Postsynaptic Proteins at Excitatory Synapses in the Brain—Relationship with Depressive Disorders
Abstract
:1. Introduction
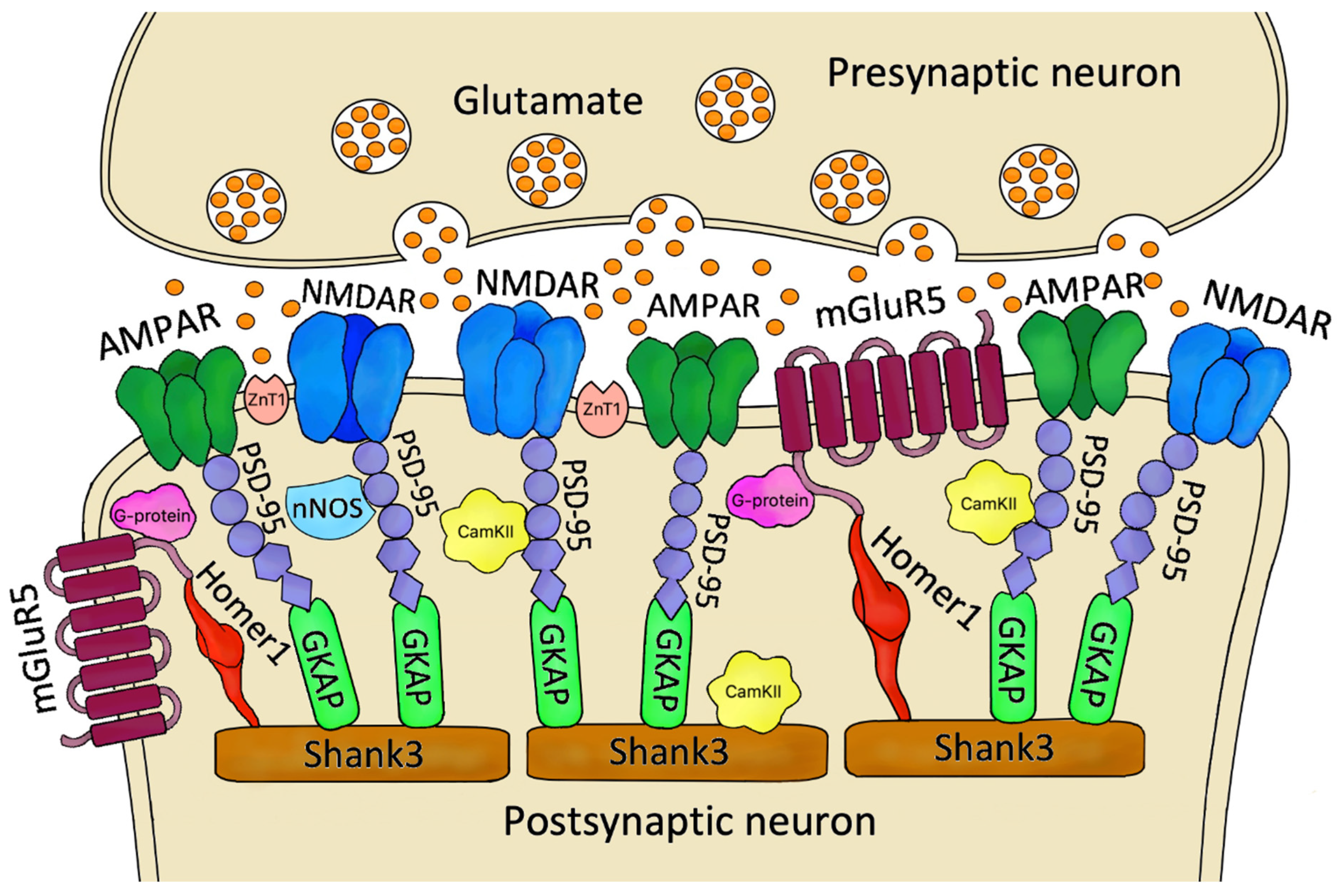
2. The Role of Postsynaptic Density Proteins in the Excitatory Synapse
3. Alterations in Postsynaptic Density Proteins in Depressive Disorders
3.1. Human Studies
3.2. Animal Studies
4. Postsynaptic Density Proteins as Therapeutic Targets for DDs
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- World Health Organization (WHO). Depression. Available online: https://www.who.int/news-room/fact-sheets/detail/depression (accessed on 9 June 2022).
- Zenebe, Y.; Akele, B.; Woldeselassie Gebre, M.; Necho, M. Prevalence and determinants of depression among old age: A systematic review and meta-analysis. Ann. Gen. Psychiatry 2021, 20, 55. [Google Scholar] [CrossRef] [PubMed]
- Duman, R.S.; Sanacora, G.; Krystal, J.H. Altered Connectivity in Depression: GABA and Glutamate Neurotransmitter Deficits and Reversal by Novel Treatments. Neuron 2019, 102, 75–90. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization (WHO). Adolescent Mental Health. Available online: https://www.who.int/news-room/fact-sheets/detail/adolescent-mental-health (accessed on 9 June 2022).
- Hossain, M.M.; Nesa, F.; Das, J.; Aggad, R.; Tasnim, S.; Bairwa, M.; Ma, P.; Ramirez, G. Global burden of mental health problems among children and adolescents during COVID-19 pandemic: An umbrella review. Psychiatry Res. 2022, 317, 114814. [Google Scholar] [CrossRef] [PubMed]
- Thapar, A.; Eyre, O.; Patel, V.; Brent, D. Depression in young people. Lancet 2022, 400, 617–631. [Google Scholar] [CrossRef]
- Thompson, S.M.; Kallarackal, A.J.; Kvarta, M.D.; Van Dyke, A.M.; LeGates, T.A.; Cai, X. An excitatory synapse hypothesis of depression. Trends Neurosci. 2015, 38, 279–294. [Google Scholar] [CrossRef]
- Tian, H.; Hu, Z.; Xu, J.; Wang, C. The molecular pathophysiology of depression and the new therapeutics. MedComm 2022, 3, e156. [Google Scholar] [CrossRef]
- Blackburn, T.P. Depressive disorders: Treatment failures and poor prognosis over the last 50 years. Pharmacol. Res. Perspect. 2019, 7, e00472. [Google Scholar] [CrossRef]
- Voineskos, D.; Daskalakis, Z.J.; Blumberger, D.M. Management of Treatment-Resistant Depression: Challenges and Strategies. Neuropsychiatr. Dis. Treat. 2020, 16, 221–234. [Google Scholar] [CrossRef]
- Adu, M.K.; Shalaby, R.; Chue, P.; Agyapong, V.I.O. Repetitive Transcranial Magnetic Stimulation for the Treatment of Resistant Depression: A Scoping Review. Behav. Sci. 2022, 12, 195. [Google Scholar] [CrossRef]
- Papalexi, E.; Galanopoulos, A.; Roukas, D.; Argyropoulos, I.; Michopoulos, I.; Douzenis, A.; Gkolia, I.; Fotiadis, P.; Kontis, D.; Zervas, I.M. Residual cognitive and psychosocial functional impairment in outpatients in Greece who responded to conventional antidepressant monotherapy treatments for major depressive disorder (MDD). J. Affect. Disord. 2022, 314, 185–192. [Google Scholar] [CrossRef]
- Saez, E.; Erkoreka, L.; Moreno-Calle, T.; Berjano, B.; Gonzalez-Pinto, A.; Basterreche, N.; Arrue, A. Genetic variables of the glutamatergic system associated with treatment-resistant depression: A review of the literature. World J. Psychiatry 2022, 12, 884–896. [Google Scholar] [CrossRef] [PubMed]
- Chevance, A.; Tomlinson, A.; Ravaud, P.; Touboul, S.; Henshall, C.; Tran, V.T.; Cipriani, A. Important adverse events to be evaluated in antidepressant trials and meta-analyses in depression: A large international preference study including patients and healthcare professionals. Evid. Based Ment. Health 2022. [Google Scholar] [CrossRef] [PubMed]
- Pilc, A.; Machaczka, A.; Kawalec, P.; Smith, J.L.; Witkin, J.M. Where do we go next in antidepressant drug discovery? A new generation of antidepressants: A pivotal role of AMPA receptor potentiation and mGlu2/3 receptor antagonism. Expert Opin. Drug Discov. 2022, 22, 1–16. [Google Scholar] [CrossRef]
- Hartig, J.; Nemeș, B. BDNF-related mutations in major depressive disorder: A systematic review. Acta Neuropsychiatr. 2022, 22, 1–61. [Google Scholar] [CrossRef] [PubMed]
- Ramos-da-Silva, L.; Carlson, P.T.; Silva-Costa, L.C.; Martins-de-Souza, D.; de Almeida, V. Molecular Mechanisms Associated with Antidepressant Treatment on Major Depression. Complex Psychiatry 2021, 7, 49–59. [Google Scholar] [CrossRef]
- Bijata, M.; Bączyńska, E.; Wlodarczyk, J. A chronic unpredictable stress protocol to model anhedonic and resilient behaviors in C57BL/6J mice. STAR Protoc. 2022, 3, 101659. [Google Scholar] [CrossRef]
- Koert, A.; Ploeger, A.; Bockting, C.L.H.; Schmidt, M.V.; Lucassen, P.J.; Schrantee, A.; Mul, J.D. The social instability stress paradigm in rat and mouse: A systematic review of protocols, limitations, and recommendations. Neurobiol. Stress 2021, 15, 100410. [Google Scholar] [CrossRef]
- Kim, I.B.; Lee, J.H.; Park, S.C. The Relationship between Stress, Inflammation, and Depression. Biomedicines 2022, 10, 1929. [Google Scholar] [CrossRef]
- Willner, P.; Gruca, P.; Lason, M.; Tota-Glowczyk, K.; Litwa, E.; Niemczyk, M.; Papp, M. Validation of chronic mild stress in the Wistar-Kyoto rat as an animal model of treatment-resistant depression. Behav. Pharmacol. 2019, 30, 239–250. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhang, Q.; Shao, X.; Ouyang, L.; Wang, X.; Zhu, K.; Chen, L. Decreased Glycogen Content Might Contribute to Chronic Stress-Induced Atrophy of Hippocampal Astrocyte volume and Depression-like Behavior in Rats. Sci. Rep. 2017, 7, 43192. [Google Scholar] [CrossRef] [Green Version]
- Evans, J.W.; Szczepanik, J.; Brutsché, N.; Park, L.T.; Nugent, A.C.; Zarate, C.A., Jr. Default Mode Connectivity in Major Depressive Disorder Measured Up to 10 Days After Ketamine Administration. Biol. Psychiatry 2018, 84, 582–590. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Han, K.M.; Choi, K.W.; Tae, W.S.; Kang, W.; Kang, Y.; Kim, A.; Ham, B.J. Volumetric alterations in subregions of the amygdala in adults with major depressive disorder. J. Affect. Disord. 2021, 295, 108–115. [Google Scholar] [CrossRef] [PubMed]
- Belleau, E.L.; Treadway, M.T.; Pizzagalli, D.A. The Impact of Stress and Major Depressive Disorder on Hippocampal and Medial Prefrontal Cortex Morphology. Biol. Psychiatry 2019, 85, 443–453. [Google Scholar] [CrossRef] [PubMed]
- Chenani, A.; Weston, G.; Ulivi, A.F.; Castello-Waldow, T.P.; Huettl, R.E.; Chen, A.; Attardo, A. Repeated stress exposure leads to structural synaptic instability prior to disorganization of hippocampal coding and impairments in learning. Transl. Psychiatry 2022, 12, 381. [Google Scholar] [CrossRef] [PubMed]
- Brosch, K.; Stein, F.; Schmitt, S.; Pfarr, J.K.; Ringwald, K.G.; Thomas-Odenthal, F.; Meller, T.; Steinsträter, O.; Waltemate, L.; Lemke, H.; et al. Reduced hippocampal gray matter volume is a common feature of patients with major depression, bipolar disorder, and schizophrenia spectrum disorders. Mol. Psychiatry 2022. [Google Scholar] [CrossRef]
- Yang, Y.; Li, X.; Cui, Y.; Liu, K.; Qu, H.; Lu, Y.; Li, W.; Zhang, L.; Zhang, Y.; Song, J.; et al. Reduced Gray Matter Volume in Orbitofrontal Cortex Across Schizophrenia, Major Depressive Disorder, and Bipolar Disorder: A Comparative Imaging Study. Front. Neurosci. 2022, 16, 919272. [Google Scholar] [CrossRef]
- Phillips, C. Brain-Derived Neurotrophic Factor, Depression, and Physical Activity: Making the Neuroplastic Connection. Neural. Plast. 2017, 2017, 7260130. [Google Scholar] [CrossRef]
- Gałecki, P.; Talarowska, M. Inflammatory theory of depression. Psychiatr. Pol. 2018, 52, 437–447. [Google Scholar] [CrossRef]
- Xu, L.; Sun, H.; Qu, C.; Shen, J.; Qu, C.; Song, H.; Li, T.; Zheng, J.; Zhang, J. The environmental enrichment ameliorates chronic unpredictable mild stress-induced depressive-like behaviors and cognitive decline by inducing autophagy-mediated inflammation inhibition. Brain Res. Bull. 2022, 187, 98–110. [Google Scholar] [CrossRef]
- Mograbi, K.M.; Suchecki, D.; da Silva, S.G.; Covolan, L.; Hamani, C. Chronic unpredictable restraint stress increases hippocampal pro-inflammatory cytokines and decreases motivated behavior in rats. Stress 2020, 23, 427–436. [Google Scholar] [CrossRef]
- Koo, J.W.; Chaudhury, D.; Han, M.H.; Nestler, E.J. Role of Mesolimbic Brain-Derived Neurotrophic Factor in Depression. Biol. Psychiatry 2019, 86, 738–748. [Google Scholar] [CrossRef] [PubMed]
- Sowa-Kućma, M.; Styczeń, K.; Siwek, M.; Misztak, P.; Nowak, R.J.; Dudek, D.; Rybakowski, J.K.; Nowak, G.; Maes, M. Lipid Peroxidation and Immune Biomarkers Are Associated with Major Depression and Its Phenotypes, Including Treatment-Resistant Depression and Melancholia. Neurotox. Res. 2018, 33, 448–460. [Google Scholar] [CrossRef]
- Popoli, M.; Yan, Z.; McEwen, B.S.; Sanacora, G. The stressed synapse: The impact of stress and glucocorticoids on glutamate transmission. Nat. Rev. Neurosci. 2011, 13, 22–37. [Google Scholar] [CrossRef] [PubMed]
- Reagan, L.P.; Reznikov, L.R.; Evans, A.N.; Gabriel, C.; Mocaër, E.; Fadel, J.R. The antidepressant agomelatine inhibits stress-mediated changes in amino acid efflux in the rat hippocampus and amygdala. Brain Res. 2012, 1466, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Reznikov, L.R.; Grillo, C.A.; Piroli, G.G.; Pasumarthi, R.K.; Reagan, L.P.; Fadel, J. Acute stress-mediated increases in extracellular glutamate levels in the rat amygdala: Differential effects of antidepressant treatment. Eur. J. Neurosci. 2007, 25, 3109–3114. [Google Scholar] [CrossRef] [PubMed]
- Blacker, C.J.; Millischer, V.; Webb, L.M.; Ho, A.M.C.; Schalling, M.; Frye, M.A.; Veldic, M. EAAT2 as a Research Target in Bipolar Disorder and Unipolar Depression: A Systematic Review. Mol. Neuropsychiatry 2020, 5 (Suppl. S1), 44–59. [Google Scholar] [CrossRef]
- Gruenbaum, B.F.; Zlotnik, A.; Frenkel, A.; Fleidervish, I.; Boyko, M. Glutamate Efflux across the Blood-Brain Barrier: New Perspectives on the Relationship between Depression and the Glutamatergic System. Metabolites 2022, 12, 459. [Google Scholar] [CrossRef]
- Sarawagi, A.; Soni, N.D.; Patel, A.B. Glutamate and GABA Homeostasis and Neurometabolism in Major Depressive Disorder. Front. Psychiatry 2021, 27, 637863. [Google Scholar] [CrossRef]
- Henter, I.D.; Park, L.T.; Zarate, C.A., Jr. Novel Glutamatergic Modulators for the Treatment of Mood Disorders: Current Status. CNS Drugs. 2021, 35, 527–543. [Google Scholar] [CrossRef]
- Jesus-Nunes, A.P.; Leal, G.C.; Correia-Melo, F.S.; Vieira, F.; Mello, R.P.; Caliman-Fontes, A.T.; Echegaray, M.V.F.; Marback, R.F.; Guerreiro-Costa, L.N.F.; Souza-Marques, B.; et al. Clinical predictors of depressive symptom remission and response after racemic ketamine and esketamine infusion in treatment-resistant depression. Hum. Psychopharmacol. Clin. Exp. 2022, 37, e2836. [Google Scholar] [CrossRef]
- Abbar, M.; Demattei, C.; El-Hage, W.; Llorca, P.M.; Samalin, L.; Demaricourt, P.; Gaillard, R.; Courtet, P.; Vaiva, G.; Gorwood, P.; et al. Ketamine for the acute treatment of severe suicidal ideation: Double blind, randomised placebo controlled trial. BMJ 2022, 376, e067194. [Google Scholar] [CrossRef] [PubMed]
- Hochschild, A.; Keilp, J.G.; Madden, S.P.; Burke, A.K.; Mann, J.J.; Grunebaum, M.F. Ketamine vs midazolam: Mood improvement reduces suicidal ideation in depression. J. Affect. Disord. 2022, 300, 10–16. [Google Scholar] [CrossRef] [PubMed]
- Pochwat, B.; Nowak, G.; Szewczyk, B. An update on NMDA antagonists in depression. Expert Rev. Neurother. 2019, 19, 1055–1067. [Google Scholar] [CrossRef] [PubMed]
- Satarker, S.; Bojja, S.L.; Gurram, P.C.; Mudgal, J.; Arora, D.; Nampoothiri, M. Astrocytic Glutamatergic Transmission and Its Implications in Neurodegenerative Disorders. Cells. 2022, 11, 1139. [Google Scholar] [CrossRef]
- Chen, Q.Y.; Li, X.H.; Zhuo, M. NMDA receptors and synaptic plasticity in the anterior cingulate cortex. Neuropharmacology 2021, 197, 108749. [Google Scholar] [CrossRef]
- Micheli, P.; Ribeiro, R.; Giorgetti, A. A Mechanistic Model of NMDA and AMPA Receptor-Mediated Synaptic Transmission in Individual Hippocampal CA3-CA1 Synapses: A Computational Multiscale Approach. Int. J. Mol. Sci. 2021, 22, 1536. [Google Scholar] [CrossRef]
- Purkey, A.M.; Dell’Acqua, M.L. Phosphorylation-Dependent Regulation of Ca2+-Permeable AMPA Receptors During Hippocampal Synaptic Plasticity. Front. Synaptic Neurosci. 2020, 12, 8. [Google Scholar] [CrossRef]
- Lira, M.; Mira, R.G.; Carvajal, F.J.; Zamorano, P.; Inestrosa, N.C.; Cerpa, W. Glutamatergic Receptor Trafficking and Delivery: Role of the Exocyst Complex. Cells. 2020, 9, 2402. [Google Scholar] [CrossRef]
- Yang, X.; Gong, R.; Qin, L.; Bao, Y.; Fu, Y.; Gao, S.; Yang, H.; Ni, J.; Yuan, T.F.; Lu, W. Trafficking of NMDA receptors is essential for hippocampal synaptic plasticity and memory consolidation. Cell Rep. 2022, 40, 111217. [Google Scholar] [CrossRef]
- Wu, Q.L.; Gao, Y.; Li, J.T.; Ma, W.Y.; Chen, N.H. The Role of AMPARs Composition and Trafficking in Synaptic Plasticity and Diseases. Cell. Mol. Neurobiol. 2021. [Google Scholar] [CrossRef]
- Sumi, T.; Harada, K. Mechanism underlying hippocampal long-term potentiation and depression based on competition between endocytosis and exocytosis of AMPA receptors. Sci. Rep. 2020, 10, 14711. [Google Scholar] [CrossRef] [PubMed]
- Feng, Z.; Chen, X.; Zeng, M.; Zhang, M. Phase separation as a mechanism for assembling dynamic postsynaptic density signalling complexes. Curr. Opin. Neurobiol. 2019, 57, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Sathler, M.F.; Khatri, L.; Roberts, J.P.; Schmidt, I.G.; Zaytseva, A.; Kubrusly, R.C.C.; Ziff, E.B.; Kim, S. Phosphorylation of the AMPA receptor subunit GluA1 regulates clathrin-mediated receptor internalization. J. Cell Sci. 2021, 134, jcs257972. [Google Scholar] [CrossRef] [PubMed]
- Giese, K.P. The role of CaMKII autophosphorylation for NMDA receptor-dependent synaptic potentiation. Neuropharmacology 2021, 193, 108616. [Google Scholar] [CrossRef] [PubMed]
- Pandey, S.; Ramsakha, N.; Sharma, R.; Gulia, R.; Ojha, P.; Lu, W.; Bhattacharyya, S. The post-synaptic scaffolding protein tamalin regulates ligand-mediated trafficking of metabotropic glutamate receptors. J. Biol. Chem. 2020, 295, 8575–8588. [Google Scholar] [CrossRef]
- Zhang, Y.; Matt, L.; Patriarchi, T.; Malik, Z.A.; Chowdhury, D.; Park, D.K.; Renieri, A.; Ames, J.B.; Hell, J.W. Capping of the N-terminus of PSD-95 by calmodulin triggers its postsynaptic release. EMBO J. 2014, 33, 1341–1353. [Google Scholar] [CrossRef]
- Thorsen., T.S.; Madsen, K.L.; Rebola, N.; Rathje, M.; Anggono, V.; Bach, A.; Moreira, I.S.; Stuhr-Hansen, N.; Dyhring, T.; Peters, D.; et al. Identification of a small-molecule inhibitor of the PICK1 PD domain that inhibits hippocampal LTP and LTD. Proc. Natl. Acad. Sci. USA 2010, 107, 413–418. [Google Scholar] [CrossRef]
- Bockaert, J.; Perroy, J.; Ango, F. The Complex Formed by Group I Metabotropic Glutamate Receptor (mGluR) and Homer1a Plays a Central Role in Metaplasticity and Homeostatic Synaptic Scaling. J. Neurosci. 2021, 41, 5567–5578. [Google Scholar] [CrossRef]
- Fukaya, M.; Sugawara, T.; Hara, Y.; Itakura, M.; Watanabe, M.; Sakagami, H. BRAG2a Mediates mGluR-Dependent AMPA Receptor Internalization at Excitatory Postsynapses through the Interaction with PSD-95 and Endophilin 3. J. Neurosci. 2020, 40, 4277–4296. [Google Scholar] [CrossRef]
- Kaizuka, T.; Takumi, T. Postsynaptic density proteins and their involvement in neurodevelopmental disorders. J. Biochem. 2018, 163, 447–455. [Google Scholar] [CrossRef]
- Tao, C.L.; Liu, Y.T.; Sun, R.; Zhang, B.; Qi, L.; Shivakoti, S.; Tian, C.L.; Zhang, P.; Lau, P.M.; Zhou, Z.H.; et al. Differentiation and Characterization of Excitatory and Inhibitory Synapses by Cryo-electron Tomography and Correlative Microscopy. J. Neurosci. 2018, 38, 1493–1510. [Google Scholar] [CrossRef] [PubMed]
- Tomasetti, C.; Iasevoli, F.; Buonaguro, E.F.; De Berardis, D.; Fornaro, M.; Fiengo, A.L.; Martinotti, G.; Orsolini, L.; Valchera, A.; Di Giannantonio, M.; et al. Treating the Synapse in Major Psychiatric Disorders: The Role of Postsynaptic Density Network in Dopamine-Glutamate Interplay and Psychopharmacologic Drugs Molecular Actions. Int. J. Mol. Sci. 2017, 18, 135. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, B.; Coba, M.P. Molecular architecture of postsynaptic Interactomes. Cell. Signal. 2020, 76, 109782. [Google Scholar] [CrossRef]
- Moraes, B.J.; Coelho, P.; Fão, L.; Ferreira, I.L.; Rego, A.C. Modified Glutamatergic Postsynapse in Neurodegenerative Disorders. Neuroscience 2021, 454, 116–139. [Google Scholar] [CrossRef]
- Baucum, A.J., 2nd. Proteomic Analysis of Postsynaptic Protein Complexes Underlying Neuronal Plasticity. ACS Chem. Neurosci. 2017, 8, 689–701. [Google Scholar] [CrossRef]
- Gray, A.L.; Hyde, T.M.; Deep-Soboslay, A.; Kleinman, J.E.; Sodhi, M.S. Sex differences in glutamate receptor gene expression in major depression and suicide. Mol. Psychiatry 2015, 20, 1057–1068. [Google Scholar] [CrossRef] [PubMed]
- Sanz-Clemente, A.; Nicoll, R.A.; Roche, K.W. Diversity in NMDA receptor composition: Many regulators, many consequences. Neuroscientist 2013, 19, 62–75. [Google Scholar] [CrossRef]
- Ryan, T.J.; Emes, R.D.; Grant, S.G.; Komiyama, N.H. Evolution of NMDA receptor cytoplasmic interaction domains: Implications for organisation of synaptic signalling complexes. BMC Neurosci. 2008, 9, 6. [Google Scholar] [CrossRef]
- Petrillil, M.A.; Kranz, T.M.; Kleinhaus, K.; Joe, P.; Getz, M.; Johnson, P.; Chao, M.V.; Malaspina, D. The Emerging Role for Zinc in Depression and Psychosis. Front. Pharmacol. 2017, 8, 414. [Google Scholar] [CrossRef]
- Sindreu, C.; Bayés, Á.; Altafaj, X.; Pérez-Clausell, J. Zinc transporter-1 concentrates at the postsynaptic density of hippocampal synapses. Mol Brain. 2014, 7, 16. [Google Scholar] [CrossRef] [Green Version]
- Doboszewska, U.; Szewczyk, B.; Sowa-Kućma, M.; Młyniec, K.; Rafało, A.; Ostachowicz, B.; Lankosz, M.; Nowak, G. Antidepressant activity of fluoxetine in the zinc deficiency model in rats involves the NMDA receptor complex. Behav. Brain Res. 2015, 287, 323–330. [Google Scholar] [CrossRef] [PubMed]
- Rafalo-Ulinska, A.; Piotrowska, J.; Kryczyk, A.; Opoka, W.; Sowa-Kucma, M.; Misztak, P.; Rajkowska, G.; Stockmeier, C.A.; Datka, W.; Nowak, G.; et al. Zinc transporters protein level in postmortem brain of depressed subjects and suicide victims. J. Psychiatr. Res. 2016, 83, 220–229. [Google Scholar] [CrossRef] [PubMed]
- Chater, T.E.; Goda, Y. The role of AMPA receptors in postsynaptic mechanisms of synaptic plasticity. Front. Cell. Neurosci. 2014, 8, 401. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Chen, S.; Swensen, A.C.; Qian, W.J.; Gouaux, E. Architecture and subunit arrangement of native AMPA receptors elucidated by cryo-EM. Science 2019, 364, 355–362. [Google Scholar] [CrossRef]
- Swanson, G.T.; Kamboj, S.K.; Cull-Candy, S.G. Single-channel properties of recombinant AMPA receptors depend on RNA editing, splice variation, and subunit composition. J. Neurosci. 1997, 17, 58–69. [Google Scholar] [CrossRef]
- Man, H.Y. GluA2-lacking, calcium-permeable AMPA receptors—Inducers of plasticity? Curr. Opin. Neurobiol. 2011, 21, 291–298. [Google Scholar] [CrossRef]
- Kawahara, Y.; Ito, K.; Sun, H.; Kanazawa, I.; Kwak, S. w editing efficiency of GluR2 mRNA is associated with a low relative abundance of ADAR2 mRNA in white matter of normal human brain. Eur. J. Neurosci. 2003, 18, 23–33. [Google Scholar] [CrossRef]
- Guo, C.; Ma, Y.Y. Calcium Permeable-AMPA Receptors and Excitotoxicity in Neurological Disorders. Front. Neural. Circuits 2021, 15, 711564. [Google Scholar] [CrossRef]
- Schwenk, J.; Baehrens, D.; Haupt, A.; Bildl, W.; Boudkkazi, S.; Roeper, J.; Fakler, B.; Schulte, U. Regional diversity and developmental dynamics of the AMPA-receptor proteome in the mammalian brain. Neuron 2014, 84, 41–54. [Google Scholar] [CrossRef]
- Kamalova, A.; Nakagawa, T. AMPA receptor structure and auxiliary subunits. J. Physiol. 2021, 599, 453–469. [Google Scholar] [CrossRef]
- Greger, I.H.; Watson, J.F.; Cull-Candy, S.G. Structural and Functional Architecture of AMPA-Type Glutamate Receptors and Their Auxiliary Proteins. Neuron 2017, 94, 713–730. [Google Scholar] [CrossRef] [PubMed]
- Laje, G.; Paddock, S.; Manji, H.; Rush, A.J.; Wilson, A.F.; Charney, D.; McMahon, F.J. Genetic markers of suicidal ideation emerging during citalopram treatment of major depression. Am. J. Psychiatry 2007, 164, 1530–1538. [Google Scholar] [CrossRef] [PubMed]
- Smart, T.G.; Paoletti, P. Synaptic neurotransmitter-gated receptors. Cold Spring Harb. Perspect. Biol. 2012, 4, a009662. [Google Scholar] [CrossRef] [PubMed]
- Shor, C.; Zuo, W.; Eloy, J.D.; Ye, J.H. The Emerging Role of LHb CaMKII in the Comorbidity of Depressive and Alcohol Use Disorders. Int. J. Mol. Sci. 2020, 21, 8123. [Google Scholar] [CrossRef]
- Proietti Onori, M.; van Woerden, G.M. Role of calcium/calmodulin-dependent kinase 2 in neurodevelopmental disorders. Brain Res. Bull. 2021, 171, 209–220. [Google Scholar] [CrossRef]
- Terbeck, S.; Akkus, F.; Chesterman, L.P.; Hasler, G. The role of metabotropic glutamate receptor 5 in the pathogenesis of mood disorders and addiction: Combining preclinical evidence with human Positron Emission Tomography (PET) studies. Front. Neurosci. 2015, 9, 86. [Google Scholar] [CrossRef]
- Nicoletti, F.; Bockaert, J.; Collingridge, G.L.; Conn, P.J.; Ferraguti, F.; Schoepp, D.D.; Wroblewski, J.T.; Pin, J.P. Metabotropic glutamate receptors: From the workbench to the bedside. Neuropharmacology 2011, 60, 1017–1041. [Google Scholar] [CrossRef]
- Bhattacharyya, S. Inside story of Group I Metabotropic Glutamate Receptors (mGluRs). Int. J. Biochem. Cell Biol. 2016, 77, 205–212. [Google Scholar] [CrossRef]
- Kim, J.H.; Marton, J.; Ametamey, S.M.; Cumming, P.A. Review of Molecular Imaging of Glutamate Receptors. Molecules 2020, 25, 4749. [Google Scholar] [CrossRef]
- Koehl, A.; Hu, H.; Feng, D.; Sun, B.; Zhang, Y.; Robertson, M.J.; Chu, M.; Kobilka, T.S.; Laeremans, T.; Steyaert, J.; et al. Structural insights into the activation of metabotropic glutamate receptors. Nature 2019, 566, 79–84. [Google Scholar] [CrossRef]
- Dickson, E.J.; Falkenburger, B.H.; Hille, B. Quantitative properties and receptor reserve of the IP(3) and calcium branch of G(q)-coupled receptor signaling. J. Gen. Physiol. 2013, 141, 521–535. [Google Scholar] [CrossRef] [PubMed]
- Jin, D.Z.; Xue, B.; Mao, L.M.; Wang, J.Q. Metabotropic glutamate receptor 5 upregulates surface NMDA receptor expression in striatal neurons via CaMKII. Brain Res. 2015, 1624, 414–423. [Google Scholar] [CrossRef] [PubMed]
- Laursen, L.; Karlsson, E.; Gianni, S.; Jemth, P. Functional interplay between protein domains in a supramodular structure involving the postsynaptic density protein PSD-95. J. Biol. Chem. 2020, 295, 1992–2000. [Google Scholar] [CrossRef]
- de Bartolomeis, A.; Latte, G.; Tomasetti, C.; Iasevoli, F. Glutamatergic postsynaptic density protein dysfunctions in synaptic plasticity and dendritic spines morphology: Relevance to schizophrenia and other behavioral disorders pathophysiology, and implications for novel therapeutic approaches. Mol. Neurobiol. 2014, 49, 484–511. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Nelson, C.D.; Li, X.; Winters, C.A.; Azzam, R.; Sousa, A.A.; Leapman, R.D.; Gainer, H.; Sheng, M.; Reese, T.S. PSD-95 is required to sustain the molecular organization of the postsynaptic density. J. Neurosci. 2011, 31, 6329–6338. [Google Scholar] [CrossRef] [PubMed]
- Nikonenko, I.; Boda, B.; Steen, S.; Knott, G.; Welker, E.; Muller, D. PSD-95 promotes synaptogenesis and multiinnervated spine formation through nitric oxide signaling. J. Cell Biol. 2008, 183, 1115–1127. [Google Scholar] [CrossRef] [PubMed]
- Somani, A.; Singh, A.K.; Gupta, B.; Nagarkoti, S.; Dalal, P.K.; Dikshit, M. Oxidative and Nitrosative Stress in Major Depressive Disorder: A Case Control Study. Brain Sci. 2022, 12, 144. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Rao, W.; Zhang, C.; Zhang, C.; Liu, M.D.; Han, F.; Yao, L.B.; Han, H.; Luo, P.; Su, N.; et al. Scaffolding protein Homer1a protects against NMDA-induced neuronal injury. Cell Death Dis. 2015, 6, e1843. [Google Scholar] [CrossRef]
- Shiraishi-Yamaguchi, Y.; Furuichi, T. The Homer family proteins. Genome Biol. 2007, 8, 206. [Google Scholar] [CrossRef]
- Wang, T.; Zhang, L.; Shi, C.; Wei, R.; Yin, C. Interaction of the Homer1 EVH1 domain and skeletal muscle ryanodine receptor. Biochem. Biophys. Res. Commun. 2019, 514, 720–725. [Google Scholar] [CrossRef]
- Clifton, N.E.; Trent, S.; Thomas, K.L.; Hall, J. Regulation and Function of Activity-Dependent Homer in Synaptic Plasticity. Mol Neuropsychiatry 2019, 5, 147–161. [Google Scholar] [CrossRef] [PubMed]
- Rao, S.; Siu, C.O.; Shi, M.; Zhang, J.; Lam, M.H.B.; Yu, M.; Wing, Y.K.; Waye, M.M.Y. Associations of Homer Scaffolding Protein 1 gene and psychological correlates with suicide attempts in Chinese: A pilot study of multifactorial risk model. Gene 2018, 679, 382–388. [Google Scholar] [CrossRef] [PubMed]
- Benedetti, F.; Poletti, S.; Locatelli, C.; Mazza, E.; Lorenzi, C.; Vitali, A.; Riberto, M.; Brioschi, S.; Vai, B.; Bollettini, I.; et al. A Homer 1 gene variant influences brain structure and function, lithium effects on white matter, and antidepressant response in bipolar disorder: A multimodal genetic imaging study. Prog. Neuropsychopharmacol. Biol. Psychiatry 2018, 81, 88–95. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, M.K.; Tang, C.; Verpelli, C.; Narayanan, R.; Stearns, M.H.; Xu, R.M.; Li, H.; Sala, C.; Hayashi, Y. The postsynaptic density proteins Homer and Shank form a polymeric network structure. Cell 2009, 137, 159–171. [Google Scholar] [CrossRef]
- Delling, J.P.; Boeckers, T.M. Comparison of SHANK3 deficiency in animal models: Phenotypes, treatment strategies, and translational implications. J. Neurodev. Disord. 2021, 13, 55. [Google Scholar] [CrossRef]
- Roussignol, G.; Ango, F.; Romorini, S.; Tu, J.C.; Sala, C.; Worley, P.F.; Bockaert, J.; Fagni, L. Shank expression is sufficient to induce functional dendritic spine synapses in aspiny neurons. J. Neurosci. 2005, 25, 3560–3570. [Google Scholar] [CrossRef] [PubMed]
- Arons, M.H.; Lee, K.; Thynne, C.J.; Kim, S.A.; Schob, C.; Kindler, S.; Montgomery, J.M.; Garner, C.C. Shank3 Is Part of a Zinc-Sensitive Signaling System That Regulates Excitatory Synaptic Strength. J. Neurosci. 2016, 36, 9124–9134. [Google Scholar] [CrossRef]
- Duffney, L.J.; Wei, J.; Cheng, J.; Liu, W.; Smith, K.R.; Kittler, J.T.; Yan, Z. Shank3 deficiency induces NMDA receptor hypofunction via an actin-dependent mechanism. J. Neurosci. 2013, 33, 15767–15778. [Google Scholar] [CrossRef]
- Ortiz, R.; Niciu, M.J.; Lukkahati, N.; Saligan, L.N.; Nugent, A.C.; Luckenbaugh, D.A.; Machado-Vieira, R.; Zarate, C.A., Jr. Shank3 as a potential biomarker of antidepressant response to ketamine and its neural correlates in bipolar depression. J. Affect. Disord. 2015, 172, 307–311. [Google Scholar] [CrossRef]
- Denayer, A.; Van Esch, H.; de Ravel, T.; Frijns, J.P.; Van Buggenhout, G.; Vogels, A.; Devriendt, K.; Geutjens, J.; Thiry, P.; Swillen, A. Neuropsychopathology in 7 Patients with the 22q13 Deletion Syndrome: Presence of Bipolar Disorder and Progressive Loss of Skills. Mol. Syndromol. 2012, 3, 14–20. [Google Scholar] [CrossRef] [Green Version]
- Chandley, M.J.; Szebeni, A.; Szebeni, K.; Crawford, J.D.; Stockmeier, C.A.; Turecki, G.; Kostrzewa, R.M.; Ordway, G.A. Elevated gene expression of glutamate receptors in noradrenergic neurons from the locus coeruleus in major depression. Int. J. Neuropsychopharmacol. 2014, 17, 1569–1578. [Google Scholar] [CrossRef]
- Toro, C.; Deakin, J.F. NMDA receptor subunit NRI and postsynaptic protein PSD-95 in hippocampus and orbitofrontal cortex in schizophrenia and mood disorder. Schizophr. Res. 2005, 80, 323–330. [Google Scholar] [CrossRef]
- Beneyto, M.; Meador-Woodruff, J.H. Lamina-specific abnormalities of NMDA receptor-associated postsynaptic protein transcripts in the prefrontal cortex in schizophrenia and bipolar disorder. Neuropsychopharmacology 2008, 33, 2175–2186. [Google Scholar] [CrossRef] [PubMed]
- Dean, B.; Gibbons, A.S.; Boer, S.; Uezato, A.; Meador-Woodruff, J.; Scarr, E.; McCullumsmith, R.E. Changes in cortical N-methyl-D-aspartate receptors and post-synaptic density protein 95 in schizophrenia, mood disorders and suicide. Aust. N. Z. J. Psychiatry 2016, 50, 275–283. [Google Scholar] [CrossRef] [PubMed]
- Feyissa, A.M.; Chandran, A.; Stockmeier, C.A.; Karolewicz, B. Reduced levels of NR2A and NR2B subunits of NMDA receptor and PSD-95 in the prefrontal cortex in major depression. Prog. Neuropsychopharmacol. Biol. Psychiatry 2009, 33, 70–75. [Google Scholar] [CrossRef] [PubMed]
- Karolewicz, B.; Szebeni, K.; Gilmore, T.; Maciag, D.; Stockmeier, C.A.; Ordway, G.A. Elevated levels of NR2A and PSD-95 in the lateral amygdala in depression. Int. J. Neuropsychopharmacol. 2009, 12, 143–153. [Google Scholar] [CrossRef]
- Sowa-Kućma, M.; Szewczyk, B.; Sadlik, K.; Piekoszewski, W.; Trela, F.; Opoka, W.; Poleszak, E.; Pilc, A.; Nowak, G. Zinc, magnesium and NMDA receptor alterations in the hippocampus of suicide victims. J. Affect. Disord. 2013, 151, 924–931. [Google Scholar] [CrossRef]
- Duric, V.; Banasr, M.; Stockmeier, C.A.; Simen, A.A.; Newton, S.S.; Overholser, J.C.; Jurjus, G.J.; Dieter, L.; Duman, R.S. Altered expression of synapse and glutamate related genes in post-mortem hippocampus of depressed subjects. Int. J. Neuropsychopharmacol. 2013, 16, 69–82. [Google Scholar] [CrossRef]
- Kristiansen, L.V.; Meador-Woodruff, J.H. Abnormal striatal expression of transcripts encoding NMDA interacting PSD proteins in schizophrenia, bipolar disorder and major depression. Schizophr. Res. 2005, 78, 87–93. [Google Scholar] [CrossRef]
- Karolewicz, B.; Szebeni, K.; Stockmeier, C.A.; Konick, L.; Overholser, J.C.; Jurjus, G.; Roth, B.L.; Ordway, G.A. Low nNOS protein in the locus coeruleus in major depression. J. Neurochem. 2004, 91, 1057–1066. [Google Scholar] [CrossRef] [Green Version]
- Loeb, E.; El Asmar, K.; Trabado, S.; Gressier, F.; Colle, R.; Rigal, A.; Martin, S.; Verstuyft, C.; Fève, B.; Chanson, P.; et al. Nitric Oxide Synthase activity in major depressive episodes before and after antidepressant treatment: Results of a large case-control treatment study. Psychol. Med. 2020, 52, 80–89. [Google Scholar] [CrossRef] [PubMed]
- Dmitrzak-Weglarz, M.; Szczepankiewicz, A.; Rybakowski, J.; Kapelski, P.; Bilska, K.; Skibinska, M.; Reszka, E.; Lesicka, M.; Jablonska, E.; Wieczorek, E.; et al. Expression Biomarkers of Pharmacological Treatment Outcomes in Women with Unipolar and Bipolar Depression. Pharmacopsychiatry 2021, 54, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Ding, Q.; Li, H.; Tian, X.; Shen, Z.; Wang, X.; Mo, F.; Huang, J.; Shen, H. Zinc and imipramine reverse the depression-like behavior in mice induced by chronic restraint stress. J. Affect. Disord. 2016, 197, 100–106. [Google Scholar] [CrossRef] [PubMed]
- Popova, D.; Ágústsdóttir, A.; Lindholm, J.; Mazulis, U.; Akamine, Y.; Castrén, E.; Karpova, N.N. Combination of fluoxetine and extinction treatments forms a unique synaptic protein profile that correlates with long-term fear reduction in adult mice. Eur. Neuropsychopharmacol. 2014, 24, 1162–1174. [Google Scholar] [CrossRef]
- Stan, T.L.; Sousa, V.C.; Zhang, X.; Ono, M.; Svenningsson, P. Lurasidone and fluoxetine reduce novelty-induced hypophagia and NMDA receptor subunit and PSD-95 expression in mouse brain. Eur. Neuropsychopharmacol. 2015, 25, 1714–1722. [Google Scholar] [CrossRef]
- Zuo, C.; Cao, H.; Ding, F.; Zhao, J.; Huang, Y.; Li, G.; Huang, S.; Jiang, H.; Jiang, Y.; Wang, F. Neuroprotective efficacy of different levels of high-frequency repetitive transcranial magnetic stimulation in mice with CUMS-induced depression: Involvement of the p11/BDNF/Homer1a signaling pathway. J. Psychiatr. Res. 2020, 125, 152–163. [Google Scholar] [CrossRef]
- Asim, M.; Hao, B.; Yang, Y.H.; Fan, B.F.; Xue, L.; Shi, Y.W.; Wang, X.G.; Zhao, H. Ketamine Alleviates Fear Generalization Through GluN2B-BDNF Signaling in Mice. Neurosci. Bull. 2020, 36, 153–164. [Google Scholar] [CrossRef]
- Luo, Y.; Yu, Y.; Zhang, M.; He, H.; Fan, N. Chronic administration of ketamine induces cognitive deterioration by restraining synaptic signaling. Mol. Psychiatry 2021, 26, 4702–4718. [Google Scholar] [CrossRef]
- Karisetty, B.C.; Maitra, S.; Wahul, A.B.; Musalamadugu, A.; Khandelwal, N.; Guntupalli, S.; Garikapati, R.; Jhansyrani, T.; Kumar, A.; Chakravarty, S. Differential effect of chronic stress on mouse hippocampal memory and affective behavior: Role of major ovarian hormones. Behav. Brain Res. 2017, 318, 36–44. [Google Scholar] [CrossRef]
- Nieoczym, D.; Socała, K.; Zelek-Molik, A.; Pieróg, M.; Przejczowska-Pomierny, K.; Szafarz, M.; Wyska, E.; Nalepa, I.; Wlaź, P. Anticonvulsant effect of pterostilbene and its influence on the anxiety- and depression-like behavior in the pentetrazol-kindled mice: Behavioral, biochemical, and molecular studies. Psychopharmacology 2021, 238, 3167–3181. [Google Scholar] [CrossRef]
- Bao, H.; Ran, P.; Zhu, M.; Sun, L.; Li, B.; Hou, Y.; Nie, J.; Shan, L.; Li, H.; Zheng, S.; et al. The Prefrontal Dectin-1/AMPA Receptor Signaling Pathway Mediates The Robust and Prolonged Antidepressant Effect of Proteo-β-Glucan from Maitake. Sci. Rep. 2016, 6, 28395. [Google Scholar] [CrossRef] [PubMed]
- Li, J.M.; Liu, L.L.; Su, W.J.; Wang, B.; Zhang, T.; Zhang, Y.; Jiang, C.L. Ketamine may exert antidepressant effects via suppressing NLRP3 inflammasome to upregulate AMPA receptors. Neuropharmacology 2019, 146, 149–153. [Google Scholar] [CrossRef] [PubMed]
- Park, M.J.; Seo, B.A.; Lee, B.; Shin, H.S.; Kang, M.G. Stress-induced changes in social dominance are scaled by AMPA-type glutamate receptor phosphorylation in the medial prefrontal cortex. Sci. Rep. 2018, 8, 15008. [Google Scholar] [CrossRef] [PubMed]
- Yi, E.S.; Oh, S.; Lee, J.K.; Leem, Y.H. Chronic stress-induced dendritic reorganization and abundance of synaptosomal PKA-dependent CP-AMPA receptor in the basolateral amygdala in a mouse model of depression. Biochem. Biophys. Res. Commun. 2017, 486, 671–678. [Google Scholar] [CrossRef]
- Yu, H.; Li, M.; Zhou, D.; Lv, D.; Liao, Q.; Lou, Z.; Shen, M.; Wang, Z.; Li, M.; Xiao, X.; et al. Vesicular glutamate transporter 1 (VGLUT1)-mediated glutamate release and membrane GluA1 activation is involved in the rapid antidepressant-like effects of scopolamine in mice. Neuropharmacology 2018, 131, 209–222. [Google Scholar] [CrossRef]
- Camargo, A.; Dalmagro, A.P.; de Souza, M.M.; Zeni, A.L.B.; Rodrigues, A.L.S. Ketamine, but not guanosine, as a prophylactic agent against corticosterone-induced depressive-like behavior: Possible role of long-lasting pro-synaptogenic signaling pathway. Exp. Neurol. 2020, 334, 113459. [Google Scholar] [CrossRef]
- Li, M.X.; Li, Q.; Sun, X.J.; Luo, C.; Li, Y.; Wang, Y.N.; Chen, J.; Gong, C.Z.; Li, Y.J.; Shi, L.P.; et al. Increased Homer1-mGluR5 mediates chronic stress-induced depressive-like behaviors and glutamatergic dysregulation via activation of PERK-eIF2α. Prog Neuropsychopharmacol. Biol. Psychiatry 2019, 95, 109682. [Google Scholar] [CrossRef]
- Shen, M.; Lv, D.; Liu, X.; Li, S.; Chen, Y.; Zhang, Y.; Wang, Z.; Wang, C. Essential roles of neuropeptide VGF regulated TrkB/mTOR/BICC1 signaling and phosphorylation of AMPA receptor subunit GluA1 in the rapid antidepressant-like actions of ketamine in mice. Brain Res. Bull. 2018, 143, 58–65. [Google Scholar] [CrossRef]
- Ju, L.; Yang, J.; Zhu, T.; Liu, P.; Yang, J. BDNF-TrkB signaling-mediated upregulation of Narp is involved in the antidepressant-like effects of (2R,6R)-hydroxynorketamine in a chronic restraint stress mouse model. BMC Psychiatry 2022, 22, 182. [Google Scholar] [CrossRef]
- Xu, X.; Wu, K.; Ma, X.; Wang, W.; Wang, H.; Huang, M.; Luo, L.; Su, C.; Yuan, T.; Shi, H.; et al. mGluR5-Mediated eCB Signaling in the Nucleus Accumbens Controls Vulnerability to Depressive-Like Behaviors and Pain After Chronic Social Defeat Stress. Mol. Neurobiol. 2021, 58, 4944–4958. [Google Scholar] [CrossRef]
- Bieler, M.; Hussain, S.; Daaland, E.S.B.; Mirrione, M.M.; Henn, F.A.; Davanger, S. Changes in concentrations of NMDA receptor subunit GluN2B, Arc and syntaxin-1 in dorsal hippocampus Schaffer collateral synapses in a rat learned helplessness model of depression. J. Comp. Neurol. 2021, 529, 3194–3205. [Google Scholar] [CrossRef] [PubMed]
- Dou, M.; Gong, A.; Liang, H.; Wang, Q.; Wu, Y.; Ma, A.; Han, L. Improvement of symptoms in a rat model of depression through combined zinc and folic acid administration via up-regulation of the Trk B and NMDA. Neurosci. Lett. 2018, 683, 196–201. [Google Scholar] [CrossRef] [PubMed]
- Li, S.X.; Han, Y.; Xu, L.Z.; Yuan, K.; Zhang, R.X.; Sun, C.Y.; Xu, D.F.; Yuan, M.; Deng, J.H.; Meng, S.Q.; et al. Uncoupling DAPK1 from NMDA receptor GluN2B subunit exerts rapid antidepressant-like effects. Mol. Psychiatry 2018, 23, 597–608. [Google Scholar] [CrossRef] [PubMed]
- Treccani, G.; Gaarn du Jardin, K.; Wegener, G.; Müller, H.K. Differential expression of postsynaptic NMDA and AMPA receptor subunits in the hippocampus and prefrontal cortex of the flinders sensitive line rat model of depression. Synapse 2016, 70, 471–474. [Google Scholar] [CrossRef]
- Pochwat, B.; Sowa-Kucma, M.; Kotarska, K.; Misztak, P.; Nowak, G.; Szewczyk, B. Antidepressant-like activity of magnesium in the olfactory bulbectomy model is associated with the AMPA/BDNF pathway. Psychopharmacology 2015, 232, 355–367. [Google Scholar] [CrossRef]
- Román-Albasini, L.; Díaz-Véliz, G.; Olave, F.A.; Aguayo, F.I.; García-Rojo, G.; Corrales, W.A.; Silva, J.P.; Ávalos, A.M.; Rojas, P.S.; Aliaga, E.; et al. Antidepressant-relevant behavioral and synaptic molecular effects of long-term fasudil treatment in chronically stressed male rats. Neurobiol. Stress 2020, 13, 100234. [Google Scholar] [CrossRef] [PubMed]
- Qiao, H.; An, S.C.; Xu, C.; Ma, X.M. Role of proBDNF and BDNF in dendritic spine plasticity and depressive-like behaviors induced by an animal model of depression. Brain Res. 2017, 1663, 29–37. [Google Scholar] [CrossRef]
- Guo, F.; Zhang, B.; Fu, Z.; Ma, Y.; Gao, Y.; Shen, F.; Huang, C.; Li, Y. The rapid antidepressant and anxiolytic-like effects of YY-21 involve enhancement of excitatory synaptic transmission via activation of mTOR signaling in the mPFC. Eur. Neuropsychopharmacol. 2016, 26, 1087–1098. [Google Scholar] [CrossRef]
- Zhou, M.; Liu, Z.; Yu, J.; Li, S.; Tang, M.; Zeng, L.; Wang, H.; Xie, H.; Peng, L.; Huang, H.; et al. Quantitative Proteomic Analysis Reveals Synaptic Dysfunction in the Amygdala of Rats Susceptible to Chronic Mild Stress. Neuroscience 2018, 376, 24–39. [Google Scholar] [CrossRef]
- Tamási, V.; Petschner, P.; Adori, C.; Kirilly, E.; Ando, R.D.; Tothfalusi, L.; Juhasz, G.; Bagdy, G. Transcriptional evidence for the role of chronic venlafaxine treatment in neurotrophic signaling and neuroplasticity including also Glutamatergic- and insulin-mediated neuronal processes. PLoS ONE 2014, 9, e113662. [Google Scholar] [CrossRef] [Green Version]
- Piva, A.; Caffino, L.; Padovani, L.; Pintori, N.; Mottarlini, F.; Sferrazza, G.; Paolone, G.; Fumagalli, F.; Chiamulera, C. The metaplastic effects of ketamine on sucrose renewal and contextual memory reconsolidation in rats. Behav. Brain Res. 2020, 379, 112347. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Ju, P.; Liu, H.; Wu, X.; Niu, Z.; Zhu, Y.; Zhang, C.; Fang, Y. Ifenprodil rapidly ameliorates depressive-like behaviors, activates mTOR signaling and modulates proinflammatory cytokines in the hippocampus of CUMS rats. Psychopharmacology 2020, 237, 1421–1433. [Google Scholar] [CrossRef] [PubMed]
- Gordillo-Salas, M.; Pilar-Cuéllar, F.; Auberson, Y.P.; Adell, A. Signaling pathways responsible for the rapid antidepressant-like effects of a GluN2A-preferring NMDA receptor antagonist. Transl. Psychiatry 2018, 8, 84. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Zhang, F.; You, Y.; Wang, H.; Yuan, S.; Wu, B.; Zhu, R.; Liu, D.; Yan, F.; Wang, Z. S-Ketamine Exerts Antidepressant Effects by Regulating Rac1 GTPase Mediated Synaptic Plasticity in the Hippocampus of Stressed Rats. Cell. Mol. Neurobiol. 2022. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.W.; Chen, Y.J.; Wu, H.F.; Chung, Y.J.; Lee, Y.C.; Li, C.T.; Lin, H.C. Ketamine ameliorates severe traumatic event-induced antidepressant-resistant depression in a rat model through ERK activation. Prog. Neuropsychopharmacol. Biol. Psychiatry 2019, 93, 102–113. [Google Scholar] [CrossRef]
- Yang, M.; Luo, C.H.; Zhu, Y.Q.; Liu, Y.C.; An, Y.J.; Iqbal, J.; Wang, Z.Z.; Ma, X.M. 7,8-Dihydroxy-4-methylcoumarin reverses depression model-induced depression-like behaviors and alteration of dendritic spines in the mood circuits. Psychoneuroendocrinology 2020, 119, 104767. [Google Scholar] [CrossRef]
- Szewczyk, B.; Pochwat, B.; Rafało, A.; Palucha-Poniewiera, A.; Domin, H.; Nowak, G. Activation of mTOR dependent signaling pathway is a necessary mechanism of antidepressant-like activity of zinc. Neuropharmacology 2015, 99, 517–526. [Google Scholar] [CrossRef]
- Zhang, K.; Yamaki, V.N.; Wei, Z.; Zheng, Y.; Cai, X. Differential regulation of GluA1 expression by ketamine and memantine. Behav. Brain Res. 2017, 316, 152–159. [Google Scholar] [CrossRef]
- Li, R.; Zhao, X.; Cai, L.; Gao, W.W. Up-regulation of GluR1 in paraventricular nucleus and greater expressions of synapse related proteins in the hypothalamus of chronic unpredictable stress-induced depressive rats. Physiol. Behav. 2017, 179, 451–457. [Google Scholar] [CrossRef]
- Hou, X.Y.; Hu, Z.L.; Zhang, D.Z.; Lu, W.; Zhou, J.; Wu, P.F.; Guan, X.L.; Han, Q.Q.; Deng, S.L.; Zhang, H.; et al. Rapid Antidepressant Effect of Hydrogen Sulfide: Evidence for Activation of mTORC1-TrkB-AMPA Receptor Pathways. Antioxid. Redox Signal. 2017, 27, 472–488. [Google Scholar] [CrossRef]
- Liu, C.Y.; Chen, J.B.; Liu, Y.Y.; Zhou, X.M.; Zhang, M.; Jiang, Y.M.; Ma, Q.Y.; Xue, Z.; Zhao, Z.Y.; Li, X.J.; et al. Saikosaponin D exerts antidepressant effect by regulating Homer1-mGluR5 and mTOR signaling in a rat model of chronic unpredictable mild stress. Chin Med. 2022, 17, 60. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.T.; Hsieh, C.P.; Lee, M.Y.; Chen, L.C.; Huang, C.M.; Chen, H.H.; Chan, M.H. Betaine prevents and reverses the behavioral deficits and synaptic dysfunction induced by repeated ketamine exposure in mice. Biomed. Pharmacother. 2021, 144, 112369. [Google Scholar] [CrossRef] [PubMed]
- Leem, Y.H.; Yoon, S.S.; Jo, S.A. Imipramine Ameliorates Depressive Symptoms by Blocking Differential Alteration of Dendritic Spine Structure in Amygdala and Prefrontal Cortex of Chronic Stress-Induced Mice. Biomol. Ther 2020, 28, 230–239. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Liu, X.L.; Mu, R.H.; Wu, Y.J.; Liu, B.B.; Geng, D.; Liu, Q.; Yi, L.T. Hippocampal BDNF signaling restored with chronic asiaticoside treatment in depression-like mice. Brain Res. Bull. 2015, 114, 62–69. [Google Scholar] [CrossRef]
- Zhuang, F.; Li, M.; Gao, X.; Wang, Y.; Wang, D.; Ma, X.; Ma, T.; Gu, S. The antidepressant-like effect of alarin is related to TrkB-mTOR signaling and synaptic plasticity. Behav. Brain Res. 2016, 313, 158–171. [Google Scholar] [CrossRef]
- Misrani, A.; Tabassum, S.; Wang, M.; Chen, J.; Yang, L.; Long, C. Citalopram prevents sleep-deprivation-induced reduction in CaMKII-CREB-BDNF signaling in mouse prefrontal cortex. Brain Res. Bull. 2020, 155, 11–18. [Google Scholar] [CrossRef]
- Yin, Y.; Qian, S.; Chen, Y.; Sun, Y.; Li, Y.; Yu, Y.; Li, J.; Wu, Z.; Yu, X.; Ge, R.; et al. Latent Sex Differences in CaMKII-nNOS Signaling That Underlie Antidepressant-Like Effects of Yueju-Ganmaidazao Decoction in the Hippocampus. Front. Behav. Neurosci. 2021, 15, 640258. [Google Scholar] [CrossRef]
- Zou, Z.; Huang, J.; Yang, Q.; Zhang, Y.; Xu, B.; Wang, P.; Chen, G. Repeated Yueju, But Not Fluoxetine, Induced Sustained Antidepressant Activity in a Mouse Model of Chronic Learned Helplessness: Involvement of CaMKII Signaling in the Hippocampus. Evid.-Based Complement. Alternat. Med. 2022, 2022, 1442578. [Google Scholar] [CrossRef]
- Misrani, A.; Tabassum, S.; Chen, X.; Tan, S.Y.; Wang, J.C.; Yang, L.; Long, C. Differential effects of citalopram on sleep-deprivation-induced depressive-like behavior and memory impairments in mice. Prog. Neuropsychopharmacol. Biol. Psychiatry 2019, 88, 102–111. [Google Scholar] [CrossRef]
- Sun, L.; Verkaik-Schakel, R.N.; Biber, K.; Plösch, T.; Serchov, T. Antidepressant treatment is associated with epigenetic alterations of Homer1 promoter in a mouse model of chronic depression. J. Affect. Disord. 2021, 279, 501–509. [Google Scholar] [CrossRef]
- Rafało-Ulińska, A.; Poleszak, E.; Szopa, A.; Serefko, A.; Rogowska, M.; Sowa, I.; Wójciak, M.; Muszyńska, B.; Krakowska, A.; Gdula-Argasińska, J.; et al. Imipramine Influences Body Distribution of Supplemental Zinc Which May Enhance Antidepressant Action. Nutrients 2020, 12, 2529. [Google Scholar] [CrossRef] [PubMed]
- Gordillo-Salas, M.; Pascual-Antón, R.; Ren, J.; Greer, J.; Adell, A. Antidepressant-Like Effects of CX717, a Positive Allosteric Modulator of AMPA Receptors. Mol. Neurobiol. 2020, 57, 3498–3507. [Google Scholar] [CrossRef] [PubMed]
- Luoni, A.; Macchi, F.; Papp, M.; Molteni, R.; Riva, M.A. Lurasidone exerts antidepressant properties in the chronic mild stress model through the regulation of synaptic and neuroplastic mechanisms in the rat prefrontal cortex. Int. J. Neuropsychopharmacol. 2014, 18, pyu061. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.; Meng, X.; Zhai, Y.; Ye, T.; Zhou, P.; Nan, F.; Sun, G.; Sun, X. Antidepressant-like effects of the Guanxin Danshen formula via mediation of the CaMK II-CREB-BDNF signalling pathway in chronic unpredictable mild stress-induced depressive rats. Ann. Transl. Med. 2019, 7, 564. [Google Scholar] [CrossRef] [PubMed]
- Palmfeldt, J.; Henningsen, K.; Eriksen, S.A.; Müller, H.K.; Wiborg, O. Protein biomarkers of susceptibility and resilience to stress in a rat model of depression. Mol. Cell. Neurosci. 2016, 74, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Yoshino, Y.; Ochi, S.; Yamazaki, K.; Nakata, S.; Abe, M.; Mori, Y.; Ueno, S. Antidepressant action via the nitric oxide system: A pilot study in an acute depressive model induced by arginin. Neurosci. Lett. 2015, 599, 69–74. [Google Scholar] [CrossRef]
- Mishra, S.K.; Hidau, M.K.; Rai, S. Memantine treatment exerts an antidepressant-like effect by preventing hippocampal mitochondrial dysfunction and memory impairment via upregulation of CREB/BDNF signaling in the rat model of chronic unpredictable stress-induced depression. Neurochem. Int. 2021, 142, 104932. [Google Scholar] [CrossRef]
- Benske, T.M.; Mu, T.W.; Wang, Y.J. Protein quality control of N-methyl-D-aspartate receptors. Front. Cell. Neurosci. 2022, 16, 907560. [Google Scholar] [CrossRef]
- Paoletti, P.; Bellone, C.; Zhou, Q. NMDA receptor subunit diversity: Impact on receptor properties, synaptic plasticity and disease. Nat. Rev. Neurosci. 2013, 14, 383–400. [Google Scholar] [CrossRef]
- Lee, C.H.; Lü, W.; Michel, J.C.; Goehring, A.; Du, J.; Song, X.; Gouaux, E. NMDA receptor structures reveal subunit arrangement and pore architecture. Nature 2014, 511, 191–197. [Google Scholar] [CrossRef] [Green Version]
- Wyllie, D.J.; Livesey, M.R.; Hardingham, G.E. Influence of GluN2 subunit identity on NMDA receptor function. Neuropharmacology 2013, 74, 4–17. [Google Scholar] [CrossRef] [PubMed]
- Mellone, M.; Pelucchi, S.; Alberti, L.; Genazzani, A.A.; Di Luca, M.; Gardoni, F. Zinc transporter-1: A novel NMDA receptor-binding protein at the postsynaptic density. J. Neurochem. 2015, 132, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Babaei, P. NMDA and AMPA receptors dysregulation in Alzheimer’s disease. Eur. J. Pharmacol. 2021, 908, 174310. [Google Scholar] [CrossRef]
- Suryavanshi, P.S.; Ugale, R.R.; Yilmazer-Hanke, D.; Stairs, D.J.; Dravid, S.M. GluN2C/GluN2D subunit-selective NMDA receptor potentiator CIQ reverses MK-801-induced impairment in prepulse inhibition and working memory in Y-maze test in mice. Br. J. Pharmacol. 2014, 171, 799–809. [Google Scholar] [CrossRef] [PubMed]
- Stachowicz, K. Is PSD-95 entangled in the side effects of antidepressants? Neurochem. Int. 2022, 159, 105391. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Kim, Y.K. Animal models for the study of depressive disorder. CNS Neurosci Ther. 2021, 27, 633–642. [Google Scholar] [CrossRef]
- Planchez, B.; Surget, A.; Belzung, C. Animal models of major depression: Drawbacks and challenges. J. Neural Transm. 2019, 126, 1383–1408. [Google Scholar] [CrossRef]
- Zhang, J.; Huang, X.Y.; Ye, M.L.; Luo, C.X.; Wu, H.Y.; Hu, Y.; Zhou, Q.G.; Wu, D.L.; Zhu, L.J.; Zhu, D.Y. Neuronal nitric oxide synthase alteration accounts for the role of 5-HT1A receptor in modulating anxiety-related behaviors. J. Neurosci. 2010, 30, 2433–2441. [Google Scholar] [CrossRef]
- Wiggins, A.K.; Shen, P.J.; Gundlach, A.L. Neuronal-NOS adaptor protein expression after spreading depression: Implications for NO production and ischemic tolerance. J. Neurochem. 2003, 87, 1368–1380. [Google Scholar] [CrossRef]
- Jaso, B.A.; Niciu, M.J.; Iadarola, N.D.; Lally, N.; Richards, E.M.; Park, M.; Ballard, E.D.; Nugent, A.C.; Machado-Vieira, R.; Zarate, C.A. Therapeutic Modulation of Glutamate Receptors in Major Depressive Disorder. Curr. Neuropharmacol. 2017, 15, 57–70. [Google Scholar] [CrossRef] [Green Version]
- Berman, R.M.; Cappiello, A.; Anand, A.; Oren, D.A.; Heninger, G.R.; Charney, D.S.; Krystal, J.H. Antidepressant effects of ketamine in depressed patients. Biol. Psychiatry 2000, 47, 351–354. [Google Scholar] [CrossRef]
- Zarate, C.A., Jr.; Singh, J.B.; Carlson, P.J.; Brutsche, N.E.; Ameli, R.; Luckenbaugh, D.A.; Charney, D.S.; Manji, H.K. A randomized trial of an N-methyl-D-aspartate antagonist in treatment-resistant major depression. Arch. Gen. Psychiatry 2006, 63, 856–864. [Google Scholar] [CrossRef] [PubMed]
- Wan, L.B.; Levitch, C.F.; Perez, A.M.; Brallier, J.W.; Iosifescu, D.V.; Chang, L.C.; Foulkes, A.; Mathew, S.J.; Charney, D.S.; Murrough, J.W. Ketamine safety and tolerability in clinical trials for treatment-resistant depression. J. Clin. Psychiatry 2015, 76, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Kheirabadi, D.; Kheirabadi, G.R.; Mirlohi, Z.; Tarrahi, M.J.; Norbaksh, A. Comparison of Rapid Antidepressant and Antisuicidal Effects of Intramuscular Ketamine, Oral Ketamine, and Electroconvulsive Therapy in Patients with Major Depressive Disorder: A Pilot Study. J. Clin. Psychopharmacol 2020, 40, 588–593. [Google Scholar] [CrossRef] [PubMed]
- Lara, D.R.; Bisol, L.W.; Munari, L.R. Antidepressant, mood stabilizing and procognitive effects of very low dose sublingual ketamine in refractory unipolar and bipolar depression. Int. J. Neuropsychopharmacol. 2013, 16, 2111–2117. [Google Scholar] [CrossRef] [PubMed]
- Yavi, M.; Lee, H.; Henter, I.D.; Park, L.T.; Zarate, C.A., Jr. Ketamine treatment for depression: A review. Discov. Ment. Health 2022, 2, 9. [Google Scholar] [CrossRef]
- Hsu, T.W.; Chu, C.S.; Ching, P.Y.; Chen, G.W.; Pan, C.C. The efficacy and tolerability of memantine for depressive symptoms in major mental diseases: A systematic review and updated meta-analysis of double-blind randomized controlled trials. J. Affect. Disord. 2022, 306, 182–189. [Google Scholar] [CrossRef]
- Krzystanek, M.; Surma, S.; Pałasz, A.; Romańczyk, M.; Krysta, K. Possible Antidepressant Effects of Memantine-Systematic Review with a Case Study. Pharmaceuticals 2021, 14, 481. [Google Scholar] [CrossRef]
- Vasiliu, O. Investigational Drugs for the Treatment of Depression (Part 2): Glutamatergic, Cholinergic, Sestrin Modulators, and Other Agents. Front. Pharmacol. 2022, 13, 884155. [Google Scholar] [CrossRef]
- Poleszak, E.; Stasiuk, W.; Szopa, A.; Wyska, E.; Serefko, A.; Oniszczuk, A.; Wośko, S.; Świąder, K.; Wlaź, P. Traxoprodil, a selective antagonist of the NR2B subunit of the NMDA receptor, potentiates the antidepressant-like effects of certain antidepressant drugs in the forced swim test in mice. Metab. Brain Dis. 2016, 31, 803–814. [Google Scholar] [CrossRef] [Green Version]
- Preskorn, S.H.; Baker, B.; Kolluri, S.; Menniti, F.S.; Krams, M.; Landen, J.W. An innovative design to establish proof of concept of the antidepressant effects of the NR2B subunit selective N-methyl-D-aspartate antagonist, CP-101,606, in patients with treatment-refractory major depressive disorder. J. Clin. Psychopharmacol. 2008, 28, 631–637. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, L.; Diaz Granados, N.; Jolkovsky, L.; Brutsche, N.; Luckenbaugh, D.A.; Herring, W.J.; Potter, W.Z.; Zarate, C.A., Jr. A Randomized, placebo-controlled, crossover pilot trial of the oral selective NR2B antagonist MK-0657 in patients with treatment-resistant major depressive disorder. J. Clin. Psychopharmacol. 2012, 32, 551–557. [Google Scholar] [CrossRef] [PubMed]
- Stroebel, D.; Buhl, D.L.; Knafels, J.D.; Chanda, P.K.; Green, M.; Sciabola, S.; Mony, L.; Paoletti, P.; Pandit, J. A Novel Binding Mode Reveals Two Distinct Classes of NMDA Receptor GluN2B-selective Antagonists. Mol. Pharmacol. 2016, 89, 541–551. [Google Scholar] [CrossRef] [PubMed]
- Heresco-Levy, U.; Gelfin, G.; Bloch, B.; Levin, R.; Edelman, S.; Javitt, D.C.; Kremer, I. A randomized add-on trial of high-dose D-cycloserine for treatment-resistant depression. Int. J. Neuropsychopharmacol. 2013, 16, 501–506. [Google Scholar] [CrossRef] [PubMed]
- Pereira, V.S.; Joca, S.R.L.; Harvey, B.H.; Elfving, B.; Wegener, G. Esketamine and rapastinel, but not imipramine, have antidepressant-like effect in a treatment-resistant animal model of depression. Acta Neuropsychiatr. 2019, 31, 258–265. [Google Scholar] [CrossRef]
- Preskorn, S.; Macaluso, M.; Mehra, D.O.; Zammit, G.; Moskal, J.R.; Burch, R.M. GLYX-13 Clinical Study Group. Randomized proof of concept trial of GLYX-13, an N-methyl-D-aspartate receptor glycine site partial agonist, in major depressive disorder nonresponsive to a previous antidepressant agent. J. Psychiatr. Pract. 2015, 21, 140–149. [Google Scholar] [CrossRef]
- Pilar-Cuellar, F.; Castro, E.; Bretin, S.; Mocaer, E.; Pazos, Á.; Díaz, Á. S 47445 counteracts the behavioral manifestations and hippocampal neuroplasticity changes in bulbectomized mice. Prog. Neuropsychopharmacol. Biol. Psychiatry 2019, 93, 205–213. [Google Scholar] [CrossRef]
- Nations, K.R.; Dogterom, P.; Bursi, R.; Schipper, J.; Greenwald, S.; Zraket, D.; Gertsik, L.; Johnstone, J.; Lee, A.; Pande, Y.; et al. Examination of Org 26576, an AMPA receptor positive allosteric modulator, in patients diagnosed with major depressive disorder: An exploratory, randomized, double-blind, placebo-controlled trial. J. Psychopharmacol. 2012, 26, 1525–1539. [Google Scholar] [CrossRef] [PubMed]
| Postsynaptic Proteins | Controls [N] | Patients [N] | Samples/Brain Region | Methods | Findings | Authors’ Names |
|---|---|---|---|---|---|---|
| NMDA receptor complex | ||||||
| GluN1 GluN2A-D GluN3A | N = 19 (male = 18, female = 1) | MDD = 18 male | Locus Coeruleus (LC) Prefrontal Cortex (PFC) | qPCR | ↔GRIN1, GRIN2A mRNA in MDD-LC ↑GRIN2B mRNA in MDD-LC ↔GRIN1, GRIN2A, GRIN2B mRNA in MDD-PFC | Chandley et al. [113] |
| GluN1 | N = 15 | MDD = 15 BD = 15 | Prefrontal Cortex | Immunoautoradiography | ↔GluN1 protein in MDD and BD | Toro and Deakin [114] |
| GluN1 GluN2A-D | N = 15 | MDD = 15 BD = 15 | Prefrontal Cortex | In situ hybridization | ↓GRIN1 mRNA in MDD and BD ↓GRIN2A mRNA in MDD ↔GRIN2B-D mRNA in MDD ↔GRIN2A-D mRNA in BD | Beneyto and Meador-Woodruff [115] |
| GluN1 GluN2A-D GluN3A | N = 20 (male = 11, female = 9) N = 45 (male = 20, female = 25) | MDD = 10 (male = 5, female = 4) BD = 10 (male = 5, female = 5) Suicide = 14 (male = 8, female = 6) | Prefrontal cortex | In situ hybridization | ↑GRIN2D and ↔GRIN1, GRIN2A-C, and GRIN3A mRNA in MDD ↓GRIN2C and ↔GRIN1, GRIN2A, B, D, and GRIN3A mRNA in BD ↓GRIN2B and ↔GRIN1, GRIN2A, C, D and GRIN3A mRNA in Suicides | Dean et al. [116] |
| GluN1 GluN2A-B | N = 32 (male = 19, female = 13) MDD-non-suicide (MDD-NS) = 19 (male, female) | MDD = 53 (male = 26, female = 27) MDD-Suicide (MDD-S) = 34 (male, female) | Dorsolateral Prefrontal Cortex | qPCR | ↑GRIN2B, ↔GRIN1 and GRIN2A mRNA in MDD-S-both sexes ↑GRIN1, GRIN2A, GRIN2B mRNA in MDD-female ↑GRIN2B, ↔GRIN1 and GRIN2A mRNA in MDD-S-female ↔GRIN1, GRIN2A and GRIN2B mRNA in male MDD and MDD-S both sexes | Gray et al. [68] |
| GluN2A | N = 10 (male) | MDD = 10 (male) | Prefrontal Cortex | Western Blot | ↓GluN2A protein in MDD | Rafalo-Ulinska et al. [74] |
| GluN1 GluN2A-B | N = 14 (male = 11, female = 3) | MDD = 14 (male =11, female = 3; 8 male and 2 female were suicides) | Prefrontal Cortex | Western Blot | ↔GluN1 protein in MDD ↓GluN2A, GluN2B protein in MDD | Feyissa et al. [117] |
| GluN1 GluN2A-B | N = 14 (male = 13, female = 1) | MDD = 14 (male = 13, female = 1; 12 subjects were suicides) | Lateral amygdala | Western Blot | ↔GluN1, GluN2B protein in MDD ↑GluN2A protein in MDD | Karolewicz et al. [118] |
| GluN2A-B | N = 6 | Suicide victims = 17 | Hippocampus | Western Blot | ↑GluN2A protein in suicides ↓GluN2B protein in suicides | Sowa-Kućma et al. [119] |
| GluN1 GluN2A-D | N = 18 (female = 7, male = 11) | MDD = 21 (female = 8, male = 13) | Hippocampus: CA1 region (CA1) and Dentate gyrus (DG) | qPCR | ↔GRIN1, GRIN2A-D mRNA in CA1 and DG in MDD—both sexes | Duric et al. [120] |
| AMPA receptor complex | ||||||
| GluA1-2 | N = 19 (male = 18, female = 1) | MDD = 18 male | Locus Coeruleus (LC) Prefrontal cortex (PFC) | qPCR | ↔GRIA1, GRIA2 mRNA in MDD—LC and PFC | Chandley et al. [113] |
| GluA1-2 | N = 32 (male = 19, female = 13) MDD-non-suicide (MDD-NS) = 19 (male, female) | MDD = 53 (male = 26, female = 27) MDD-Suicide (MDD-S) = 34 (male, female) | Dorsolateral Prefrontal Cortex | qPCR | ↑GRIA2 and ↔GRIA1 mRNA in MDD—both sexes ↑GRIA2 and ↔GRIA1 mRNA in MDD-female ↔GRIA1 and GRIA2 mRNA in male MDD and MDD-S | Gray et al. [68] |
| GluA1 | N = 10 (male) | MDD = 10 (male) | Prefrontal Cortex | Western Blot | ↓GluA1 protein in MDD | Rafalo-Ulinska et al. [74] |
| GluA1-4 | N = 18 (female = 7, male = 11) | MDD = 21 (female = 8, male = 13) | Hippocampus: CA1 region (CA1) and Dentate gyrus (DG) | qPCR | ↓GRIA1 and GRIA3, ↔GRIA2 and GRIA4 mRNA in MDD-both sexes-CA1 and DG | Duric et al. [120] |
| Metabotropic glutamate receptor 5 | ||||||
| GluR5 | N = 32 (male = 19, female = 13) MDD-non-suicide (MDD-NS) = 19 (male, female) | MDD = 53 (male = 26, female = 27) MDD-Suicide (MDD-S) = 34 (male, female) | Dorsolateral Prefrontal Cortex | qPCR | ↑GRM5 mRNA in MDD-female ↓GRM5 mRNA in MDD—male ↔GRM5 mRNA in MDD-both sexes and MDD-S-both sexes | Gray et al. [68] |
| N = 19 (male = 18, female = 1) | MDD = 18 male | Locus Coeruleus (LC) Prefrontal Cortex (PFC) | qPCR | ↑GRM5 mRNA in MDD—LC ↔GRM5 mRNA in MDD—PFC | Chandley et al. [113] | |
| Postsynaptic density protein 95 | ||||||
| PSD-95 | N = 10 (male) | MDD = 10 (male) | Prefrontal Cortex | Western Blot | ↓PSD-95 protein in MDD | Rafalo-Ulinska et al. [74] |
| N = 14 (male = 11, female = 3) | MDD = 14 (male =11, female = 3; 8 male and 2 female were suicides) | Prefrontal Cortex | Western Blot | ↓PSD-95 protein level in MDD | Feyissa et al. [117] | |
| N = 14 (male = 13, female = 1) | MDD = 14 (male = 13, female = 1; 12 subjects were suicides) | Lateral amygdala | Western Blot | ↑PSD-95 protein in MDD | Karolewicz et al. [118] | |
| N = 6 | Suicide victims = 17 | Hippocampus | Western Blot | ↓PSD-95 protein in suicides | Sowa-Kućma et al. [119] | |
| N = 15 | MDD = 15 BD = 15 | Prefrontal Cortex | In situ hybridization | ↔DLG4 mRNA in MDD ↓DLG4 mRNA in BD | Kristiansen and Meador-Woodruff [121] | |
| N = 15 | MDD = 15 BD = 15 | Prefrontal Cortex | Immunoautoradiography | ↓PSD-95 protein in BD vs. MDD | Toro and Deakin [114] | |
| N = 15 | MDD = 15 BD = 15 | Prefrontal Cortex | In situ hybridization | ↔DLG4 mRNA in MDD and BD | Beneyto and Meador-Woodruff [115] | |
| N = 20 (male = 11, female = 9) N = 45 (male = 20, female = 25) | MDD = 10 (male = 5, female = 4) BD = 10 (male = 5, female = 5) Suicide = 14 (male = 8, female = 6) | Prefrontal cortex | Western Blot | ↔ PSD-95 protein in MDD and BD ↓PSD-95 protein in suicides | Dean et al. [116] | |
| Zinc transporters | ||||||
| ZnT-1 | N = 10 (male) N = 8 (male, female) | MDD = 10 (male) Suicide victims = 11 (male, female) | Prefrontal Cortex | Western Blot | ↑ZnT-1 protein level in MDD and suicides | Rafalo-Ulinska et al. [74] |
| Nitric oxide synthases | ||||||
| nNOS | N = 27 (controls) N = 27 (first- degree relatives) | MDD = 29 | Neutrophils (venous blood samples) | qPCR | ↑NOS1 mRNA in MDD ↔NOS1 mRNA in first-degree relatives | Somani et al. [99] |
| N = 14 (male = 13, female = 1) | MDD = 14 (male = 13, female = 1; 12 subjects were suicides) | Lateral amygdala | Western Blot | ↔nNOS protein in MDD | Karolewicz et al. [118] | |
| N = 12 (male = 10, female = 2) | MDD = 12 (male = 9, female = 3; 12 subjects were suicides) | Locus coeruleus (LC) Cerebellum (CER) | Western blot | ↓nNOS protein in MDD-LC ↔nNOS protein in MDD-CER | Karolewicz et al. [122] | |
| NOS | N = 895 | MDD = 460 (drug-free = 104; antidepressant group = 356: SSRI = 138; SNRI = 137; another = 81) | Blood plasma | LC-MS | ↓NOS activity (L-Citrulline/L-Arginine ratio) in whole MDD group ↓NOS activity in drug-free MDD ↑NOS activity in whole MDD group at 3 months and 6 months ↑NOS activity in MDD responders group at 3 months and 6 months ↔NOS activity in MDD non-responders group at 3 months and 6 months | Loeb et al. [123] |
| SH3 and multiple ankyrin repeat domains 3 | ||||||
| Shank3 | No control | MDD = 24 women BD = 32 women (All participants were treated with antidepressants and/or mood stabilizers of the 1st and 2nd generation) | Peripheral blood mononuclear cells (PBMCs) | Microarray | ↑SHANK3 mRNA in MDD after treatment | Dmitrzak-Weglarz et al. [124] |
| Postsynaptic Proteins | Species/Strain | Model/Treatment/Groups | Samples/Brain Region | Methods | Findings | Authors’ Names |
|---|---|---|---|---|---|---|
| NMDA receptor complex | ||||||
| NMDAR (no information on subunits) | ICR male mice |
| Hippocampus | qPCR | ↑NMDAR mRNA after CRS, CRS + Zn15, CRS + IMI5 and ↓NMDAR mRNA after CRS + Zn30, CRS + IMI20, and CRS + IMI5 + Zn15 | Ding et al. [125] |
| GluN2A | C57BL/6J male mice |
| Infralimbic Cortex (IL), Prelimbic cortex (PL), Basolateral Amygdala (BLA), Lateral Amygdala (LA), Central Amygdala (CEA) CA1 Strata Oriens (OR), CA1 Pyramidal (PYR), CA1 Radiatum (RAD) and CA1 Lacunosum-moleculare layers (LM) | Fluorescence Immunohistochemistry | ↔ GluN2A protein in all brain region after FC + FLU ↑GluN2A protein in BLA, LA, CEA, CA1 PYR after FC+FLU+EXT | Popova et al. [126] |
| GluN2A-B | C57Bl/6J male mice |
| Hippocampus (HP) Prefrontal Cortex (PFC) | Western Blot | ↓GluN2A, GluN2B protein after LUR10 and FLU20—HP and PFC | Stan et al. [127] |
| GluN2A-B | C57BL/6J male mice |
| Hippocampus (HP) Prefrontal Cortex (PFC) | Western Blot | ↓GluN2A, GluN2B and ↔GluN1 protein level after CUMS—HP ↔GluN1, GluN2A and GluN2B protein after CUMS—PFC ↔GluN2A protein and CUMS + 25 Hz HF-rTMS and ↔GluN2B protein after CUMS+15 Hz HF-rTMS and CUMS + 25 Hz HF-rTMS—HP ↔GluN1, GluN2A, GluN2B protein after CUMS+15 Hz HF-rTMS, CUMS + 25 Hz HF-rTMS | Zuo et al. [128] |
| GluN2A-B | C57BL/6J male mice |
| Basolateral Amygdala (BLA) Inferior-Limbic Prefrontal Cortex (IL-PFC) | Western Blot | ↔GluN2A and ↓GluN2B protein in BLA and IL-PFC after KET | Asim et al. [129] |
| pS1325-GluN2A pS-1303-GluN2B GluN2A-B GluN1 | C57BL/6J male mice |
| Hippocampus | Western Blot qPCR | ↔GluN1, GluN2A, GluN2B, protein after KET1, 3, 5, 10 ↓GluN2A, GluN2B and ↔GluN1 protein after KET28 ↓GluN2A and GluN2B protein after KET_10 KET_20 and KET_30 pS1325-GluN2A, pS-1303-GluN2B, GluN2A, GluN2B (total, surface, intracellular) protein after KET_30 ↓Grin2A, Grin2B and ↔ Grin1 mRNA after KET_30 | Luo et al. [130] |
| GluN2A-B | C57BL/6J female mice |
| Hippocampus | qPCR | ↑Grin2A and Grin2B mRNA after CUS Grin2A and Grin2B mRNA after OVX + CUS ↔Grin2A and ↑Grin2B mRNA after OVX + CUS + P ↑Grin2A and Grin2B mRNA after OVX + CUS+ E | Karisetty et al. [131] |
| GluN2B | Swiss male mice |
before each PTZ injection | Prefrontal Cortex (PFC) Hippocampus (HP) | qPCR | ↔Grin2B mRNA level after PTE100, PTE200 + PTZ—PFC ↓Grin2B mRNA after VPA + PTZ—PFC ↔Grin2B mRNA after VPA, PTE100, 200 + PTZ in HP | Nieoczym et al. [132] |
| AMPA receptor complex | ||||||
| GluA1-pS845 GluA1-3 | CD-1 male mice |
| Prefrontal Cortex—whole tissue lysate Prefrontal Cortex—synaptic fraction | Western Blot | After 60 min of injection: ↔GluA1 and ↑GluA1-pS845 protein after PGM5, 8, 12.5 and IMI ↔GluA2 and GluA3 protein after PGM5, 8, 12.5 and IMI After 5 days of injection: ↑GluA1 protein after PGM 8 and 12.5 and ↑GluA1-pS845protein after PGM5, 8, 12.5 and IMI ↔GluA2 and GluA3 protein after PGM5, 8, 12.5 and IMI After 60 min of injection: ↔GluA1 and ↑GluA1-pS845 protein after PGM5, 8, 12.5 and IMI ↔GluA2 and GluA3 protein after PGM5, 8, 12.5 and IMI After 5 days of injection: ↑GluA1 and GluA1-pS845 protein after PGM5, 8, 12.5 and IMI ↑GluA2 protein after PGM8, 12.5 and IMI ↑GluA3 protein after PGM5, 8, 12.5 and IMI | Bao et al. [133] |
| GluA1 | C57BL/6J male mice |
| Hippocampus | Western Blot | ↑GluA1 protein after LPS+KET and LPS +YVAdD ↓GluA1 protein level after LPS ↔GluA1 protein level after LPS+YVAdD+KET | Li et al. [134] |
| GluA1 GluA1-pS818 GluA1-pS831 GluA1-pS845 GluA2 GluA2-pS880 | C57BL/6J male mice |
| Medial Prefrontal Cortex | Western Blot | ↔GluA1, ↓GluA1-pS831 and GluA2, and ↔GluA2-pS880 protein after CRS ↔ GluA1-pS818, ↑GluA1-pS831, ↔GluA2 and ↑ GluA2-pS880 after CRS+FLU | Park et al. [135] |
| GluA1-2 | C57BL/6J male mice |
| Infralimbic Cortex (IL), Prelimbic cortex (PL), Basolateral Amygdala (BLA), Lateral Amygdala (LA), Central Amygdala (CEA) CA1 Strata Oriens (OR), CA1 Pyramidal (PYR), CA1 Radiatum (RAD) and CA1 Lacunosum-moleculare layers (LM) | Fluorescence Immunohistochemistry | ↓GluA1 protein in CA1 OR after FC+WAT+EXT ↓GluA1 protein in CA1 PYR after FC + FLU + EXT ↑GluA2 protein in all brain region after FC+WAT+EXT and FC+FLU+EXT | Popova et al. [126] |
| GluA1 GluA1-pS845 GluA2 | C57BL/6J male mice |
| Basolateral Amygdala | Western Blot | ↑GluA1 and GluA1-pS845, and ↓GluA2 protein after CRS ↑GluA1, ↓GluA1-pS845 and ↑GluA2 protein after CRS + FLU | Yi et al. [136] |
| GluA1 GluA1-pS845 | C57BL/6J male mice |
| Western Blot | ↓GluA1 and GluA1-pS845 protein after CUS ↑GluA1 and GluA1-pS845 after CUS + SCO 25 and 50 | Yu et al. [137] | |
| GluA1 | Swiss male mice |
| Hippocampus (HP) Prefrontal Cortex (PFC) | Western Blot | ↔ GluA1 protein in Hp after KET5, GUO5 ↓GluA1 protein in HP after CORT, CORT+GUO5 ↑GluA1 protein in HP after CORT+KET ↔GluA1 protein in PFC after KET5, GUO5, CORT, CORT +KET5, CORT + GUO5 ↔GluA1 protein in HP after FLU ↓GluA1 protein in HP after CORT ↓GluA1 protein in HP after CORT+FLU ↔GluA1 protein in PFC after FLU, CORT, CORT + FLU ↔GluA1 protein in HP after KET1+GUO5 ↓GluA1 protein in HP after CORT ↓GluA1 protein in HP after CORT+KET1+GUO5 ↔GluA1 protein in PFC after KET1 +GUO5, CORT, CORT+KET1+GUO5 | Camargo et al. [138] |
| GluA1-2 | C57BL/6J male mice |
| Hippocampus | Surface receptor cross-linking with BS3 and Western Blot | ↔GluA1, GluA2 protein in Susceptible and Resilient mice ↓GluA1, GluA2 protein after coss-linking in Susceptible mice | Li et al. [139] |
| GluA1-pS845 | C57BL/6J male mice |
| Prefrontal Cortex | Western Blot Fluorescence Immunohistochemistry | ↑GluA1-pS845 protein after NC shRNA+KET ↓GluA1-pS845 protein after VGF shRNA and VGF shRNA+KET ↑GluA1-pS845 protein after NC shRNA+KET ↓GluA1-pS845 protein after VGF shRNA and VGF shRNA+KET | Shen et al. [140] |
| GluA1-2 | C57BL/6J male mice |
| Hippocampus | Western Blot | ↓GluA1,GluA2 protein level after CRS, CRS+HNK, CRS+HNK+NBQX, CRS+HNK+ANA-12 | Ju et al. [141] |
| pS831-GluA1 pS880-GluA2 GluA1-2 | C57BL/6J male mice |
| Hippocampus | Western BlotqPCR | ↔GluA1,GluA2 protein after KET1, 3, 5, 10 and KET_30 ↓GluA1, GluA2 protein after KET_10, 20 and 30 ↓GluA1,GluA2 after KET.10, KET.20, KET.30 ↓p-S831-GluA1 and p-S880-GluA2 (total, surface, intracellular) protein after KET_30 ↓Gria1, Gria2 mRNA after KET_30 | Luo et al. [130] |
| Metabotropic glutamate receptor 5 | ||||||
| mGluR5 | C57BL/6J male mice |
| Nucleus Accumbens (NAc) | Western Blot | ↓mGluR5 protein in Susceptible mice ↑mGluR5 protein in Resilient mice | Xu et al. [142] |
| C57BL/6J male mice |
| Hippocampus | qPCR | ↑Grm5 mRNA in Susceptible mice ↑Grm5 mRNA after CRS | Li et al. [139] | |
| Postsynaptic Proteins | Species/Strain | Model/Treatment/Groups | Brain Region | Methods | Findings | Authors’ Names |
|---|---|---|---|---|---|---|
| NMDA receptor complex | ||||||
| GluN2A-B | Sprague Dawley male and female rats |
| Hippocampus | Western blot Post-embedding Immunogold | ↓GluN2A/GluN2B protein in LH compared to NLH ↑GluN2B protein in LH compared to WT ↔GluN2B protein in NLH compared to WT | Bieler et al. [143] |
| NMDAR (no information on subunits) | Sprague Dawley male rats |
| Frontal cortex | qPCR | ↓NMDAR mRNA after CUMS ↑NMDAR mRNA n after CUMS+ PAR, CUMS+ Zn+ FA, CUMS+ Zn+ FA+ PAR | Dou et al. [144] |
| GluN1 GluN2A pT1325-GluN2A GluN2B pS1303-GluN2B | Sprague Dawley male rats |
| Medial prefrontal cortex (mPFC) | Western blot | ↑GluN1, GluN2B, p-GluN2B and ↔ GluN2A, p-GluN2A protein after CUS ↔GluN2A, p-GluN2A, GluN2B and ↓p-GluN2B protein after CUS+ IFE ↑GluN1 protein after CUS+ IFE ↔GluN1 protein level after CUS+TC-DAPK 6 ↔GluN2B and ↓p-GluN2B protein level after CUS+TC-DAPK 6 ↔GluN1 protein level after CUS+ AAV-shDAPK1 ↔GluN2B protein after CUS+ AAV-shDAPK1 ↓p-GluN2B protein after CUS+ AAV-shDAPK1 | Li et al. [145] |
| GluN2A-B | Flinders Sensitive Line (FSL) male rats Flinders Resistant Line (FRL) male rats |
| Hippocampus (HP) Prefrontal cortex (PFC) | Western blot | ↓GluN2A and ↑GluN2B protein in FSL—HP ↔GluN2A and GluN2B protein in FSL—PFC | Treccani et al. [146] |
| GluN2A-B | Sprague Dawley male rats |
| Prefrontal cortex (PFC) Hippocampus (HP) Amygdala (AMY) |
Western blot qPCR | ↔GluN2A protein after OB in PFC vs. Sham ↔GluN2A protein after OB+ Mg15 in PFC vs. OB ↔GluN2B protein after OB in PFC vs. Sham ↔GluN2B protein after OB+ Mg15 in PFC vs. OB ↔GluN2A protein after OB in HP vs. Sham ↔GluN2A protein after OB+ Mg15 in HP vs. OB ↔GluN2B protein after OB in HP vs. Sham ↔GluN2B protein after OB+ Mg15 in HP vs. OB ↔GluN2A protein after OB in AMY vs. Sham ↔GluN2A protein after OB+ Mg15 in AMY vs. OB ↔GluN2B protein after OB in AMY vs. Sham ↑GluN2B protein after OB+ Mg15 in AMY vs. OB ↔Grin2A mRNA after OB in PFC vs. Sham ↔Grin2A mRNA after OB+ Mg15 in PFC vs. OB ↔Grin2B mRNA after OB in PFC vs. Sham ↑Grin2B mRNA after OB+ 15 Mg15 in PFC vs. OB ↓Grin2A mRNA after OB in HP vs. Sham ↔Grin2A mRNA after OB+ Mg15 in HP vs. OB ↓Grin2B mRNA after OB in HP vs. Sham ↑Grin2B mRNA after OB+ Mg15 in HP vs. OB | Pochwat et al. [147] |
| GluN1 GluN2A-B | Sprague Dawley male rats |
| Hippocampus | Western blot | ↑GluN1, GluN2A and GluN2B proteins after ZnD vs. ZnA ↓GluN1, GluN2A and GluN2B proteins after ZnD+ FLU vs. ZnD | Doboszewska et al. [73] |
| GluN2A-B | Sprague Dawley male rats |
| Hippocampus (whole tissue lysate) Hippocampus (synaptoneurosomes) | Western blot | ↓GluN2A and ↔GluN2B protein after CRS ↔ GluN2A, ↔GluN2B protein after CRS+ FAS ↔GluN2A, GluN2B protein after CRS ↔GluN2A, GluN2B protein after CRS+ FAS | Román-Albasini et al. [148] |
| GluN2B | Sprague Dawley male rats |
| Hippocampus | Western blot | ↔ GluN2B protein after CUMS ↑GluN2B protein after CUMS+BDNF ↓GluN2B protein after proBDNF | Qiao et al. [149] |
| GluN2B | Sprague Dawley male rats |
| Medial prefrontal cortex | Western blot | ↓GluN2B protein after CUMS ↑GluN2B protein after CUMS+ YY-21 ↑GluN2B protein after CUMS+ FLU | Guo et al. [150] |
| GluN2A-B | Sprague Dawley male rats |
| Amygdala | Western blot | ↑GluN2A and GluN2B protein after CMS | Zhou et al. [151] |
| GluN2A-B | Dark Agouti male rats |
| Frontal cortex | qPCR | ↑Grin2A and Grin2B mRNA after VLX | Tamási et al. [152] |
| GluN2B | Sprague Dawley male rats |
| Hippocampus (HP) Nucleus accumbens (NAc) Amygdala (AMY) | Western blot | ↓GluN2B protein after KET in HP ↑GluN2B protein after KET in NAc ↑GluN2B protein after KET in AMY | Piva et al. [153] |
| AMPA receptor complex | ||||||
| GluA1 | Sprague Dawley male rats |
| Hippocampus (HP) Medial prefrontal cortex (mPFC) | Western blot | ↓GluA1 protein after CUMS and ↑GluA1 protein after CUMS+ IFE in HP ↓GluA1 protein after CUMS and ↑GluA1 protein after CUMS+ IFE in mPFC | Yao et al. [154] |
| GluA1 | Sprague Dawley male rats |
| Medial prefrontal cortex | Western blot | ↑GluA1 protein after NVP-AAM077 (only 30 min after administration) | Gordillo-Salas et al. [155] |
| GluA1 | Sprague Dawley male rats |
| Medial prefrontal cortex | Western blot | ↓GluA1 protein after CUS ↑GluA1 protein after CUS+IFE | Li et al. [145] |
| GluA1 pS831-GluA1 pS845-GluA1 | Sprague Dawley male rats |
| Prefrontal cortex (PFC) Hippocampus (HP) Amygdala (AMY) | Western blot | ↔GluA1, pS831-GluA1, pS845-GluA1 proteins after OB in PFC vs. Sham ↔GluA1,↑pS831-GluA1 and pS845-GluA1 protein after OB+ Mg15 in PFC vs. OB ↔GluA1, pS831-GluA1 and ↑pS845-GluA1 protein after OB in HP vs. Sham ↔GluA1, pS831-GluA1 and ↓pS845-GluA1 protein after OB+ Mg15 in HP vs. OB ↔GluA1, pS831-GluA1 and pS845-GluA1 protein after OB in AMY vs. Sham ↔GluA1, pS831-GluA1 and pS845-GluA1 protein after OB+ Mg15 in AMY vs. OB | Pochwat et al. [147] |
| GluA1 pS831-GluA1 pS845-GluA1 GluA2 | Sprague Dawley male rats |
| Hippocampus (whole tissue lysate) Hippocampus (synaptoneurosomes) | Western blot | ↓GluA1 and ↔pS831-GluA1, pS845-GluA1, GluA2 protein level after CRS ↔GluA1, pS831-GluA1, pS845-GluA1, GluA2 protein after CRS+ FAS ↔GluA1, pS831-GluA1, pS845-GluA1, GluA2 protein after CRS ↔GluA1, pS845-GluA1, GluA2 and ↓pS831-GluA1 proteins after CRS+ FAS | Román-Albasini et al. [148] |
| GluA3 | Dark Agouti male rats |
| Frontal cortex | qPCR | ↑Gria3 mRNA after VLX | Tamási et al. [152] |
| GluA1 | Sprague Dawley male rats |
| Hippocampus | Western blot | ↓GluA1 protein after CUMS ↑GluA1 protein after CUMS+S-KET ↔GluA1 protein after CUMS+ NSC23766 + S-KET ↔GluA1 protein after CUMS + NSC23766 | Zhu et al. [156] |
| GluA1 | Sprague Dawley rats |
| Amygdala (AMY) Prefrontal cortex (PFC) | Western blot | ↑GluA1 protein after 3 CS-US in AMY ↔GluA1 protein after 6 CS-US in AMY ↑GluA1 protein after 10 CS-US in AMY ↓GluA1 protein after 10 CS-US+ KET in AMY ↔GluA1 protein after 10 CS-US + FLU in AMY ↓GluA1 protein after 3 CS-US in PFC ↓GluA1 protein after 6 CS-US in PFC ↓GluA1 protein after 10 CS-US in PFC ↑GluA1 protein after 10 CS-US+ KET in PFC ↔GluA1 protein after 10 CS-US + FLU in PFC | Lee et al. [157] |
| GluA1 | Sprague Dawley male rats |
| Hippocampus | Western blot | ↓GluA1 protein after CUMS ↑GluA1 protein after CUMS+ VLX ↑GluA1 protein after CUMS+ Dhmc | Yang et al. [158] |
| GluA1 pS845-GluA1 | Sprague Dawley male rats |
| Medial prefrontal cortex | Western blot | ↔GluA1 and ↓pS845-GluA1 proteins after CUMS ↔GluA1 and ↑pS845-GluA1 proteins after CUMS+ YY-21 ↔GluA1 and ↑pS845-GluA1 proteins after CUMS+ FLU | Guo et al. [150] |
| GluA1 | Sprague Dawley male rats |
| Prefrontal cortex | Western blot | ↔GluA1 protein 30 min after Zn5 treatment ↑GluA1 protein 3 h after Zn5 treatment ↑GluA1 protein 24h after Zn treatment | Szewczyk et al. [159] |
| GluA1 pS845-GluA1 | Sprague Dawley male and female rats |
| Hippocampus | Western blot | ↑GluA1 and pS845-GluA1 protein after KET ↔GluA1 and ↑pS845-GluA1 protein after MEM | Zhang et al. [160] |
| GluA1 | Sprague Dawley male rats |
| Hypothalamic paraventricular nucleus (hPVN) Hypothalamus (HY) | Western blot qPCR | ↑ GluA1 protein after CUS in hPVN ↑GluA1 protein and ↑Gria1 mRNA after CUS in HY | Li et al. [161] |
| GluA1-2 | Sprague Dawley male rats |
| Hippocampus | Western blot | ↓GluA1 and GluA2 protein after CUMS ↑GluA1 protein after CUMS+ NaHS ↑GluA2 protein after CUMS+ NaHS | Hou et al. [162] |
| GluA1 | Sprague Dawley male rats |
| Hippocampus (HP) Nucleus accumbens (NAc) Amygdala (AMY) | Western blot | ↓GluA1 protein after KET in HP ↓GluA1 protein after KET in NAc ↔GluA1 protein after KET in AMY | Piva et al. [153] |
| Metabotropic glutamate receptor 5 | ||||||
| mGluR5 | Sprague Dawley male rats |
| Hippocampus | Western blot Immunohistochemistry qPCR | ↓mGluR5 protein after CUMS ↑mGluR5 protein after CUMS+ FLU ↑mGluR5 protein after CUMS+ SSDH ↑mGluR5 protein after CUMS+ SSDL ↓mGluR5 protein after CUMS ↑mGluR5 protein after CUMS+ FLU ↑mGluR5 protein after CUMS+ SSDH ↑mGluR5 protein after CUMS+ SSDL ↓Grm5 mRNA after CUMS ↑Grm5mRNA after CUMS+ FLU ↑Grm5 mRNA after CUMS+ SSDH ↑Grm5 mRNA after CUMS+ SDDL | Liu et al. [163] |
| mGluR5 | Sprague Dawley male rats |
| Hippocampus (HP) Nucleus accumbens (NAc) Amygdala (AMY) | Western blot | ↓mGluR5 protein after KET in HP ↓mGluR5 protein after KET in NAc ↑mGluR5 protein after KET in AMY | Piva et al. [153] |
| Postsynaptic Proteins | Species/Strain | Model/Treatment/Groups | Brain Region | Methods | Findings | Authors’ Names |
|---|---|---|---|---|---|---|
| Postsynaptic density protein 95 | ||||||
| PSD-95 | ICR male, female mice |
| Hippocampus—CA1 | Immunofluorescence | ↓PSD-95 protein after KET ↑PSD-95 protein after BET100, KE+BET30 and KE+BET100 | Chen et al. [164] |
| Swiss male mice |
| Hippocampus (HP) Prefrontal Cortex (PFC) | Western Blot | ↔PSD-95 protein in HP after KET5, GUO5↓PSD-95 protein in HP after CORT, CORT+GUO5 ↑PSD-95 protein in HP after CORT+KET5 ↔PSD-95 protein in PFC after KET5, GUO5, CORT, CORT+KET5, CORT+GUO5 ↔PSD-95 protein in HP after FLU ↓PSD-95 protein in HP after CORT ↓PSD-95 protein in HP after CORT +FLU ↔PSD-95 protein in PFC after FLU, CORT, CORT + FLU ↔PSD-95 protein in HP after KET1+GUO5 ↓PSD-95 protein in HP after CORT, CORT+KET1+GUO5 ↔PSD-95 protein in PFC after KET1 + GUO5, CORT, CORT + KET1 + GUO5 | Camargo et al. [138] | |
| C57BL/6 male mice |
| Basal nuclei (BLA) Lateral nuclei (LAT) Medial Prefrontal Cortex (mPFC) | Immunohistochemistry | ↑PSD-95 protein in LAT and BLA after CRS ↓PSD-95 protein in LAT and BLA after CRS+IMI ↑PSD-95 expression in mPFC after CRS+IMI ↓PSD-95 expression in mPFC after CRS | Leem et al. [165] | |
| C57BL/6 male mice |
| Frontal Cortex Hippocampus | Western Blot | ↔PSD-95 protein after CUMS, CUMS + FLU, CUMS+ASI10, 20, 40 ↓PSD-95 protein after CUMS ↑PSD-95 protein after CUMS+ASI10, CUMS + ASI20, CUMS +ASI40, CUMS+FLU | Luo et al. [166] | |
| C57Bl/6J male mice |
| Hippocampus (HP) Prefrontal Cortex (PFC) | Western Blot | ↓PSD-95 protein after LUR10 and FLU20 ↓PSD-95 protein after LUR10 and FLU20 | Stan et al. [127] | |
| C57Bl/6J male mice |
| Basolateral Amygdala | Western Blot | ↑PSD-95 protein after CRS ↓PSD-95 protein after CRS + FLU | Yi et al. [136] | |
| C57BL/6J male mice |
| Hippocampus | Western Blot | ↓PSD-95 protein after CRS, CRS+HNK, CRS+HNK+NBQX, CRS+HNK+ANA-12 | Ju et al. [141] | |
| KM (Kunming) male mice |
| Prefrontal Cortex (PFC) Hippocampus (HP) Hypothalamus (HY) Olfactory Bulb (AL) | qPCR Western Blot | ↓Dlg4 mRNA after UCMS, UCMS + Ala2+ Rapa in PFC, HP, Hy, AL. ↑Dlg4mRNA after UCMS+ Ala1 Ala2 in PFC, HP ↔Dlg4 mRNA after UCMS + Ala1 in HY, AL ↑Dlg4 mRNA after UCMS + Ala2 in HY, AL ↓PSD-95 protein after UCMS, UCMS+Ala2+Rapa in PFC ↑PSD-95 protein after UCMS+Ala2 in PFC ↔PSD-95 protein after UCMS+Ala1 in PFC ↓PSD-95 protein after UCMS, UCMS + Ala2 + Rapa in Hp ↑ PSD-95 protein after UCMS+Ala1, Ala2 in Hp ↓PSD-95 protein after UCMS in Hy ↔PSD-95 protein after UCMS + Ala1, Ala2, Ala2 + Rapa in HY ↓PSD-95 protein after UCMS in AL ↑PSD-95 protein after UCMS + Ala2 in AL ↔PSD-95 protein after UCMS + Ala1, Ala2+Rapa | Zhuang et al. [167] | |
| Calcium/calmodulin-dependent protein kinase II | ||||||
| p-CaMKII | C57BL/6 male mice |
| Basal nuclei (BA) Lateral nuclei (LAT) Medial Prefrontal Cortex (mPFC) | Immunohistochemistry | ↑p-CamKII protein in LAT, BA after CRS and ↓after CRS+IMI ↓p-CamKII protein after CRS in mPFC ↑p-CamKII protein after CRS+IMI in mPFC | Leem et al. [165] |
| p-T286-CaMKII t-CaMKII | C57BL/6J male, female mice |
| Prefrontal Cortex | Western Blot | ↓CaMKII-p-T286 and ↔ t-CaMKII protein after SD ↑CaMKII-p-T286 protein after CTM and CTM + SD | Misrani et al. [168] |
| CaMKII | ICR male, female mice |
| Hippocampus | Western Blot | ↔CaMKII protein after YG, ES, YG+LPS, ES+LPS (both sexes) ↓CaMKII (female) and ↑CaMKII (male) protein after CUS ↑CaMKII (female) and ↓CaMKII (male) protein after CUS+YG, CUS+ES | Yin et al. [169] |
| p-CaMKII (no information about the site of phosphorylation) | ICR male, female mice |
| Hippocampus | Western Blot | ↓p-CAMKII protein after cLH and cLH+YJ (at day 7) ↔ p-CAMKII protein after cLH+FLU (at day 7) ↓p-CAMKII protein after cLH and cLH+FLU (at day 26—after 5 days break in drug administration) | Zou et al. [170] |
| p-T286- CaMKII | C57BL/6J male, female mice |
| Hippocampus | Western Blot | ↓p-T286-CAMKII protein after SD ↑p-T286-CAMKII protein after CTM, CTM + SD | Misrani et al. [171] |
| CaMKIIα | C57BL/6J male mice |
| Basolateral Amygdala (BLA) Inferior-Limbic Prefrontal Cortex (IL-PFC) | Western Blot | ↔CAMKIIα protein after KET in BLA and IL-PFC | Asim et al. [129] |
| Homer scaffold protein 1 | ||||||
| Homer 1a | C57BL/6J male mice |
| Hippocampus (HP) Prefrontal Cortex (PFC) | Western Blot | ↓Homer 1a protein in HP and PFC after CUMS ↔Homer 1a protein in HP and PFC after CUMS+15 Hz HF-rTMS, CUMS + 25 Hz HF-rTMS | Zuo et al. [128] |
| Homer 1a Homer 1b/c | C57BL/6 female mice |
| Cortex | qPCR | ↓Homer1a and Homer1b/c mRNA expression after CDM ↑Homer1a and Homer1b/c mRNA expression after IMI, FLU, KET | Sun et al. [172] |
| Homer 1a Homer 1b/c | C57BL/6 J, CD1 male mice |
| Medial Prefrontal Cortex (mPFC) Hippocampus (HP) Amygdala (AMY) | qPCR Western Blot | ↔Homer1a mRNA in Susceptible, and Resilient mice after CSDS—all brain regions ↑Homer1b/c mRNA in Susceptible mice after CSDS—HP ↔Homer 1a protein in Susceptible and Resilient mice after CSDS—all brain regions ↑Homer 1b/c protein in Susceptible mice after CSDS—HP | Li et al. [139] |
| Zinc transporters | ||||||
| ZnT-1 | Albino Swiss male mouse |
| Prefrontal Cortex Hippocampus | Western Blot | ↓ZnT-1 protein after Zn ↔ZnT-1 protein after IMI and Zn + IMI ↓ZnT-1 protein after Zn ↔ ZnT-1 protein level after IMI ↑ ZnT-1 protein level after Zn + IMI | Rafało-Ulińska et al. [173] |
| Nitric oxide synthases | ||||||
| nNOS | ICR male, female mouse |
| Hippocampus | Western Blot | ↔ nNOS protein after YG, ES, YG+LPS, ES+LPS (both sexes) ↓nNOS protein after CUS (female) ↑nNOS protein after CUS+YG, CUS+ES (female) ↑nNOS protein after CUS (male) ↓nNOS protein after CUS+YG, CUS+ES (male) | Yin et al. [169] |
| Postsynaptic Proteins | Species/Strain | Model/Treatment | Brain Region | Methods | Findings | Authors’ Names |
|---|---|---|---|---|---|---|
| Postsynaptic density protein 95 | ||||||
| PSD-95 | Sprague Dawley male rats |
| Medial prefrontal cortex | Western blot | ↓PSD-95 protein after CUS ↑PSD-95 protein after CUS+ IFE | Li et al. [145] |
| Sprague Dawley male rats |
| Hippocampus (CA1 and CA3 region) Dentate gyrus | Western blot | ↓PSD-95 protein after CUMS in all regions ↑PSD-95 protein after CUMS+ NaHS in all regions | Hou et al. [162] | |
| Sprague Dawley male rats |
| Hypothalamus | Western blot qPCR | ↑PSD-95 protein after CUS ↑Dlg4 mRNA after CUS | Li et al. [161] | |
| Sprague Dawley male rats |
| Medial prefrontal cortex | Western blot | ↑PSD-95 protein after CX717 (only 2 h after administration) | Gordillo-Salas et al. [174] | |
| Sprague Dawley male rats |
| Hippocampus | Western blot | ↓PSD-95 protein after CUMS ↑PSD-95 protein after CUMS+ S-KET ↑PSD-95 protein after CUMS+ NSC23766 + S-KET ↑PSD-95 protein after CUMS+ NSC23766 | Zhu et al. [156] | |
| Sprague Dawley male rats |
| Amygdala (AMY) Prefrontal cortex (PFC) | Western blot | ↑PSD-95 protein after 3 CS-US in AMY ↔PSD-95 protein after 6 CS-US in AMY ↑PSD-95 protein after 10 CS-US in AMY ↔PSD-95 protein after 10 CS-US+ KET in AMY ↔PSD-95 protein after 10 CS-US+ FLU in AMY ↓PSD-95 protein after 3 CS-US in PFC ↓PSD-95 protein after 6 CS-US in PFC ↓PSD-95 protein after 10 CS-US in PFC ↑PSD-95 protein after 10 CS-US+ KET in PFC ↔PSD-95 protein after 10 CS-US + FLU in PFC | Lee et al. [157] | |
| Sprague Dawley male rats |
| Medial prefrontal cortex | Western blot | ↓PSD-95 protein after CUMS ↑PSD-95 protein after CUMS+ YY-21 ↔PSD-95 protein after CUMS+ FLU | Guo et al. [150] | |
| Sprague Dawley male rats |
| Prefrontal cortex | Western blot | ↔PSD95 protein 30min after Zn treatment ↑PSD95 protein level 3 h after Zn treatment ↔PSD95 protein level 24h after Zn treatment | Szewczyk et al. [159] | |
| Wistar male rats |
| Prefrontal cortex | qPCR | ↓Dlg4 mRNA after CMS (stress-reactive group) ↑Dlg4mRNA after CMS+ LUR | Luoni et al. [175] | |
| PSD-95 | Sprague Dawley male rats |
| Hippocampus | Western blot Immunohistochemistry qPCR | ↓PSD-95 protein after CUMS ↑PSD95 protein CUMS+ FLU ↑PSD95 protein CUMS+ SDDH ↔PSD95 protein CUMS+ SDDL ↓PSD-95 protein after CUMS ↑PSD95 protein CUMS+ FLU ↑PSD95 protein CUMS+ SDDH ↔PSD95 protein CUMS+ SDDL ↓Dlg4 mRNA after CUMS ↑Dlg4 mRNA after CUMS+ FLU ↑Dlg4 mRNA after CUMS+ SDDH ↑Dlg4 mRNA after CUMS+ SDDL | Liu et al. [163] |
| Calcium/calmodulin-dependent protein kinase II | ||||||
| CaMKII p-T286-CaMKII p-T286-CaMKIIα p-T286-CaMKIIβ | Sprague Dawley male rats |
| Hippocampus | Western blot | ↔CaMKII protein after CUMS ↔CaMKII protein after CUMS+ GXDSF in all groups ↑p-CaMKII protein after CUMS ↓p-CaMKII protein after CUMS+ GXDSF in all groups ↑p-CaMKIIα protein after CUMS ↓p-CaMKIIα protein after CUMS+ GXDSF in all groups ↑p-CaMKIIβ protein after CUMS ↓ p-CaMKIIβ protein after CUMS+ GXDSF in all groups | Xie et al. [176] |
| CaMKIIγ CaMKIIβ | Dark Agouti male rats |
| Frontal cortex | qPCR | ↑Camk2g mRNA after VLX ↑Camk2b mRNA after VLX | Tamási et al. [152] |
| Homer scaffold protein 1 | ||||||
| Homer 1 | Wistar male rats |
| Prefrontal cortex | Western blot | ↑Homer1 protein in resilient group compared to anhedonic | Palmfeldt et al. [177] |
| Homer 1 | Sprague Dawley male rats |
| Hippocampus (synaptoneurosomes) | Western blot | ↔Homer1 protein after CRS ↔Homer1 protein after CRS+ FAS | Román-Albasini et al. [148] |
| Homer 1 | Sprague Dawley male rats |
| Hippocampus | Western blot Immunohistochemistry qPCR | ↑Homer1b/c protein after CUMS ↓Homer-1b/c protein after CUMS+ FLU ↓Homer1b/c protein after CUMS+ SSDH ↓Homer1b/c protein after CUMS+ SSDL ↑Homer1b/c protein after CUMS ↓Homer1b/c protein after CUMS+ FLU ↓Homer1b/c protein after CUMS+ SSDH ↓Homer1b/c protein after CUMS+ SSDL ↑Homer1 mRNA after CUMS ↓Homer1 mRNA after CUMS+ FLU ↓Homer1 mRNA after CUMS+ SDDH ↓Homer1 mRNA after CUMS+ SDDL | Liu et al. [163] |
| Nitric oxide synthases | ||||||
| nNOS iNOS eNOS | Wistar male rats |
| Cerebellum (CER) Frontal cortex (FC) Midbrain (MID) Olfactory bulb (OB) Pons Striatum Temporal cortex (TC) Thalamus (TH) Hippocampus (HP) | qPCR | ↔Nos1 mRNA after AR ↔Nos1 mRNA after AR+ FLU, AR+MIL, AR+MIR—all brain regions ↑Nos2 mRNA after AR—CER, FC, MID, OB, pons, striatum, TC, TH ↔Nos2 mRNA after AR—HP ↔Nos2 mRNA after AR+ FLU, AR+MIL, AR+MIR—all brain regions ↔Nos3 mRNA after AR ↑Nos3 mRNA after AR+ FLU—CER, Hp, midbrain, pons, striatum, thalamus, FC ↔Nos3 mRNA after AR+ FLU—OB, TC; after AR+ MIL and AR+ MIR—all brain regions | Yoshino et al. [178] |
| nNOS | Sprague Dawley male rats |
| Hippocampus | Western blot | ↑nNOS protein after CUS ↓nNOS protein after CUS + MEM | Mishra et al. [179] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Samojedny, S.; Czechowska, E.; Pańczyszyn-Trzewik, P.; Sowa-Kućma, M. Postsynaptic Proteins at Excitatory Synapses in the Brain—Relationship with Depressive Disorders. Int. J. Mol. Sci. 2022, 23, 11423. https://doi.org/10.3390/ijms231911423
Samojedny S, Czechowska E, Pańczyszyn-Trzewik P, Sowa-Kućma M. Postsynaptic Proteins at Excitatory Synapses in the Brain—Relationship with Depressive Disorders. International Journal of Molecular Sciences. 2022; 23(19):11423. https://doi.org/10.3390/ijms231911423
Chicago/Turabian StyleSamojedny, Sylwia, Ewelina Czechowska, Patrycja Pańczyszyn-Trzewik, and Magdalena Sowa-Kućma. 2022. "Postsynaptic Proteins at Excitatory Synapses in the Brain—Relationship with Depressive Disorders" International Journal of Molecular Sciences 23, no. 19: 11423. https://doi.org/10.3390/ijms231911423





