Integrated Metabolome and Transcriptome Analysis Reveals a Regulatory Network of Fruit Peel Pigmentation in Eggplant (Solanum melongena L.)
Abstract
:1. Introduction
2. Results
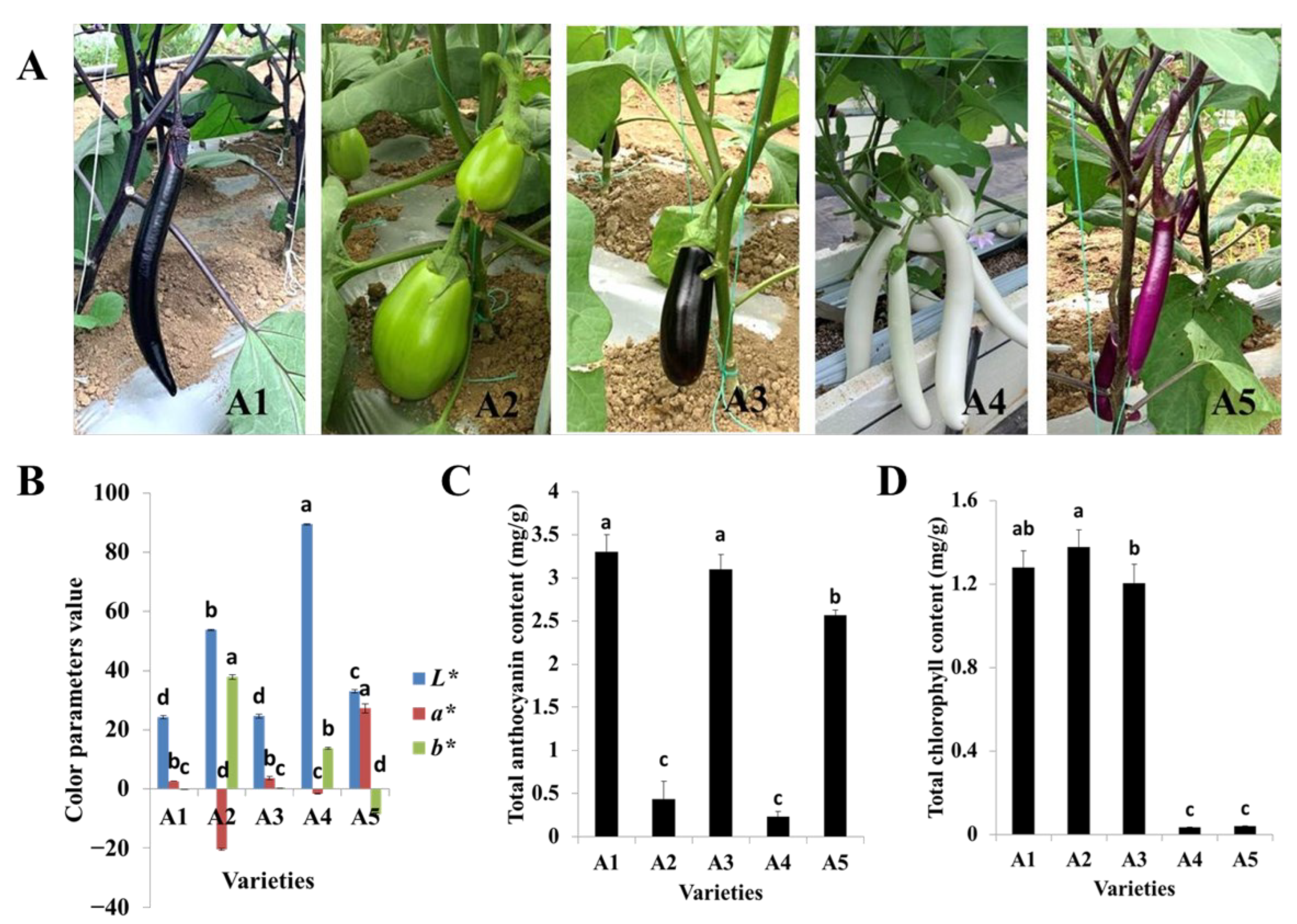
2.1. Variation in Fruit Peel Color and Total Anthocyanin and Chlorophyll Contents
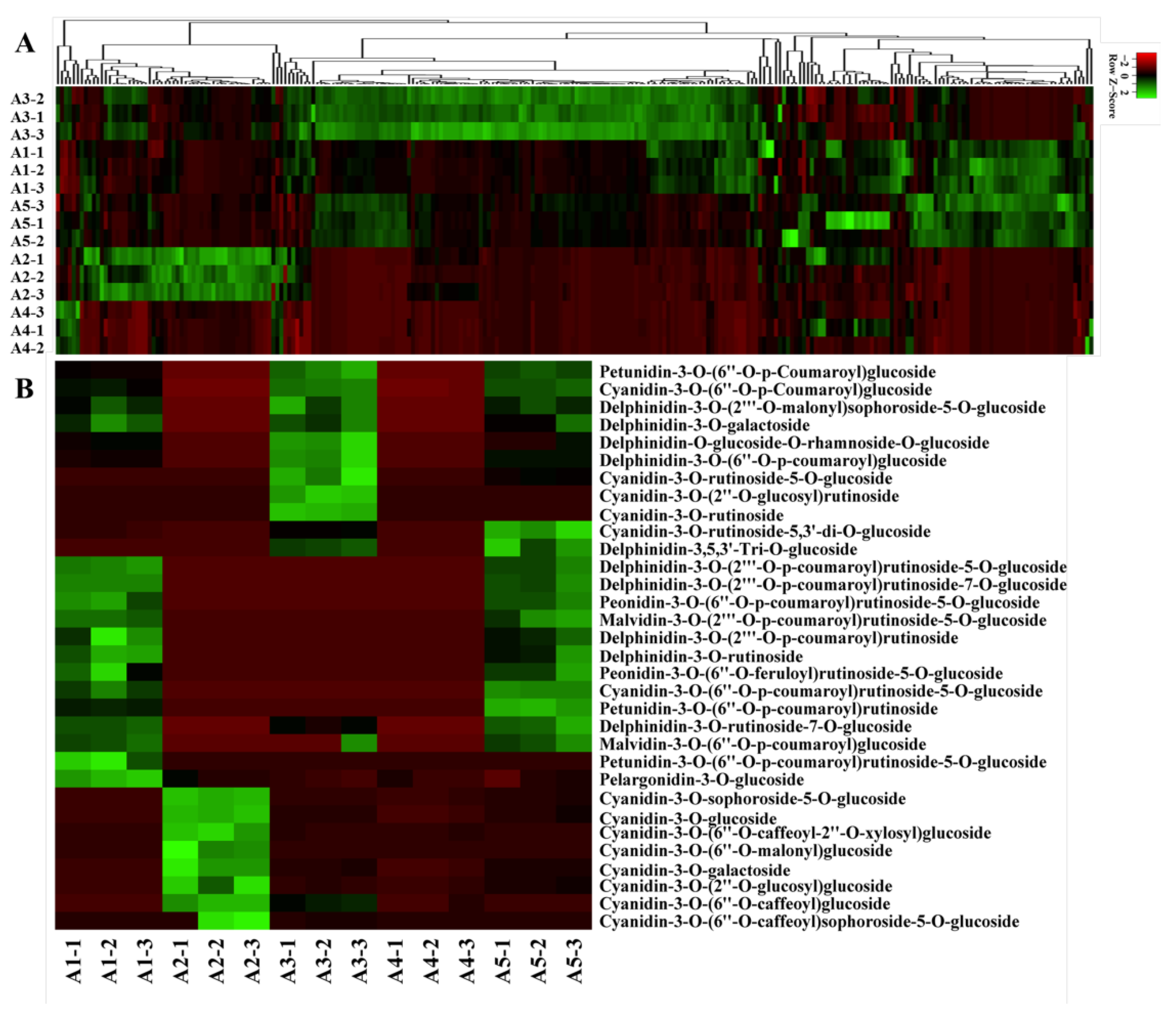
2.2. Flavonoid Metabolic Differences
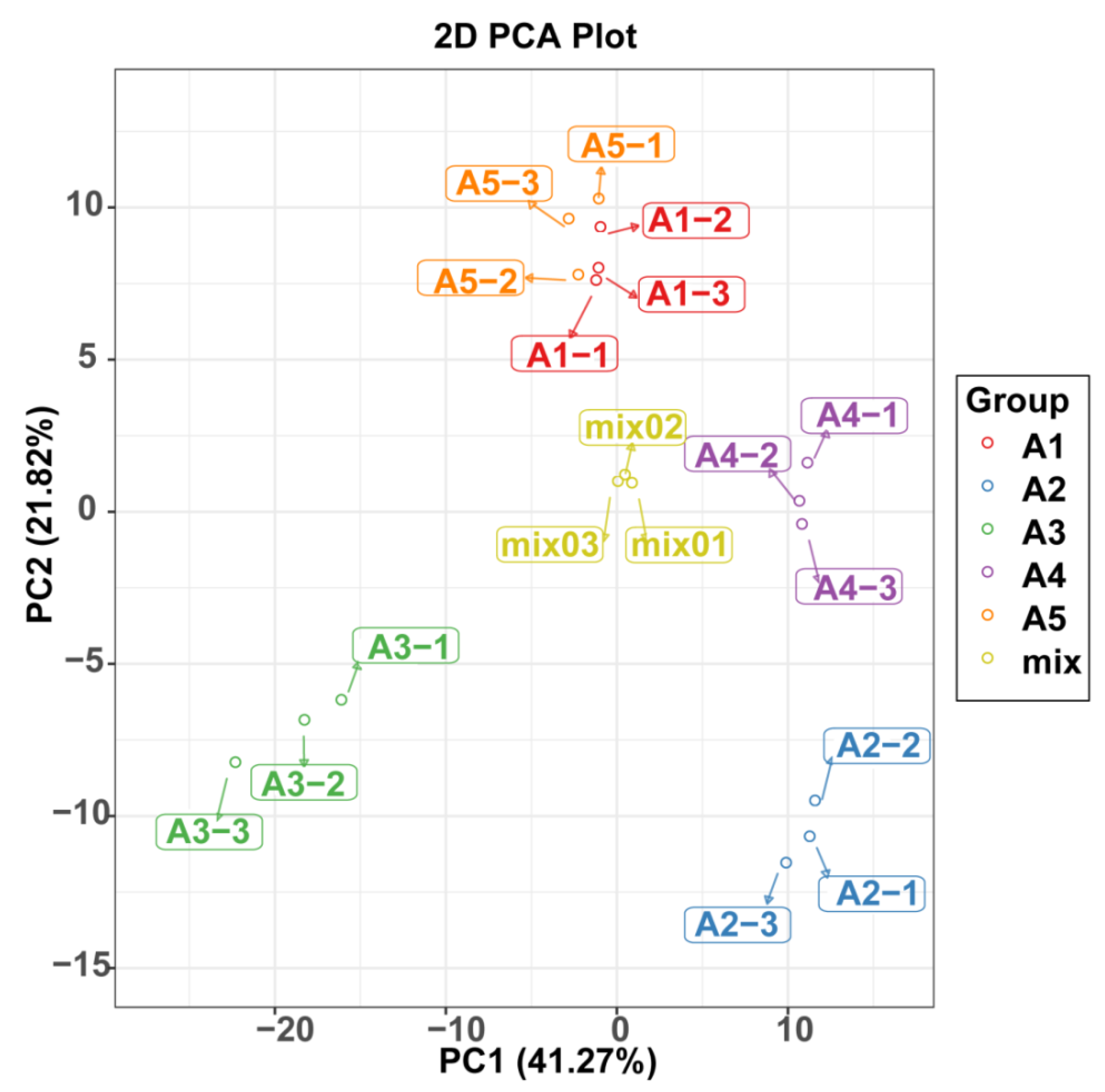
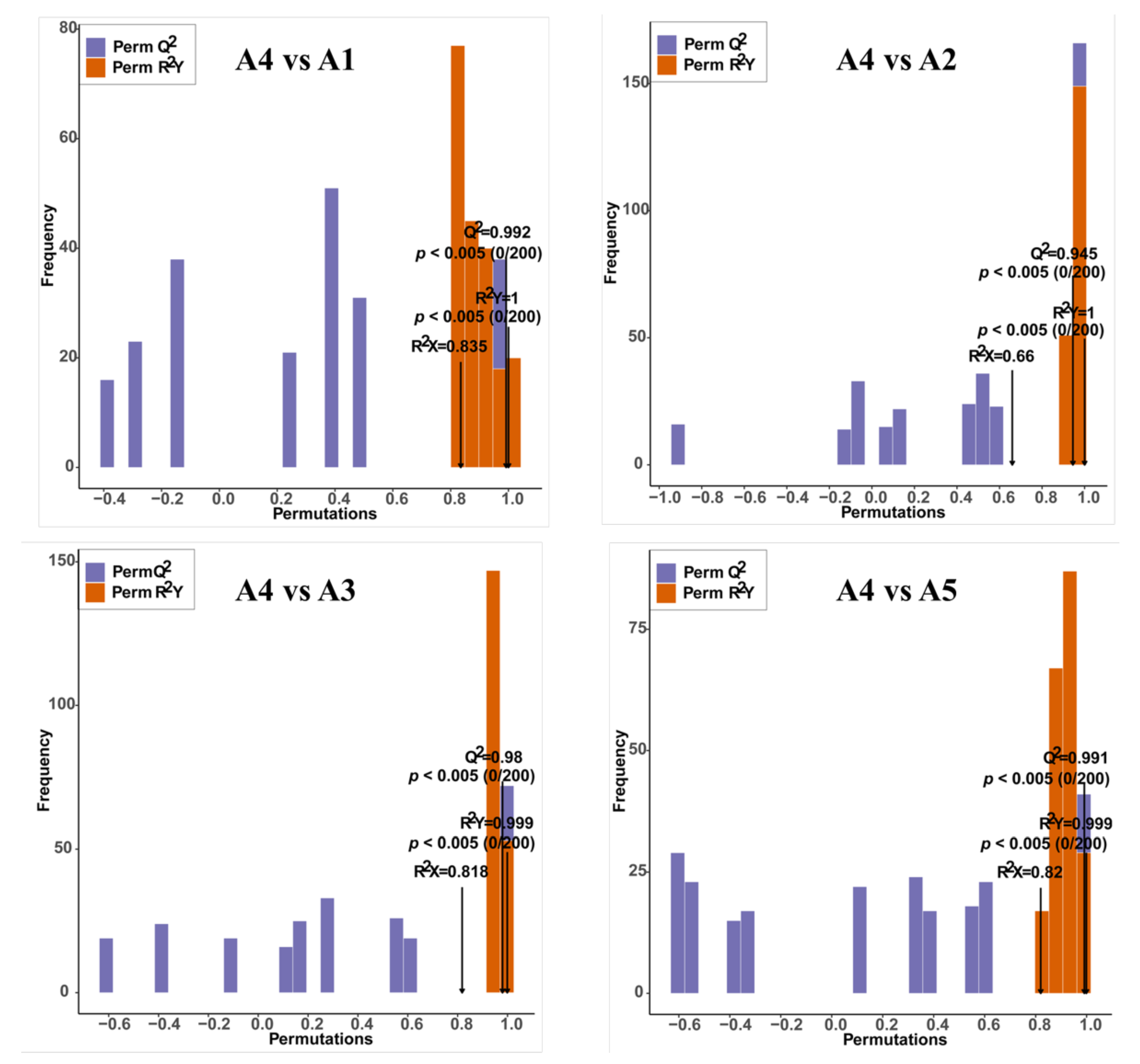
2.3. Comparison of Differentially Accumulated Metabolites among the Five Eggplant Varieties
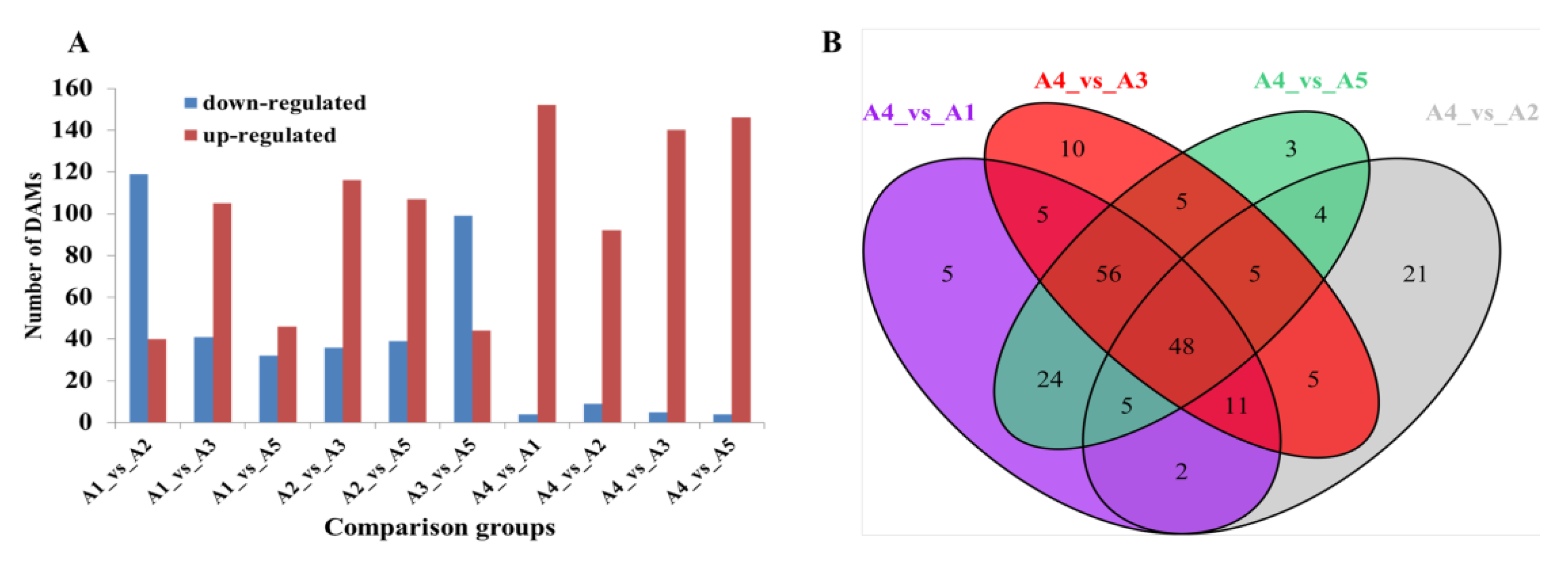
2.4. Comparative Transcriptome of Fruit Peels among the Five Eggplant Varieties
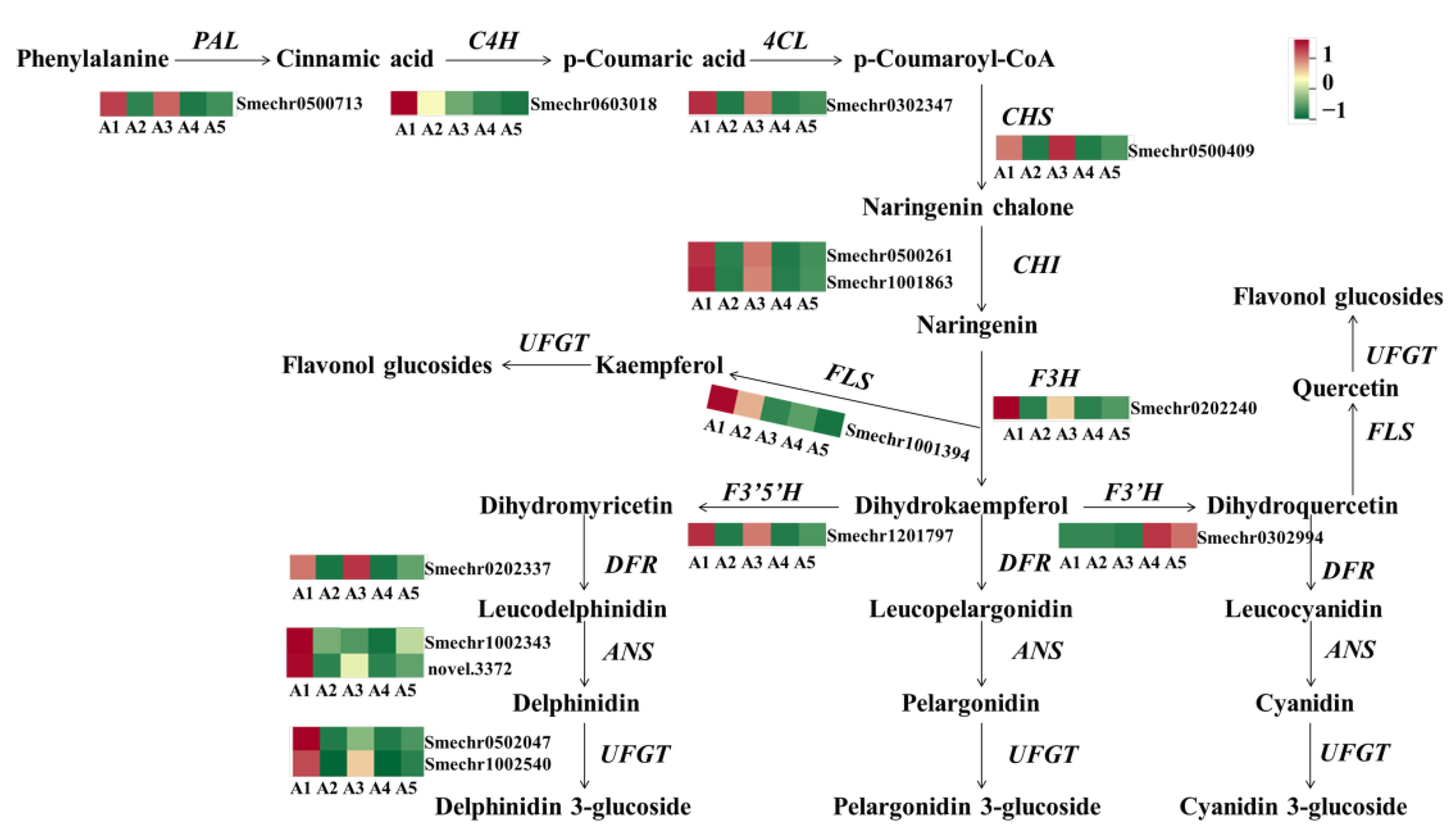
2.5. Identification of DEGs Involved in Flavonoid Biosynthesis
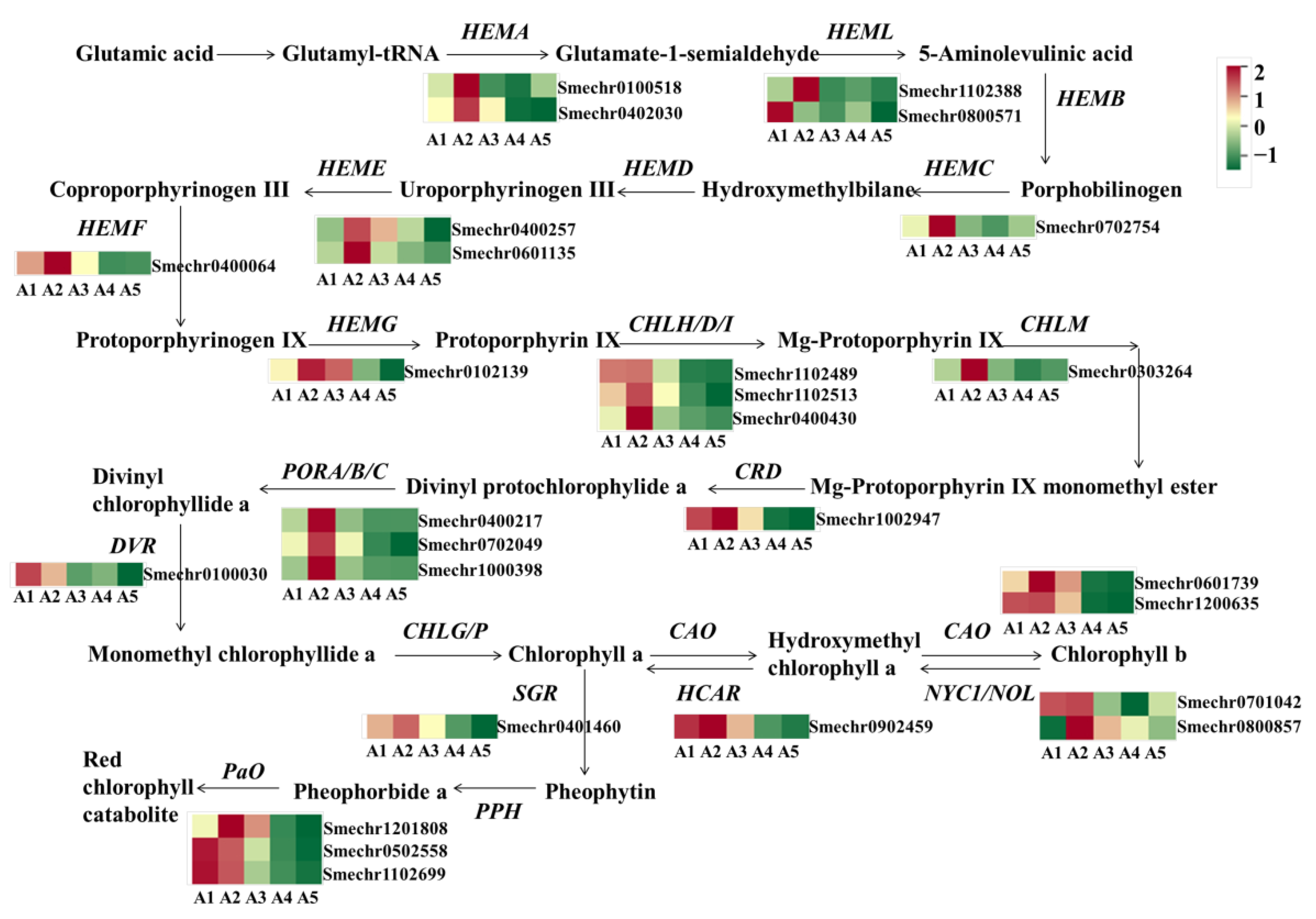
2.6. Identification of DEGs Involved in Chlorophyll Metabolism
2.7. Identification of DEGs Involved in Hormone-Mediated Signaling Pathways
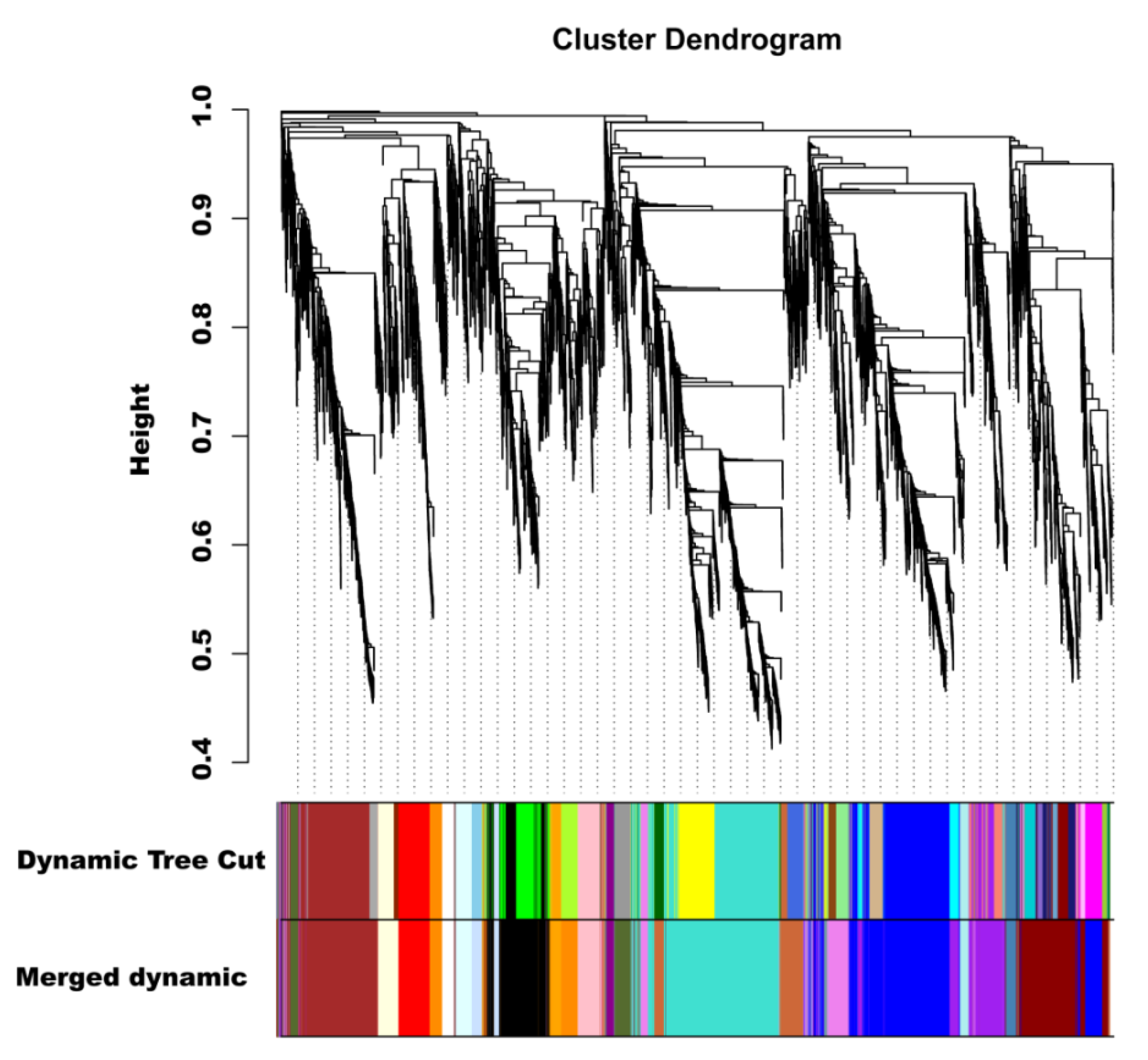
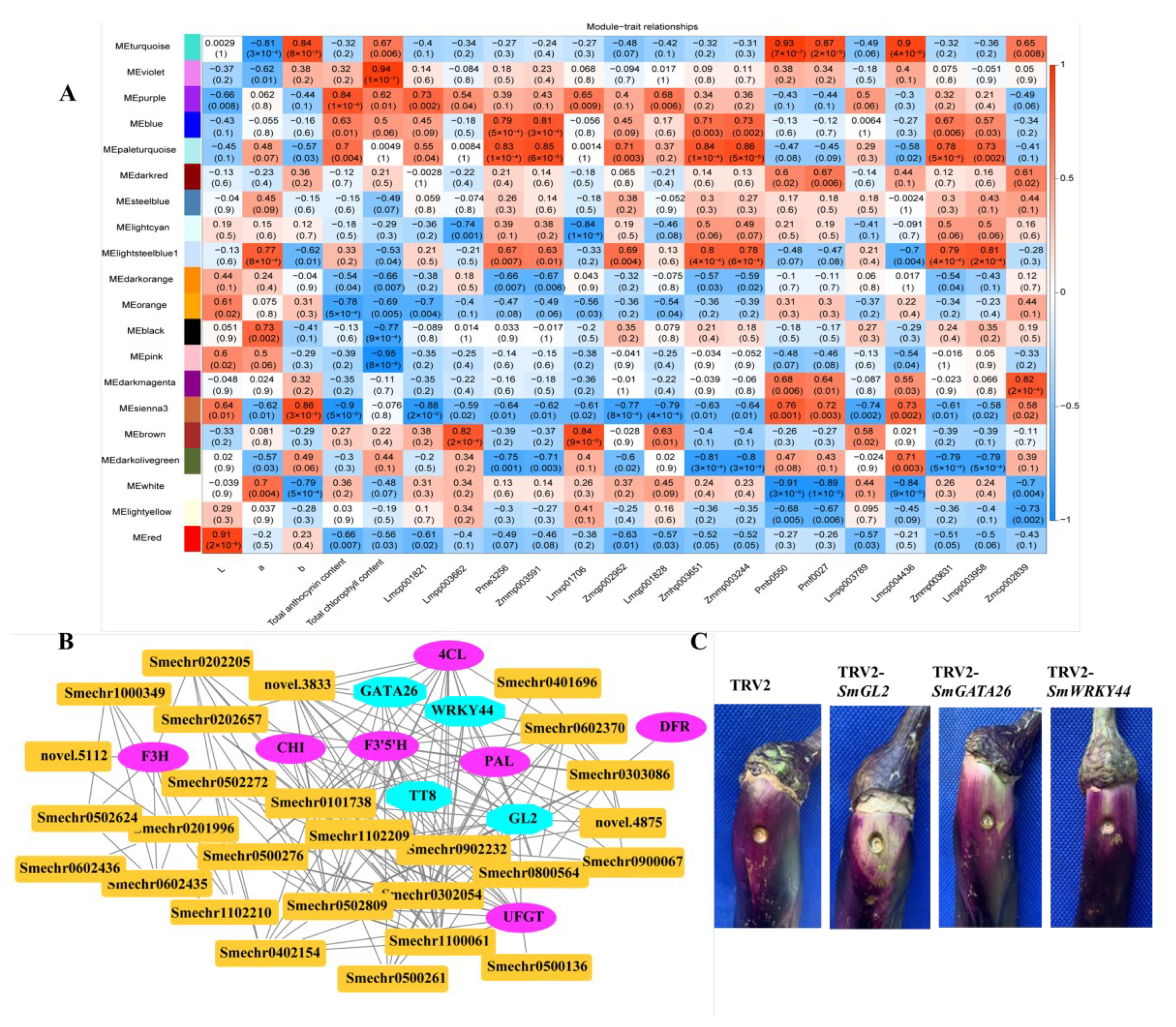
2.8. Identification of Coexpressed Gene Networks and Key Candidates
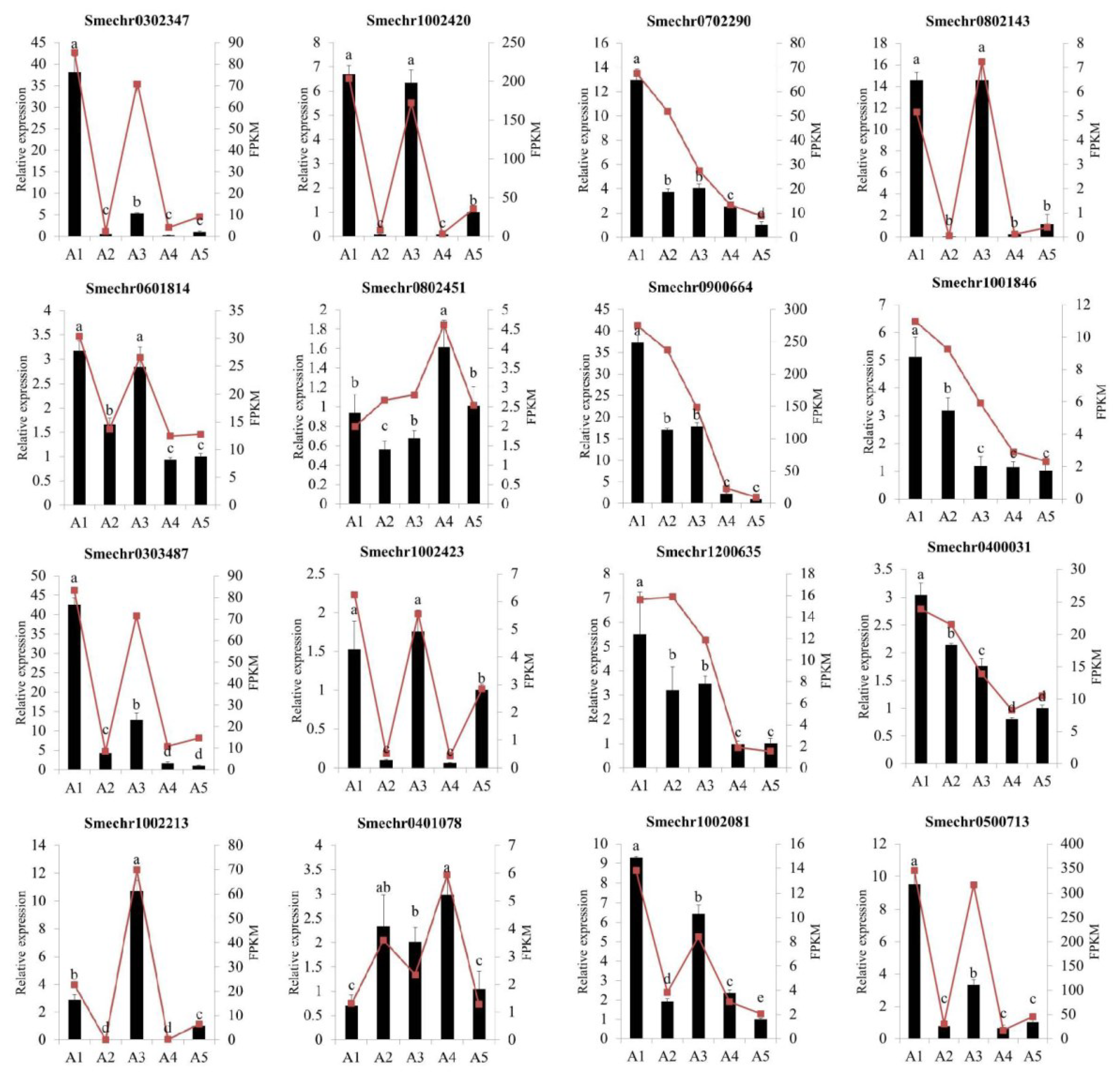
2.9. Validation of Differentially Expressed Genes by Quantitative Real-Time PCR
3. Discussion
3.1. Pigment Accumulation in Different Colors of Eggplant Fruit Peels
3.2. The Activation of Early Phenylpropanoid/Flavonoid Biosynthesis Genes for Anthocyanin Accumulation in Fruit Peels
3.3. Candidate Transcription Factors Modulated Anthocyanin Accumulation in Eggplant Fruit Peels
3.4. Plant Hormone Signaling Pathway for Color Variation in Eggplant Fruit Skins
4. Materials and Methods
4.1. Plant Materials
4.2. Total Anthocyanin and Chlorophyll Analysis
4.3. Fruit Peel Color Intensity Measurement
4.4. Flavonoid Identification and Quantification
4.5. RNA-Seq Analysis
4.6. Bioinformatic Analysis
4.7. Coexpression Network Analysis
4.8. Quantitative Real-Time PCR Analysis
4.9. Vector Construction and Transformation of Eggplant
4.10. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Shen, N.; Wang, T.F.; Gan, Q.; Liu, S.; Wang, L.; Jin, B. Plant flavonoids: Classification, distribution, biosynthesis, and antioxidant activity. Food Chem. 2022, 383, 132531. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.X.; Feng, Y.; Yu, S.H.; Fan, Z.Q.; Li, X.L.; Li, J.Y.; Yin, H.F. The flavonoid biosynthesis network in plants. Int. J. Mol. Sci. 2021, 22, 12824. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yang, Z.; Zeng, Q.W.; Wang, S.B.; Luo, Y.W.; Huang, Y.; Xin, Y.C.; He, N.J. Abnormal expression of bHLH3 disrupts a flavonoid homeostasis network, causing differences in pigment composition among mulberry fruits. Hortic. Res. 2020, 7, 83. [Google Scholar] [CrossRef]
- Mizuno, T.; Yabuya, T.; Kitajima, J.; Iwashina, T. Identification of novel Cglycosylflavones and their contribution to flower colour of the Dutch iris cultivars. Plant Physiol. Biochem. 2013, 72, 116–124. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Simal-Gandara, J.; Mei, J.F.; Ma, W.J.; Peng, Q.H.; Shi, Y.L.; Xu, Q.; Lin, Z.; Lv, H.P. Insight into the pigmented anthocyanins and the major potential co-pigmented flavonoids in purple-coloured leaf teas. Food Chem. 2021, 363, 130278. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.H.; Liu, S.Y.; Liu, Y.P.; Liu, J.; Yang, Y.; Wang, D.; Zhuang, Y. Extensive transcriptome changes underlying the fruit skin colour intensity variation in purple eggplant. Not. Bot. Horti. Agrobo. 2021, 49, 12434. [Google Scholar] [CrossRef]
- Xu, W.; Dubos, C.; Lepiniec, L. Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci. 2015, 20, 176–185. [Google Scholar] [CrossRef]
- An, J.P.; Zhang, X.W.; You, C.X.; Bi, S.Q.; Wang, X.F.; Hao, Y.J. MdWRKY40 promotes wounding-induced anthocyanin biosynthesis in association with MdMYB1 and undergoes MdBT2-mediated degradation. New Phytol. 2019, 224, 380–395. [Google Scholar] [CrossRef]
- Liu, H.N.; Su, J.; Zhu, Y.F.; Yao, G.F.; Allan, A.C.; Ampomah-Dwamena, A.; Shu, Q.; Lin-Wang, K.; Zhang, S.L.; Wu, J. The involvement of PybZIPa in light-induced anthocyanin accumulation via the activation of PyUFGT through binding to tandem G-boxes in its promoter. Hortic. Res. 2019, 6, 134. [Google Scholar] [CrossRef] [Green Version]
- Ni, J.B.; Premathilake, A.T.; Gao, Y.H.; Yu, W.J.; Tao, R.Y.; Teng, Y.W.; Bai, S.L. Ethylene-activated PpERF105 induces the expression of the repressor-type R2R3-MYB gene PpMYB140 to inhibit anthocyanin biosynthesis in red pear fruit. Plant J. 2021, 105, 167–181. [Google Scholar] [CrossRef]
- Toppino, L.; Barchi, L.; Mercati, F.; Acciarri, N.; Perrone, D.; Martina, M.; Gattolin, S.; Sala, T.; Fadda, S.; Mauceri, A.; et al. A new intra-specific and high-resolution genetic map of eggplant based on a RIL population, and location of QTLs related to plant anthocyanin pigmentation and seed vigour. Genes 2020, 11, 745. [Google Scholar] [CrossRef] [PubMed]
- Gürbüz, N.; Uluişik, S.; Frary, A.; Frary, A.; Doğanlar, S. Health benefits and bioactive compounds of eggplant. Food Chem. 2018, 268, 602–610. [Google Scholar] [CrossRef] [PubMed]
- Stommel, J.R.; Dumm, J.M. Coordinated regulation of biosynthetic and regulatory genes coincides with anthocyanin accumulation in developing eggplant fruit. J. Am. Soc. Hortic. Sci. 2015, 140, 129–135. [Google Scholar] [CrossRef]
- Jiang, M.M.; Ren, L.; Lian, H.L.; Liu, Y.; Chen, H.Y. Novel insight into the mechanism underlying light-controlled anthocyanin accumulation in eggplant (Solanum melongena L.). Plant Sci. 2016, 249, 46–58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, J.; Ren, L.; Gao, Z.; Jiang, M.M.; Liu, Y.; Zhou, L.; He, Y.J.; Chen, H.Y. Combined transcriptomic and proteomic analysis constructs a new model for light-induced anthocyanin biosynthesis in eggplant (Solanum melongena L.). Plant Cell Environ. 2017, 40, 3069–3087. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Chen, H.; Zhou, L.; Liu, Y.; Chen, H. Comparative transcription analysis of photosensitive and non-photosensitive eggplants to identify genes involved in dark regulated anthocyanin synthesis. BMC Genom. 2019, 20, 678. [Google Scholar] [CrossRef]
- Li, Y.; Chen, Y.; Zhou, L.; You, S.; Deng, H.; Chen, Y.; Alseekh, S.; Yuan, Y.; Fu, R.; Zhang, Z.; et al. MicroTom metabolic network: Rewiring tomato metabolic regulatory network throughout the growth cycle. Mol. Plant 2020, 13, 1–16. [Google Scholar] [CrossRef]
- Wang, S.; Li, Y.; He, L.; Yang, J.; Fernie, A.R.; Luo, J. Natural variance at the interface of plant primary and specialized metabolism. Curr. Opin. Plant Biol. 2022, 67, 102201. [Google Scholar] [CrossRef]
- Sadilova, E.; Stintzing, F.C.; Carle, R. Anthocyanins, colour and antioxidant properties of eggplant (Solanum melongena L.) and violet pepper (Capsicum annuum L.) peel extracts. Z. Für Nat. C 2006, 61c, 527–535. [Google Scholar] [CrossRef]
- Azuma, K.; Ohyama, A.; Ippoushi, K.; Ichiyanagi, T.; Takeuchi, A.; Saito, T.; Fukuoka, H. Structures and antioxidant activity of anthocyanins in many accessions of eggplant and its related species. J. Agric. Food Chem. 2008, 56, 10154–10159. [Google Scholar] [CrossRef]
- Zhao, C.L.; Yu, Y.Q.; Chen, Z.J.; Wen, G.S.; Wei, F.G.; Zheng, Q.; Wang, C.D.; Xiao, X.L. Stability-increasing effects of anthocyanin glycosyl acylation. Food Chem. 2017, 214, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Li, W.F.; Mao, J.; Yang, S.J.; Guo, Z.G.; Ma, Z.H.; Dawuda, M.M.; Zuo, C.W.; Chu, M.Y.; Chen, B.H. Anthocyanin accumulation correlates with hormones in the fruit skin of ‘Red Delicious’ and its four generation bud sport mutants. BMC Plant Biol. 2018, 18, 363. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, X.; Gao, G.; Yu, C.; Zhu, A.; Chen, J.; Chen, K.; Wang, X.; Abubakar, A.S.; Chen, P. Transcriptome and metabolome analysis reveals anthocyanin biosynthesis pathway associated with ramie (Boehmeria nivea (L.) Gaud.) leaf color formation. BMC Genom. 2021, 22, 684. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.B.; Li, L.J.; Wei, M.; Li, J.; Yang, F.J. SmMYB113 is a key transcription factor responsible for compositional variation of anthocyanin and color diversity among eggplant peels. Front. Plant Sci. 2022, 13, 843996. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Wang, L.; Liu, Z.; Zhao, Z.; Zhao, J.; Wang, Z.; Zhou, G.; Liu, P.; Liu, M. Transcriptome and metabolome profiling unveil the mechanisms of Ziziphus jujube Mill. peel coloration. Food Chem. 2020, 312, 125903. [Google Scholar] [CrossRef]
- Shi, Q.; Du, J.; Zhu, D.; Li, X.; Li, X. Metabolomic and transcriptomic analyses of anthocyanin biosynthesis mechanisms in the color mutant Ziziphus jujube cv. Tailihong. J. Agric. Food Chem. 2020, 68, 15186–15198. [Google Scholar] [CrossRef]
- Meng, R.; Qu, D.; Zhao, Z.; Gao, Z.; Liu, Y.; Yang, H.; Shi, X.; Zhao, Z. Anthocyanin accumulation and related gene family expression in the skin of dark-grown red and non-red apples (Malus domestica Borkh.) in response to sunlight. Sci. Hortic. 2015, 189, 66–73. [Google Scholar] [CrossRef]
- Li, L.Z.; He, Y.J.; Ge, H.Y.; Liu, Y.; Chen, H.Y. Functional characterization of SmMYB86, a negative regulator of anthocyanin biosynthesis in eggplant (Solanum melongena L.). Plant Sci. 2021, 302, 110696. [Google Scholar] [CrossRef]
- Shi, S.L.; Liu, Y.; He, Y.J.; Li, L.Z.; Li, D.L.; Chen, H.Y. R2R3-MYB transcription factor SmMYB75 promotes anthocyanin biosynthesis in eggplant (Solanum melongena L.). Sci. Hortic. 2021, 282, 110020. [Google Scholar] [CrossRef]
- Li, L.Z.; Li, S.H.; Ge, H.Y.; Shi, S.L.; Li, D.L.; Liu, Y.; Chen, H.Y. A light-responsive transcription factor SmMYB35 enhances anthocyanin biosynthesis in eggplant (Solanum melongena L.). Planta 2022, 255, 12. [Google Scholar] [CrossRef]
- Duan, Z.F.; Tian, S.Y.; Yang, G.B.; Wei, M.; Li, J.; Yang, F.J. The basic helix-loop-helix transcription factor SmbHLH1 represses anthocyanin biosynthesis in eggplant. Front. Plant Sci. 2021, 12, 757936. [Google Scholar] [CrossRef] [PubMed]
- Xi, H.C.; He, Y.J.; Chen, H.Y. Functional characterization of SmbHLH13 in anthocyanin biosynthesis and flowering in eggplant. Hortic. Plant J. 2021, 7, 73–80. [Google Scholar] [CrossRef]
- Zhou, L.; He, Y.; Li, J.; Liu, Y.; Chen, H.Y. CBFs function in anthocyanin biosynthesis by interacting with MYB113 in eggplant (Solanum melongena L.). Plant Cell Physiol. 2020, 61, 416–426. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, S.; Goto-Yamamoto, N.; Hirochika, H. Retrotransposon-induced mutations in grape skin color. Science 2004, 304, 982. [Google Scholar] [CrossRef]
- Sun, Q.; Jiang, S.; Zhang, T.; Xu, H.; Fang, H.; Zhang, J.; Su, M.; Wang, Y.; Zhang, Z.; Wang, N.; et al. Apple NAC transcription factor MdNAC52 regulates biosynthesis of anthocyanin and proanthocyanidin through MdMYB9 and MdMYB11. Plant Sci. 2019, 289, 110286. [Google Scholar] [CrossRef]
- LaFountain, A.M.; Yuan, Y.W. Repressors of anthocyanin biosynthesis. New Phytol. 2021, 31, 933–949. [Google Scholar] [CrossRef]
- Wang, X.; Wang, X.; Hu, Q.; Dai, X.; Tian, H.; Zheng, K.; Wang, X.; Mao, T.; Chen, J.G.; Wang, S. Characterization of an activation-tagged mutant uncovers a role of GLABRA2 in anthocyanin biosynthesis in Arabidopsis. Plant J. 2015, 83, 300–311. [Google Scholar] [CrossRef]
- He, Y.J.; Wang, Z.W.; Ge, H.Y.; Liu, Y.; Chen, H.Y. Weighted gene co-expression network analysis identifies genes related to anthocyanin biosynthesis and functional verification of hub gene SmWRKY44. Plant Sci. 2021, 309, 110935. [Google Scholar] [CrossRef]
- Das, P.K.; Shin, D.H.; Choi, S.B.; Park, Y. Sugar-hormone cross-talk in anthocyanin biosynthesis. Mol. Cells 2012, 34, 501–507. [Google Scholar] [CrossRef] [Green Version]
- Qi, T.; Song, S.; Ren, Q.; Wu, D.; Huang, H.; Chen, Y.; Fan, M.; Peng, W.; Ren, C.; Xie, D. The jasmonate-ZIM-domain proteins interact with the WD-Repeat/bHLH/MYB complexes to regulate jasmonate-mediated anthocyanin accumulation and trichome initiation in Arabidopsis thaliana. Plant Cell 2011, 23, 1795–1814. [Google Scholar] [CrossRef]
- An, J.; Xu, R.; Liu, X.; Zhang, J.; Wang, X.; You, C.; Hao, Y. Jasmonate induces biosynthesis of anthocyanin and proanthocyanidin in apple by mediating the JAZ1-TRB1-MYB9 complex. Plant J. 2021, 106, 1414–1430. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Xu, H.; Wang, N.; Jiang, S.; Fang, H.; Zhang, Z.; Yang, G.; Wang, Y.; Su, M.; Xu, L.; et al. The ethylene response factor MdERF1B regulates anthocyanin and proanthocyanin biosynthesis in apple. Plant Mol. Biol. 2018, 98, 205–218. [Google Scholar] [CrossRef] [PubMed]
- An, J.P.; Zhang, X.W.; Liu, Y.J.; Wang, X.F.; You, C.X.; Hao, Y.J. ABI5 regulates ABA-induced anthocyanin biosynthesis by modulating the MYB1-bHLH3 complex in apple. J. Exp. Bot. 2021, 72, 1460–1472. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Hu, Z.; Chu, G.; Huang, C.; Tian, S.; Zhao, Z.; Chen, G. Anthocyanin accumulation and molecular analysis of anthocyanin biosynthesis-associated genes in eggplant (Solanum melongena L.). J. Agric. Food Chem. 2014, 62, 2906–2912. [Google Scholar] [CrossRef]
- Florea, L.; Salzberg, S.L. Thousands of exon skipping events differentiate among splicing patterns in sixteen human tissues. F1000Research 2013, 2, 188. [Google Scholar] [CrossRef] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Xie, C.; Mao, X.; Huang, J.; Ding, Y.; Wu, J.; Dong, S.; Kong, L.; Gao, G.; Li, C.; Wei, L. KOBAS 2.0: A web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011, 39, 316–322. [Google Scholar] [CrossRef] [Green Version]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinf. 2008, 9, 559. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and 2(-Delta Delta C(T)) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
| Samples | Clean Reads | Reads Mapped | Q20 (%) | Q30 (%) | GC Content (%) |
|---|---|---|---|---|---|
| A1–1 | 47409416 | 45993726 (97.01%) | 97.69 | 93.49 | 42.23 |
| A1–2 | 46685604 | 45365789 (97.17%) | 97.77 | 93.64 | 42.46 |
| A1–3 | 48592694 | 47236677 (97.21%) | 97.8 | 93.68 | 42.69 |
| A2–1 | 48679876 | 47284281 (97.13%) | 97.75 | 93.59 | 42.33 |
| A2–2 | 43769020 | 42395918 (96.86%) | 97.67 | 93.51 | 41.91 |
| A2–3 | 49045732 | 47596182 (97.04) | 97.68 | 93.41 | 42.18 |
| A3–1 | 46608676 | 45126433 (96.82) | 97.76 | 93.63 | 41.66 |
| A3–2 | 45840606 | 44446440 (96.96%) | 97.8 | 93.73 | 41.9 |
| A3–3 | 49665704 | 47998296 (96.64%) | 97.68 | 93.5 | 42.34 |
| A4–1 | 47772226 | 46365463 (97.06%) | 97.79 | 93.67 | 42.50 |
| A4–2 | 42518600 | 41148256 (96.78%) | 97.43 | 92.92 | 42.12 |
| A4–3 | 49236290 | 47743262 (96.97%) | 97.77 | 93.64 | 42.22 |
| A5–1 | 46807752 | 45518684 (97.25%) | 97.88 | 93.9 | 42.42 |
| A5–2 | 47101278 | 46077478 (97.83%) | 98.15 | 94.37 | 42.34 |
| A5–3 | 43808476 | 42891632 (97.91%) | 98.23 | 94.57 | 42.44 |
| Comparison Groups | Total DEGs | Downregulated DEGs | Upregulated DEGs |
|---|---|---|---|
| A1 vs. A2 | 4265 | 1936 | 2329 |
| A1 vs. A3 | 3987 | 2236 | 1751 |
| A1 vs. A5 | 4005 | 2431 | 1574 |
| A2 vs. A3 | 4253 | 2507 | 1746 |
| A2 vs. A5 | 3837 | 2540 | 1297 |
| A3 vs. A5 | 3629 | 2087 | 1542 |
| A4 vs. A1 | 4456 | 1679 | 2777 |
| A4 vs. A2 | 4095 | 1314 | 2781 |
| A4 vs. A3 | 2881 | 1176 | 1705 |
| A4 vs. A5 | 2152 | 1146 | 1006 |
| Module | Gene | Annotation | Connectivity |
|---|---|---|---|
| Purple | Smechr0303487 | Homobox-leucine zipper protein GLABRA2 | 164.2872382 |
| Purple | Smechr1201797 | Flavonoid 3′5′-hydroxylase | 163.1378627 |
| Purple | Smechr0302054 | Uncharacterized protein | 163.0477793 |
| Purple | Smechr0601814 | GATA transcription factor 26 | 162.7337461 |
| Purple | Smechr1002540 | Anthocyanin 3-O-glucosyltransferase | 159.0606892 |
| Purple | Smechr0500713 | Phenylalanine ammonia-lyase 2 | 157.7681631 |
| Purple | Smechr1002420 | WRKY transcription factor 44 | 157.7222576 |
| Purple | Smechr1102209 | Kirola | 157.0941481 |
| Purple | Smechr0901701 | Anthocyanin-related transcription factor TT8 | 156.9502973 |
| Purple | Smechr0302347 | 4-coumarate-CoA ligase 2 | 156.3548796 |
| Violet | Smechr1000379 | Chlorophyll a-b binding protein | 103.30002 |
| Violet | Smechr0902639 | 30S ribosomal protein S1 | 102.53807 |
| Violet | Smechr0901976 | Uncharacterized protein | 101.83604 |
| Violet | Smechr0502558 | Protein TIC55, chloroplastic | 101.50027 |
| Violet | Smechr1100719 | ABC transporter F family member 5 | 98.330714 |
| Violet | Smechr0900664 | Chlorophyll a-b binding protein | 97.539107 |
| Violet | Smechr1001174 | Uncharacterized protein | 94.912891 |
| Violet | Smechr1200635 | Chlorophyllide a oxygenase | 90.176381 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, X.; Liu, S.; Yang, Y.; Liu, J.; Zhuang, Y. Integrated Metabolome and Transcriptome Analysis Reveals a Regulatory Network of Fruit Peel Pigmentation in Eggplant (Solanum melongena L.). Int. J. Mol. Sci. 2022, 23, 13475. https://doi.org/10.3390/ijms232113475
Zhou X, Liu S, Yang Y, Liu J, Zhuang Y. Integrated Metabolome and Transcriptome Analysis Reveals a Regulatory Network of Fruit Peel Pigmentation in Eggplant (Solanum melongena L.). International Journal of Molecular Sciences. 2022; 23(21):13475. https://doi.org/10.3390/ijms232113475
Chicago/Turabian StyleZhou, Xiaohui, Songyu Liu, Yan Yang, Jun Liu, and Yong Zhuang. 2022. "Integrated Metabolome and Transcriptome Analysis Reveals a Regulatory Network of Fruit Peel Pigmentation in Eggplant (Solanum melongena L.)" International Journal of Molecular Sciences 23, no. 21: 13475. https://doi.org/10.3390/ijms232113475
APA StyleZhou, X., Liu, S., Yang, Y., Liu, J., & Zhuang, Y. (2022). Integrated Metabolome and Transcriptome Analysis Reveals a Regulatory Network of Fruit Peel Pigmentation in Eggplant (Solanum melongena L.). International Journal of Molecular Sciences, 23(21), 13475. https://doi.org/10.3390/ijms232113475





