Characterization of the SWEET Gene Family in Longan (Dimocarpus longan) and the Role of DlSWEET1 in Cold Tolerance
Abstract
:1. Introduction
2. Results
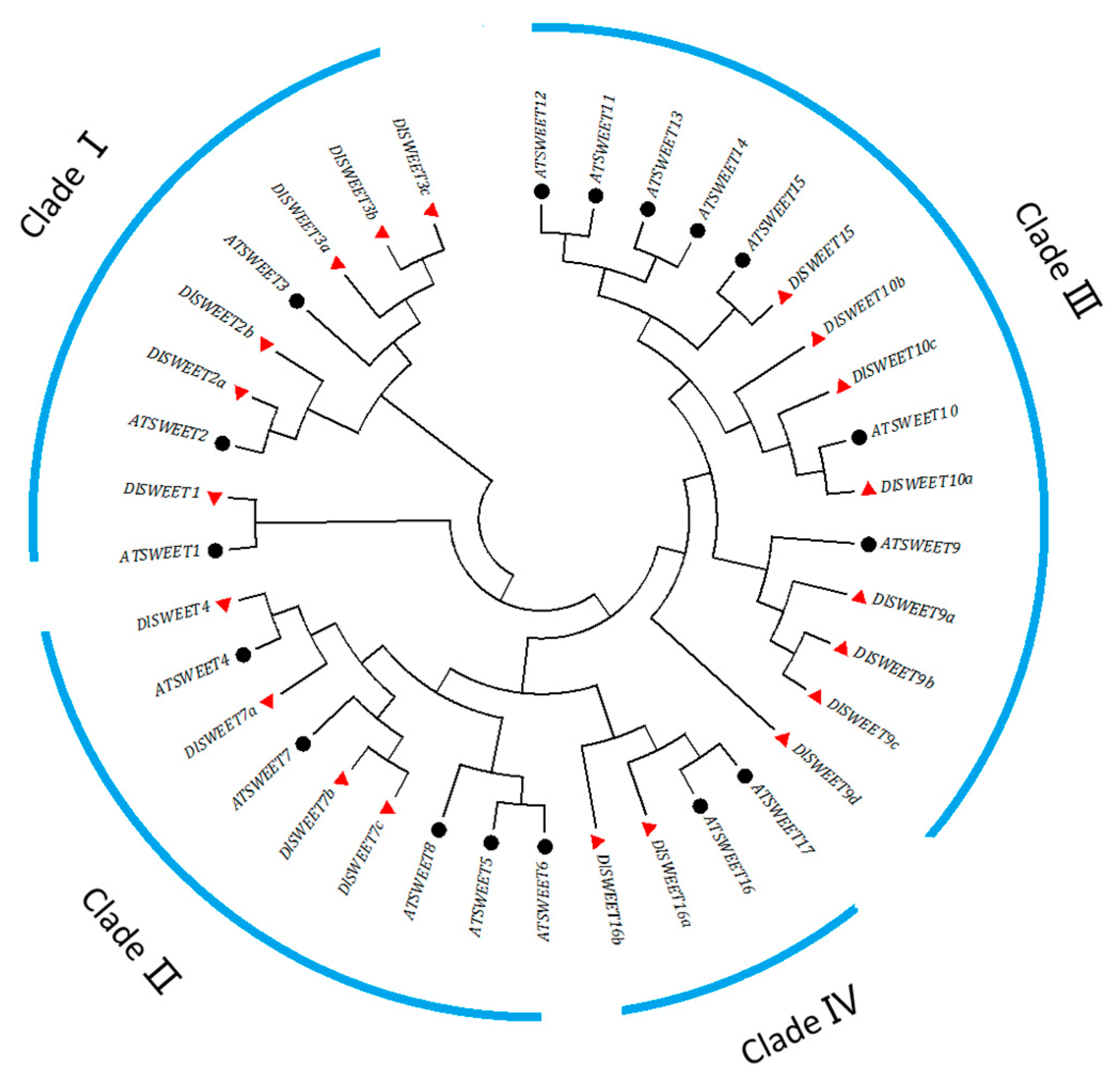
2.1. Identification and Phylogenetic Analysis of SWEET Proteins in Longan
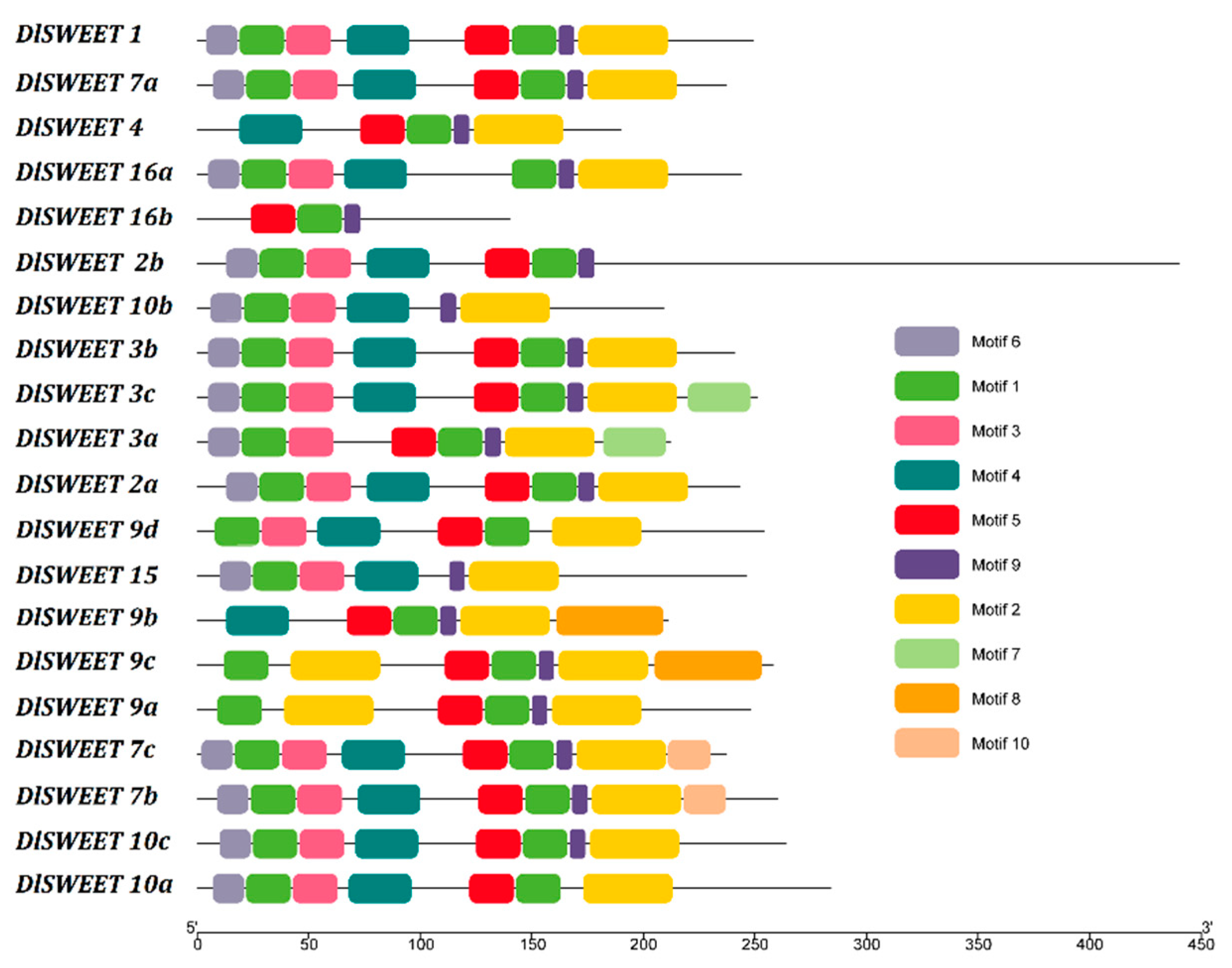
2.2. Analysis of Transmembrane Domains and Conserved Motifs
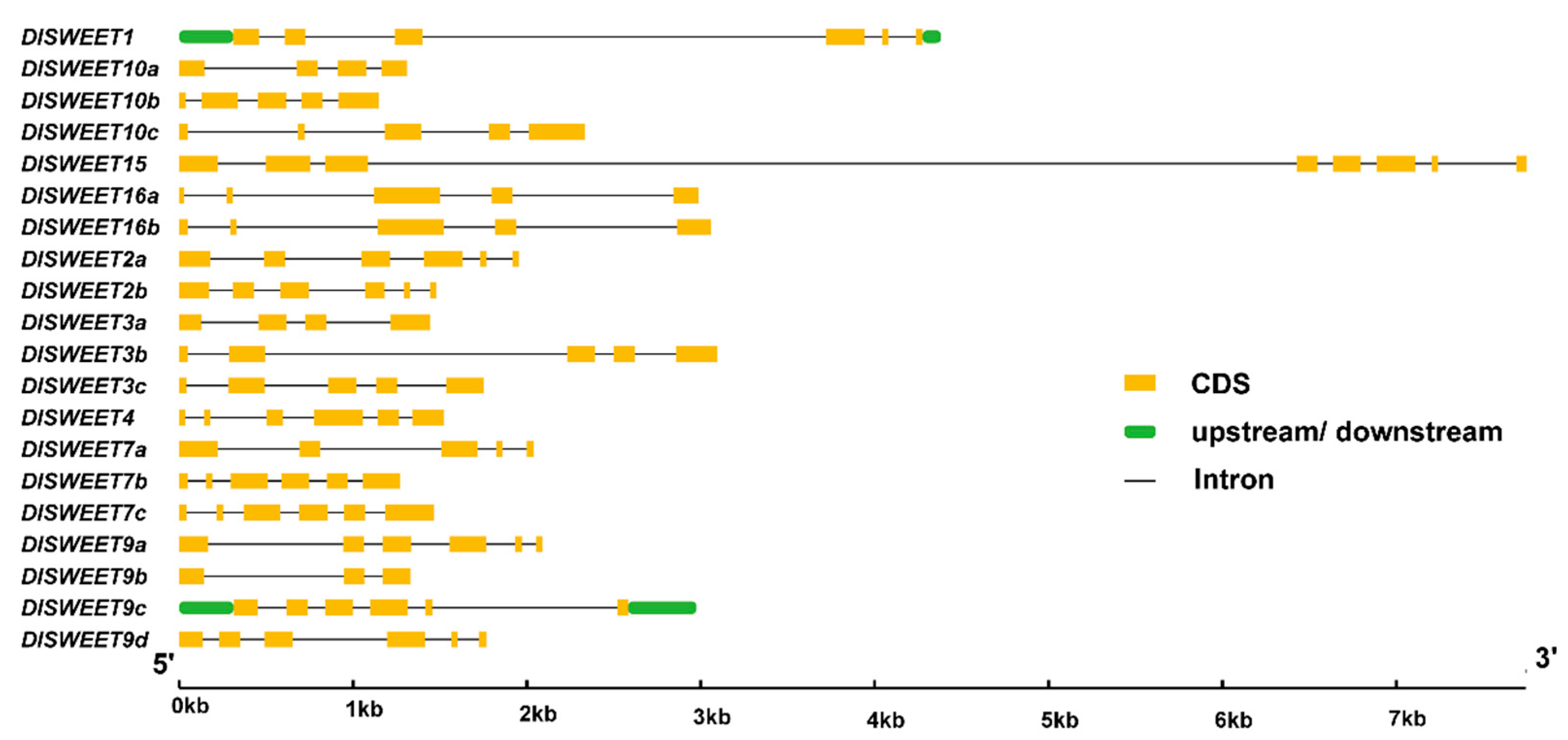
2.3. Exon–Intro Organization of DlSWEET Genes
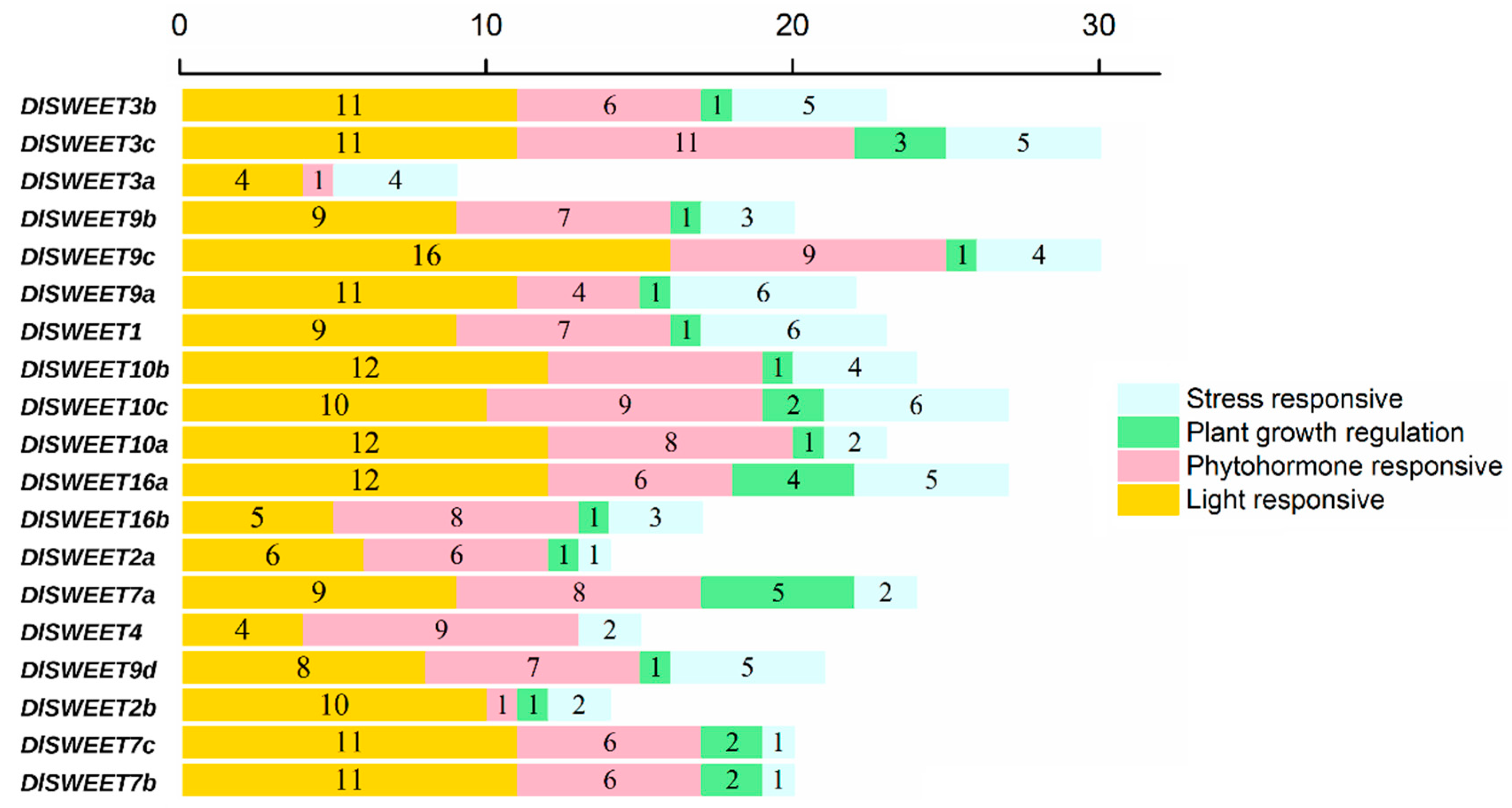
2.4. Cis-Acting Elements in the Promoters of DlSWEETs
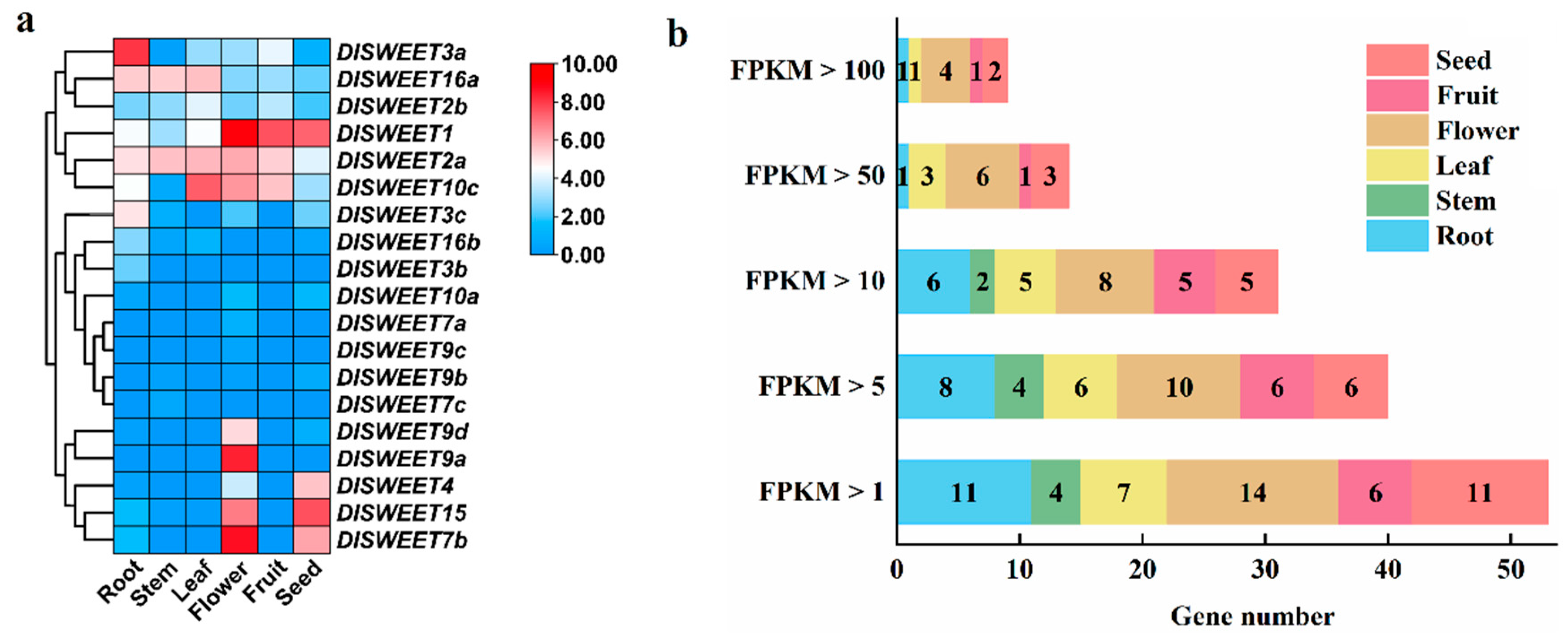
2.5. Tissue-Specific Expression Patterns of DlSWEET Genes
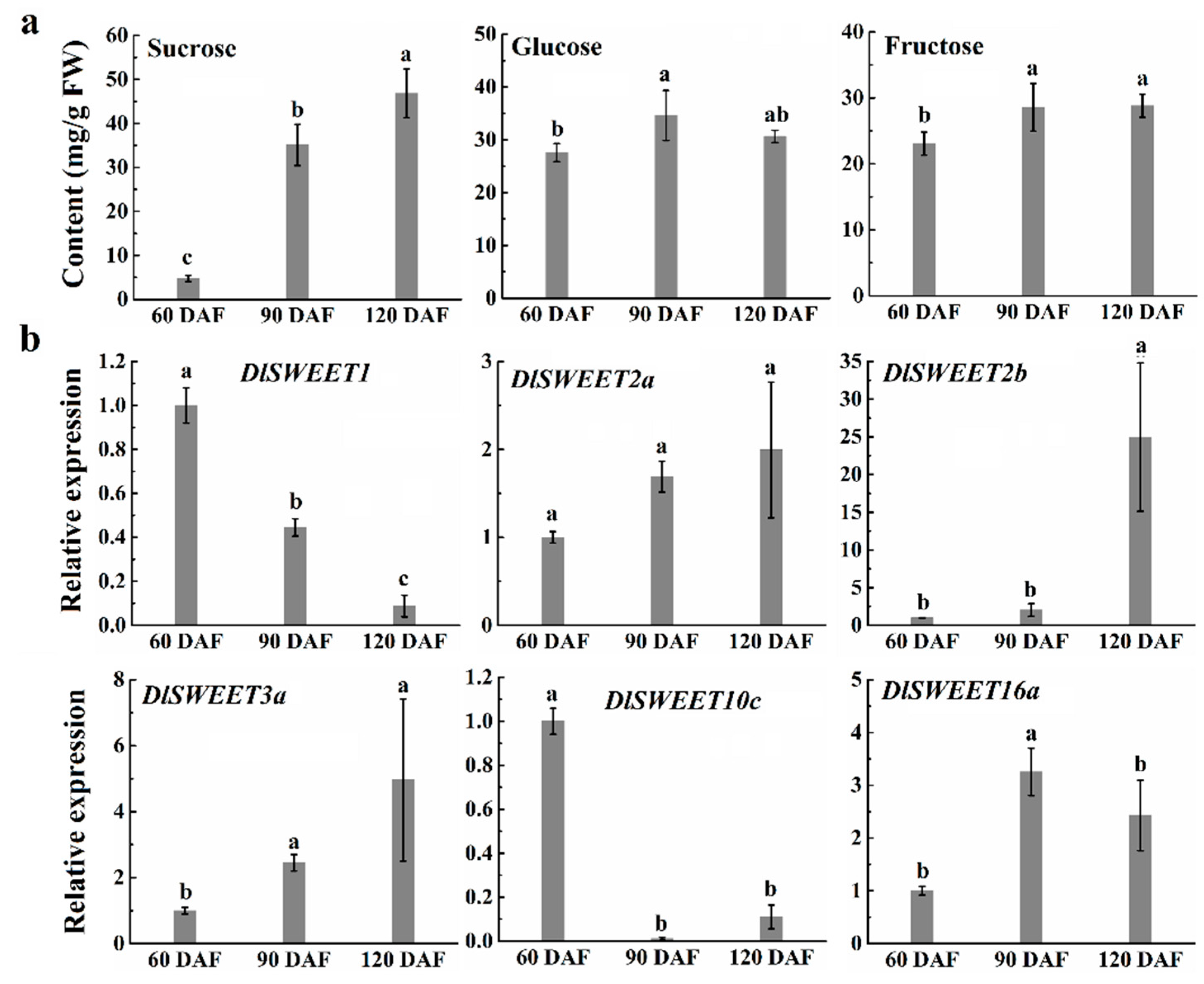
2.6. Soluble Sugar and Expression Patterns of DlSWEET Genes during Fruit Development
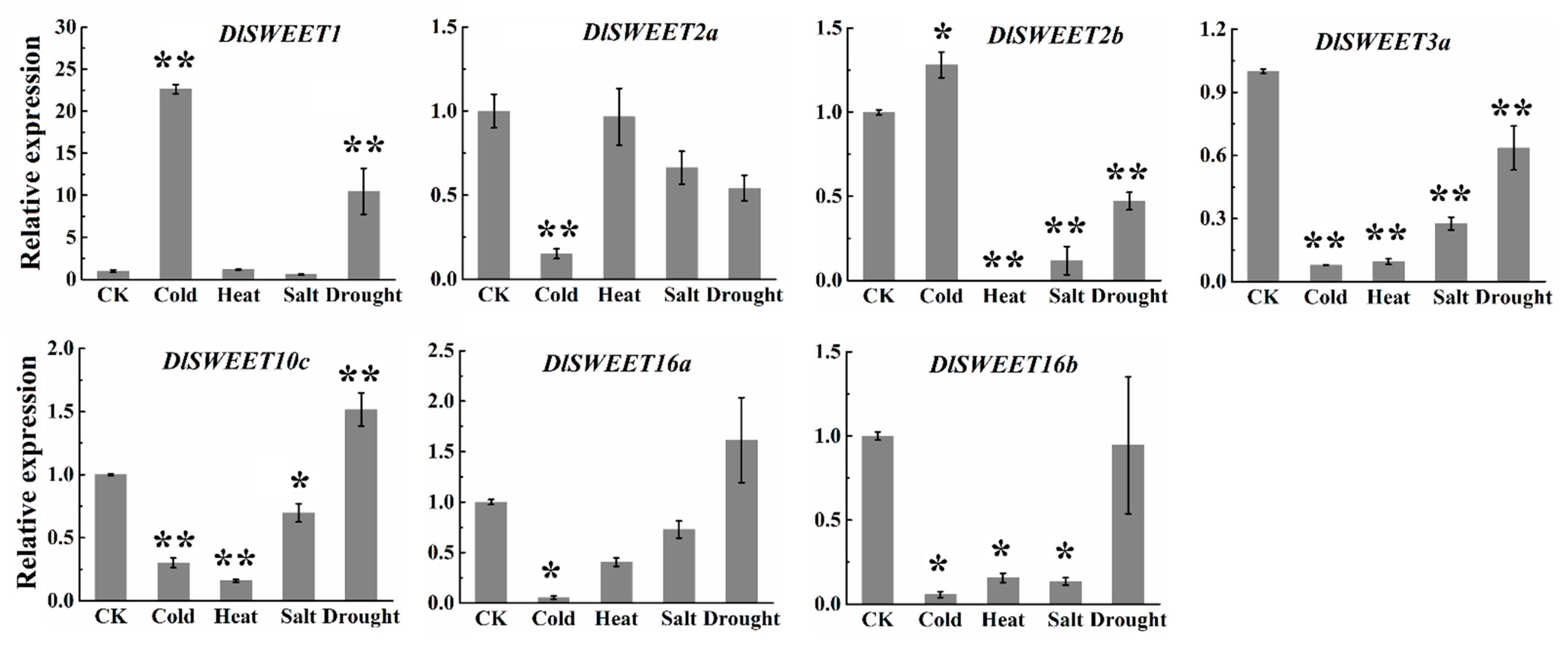
2.7. Expression Patterns of DlSWEET Genes under Abiotic Stress Condition
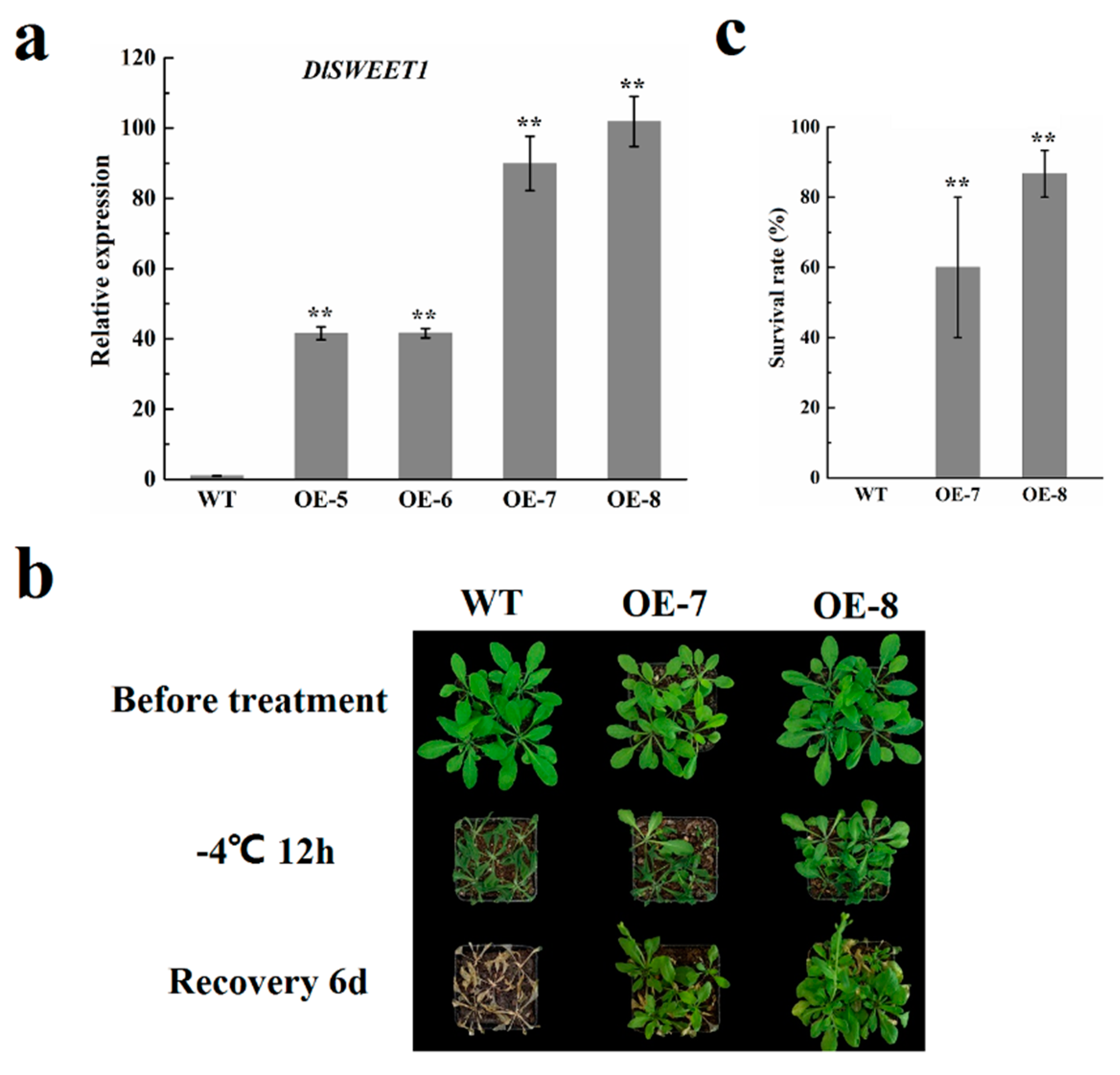
2.8. Overexpression of DlSWEET1 Enhances Cold Tolerance in Transgenic Arabidopsis Plants
3. Discussion
4. Materials and Methods
4.1. Plant Materials, Stress Treatments, and Measurement of Soluble Sugars Content
4.2. Identification, Phylogenetic Analysis, and Motifs Prediction of SWEET Protein in Longan
4.3. Cis-Elements Search and Exon–Intron Organization of DlSWEET Genes
4.4. Expression Pattern Analysis of DlSWEET Genes
4.5. RNA Isolation and qRT-PCR Analysis
4.6. Functional Analysis of DlSWEET1-Overexpressing Transgenic Arabidopsis Thaliana
4.7. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sheen, J.; Zhou, L.; Jang, J.C. Sugars as signalling molecules. Curr. Opin. Plant Biol. 1999, 2, 410–418. [Google Scholar] [CrossRef]
- Fang, T.; Peng, Y.; Rao, Y.; Li, S.; Zeng, L. Genome-wide identification and expression analysis of sugar transporter (ST) gene family in longan (Dimocarpus longan L.). Plants 2020, 9, 342. [Google Scholar] [CrossRef] [Green Version]
- Wen, S.Y.; Neuhaus, H.E.; Cheng, J.T.; Bie, Z.L. Contributions of sugar transporters to crop yield and fruit quality. J. Exp. Bot. 2022, 73, 2275–2289. [Google Scholar] [CrossRef]
- Chen, T.; Zhang, Z.; Li, B.; Qin, G.; Tian, S. Molecular basis for optimizing sugar metabolism and transport during fruit development. aBIOTECH 2021, 2, 330–340. [Google Scholar] [CrossRef]
- Zhen, Q.; Fang, T.; Peng, Q.; Liao, L.; Zhao, L.; Owiti, A.; Han, Y. Developing gene-tagged molecular markers for evaluation of genetic association of apple sweet genes with fruit sugar accumulation. Hortic. Res. 2018, 5, 14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.Q.; Cheung, L.S.; Feng, L.; Tanner, W.; Frommer, W.B. Transport of sugars. Annu. Rev. Biochem. 2015, 84, 865–894. [Google Scholar] [CrossRef]
- Huang, W.F.; Hu, B.; Liu, J.L.; Zhou, Y.; Liu, S.Q. Identification and Characterization of Tonoplast Sugar Transporter (TST) Gene Family in Cucumber. Hortic. Plant J. 2017, 6, 145–157. [Google Scholar] [CrossRef]
- Doidy, J.; Grace, E.; Kuhn, C.; Simon-Plas, F.; Casieri, L.; Wipf, D. Sugar transporters in plants and in their interactions with fungi. Trends Plant Sci. 2012, 17, 413–422. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.Q.; Hou, B.H.; Lalonde, S.; Takanaga, H.; Hartung, M.L.; Qu, X.Q.; Guo, W.J.; Kim, J.G.; Underwood, W.; Chaudhuri, B.; et al. Sugar transporters for intercellular exchange and nutrition of pathogens. Nature 2010, 468, 527–532. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xuan, Y.H.; Hu, Y.B.; Chen, L.Q.; Sosso, D.; Ducat, D.C.; Hou, B.H.; Frommer, W.B. Functional role of oligomerization for bacterial and plant SWEET sugar transporter family. Proc. Natl. Acad. Sci. USA 2013, 110, E3685–E3694. [Google Scholar] [CrossRef] [Green Version]
- Baker, R.F.; Leach, K.A.; Braun, D.M. SWEET as sugar, new sucrose effluxers in plants. Mol. Plant 2012, 5, 766–768. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.Q.; Qu, X.Q.; Hou, B.H.; Sosso, D.; Osorio, S.; Fernie, A.R.; Frommer, W.B. Sucrose efflux mediated by SWEET proteins as a key step for phloem transport. Science 2012, 335, 207–211. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; Wang, S. Rice MtN3/saliva/SWEET family genes and their homologs in cellular organisms. Mol. Plant 2013, 6, 665–674. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, Y.; Tao, Y.; Cheung, L.S.; Fan, C.; Chen, L.Q.; Xu, S.; Perry, K.; Frommer, W.B.; Feng, L. Structures of bacterial homologues of SWEET transporters in two distinct conformations. Nature 2014, 515, 448–452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, Y.B.; Sosso, D.; Qu, X.Q.; Chen, L.Q.; Ma, L.; Chermak, D.; Zhang, D.C.; Frommer, W.B. Phylogenetic evidence for a fusion of archaeal and bacterial SemiSWEETs to form eukaryotic SWEETs and identification of SWEET hexose transporters in the amphibian chytrid pathogen Batrachochytrium dendrobatidis. FASEB J. 2016, 30, 3644–3654. [Google Scholar] [CrossRef] [Green Version]
- Xie, H.; Wang, D.; Qin, Y.; Ma, A.; Zhao, J. Genome-wide identification and expression analysis of sweet gene family in Litchi chinensis reveal the involvement of Lcsweet2a/3b in early seed development. BMC Plant Biol. 2019, 19, 499. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, S.; Ren, Y.; Gan, C.; Li, B.; Fan, Y.; Zhao, X.; Yuan, Z. Identification, Analysis and Gene Cloning of the SWEET Gene Family Provide Insights into Sugar Transport in Pomegranate (Punica granatum). Int. J. Mol. Sci. 2022, 23, 2471. [Google Scholar] [CrossRef]
- Lin, I.W.; Sosso, D.; Chen, L.Q.; Gase, K.; Kim, S.G.; Kessler, D.; Klinkenberg, P.M.; Gorder, M.K.; Hou, B.U.; Qu, X.Q.; et al. Nectar secretion requires sucrose phosphate synthases and the sugar transporter SWEET9. Nature 2014, 508, 546–549. [Google Scholar] [CrossRef]
- Ko, H.Y.; Tseng, H.W.; Ho, L.H.; Wang, L.; Chang, T.F.; Lin, A.; Ruan, Y.L.; Neuhaus, H.E.; Guo, W.J. Hexose translocation mediated by SlSWEET5b is required for pollen maturation in Solanum lycopersicum. Plant Physiol. 2022, 189, 344–359. [Google Scholar] [CrossRef]
- Kanno, Y.; Oikawa, T.; Chiba, Y.; Ishimaru, Y.; Shimizu, T.; Sano, N.; Koshiba, T.; Kamiya, Y.J.; Ueda, M.; Seo, M. AtSWEET13 and AtSWEET14 regulate gibberellin-mediated physiological processes. Nat. Commun. 2016, 7, 13245. [Google Scholar] [CrossRef]
- Ninan, A.; Grant, J.; Song, J.; Jameson, P. Expression of Genes Related to Sugar and Amino Acid Transport and Cytokinin Metabolism during Leaf Development and Senescence in Pisum sativum L. Plants 2019, 8, 76. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, Y.; Wu, P.; Xu, S.; Chen, Y.; Li, M.; Wu, G.; Jiang, H. Genome-Wide Identification, Expression Patterns and Sugar Transport of the Physic Nut SWEET Gene Family and a Functional Analysis of JcSWEET16 in Arabidopsis. Int. J. Mol. Sci. 2022, 23, 5391. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Zhang, F.; Song, S.; Yu, X.; Ren, Y.; Zhao, X.; Liu, H.; Liu, G.; Wang, Y.; He, H. CsSWEET2, a Hexose Transporter from Cucumber (Cucumis sativus L.), Affects Sugar Metabolism and Improves Cold Tolerance in Arabidopsis. Int. J. Mol. Sci. 2022, 23, 3886. [Google Scholar] [CrossRef] [PubMed]
- Xuan, C.; Lan, G.; Si, F.; Zeng, Z.; Wang, C.; Yadav, V.; Wei, C.; Zhang, X. Systematic Genome-Wide Study and Expression Analysis of SWEET Gene Family, Sugar Transporter Family Contributes to Biotic and Abiotic Stimuli in Watermelon. Int. J. Mol. Sci. 2021, 22, 8407. [Google Scholar] [CrossRef] [PubMed]
- Klemens, P.A.; Patzke, K.; Deitmer, J.W.; Spinner, L.; Hir, R.L.; Bellini, C.; Bedu, M.; Chardon, F.; Krapp, A.; Neuhaus, H.E. Overexpression of the Vacuolar Sugar Carrier AtSWEET16 Modifies Germination, Growth, and Stress Tolerance in Arabidopsis. Plant Physiol. 2013, 163, 1338–1352. [Google Scholar] [CrossRef] [Green Version]
- Phukan, U.J.; Jeena, G.S.; Tripathi, V.; Shukla, R.K. MaRAP2-4, a waterlogging-responsive ERF from Mentha, regulates bidirectional sugar transporter AtSWEET10 to modulate stress response in Arabidopsis. Plant Biotechnol. J. 2017, 16, 221–233. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Yao, L.; Hao, X.; Li, N.; Qian, W.; Yue, C.; Ding, C.; Zeng, J.; Yang, Y.; Wang, X. Tea plant SWEET transporters, expression profiling, sugar transport, and the involvement of CsSWEET16 in modifying cold tolerance in Arabidopsis. Plant Mol. Biol. 2018, 96, 577–592. [Google Scholar] [CrossRef]
- Jiang, S.; Balan, B.; Assis, R.d.A.B.; Sagawa, C.H.D.; Wan, X.; Han, S.; Wang, L.; Zhang, L.; Zaini, P.A.; Walawage, S.L.; et al. Genome-Wide Profiling and Phylogenetic Analysis of the SWEET Sugar Transporter Gene Family in Walnut and Their Lack of Responsiveness to Xanthomonas arboricola pv. juglandis Infection. Int. J. Mol. Sci. 2020, 21, 1251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Streubel, J.; Pesce, C.; Hutin, M.; Koebnik, R.; Boch, J.; Szurek, B. Five phylogenetically close rice SWEET genes confer TAL effector-mediated susceptibility to Xanthomonas oryzae pv. oryzae. New Phytol. 2013, 200, 808–819. [Google Scholar] [CrossRef]
- Cohn, M.; Bart, R.S.; Shybut, M.; Dahlbeck, D.; Gomez, M.; Morbitzer, R.; Hou, B.H.; Frommer, W.B.; Lahaye, T.; Staskawicz, B.J. Xanthomonas axonopodis virulence is promoted by a transcription activator-like effector-mediated induction of a SWEET sugar transporter in cassava. Mol. Plant Microbe Interact. 2014, 27, 1186–1198. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, W.; Wang, S.; Yu, F.; Tang, J.; Shan, X.; Bao, K.; Yu, L.; Wang, H.; Fei, Z.; Li, J. Genome-wide characterization and expression profiling of SWEET genes in cabbage (Brassica oleracea var. capitata L.) reveal their roles in chilling and club root disease responses. BMC Genom. 2019, 20, 93. [Google Scholar]
- Sosso, D.; Luo, D.; Li, Q.B.; Sasse, J.; Yang, J.; Gendrot, G.; Suzuki, M.; Koch, K.K.; McCarty, D.R.; Chourey, P.S.; et al. Seed filling in domesticated maize and rice depends on sweet-mediated hexose transport. Nat. Genet. 2015, 47, 1489–1493. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Li, H.; Xia, X.; Liu, X.; Yang, L. Functional and evolution characterization of SWEET sugar transporters in Ananas comosus. Biochem. Biophys. Res. Commun. 2018, 496, 407–414. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zou, L.; Ren, C.; Ren, F.; Wang, Y.; Fan, P.; Li, S.; Liang, Z. VvSWEET10 mediates sugar accumulation in grapes. Genes 2019, 10, 255. [Google Scholar] [CrossRef] [Green Version]
- Rangkadilok, N.; Worasuttayangkurn, L.; Bennett, R.N.; Satayavivad, J. Identification and quantification of polyphenolic compounds in Longan (Euphoria longana Lam.) fruit. J. Agric. Food Chem. 2005, 53, 1387–1392. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Zhou, C.; Wan, Z. Analysis on the development status of lychee industry in Guangdong province in 2010. Guangdong Agric. Sci. 2011, 4, 16–18. [Google Scholar]
- Yang, X.; Wang, R.; Jing, H.; Chen, Q.; Fu, J. Three novel c-repeat binding factor genes of dimocarpus longan regulate cold stress response in Arabidopsis. Front. Plant Sci. 2020, 11, 1026. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Min, J.; Lai, R.; Wu, Z.; Chen, Y.; Yu, L.; Cheng, C.; Jin, Y.; Tian, Q.; Liu, Q.; et al. Genome-wide sequencing of longan (Dimocarpus longan Lour.) provides insights into molecular basis of its polyphenol-rich characteristics. Gigascience 2017, 6, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Chen, L.Q. SWEET sugar transporters for phloem transport and pathogen nutrition. New Phytol. 2014, 201, 1150–1155. [Google Scholar] [CrossRef]
- Eom, J.S.; Chen, L.Q.; Sosso, D.; Julius, B.T.; Lin, I.W.; Qu, X.Q.; Braun, D.M.; Frommer, W.B. SWEETs, transporters for intracellular and intercellular sugar translocation. Curr. Opin. Plant Biol. 2015, 25, 53–62. [Google Scholar] [CrossRef] [Green Version]
- Patil, G.; Valliyodan, B.; Deshmukh, R.K.; Prince, S.J.; Nicander, B.; Zhao, M.; Sonah, H.; Song, L.; Lin, L.; Chaudhary, J.; et al. Soybean (Glycine max) SWEET gene family, insights through comparative genomics, transcriptome profiling and whole genome re-sequence analysis. BMC Genom. 2015, 16, 520. [Google Scholar] [CrossRef] [Green Version]
- Jia, B.; Zhu, X.F.; Pu, Z.J.; Duan, Y.X.; Hao, L.J.; Zhang, J.; Chen, L.Q.; Jeon, C.O.; Xuan, Y.H. Integrative view of the diversity and evolution of SWEET and SemiSWEET sugar transporters. Front. Plant Sci. 2017, 8, 2178. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gautam, T.; Saripalli, G.; Gahlaut, V.; Kumar, A.; Sharma, P.K.; Balyan, H.S.; Gupta, P.K. Further studies on sugar transporter (SWEET) genes in wheat (Triticum aestivum L.). Mol. Biol. Rep. 2019, 46, 2327–2353. [Google Scholar] [CrossRef]
- Chong, J.; Piron, M.C.; Meyer, S.; Merdinoglu, D.; Bertsch, C.; Mestre, P. The sweet family of sugar transporters in grapevine, VvSWEET4 is involved in the interaction with Botrytis cinerea. J. Exp. Bot. 2014, 65, 6589–6601. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miao, H.; Sun, P.; Liu, Q.; Miao, Y.; Liu, J.; Zhang, K.; Hu, W.; Zhang, J.; Wang, J.; Wang, Z.; et al. Genome wide analyses of SWEET family proteins reveal involvement in fruit development and abiotic/biotic stress responses in banana. Sci. Rep. 2017, 7, 3536. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, B.; Wu, H.; Huang, W.; Song, J.; Zhou, Y.; Lin, Y. SWEET Gene Family in Medicago truncatula, Genome-Wide Identification, Expression and Substrate Specificity Analysis. Plants 2019, 8, 338. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.; Yokosho, K.; Guo, R.; Whelan, J.; Ruan, Y.L.; Ma, J.F.; Shou, H. The soybean sugar transporter GmSWEET15 mediates sucrose export from endosperm to early embryo. Plant Physiol. 2019, 180, 2133–2141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, Y.; Wang, Y.; Shan, Y.; Qin, Q. Characterization of SWEET family members from loquat and their responses to exogenous induction. Tree Genet. Genomes 2017, 13, 123. [Google Scholar] [CrossRef]
- Feng, C.Y.; Han, J.X.; Han, X.X.; Jiang, J. Genome-wide identification, phylogeny, and expression analysis of the SWEET gene family in tomato. Gene 2015, 573, 261–272. [Google Scholar] [CrossRef]
- Li, J.; Qin, M.; Qiao, X.; Cheng, Y.; Li, X.; Zhang, H.; Wu, J. A new insight into the evolution and functional divergence of sweet transporters in Chinese white pear (Pyrus bretschneideri). Plant Cell Physiol. 2017, 58, 839–850. [Google Scholar] [CrossRef] [Green Version]
- Hu, L.; Zhang, F.; Song, S.; Tang, X.; Xu, X.; Liu, G.; Wang, Y.; He, H. Genome-wide identification, characterization, and expression analysis of the SWEET gene family in cucumber. J. Integr. Agric. 2017, 16, 60345–60347. [Google Scholar] [CrossRef] [Green Version]
- An, Z.Y.; Fang, R.; Huang, X.W.; Yao, J.Y.; Wei, S.T. Cloning and Expression Analysis of a SWEET Family Gene from Annona squamosa L. Chin. J. Trop. Crops 2020, 41, 2143–2148. [Google Scholar]
- Chen, L.Q.; Lin, I.W.; Qu, X.Q.; Sosso, D.; McFarlane, H.E.; Londoño, A.; Samuels, A.L.; Frommer, W.B. A cascade of sequentially expressed sucrose transporters in the seed coat and endosperm provides nutrition for the Arabidopsis embryo. Plant Cell 2015, 27, 607–619. [Google Scholar] [CrossRef] [Green Version]
- Gibson, S.I. Control of plant development and gene expression by sugar signaling. Curr. Opin. Plant Biol. 2005, 8, 93–102. [Google Scholar] [CrossRef]
- Yamada, K.; Osakabe, Y.; Mizoi, J.; Nakashima, K.; Fujita, Y.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Functional analysis of an Arabidopsis thaliana abiotic stress-inducible facilitated diffusion transporter for monosaccharides. J. Biol. Chem. 2010, 285, 1138–1146. [Google Scholar] [CrossRef] [Green Version]
- Yue, C.; Cao, H.; Wang, L.; Zhou, Y.; Huang, Y.; Hao, X.; Wang, Y.; Wang, B.; Yang, Y.; Wang, X. Effects of cold acclimation on sugar metabolism and sugar-related gene expression in tea plant during the winter season. Plant Mol. Biol. 2015, 88, 591–608. [Google Scholar] [CrossRef]
- Fang, T.; Cai, Y.; Yang, Q.; Ogutu, C.; Liao, L.; Han, Y. Analysis of sorbitol content variation in wild and cultivated apples. J. Sci. Food Agric. 2020, 100, 139–144. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6, Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [Green Version]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar] [PubMed]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0, An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.; Frank, M.; He, Y.; Xia, R. TBtools—An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
| Gene ID | Gene Name | Clade | Location | gDNA (bp) | CDS (bp) | Protein (aa) | PI | MV (KDa) | GRAVY | TMHs |
|---|---|---|---|---|---|---|---|---|---|---|
| Dlo_004842.1 | DlSWEET1 | I | scaffold132:1222727..1224250(+) | 1524 | 750 | 249 | 9.44 | 27.20 | 0.678 | 7 |
| Dlo_011035.1 | DlSWEET2a | I | scaffold208:248691..251666(−) | 2976 | 732 | 243 | 9.46 | 27.10 | 0.77 | 7 |
| Dlo_031144.1 | DlSWEET2b | I | scaffold81:587713..595463(−) | 7751 | 1323 | 440 | 9.32 | 49.29 | 0.236 | 6 |
| Dlo_001364.1 | DlSWEET3a | I | scaffold105:240059..241538(−) | 1480 | 639 | 212 | 9.24 | 24.07 | 0.509 | 6 |
| Dlo_001358.3 | DlSWEET3b | I | scaffold105:129309..133690(−) | 4382 | 726 | 241 | 8.54 | 26.49 | 0.559 | 7 |
| Dlo_001362.1 | DlSWEET3c | I | scaffold105:227382..229337(−) | 1956 | 756 | 251 | 9.06 | 28.19 | 0.459 | 7 |
| Dlo_016654.1 | DlSWEET4 | II | scaffold32:314489..315798(−) | 1310 | 573 | 190 | 9.54 | 21.48 | 0.729 | 5 |
| Dlo_012330.1 | DlSWEET7a | II | scaffold23:111069..112837(−) | 1769 | 714 | 237 | 9.28 | 26.51 | 0.833 | 7 |
| Dlo_036665.1 | DlSWEET7b | II | scaffold1269:54809..57866(+) | 3058 | 780 | 260 | 9.1 | 29.08 | 0.767 | 7 |
| Dlo_035889.1 | DlSWEET7c | II | scaffold1269:54830..57818(+) | 2989 | 711 | 237 | 9 | 26.66 | 0.914 | 6 |
| Dlo_002779.1 | DlSWEET9a | III | scaffold1141:110734..112484(+) | 1751 | 747 | 248 | 9.38 | 27.76 | 0.496 | 7 |
| Dlo_002777.1 | DlSWEET9b | III | scaffold1141:86225..87669(+) | 1445 | 636 | 211 | 9.15 | 23.77 | 0.602 | 5 |
| Dlo_002778.1 | DlSWEET9c | III | scaffold1141:97643..100738(+) | 3096 | 777 | 258 | 9.28 | 28.98 | 0.416 | 7 |
| Dlo_024131.1 | DlSWEET9d | III | scaffold539:83282..84431(+) | 1150 | 765 | 254 | 9.01 | 28.42 | 0.531 | 6 |
| Dlo_006037.1 | DlSWEET10a | III | scaffold1436:78359..79825(+) | 1467 | 855 | 284 | 9.4 | 32.10 | 0.603 | 7 |
| Dlo_005388.1 | DlSWEET10b | III | scaffold139:126837..128875(−) | 2039 | 630 | 209 | 8.4 | 23.76 | 0.482 | 5 |
| Dlo_006035.1 | DlSWEET10c | III | scaffold1436:44684..45953(+) | 1270 | 810 | 269 | 9.17 | 29.76 | 0.645 | 7 |
| Dlo_026392.1 | DlSWEET15 | III | scaffold601:142..2474(+) | 2333 | 741 | 246 | 5.77 | 27.80 | 0.574 | 5 |
| Dlo_006477.1 | DlSWEET16a | IV | scaffold148:330655..332743(−) | 2089 | 735 | 244 | 8.56 | 27.11 | 0.645 | 6 |
| Dlo_006478.1 | DlSWEET16b | IV | scaffold148:336807..338137(−) | 1331 | 423 | 140 | 6.17 | 15.48 | 0.344 | 3 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fang, T.; Rao, Y.; Wang, M.; Li, Y.; Liu, Y.; Xiong, P.; Zeng, L. Characterization of the SWEET Gene Family in Longan (Dimocarpus longan) and the Role of DlSWEET1 in Cold Tolerance. Int. J. Mol. Sci. 2022, 23, 8914. https://doi.org/10.3390/ijms23168914
Fang T, Rao Y, Wang M, Li Y, Liu Y, Xiong P, Zeng L. Characterization of the SWEET Gene Family in Longan (Dimocarpus longan) and the Role of DlSWEET1 in Cold Tolerance. International Journal of Molecular Sciences. 2022; 23(16):8914. https://doi.org/10.3390/ijms23168914
Chicago/Turabian StyleFang, Ting, Ya Rao, Mengzhen Wang, Yun Li, Yujun Liu, Pengpeng Xiong, and Lihui Zeng. 2022. "Characterization of the SWEET Gene Family in Longan (Dimocarpus longan) and the Role of DlSWEET1 in Cold Tolerance" International Journal of Molecular Sciences 23, no. 16: 8914. https://doi.org/10.3390/ijms23168914
APA StyleFang, T., Rao, Y., Wang, M., Li, Y., Liu, Y., Xiong, P., & Zeng, L. (2022). Characterization of the SWEET Gene Family in Longan (Dimocarpus longan) and the Role of DlSWEET1 in Cold Tolerance. International Journal of Molecular Sciences, 23(16), 8914. https://doi.org/10.3390/ijms23168914







