Normal and Aberrant TALE-Class Homeobox Gene Activities in Pro-B-Cells and B-Cell Precursor Acute Lymphoblastic Leukemia
Abstract
1. Introduction
2. Results
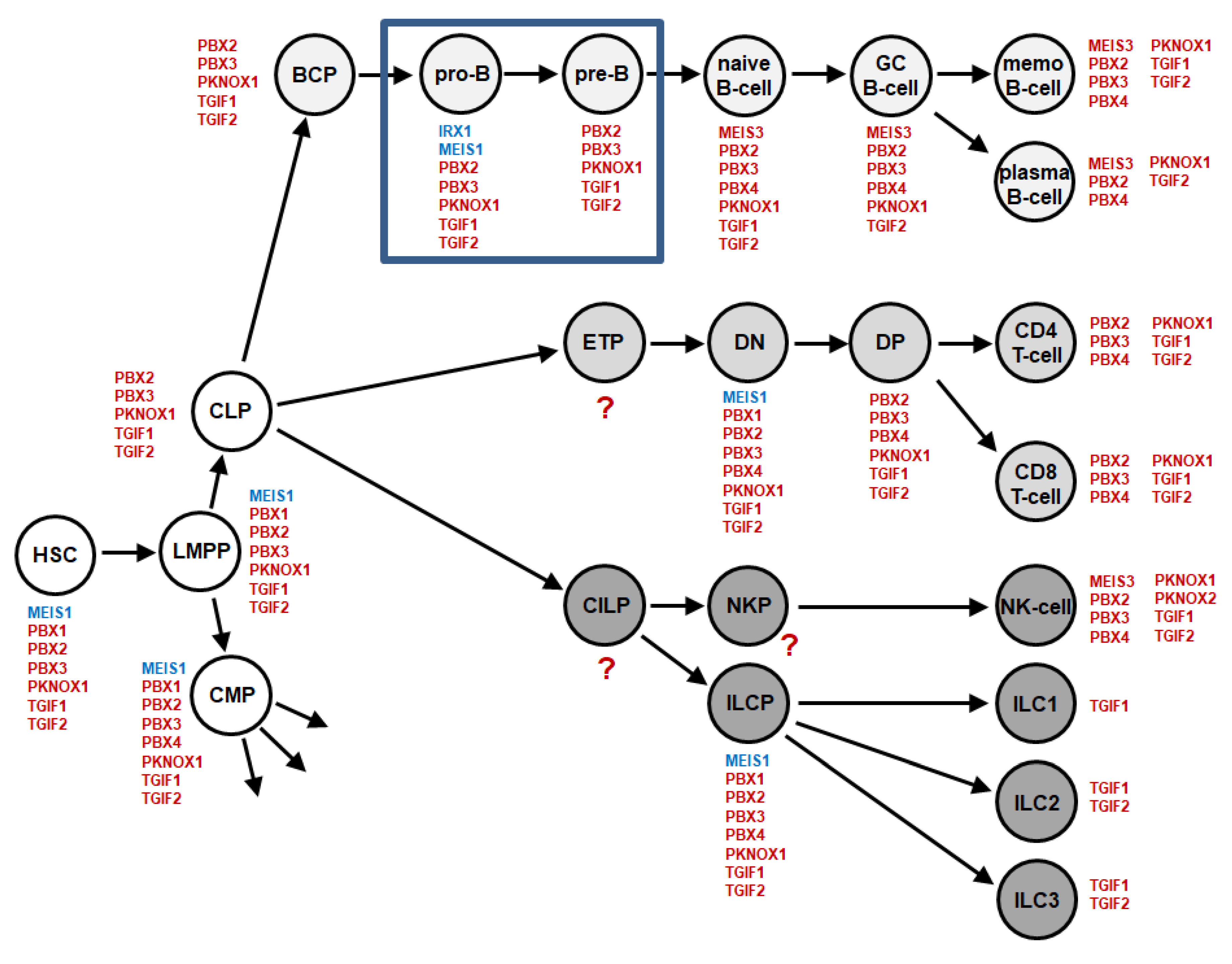
2.1. TALE Homeobox Gene Activities in Early B-Cell Development
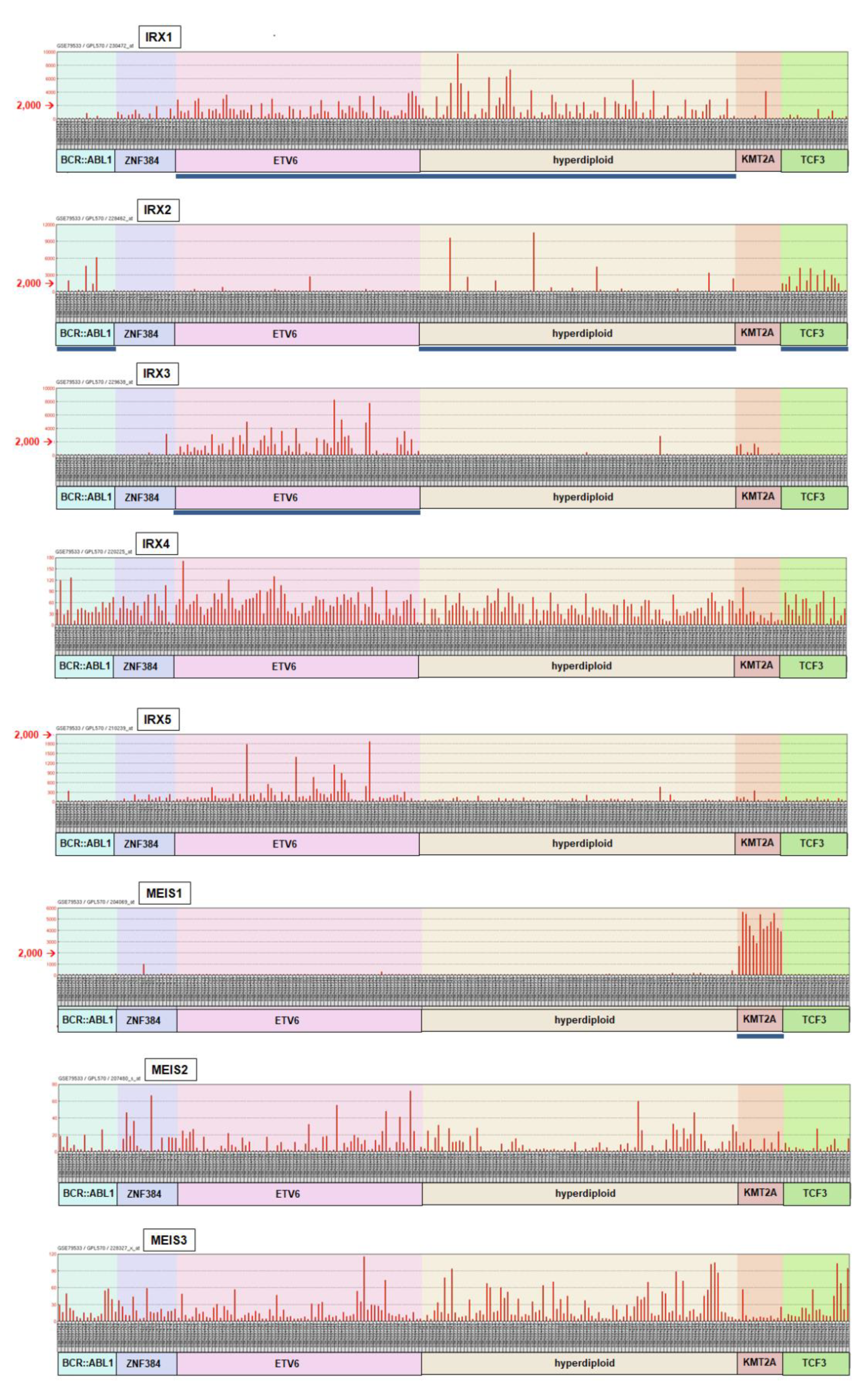
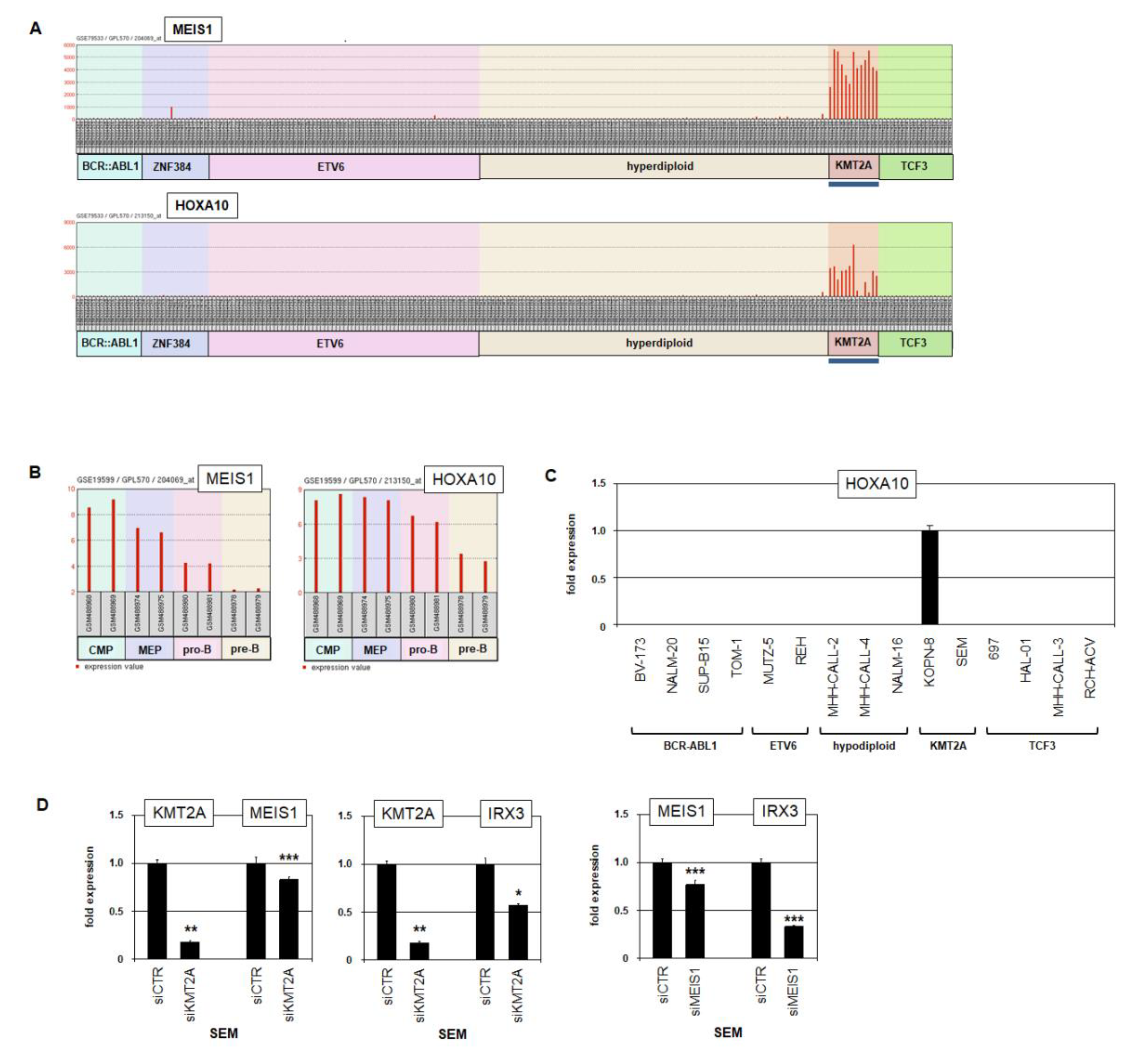
2.2. Aberrant Expression of IRX2, IRX3 and MEIS1 in BCP-ALL Patients and Cell Lines
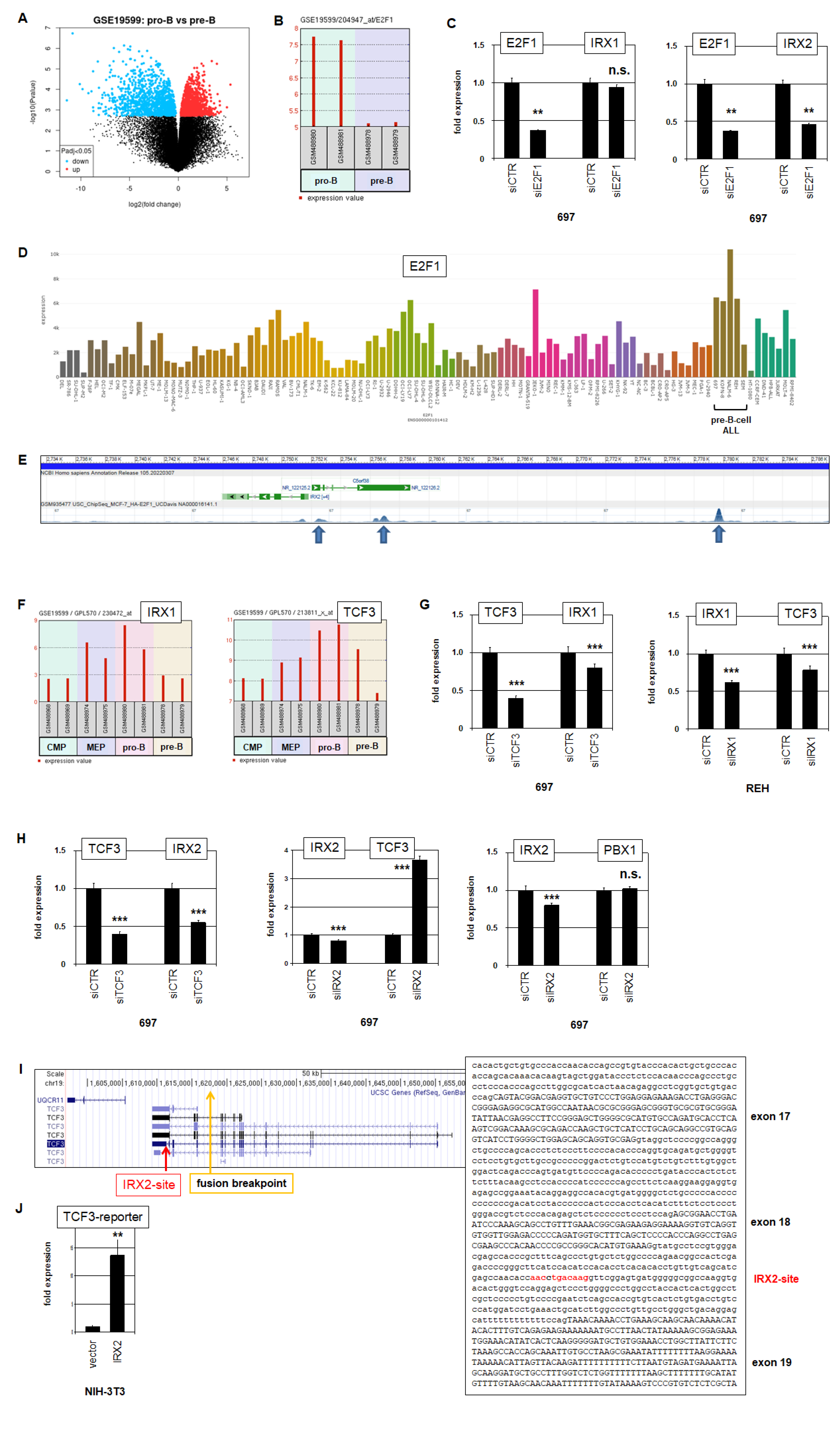
2.3. E2F1 and TCF3 Are Regulators and Targets of IRX1 and IRX2 in BCP-ALL
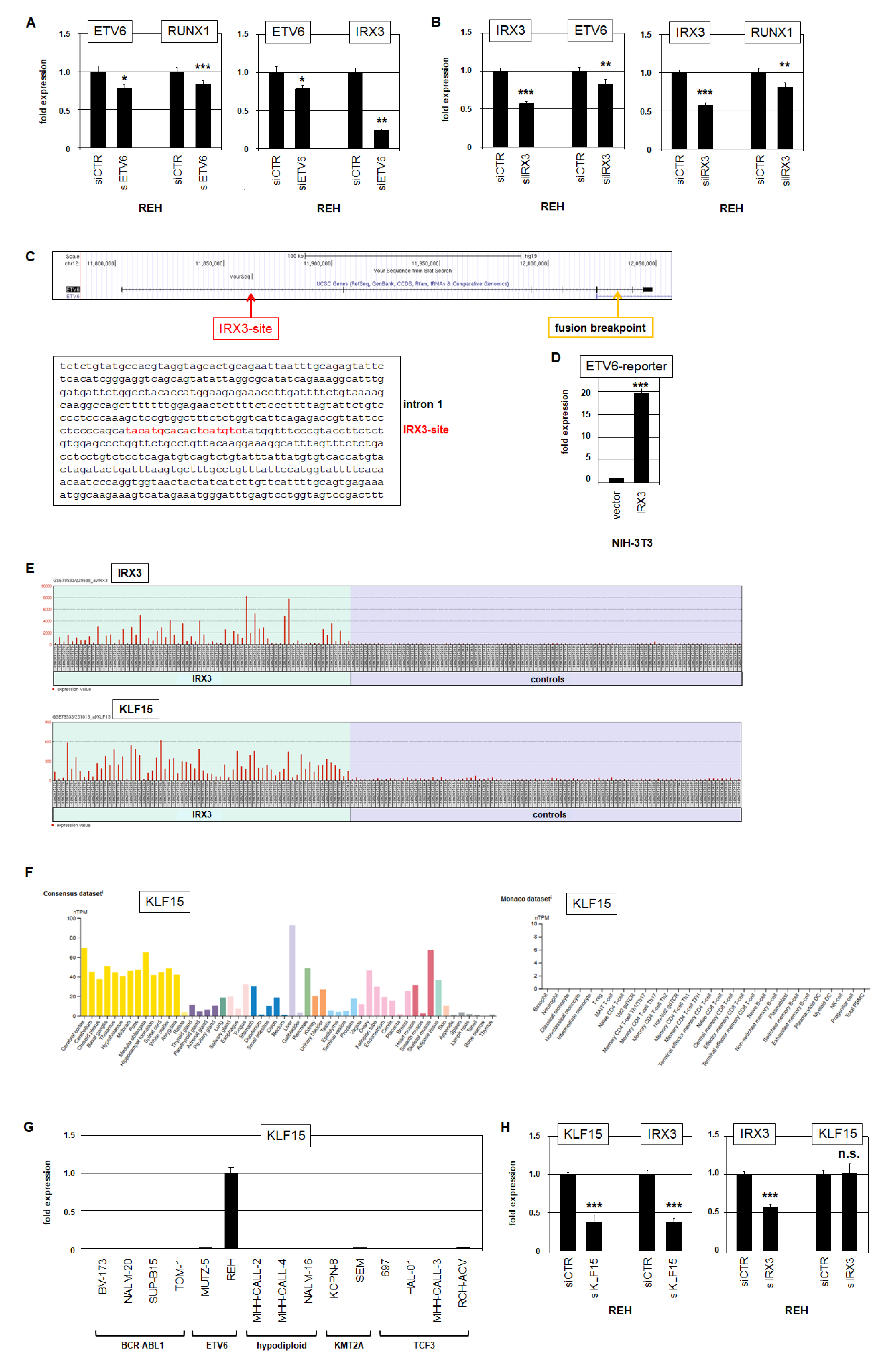
2.4. IRX3 Regulates ETV6 and KLF15 Regulates IRX3 in BCP-ALL
2.5. MEIS1 Activates Expression of IRX3 in BCP-ALL
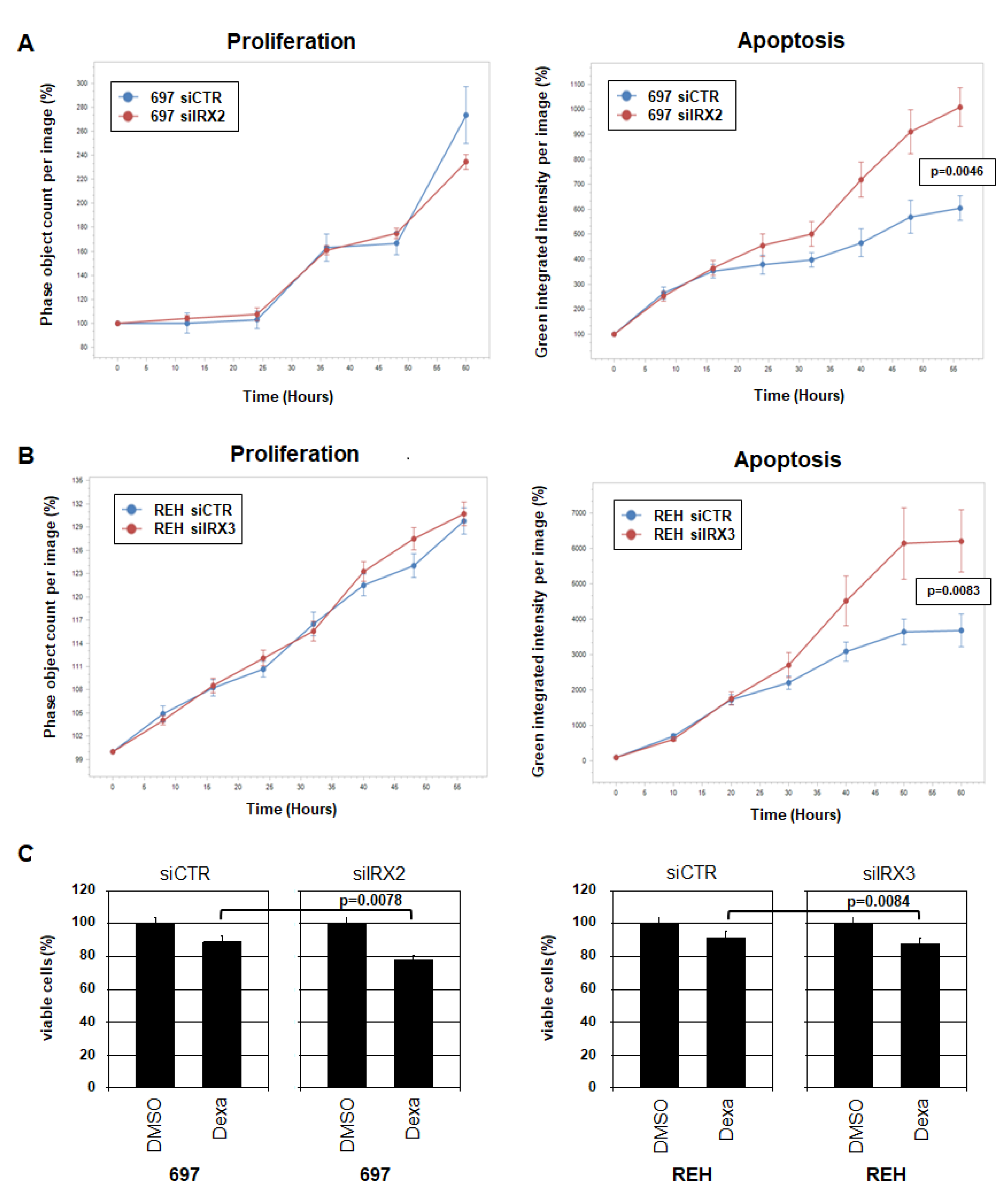
2.6. Functional Analysis of IRX2 and IRX3 in BCP-ALL
3. Discussion
4. Materials and Methods
4.1. Analysis of Expression Profiling and RNA-Seq Data
4.2. Cell Lines and Treatments
4.3. Polymerase Chain Reaction (PCR) Analyses
4.4. Reporter Gene Assays
4.5. Protein Analysis
4.6. Genomic Profiling Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hunger, S.P.; Mullighan, C.G. Acute lymphoblastic leukemia in children. N. Engl. J. Med. 2015, 373, 1541–1552. [Google Scholar] [CrossRef]
- Lilljebjörn, H.; Fioretos, T. New oncogenic subtypes in pediatric B-cell precursor acute lymphoblastic leukemia. Blood 2017, 130, 1395–1401. [Google Scholar] [CrossRef] [PubMed]
- Hirabayashi, S.; Ohki, K.; Nakabayashi, K.; Ichikawa, H.; Momozawa, Y.; Okamura, K.; Yaguchi, A.; Terada, K.; Saito, Y.; Yoshimi, A.; et al. ZNF384-related fusion genes define a subgroup of childhood B-cell precursor acute lymphoblastic leukemia with a characteristic immunotype. Haematologica 2017, 102, 118–129. [Google Scholar] [CrossRef]
- Li, J.F.; Dai, Y.T.; Lilljebjörn, H.; Shen, S.H.; Cui, B.W.; Bai, L.; Liu, Y.F.; Qian, M.X.; Kubota, Y.; Kiyoi, H.; et al. Transcriptional landscape of B cell precursor acute lymphoblastic leukemia based on an international study of 1,223 cases. Proc. Natl. Acad. Sci. USA 2018, 115, E11711–E11720. [Google Scholar] [CrossRef] [PubMed]
- Brady, S.W.; Roberts, K.G.; Gu, Z.; Shi, L.; Pounds, S.; Pei, D.; Cheng, C.; Dai, Y.; Devidas, M.; Qu, C.; et al. The genomic landscape of pediatric acute lymphoblastic leukemia. Nat. Genet. 2022, 54, 1376–1389. [Google Scholar] [CrossRef] [PubMed]
- Malouf, C.; Ottersbach, K. Molecular processes involved in B cell acute lymphoblastic leukaemia. Cell Mol. Life Sci. 2018, 75, 417–446. [Google Scholar] [CrossRef]
- Wilson, N.K.; Foster, S.D.; Wang, X.; Knezevic, K.; Schütte, J.; Kaimakis, P.; Chilarska, P.M.; Kinston, S.; Ouwehand, W.H.; Dzierzak, E.; et al. Combinatorial transcriptional control in blood stem/progenitor cells: Genome-wide analysis of ten major transcriptional regulators. Cell Stem Cell 2010, 7, 532–544. [Google Scholar] [CrossRef]
- Kucinski, I.; Wilson, N.K.; Hannah, R.; Kinston, S.J.; Cauchy, P.; Lenaerts, A.; Grosschedl, R.; Göttgens, B. Interactions between lineage-associated transcription factors govern haematopoietic progenitor states. EMBO J. 2020, 39, e104983. [Google Scholar] [CrossRef] [PubMed]
- Rothenberg, E.V. Transcriptional control of early T and B cell developmental choices. Annu. Rev. Immunol. 2014, 32, 283–321. [Google Scholar] [CrossRef]
- Bürglin, T.R.; Affolter, M. Homeodomain proteins: An update. Chromosoma 2016, 125, 497–521. [Google Scholar] [CrossRef]
- Holland, P.W.; Booth, H.A.; Bruford, E.A. Classification and nomenclature of all human homeobox genes. BMC Biol. 2007, 5, 47. [Google Scholar] [CrossRef]
- Bürglin, T.R. Analysis of TALE superclass homeobox genes (MEIS, PBC, KNOX, Iroquois, TGIF) reveals a novel domain conserved between plants and animals. Nucleic Acids Res. 1997, 25, 4173–4180. [Google Scholar] [CrossRef]
- Mukherjee, K.; Bürglin, T.R. Comprehensive analysis of animal TALE homeobox genes: New conserved motifs and cases of accelerated evolution. J. Mol. Evol. 2007, 65, 137–153. [Google Scholar] [CrossRef]
- Bosse, A.; Zülch, A.; Becker, M.B.; Torres, M.; Gómez-Skarmeta, J.L.; Modolell, J.; Gruss, P. Identification of the vertebrate Iroquois homeobox gene family with overlapping expression during early development of the nervous system. Mech. Dev. 1997, 69, 169–181. [Google Scholar] [CrossRef]
- Zhang, P.; Yang, H.; Wang, X.; Wang, L.; Cheng, Y.; Zhang, Y.; Tu, Y. The genomic organization and function of IRX1 in tumorigenesis and development. Cancer Transl. Med. 2017, 3, 29–33. [Google Scholar]
- Kühn, A.; Löscher, D.; Marschalek, R. The IRX1/HOXA connection: Insights into a novel t(4;11)- specific cancer mechanism. Oncotarget 2016, 7, 35341–35352. [Google Scholar] [CrossRef] [PubMed]
- Somerville, T.D.D.; Simeoni, F.; Chadwick, J.A.; Williams, E.L.; Spencer, G.J.; Boros, K.; Wirth, C.; Tholouli, E.; Byers, R.J.; Somervaille, T.C.P. Derepression of the Iroquois homeodomain transcription factor gene IRX3 confers differentiation block in acute leukemia. Cell Rep. 2018, 22, 638–652. [Google Scholar] [CrossRef] [PubMed]
- Nagel, S.; Pommerenke, C.; Meyer, C.; MacLeod, R.A.F. The hematopoietic TALE-code shows normal activity of IRX1 in myeloid progenitors and reveals ectopic expression of IRX3 and IRX5 in acute myeloid leukemia. Int. J. Mol. Sci. 2022, 23, 3192. [Google Scholar] [CrossRef]
- Longobardi, E.; Penkov, D.; Mateos, D.; De Florian, G.; Torres, M.; Blasi, F. Biochemistry of the tale transcription factors PREP, MEIS, and PBX in vertebrates. Dev. Dyn. 2014, 243, 59–75. [Google Scholar] [CrossRef] [PubMed]
- Yao, M.; Gu, Y.; Yang, Z.; Zhong, K.; Chen, Z. MEIS1 and its potential as a cancer therapeutic target (Review). Int. J. Mol. Med. 2021, 48, 181. [Google Scholar] [CrossRef]
- Slany, R.K. The molecular biology of mixed lineage leukemia. Haematologica 2009, 94, 984–993. [Google Scholar] [CrossRef]
- Ferrando, A.A.; Armstrong, S.A.; Neuberg, D.S.; Sallan, S.E.; Silverman, L.B.; Korsmeyer, S.J.; Look, A.T. Gene expression signatures in MLL-rearranged T-lineage and B-precursor acute leukemias: Dominance of HOX dysregulation. Blood 2003, 102, 262–268. [Google Scholar] [CrossRef]
- Graux, C.; Cools, J.; Michaux, L.; Vandenberghe, P.; Hagemeijer, A. Cytogenetics and molecular genetics of T-cell acute lymphoblastic leukemia: From thymocyte to lymphoblast. Leukemia 2006, 20, 1496–1510. [Google Scholar] [CrossRef]
- Alharbi, R.A.; Pettengell, R.; Pandha, H.S.; Morgan, R. The role of HOX genes in normal hematopoiesis and acute leukemia. Leukemia 2013, 27, 1000–1008. [Google Scholar] [CrossRef]
- Nagel, S. NKL-code in normal and aberrant hematopoiesis. Cancers 2021, 13, 1961. [Google Scholar] [CrossRef]
- Nagel, S.; Pommerenke, C.; Meyer, C.; MacLeod, R.A.F.; Drexler, H.G. Establishment of the TALE-code reveals aberrantly activated homeobox gene PBX1 in Hodgkin lymphoma. PLoS ONE 2021, 16, e0246603. [Google Scholar] [CrossRef]
- Andersson, A.; Edén, P.; Olofsson, T.; Fioretos, T. Gene expression signatures in childhood acute leukemias are largely unique and distinct from those of normal tissues and other malignancies. BMC Med. Genom. 2010, 3, 6. [Google Scholar] [CrossRef]
- Fischer, U.; Yang, J.J.; Ikawa, T.; Hein, D.; Vicente-Dueñas, C.; Borkhardt, A.; Sánchez-García, I. Cell Fate Decisions: The role of transcription factors in early B-cell development and leukemia. Blood Cancer Discov. 2020, 1, 224–233. [Google Scholar] [CrossRef]
- Bialkowska, A.B.; Yang, V.W.; Mallipattu, S.K. Krüppel-like factors in mammalian stem cells and development. Development 2017, 144, 737–754. [Google Scholar] [CrossRef]
- Tallack, M.R.; Perkins, A.C. KLF1 directly coordinates almost all aspects of terminal erythroid differentiation. IUBMB Life 2010, 62, 886–890. [Google Scholar]
- Mamo, A.; Krosl, J.; Kroon, E.; Bijl, J.; Thompson, A.; Mayotte, N.; Girard, S.; Bisaillon, R.; Beslu, N.; Featherstone, M.; et al. Molecular dissection of Meis1 reveals 2 domains required for leukemia induction and a key role for Hoxa gene activation. Blood 2006, 108, 622–629. [Google Scholar] [CrossRef]
- Teachey, D.T.; Pui, C.H. Comparative features and outcomes between paediatric T-cell and B-cell acute lymphoblastic leukaemia. Lancet Oncol. 2019, 20, e142–e154. [Google Scholar] [CrossRef]
- Nagel, S.; Meyer, C. Establishment of the TBX-code reveals aberrantly activated T-box gene TBX3 in Hodgkin lymphoma. PLoS ONE 2021, 16, e0259674. [Google Scholar] [CrossRef]
- Kwon, K.; Hutter, C.; Sun, Q.; Bilic, I.; Cobaleda, C.; Malin, S.; Busslinger, M. Instructive role of the transcription factor E2A in early B lymphopoiesis and germinal center B cell development. Immunity 2008, 28, 751–762. [Google Scholar] [CrossRef]
- Lin, Y.C.; Jhunjhunwala, S.; Benner, C.; Heinz, S.; Welinder, E.; Mansson, R.; Sigvardsson, M.; Hagman, J.; Espinoza, C.A.; Dutkowski, J.; et al. A global network of transcription factors, involving E2A, EBF1 and Foxo1, that orchestrates B cell fate. Nat. Immunol. 2010, 11, 635–643. [Google Scholar] [CrossRef]
- Somasundaram, R.; Prasad, M.A.; Ungerbäck, J.; Sigvardsson, M. Transcription factor networks in B-cell differentiation link development to acute lymphoid leukemia. Blood 2015, 126, 144–152. [Google Scholar] [CrossRef]
- Neveu, B.; Caron, M.; Lagacé, K.; Richer, C.; Sinnett, D. Genome wide mapping of ETV6 binding sites in pre-B leukemic cells. Sci. Rep. 2018, 8, 15526. [Google Scholar] [CrossRef]
- Meyer, C.; MacLeod, R.A.; Quentmeier, H.; Janssen, J.W.; Coignet, L.J.; Dyer, M.J.; Drexler, H.G. Establishment of the B cell precursor acute lymphoblastic leukemia cell line MUTZ-5 carrying a (12:13) translocation. Leukemia 2001, 15, 1471–1474. [Google Scholar] [CrossRef]
- Poppy Roworth, A.; Ghari, F.; La Thangue, N.B. To live or let die—Complexity within the E2F1 pathway. Mol. Cell Oncol. 2015, 2, e970480. [Google Scholar] [CrossRef]
- Gala, S.; Marreiros, A.; Stewart, G.J.; Williamson, P. Overexpression of E2F-1 leads to cytokine-independent proliferation and survival in the hematopoietic cell line BaF-B03. Blood 2001, 97, 227–234. [Google Scholar] [CrossRef]
- Uhlen, M.; Karlsson, M.J.; Zhong, W.; Tebani, A.; Pou, C.; Mikes, J.; Lakshmikanth, T.; Forsström, B.; Edfors, F.; Odeberg, J.; et al. A genome-wide transcriptomic analysis of protein-coding genes in human blood cells. Science 2019, 366, eaax9198. [Google Scholar] [CrossRef]
- Quentmeier, H.; Pommerenke, C.; Dirks, W.G.; Eberth, S.; Koeppel, M.; MacLeod, R.A.F.; Nagel, S.; Steube, K.; Uphoff, C.C.; Drexler, H.G. The LL-100 panel: 100 cell lines for blood cancer studies. Sci. Rep. 2019, 9, 8218. [Google Scholar] [CrossRef]
- Bungaro, S.; Dell’Orto, M.C.; Zangrando, A.; Basso, D.; Gorletta, T.; Lo Nigro, L.; Leszl, A.; Young, B.D.; Basso, G.; Bicciato, S.; et al. Integration of genomic and gene expression data of childhood ALL without known aberrations identifies subgroups with specific genetic hallmarks. Genes Chromosomes Cancer. 2009, 48, 22–38. [Google Scholar] [CrossRef]
- Davis, S.; Meltzer, P.S. GEOquery: A bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 2007, 23, 1846–1867. [Google Scholar] [CrossRef]
- Van Dongen, J.J.; Macintyre, E.A.; Gabert, J.A.; Delabesse, E.; Rossi, V.; Saglio, G.; Gottardi, E.; Rambaldi, A.; Dotti, G.; Griesinger, F.; et al. Standardized RT-PCR analysis of fusion gene transcripts from chromosome aberrations in acute leukemia for detection of minimal residual disease. Report of the BIOMED-1 Concerted Action: Investigation of minimal residual disease in acute leukemia. Leukemia 1999, 13, 1901–1928. [Google Scholar] [CrossRef]
- Weirauch, M.T.; Yang, A.; Albu, M.; Cote, A.G.; Montenegro-Montero, A.; Drewe, P.; Najafabadi, H.S.; Lambert, S.A.; Mann, I.; Cook, K.; et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 2014, 158, 1431–1443. [Google Scholar] [CrossRef]
- Nagel, S.; Schneider, B.; Meyer, C.; Kaufmann, M.; Drexler, H.G.; Macleod, R.A. Transcriptional deregulation of homeobox gene ZHX2 in Hodgkin lymphoma. Leuk. Res. 2012, 36, 646–655. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nagel, S.; Meyer, C. Normal and Aberrant TALE-Class Homeobox Gene Activities in Pro-B-Cells and B-Cell Precursor Acute Lymphoblastic Leukemia. Int. J. Mol. Sci. 2022, 23, 11874. https://doi.org/10.3390/ijms231911874
Nagel S, Meyer C. Normal and Aberrant TALE-Class Homeobox Gene Activities in Pro-B-Cells and B-Cell Precursor Acute Lymphoblastic Leukemia. International Journal of Molecular Sciences. 2022; 23(19):11874. https://doi.org/10.3390/ijms231911874
Chicago/Turabian StyleNagel, Stefan, and Corinna Meyer. 2022. "Normal and Aberrant TALE-Class Homeobox Gene Activities in Pro-B-Cells and B-Cell Precursor Acute Lymphoblastic Leukemia" International Journal of Molecular Sciences 23, no. 19: 11874. https://doi.org/10.3390/ijms231911874
APA StyleNagel, S., & Meyer, C. (2022). Normal and Aberrant TALE-Class Homeobox Gene Activities in Pro-B-Cells and B-Cell Precursor Acute Lymphoblastic Leukemia. International Journal of Molecular Sciences, 23(19), 11874. https://doi.org/10.3390/ijms231911874





