Improved Salinity Tolerance-Associated Variables Observed in EMS Mutagenized Wheat Lines
Abstract
1. Introduction
2. Results
2.1. Selection of Mutagenized Lines for In-Depth Study
2.2. Mutagenized Wheat Lines Differed in Shoot Size in Response to Salinity
Smoothed Estimated Plant Biomass (sEPB) Absolute Growth Rate (AGR)
2.3. Large Variation Observed in Leaf Na+, K+, and Cl− Content on a Dry Weight Basis within the Selected Mutagenized Lines
2.3.1. K+ Content Based on Dry Weight
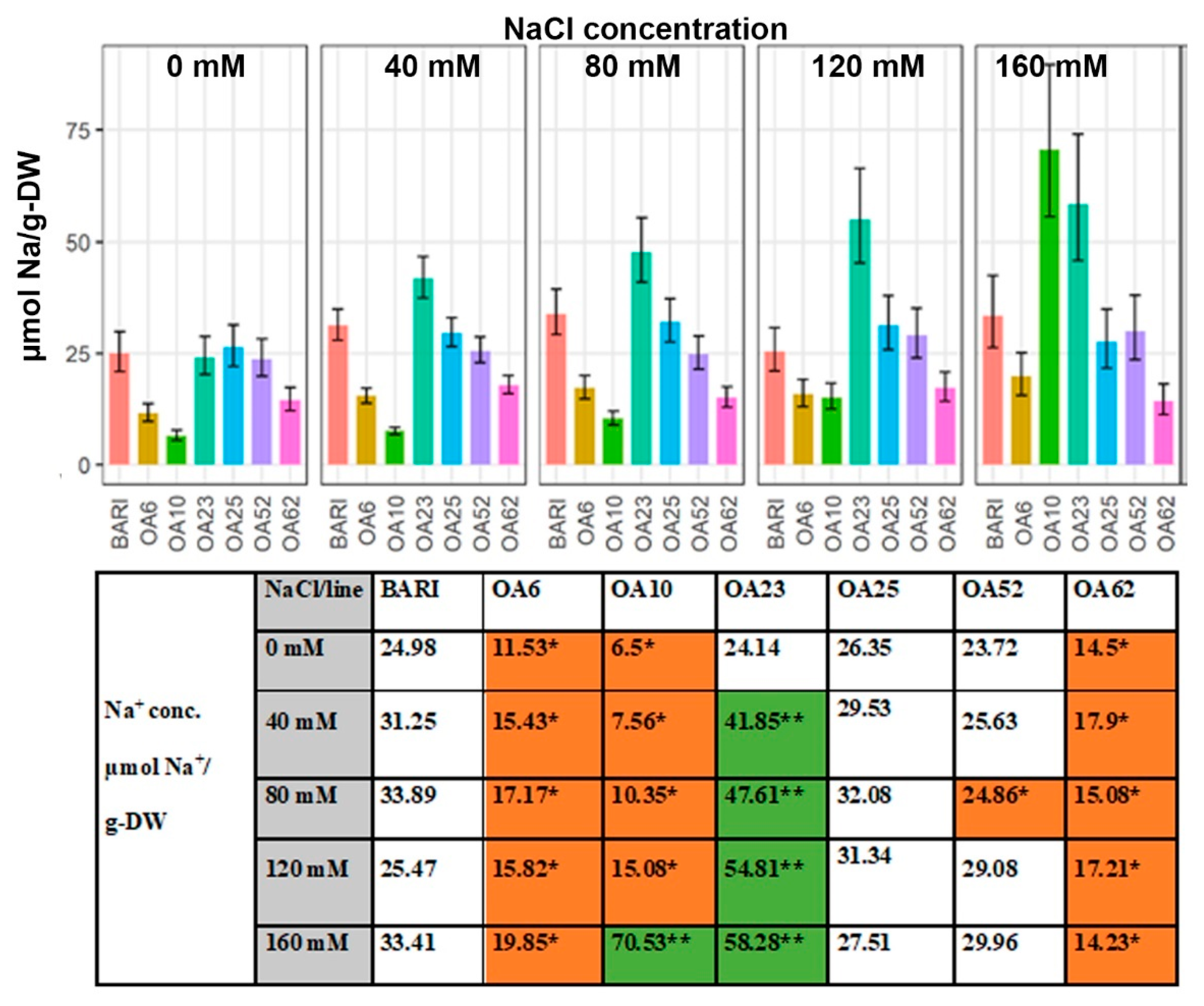
2.3.2. Na+ Content Based on Dry Weight
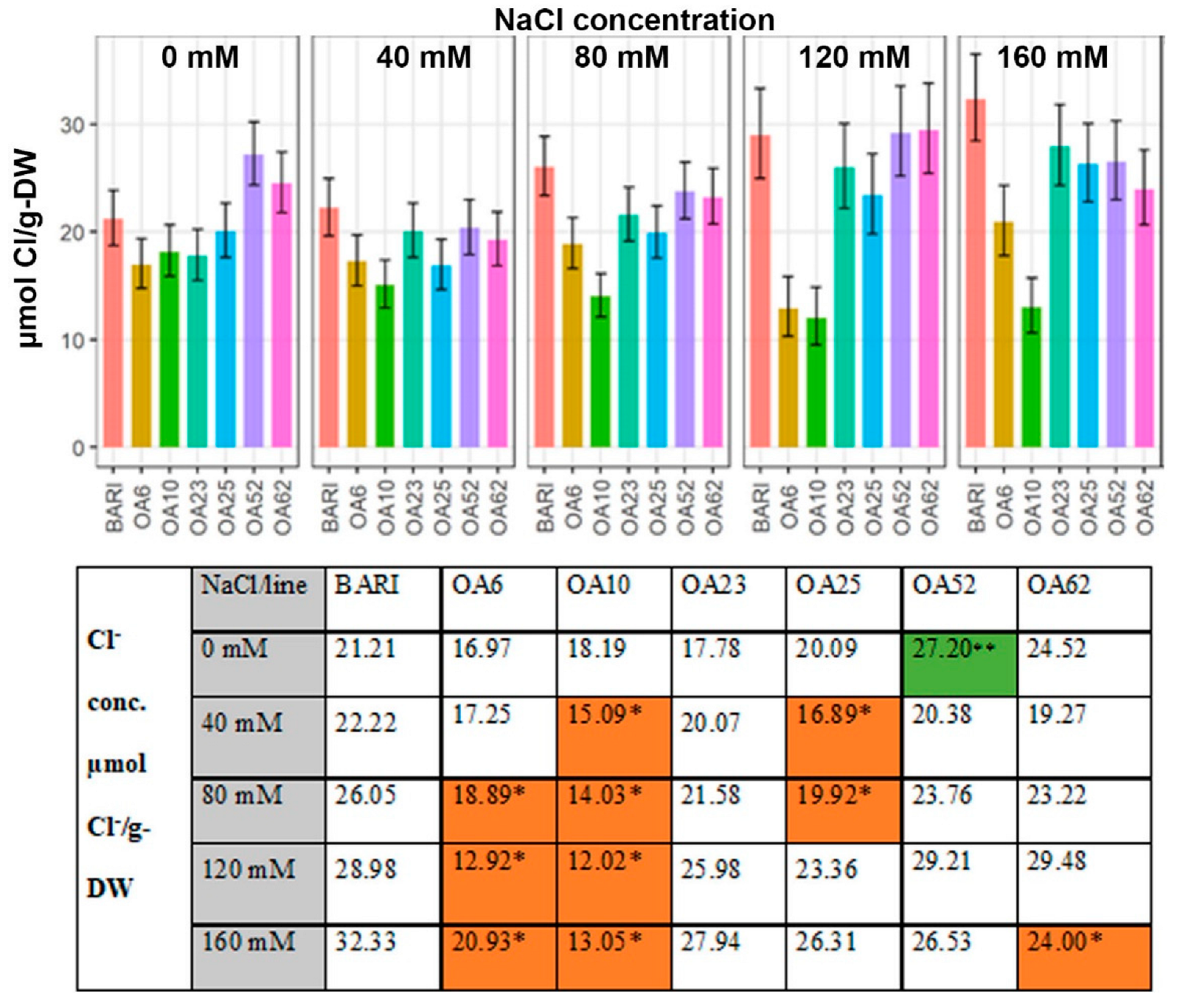
2.3.3. Cl− Content Based on the Dry Weight
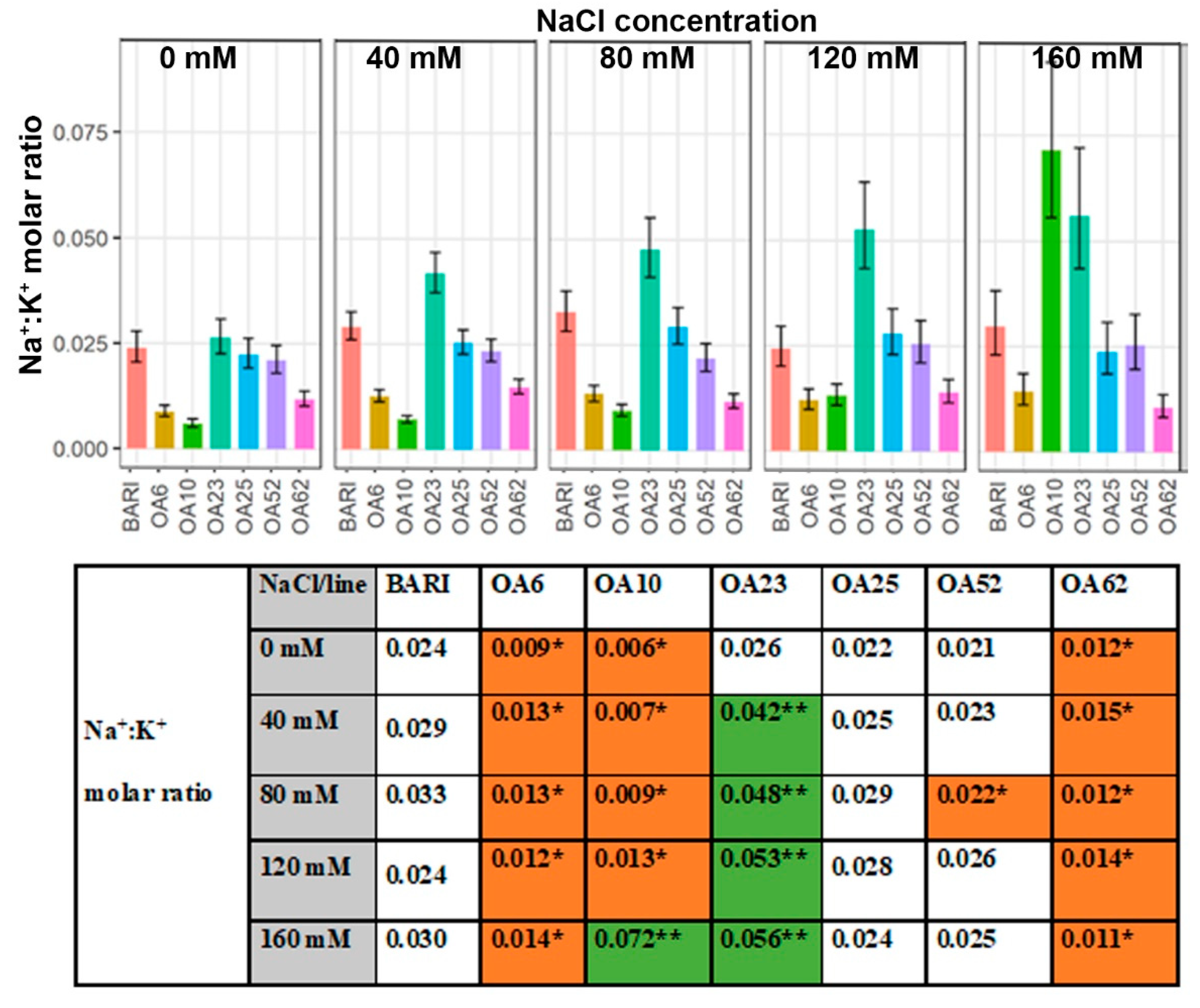
2.3.4. Leaf Na+:K+ Ratio
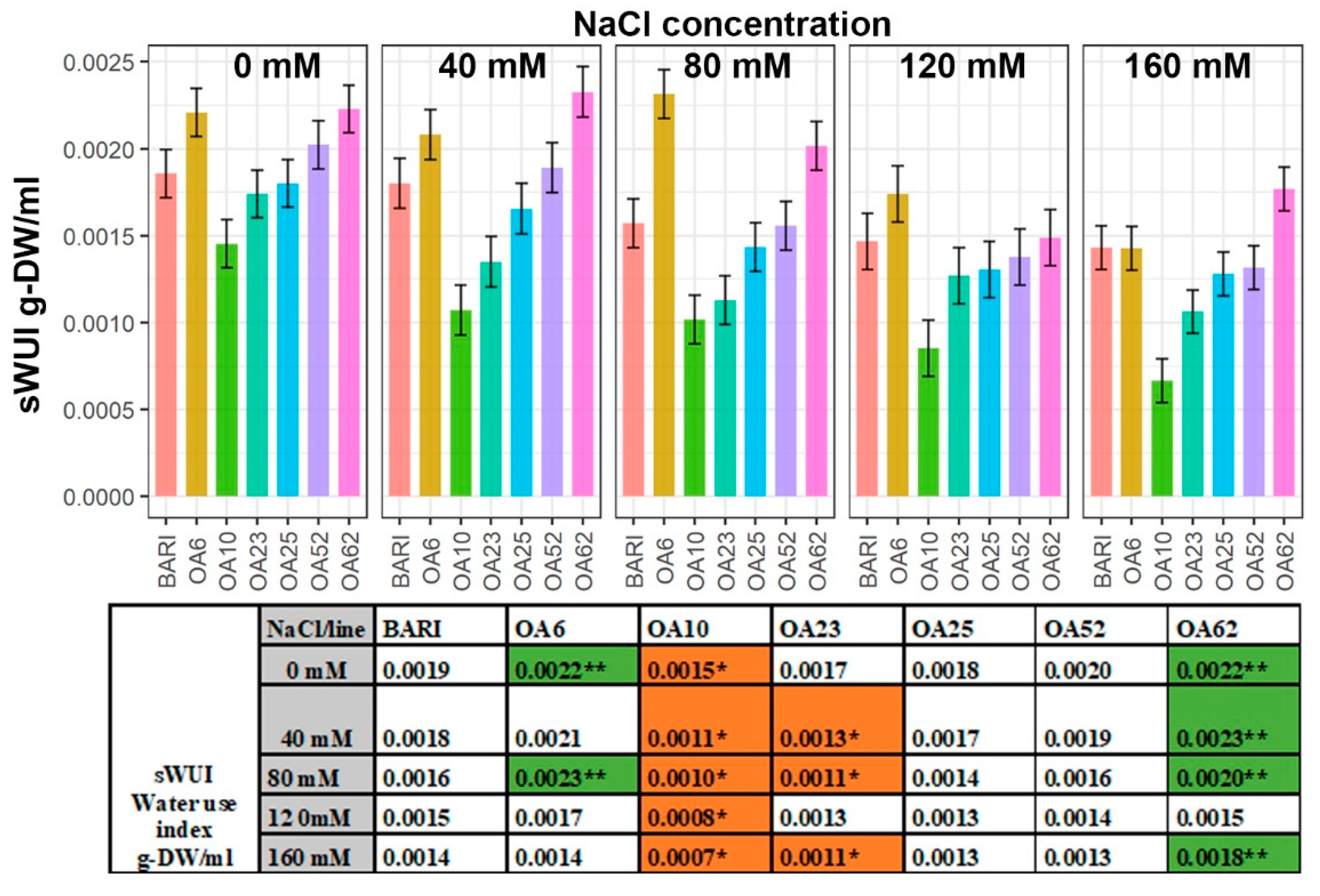
2.4. The Water Use Index (sWUI) Was Greater in Two Lines, whereas One Line Was Less Efficient
2.5. Several Lines Had Higher TKW Than the Control at Higher Salt Levels
3. Discussion
3.1. OA6 and OA62 Had Improved Growth during Osmotic and Ion Stress
3.2. Increased Biomass of OA6 and OA62 Correlated Well with Efficient Water Use
3.3. Ion Concentration/Homeostasis
3.3.1. OA23 Retained High Na+ Levels and Maintained Performance
3.3.2. Cl− Levels Were Low in All Mutant Lines Relative to the Control
3.3.3. K+ Levels Were Generally Higher Than the Control except for OA23
3.3.4. Ion Ratios—Increased K+ Levels Manage Na+ Accumulation in Three Lines
3.4. Mutagenized Lines Overall Had Relatively High TKW Relative to the Controls in Saline Conditions
3.5. Variation in the Improved Salinity Tolerance-Associated Variables Observed in the Characterized Lines
4. Materials and Methods
4.1. Plant Material
4.2. Experiment Description
4.3. Preparation of Image Data
4.4. Water Use
4.5. Harvest (Leaf) Data
4.6. Measurement of Leaf Ion Concentration
4.7. Thousand Kernel Weight (TKW)
4.8. Statistical Analyses
4.9. Rationale behind Selection of Lines for in Depth Studies
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- FAO. Global Map of Salt-Affected Soils; FAO: Rome, Italy, 2021; pp. 1–20. [Google Scholar]
- Bodirsky, B.L.; Rolinski, S.; Biewald, A.; Weindl, I.; Popp, A.; Lotze-Campen, H. Global food demand scenarios for the 21st century. PLoS ONE 2015, 10, e0139201. [Google Scholar] [CrossRef]
- Miransari, M.; Smith, D. Sustainable wheat (Triticum aestivum L.) production in saline fields: A review. Crit. Rev. Biotechnol. 2019, 39, 999–1014. [Google Scholar] [CrossRef]
- Hassani, A.; Azapagic, A.; Shokri, N. Global predictions of primary soil salinization under changing climate in the 21st century. Nat. Commun. 2021, 12, 6663. [Google Scholar] [CrossRef]
- Gould, I.; De Waegemaeker, J.; Tzemi, D.; Wright, I.; Pearson, S.; Ruto, E.; Karrasch, L.; Christensen, L.S.; Aronsson, H.; Eich-Greatorex, S. Salinization threats to agriculture across the north sea region. In Future of Sustainable Agriculture in Saline Environments; Negacz, K., Vellinga, P., Barrett-Lennard, E., Choukr-Allah, R., Elzenga, T., Eds.; CRC Press, Taylor and Francis Group: Boca Raton, FL, USA, 2021; pp. 71–92. [Google Scholar]
- Hussain, S.; Shaukat, M.; Ashraf, M.; Zhu, C.; Jin, Q.; Zhang, J. Salinity stress in arid and semi-arid climates: Effects and management in field crops. In Climate Change and Agriculture; Hussain, S., Ed.; IntechOpen: London, UK, 2019; Volume 13. [Google Scholar]
- Lethin, J.; Shakil, S.S.M.; Hassan, S.; Sirijovski, N.; Töpel, M.; Olsson, O.; Aronsson, H. Development and characterization of an ems-mutagenized wheat population and identification of salt-tolerant wheat lines. BMC Plant Biol. 2020, 20, 18. [Google Scholar] [CrossRef]
- Malaker, P.; Reza, M.M.A. Resistance to rusts in bangladeshi wheat. Czech J. Genet. Plant Breed. 2011, 47, S155–S159. [Google Scholar] [CrossRef]
- Department of Crop Botany, Bangladesh. Digital Herbarium of Crop Plants. Available online: http://dhcrop.bsmrau.net/ (accessed on 11 January 2022).
- Barma, N.; Saifuzzaman, M.; Hossain, A.; Rahman, M.M.; Saha, N.K.; Rawson, H.J. Breeding wheat for heat and salt tolerance in southern bangladesh. In Sustainable Intensification of Rabi Cropping in Southern Bangladesh Using Wheat and Mungbean; Rawson, H.M., Ed.; ACIAR Technical Reports: Canberra, Australia, 2011; p. 170. [Google Scholar]
- Haque, S.A. Salinity problems and crop production in coastal regions of bangladesh. Pak. J. Bot. 2006, 38, 1359–1365. [Google Scholar]
- Mousavi, H.; Bruning, B.; Van Straten, G.; Almås, Å.R.; Lethin, J.; Naik, N.; Hassan, S.; Olsson, O.; Aronsson, H. Effects of increasing salinity by drip irrigation on total grain weight show high yield potential of putative salt-tolerant mutagenized wheat lines. Sustainability 2022, 14, 5061. [Google Scholar] [CrossRef]
- Neilson, E.H.; Edwards, A.M.; Blomstedt, C.; Berger, B.; Møller, B.L.; Gleadow, R.M. Utilization of a high-throughput shoot imaging system to examine the dynamic phenotypic responses of a c4 cereal crop plant to nitrogen and water deficiency over time. J. Exp. Bot. 2015, 66, 1817–1832. [Google Scholar] [CrossRef]
- Al-Tamimi, N.; Brien, C.; Oakey, H.; Berger, B.; Saade, S.; Ho, Y.S.; Schmöckel, S.M.; Tester, M.; Negrão, S. Salinity tolerance loci revealed in rice using high-throughput non-invasive phenotyping. Nat. Commun. 2016, 7, 13342. [Google Scholar] [CrossRef]
- Berger, B.; Parent, B.; Tester, M. High-throughput shoot imaging to study drought responses. J. Exp. Bot. 2010, 61, 3519–3528. [Google Scholar] [CrossRef]
- Tilbrook, J.; Schilling, R.K.; Berger, B.; Garcia, A.F.; Trittermann, C.; Coventry, S.; Rabie, H.; Brien, C.; Nguyen, M.; Tester, M.; et al. Variation in shoot tolerance mechanisms not related to ion toxicity in barley. Funct. Plant Biol. 2017, 44, 1194–1206. [Google Scholar] [CrossRef]
- Atieno, J.; Li, Y.; Langridge, P.; Dowling, K.; Brien, C.; Berger, B.; Varshney, R.K.; Sutton, T. Exploring genetic variation for salinity tolerance in chickpea using image-based phenotyping. Sci. Rep. 2017, 7, 1–11. [Google Scholar]
- Liang, W.; Ma, X.; Wan, P.; Liu, L. Plant salt-tolerance mechanism: A review. Biochem. Biophys. Res. Commun. 2018, 495, 286–291. [Google Scholar] [CrossRef]
- Deinlein, U.; Stephan, A.B.; Horie, T.; Luo, W.; Xu, G.; Schroeder, J.I. Plant salt-tolerance mechanisms. Trends Plant Sci. 2014, 19, 371–379. [Google Scholar] [CrossRef]
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef]
- Munns, R.; James, R.A.; Xu, B.; Athman, A.; Conn, S.J.; Jordans, C.; Byrt, C.S.; Hare, R.A.; Tyerman, S.D.; Tester, M.; et al. Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat. Biotechnol. 2012, 30, 360–364. [Google Scholar] [CrossRef]
- Munns, R.; Schachtman, D.; Condon, A. The significance of a two-phase growth response to salinity in wheat and barley. Aust. J. Plant Physiol. 1995, 22, 561–569. [Google Scholar] [CrossRef]
- Munns, R.; Passioura, J.B.; Colmer, T.D.; Byrt, C.S. Osmotic adjustment and energy limitations to plant growth in saline soil. New Phytol. 2020, 225, 1091–1096. [Google Scholar] [CrossRef]
- Wakeel, A.; Farooq, M.; Qadir, M.; Schubert, S. Potassium substitution by sodium in plants. Crit. Rev. Plant Sci. 2011, 30, 401–413. [Google Scholar] [CrossRef]
- Fowler, J. Interaction of salinity and temperature on the germination of crambe. Agron. J. 1991, 83, 169–172. [Google Scholar] [CrossRef]
- Flowers, T.; Yeo, A. Breeding for salinity resistance in crop plants: Where next? Aust. J. Plant Physiol. 1995, 22, 875–884. [Google Scholar] [CrossRef]
- Campbell, M.T.; Knecht, A.C.; Berger, B.; Brien, C.J.; Wang, D.; Walia, H. Integrating image-based phenomics and association analysis to dissect the genetic architecture of temporal salinity responses in rice. Plant Physiol. 2015, 168, 1476–1489. [Google Scholar] [CrossRef] [PubMed]
- Rajendran, K.; Tester, M.; Roy, S. Quantifying the three main components of salinity tolerance in cereals. Plant Cell Environ. 2009, 32, 237–249. [Google Scholar] [CrossRef]
- Takahashi, F.; Tilbrook, J.; Trittermann, C.; Berger, B.; Roy, S.J.; Seki, M.; Shinozaki, K.; Tester, M. Comparison of leaf sheath transcriptome profiles with physiological traits of bread wheat cultivars under salinity stress. PLoS ONE 2015, 10, e0133322. [Google Scholar] [CrossRef] [PubMed]
- Hairmansis, A.; Berger, B.; Tester, M.; Roy, S.J. Image-based phenotyping for non-destructive screening of different salinity tolerance traits in rice. Rice 2014, 7, 16. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, K.; Ishimori, M.; Kajiya-Kanegae, H.; Takanashi, H.; Fujimoto, M.; Yoneda, J.-I.; Yano, K.; Koshiba, T.; Tanaka, R.; Iwata, H.; et al. Effect of salt tolerance on biomass production in a large population of sorghum accessions. Breed. Sci. 2020, 70, 167–175. [Google Scholar] [CrossRef] [PubMed]
- Saddiq, M.S.; Iqbal, S.; Hafeez, M.B.; Ibrahim, A.M.H.; Raza, A.; Fatima, E.M.; Baloch, H.; Jahanzaib; Woodrow, P.; Ciarmiello, L.F. Effect of salinity stress on physiological changes in winter and spring wheat. Agronomy 2021, 11, 1193. [Google Scholar] [CrossRef]
- Ma, Y.; Dias, M.C.; Freitas, H. Drought and salinity stress responses and microbe-induced tolerance in plants. Front. Plant Sci. 2020, 11, 591911. [Google Scholar] [CrossRef]
- Liu, J.; Fu, C.; Li, G.; Khan, M.N.; Wu, H. Ros homeostasis and plant salt tolerance: Plant nanobiotechnology updates. Sustainability 2021, 13, 3552. [Google Scholar] [CrossRef]
- Zahra, N.; Al Hinai, M.S.; Hafeez, M.B.; Rehman, A.; Wahid, A.; Siddique, K.H.; Farooq, M. Regulation of photosynthesis under salt stress and associated tolerance mechanisms. Plant Physiol. Biochem. 2022, 178, 55–69. [Google Scholar] [CrossRef]
- Cheng, X.; Liu, X.; Mao, W.; Zhang, X.; Chen, S.; Zhan, K.; Bi, H.; Xu, H. Genome-wide identification and analysis of hak/kup/kt potassium transporters gene family in wheat (Triticum aestivum L.). Int. J. Mol. Sci. 2018, 19, 3969. [Google Scholar] [CrossRef] [PubMed]
- Genc, Y.; Taylor, J.; Lyons, G.; Li, Y.; Cheong, J.; Appelbee, M.; Oldach, K.; Sutton, T. Bread wheat with high salinity and sodicity tolerance. Front. Plant Sci. 2019, 10, 1280. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Qiu, C.-W.; Fan, Y.; Huang, X.; Khan, W.; Wu, F.; Zhou, M.; Wang, Y.; Cao, F. Genome-wide association and transcriptome analysis reveals candidate genes for potassium transport under salinity stress in wheat. Environ. Exp. Bot. 2022, 202, 105034. [Google Scholar] [CrossRef]
- Xue, Q.; Stewart, B.; Lazar, M.D.; Piccinni, G.; Salisbury, C.D. Genotypic variation of osmotic adjustment, water-use and transpiration efficiency among closely related wheat lines. J. Crop Improv. 2012, 26, 258–281. [Google Scholar] [CrossRef]
- Hasanuzzaman, M.; Nahar, K.; Rahman, A.; Islam Anee, T.; Ul Alam, M.; Farha Bhuiyan, T.; Oku, H.; Fujita, M. Approaches to enhance salt stress tolerance in wheat. In Wheat Improvement, Management, and Utilization; Wanyera, R., Owuoche, J., Eds.; IntechOpen Book Series; IntechOpen: London, UK, 2017. [Google Scholar]
- Li, P.F.; Ma, B.L.; Guo, S.; Ding, T.T.; Xiong, Y.C. Bottom-up redistribution of biomass optimizes energy allocation, water use and yield formation in dryland wheat improvement. J. Sci. Food Agric. 2022, 102, 3336–3349. [Google Scholar] [CrossRef]
- Parveen, A.; Ahmar, S.; Kamran, M.; Malik, Z.; Ali, A.; Riaz, M.; Abbasi, G.H.; Khan, M.; Sohail, A.B.; Rizwan, M. Abscisic acid signaling reduced transpiration flow, regulated Na+ ion homeostasis and antioxidant enzyme activities to induce salinity tolerance in wheat (Triticum aestivum L.) seedlings. Environ. Technol. Innov. 2021, 24, 101808. [Google Scholar] [CrossRef]
- Genc, Y.; Mcdonald, G.K.; Tester, M. Reassessment of tissue Na+ concentration as a criterion for salinity tolerance in bread wheat. Plant Cell Environ. 2007, 30, 1486–1498. [Google Scholar] [CrossRef]
- Tavakkoli, E.; Fatehi, F.; Coventry, S.; Rengasamy, P.; McDonald, G.K. Additive effects of Na+ and Cl− ions on barley growth under salinity stress. J. Exp. Bot. 2011, 62, 2189–2203. [Google Scholar] [CrossRef]
- Flowers, T.; Dalmond, D. Protein synthesis in halophytes: The influence of potassium, sodium and magnesium in vitro. Plant Soil 1992, 146, 153–161. [Google Scholar] [CrossRef]
- Qin, Y.; Tian, Y.; Wang, J.; Han, L. Cloning and expression of a salinity tolerance related gene tahak1 from wheat. J. Triticeae Crops 2011, 31, 1014–1019. [Google Scholar]
- Ai, X.H. Functional Study of Transporter Genes TaBaSS2 and TaHaK11 in Wheat; Shandong University: Jinan, China, 2013. [Google Scholar]
- Garbarino, J.; Dupont, F. NaCl inuces a Na+/H+ antiport in tonoplast vesicles from barley roots. Plant Physiol. 1988, 86, 231–236. [Google Scholar] [CrossRef] [PubMed]
- Isayenkov, S.V.; Maathuis, F.J. Plant salinity stress: Many unanswered questions remain. Front. Plant Sci. 2019, 10, 80. [Google Scholar] [CrossRef] [PubMed]
- Assaha, D.V.M.; Ueda, A.; Saneoka, H.; Al-Yahyai, R.; Yaish, M.W. The role of Na+ and K+ transporters in salt stress adaptation in glycophytes. Front. Physiol. 2017, 8, 509. [Google Scholar] [CrossRef] [PubMed]
- Hassan, S.; Lethin, J.; Blomberg, R.; Mousavi, H.; Aronsson, H. In silico based screening of wrky genes for identifying functional genes regulated by wrky under salt stress. Comput. Biol. Chem. 2019, 83, 107131. [Google Scholar] [CrossRef] [PubMed]
- Hassan, S.; Berk, K.; Aronsson, H. Evolution and identification of dreb transcription factors in the wheat genome: Modeling, docking and simulation of dreb proteins associated with salt stress. J. Biomol. Struct. Dyn. 2021, 40, 7191–7204. [Google Scholar] [CrossRef]
- Hussain, N.; Ghaffar, A.; Zafar, Z.U.; Javed, M.; Shah, K.H.; Noreen, S.; Manzoor, H.; Iqbal, M.; Hassan, I.F.Z.; Bano, H.; et al. Identification of novel source of salt tolerance in local bread wheat germplasm using morpho-physiological and biochemical attributes. Sci. Rep. 2021, 11, 10854. [Google Scholar] [CrossRef]
- Brien, C. dae: Functions Useful in the Design and Anova of Experiments, R Package Version 3.1-16; 2019. Available online: https://cran.r-project.org/web/packages/dae/index.html (accessed on 20 September 2022).
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. Available online: https://www.r-project.org (accessed on 20 September 2022).
- Butler, D.G. od: Generate Optimal Experimental Designs, Version 2.0.0; 2018. Available online: https://mmade.org/ (accessed on 20 September 2022).
- Berger, B.; de Regt, B.; Tester, M. Trait dissection of salinity tolerance with plant phenomics. In Plant Salt Tolerance; Shabala, S., Cuin, T.A., Eds.; Springer: Berlin/Heidelberg, Germany, 2012; pp. 399–413. [Google Scholar]
- Brien, C.; Jewell, N.; Watts-Williams, S.J.; Garnett, T.; Berger, B. Smoothing and extraction of traits in the growth analysis of noninvasive phenotypic data. Plant Methods 2020, 16, 36. [Google Scholar] [CrossRef]
- Brien, C. growthPheno: Plotting, Smoothing and Growth Trait Extraction for Longitudinal Data—Version: 1.0-26. 2020. Available online: http://cran.at.r-project.org/package=growthPheno (accessed on 20 September 2022).
- Butler, D.; Cullis, B.; Gilmour, A.; Gogel, B.; Thompson, R. ASReml-R Reference Manual Version 4, 2009. Available online: http://asreml.org (accessed on 20 September 2022).
- Brien, C. asremlPlus: Augments Asreml-R in Fitting Mixed Models and Packages Generally in Exploring Prediction Differences, R Package Version 4.1–28; 2019. Available online: http://cran.r-project.org/package=asremlPlus/ (accessed on 17 June 2019).


| NaCl /Line | BARI | OA6 | OA10 | OA23 | OA25 | OA52 | OA62 |
|---|---|---|---|---|---|---|---|
| 0 mM | 51.65 ± 0.85 | 53.82 ± 4.40 | 29.50 ± 1.29 | 51.84 ± 3.45 | 54.54 ± 1.89 | 52.79 ± 0.90 | 46.48 ± 3.62 |
| 40 mM | 47.25 ± 2.41 | 45.82 ± 8.66 | 26.27 ± 2.43 | 57.98 ± 7.94 | 51.90 ± 5.06 | 46.56 ± 4.51 | 44.75 ± 0.90 |
| 80 mM | 48.60 ± 5.28 | 49.73 ± 12.6 | 28.76 ± 2.40 | 62.43 ± 5.58 | 50.21 ± 5.55 | 50.43 ± 5.91 | 44.05 ± 2.42 |
| 120 mM | 40.81 ± 3.41 | 47.74 ± 2.93 | 26.15 ± 1.51 | 51.13 ± 4.68 | 58.11 ± 0.19 | 58.26 ± 21.3 | 41.57 ± 2.10 |
| 160 mM | 42.36 ± 4.11 | 46.76 ± 2.82 | 25.86 ± 2.46 | 55.42 ± 8.03 | 47.20 ± 5.06 | 42.87 ± 3.97 | 40.89 ± 1.24 |
| Mean | 46.13 | 48.77 | 27.31 | 55.76 | 52.39 | 50.18 | 43.55 |
| Group | B | AB | C | A | A | AB | B |
| Parameter/Line | OA6 | OA10 | OA23 | OA25 | OA52 | OA62 |
|---|---|---|---|---|---|---|
| sEPB AGR | ||||||
| K+ | ||||||
| Na+ | ||||||
| Cl− | ||||||
| Na+:K+ | ||||||
| sWUI | ||||||
| TKW |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lethin, J.; Byrt, C.; Berger, B.; Brien, C.; Jewell, N.; Roy, S.; Mousavi, H.; Sukumaran, S.; Olsson, O.; Aronsson, H. Improved Salinity Tolerance-Associated Variables Observed in EMS Mutagenized Wheat Lines. Int. J. Mol. Sci. 2022, 23, 11386. https://doi.org/10.3390/ijms231911386
Lethin J, Byrt C, Berger B, Brien C, Jewell N, Roy S, Mousavi H, Sukumaran S, Olsson O, Aronsson H. Improved Salinity Tolerance-Associated Variables Observed in EMS Mutagenized Wheat Lines. International Journal of Molecular Sciences. 2022; 23(19):11386. https://doi.org/10.3390/ijms231911386
Chicago/Turabian StyleLethin, Johanna, Caitlin Byrt, Bettina Berger, Chris Brien, Nathaniel Jewell, Stuart Roy, Hesam Mousavi, Selvakumar Sukumaran, Olof Olsson, and Henrik Aronsson. 2022. "Improved Salinity Tolerance-Associated Variables Observed in EMS Mutagenized Wheat Lines" International Journal of Molecular Sciences 23, no. 19: 11386. https://doi.org/10.3390/ijms231911386
APA StyleLethin, J., Byrt, C., Berger, B., Brien, C., Jewell, N., Roy, S., Mousavi, H., Sukumaran, S., Olsson, O., & Aronsson, H. (2022). Improved Salinity Tolerance-Associated Variables Observed in EMS Mutagenized Wheat Lines. International Journal of Molecular Sciences, 23(19), 11386. https://doi.org/10.3390/ijms231911386









