Loss of Protein Stability and Function Caused by P228L Variation in NADPH-Cytochrome P450 Reductase Linked to Lower Testosterone Levels
Abstract
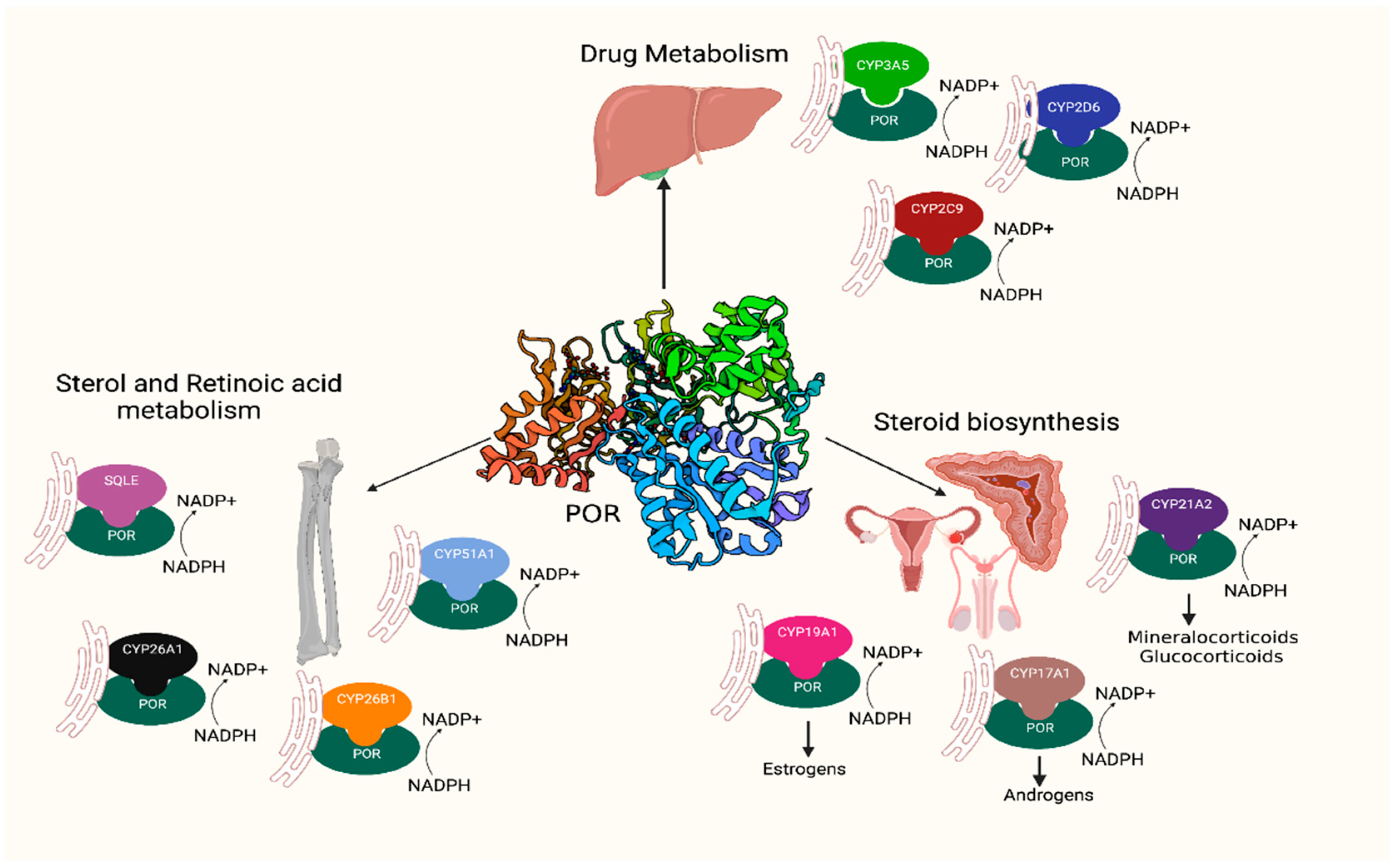
1. Introduction
2. Results
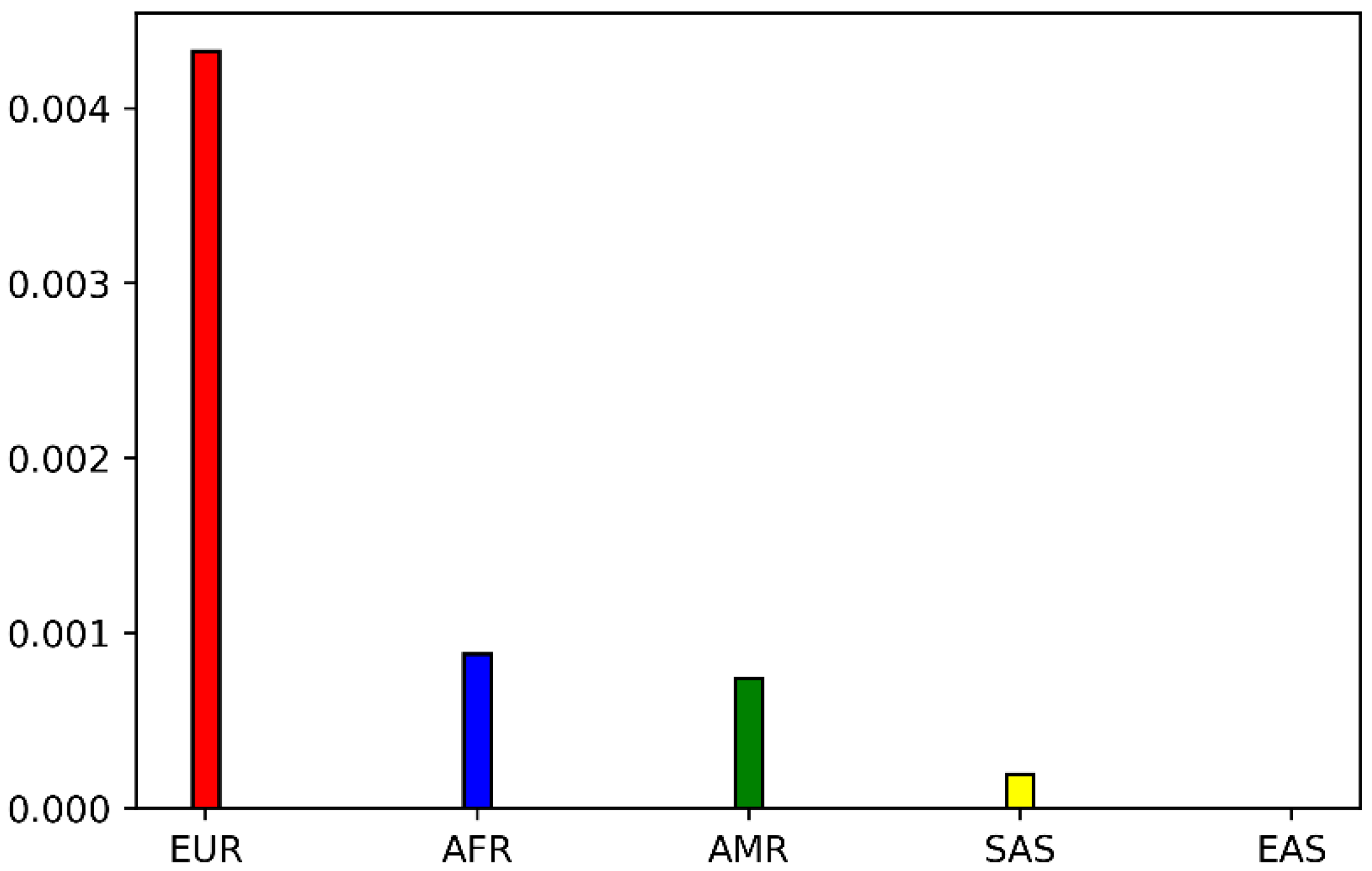
2.1. Genetic Distribution of POR P228L
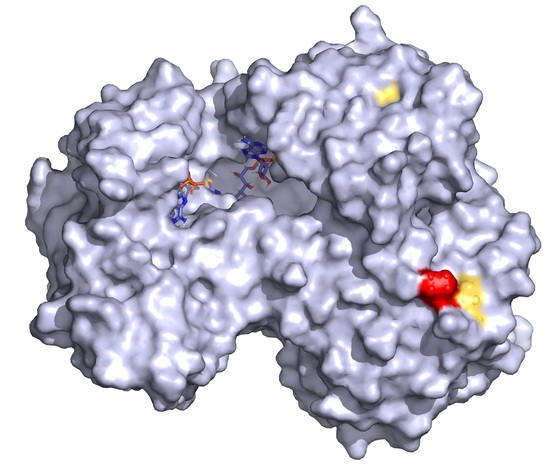
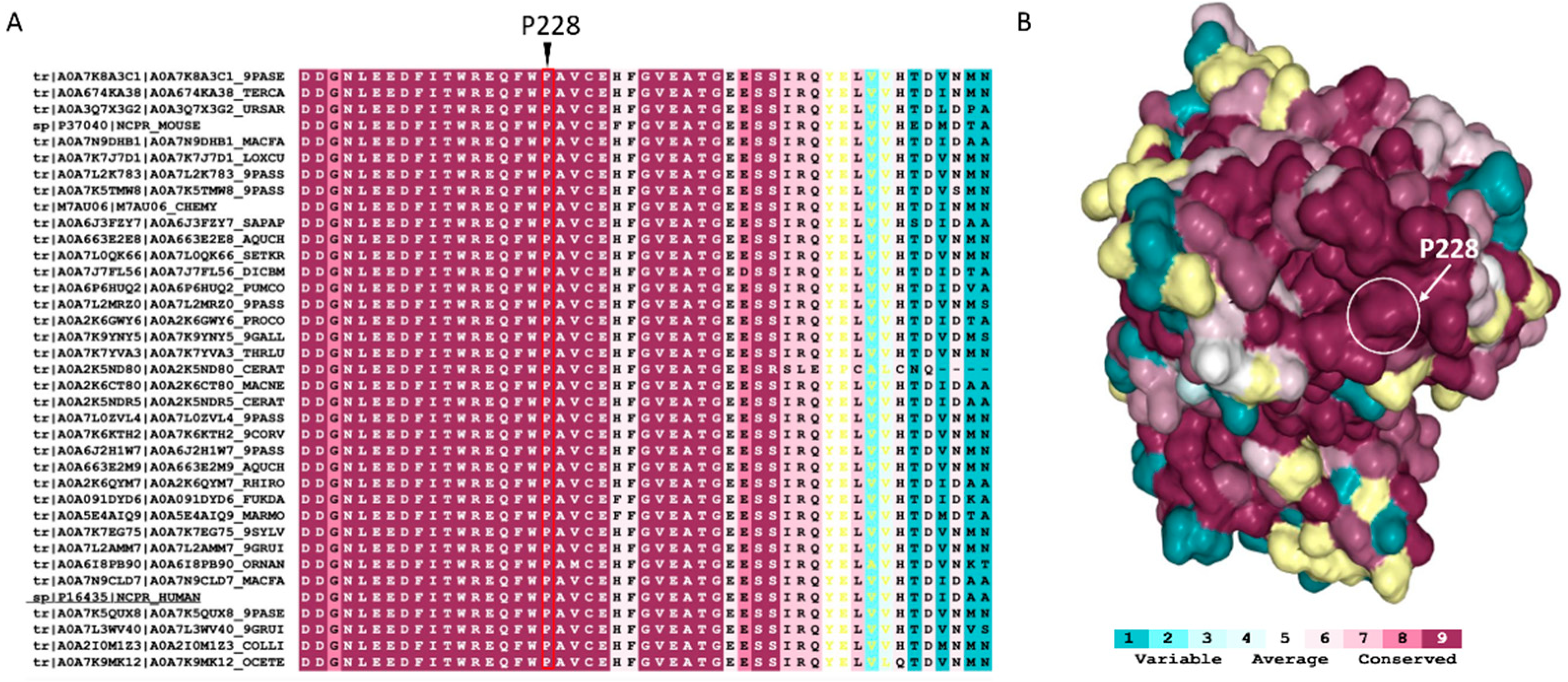
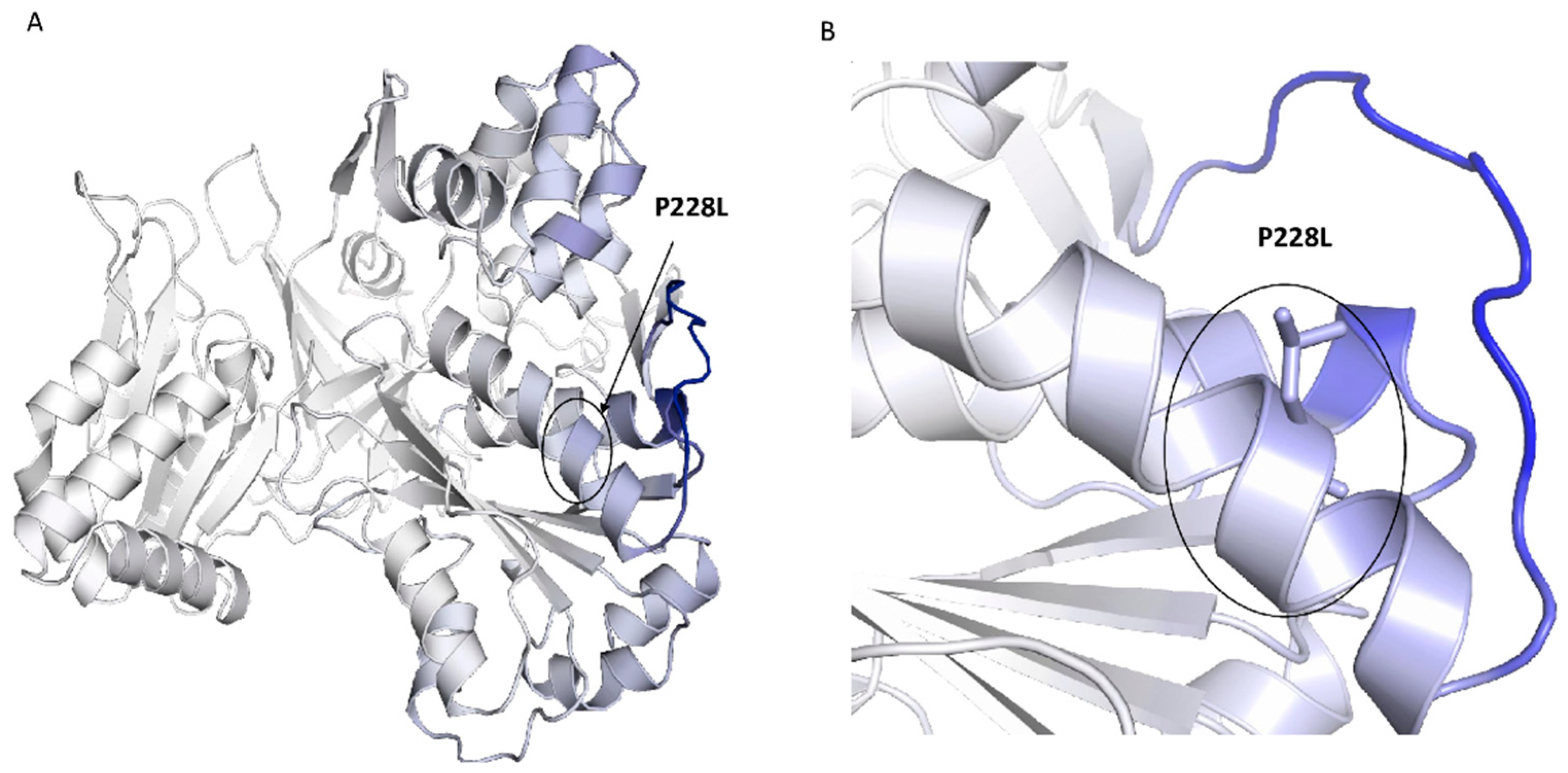
2.2. Conservation and Structural Analysis of POR
2.3. In Silico Predictions of POR P228L Stability
2.4. Flavin Content of WT and Mutant P228L Proteins
2.5. Stability of Mutant P228L Proteins
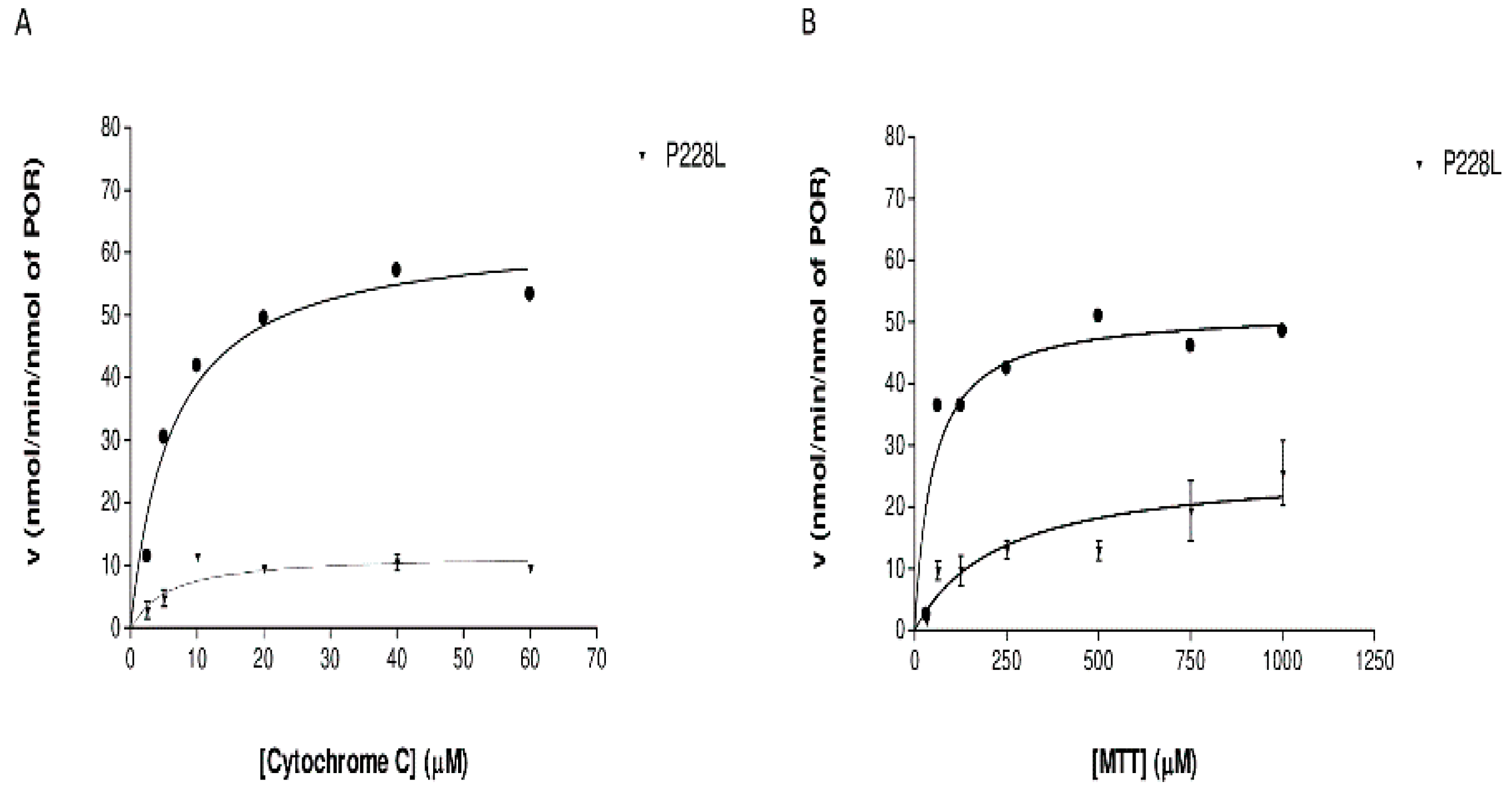
2.6. Effect of POR Variant P228L on Cytochrome c and MTT Reduction Activities
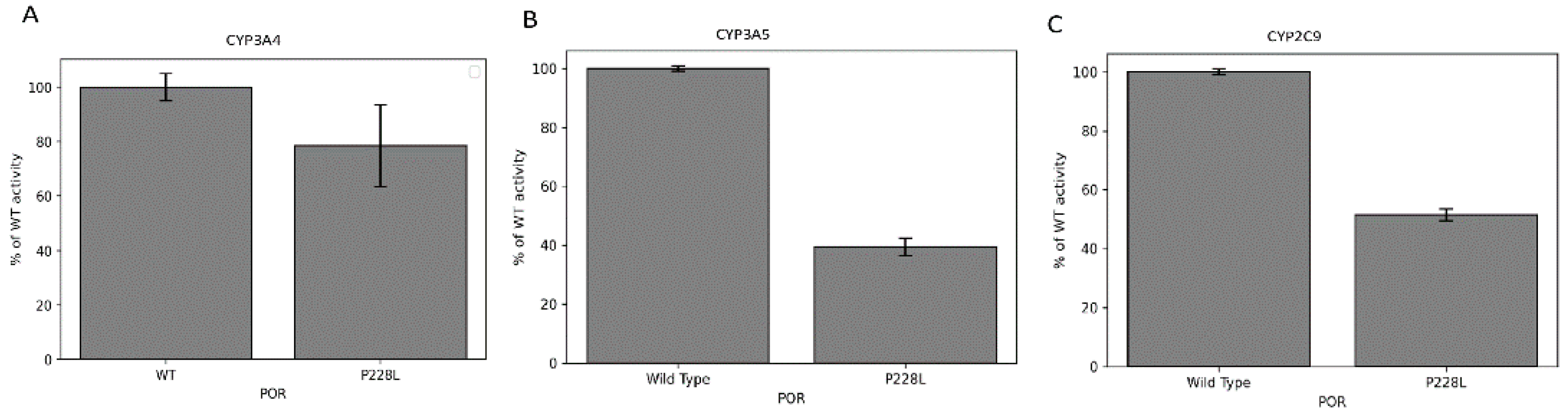
2.7. Activities of CYP3A4, CYP3A5, and CYP2C9 Enzymes with POR P228L
2.8. Activities of Steroid Metabolising CYP17A1 and CYP19A1 Enzymes with POR P228L
3. Discussion
4. Materials and Methods
4.1. Analysis of POR P228L Allele Frequencies
4.2. Conservation Analysis
4.3. In Silico Predictions of POR P228L Stability
4.4. Protein Structure Analysis of POR Variants
4.5. Expression of POR in E. coli
4.6. Purification of Recombinant Human POR from Isolated E. coli Membranes
4.7. Flavin Content
4.8. Fast Proteolysis Assay (FASTpp)
4.9. CPM Fluorescence Assay
4.10. POR Assays with Cytochrome c and MTT
4.11. Assay of Cytochrome P450 CYP3A4, CYP3A5, and CYP2C9 Activity in Reconstituted Liposomes
4.12. Assay of Cytochrome P450 CYP17A1 and CYP19A1
4.13. Statistical Analysis of Results
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pandey, A.V.; Flück, C.E. NADPH P450 oxidoreductase: Structure, function, and pathology of diseases. Pharm. Ther. 2013, 138, 229–254. [Google Scholar] [CrossRef] [PubMed]
- Miller, W.L.; Huang, N.; Flück, C.E.; Pandey, A.V. P450 oxidoreductase deficiency. Lancet 2004, 364, 1663. [Google Scholar] [CrossRef]
- Flück, C.E.; Tajima, T.; Pandey, A.V.; Arlt, W.; Okuhara, K.; Verge, C.F.; Jabs, E.W.; Mendonca, B.B.; Fujieda, K.; Miller, W.L. Mutant P450 oxidoreductase causes disordered steroidogenesis with and without Antley-Bixler syndrome. Nat. Genet. 2004, 36, 228–230. [Google Scholar] [CrossRef] [PubMed]
- Huang, N.; Pandey, A.V.; Agrawal, V.; Reardon, W.; Lapunzina, P.D.; Mowat, D.; Jabs, E.W.; Van Vliet, G.; Sack, J.; Flück, C.E.; et al. Diversity and function of mutations in p450 oxidoreductase in patients with Antley-Bixler syndrome and disordered steroidogenesis. Am. J. Hum. Genet. 2005, 76, 729–749. [Google Scholar] [CrossRef] [PubMed]
- Burkhard, F.Z.; Parween, S.; Udhane, S.S.; Flück, C.E.; Pandey, A.V. P450 Oxidoreductase deficiency: Analysis of mutations and polymorphisms. J. Steroid Biochem. Mol. Biol. 2017, 165, 38–50. [Google Scholar] [CrossRef] [PubMed]
- Jensen, S.B.; Thodberg, S.; Parween, S.; Moses, M.E.; Hansen, C.C.; Thomsen, J.; Sletfjerding, M.B.; Knudsen, C.; Del Giudice, R.; Lund, P.M.; et al. Biased cytochrome P450-mediated metabolism via small-molecule ligands binding P450 oxidoreductase. Nat. Commun. 2021, 12, 2260. [Google Scholar] [CrossRef] [PubMed]
- Iijima, S.; Ohishi, A.; Ohzeki, T. Cytochrome P450 oxidoreductase deficiency with Antley-Bixler syndrome: Steroidogenic capacities. J. Pediatr. Endocrinol. Metab. 2009, 22, 469–475. [Google Scholar] [CrossRef]
- Fagerberg, L.; Hallstrom, B.M.; Oksvold, P.; Kampf, C.; Djureinovic, D.; Odeberg, J.; Habuka, M.; Tahmasebpoor, S.; Danielsson, A.; Edlund, K.; et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell Proteom. 2014, 13, 397–406. [Google Scholar] [CrossRef]
- Fukami, M.; Nishimura, G.; Homma, K.; Nagai, T.; Hanaki, K.; Uematsu, A.; Ishii, T.; Numakura, C.; Sawada, H.; Nakacho, M.; et al. Cytochrome P450 Oxidoreductase Deficiency: Identification and Characterization of Biallelic Mutations and Genotype-Phenotype Correlations in 35 Japanese Patients. J. Clin. Endocrinol. Metab. 2009, 94, 1723–1731. [Google Scholar] [CrossRef]
- Adachi, M.; Tachibana, K.; Asakura, Y.; Yamamoto, T.; Hanaki, K.; Oka, A. Compound heterozygous mutations of cytochrome P450 oxidoreductase gene (POR) in two patients with Antley-Bixler syndrome. Am. J. Med. Genet. A 2004, 128A, 333–339. [Google Scholar] [CrossRef]
- Pandey, A.V.; Fluck, C.E.; Huang, N.; Tajima, T.; Fujieda, K.; Miller, W.L. P450 oxidoreductase deficiency: A new disorder of steroidogenesis affecting all microsomal P450 enzymes. Endocr Res. 2004, 30, 881–888. [Google Scholar] [CrossRef] [PubMed]
- McCammon, K.M.; Panda, S.P.; Xia, C.; Kim, J.J.; Moutinho, D.; Kranendonk, M.; Auchus, R.J.; Lafer, E.M.; Ghosh, D.; Martasek, P.; et al. Instability of the Human Cytochrome P450 Reductase A287P Variant Is the Major Contributor to Its Antley-Bixler Syndrome-like Phenotype. J. Biol. Chem. 2016, 291, 20487–20502. [Google Scholar] [CrossRef] [PubMed]
- Riddick, D.S.; Ding, X.; Wolf, C.R.; Porter, T.D.; Pandey, A.V.; Zhang, Q.Y.; Gu, J.; Finn, R.D.; Ronseaux, S.; McLaughlin, L.A.; et al. NADPH-cytochrome P450 oxidoreductase: Roles in physiology, pharmacology, and toxicology. Drug Metab. Dispos. 2013, 41, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Parween, S.; Rojas Velazquez, M.N.; Udhane, S.S.; Kagawa, N.; Pandey, A.V. Variability in Loss of Multiple Enzyme Activities Due to the Human Genetic Variation P284T Located in the Flexible Hinge Region of NADPH Cytochrome P450 Oxidoreductase. Front. Pharmacol. 2019, 10, 1187. [Google Scholar] [CrossRef] [PubMed]
- Parween, S.; Roucher-Boulez, F.; Flück, C.E.; Lienhardt-Roussie, A.; Mallet, D.; Morel, Y.; Pandey, A.V. P450 Oxidoreductase Deficiency: Loss of Activity Caused by Protein Instability From a Novel L374H Mutation. J. Clin. Endocrinol. Metab. 2016, 101, 4789–4798. [Google Scholar] [CrossRef]
- Kumar, R.; Jayaraman, M.; Ramadas, K.; Chandrasekaran, A. Insight into the structural and functional analysis of the impact of missense mutation on cytochrome P450 oxidoreductase. J. Mol. Graph. Model. 2020, 100, 107708. [Google Scholar] [CrossRef]
- Flück, C.E.; Mullis, P.E.; Pandey, A.V. Reduction in hepatic drug metabolizing CYP3A4 activities caused by P450 oxidoreductase mutations identified in patients with disordered steroid metabolism. Biochem. Biophys. Res. Commun. 2010, 401, 149–153. [Google Scholar] [CrossRef]
- Flück, C.E.; Pandey, A.V. Impact on CYP19A1 activity by mutations in NADPH cytochrome P450 oxidoreductase. J. Steroid Biochem. Mol. Biol. 2017, 165 Pt A, 64–70. [Google Scholar] [CrossRef]
- Flück, C.E.; Parween, S.; Rojas Velazquez, M.N.; Pandey, A.V. Inhibition of placental CYP19A1 activity remains as a valid hypothesis for 46,XX virilization in P450 oxidoreductase deficiency. Proc. Natl. Acad. Sci. USA 2020, 117, 14632–14633. [Google Scholar] [CrossRef]
- Parween, S.; DiNardo, G.; Baj, F.; Zhang, C.; Gilardi, G.; Pandey, A.V. Differential effects of variations in human P450 oxidoreductase on the aromatase activity of CYP19A1 polymorphisms R264C and R264H. J. Steroid Biochem. Mol. Biol. 2020, 196, 105507. [Google Scholar] [CrossRef]
- Udhane, S.S.; Parween, S.; Kagawa, N.; Pandey, A.V. Altered CYP19A1 and CYP3A4 Activities Due to Mutations A115V, T142A, Q153R and P284L in the Human P450 Oxidoreductase. Front. Pharmacol. 2017, 8, 580. [Google Scholar] [CrossRef] [PubMed]
- Ruth, K.S.; Day, F.R.; Tyrrell, J.; Thompson, D.J.; Wood, A.R.; Mahajan, A.; Beaumont, R.N.; Wittemans, L.; Martin, S.; Busch, A.S.; et al. The Endometrial Cancer Association, C., Using human genetics to understand the disease impacts of testosterone in men and women. Nat. Med. 2020, 26, 252–258. [Google Scholar] [CrossRef] [PubMed]
- Sinnott-Armstrong, N.; Tanigawa, Y.; Amar, D.; Mars, N.; Benner, C.; Aguirre, M.; Venkataraman, G.R.; Wainberg, M.; Ollila, H.M.; Kiiskinen, T.; et al. Genetics of 35 blood and urine biomarkers in the UK Biobank. Nat. Genet. 2021, 53, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Ashkenazy, H.; Abadi, S.; Martz, E.; Chay, O.; Mayrose, I.; Pupko, T.; Ben-Tal, N. ConSurf 2016: An improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 2016, 44, W344–W350. [Google Scholar] [CrossRef]
- Aigrain, L.; Pompon, D.; Morera, S.; Truan, G. Structure of the open conformation of a functional chimeric NADPH cytochrome P450 reductase. EMBO Rep. 2009, 10, 742–747. [Google Scholar] [CrossRef]
- Xia, C.; Shen, A.L.; Duangkaew, P.; Kotewong, R.; Rongnoparut, P.; Feix, J.; Kim, J.P. Structural and Functional Studies of the Membrane-Binding Domain of NADPH-Cytochrome P450 Oxidoreductase. Biochemistry 2019, 58, 2408–2418. [Google Scholar] [CrossRef]
- Marohnic, C.C.; Panda, S.P.; Martasek, P.; Masters, B.S. Diminished FAD binding in the Y459H and V492E Antley-Bixler syndrome mutants of human cytochrome P450 reductase. J. Biol. Chem. 2006, 281, 35975–35982. [Google Scholar] [CrossRef]
- Zanger, U.M.; Schwab, M. Cytochrome P450 enzymes in drug metabolism: Regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacol. Ther. 2013, 138, 103–141. [Google Scholar] [CrossRef]
- Minde, D.P.; Maurice, M.M.; Rudiger, S.G. Determining biophysical protein stability in lysates by a fast proteolysis assay, FASTpp. PLoS ONE 2012, 7, e46147. [Google Scholar] [CrossRef]
- Park, C.; Marqusee, S. Pulse proteolysis: A simple method for quantitative determination of protein stability and ligand binding. Nat Methods. 2005, 2, 207–212. [Google Scholar] [CrossRef]
- Rwere, F.; Xia, C.; Im, S.; Haque, M.M.; Stuehr, D.J.; Waskell, L.; Kim, J.J. Mutants of Cytochrome P450 Reductase Lacking Either Gly-141 or Gly-143 Destabilize Its FMN Semiquinone. J. Biol. Chem. 2016, 291, 14639–14661. [Google Scholar] [CrossRef] [PubMed]
- Pandey, A.V.; Sproll, P. Pharmacogenomics of human P450 oxidoreductase. Front. Pharmacol. 2014, 5, 103. [Google Scholar] [CrossRef] [PubMed]
- Velazquez, M.N.R.; Parween, S.; Udhane, S.S.; Pandey, A.V. Variability in human drug metabolizing cytochrome P450 CYP2C9, CYP2C19 and CYP3A5 activities caused by genetic variations in cytochrome P450 oxidoreductase. Biochem. Biophys. Res. Commun. 2019, 515, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Pandey, A.V.; Flück, C.E.; Mullis, P.E. Altered heme catabolism by heme oxygenase-1 caused by mutations in human NADPH cytochrome P450 reductase. Biochem. Biophys. Res. Commun. 2010, 400, 374–378. [Google Scholar] [CrossRef] [PubMed]
- Flück, C.E.; Nicolo, C.; Pandey, A.V. Clinical, structural and functional implications of mutations and polymorphisms in human NADPH P450 oxidoreductase. Fund Clin. Pharm. 2007, 21, 399–410. [Google Scholar] [CrossRef]
- Bendl, J.; Musil, M.; Stourac, J.; Zendulka, J.; Damborsky, J.; Brezovsky, J. PredictSNP2: A Unified Platform for Accurately Evaluating SNP Effects by Exploiting the Different Characteristics of Variants in Distinct Genomic Regions. PLoS Comput. Biol. 2016, 12, e1004962. [Google Scholar] [CrossRef]
- Prado, M.J.; Singh, S.; Ligabue-Braun, R.; Meneghetti, B.V.; Rispoli, T.; Kopacek, C.; Monteiro, K.; Zaha, A.; Rossetti, M.L.R.; Pandey, A.V. Characterization of Mutations Causing CYP21A2 Deficiency in Brazilian and Portuguese Populations. Int. J. Mol. Sci. 2021, 23, 296. [Google Scholar] [CrossRef]
- Rodrigues, C.H.; Pires, D.E.; Ascher, D.B. DynaMut: Predicting the impact of mutations on protein conformation, flexibility and stability. Nucleic Acids Res. 2018, 46, W350–W355. [Google Scholar] [CrossRef]
- Studier, F.W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 2005, 41, 207–234. [Google Scholar] [CrossRef]
- Pandey, A.V.; Kempna, P.; Hofer, G.; Mullis, P.E.; Flück, C.E. Modulation of human CYP19A1 activity by mutant NADPH P450 oxidoreductase. Mol. Endocrinol. 2007, 21, 2579–2595. [Google Scholar] [CrossRef]
- Faeder, E.J.; Siegel, L.M. A rapid micromethod for determination of FMN and FAD in mixtures. Anal. Biochem. 1973, 53, 332–336. [Google Scholar] [CrossRef]
- Ashok, Y.; Nanekar, R.; Jaakola, V.P. Defining thermostability of membrane proteins by western blotting. Protein Eng. Des. Sel. 2015, 28, 539–542. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Alexandrov, A.I.; Mileni, M.; Chien, E.Y.; Hanson, M.A.; Stevens, R.C. Microscale fluorescent thermal stability assay for membrane proteins. Structure 2008, 16, 351–359. [Google Scholar] [CrossRef]
- Parween, S.; Fernandez-Cancio, M.; Benito-Sanz, S.; Camats, N.; Rojas Velazquez, M.N.; Lopez-Siguero, J.P.; Udhane, S.S.; Kagawa, N.; Fluck, C.E.; Audi, L.; et al. Molecular Basis of CYP19A1 Deficiency in a 46,XX Patient With R550W Mutation in POR: Expanding the PORD Phenotype. J. Clin. Endocrinol. Metab. 2020, 105, e1272–e1290. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez Castano, P.; Parween, S.; Pandey, A.V. Bioactivity of Curcumin on the Cytochrome P450 Enzymes of the Steroidogenic Pathway. Int. J. Mol. Sci. 2019, 20, 4606. [Google Scholar] [CrossRef]
- Agrawal, V.; Huang, N.; Miller, W.L. Pharmacogenetics of P450 oxidoreductase: Effect of sequence variants on activities of CYP1A2 and CYP2C19. Pharm. Genom. 2008, 18, 569–576. [Google Scholar] [CrossRef] [PubMed]
- Huang, N.; Agrawal, V.; Giacomini, K.M.; Miller, W.L. Genetics of P450 oxidoreductase: Sequence variation in 842 individuals of four ethnicities and activities of 15 missense mutations. Proc. Natl. Acad. Sci. USA 2008, 105, 1733–1738. [Google Scholar] [CrossRef] [PubMed]

| Project | Reference Allele Frequency | Alternative Allele Frequency | Allele Count | Allele Number |
|---|---|---|---|---|
| GnomAD_exome | C = 0.99752 | T = 0.002480 | 610 | 245,926 |
| ALFA | C = 0.996689 | T = 0.003311 | 474 | 143,178 |
| GnomAD | C = 0.997291 | T = 0.002709 | 380 | 140,266 |
| ExAC | C = 0.997025 | T = 0.002915 | 316 | 108,412 |
| PAGE_STUDY | C = 0.00896 | T = 0.00104 | 82 | 78,700 |
| GO-ESP | C = 0.9654 | T = 0.00346 | 43 | 12,434 |
| 1000 G | C = 0.9978 | T = 0.0022 | 11 | 5008 |
| Method | Prediction | Accuracy |
|---|---|---|
| PredicSNP2 | Deleterious | 0.89 |
| CADD | Deleterious | 0.52 |
| DANN | Deleterious | 0.89 |
| FATHMM | Deleterious | 0.67 |
| FunSeq2 | Deleterious | 0.61 |
| GWAVA | Deleterious | 0.47 |
| Meta-SNP | Deleterious | 0.6 |
| Vmax (nmol/min/mmol of POR) | Km (µM) | Vmax/Km (% of WT) | |
|---|---|---|---|
| Cytochrome c reduction activity | |||
| WT | 633 | 6.2 | 102.1 (100) |
| P228L | 119 | 5.6 | 27 (27) |
| MTT reduction activity | |||
| WT | 519 | 42.2 | 12.3 (100) |
| P228L | 267 | 235 | 1.1 (9) |
| Vmax (pmol/min*mg) | Km (µM) | Vmax/Km (% of WT) | |
|---|---|---|---|
| CYP17A1: 17-hydroxylase activity (Prog to 17OHP) | |||
| POR-WT | 257 ± 32 | 4.1 ± 0.5 | 63 (100) |
| POR-P228L | 193 ± 19 | 5.3 ± 0.7 | 36 (57) |
| CYP17A1: lyase activity (17OHPreg to DHEA) | |||
| POR-WT | 59.32 ± 12.35 | 1.34 ± 0.22 | 44 (100) |
| POR-P228L | 34.13 ± 7.09 | 2.82 ± 0.39 | 12 (27) |
| CYP19A1: aromatase (D4A to E1) | |||
| POR-WT | 12.35 ± 1.65 | 0.27 ± 0.06 | 46 (100) |
| POR-P228L | 7.62 ± 1.64 | 0.37 ± 0.08 | 21 (46) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rojas Velazquez, M.N.; Noebauer, M.; Pandey, A.V. Loss of Protein Stability and Function Caused by P228L Variation in NADPH-Cytochrome P450 Reductase Linked to Lower Testosterone Levels. Int. J. Mol. Sci. 2022, 23, 10141. https://doi.org/10.3390/ijms231710141
Rojas Velazquez MN, Noebauer M, Pandey AV. Loss of Protein Stability and Function Caused by P228L Variation in NADPH-Cytochrome P450 Reductase Linked to Lower Testosterone Levels. International Journal of Molecular Sciences. 2022; 23(17):10141. https://doi.org/10.3390/ijms231710141
Chicago/Turabian StyleRojas Velazquez, Maria Natalia, Mathias Noebauer, and Amit V. Pandey. 2022. "Loss of Protein Stability and Function Caused by P228L Variation in NADPH-Cytochrome P450 Reductase Linked to Lower Testosterone Levels" International Journal of Molecular Sciences 23, no. 17: 10141. https://doi.org/10.3390/ijms231710141
APA StyleRojas Velazquez, M. N., Noebauer, M., & Pandey, A. V. (2022). Loss of Protein Stability and Function Caused by P228L Variation in NADPH-Cytochrome P450 Reductase Linked to Lower Testosterone Levels. International Journal of Molecular Sciences, 23(17), 10141. https://doi.org/10.3390/ijms231710141