Regulation of Circadian Genes Nr1d1 and Nr1d2 in Sex-Different Manners during Liver Aging
Abstract
:1. Introduction
2. Results
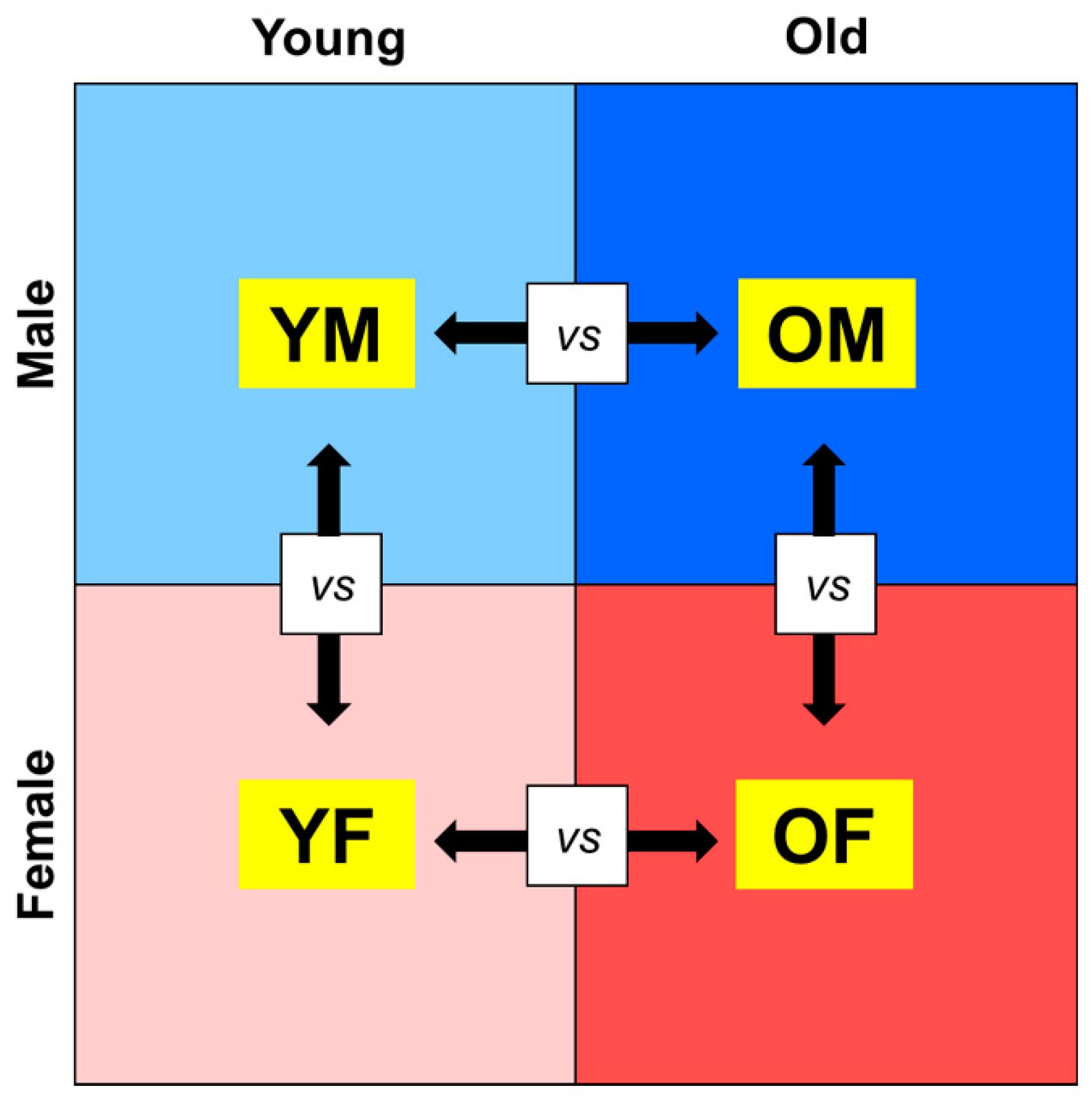
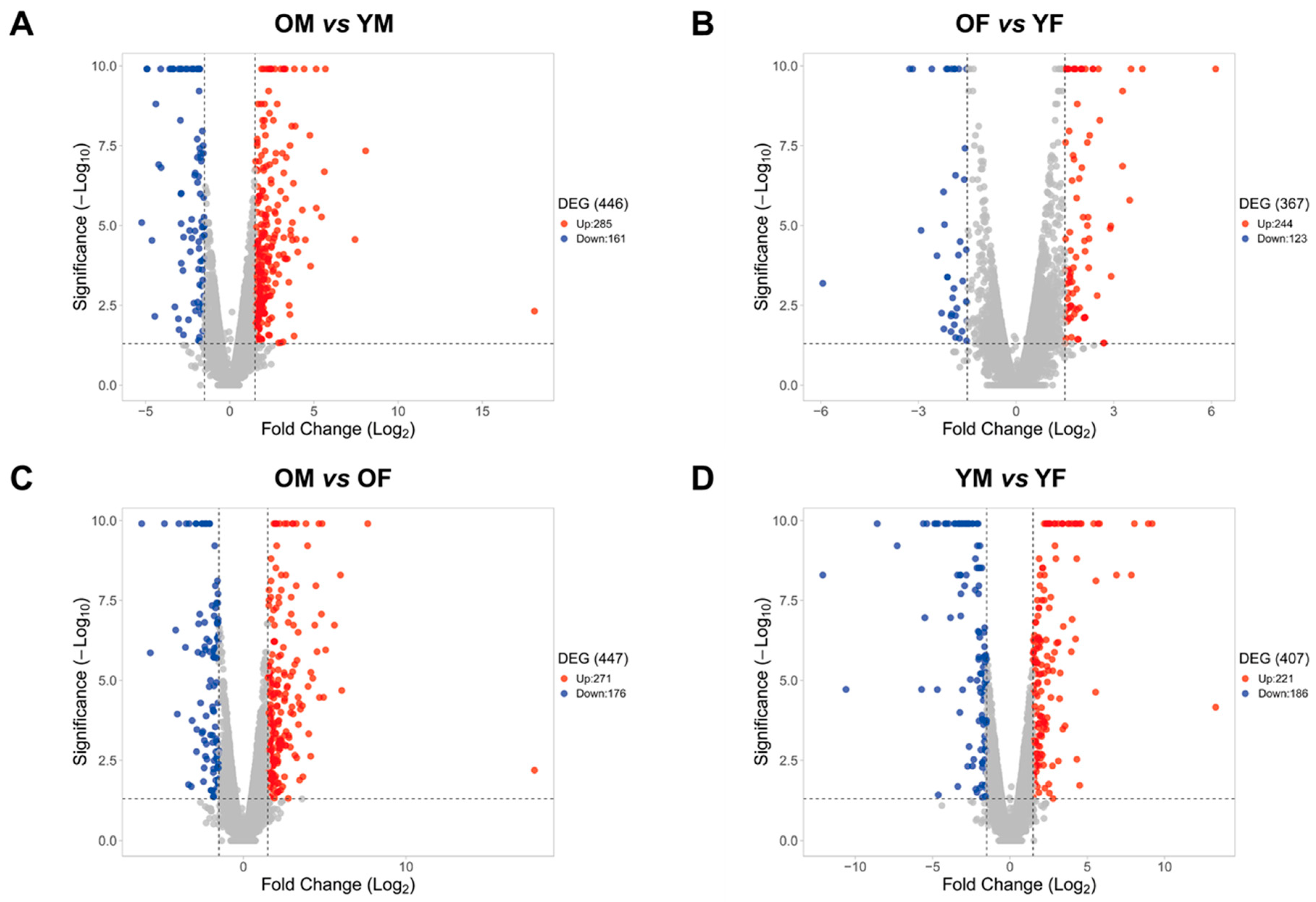
2.1. Age- and Sex-Related Changes in Gene Expression in Rat Liver
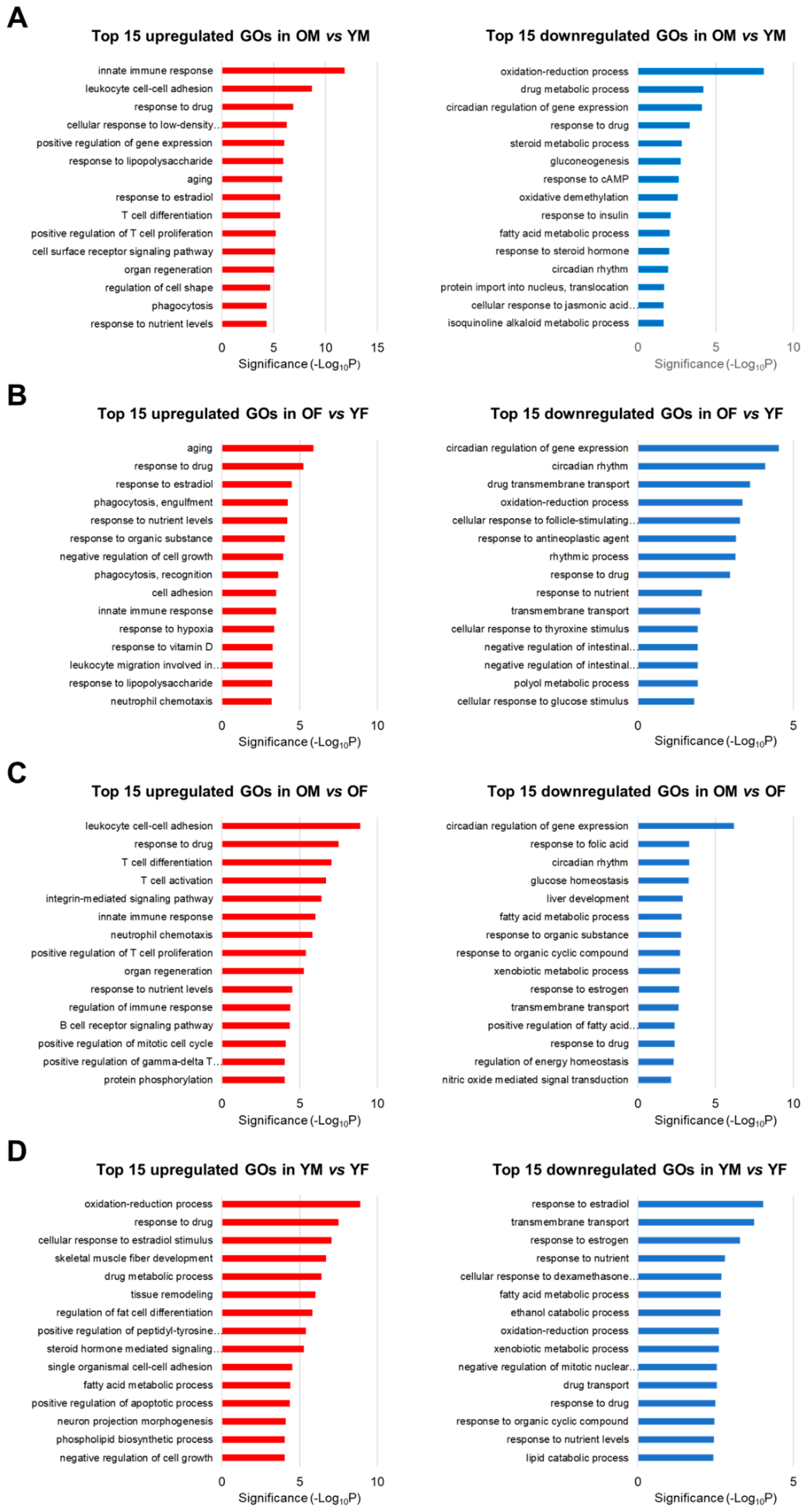
2.2. Biological Functions of Genes Whose Expression Was Changed According to Age and Sex in Rat Livers, as Detected by GO Enrichment Analysis
2.3. Biological Pathways of Genes Whose Expression Was Changed According to Age and Sex in Rat Liver, as Detected by KEGG Enrichment Analysis
2.4. Gene Expression of Circadian Genes That Were Differentially Expressed According to Age and Sex in Rat Livers, as Detected by Transcriptomic Analysis
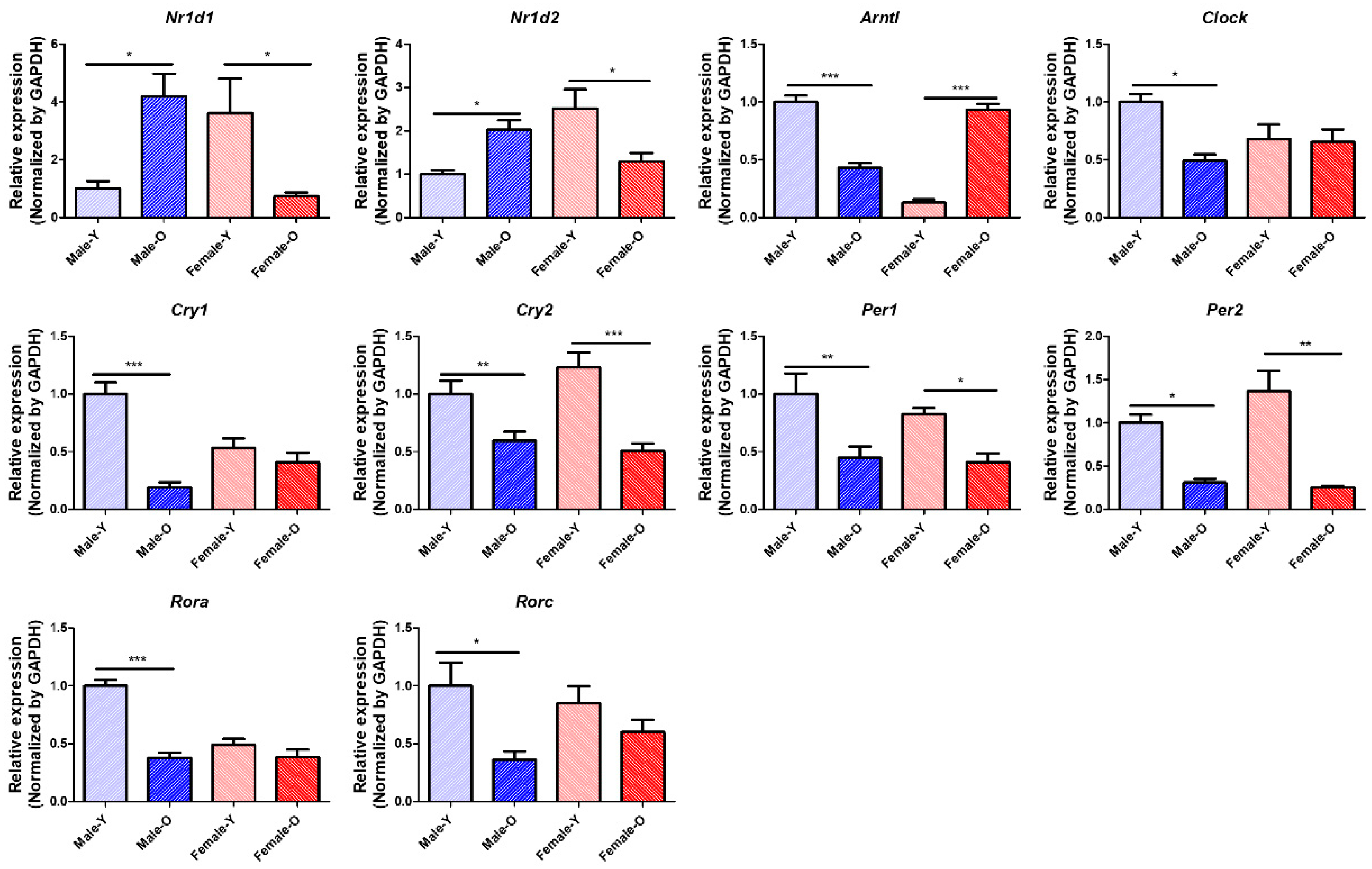
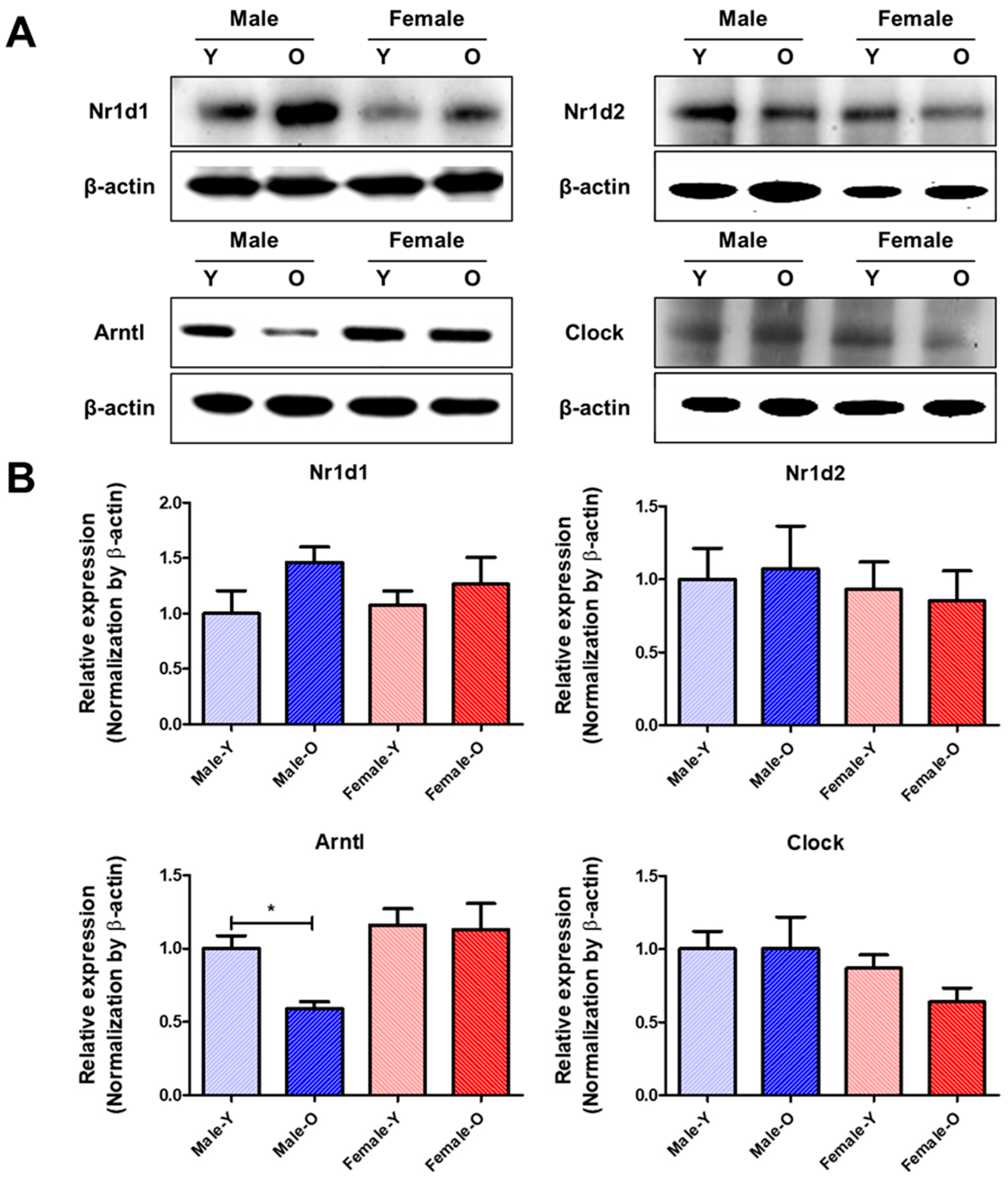
2.5. Gene Expression of Circadian Genes That Were Differentially Expressed According to Age and Sex in Rat Livers, as Determined via qRT-PCR and Western Blotting
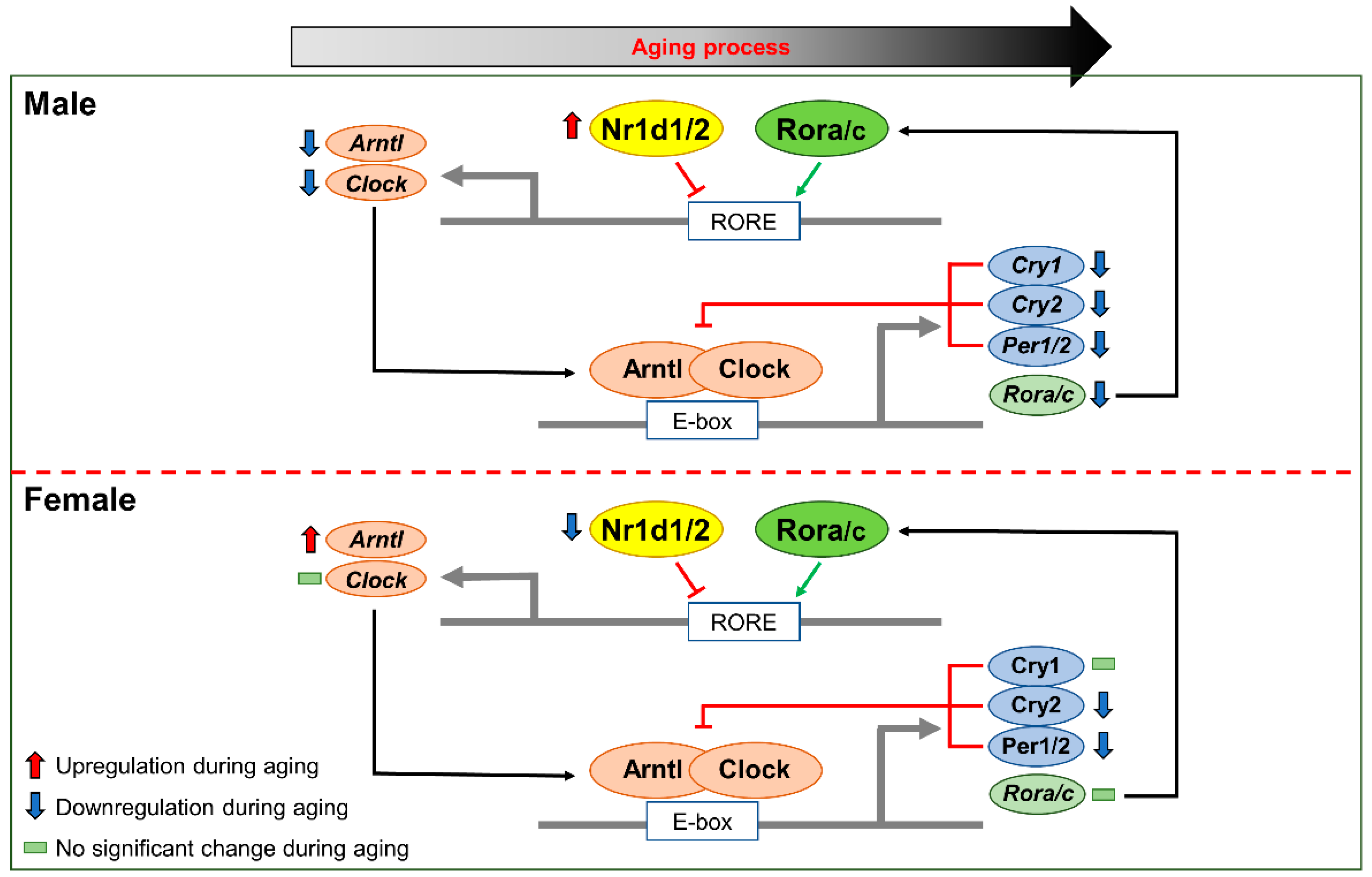
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. RNA Sequencing (RNA-Seq)
4.3. Differential Expressed Gene (DEG) Analysis
4.4. Enrichment Analysis of the DEGs
4.5. Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
4.6. Cytosolic Protein Extraction from Liver Tissues
4.7. Western Blotting
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kondratova, A.A.; Kondratov, R.V. The circadian clock and pathology of the ageing brain. Nat. Rev. Neurosci. 2012, 13, 325–335. [Google Scholar] [CrossRef]
- Welz, P.S.; Benitah, S.A. Molecular Connections Between Circadian Clocks and Aging. J. Mol. Biol. 2020, 432, 3661–3679. [Google Scholar] [CrossRef] [PubMed]
- Acosta-Rodriguez, V.A.; Rijo-Ferreira, F.; Green, C.B.; Takahashi, J.S. Importance of circadian timing for aging and longevity. Nat. Commun. 2021, 12, 2862. [Google Scholar] [CrossRef] [PubMed]
- Barth, E.; Srivastava, A.; Wengerodt, D.; Stojiljkovic, M.; Axer, H.; Witte, O.W.; Kretz, A.; Marz, M. Age-dependent expression changes of circadian system-related genes reveal a potentially conserved link to aging. Aging 2021, 13, 25694–25716. [Google Scholar] [CrossRef] [PubMed]
- Yanar, K.; Simsek, B.; Cakatay, U. Integration of Melatonin Related Redox Homeostasis, Aging, and Circadian Rhythm. Rejuvenation Res. 2019, 22, 409–419. [Google Scholar] [CrossRef]
- Sharma, A.; Sethi, G.; Tambuwala, M.M.; Aljabali, A.A.A.; Chellappan, D.K.; Dua, K.; Goyal, R. Circadian Rhythm Disruption and Alzheimer’s Disease: The Dynamics of a Vicious Cycle. Curr. Neuropharmacol. 2021, 19, 248–264. [Google Scholar] [CrossRef]
- Hood, S.; Amir, S. The aging clock: Circadian rhythms and later life. J. Clin. Investig. 2017, 127, 437–446. [Google Scholar] [CrossRef]
- De Nobrega, A.K.; Lyons, L.C. Aging and the clock: Perspective from flies to humans. Eur. J. Neurosci. 2020, 51, 454–481. [Google Scholar] [CrossRef]
- Banks, G.; Nolan, P.M.; Peirson, S.N. Reciprocal interactions between circadian clocks and aging. Mamm. Genome 2016, 27, 332–340. [Google Scholar] [CrossRef]
- Nicolaides, N.C.; Chrousos, G.P. Sex differences in circadian endocrine rhythms: Clinical implications. Eur. J. Neurosci. 2020, 52, 2575–2585. [Google Scholar] [CrossRef]
- Hirao, J.; Nishimura, M.; Arakawa, S.; Niino, N.; Mori, K.; Furukawa, T.; Sanbuissho, A.; Manabe, S.; Nishihara, M.; Mori, Y. Sex and circadian modulatory effects on rat liver as assessed by transcriptome analyses. J. Toxicol. Sci. 2011, 36, 9–22. [Google Scholar] [CrossRef] [PubMed]
- Bailey, M.; Silver, R. Sex differences in circadian timing systems: Implications for disease. Front. Neuroendocr. 2014, 35, 111–139. [Google Scholar] [CrossRef] [PubMed]
- Spitschan, M.; Santhi, N.; Ahluwalia, A.; Fischer, D.; Hunt, L.; Karp, N.A.; Levi, F.; Pineda-Torra, I.; Vidafar, P.; White, R. Sex differences and sex bias in human circadian and sleep physiology research. Elife 2022, 11, e65419. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Julius, A.A.; Wen, J.T. Optimization of light exposure and sleep schedule for circadian rhythm entrainment. PLoS ONE 2021, 16, e0251478. [Google Scholar] [CrossRef]
- Emens, J.S.; Burgess, H.J. Effect of Light and Melatonin and Other Melatonin Receptor Agonists on Human Circadian Physiology. Sleep Med. Clin. 2015, 10, 435–453. [Google Scholar] [CrossRef] [PubMed]
- Adafer, R.; Messaadi, W.; Meddahi, M.; Patey, A.; Haderbache, A.; Bayen, S.; Messaadi, N. Food Timing, Circadian Rhythm and Chrononutrition: A Systematic Review of Time-Restricted Eating’s Effects on Human Health. Nutrients 2020, 12, 3770. [Google Scholar] [CrossRef]
- Page, A.J. Gastrointestinal Vagal Afferents and Food Intake: Relevance of Circadian Rhythms. Nutrients 2021, 13, 844. [Google Scholar] [CrossRef]
- Yamamuro, D.; Takahashi, M.; Nagashima, S.; Wakabayashi, T.; Yamazaki, H.; Takei, A.; Takei, S.; Sakai, K.; Ebihara, K.; Iwasaki, Y.; et al. Peripheral circadian rhythms in the liver and white adipose tissue of mice are attenuated by constant light and restored by time-restricted feeding. PLoS ONE 2020, 15, e0234439. [Google Scholar] [CrossRef]
- Roenneberg, T.; Merrow, M. Entrainment of the human circadian clock. Cold Spring Harb. Symp. Quant. Biol. 2007, 72, 293–299. [Google Scholar] [CrossRef]
- Kuljis, D.A.; Loh, D.H.; Truong, D.; Vosko, A.M.; Ong, M.L.; McClusky, R.; Arnold, A.P.; Colwell, C.S. Gonadal- and sex-chromosome-dependent sex differences in the circadian system. Endocrinology 2013, 154, 1501–1512. [Google Scholar] [CrossRef]
- Chen, X.; Wang, L.; Loh, D.H.; Colwell, C.S.; Tache, Y.; Reue, K.; Arnold, A.P. Sex differences in diurnal rhythms of food intake in mice caused by gonadal hormones and complement of sex chromosomes. Horm. Behav. 2015, 75, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Okyar, A.; Kumar, S.A.; Filipski, E.; Piccolo, E.; Ozturk, N.; Monje, H.X.; Pala, Z.; Abraham, K.; Gomes, A.; Orman, M.N.; et al. Sex-, feeding-, and circadian time-dependency of P-glycoprotein expression and activity-implications for mechanistic pharmacokinetics modeling. Sci. Rep. 2019, 9, 10505. [Google Scholar] [CrossRef] [PubMed]
- Textoris, J.; Ban, L.H.; Capo, C.; Raoult, D.; Leone, M.; Mege, J.L. Sex-Related Differences in Gene Expression Following Coxiella burnetii Infection in Mice: Potential Role of Circadian Rhythm. PLoS ONE 2010, 5, e12190. [Google Scholar] [CrossRef] [PubMed]
- Eftekhar Vaghefi, S.S.; Mousavi, F.; Khaksari, M.; Asadikaram, G.; Soltani, Z. Sex-related Changes in Circadian Rhythm of Inflammatory and Oxidative Stress Markers in CKD. Iran. J. Kidney Dis. 2021, 15, 351–363. [Google Scholar]
- Bur, I.M.; Cohen-Solal, A.M.; Carmignac, D.; Abecassis, P.Y.; Chauvet, N.; Martin, A.O.; van der Horst, G.T.; Robinson, I.C.; Maurel, P.; Mollard, P.; et al. The circadian clock components CRY1 and CRY2 are necessary to sustain sex dimorphism in mouse liver metabolism. J. Biol. Chem. 2009, 284, 9066–9073. [Google Scholar] [CrossRef]
- Cox, K.H.; Takahashi, J.S. Circadian clock genes and the transcriptional architecture of the clock mechanism. J. Mol. Endocrinol. 2019, 63, R93–R102. [Google Scholar] [CrossRef]
- Trott, A.J.; Menet, J.S. Regulation of circadian clock transcriptional output by CLOCK: BMAL1. PLoS Genet. 2018, 14, e1007156. [Google Scholar] [CrossRef]
- Partch, C.L.; Green, C.B.; Takahashi, J.S. Molecular architecture of the mammalian circadian clock. Trends Cell Biol. 2014, 24, 90–99. [Google Scholar] [CrossRef]
- Cao, X.M.; Yang, Y.Y.; Selby, C.P.; Liu, Z.X.; Sancar, A. Molecular mechanism of the repressive phase of the mammalian circadian clock. Proc. Natl. Acad. Sci. USA 2020, 118, e2021174118. [Google Scholar] [CrossRef]
- Takahashi, J.S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 2017, 18, 164–179. [Google Scholar] [CrossRef]
- Cho, H.; Zhao, X.; Hatori, M.; Yu, R.T.; Barish, G.D.; Lam, M.T.; Chong, L.W.; DiTacchio, L.; Atkins, A.R.; Glass, C.K.; et al. Regulation of circadian behaviour and metabolism by REV-ERB-alpha and REV-ERB-beta. Nature 2012, 485, 123–127. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Solt, L.A.; Kojetin, D.J.; Burris, T.P. The REV-ERBs and RORs: Molecular links between circadian rhythms and lipid homeostasis. Future Med. Chem. 2011, 3, 623–638. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.L.; Li, L. Circadian Clock Regulates Inflammation and the Development of Neurodegeneration. Front. Cell. Infect. Microbiol. 2021, 11, 696554. [Google Scholar] [CrossRef] [PubMed]
- Gombert, M.; Carrasco-Luna, J.; Pin-Arboledas, G.; Codoner-Franch, P. The connection of circadian rhythm to inflammatory bowel disease. Transl. Res. 2019, 206, 107–118. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Huang, L.; Zhao, J.; Chen, S.; Liu, J.; Li, G. The circadian clock and inflammation: A new insight. Clin. Chim. Acta 2021, 512, 12–17. [Google Scholar] [CrossRef]
- Hergenhan, S.; Holtkamp, S.; Scheiermann, C. Molecular Interactions between Components of the Circadian Clock and the Immune System. J. Mol. Biol. 2020, 432, 3700–3713. [Google Scholar] [CrossRef]
- Curtis, A.M.; Bellet, M.M.; Sassone-Corsi, P.; O’Neill, L.A. Circadian clock proteins and immunity. Immunity 2014, 40, 178–186. [Google Scholar] [CrossRef]
- Huang, W.; Ramsey, K.M.; Marcheva, B.; Bass, J. Circadian rhythms, sleep, and metabolism. J. Clin. Investig. 2011, 121, 2133–2141. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, J. Bile acid metabolism and circadian rhythms. Am. J. Physiol. Liver Physiol. 2020, 319, G549–G563. [Google Scholar] [CrossRef]
- Froy, O. The circadian clock and metabolism. Clin. Sci. 2011, 120, 65–72. [Google Scholar] [CrossRef]
- Panda, S. Circadian physiology of metabolism. Science 2016, 354, 1008–1015. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tong, X.; Yin, L. Circadian rhythms in liver physiology and liver diseases. Compr. Physiol. 2013, 3, 917–940. [Google Scholar] [CrossRef] [PubMed]
- Serin, Y.; Acar Tek, N. Effect of Circadian Rhythm on Metabolic Processes and the Regulation of Energy Balance. Ann. Nutr. Metab. 2019, 74, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Huang, Y.; Chu, F.; Ji, S.; Liao, K.; Cui, Z.; Chen, J.; Tang, S. Clock Gene Nr1d1 Alleviates Retinal Inflammation Through Repression of Hmga2 in Microglia. J. Inflamm. Res. 2021, 14, 5901–5918. [Google Scholar] [CrossRef]
- Liu, H.; Zhu, Y.; Gao, Y.; Qi, D.; Zhao, L.; Zhao, L.; Liu, C.; Tao, T.; Zhou, C.; Sun, X.; et al. NR1D1 modulates synovial inflammation and bone destruction in rheumatoid arthritis. Cell Death Dis. 2020, 11, 129. [Google Scholar] [CrossRef]
- Pourcet, B.; Zecchin, M.; Ferri, L.; Beauchamp, J.; Sitaula, S.; Billon, C.; Delhaye, S.; Vanhoutte, J.; Mayeuf-Louchart, A.; Thorel, Q.; et al. Nuclear Receptor Subfamily 1 Group D Member 1 Regulates Circadian Activity of NLRP3 Inflammasome to Reduce the Severity of Fulminant Hepatitis in Mice. Gastroenterology 2018, 154, 1449–1464.e20. [Google Scholar] [CrossRef]
- Scheiermann, C.; Kunisaki, Y.; Frenette, P.S. Circadian control of the immune system. Nat. Rev. Immunol. 2013, 13, 190–198. [Google Scholar] [CrossRef]
- Griffin, P.; Dimitry, J.M.; Sheehan, P.W.; Lananna, B.V.; Guo, C.; Robinette, M.L.; Hayes, M.E.; Cedeno, M.R.; Nadarajah, C.J.; Ezerskiy, L.A.; et al. Circadian clock protein Rev-erbalpha regulates neuroinflammation. Proc. Natl. Acad. Sci. USA 2019, 116, 5102–5107. [Google Scholar] [CrossRef]
- Sato, S.; Sakurai, T.; Ogasawara, J.; Takahashi, M.; Izawa, T.; Imaizumi, K.; Taniguchi, N.; Ohno, H.; Kizaki, T. A circadian clock gene, Rev-erbalpha, modulates the inflammatory function of macrophages through the negative regulation of Ccl2 expression. J. Immunol. 2014, 192, 407–417. [Google Scholar] [CrossRef]
- Hunter, A.L.; Pelekanou, C.E.; Barron, N.J.; Northeast, R.C.; Grudzien, M.; Adamson, A.D.; Downton, P.; Cornfield, T.; Cunningham, P.S.; Billaud, J.N.; et al. Adipocyte NR1D1 dictates adipose tissue expansion during obesity. Elife 2021, 10, e63324. [Google Scholar] [CrossRef]
- Dai, L.; Zhang, Q.; Shi, J.; Bai, X.; An, X.; Zhang, B.; Zhang, Y.; Zhao, X. The Distribution, Expression Patterns and Functional Analysis of NR1D1 and NR4A2 in the Reproductive Axis Tissues of the Male Tianzhu White Yak. Animals 2021, 11, 3117. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Zhao, M.; Lu, D.; Wang, S.; Yu, F.; Guo, L.; Wen, S.; Wu, B. REV-ERBalpha Regulates CYP7A1 Through Repression of Liver Receptor Homolog-1. Drug Metab. Dispos. 2018, 46, 248–258. [Google Scholar] [CrossRef] [PubMed]
- Woldt, E.; Sebti, Y.; Solt, L.A.; Duhem, C.; Lancel, S.; Eeckhoute, J.; Hesselink, M.K.; Paquet, C.; Delhaye, S.; Shin, Y.; et al. Rev-erb-alpha modulates skeletal muscle oxidative capacity by regulating mitochondrial biogenesis and autophagy. Nat. Med. 2013, 19, 1039–1046. [Google Scholar] [CrossRef] [PubMed]
- Grant, D.; Yin, L.; Collins, J.L.; Parks, D.J.; Orband-Miller, L.A.; Wisely, G.B.; Joshi, S.; Lazar, M.A.; Willson, T.M.; Zuercher, W.J. GSK4112, a small molecule chemical probe for the cell biology of the nuclear heme receptor Rev-erbalpha. ACS Chem. Biol. 2010, 5, 925–932. [Google Scholar] [CrossRef]
- Solt, L.A.; Wang, Y.; Banerjee, S.; Hughes, T.; Kojetin, D.J.; Lundasen, T.; Shin, Y.; Liu, J.; Cameron, M.D.; Noel, R.; et al. Regulation of circadian behaviour and metabolism by synthetic REV-ERB agonists. Nature 2012, 485, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Bunger, M.K.; Walisser, J.A.; Sullivan, R.; Manley, P.A.; Moran, S.M.; Kalscheur, V.L.; Colman, R.J.; Bradfield, C.A. Progressive arthropathy in mice with a targeted disruption of the Mop3/Bmal-1 locus. Genesis 2005, 41, 122–132. [Google Scholar] [CrossRef]
- Wyse, C.A.; Coogan, A.N. Impact of aging on diurnal expression patterns of CLOCK and BMAL1 in the mouse brain. Brain Res. 2010, 1337, 21–31. [Google Scholar] [CrossRef]
- Marcheva, B.; Ramsey, K.M.; Buhr, E.D.; Kobayashi, Y.; Su, H.; Ko, C.H.; Ivanova, G.; Omura, C.; Mo, S.; Vitaterna, M.H.; et al. Disruption of the clock components CLOCK and BMAL1 leads to hypoinsulinaemia and diabetes. Nature 2010, 466, 627–631. [Google Scholar] [CrossRef] [PubMed]
- Jia, M.; Su, B.; Mo, L.; Qiu, W.; Ying, J.; Lin, P.; Yang, B.; Li, D.; Wang, D.; Xu, L.; et al. Circadian clock protein CRY1 prevents paclitaxelinduced senescence of bladder cancer cells by promoting p53 degradation. Oncol. Rep. 2021, 45, 1033–1043. [Google Scholar] [CrossRef]
- Bekki, H.; Duffy, T.; Okubo, N.; Olmer, M.; Alvarez-Garcia, O.; Lamia, K.; Kay, S.; Lotz, M. Suppression of circadian clock protein cryptochrome 2 promotes osteoarthritis. Osteoarthr. Cartil. 2020, 28, 966–976. [Google Scholar] [CrossRef]
- Baburski, A.Z.; Sokanovic, S.J.; Bjelic, M.M.; Radovic, S.M.; Andric, S.A.; Kostic, T.S. Circadian rhythm of the Leydig cells endocrine function is attenuated during aging. Exp. Gerontol. 2016, 73, 5–13. [Google Scholar] [CrossRef] [PubMed]
- Morioka, N.; Saeki, M.; Sugimoto, T.; Higuchi, T.; Zhang, F.F.; Nakamura, Y.; Hisaoka-Nakashima, K.; Nakata, Y. Downregulation of the spinal dorsal horn clock gene Per1 expression leads to mechanical hypersensitivity via c-jun N-terminal kinase and CCL2 production in mice. Mol. Cell. Neurosci. 2016, 72, 72–83. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Nah, S.S.; Chang, S.H.; Kim, H.K.; Kwon, J.T.; Lee, S.; Cho, I.H.; Lee, S.W.; Kim, Y.O.; Hong, S.J.; et al. PER2 is downregulated by the LPS-induced inflammatory response in synoviocytes in rheumatoid arthritis and is implicated in disease susceptibility. Mol. Med. Rep. 2017, 16, 422–428. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Yuan, X.; Li, Y.; Chen, J.; Wu, S.; Jiang, A.; Miao, X.; Shu, Q. Circulating exosomal microRNA-18a-5p accentuates intestinal inflammation in Hirschsprung-associated enterocolitis by targeting RORA. Am. J. Transl. Res. 2021, 13, 4182–4196. [Google Scholar]
- Han, S.; Li, Z.; Han, F.; Jia, Y.; Qi, L.; Wu, G.; Cai, W.; Xu, Y.; Li, C.; Zhang, W.; et al. ROR alpha protects against LPS-induced inflammation by down-regulating SIRT1/NF-kappa B pathway. Arch. Biochem. Biophys. 2019, 668, 1–8. [Google Scholar] [CrossRef]
- Chai, C.; Cox, B.; Yaish, D.; Gross, D.; Rosenberg, N.; Amblard, F.; Shemuelian, Z.; Gefen, M.; Korach, A.; Tirosh, O.; et al. Agonist of RORA Attenuates Nonalcoholic Fatty Liver Progression in Mice via Up-regulation of MicroRNA 122. Gastroenterology 2020, 159, 999–1014.e9. [Google Scholar] [CrossRef]
- El-Karaksy, S.M.; Raafat, H.A.; Abadir, M.N.; Hanna, M.O. Down-regulation of expression of retinoid acid-related orphan receptor C (RORC) in systemic lupus erythematosus. J. Recept. Signal Transduct. 2016, 36, 207–212. [Google Scholar] [CrossRef]
- Billon, C.; Murray, M.H.; Avdagic, A.; Burris, T.P. RORgamma regulates the NLRP3 inflammasome. J. Biol. Chem. 2019, 294, 10–19. [Google Scholar] [CrossRef]
- Nadeem, A.; Al-Harbi, N.O.; Al-Harbi, M.M.; El-Sherbeeny, A.M.; Ahmad, S.F.; Siddiqui, N.; Ansari, M.A.; Zoheir, K.M.A.; Attia, S.M.; Al-Hosaini, K.A.; et al. Imiquimod-induced psoriasis-like skin inflammation is suppressed by BET bromodomain inhibitor in mice through RORC/IL-17A pathway modulation. Pharmacol. Res. 2015, 99, 248–257. [Google Scholar] [CrossRef]
- Xiao, L.; Zhang, Z.; Luo, X.; Yang, H.; Li, F.; Wang, N. Retinoid acid receptor-related orphan receptor alpha (RORalpha) regulates macrophage M2 polarization via activation of AMPKalpha. Mol. Immunol. 2016, 80, 17–23. [Google Scholar] [CrossRef]
- Auclair, M.; Roblot, N.; Capel, E.; Feve, B.; Antoine, B. Pharmacological modulation of RORalpha controls fat browning, adaptive thermogenesis, and body weight in mice. Am. J. Physiol. Endocrinol. Metab. 2021, 320, E219–E233. [Google Scholar] [CrossRef] [PubMed]
- Kumar, N.; Lyda, B.; Chang, M.R.; Lauer, J.L.; Solt, L.A.; Burris, T.P.; Kamenecka, T.M.; Griffin, P.R. Identification of SR2211: A potent synthetic RORgamma-selective modulator. ACS Chem. Biol. 2012, 7, 672–677. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fauber, B.P.; Gobbi, A.; Robarge, K.; Zhou, A.H.; Barnard, A.; Cao, J.H.; Deng, Y.Z.; Eidenschenk, C.; Everett, C.; Ganguli, A.; et al. Discovery of imidazo [1,5-a]pyridines and -pyrimidines as potent and selective RORc inverse agonists. Bioorg. Med. Chem. Lett. 2015, 25, 2907–2912. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Su, M.; Wu, X.Z.; Wang, D.Z.; Kang, Y.Y.; Wang, C.G.; Assani, I.; Wang, M.X.; Zhao, S.F.; Lv, S.M.; et al. Discovery of 2H-chromone-4-one based sulfonamide derivatives as potent retinoic acid receptor-related orphan receptor gammat inverse agonists. Eur. J. Med. Chem. 2022, 229, 114065. [Google Scholar] [CrossRef]
- Guillaumond, F.; Dardente, H.; Giguere, V.; Cermakian, N. Differential control of Bmal1 circadian transcription by REV-ERB and ROR nuclear receptors. J. Biol. Rhythms. 2005, 20, 391–403. [Google Scholar] [CrossRef]
- Rohr, K.E.; Telega, A.; Savaglio, A.; Evans, J.A. Vasopressin regulates daily rhythms and circadian clock circuits in a manner influenced by sex. Horm. Behav. 2021, 127, 104888. [Google Scholar] [CrossRef]
- Iwahana, E.; Karatsoreos, I.; Shibata, S.; Silver, R. Gonadectomy reveals sex differences in circadian rhythms and suprachiasmatic nucleus androgen receptors in mice. Horm. Behav. 2008, 53, 422–430. [Google Scholar] [CrossRef]
- Kawamura, M.; Tasaki, H.; Misawa, I.; Chu, G.; Yamauchi, N.; Hattori, M.A. Contribution of testosterone to the clock system in rat prostate mesenchyme cells. Andrology 2014, 2, 225–233. [Google Scholar] [CrossRef]
- Zhao, L.; Zhang, J.; Yang, L.; Zhang, H.; Zhang, Y.; Gao, D.; Jiang, H.; Li, Y.; Dong, H.; Ma, T.; et al. Glyphosate exposure attenuates testosterone synthesis via NR1D1 inhibition of StAR expression in mouse Leydig cells. Sci. Total Environ. 2021, 785, 147323. [Google Scholar] [CrossRef]
- Isayama, K.; Chen, H.; Yamauchi, N.; Hattori, M.A. REV-ERBalpha inhibits the PTGS2 expression in bovine uterus endometrium stromal and epithelial cells exposed to ovarian steroids. J. Reprod. Dev. 2014, 60, 362–370. [Google Scholar] [CrossRef]
- Wang, L.; Li, J.; Zhang, L.; Shi, S.; Zhou, X.; Hu, Y.; Gao, L.; Yang, G.; Pang, W.; Chen, H.; et al. NR1D1 targeting CYP19A1 inhibits estrogen synthesis in ovarian granulosa cells. Theriogenology 2022, 180, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Astafev, A.A.; Patel, S.A.; Kondratov, R.V. Calorie restriction effects on circadian rhythms in gene expression are sex dependent. Sci. Rep. 2017, 7, 9716. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, V.W.; Sarachana, T.; Sherrard, R.M.; Kocher, K.M. Investigation of sex differences in the expression of RORA and its transcriptional targets in the brain as a potential contributor to the sex bias in autism. Mol. Autism 2015, 6, 7. [Google Scholar] [CrossRef] [PubMed]
- Gardner, R.S.; Newman, L.A.; Mohler, E.G.; Tunur, T.; Gold, P.E.; Korol, D.L. Aging is not equal across memory systems. Neurobiol. Learn. Mem. 2020, 172, 107232. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Yu, S.H.; Kim, S.-O. Decreased urothelial expression of caveolin 1 and 2 in aging rats showing detrusor overactivity: Potential association with aging bladder. Investig. Clin. Urol. 2021, 62, 690–696. [Google Scholar] [CrossRef] [PubMed]
- Davis, K.; Pearl, C.A. Effects of Estrogen Treatment on Aging in the Rat Epididymis. Anat. Rec. 2019, 302, 1447–1457. [Google Scholar] [CrossRef] [PubMed]
- Andreollo, N.A.; Santos, E.F.; Araujo, M.R.; Lopes, L.R. Rat’s age versus human’s age: What is the relationship? Arq. Bras. Cir. Dig. 2012, 25, 49–51. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Sun, J.Q.; Nishiyama, T.; Shimizu, K.; Kadota, K. TCC: An R package for comparing tag count data with robust normalization strategies. BMC Bioinform. 2013, 14, 219. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goedhart, J.; Luijsterburg, M.S. VolcaNoseR is a web app for creating, exploring, labeling and sharing volcano plots. Sci. Rep. 2020, 10, 20560. [Google Scholar] [CrossRef] [PubMed]
| Dataset | Up | Down |
|---|---|---|
| OM vs. YM | Natural killer cell-mediated cytotoxicity, Hematopoietic cell lineage, Rheumatoid arthritis, Cell adhesion molecules (CAMs), Fc epsilon RI signaling pathway, Antigen processing and presentation, Asthma, Jak-STAT signaling pathway, Cytokine—cytokine receptor interaction, Leukocyte transendothelial migration, Inflammatory bowel disease (IBD), B cell receptor signaling pathway, Type I diabetes mellitus, T cell receptor signaling pathway, Retinol metabolism, Primary immunodeficiency, Central carbon metabolism in cancer | Steroid hormone biosynthesis, Metabolic pathways, Retinol metabolism, Linoleic acid metabolism, Drug metabolism—other enzymes, Circadian rhythm |
| OF vs. YF | Maturity onset diabetes of the young, CAMs, Glycine, serine and threonine metabolism, Metabolic pathways | Drug metabolism—cytochrome P450, Nicotinate and nicotinamide metabolism, Glutathione metabolism, Metabolism of xenobiotics by cytochrome P450, Metabolic pathways, Pentose and glucuronate interconversions, Circadian rhythm, Steroid hormone biosynthesis, Galactose metabolism, Retinol metabolism, Fructose and mannose metabolism, Tyrosine metabolism |
| OM vs. OF | Natural killer cell-mediated cytotoxicity, CAMs, Rheumatoid arthritis, Leukocyte transendothelial migration, Hematopoietic cell lineage, Fc epsilon RI signaling pathway, T cell receptor signaling pathway, Primary immunodeficiency, Antigen processing and presentation, Intestinal immune network for IgA production, Asthma, Inflammatory bowel disease (IBD), B cell receptor signaling pathway | Retinol metabolism, Circadian rhythm, Ascorbate and aldarate metabolism, Fatty acid elongation, Biosynthesis of unsaturated fatty acids, Metabolic pathways, Metabolism of xenobiotics by cytochrome P450, Drug metabolism—cytochrome P450, Steroid hormone biosynthesis, Drug metabolism—other enzymes, Histidine metabolism, Pentose and glucuronate interconversions, PPAR signaling pathway |
| YM vs. YF | Steroid hormone biosynthesis, Metabolic pathways, PPAR signaling pathway, Choline metabolism in cancer | Drug metabolism—cytochrome P450, Steroid hormone biosynthesis, Retinol metabolism, Butanoate metabolism, Metabolic pathways, Metabolism of xenobiotics by cytochrome P450, Drug metabolism—other enzymes, Ascorbate and aldarate metabolism, Biosynthesis of unsaturated fatty acids, Fatty acid elongation |
| Gene | FC (OM vs. YM) | FC (OF vs. YF) | FC (OM vs. OF) |
|---|---|---|---|
| Nr1d1 | 6.680703355 | −2.056227653 | 3.680750602 |
| Nr1d2 | 2.602683711 | −2.345669898 | 1.607701981 |
| Arntl (or Bmal1) | −2.114036081 | 3.972369982 | −1.969732886 |
| Clock (or Npas2) | −1.859609885 | 29.65081798 | −2.602683711 |
| Cry1 | −3.226567037 | 1.155886707 | −2.411615655 |
| Cry2 | −1.209994089 | −1.317679952 | −1.112650121 |
| Per1 | −1.441928871 | −1.009471374 | −1.783857039 |
| Per2 | −2.042024251 | −2.02791896 | −1.822602561 |
| Rora | −2.789487333 | 1.340712592 | −2.462288827 |
| Rorc | −1.821339667 | 1.134455485 | −1.876442393 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Noh, S.G.; Jung, H.J.; Kim, S.; Arulkumar, R.; Kim, D.H.; Park, D.; Chung, H.Y. Regulation of Circadian Genes Nr1d1 and Nr1d2 in Sex-Different Manners during Liver Aging. Int. J. Mol. Sci. 2022, 23, 10032. https://doi.org/10.3390/ijms231710032
Noh SG, Jung HJ, Kim S, Arulkumar R, Kim DH, Park D, Chung HY. Regulation of Circadian Genes Nr1d1 and Nr1d2 in Sex-Different Manners during Liver Aging. International Journal of Molecular Sciences. 2022; 23(17):10032. https://doi.org/10.3390/ijms231710032
Chicago/Turabian StyleNoh, Sang Gyun, Hee Jin Jung, Seungwoo Kim, Radha Arulkumar, Dae Hyun Kim, Daeui Park, and Hae Young Chung. 2022. "Regulation of Circadian Genes Nr1d1 and Nr1d2 in Sex-Different Manners during Liver Aging" International Journal of Molecular Sciences 23, no. 17: 10032. https://doi.org/10.3390/ijms231710032
APA StyleNoh, S. G., Jung, H. J., Kim, S., Arulkumar, R., Kim, D. H., Park, D., & Chung, H. Y. (2022). Regulation of Circadian Genes Nr1d1 and Nr1d2 in Sex-Different Manners during Liver Aging. International Journal of Molecular Sciences, 23(17), 10032. https://doi.org/10.3390/ijms231710032







