S.; Homesley, H.D.; Fowler, J.; Greer, B.E.; et al. Incorporation of Bevacizumab in the Primary Treatment of Ovarian Cancer. N. Engl. J. Med. 2011, 365, 2473–2483. [Google Scholar] [CrossRef]
- Liu, C.; Barger, C.J.; Karpf, A.R. Foxm1: A Multifunctional Oncoprotein and Emerging Therapeutic Target in Ovarian Cancer. Cancers 2021, 13, 3065. [Google Scholar] [CrossRef]
- Oturkar, C.C.; Gandhi, N.; Rao, P.; Eng, K.H.; Miller, A.; Singh, P.K.; Zsiros, E.; Odunsi, K.O.; Das, G.M. Estrogen Receptor-Beta2 (ERβ2)–Mutant P53–FOXM1 Axis: A Novel Driver of Proliferation, Chemoresistance, and Disease Progression in High Grade Serous Ovarian Cancer (HGSOC). Cancers 2022, 14, 1120. [Google Scholar] [CrossRef]
- Chiaramonte, R.; Colombo, M.; Bulfamante, G.; Falleni, M.; Tosi, D.; Garavelli, S.; de Simone, D.; Vigolo, E.; Todoerti, K.; Neri, A.; et al. Notch Pathway Promotes Ovarian Cancer Growth and Migration via CXCR4/SDF1α Chemokine System. Int. J. Biochem. Cell Biol. 2015, 66, 134–140. [Google Scholar] [CrossRef]
- Huang, C.C.; Cheng, S.H.; Wu, C.H.; Li, W.Y.; Wang, J.S.; Kung, M.L.; Chu, T.H.; Huang, S.T.; Feng, C.T.; Huang, S.C.; et al. Delta-like 1 Homologue Promotes Tumorigenesis and Epithelial-Mesenchymal Transition of Ovarian High-Grade Serous Carcinoma through Activation of Notch Signaling. Oncogene 2019, 38, 3201–3215. [Google Scholar] [CrossRef] [PubMed]
- de Felice, F.; Vertechy, L.; Giudice, E.; Ergasti, R.; Boccia, S.; Fagotti, A.; Scambia, G.; Marchetti, C. Evolution of Clinical Trials in Ovarian Cancer Management over the Past 20 Years: Never Settle Down, Always Go Beyond. J. Oncol. 2021, 2021, 1682532. [Google Scholar] [CrossRef]
- Home—ClinicalTrials.Gov. Available online: https://clinicaltrials.gov/ct2/home (accessed on 11 August 2022).
- Lawton, F.G.; Pavlik, E.J. Perspectives on Ovarian Cancer 1809 to 2022 and Beyond. Diagnostics 2022, 12, 791. [Google Scholar] [CrossRef]
- Menon, U.; Karpinskyj, C.; Gentry-Maharaj, A. Ovarian Cancer Prevention and Screening. Obstet. Gynecol. 2018, 131, 909–927. [Google Scholar] [CrossRef] [PubMed]
- Stewart, C.; Ralyea, C.; Lockwood, S. Ovarian Cancer: An Integrated Review. Semin. Oncol. Nurs. 2019, 35, 151–156. [Google Scholar] [CrossRef]
- Zhu, J.W.; Charkhchi, P.; Akbari, M.R. Potential Clinical Utility of Liquid Biopsies in Ovarian Cancer. Mol. Cancer 2022, 21, 114. [Google Scholar] [CrossRef]
- Heitzer, E.; Haque, I.S.; Roberts, C.E.S.; Speicher, M.R. Current and Future Perspectives of Liquid Biopsies in Genomics-Driven Oncology. Nat. Rev. Genet. 2019, 20, 71–88. [Google Scholar] [CrossRef]
- Yang, J.; Ma, J.; Jin, Y.; Cheng, S.; Huang, S.; Zhang, N.; Wang, Y. Development and Validation for Prognostic Nomogram of Epithelial Ovarian Cancer Recurrence Based on Circulating Tumor Cells and Epithelial–Mesenchymal Transition. Sci. Rep. 2021, 11, 6540. [Google Scholar] [CrossRef]
- Zhang, X.; Li, H.; Yu, X.; Li, S.; Lei, Z.; Li, C.; Zhang, Q.; Han, Q.; Li, Y.; Zhang, K.; et al. Analysis of Circulating Tumor Cells in Ovarian Cancer and Their Clinical Value as a Biomarker. Cell. Physiol. Biochem. 2018, 48, 1983–1994. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Cheng, S.; Zhang, N.; Jin, Y.; Wang, Y. Liquid Biopsy for Ovarian Cancer Using Circulating Tumor Cells: Recent Advances on the Path to Precision Medicine. Biochim. Biophys. Acta Rev. Cancer 2022, 1877, 188660. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.-X.; Hong Neoh, K.; Chang, X.-H.; Sun, Y.; Cheng, H.-Y.; Ye, X.; Ma, R.-Q.; Han, R.P.; Cui, H. Diagnostic Value of HE4+ Circulating Tumor Cells in Patients with Suspicious Ovarian Cancer. Oncotarget 2018, 9, 7522–7533. [Google Scholar] [CrossRef]
- Pearl, M.L.; Zhao, Q.; Yang, J.; Dong, H.; Tulley, S.; Zhang, Q.; Golightly, M.; Zucker, S.; Chen, W.T. Prognostic Analysis of Invasive Circulating Tumor Cells (ICTCs) in Epithelial Ovarian Cancer. Gynecol. Oncol. 2014, 134, 581–590. [Google Scholar] [CrossRef] [PubMed]
- Lou, E.; Vogel, R.I.; Teoh, D.; Hoostal, S.; Grad, A.; Gerber, M.; Monu, M.; Lukaszewski, T.; Deshpande, J.; Linden, M.A.; et al. Assessment of Circulating Tumor Cells as a Predictive Biomarker of Histology in Women with Suspected Ovarian Cancer. Lab. Med. 2018, 49, 134–139. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Yang, J.; Jin, Y.; Cheng, S.; Huang, S.; Zhang, N.; Wang, Y. Artificial Intelligence Based on Blood Biomarkers Including Ctcs Predicts Outcomes in Epithelial Ovarian Cancer: A Prospective Study. OncoTargets Ther. 2021, 14, 3267–3280. [Google Scholar] [CrossRef] [PubMed]
- Schuster, E.; Taftaf, R.; Reduzzi, C.; Albert, M.K.; Romero-Calvo, I.; Liu, H. Better Together: Circulating Tumor Cell Clustering in Metastatic Cancer. Trends Cancer 2021, 7, 1020–1032. [Google Scholar] [CrossRef]
- Zhou, Y.; Bian, B.; Yuan, X.; Xie, G.; Ma, Y.; Shen, L. Prognostic Value of Circulating Tumor Cells in Ovarian Cancer: A Meta-Analysis. PLoS ONE 2015, 10, e130873. [Google Scholar] [CrossRef] [PubMed]
- Trinidad, C.V.; Tetlow, A.L.; Bantis, L.E.; Godwin, A.K. Reducing Ovarian Cancer Mortality through Early Detection: Approaches Using Circulating Biomarkers. Cancer Prev. Res. 2020, 13, 241–252. [Google Scholar] [CrossRef]
- Keller, L.; Belloum, Y.; Wikman, H.; Pantel, K. Clinical Relevance of Blood-Based CtDNA Analysis: Mutation Detection and Beyond. Br. J. Cancer 2021, 124, 345–358. [Google Scholar] [CrossRef]
- Barbosa, A.; Peixoto, A.; Pinto, P.; Pinheiro, M.; Teixeira, M.R. Potential Clinical Applications of Circulating Cell-Free DNA in Ovarian Cancer Patients. Expert Rev. Mol. Med. 2018, 20, e6. [Google Scholar] [CrossRef]
- Mari, R.; Mamessier, E.; Lambaudie, E.; Provansal, M.; Birnbaum, D.; Bertucci, F.; Sabatier, R. Liquid Biopsies for Ovarian Carcinoma: How Blood Tests May Improve the Clinical Management of a Deadly Disease. Cancers 2019, 11, 774. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Pu, K.; Ge, L.; Wu, X. Diagnostic Significance Assessment of the Circulating Cell-Free DNA in Ovarian Cancer: An Updated Meta-Analysis. Gene 2019, 714, 143993. [Google Scholar] [CrossRef] [PubMed]
- Zill, O.A.; Banks, K.C.; Fairclough, S.R.; Mortimer, S.A.; Vowles, J.V.; Mokhtari, R.; Gandara, D.R.; Mack, P.C.; Odegaard, J.I.; Nagy, R.J.; et al. The Landscape of Actionable Genomic Alterations in Cell-Free Circulating Tumor DNA from 21,807 Advanced Cancer Patients. Clin. Cancer Res. 2018, 24, 3528–3538. [Google Scholar] [CrossRef]
- Parkinson, C.A.; Gale, D.; Piskorz, A.M.; Biggs, H.; Hodgkin, C.; Addley, H.; Freeman, S.; Moyle, P.; Sala, E.; Sayal, K.; et al. Exploratory Analysis of TP53 Mutations in Circulating Tumour DNA as Biomarkers of Treatment Response for Patients with Relapsed High-Grade Serous Ovarian Carcinoma: A Retrospective Study. PLoS Med. 2016, 13, e1002198. [Google Scholar] [CrossRef] [PubMed]
- Bettegowda, C.; Sausen, M.; Leary, R.J.; Kinde, I.; Wang, Y.; Agrawal, N.; Bartlett, B.R.; Wang, H.; Luber, B.; Alani, R.M.; et al. Detection of Circulating Tumor DNA in Early-and Late-Stage Human Malignancies. Sci. Transl. Med. 2014, 6, 224ra24. [Google Scholar] [CrossRef]
- Elias, K.M.; Guo, J.; Bast, R.C. Early Detection of Ovarian Cancer. Hematol. Oncol. Clin. N. Am. 2018, 32, 903–914. [Google Scholar] [CrossRef]
- Cristiano, S.; Leal, A.; Phallen, J.; Fiksel, J.; Adleff, V.; Bruhm, D.C.; Jensen, S.Ø.; Medina, J.E.; Hruban, C.; White, J.R.; et al. Genome-Wide Cell-Free DNA Fragmentation in Patients with Cancer. Nature 2019, 570, 385–389. [Google Scholar] [CrossRef]
- Wang, B.; Yu, L.; Luo, X.; Huang, L.; Li, Q.S.; Shao, X.S.; Liu, Y.; Fan, Y.; Yang, G.Z. Detection of OPCML Methylation, a Possible Epigenetic Marker, from Free Serum Circulating DNA to Improve the Diagnosis of Early-Stage Ovarian Epithelial Cancer. Oncol. Lett. 2017, 14, 217–223. [Google Scholar] [CrossRef]
- Zhang, Q.; Hu, G.; Yang, Q.; Dong, R.; Xie, X.; Ma, D.; Shen, K.; Kong, B. A Multiplex Methylation-Specific PCR Assay for the Detection of Early-Stage Ovarian Cancer Using Cell-Free Serum DNA. Gynecol. Oncol. 2013, 130, 132–139. [Google Scholar] [CrossRef]
- Giannopoulou, L.; Chebouti, I.; Pavlakis, K.; Kasimir-Bauer, S.; Lianidou, E.S. RASSF1A Promoter Methylation in High-Grade Serous Ovarian Cancer: A Direct Comparison Study in Primary Tumors, Adjacent Morphologically Tumor Cell-Free Tissues and Paired Circulating Tumor DNA. Oncotarget 2017, 8, 21429–21443. [Google Scholar] [CrossRef]
- Widschwendter, M.; Zikan, M.; Wahl, B.; Lempiäinen, H.; Paprotka, T.; Evans, I.; Jones, A.; Ghazali, S.; Reisel, D.; Eichner, J.; et al. The Potential of Circulating Tumor DNA Methylation Analysis for the Early Detection and Management of Ovarian Cancer. Genome Med. 2017, 9, 116. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Gupta, S.; Badarukhiya, J.A.; Sachan, M. Detection of Aberrant Methylation of HOXA9 and HIC1 through Multiplex MethyLight Assay in Serum DNA for the Early Detection of Epithelial Ovarian Cancer. Int. J. Cancer 2020, 147, 1740–1752. [Google Scholar] [CrossRef]
- Asante, D.B.; Calapre, L.; Ziman, M.; Meniawy, T.M.; Gray, E.S. Liquid Biopsy in Ovarian Cancer Using Circulating Tumor DNA and Cells: Ready for Prime Time? Cancer Lett. 2020, 468, 59–71. [Google Scholar] [CrossRef]
- Barbosa, A.; Pinto, P.; Peixoto, A.; Guerra, J.; Pinheiro, M.; Santos, C.; Pinto, C.; Escudeiro, C.; Bartosch, C.; Santos, R.; et al. Next Generation Sequencing of Tumor and Matched Plasma Samples: Identification of Somatic Variants in CtDNA From Ovarian Cancer Patients. Front. Oncol. 2021, 11, 754094. [Google Scholar] [CrossRef]
- Phallen, J.; Sausen, M.; Adleff, V.; Leal, A.; Hruban, C.; White, J.; Anagnostou, V.; Fiksel, J.; Cristiano, S.; Papp, E.; et al. Direct Detection of Early-Stage Cancers Using Circulating Tumor DNA. Sci. Transl. Med. 2017, 9, eaan2415. [Google Scholar] [CrossRef]
- Zhou, Q.; Li, W.; Leng, B.; Zheng, W.; He, Z.; Zuo, M.; Chen, A. Circulating Cell Free DNA as the Diagnostic Marker for Ovarian Cancer: A Systematic Review and Meta-Analysis. PLoS ONE 2016, 11, e155495. [Google Scholar] [CrossRef] [PubMed]
- Thusgaard, C.F.; Korsholm, M.; Koldby, K.M.; Kruse, T.A.; Thomassen, M.; Jochumsen, K.M. Epithelial Ovarian Cancer and the Use of Circulating Tumor DNA: A Systematic Review. Gynecol. Oncol. 2021, 161, 884–895. [Google Scholar] [CrossRef] [PubMed]
- Cohen, J.D.; Li, L.; Wang, Y.; Thoburn, C.; Afsari, B.; Danilova, L.; Douville, C.; Javed, A.A.; Wong, F.; Mattox, A.; et al. Detection and Localization of Surgically Resectable Cancers with a Multi-Analyte Blood Test. Science 2018, 359, 926–930. [Google Scholar] [CrossRef] [PubMed]
- Mathios, D.; Johansen, J.S.; Cristiano, S.; Medina, J.E.; Phallen, J.; Larsen, K.R.; Bruhm, D.C.; Niknafs, N.; Ferreira, L.; Adleff, V.; et al. Detection and Characterization of Lung Cancer Using Cell-Free DNA Fragmentomes. Nat. Commun. 2021, 12, 5060. [Google Scholar] [CrossRef]
- Drag, M.H.; Kilpeläinen, T.O. Cell-Free Dna and Rna—Measurement and Applications in Clinical Diagnostics with Focus on Metabolic Disorders. Physiol Genom. 2021, 53, 33–46. [Google Scholar] [CrossRef]
- Joosse, S.A.; Müller, V.; Steinbach, B.; Pantel, K.; Schwarzenbach, H. Circulating Cell-Free Cancer-Testis MAGE-A RNA, BORIS RNA, Let-7b and MiR-202 in the Blood of Patients with Breast Cancer and Benign Breast Diseases. Br. J. Cancer 2014, 111, 909–917. [Google Scholar] [CrossRef] [PubMed]
- Larson, M.H.; Pan, W.; Kim, H.J.; Mauntz, R.E.; Stuart, S.M.; Pimentel, M.; Zhou, Y.; Knudsgaard, P.; Demas, V.; Aravanis, A.M.; et al. A Comprehensive Characterization of the Cell-Free Transcriptome Reveals Tissue- and Subtype-Specific Biomarkers for Cancer Detection. Nat. Commun. 2021, 12, 2357. [Google Scholar] [CrossRef] [PubMed]
- Yokoi, A.; Yoshioka, Y.; Hirakawa, A.; Yamamoto, Y.; Ishikawa, M.; Ikeda, S.-I.; Kato, T.; Niimi, K.; Kajiyama, H.; Kikkawa, F.; et al. A Combination of Circulating MiRNAs for the Early Detection of Ovarian Cancer. Oncotarget 2017, 8, 89811–89823. [Google Scholar] [CrossRef]
- Alshamrani, A.A. Roles of MicroRNAs in Ovarian Cancer Tumorigenesis: Two Decades Later, What Have We Learned? Front. Oncol. 2020, 10, 1084. [Google Scholar] [CrossRef] [PubMed]
- Todeschini, P.; Salviato, E.; Paracchini, L.; Ferracin, M.; Petrillo, M.; Zanotti, L.; Tognon, G.; Gambino, A.; Calura, E.; Caratti, G.; et al. Circulating MiRNA Landscape Identifies MiR-1246 as Promising Diagnostic Biomarker in High-Grade Serous Ovarian Carcinoma: A Validation across Two Independent Cohorts. Cancer Lett. 2017, 388, 320–327. [Google Scholar] [CrossRef]
- Gong, J.; Xu, X.; Zhang, X.; Zhou, Y. LncRNA MIR4435-2HG Is a Potential Early Diagnostic Marker for Ovarian Carcinoma. Acta Biochim. Biophys. Sin. 2019, 51, 953–959. [Google Scholar] [CrossRef]
- Cui, Y.; Shen, G.; Zhou, D.; Wu, F. CASC11 Overexpression Predicts Poor Prognosis and Regulates Cell Proliferation and Apoptosis in Ovarian Carcinoma. Cancer Manag. Res. 2020, 12, 523–529. [Google Scholar] [CrossRef]
- Hulstaert, E.; Morlion, A.; Levanon, K.; Vandesompele, J.; Mestdagh, P. Candidate RNA Biomarkers in Biofluids for Early Diagnosis of Ovarian Cancer: A Systematic Review. Gynecol. Oncol. 2021, 160, 633–642. [Google Scholar] [CrossRef]
- Ye, S.-B.; Li, Z.-L.; Luo, D.-H.; Huang, B.-J.; Chen, Y.-S.; Zhang, X.-S.; Cui, J.; Zeng, Y.-X.; Li, J. Tumor-Derived Exosomes Promote Tumor Progression and T-Cell Dysfunction through the Regulation of Enriched Exosomal MicroRNAs in Human Nasopharyngeal Carcinoma. Oncotarget 2014, 5, 5439. [Google Scholar] [CrossRef]
- Feng, W.; Dean, D.C.; Hornicek, F.J.; Shi, H.; Duan, Z. Exosomes Promote Pre-Metastatic Niche Formation in Ovarian Cancer. Mol. Cancer 2019, 18, 124. [Google Scholar] [CrossRef] [PubMed]
- Au Yeung, C.L.; Co, N.N.; Tsuruga, T.; Yeung, T.L.; Kwan, S.Y.; Leung, C.S.; Li, Y.; Lu, E.S.; Kwan, K.; Wong, K.K.; et al. Exosomal Transfer of Stroma-Derived MiR21 Confers Paclitaxel Resistance in Ovarian Cancer Cells through Targeting APAF1. Nat. Commun. 2016, 7, 11150. [Google Scholar] [CrossRef]
- Croft, P.K.D.; Sharma, S.; Godbole, N.; Rice, G.E.; Salomon, C. Ovarian-Cancer-Associated Extracellular Vesicles: Microenvironmental Regulation and Potential Clinical Applications. Cells 2021, 10, 2272. [Google Scholar] [CrossRef]
- Taylor, D.D.; Gercel-Taylor, C. MicroRNA Signatures of Tumor-Derived Exosomes as Diagnostic Biomarkers of Ovarian Cancer. Gynecol. Oncol. 2008, 110, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Choi, M.C.; Jeong, J.Y.; Hwang, S.; Jung, S.G.; Joo, W.D.; Park, H.; Song, S.H.; Lee, C.; Kim, T.H.; et al. Serum Exosomal MiRNA-145 and MiRNA-200c as Promising Biomarkers for Preoperative Diagnosis of Ovarian Carcinomas. J. Cancer 2019, 10, 1958–1967. [Google Scholar] [CrossRef] [PubMed]
- Maeda, K.; Sasaki, H.; Ueda, S.; Miyamoto, S.; Terada, S.; Konishi, H.; Kogata, Y.; Ashihara, K.; Fujiwara, S.; Tanaka, Y.; et al. Serum Exosomal MicroRNA-34a as a Potential Biomarker in Epithelial Ovarian Cancer. J. Ovarian Res. 2020, 13, 47. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Ou, X.; Wu, X. Proteomics Profiling of Plasma Exosomes in Epithelial Ovarian Cancer: A Potential Role in the Coagulation Cascade, Diagnosis and Prognosis. Int. J. Oncol. 2019, 54, 1719–1733. [Google Scholar] [CrossRef]
- Hinestrosa, J.P.; Kurzrock, R.; Lewis, J.M.; Schork, N.J.; Schroeder, G.; Kamat, A.M.; Lowy, A.M.; Eskander, R.N.; Perrera, O.; Searson, D.; et al. Early-Stage Multi-Cancer Detection Using an Extracellular Vesicle Protein-Based Blood Test. Commun. Med. 2022, 2, 29. [Google Scholar] [CrossRef]
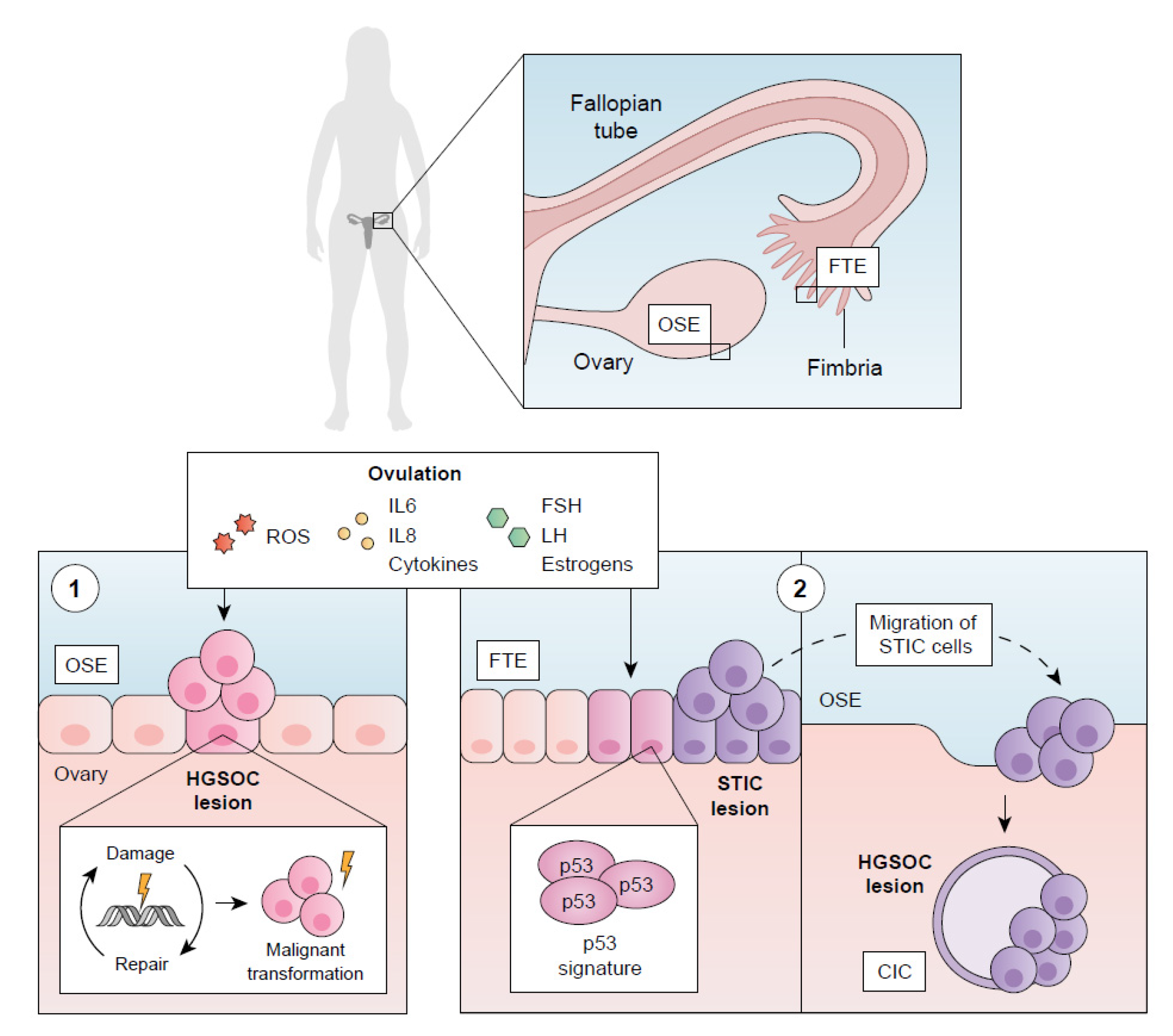
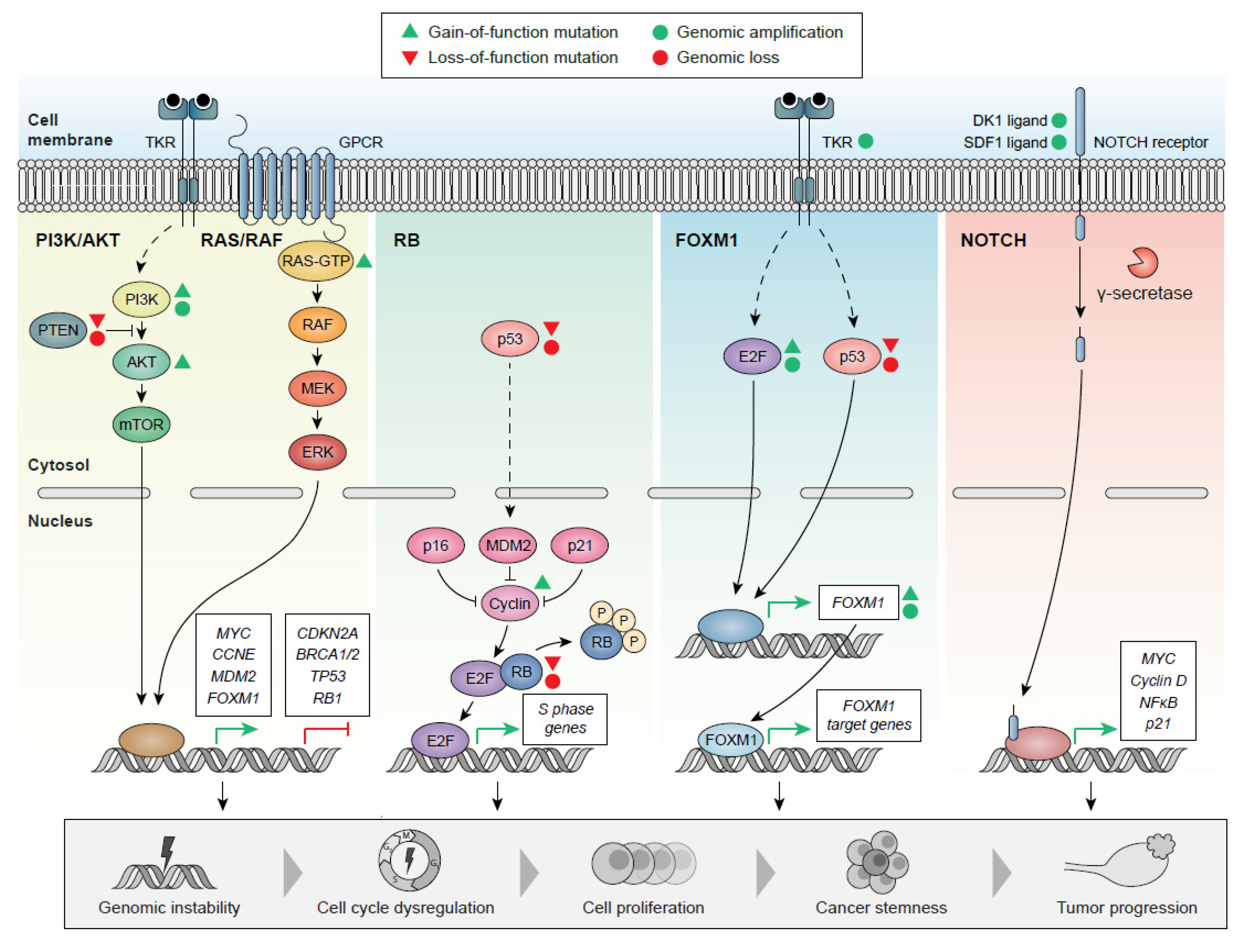
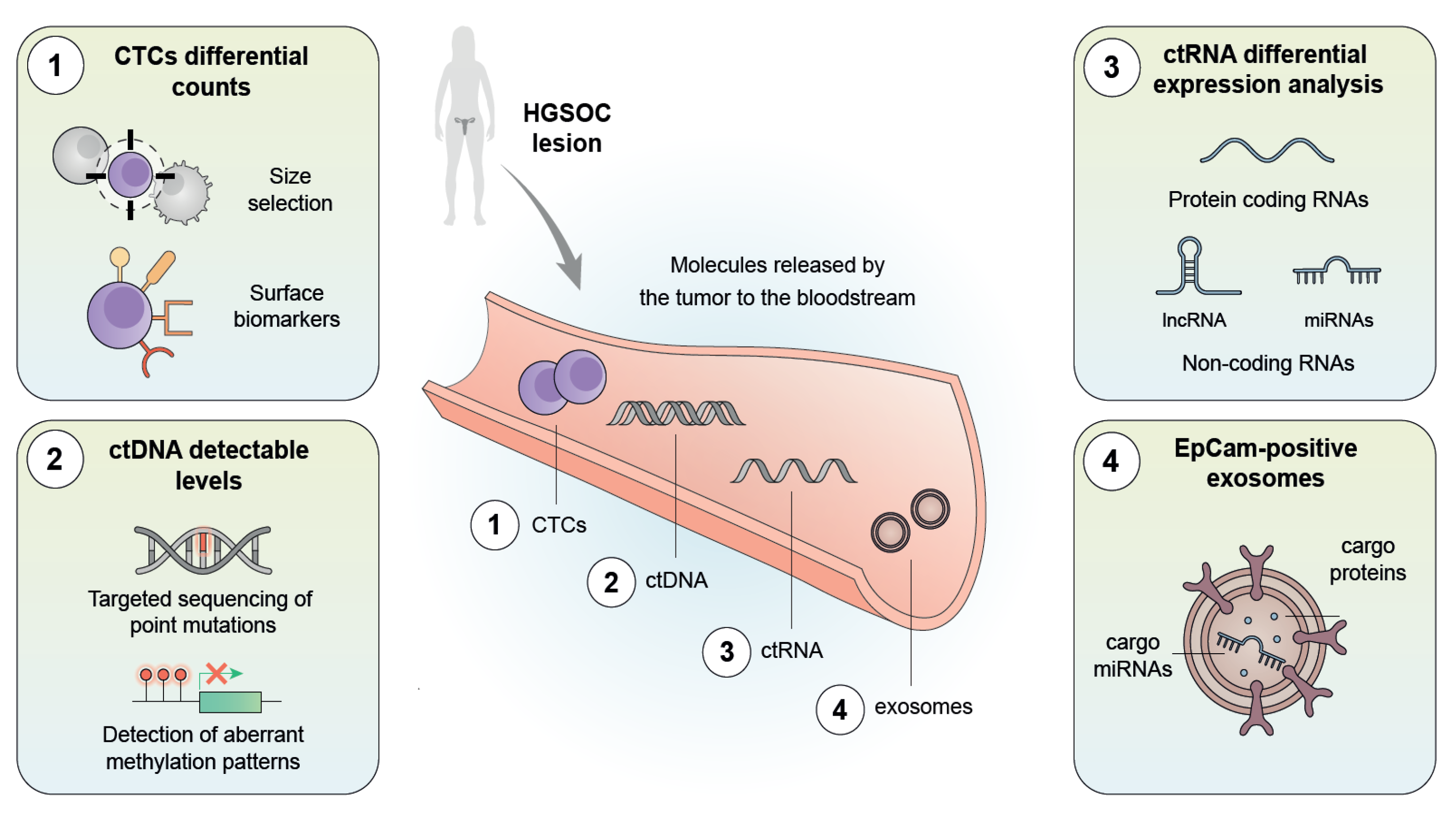
| Inherited Mutations | Mutations | Susceptible Loci | eQTL Associations | [82,166,167,168,169] | |||
| BRCA1/2, CHEK2, RAD51, MRE11A, BRIP1, RAD50, PALB2, ATM | 3q28, 10q24.33, 4q32.3, 18q11.2, 8q21.11, 22q12 | HOXD9, STRCP1, CDC42, CPNE1, CDA8, HEATR3, CCNE1 | |||||
| Somatic Mutations | Universal Mutation | Other Mutations | [82,91,170,171] | ||||
| TP53, in almost all cases | PTEN, KRAS, FAT3, CSDM, MLH3, RB1 | ||||||
| Chromosomal Aberrations | Chromosomal Gains | Chromosomal Losses | Focal Amplification Areas | Focal Deletion Areas | CLP | [82,172,173,174] | |
| Chr 1, 3, 7, 8, 12, 20 | Chr 5, 6, 11, 16, 17, 18, 19 | CNNE1, MYC, MECOM | PTEN, RB1, NF1 | MLF1, ARID1A, BRCA1, MYC, CCNE1, PIK3CA, TP53 | |||
| Epigenetic Alterations | DNA methylation | Histone Modifications | miRNAs | lncRNAs | [175,176,177,178,179,180,181,182] | ||
| Hypermethylation: ↓ transcription BRCA1 promoter | Methyltransferase overexpression (CARM1) | miR-1290, miR27-a-3, miR23a, miR200 family, miR025 | CTBP1-DT | ||||
| Hypomethylation: ↑ oncogene transcription | Histone deacetylase overexpression | ||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Punzón-Jiménez, P.; Lago, V.; Domingo, S.; Simón, C.; Mas, A. Molecular Management of High-Grade Serous Ovarian Carcinoma. Int. J. Mol. Sci. 2022, 23, 13777. https://doi.org/10.3390/ijms232213777
Punzón-Jiménez P, Lago V, Domingo S, Simón C, Mas A. Molecular Management of High-Grade Serous Ovarian Carcinoma. International Journal of Molecular Sciences. 2022; 23(22):13777. https://doi.org/10.3390/ijms232213777
Chicago/Turabian StylePunzón-Jiménez, Paula, Victor Lago, Santiago Domingo, Carlos Simón, and Aymara Mas. 2022. "Molecular Management of High-Grade Serous Ovarian Carcinoma" International Journal of Molecular Sciences 23, no. 22: 13777. https://doi.org/10.3390/ijms232213777
APA StylePunzón-Jiménez, P., Lago, V., Domingo, S., Simón, C., & Mas, A. (2022). Molecular Management of High-Grade Serous Ovarian Carcinoma. International Journal of Molecular Sciences, 23(22), 13777. https://doi.org/10.3390/ijms232213777







