Identification and Characterization of AUXIN Response Factor Gene Family Reveals Their Regulatory Network to Respond the Multi-Hormones Crosstalk during GA-Induced Grape Parthenocarpic Berry
Abstract
1. Introduction
2. Results
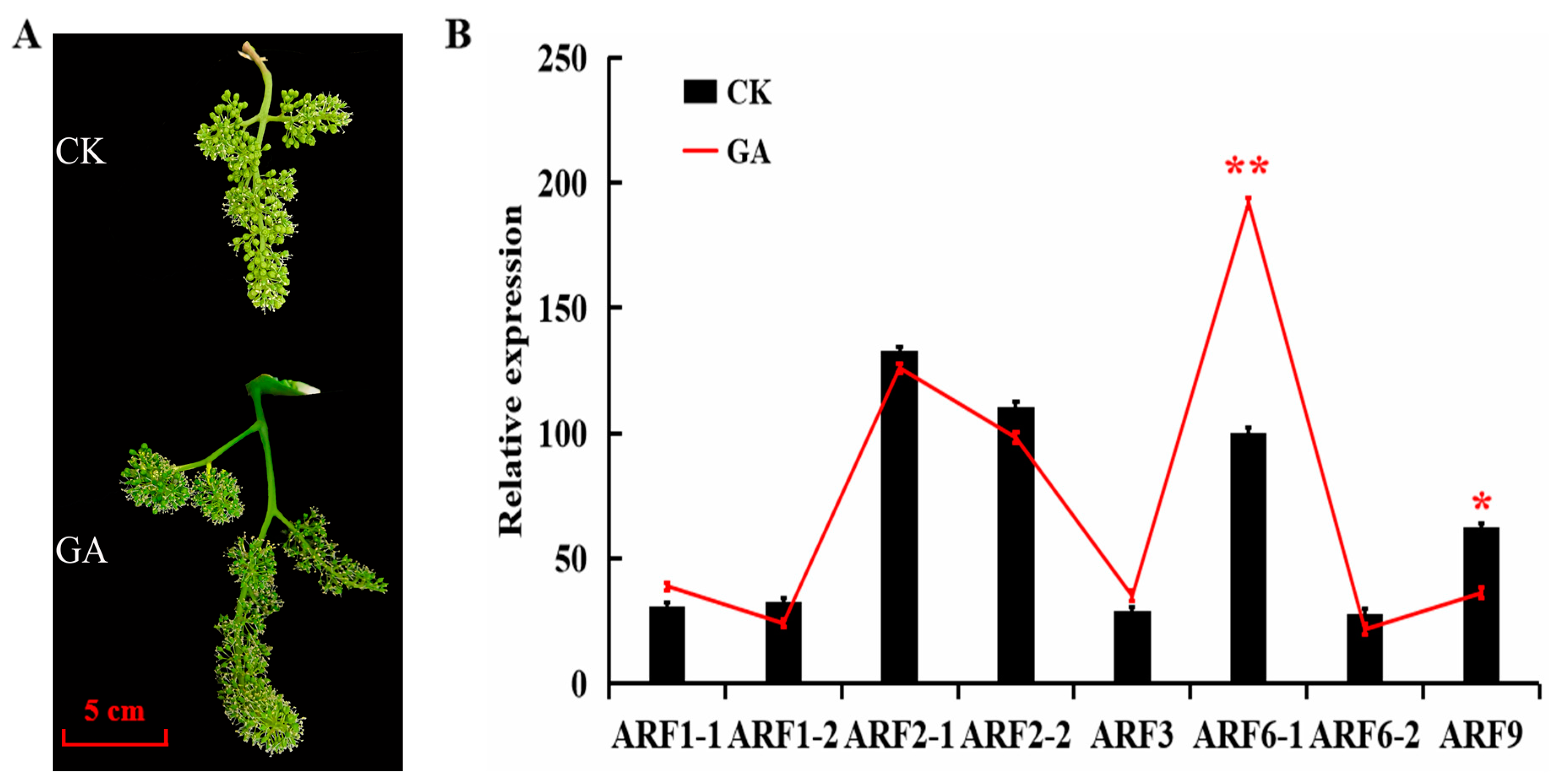
2.1. Changes of Berry Development and Quality during GA-Induced Parthenocarpy in Grapes
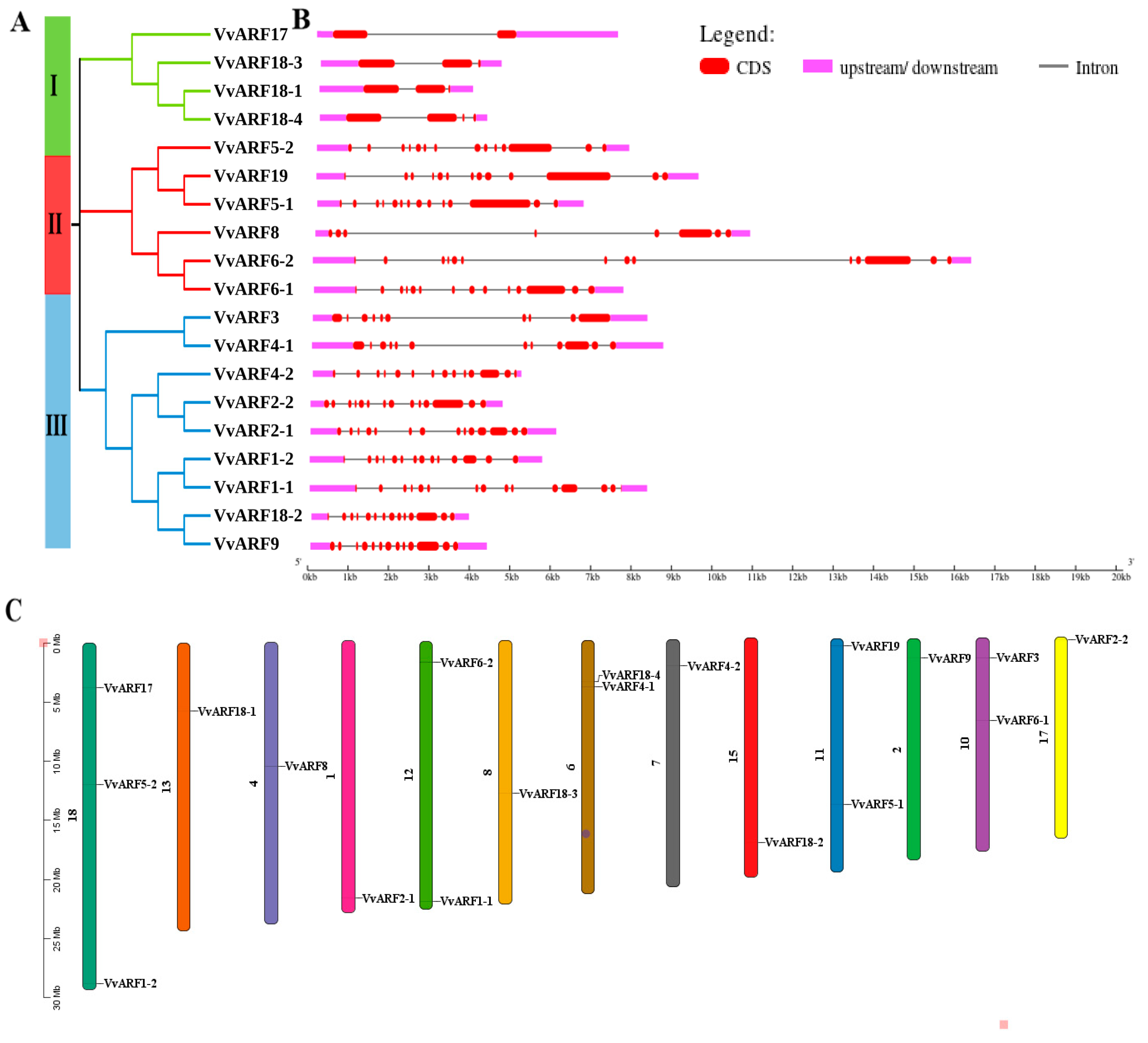
2.2. Identification and Characterization of ARF Family in Grapes
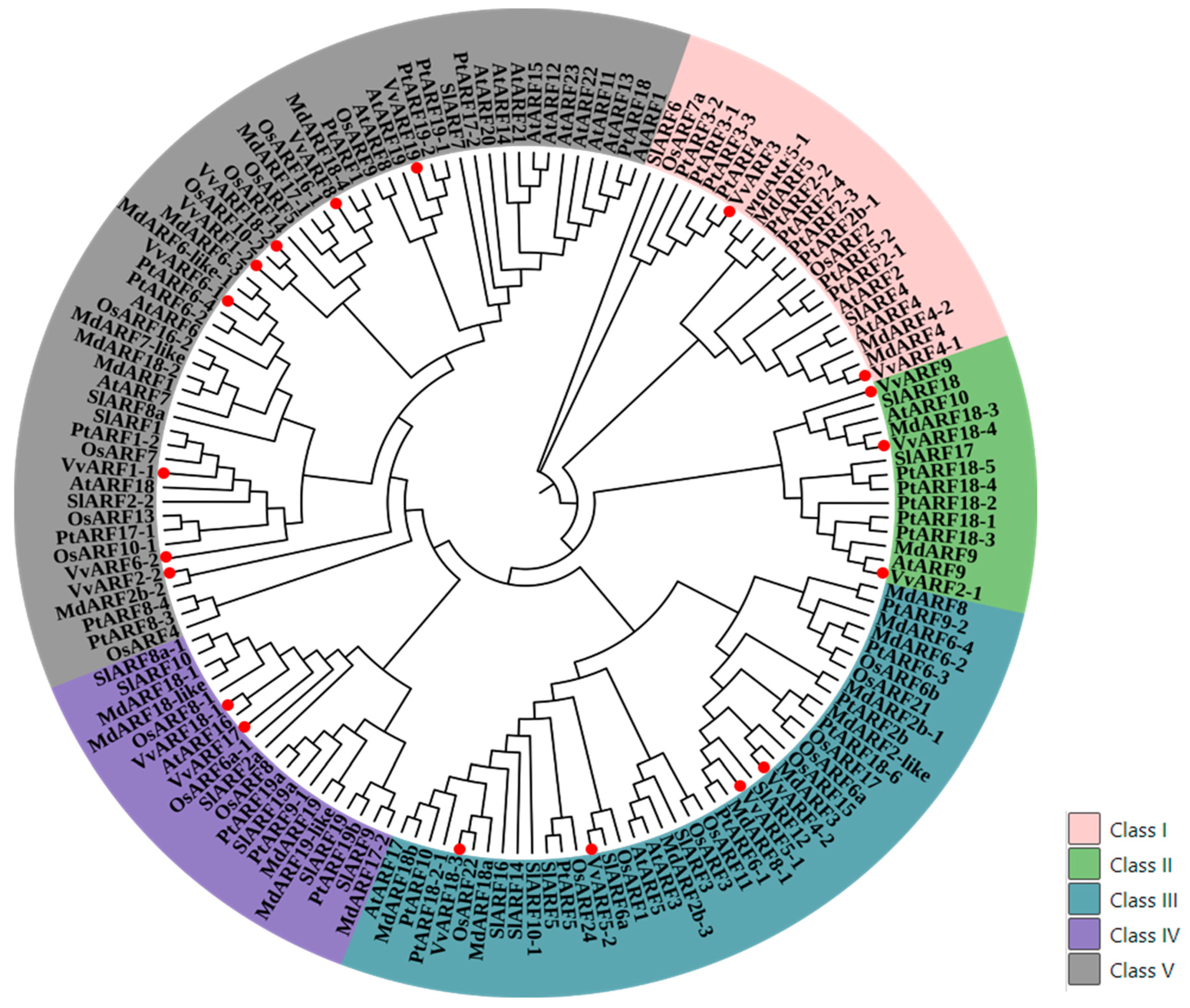
2.3. Conservative and Evolutionary Analysis of VvARF Family across Diverse Plant Species
2.4. Screening of Hormone Cis-Elements in the Promoters of VvARFs
2.5. Gene Ontology and Pathway Mapping of VvARFs
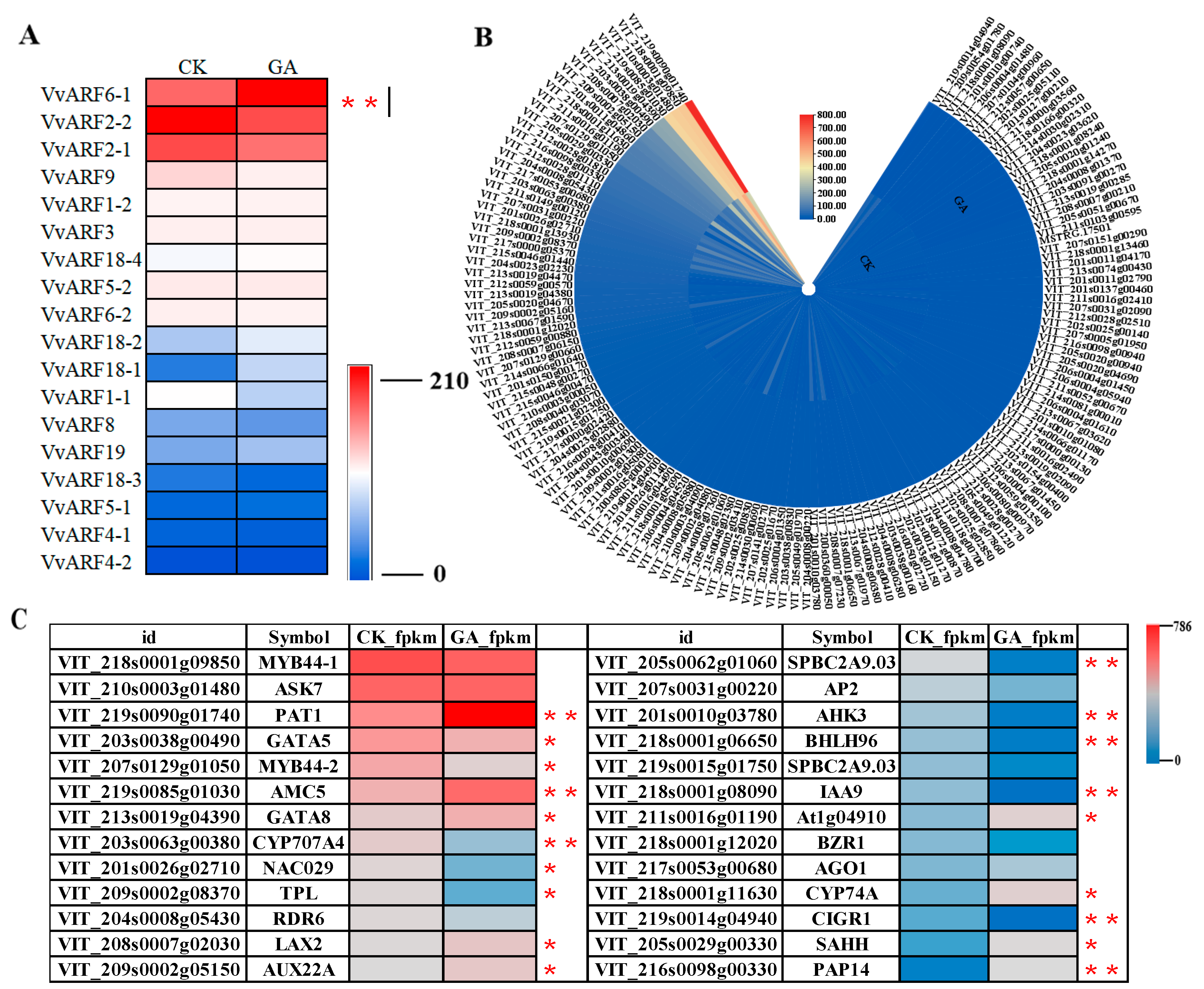
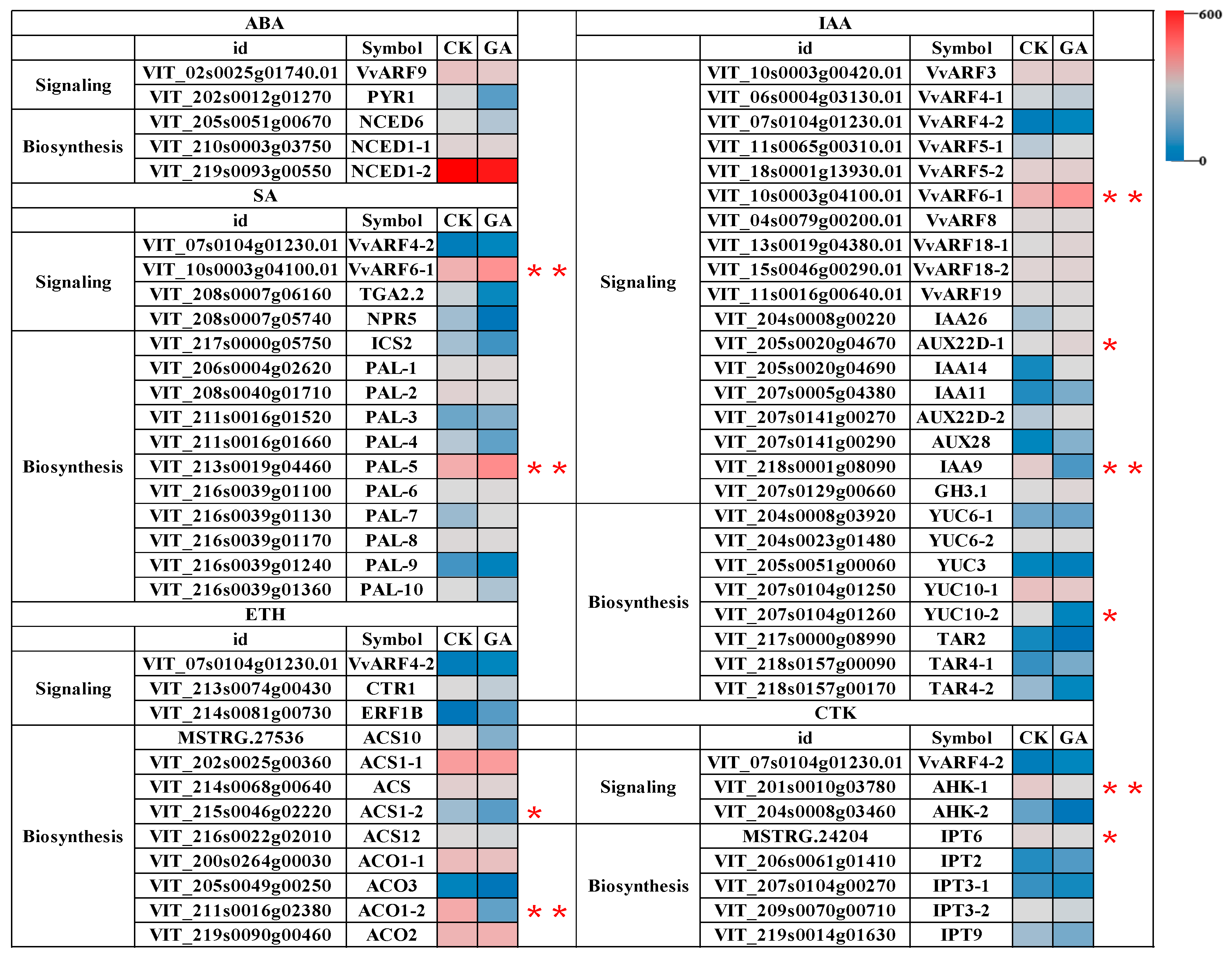
2.6. Differential Expression Profiles of VvARFs and Their Interacted Genes during GA-Induced Grape Parthenocarpy
2.7. Regulatory Network of VvARFs in Response to GA-Mediated Multi-Hormone Signals
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Determination of Total Soluble Solids, Sugar and Acidity Content
4.3. Phylogenetic Analysis
4.4. Determination of Exon/Intron Structures of VvARFs
4.5. Analysis of Cis-Acting Elements in Promoter Regions of VvARFs
4.6. RNA Extraction, Library Construction and Sequencing
4.7. Analysis of Differentially Expressed Genes
4.8. Functional Annotation and Enrichment Analysis of Differentially Expressed Genes in VvARFs
4.9. Predication of Protein-Protein Interaction
4.10. Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR) Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhang, W.; Wang, C.; Zhu, X.; Ma, C.; Wang, W.; Leng, X.; Zheng, T.; Fang, J. Identification of the whole grape genome DELLA protein gene family and its response to exogenous gibberellin to regulate grape fruit development. China Agric. Sci. 2018, 51, 3130–3146. [Google Scholar]
- Shiri, Y.; Solouki, M.; Ebrahimie, E.; Emamjomeh, A.; Zahiri, J. Gibberellin causes wide transcriptional modifications in the early stage of grape cluster development. Genomics 2020, 112, 820–830. [Google Scholar] [CrossRef]
- Wang, X.; Wang, C.; Fang, J.; Shangguan, L.; Liu, H.; Han, J. Research progress in the regulation of parthenocarpy in fruit trees by plant hormones. Chin. J. Plant Physiol. 2012, 48, 629–636. [Google Scholar] [CrossRef]
- Hu, J.; Israeli, A.; Ori, N.; Sun, T.P. The Interaction between DELLA and ARF/IAA Mediates Crosstalk between Gibberellin and Auxin Signaling to Control Fruit Initiation in Tomato. Plant Cell 2018, 30, 1710–1728. [Google Scholar] [CrossRef]
- Kim, J.S.; Ezura, K.; Lee, J.; Kojima, M.; Takebayashi, Y.; Sakakibara, H.; Ariizumi, T.; Ezura, H. The inhibition of SlIAA9 mimics an increase in endogenous auxin and mediates changes in auxin and gibberellin signalling during parthenocarpic fruit development in tomato. J. Plant Physiol. 2020, 252, 153238. [Google Scholar] [CrossRef]
- Zhang, W.; Abdelrahman, M.; Jiu, S.; Guan, L.; Han, J.; Zheng, T.; Jia, H.; Song, C.; Fang, J.; Wang, C. VvmiR160s/VvARFs interaction and their spatio-temporal expression/cleavage products during GA-induced grape parthenocarpy. BMC Plant Biol. 2019, 19, 111. [Google Scholar] [CrossRef]
- Cancé, C.; Martin-Arevalillo, R.; Boubekeur, K.; Dumas, R. Auxin response factors are keys to the many auxin doors. New Phytol. 2022, 235, 402–419. [Google Scholar] [CrossRef]
- Hagen, G.; Guilfoyle, T. Auxin-responsive gene expression: Genes, promoters and regulatory factors. Plant Mol. Biol. 2002, 49, 373–385. [Google Scholar] [CrossRef]
- Roosjen, M.; Paque, S.; Weijers, D. Auxin Response Factors: Output control in auxin biology. J. Exp. Bot. 2018, 69, 179–188. [Google Scholar] [CrossRef]
- Tiwari, S.B.; Hagen, G.; Guilfoyle, T. The roles of auxin response factor domains in auxin-responsive transcription. Plant Cell. 2003, 15, 533–543. [Google Scholar] [CrossRef]
- Da Silva, E.M.; Gffe Silva Bidoia, D.B.; da Silva Azevedo, M.; de Jesus, F.A.; Pino, L.E.; Peres, L.E.P.; Carrera, E.; López-Díaz, I.; Nogueira, F.T.S. microRNA159-targeted SlGAMYB transcription factors are required for fruit set in tomato. Plant J. 2017, 92, 95–109. [Google Scholar] [CrossRef] [PubMed]
- Goetz, M.; Hooper, L.C.; Johnson, S.D.; Rodrigues, J.C.; Vivian-Smith, A.; Koltunow, A.M. Expression of aberrant forms of AUXIN RESPONSE FACTOR8 stimulates parthenocarpy in Arabidopsis and tomato. Plant Physiol. 2007, 145, 351–366. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y. Cloning and Identification of Tomato Sly-miR167 and Its Effect on Fruit Setting and Parthenocarpy; Chongqing University: Chongqing, China, 2010. [Google Scholar]
- Hu, J.; Su, H.; Cao, H.; Wei, H.; Fu, X.; Jiang, X.; Song, Q.; He, X.; Xu, C.; Luo, K. AUXIN RESPONSE FACTOR7 integrates gibberellin and auxin signaling via interactions between DELLA and AUX/IAA proteins to regulate cambial activity in poplar. Plant Cell 2022, 34, 2688–2707. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Sittmann, J.; Guo, L.; Xiao, Y.; Huang, X.; Pulapaka, A.; Liu, Z. Gibberellin and auxin signaling genes RGA1 and ARF8 repress accessory fruit initiation in diploid strawberry. Plant Physiol. 2021, 185, 1059–1075. [Google Scholar] [CrossRef]
- Diao, D.; Hu, X.; Guan, D.; Wang, W.; Yang, H.; Liu, Y. Genome-wide identification of the ARF (auxin response factor) gene family in peach and their expression analysis. Mol. Biol. Rep. 2020, 47, 4331–4344. [Google Scholar] [CrossRef]
- Kalluri, U.; Difazio, C.S.P.; Brunner, A.M.; Tuskan, G.A. Genome-wide analysis of Aux/IAA and ARF gene families in Populus trichocarpa. BMC Plant Biol. 2007, 7, 59. [Google Scholar] [CrossRef]
- Okushima, Y.; Overvoorde, P.J.; Arima, K.; Alonso, J.M.; Chan, A.; Chang, C.; Ecker, J.R.; Hughes, B.; Lui, A.; Nguyen, D.; et al. Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: Unique and overlapping functions of ARF7 and ARF19. Plant Cell. 2005, 17, 444–463. [Google Scholar] [CrossRef]
- Wang, D.; Pei, K.; Fu, Y.; Sun, Z.; Li, S.; Liu, H.; Tang, K.; Han, B.; Tao, Y. Genome-wide analysis of the auxin response factors (ARF) gene family in rice (Oryza sativa). Gene 2007, 394, 13–24. [Google Scholar] [CrossRef]
- Wu, J.; Wang, F.; Cheng, L.; Kong, F.; Peng, Z.; Liu, S.; Yu, X.; Lu, G. Identification, isolation and expression analysis of auxin response factor (ARF) genes in Solanum lycopersicum. Plant Cell Rep. 2011, 30, 2059–2073. [Google Scholar] [CrossRef]
- Liu, Z.; Miao, L.; Huo, R.; Song, X.; Johnson, C.; Kong, L.; Sundaresan, V.; Yu, X. ARF2-ARF4 and ARF5 are Essential for Female and Male Gametophyte Development in Arabidopsis. Plantand Cell Physiol. 2018, 59, 179–189. [Google Scholar] [CrossRef]
- Promchuea, S.; Zhu, Y.; Chen, Z.; Zhang, J.; Gong, Z. ARF2 coordinates with PLETHORAs and PINs to orchestrate ABA-mediated root meristem activity in Arabidopsis. J. Integr. Plant Biol. 2017, 59, 30–43. [Google Scholar] [CrossRef]
- Kalve, S.; Sizani, B.L.; Markakis, M.N.; Helsmoortel, C.; Vandeweyer, G.; Laukens, K.; Sommen, M.; Naulaerts, S.; Vissenberg, K.; Prinsen, E.; et al. Osmotic stress inhibits leaf growth of Arabidopsis thaliana by enhancing ARF-mediated auxin responses. New Phytol. 2020, 226, 1766–1780. [Google Scholar] [CrossRef] [PubMed]
- Schruff, M.C.; Spielman, M.; Tiwari, S.; Adams, S.; Fenby, N.; Scott, R.J. The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 2006, 133, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Reed, J.W.; Wu, M.F.; Reeves, P.H.; Hodgens, C.; Yadav, V.; Hayes, S.; Pierik, R. Three Auxin Response Factors Promote Hypocotyl Elongation. Plant Physiol. 2018, 178, 864–875. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Jin, S.; Li, P.; Jiang, X.; Li, Y.; Hou, B. Ectopic expression of UGT84A2 delayed flowering by indole-3-butyric acid-mediated transcriptional repression of ARF6 and ARF8 genes in Arabidopsis. Plant Cell Rep. 2017, 36, 1995–2006. [Google Scholar] [CrossRef]
- De Jong, M.; Mariani, C.; Vriezen, W.H. The role of auxin and gibberellin in tomato fruit set. J. Exp. Bot. 2009, 60, 1523–1532. [Google Scholar] [CrossRef]
- Nagpal, P.; Ellis, C.M.; Weber, H.; Ploense, S.E.; Barkawi, L.S.; Guilfoyle, T.J.; Hagen, G.; Alonso, J.M.; Cohen, J.D.; Farmer, E.E.; et al. Auxin response factors ARF6 and ARF8 promote jasmonic acid production and flower maturation. Development 2005, 132, 4107–4118. [Google Scholar] [CrossRef]
- Liu, K.; Li, Y.; Chen, X.; Li, L.; Liu, K.; Zhao, H.; Wang, Y.; Han, S. ERF72 interacts with ARF6 and BZR1 to regulate hypocotyl elongation in Arabidopsis. J. Exp. Bot. 2018, 69, 3933–3947. [Google Scholar] [CrossRef]
- Zhang, T.; Li, W.; Xie, R.; Xu, L.; Zhou, Y.; Li, H.; Yuan, C.; Zheng, X.; Xiao, L.; Liu, K. CpARF2 and CpEIL1 interact to mediate auxin-ethylene interaction and regulate fruit ripening in papaya. Plant J. Cell Mol. Biol. 2020, 103, 1318–1337. [Google Scholar] [CrossRef]
- Lu, X.; Zhang, X.; Zhang, D.; Ma, M.T.; Wu, D. Effects of harvest time and fruit bagging treatments on the fruit quality of ‘Red Fuji’ grown in the cold highland in southwest China. J. Fruit Sci. 2017, 34, 8. [Google Scholar]
- Robinson, M.; McCarthy, D.; Smyth, G. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- Singh, V.K.; Rajkumar, M.S.; Garg, R.; Jain, M. Genome-wide identification and co-expression network analysis provide insights into the roles of auxin response factor gene family in chickpea. Sci. Rep. 2017, 7, 10895. [Google Scholar] [CrossRef]
- Wan, S.; Li, W.; Zhu, Y.; Liu, Z.; Huang, W.; Zhan, J. Genome-wide identification, characterization and expression analysis of the auxin response factor gene family in Vitis vinifera. Plant Cell Rep. 2014, 33, 1365–1375. [Google Scholar] [CrossRef] [PubMed]
- Oh, E.; Zhu, J.; Bai, M.; Arenhart, R.A.; Sun, Y.; Wang, Z.Y. Cell elongation is regulated through a central circuit of interacting transcription factors in the Arabidopsis hypocotyl. Elife 2014, 3, e03031. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Xie, Z.; Hu, C.; Zhang, J. A Review of Auxin Response Factors (ARFs) in Plants. Front. Plant Sci. 2016, 7, 47. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Fang, T.; Zhang, Y.; Zhang, M.; Zeng, L. Genome-Wide Identification and Expression Analysis of Auxin Response Factor (ARF) Gene Family in Longan (Dimocarpus longan L.). Plants 2020, 9, 221. [Google Scholar] [CrossRef]
- Li, W.; Chen, F.; Wang, Y.; Zheng, H.; Yi, Q.; Ren, Y.; Gao, J. Genome-wide identification and functional analysis of ARF transcription factors in Brassica juncea var. tumida. PLoS ONE. 2020, 15, e0232039. [Google Scholar] [CrossRef]
- Galvão, V.C.; Collani, S.; Horrer, D.; Schmid, M. Gibberellic acid signaling is required for ambient temperature-mediated induction of flowering in Arabidopsis thaliana. Plant J. 2015, 84, 949–962. [Google Scholar] [CrossRef]
- Niwa, T.; Suzuki, T.; Takebayashi, Y.; Ishiguro, R.; Higashiyama, T.; Sakakibara, H.; Ishiguro, S. Jasmonic acid facilitates flower opening and floral organ development through the upregulated expression of SlMYB21 transcription factor in tomato. Biosci. Biotechnol. Biochem. 2018, 82, 292–303. [Google Scholar] [CrossRef]
- Yamaguchi, N.; Winter, C.M.; Wu, M.F.; Kanno, Y.; Yamaguchi, A.; Seo, M.; Wagner, D. Gibberellin acts positively then negatively to control onset of flower formation in Arabidopsis. Science 2014, 344, 638–641. [Google Scholar] [CrossRef]
- Adenot, X.; Elmayan, T.; Lauressergues, D.; Boutet, S.; Bouché, N.; Gasciolli, V.; Vaucheret, H. DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr. Biol. 2006, 16, 927–932. [Google Scholar] [CrossRef] [PubMed]
- Hobecker, K.V.; Reynoso, M.A.; Bustos-Sanmamed, P.; Wen, J.; Mysore, K.S.; Crespi, M.; Blanco, F.A.; Zanetti, M.E. The MicroRNA390/TAS3 Pathway Mediates Symbiotic Nodulation and Lateral Root Growth. Plant Physiol. 2017, 174, 2469–2486. [Google Scholar] [CrossRef] [PubMed]
- Lavenus, J.; Goh, T.; Roberts, I.; Guyomarc’h, S.; Lucas, M.; De Smet, I.; Fukaki, H.; Beeckman, T.; Bennett, M.; Laplaze, L. Lateral root development in Arabidopsis: Fifty shades of auxin. Trends Plant Sci. 2013, 18, 450–458. [Google Scholar] [CrossRef] [PubMed]
- Marin, E.; Jouannet, V.; Herz, A.; Lokerse, A.S.; Weijers, D.; Vaucheret, H.; Nussaume, L.; Crespi, M.D.; Maizel, A. miR390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell. 2010, 22, 1104–1117. [Google Scholar] [CrossRef]
- Wang, H.; Jones, B.; Li, Z.; Frasse, P.; Delalande, C.; Regad, F.; Chaabouni, S.; Latché, A.; Pech, J.C.; Bouzayen, M. The tomato Aux/IAA transcription factor IAA9 is involved in fruit development and leaf morphogenesis. Plant Cell. 2005, 17, 2676–2692. [Google Scholar] [CrossRef]
- Vivian-Smith, A.; Luo, M.; Chaudhury, A.; Koltunow, A. Fruit development is actively restricted in the absence of fertilization in Arabidopsis. Development 2001, 128, 2321–2331. [Google Scholar] [CrossRef]
- Du, L.; Bao, C.; Hu, T.; Zhu, Q.; Hu, H.; He, Q.; Mao, W. SmARF8, a transcription factor involved in parthenocarpy in eggplant. Mol. Genet. Genomics. 2016, 291, 93–105. [Google Scholar] [CrossRef]
- Dos Santos, R.C.; Nietsche, S.; Pereira, M.C.T.; Ribeiro, L.M.; Mercadante-Simões, M.O.; Carneiro Dos Santos, B.H. Atemoya fruit development and cytological aspects of GA3-induced growth and parthenocarpy. J. Protoplasma 2019, 256, 1345–1360. [Google Scholar] [CrossRef]
- Wang, C.; Jogaiah, S.; Zhang, W.; Abdelrahman, M.; Fang, J. Spatio-temporal expression of miRNA159 family members and their GAMYB target gene during the modulation of gibberellin-induced grapevine parthenocarpy. J. Exp. Bot. 2018, 69, 3639–3650. [Google Scholar] [CrossRef]
- Bai, Y.; Wang, W.; Dong, T.; Guan, L.; Su, Z.; Jia, H.; Fang, J. Vvi-miR160s Mediates VvARF18 Response to Gibberellin Regulating Grape Seed Development. Sci. Agric. Sin. 2020, 53, 14. [Google Scholar]
- Wang, H.; Wu, T.; Liu, J.; Cong, L.; Zhu, Y.; Zhai, R.; Yang, C.; Wang, Z.; Ma, F.; Xu, L. PbGA20ox2 Regulates Fruit Set and Induces Parthenocarpy by Enhancing GA 4 Content. Front. Plant Sci. 2020, 11, 113. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Yamamuro, C. Interplays between auxin and GA signaling coordinate early fruit development. Hortic. Res. 2022, 9, uhab078. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Chen, B.; Xia, X.; Mao, W.; Shi, K.; Zhou, Y.; Yu, J. Cytokinin-induced parthenocarpic fruit development in tomato is partly dependent on enhanced gibberellin and auxin biosynthesis. PLoS ONE. 2013, 8, e70080. [Google Scholar] [CrossRef] [PubMed]
- De Jong, M.; Wolters-Arts, M.; Feron, R.; Mariani, C.; Vriezen, W.H. The Solanum lycopersicum auxin response factor 7 (SlARF7) regulates auxin signaling during tomato fruit set and development. Plant J. 2009, 57, 160–170. [Google Scholar] [CrossRef]
- Liu, N.; Wu, S.; Van Houten, J.; Wang, Y.; Ding, B.; Fei, Z.; Clarke, T.H.; Reed, J.W.; Knaap, E. Down-regulation of AUXIN RESPONSE FACTORS 6 and 8 by microRNA 167 leads to floral development defects and female sterility in tomato. J. Exp. Bot. 2014, 65, 2507–2520. [Google Scholar] [CrossRef]
- Clepet, C.; Devani, R.S.; Boumlik, R.; Hao, Y.; Morin, H.; Marcel, F.; Verdenaud, M.; Mania, B.; Brisou, G.; Citerne, S.; et al. The miR166-SlHB15A regulatory module controls ovule development and parthenocarpic fruit set under adverse temperatures in tomato. Mol Plant. 2021, 14, 1185–1198. [Google Scholar] [CrossRef]
- Cao, J.; Jiang, W.; Zhao, Y. Post Harvest Physiological and Biochemical Experiment Guidance of Fruits and Vegetables; Woodhead Publishing: Cambridge, UK, 2007. [Google Scholar]
- Zhang, Y.; Wang, C.; Yu, H.; Cai, B.; Fang, J. Screening of RNA Extraction Methods for Various Grapevine Organs and Tissues. Acta Agric. Boreali-Occident. Sin. 2010, 19, 135–140. [Google Scholar]




| Gene ID | Symbol | Pathway | K_ID | GO Process | |
|---|---|---|---|---|---|
| I | VIT_18s0001g04180 | VvARF17 | GO:0009987//cellular process | ||
| VIT_208s0040g01810 | VvARF18-3 | GO:0009755//hormone-mediated signaling pathway | |||
| VIT_213s0019g04380 | VvARF18-1 | ko04075//Plant hormone signal transduction | K14486 | GO:0007275//multicellular organism development | |
| VIT_206s0004g02750 | VvARF18-4 | GO:0009718//anthocyanin-containing compound biosynthetic process | |||
| II | VIT_218s0001g13930 | VvARF5-2 | ko04075//Plant hormone signal transduction | K14486 | GO:0000578//embryonic axis specification |
| VIT_211s0016g00640 | VvARF19 | ko04075//Plant hormone signal transduction | K14486 | GO:0009755//hormone-mediated signaling pathway | |
| VIT_211s0065g00310 | VvARF5-1 | ko04075//Plant hormone signal transduction | K14486 | GO:0009755//hormone-mediated signaling pathway | |
| VIT_204s0079g00200 | VvARF8 | - | |||
| VIT_212s0028g01170 | VvARF6-2 | ko04075//Plant hormone signal transduction | K14486 | GO:0001101//response to acid chemical | |
| VIT_210s0003g04100 | VvARF6-1 | GO:0003002//regionalization | |||
| III | VIT_210s0003g00420 | VvARF3 | ko04075//Plant hormone signal transduction | K14486 | GO:0001708//cell fate specification |
| VIT_206s0004g03130 | VvARF4-1 | GO:0001708//cell fate specification | |||
| VIT_207s0104g01230 | VvARF4-2 | GO:0009987//cellular process | |||
| VIT_217s0000g00320 | VvARF2-2 | GO:0001101//response to acid chemical | |||
| VIT_201s0244g00150 | VvARF2-1 | GO:0006355//regulation of transcription, DNA-templated | |||
| VIT_218s0089g00910 | VvARF1-2 | ko04075//Plant hormone signal transduction | K14486 | GO:0009755//hormone-mediated signaling pathway | |
| VIT_212s0035g01800 | VvARF1-1 | ko04075//Plant hormone signal transduction | K14486 | GO:0009755//hormone-mediated signaling pathway | |
| VIT_215s0046g00290 | VvARF18-2 | GO:0009725//response to hormone | |||
| VIT_202s0025g01740 | VvARF9 | ko04075//Plant hormone signal transduction | K14486 | GO:0009755//hormone-mediated signaling pathway |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sheng, Z.; Xuan, X.; Wang, F.; Sadeghnezhad, E.; Gong, P.; Xiao, Y.; Dong, T.; Zhang, P.; Wang, X.; Fang, J.; et al. Identification and Characterization of AUXIN Response Factor Gene Family Reveals Their Regulatory Network to Respond the Multi-Hormones Crosstalk during GA-Induced Grape Parthenocarpic Berry. Int. J. Mol. Sci. 2022, 23, 11108. https://doi.org/10.3390/ijms231911108
Sheng Z, Xuan X, Wang F, Sadeghnezhad E, Gong P, Xiao Y, Dong T, Zhang P, Wang X, Fang J, et al. Identification and Characterization of AUXIN Response Factor Gene Family Reveals Their Regulatory Network to Respond the Multi-Hormones Crosstalk during GA-Induced Grape Parthenocarpic Berry. International Journal of Molecular Sciences. 2022; 23(19):11108. https://doi.org/10.3390/ijms231911108
Chicago/Turabian StyleSheng, Zilu, Xuxian Xuan, Fei Wang, Ehsan Sadeghnezhad, Peijie Gong, Yingke Xiao, Tianyu Dong, Peian Zhang, Xicheng Wang, Jinggui Fang, and et al. 2022. "Identification and Characterization of AUXIN Response Factor Gene Family Reveals Their Regulatory Network to Respond the Multi-Hormones Crosstalk during GA-Induced Grape Parthenocarpic Berry" International Journal of Molecular Sciences 23, no. 19: 11108. https://doi.org/10.3390/ijms231911108
APA StyleSheng, Z., Xuan, X., Wang, F., Sadeghnezhad, E., Gong, P., Xiao, Y., Dong, T., Zhang, P., Wang, X., Fang, J., & Wang, C. (2022). Identification and Characterization of AUXIN Response Factor Gene Family Reveals Their Regulatory Network to Respond the Multi-Hormones Crosstalk during GA-Induced Grape Parthenocarpic Berry. International Journal of Molecular Sciences, 23(19), 11108. https://doi.org/10.3390/ijms231911108






