Identification and Characterization of the Core Region of ZmDi19-5 Promoter Activity and Its Upstream Regulatory Proteins
Abstract
:1. Introduction
2. Results
2.1. Identification of ZmDi19 Genes
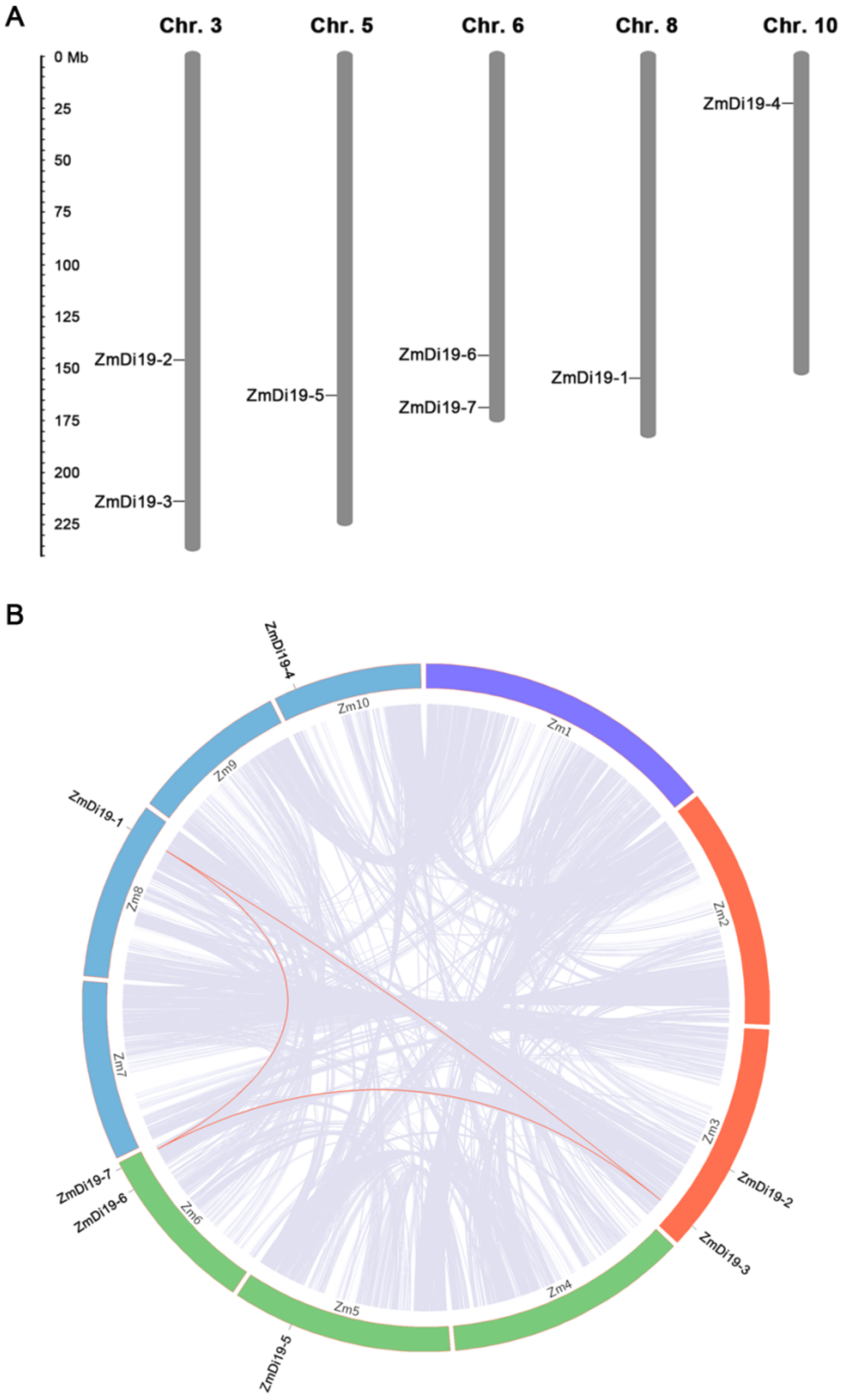
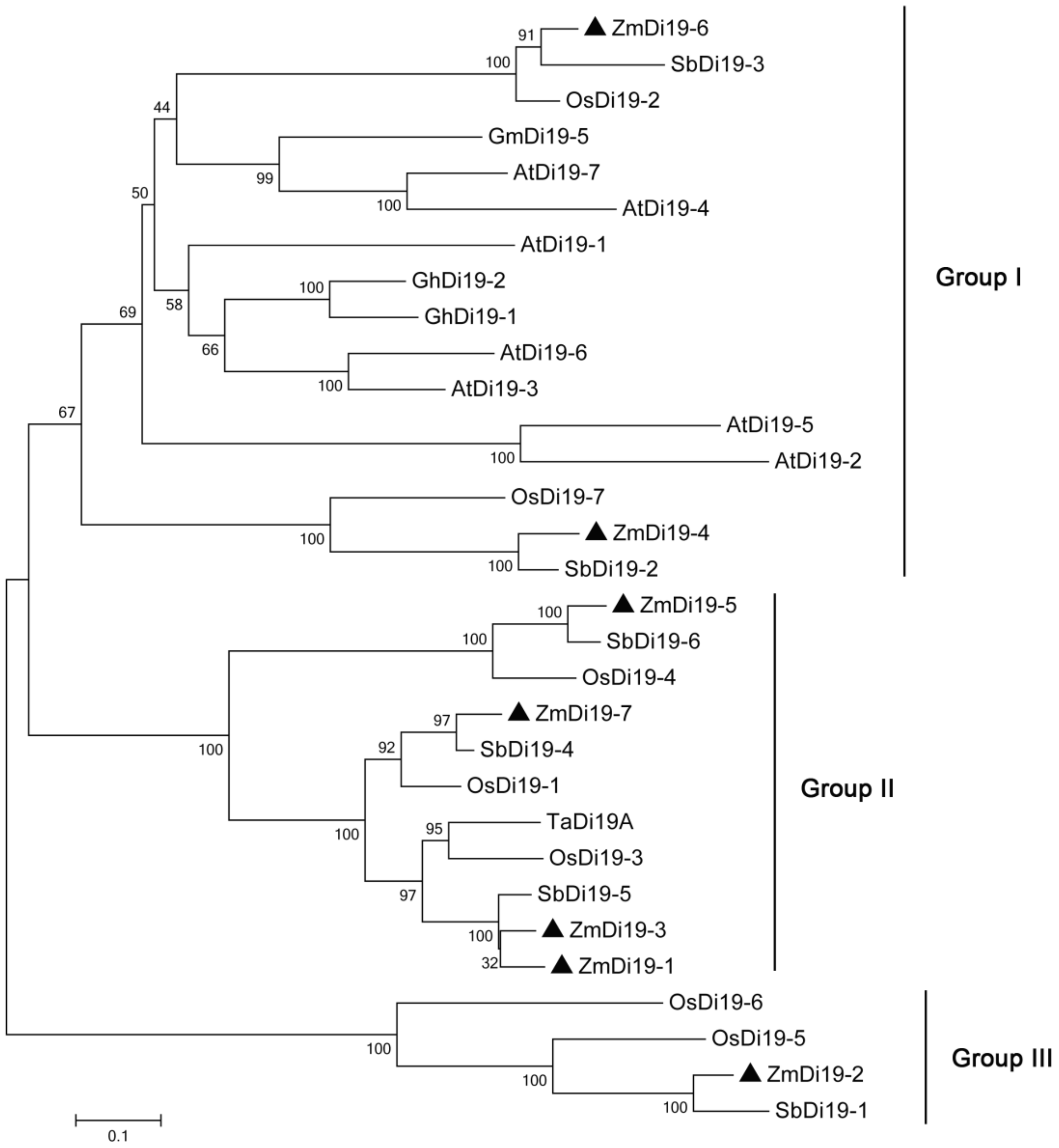
2.2. Phylogenetic Relationships, Gene Structure, Chromosomal Location, and Gene Duplication
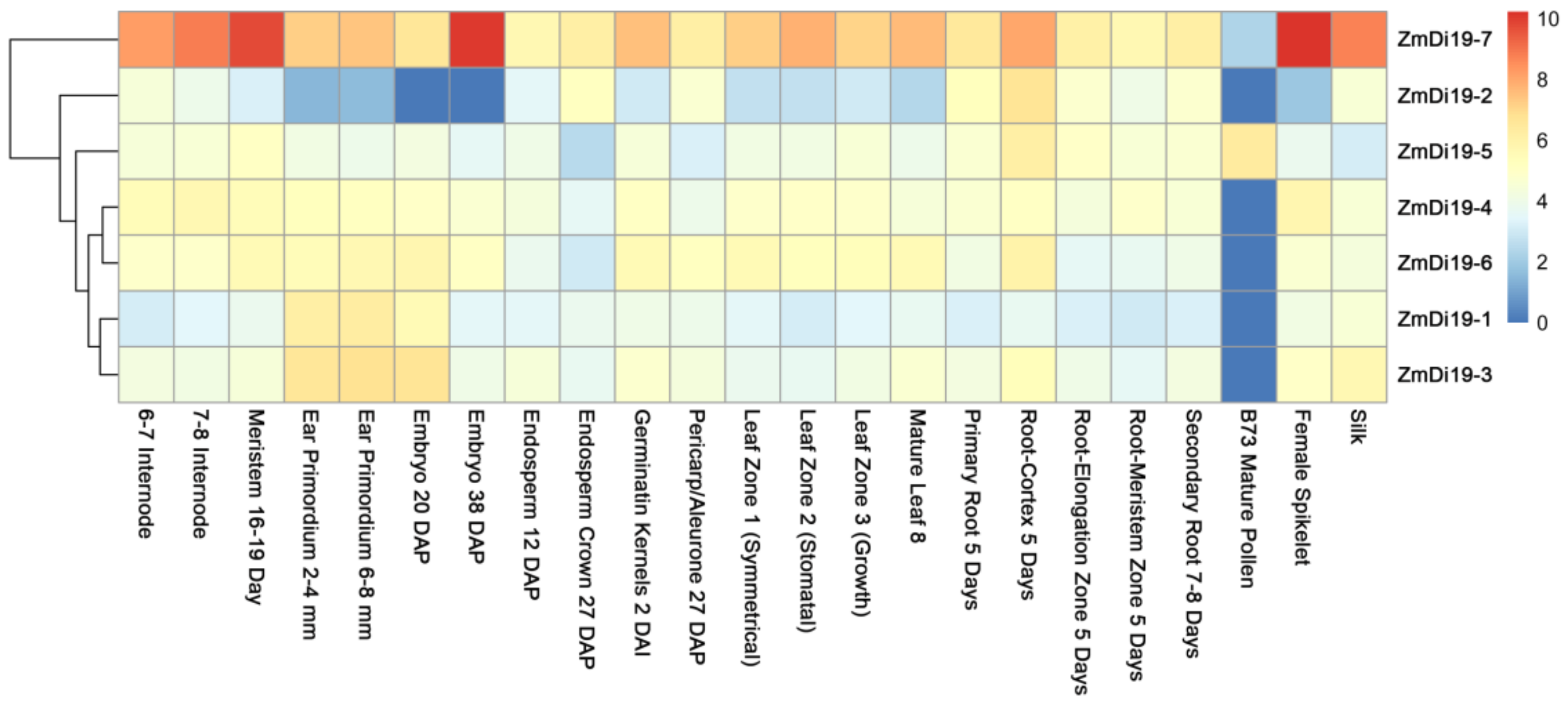
2.3. Expression Pattern Analysis of ZmDi19 Genes at Different Developmental Stages
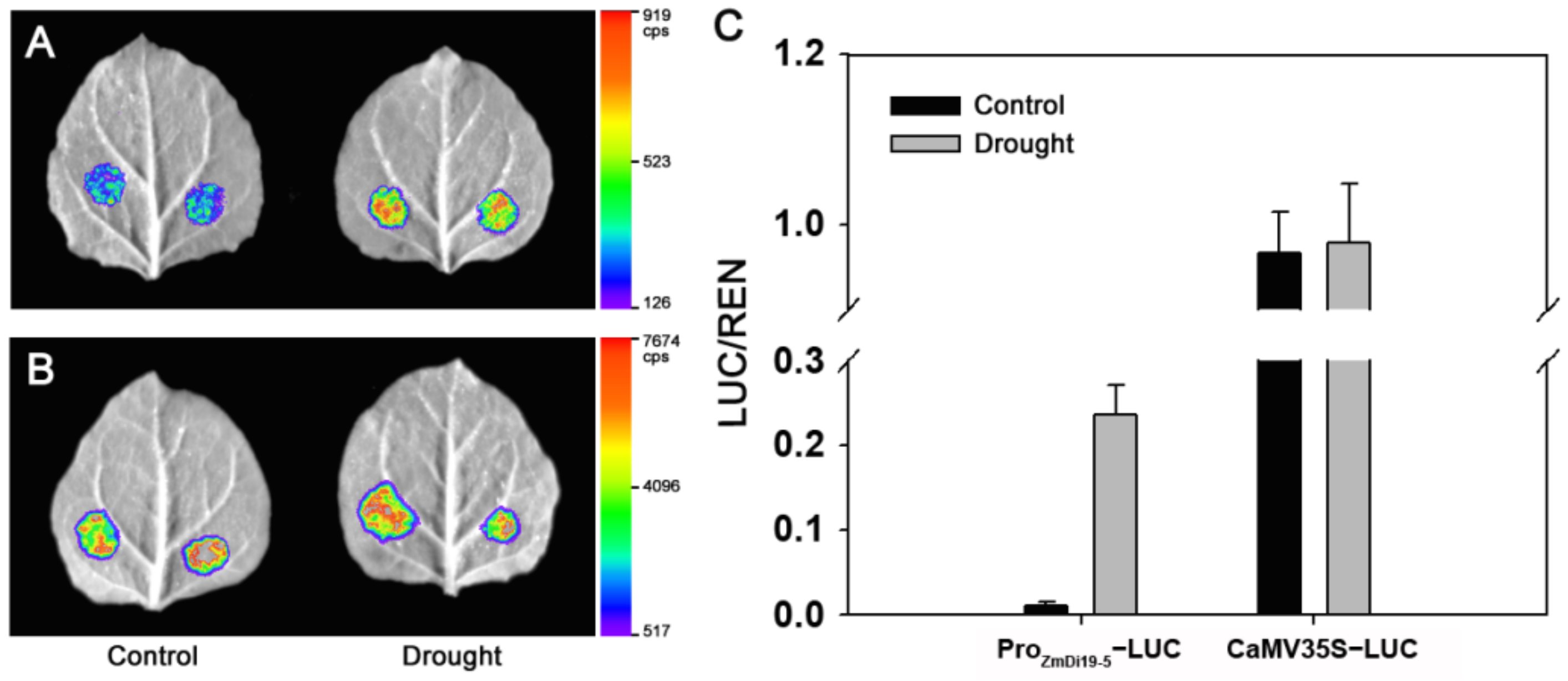
2.4. Sequence Analysis and Expressed Activity of ZmDi19-5 Promoter under Drought Stress
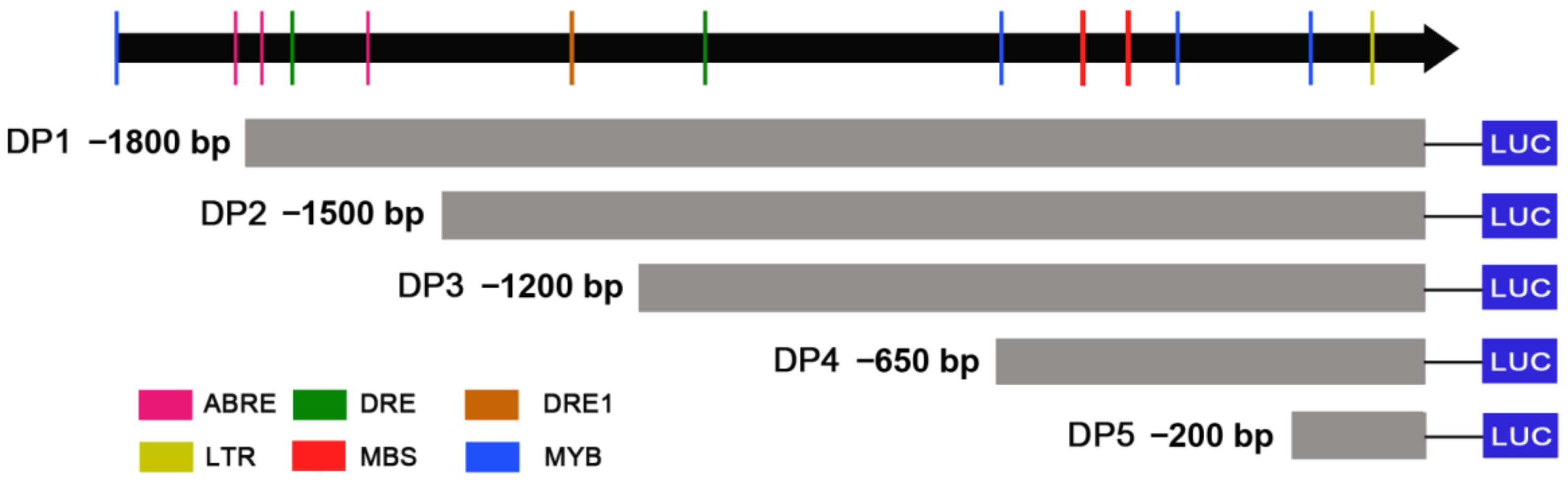
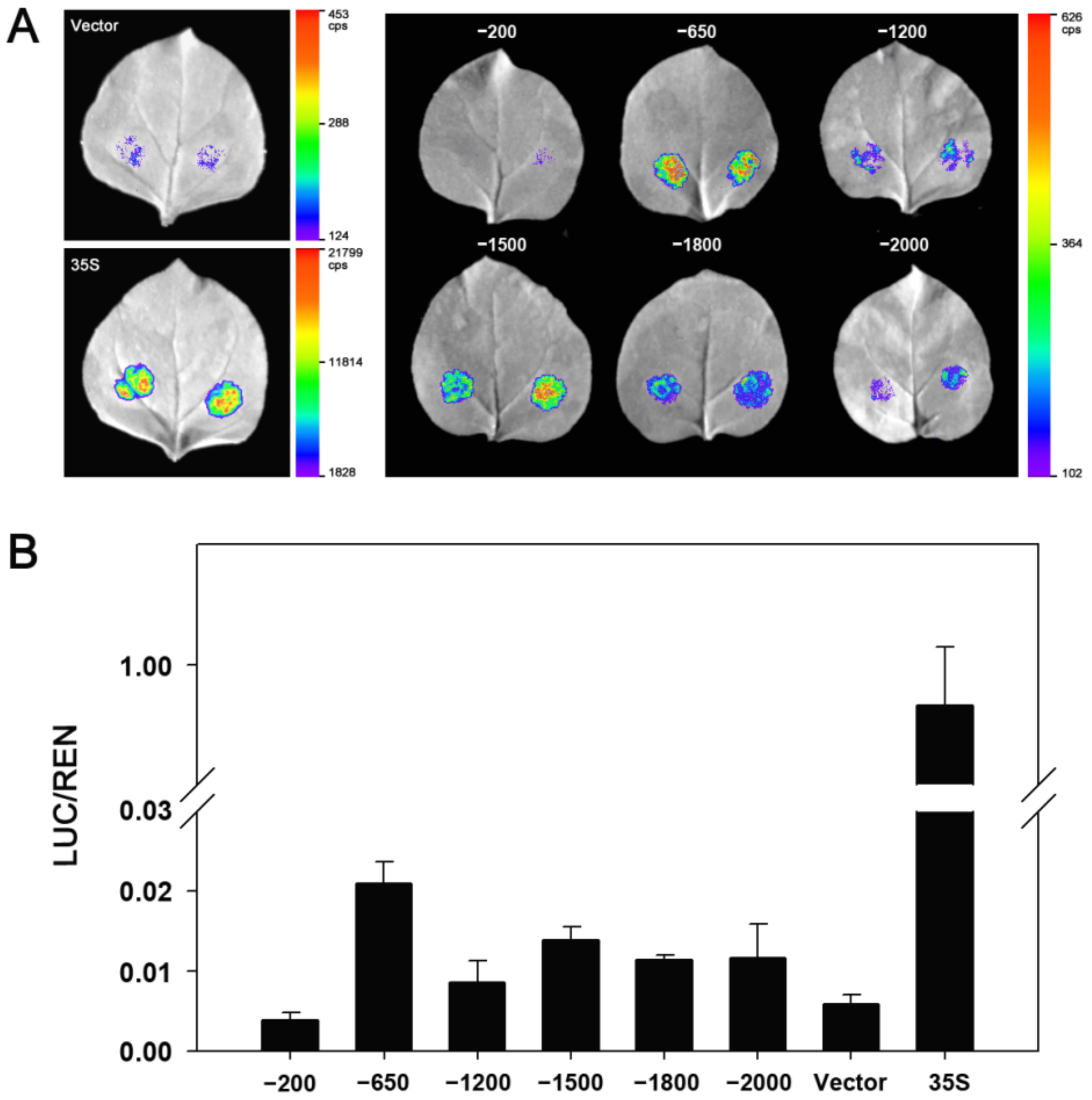
2.5. Identification of Core Region of ZmDi19-5 Promoter
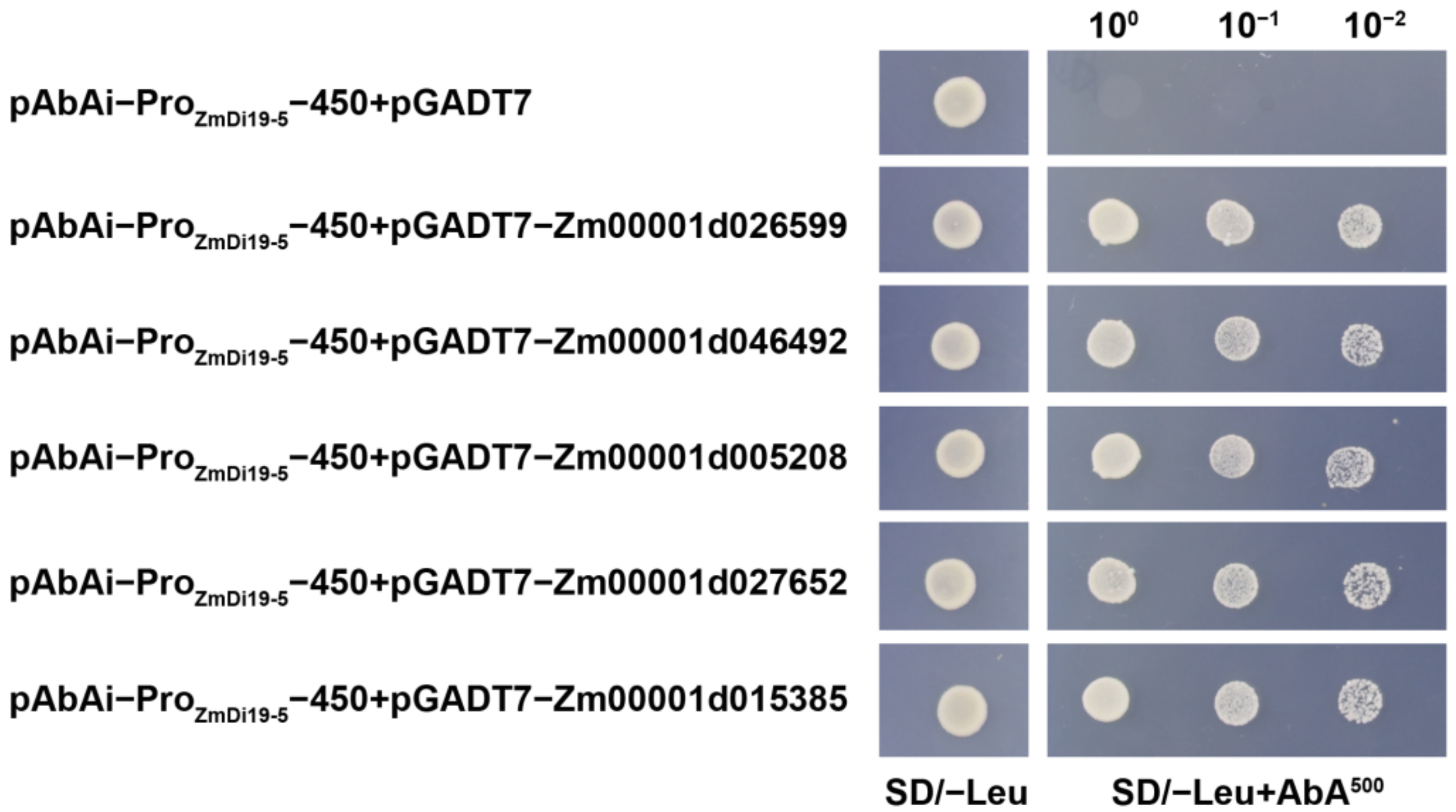
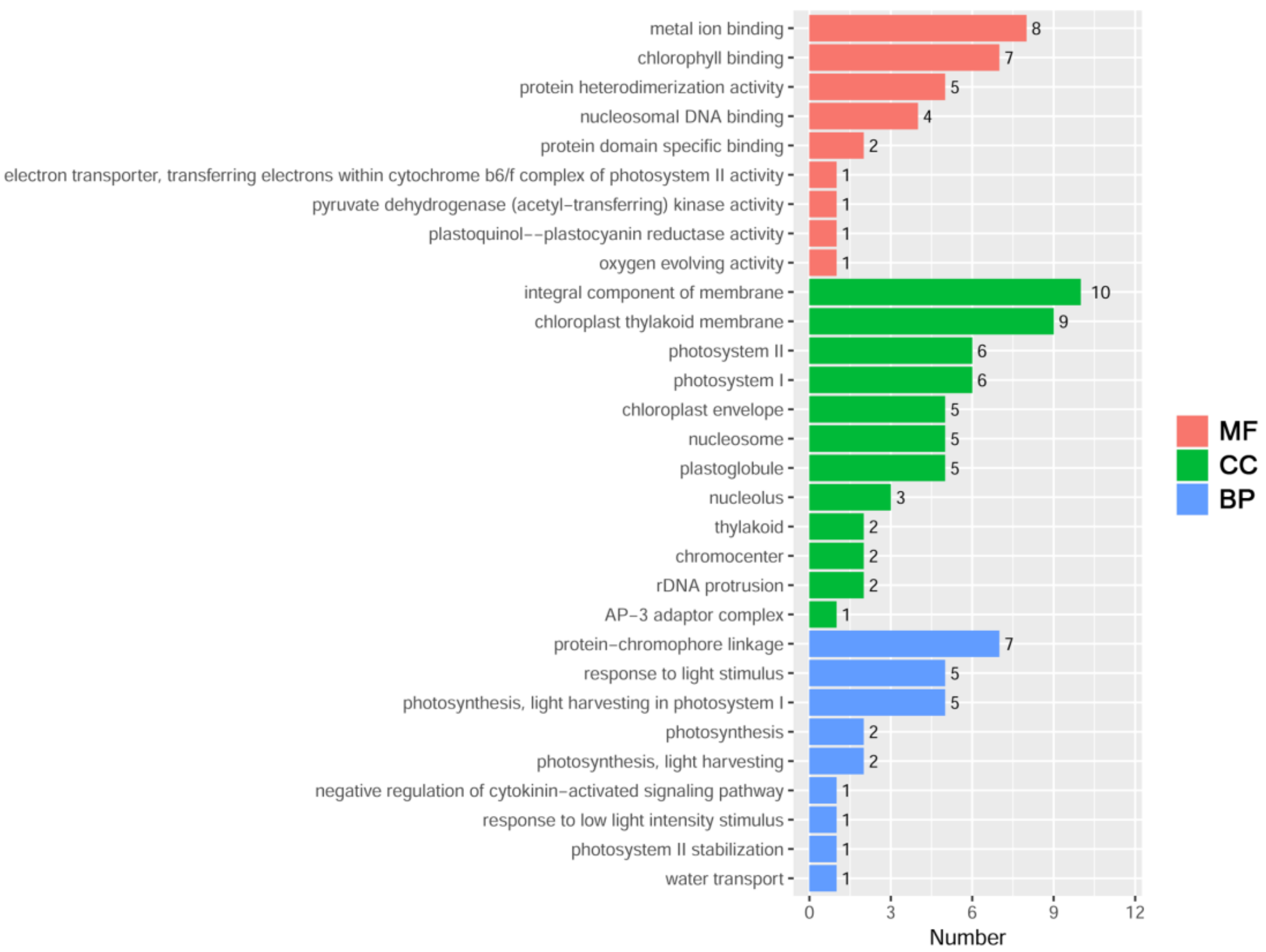
2.6. Identification of Upstream Regulatory Proteins of ZmDi19-5
3. Discussion
4. Materials and Methods
4.1. Identification and Sequence Analysis of Di19 Genes in Maize
4.2. Phylogenetic Analysis
4.3. Expression Pattern Analysis Using Transcriptome Data
4.4. Cloning and Sequence Analysis of ZmDi19-5 Promoter
4.5. Transient Transformation in Tobacco
4.6. Determination of LUC Activity and Identification of Promoter Core Region
4.7. Yeast One-Hybrid Library Screening
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Xiong, L.; Schumaker, K.S.; Zhu, J.K. Cell signaling during cold, drought, and salt stress. Plant Cell 2002, 14, S165–S183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, J.K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xiang, Y.; Tang, N.; Du, H.; Ye, H.Y.; Xiong, L.Z. Characterization of OsbZIP23 as a Key Player of the Basic Leucine Zipper Transcription Factor Family for Conferring Abscisic Acid Sensitivity and Salinity and Drought Tolerance in Rice. Plant Physiol. 2008, 148, 1938–1952. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pastori, G.M.; Foyer, C.H. Common components, networks, and pathways of cross-tolerance to stress. The central role of “redox” and abscisic acid-mediated controls. Plant Physiol. 2002, 129, 460–468. [Google Scholar] [CrossRef] [Green Version]
- Shinozaki, K.; Yamaguchi-Shinozaki, K. Molecular responses to drought and cold stress. Curr. Opin. Biotechnol. 1996, 7, 161–167. [Google Scholar] [CrossRef]
- Shinozaki, K.; Yamaguchi-Shinozaki, K.; Seki, M. Regulatory network of gene expression in the drought and cold stress responses. Curr. Opin. Plant Biol. 2003, 6, 410–417. [Google Scholar] [CrossRef]
- Ramachandran, S.; Hiratsuka, K.; Chua, N.H. Transcription factors in plant growth and development. Curr. Opin. Genet. Dev. 1994, 4, 642–646. [Google Scholar] [CrossRef]
- Spitz, F.; Furlong, E.E. Transcription factors: From enhancer binding to developmental control. Nat. Rev. Genet. 2012, 13, 613–626. [Google Scholar] [CrossRef]
- Singh, K.; Foley, R.C.; Onate-Sanchez, L. Transcription factors in plant defense and stress responses. Curr. Opin. Plant Biol. 2002, 5, 430–436. [Google Scholar] [CrossRef]
- Yamasaki, K.; Kigawa, T.; Inoue, M.; Watanabe, S.; Tateno, M.; Seki, M.; Shinozaki, K.; Yokoyama, S. Structures and evolutionary origins of plant-specific transcription factor DNA-binding domains. Plant Physiol. Biochem. 2008, 46, 394–401. [Google Scholar] [CrossRef]
- Hichri, I.; Muhovski, Y.; Clippe, A.; Zizkova, E.; Dobrev, P.I.; Motyka, V.; Lutts, S. SlDREB2, a tomato dehydration-responsive element-binding 2 transcription factor, mediates salt stress tolerance in tomato and Arabidopsis. Plant Cell Environ. 2016, 39, 62–79. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.Y.; Wang, Y.C.; Wang, L.Q.; Hu, P.; Wang, Y.M.; Jia, Y.Y.; Zhang, C.R.; Zhang, Y.; Zhang, Y.M.; Wang, C.; et al. Expression of the MYB transcription factor gene BplMYB46 affects abiotic stress tolerance and secondary cell wall deposition in Betula platyphylla. Plant Biotechnol. J. 2017, 15, 107–121. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Hong, Y.; Zhang, H.; Li, D.; Song, F. Rice NAC transcription factor ONAC095 plays opposite roles in drought and cold stress tolerance. BMC Plant Biol. 2016, 16, 203. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Ma, Q.; Jin, X.; Peng, X.; Liu, J.; Deng, L.; Yan, H.; Sheng, L.; Jiang, H.; Cheng, B. A novel maize homeodomain-leucine zipper (HD-Zip) I gene, Zmhdz10, positively regulates drought and salt tolerance in both rice and Arabidopsis. Plant Cell Physiol. 2014, 55, 1142–1156. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, H.; Wang, Y.; Xu, P.; Zhang, Z. Overexpression of a WRKY Transcription Factor TaWRKY2 Enhances Drought Stress Tolerance in Transgenic Wheat. Front. Plant Sci. 2018, 9, 997. [Google Scholar] [CrossRef] [Green Version]
- Milla, M.A.; Townsend, J.; Chang, I.F.; Cushman, J.C. The Arabidopsis AtDi19 gene family encodes a novel type of Cys2/His2 zinc-finger protein implicated in ABA-independent dehydration, high-salinity stress and light signaling pathways. Plant Mol. Biol. 2006, 61, 13–30. [Google Scholar] [CrossRef]
- Liu, W.X.; Zhang, F.C.; Zhang, W.Z.; Song, L.F.; Wu, W.H.; Chen, Y.F. Arabidopsis Di19 Functions as a Transcription Factor and Modulates PR1, PR2, and PR5 Expression in Response to Drought Stress. Mol. Plant 2013, 6, 1487–1502. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.L.; Yu, C.C.; Chen, C.; He, C.L.; Zhu, Y.G.; Huang, W.C. Identification of rice Di19 family reveals OsDi19-4 involved in drought resistance. Plant Cell Rep. 2014, 33, 2047–2062. [Google Scholar] [CrossRef]
- Brown, R.S.; Sander, C.; Argos, P. The primary structure of transcription factor TFIIIA has 12 consecutive repeats. FEBS Lett. 1985, 186, 271–274. [Google Scholar] [CrossRef] [Green Version]
- Miller, J.; McLachlan, A.D.; Klug, A. Repetitive zinc-binding domains in the protein transcription factor IIIA from Xenopus oocytes. EMBO J. 1985, 4, 1609–1614. [Google Scholar] [CrossRef]
- Pabo, C.O.; Peisach, E.; Grant, R.A. Design and selection of novel Cys2His2 zinc finger proteins. Annu. Rev. Biochem. 2001, 70, 313–340. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.X.; Li, Y.; Li, D.D.; Xu, W.L.; Zheng, Y.; Li, X.B. Arabidopsis drought-induced protein Di19-3 participates in plant response to drought and high salinity stresses. Plant Mol. Biol. 2014, 86, 609–625. [Google Scholar] [CrossRef] [PubMed]
- Feng, Z.J.; Cui, X.Y.; Cui, X.Y.; Chen, M.; Yang, G.X.; Ma, Y.Z.; He, G.Y.; Xu, Z.S. The soybean GmDi19-5 interacts with GmLEA3.1 and increases sensitivity of transgenic plants to abiotic stresses. Front. Plant Sci. 2015, 6, 179. [Google Scholar] [CrossRef] [Green Version]
- Li, S.; Xu, C.; Yang, Y.; Xia, G. Functional analysis of TaDi19A, a salt-responsive gene in wheat. Plant Cell Environ. 2010, 33, 117–129. [Google Scholar]
- Li, G.; Tai, F.J.; Zheng, Y.; Luo, J.; Gong, S.Y.; Zhang, Z.T.; Li, X.B. Two cotton Cys2/His2-type zinc-finger proteins, GhDi19-1 and GhDi19-2, are involved in plant response to salt/drought stress and abscisic acid signaling. Plant Mol. Biol. 2010, 74, 437–452. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.X.; Nie, X.Y.; Hu, R.; Li, G.; Xu, W.L.; Li, X.B. Phosphorylation of serine residue modulates cotton Di19-1 and Di19-2 activities for responding to high salinity stress and abscisic acid signaling. Sci. Rep. 2016, 6, 20371. [Google Scholar] [CrossRef]
- Dong, Q.; Jiang, H.; Xu, Q.; Li, X.; Peng, X.; Yu, H.; Xiang, Y.; Cheng, B. Cloning and characterization of a multifunctional promoter from maize (Zea mays L.). Appl. Biochem. Biotechnol. 2015, 175, 1344–1357. [Google Scholar] [CrossRef]
- Hernandez-Garcia, C.M.; Finer, J.J. Identification and validation of promoters and cis-acting regulatory elements. Plant Sci. 2014, 217, 109–119. [Google Scholar] [CrossRef] [Green Version]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci. 2005, 10, 88–94. [Google Scholar] [CrossRef]
- Walley, J.W.; Sartor, R.C.; Shen, Z.; Schmitz, R.J.; Wu, K.J.; Urich, M.A.; Nery, J.R.; Smith, L.G.; Schnable, J.C.; Ecker, J.R.; et al. Integration of omic networks in a developmental atlas of maize. Science 2016, 353, 814–818. [Google Scholar] [CrossRef] [Green Version]
- Sheng, L.; Jiang, H.; Yan, H.; Li, X.; Lin, Y.; Ye, H.; Cheng, B. MGFD: The maize gene families database. Database 2016, 2016, baw004. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moore, R.C.; Purugganan, M.D. The early stages of duplicate gene evolution. Proc. Natl. Acad. Sci. USA 2003, 100, 15682–15687. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.G.; Cai, H.L.; Lu, M.; Wei, Q.Y.; Xu, L.J.; Bo, C.; Ma, Q.; Zhao, Y.; Cheng, B.J. A maize stress-responsive Di19 transcription factor, ZmDi19-1, confers enhanced tolerance to salt in transgenic Arabidopsis. Plant Cell Rep. 2019, 38, 1563–1578. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Wang, M.; Guo, Z.; Zhu, X.; Xia, Z. Identification of a 119-bp Promoter of the Maize Sulfite Oxidase Gene (ZmSO) That Confers High-Level Gene Expression and ABA or Drought Inducibility in Transgenic Plants. Int. J. Mol. Sci. 2019, 20, 3326. [Google Scholar] [CrossRef] [Green Version]
- Chaves, M.M.; Flexas, J.; Pinheiro, C. Photosynthesis under drought and salt stress: Regulation mechanisms from whole plant to cell. Ann. Bot. 2009, 103, 551–560. [Google Scholar] [CrossRef] [Green Version]
- Dalal, V.K. Modulation of photosynthesis and other proteins during water-stress. Mol. Biol. Rep. 2021, 48, 3681–3693. [Google Scholar] [CrossRef]
- Ma, H.; Li, P.; Liu, X.; Li, C.; Zhang, S.; Wang, X.; Tao, X. Poly-γ-glutamic acid enhanced the drought resistance of maize by improving photosynthesis and affecting the rhizosphere microbial community. BMC Plant Biol. 2022, 22, 11. [Google Scholar] [CrossRef]
- Bose, J.; Xie, Y.; Shen, W.; Shabala, S. Haem oxygenase modifies salinity tolerance in Arabidopsis by controlling K+ retention via regulation of the plasma membrane H+-ATPase and by altering SOS1 transcript levels in roots. J. Exp. Bot. 2013, 64, 471–481. [Google Scholar] [CrossRef]
- Hu, H.; Dai, M.; Yao, J.; Xiao, B.; Li, X.; Zhang, Q.; Xiong, L. Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc. Natl. Acad. Sci. USA 2006, 103, 12987–12992. [Google Scholar] [CrossRef] [Green Version]
- Liang, W.H.; Li, L.; Zhang, F.; Liu, Y.X.; Li, M.M.; Shi, H.H.; Li, H.; Shang, F.; Chen, L.; Lin, Q.T.; et al. Effects of abiotic stress, light, phytochromes and phytohormones on the expression of OsAQP, a rice aquaporin gene. Plant Growth Regul. 2013, 69, 21–27. [Google Scholar] [CrossRef]
- Jung, C.G.; Hwang, S.G.; Park, Y.C.; Park, H.M.; Kim, D.S.; Park, D.H.; Jang, C.S. Molecular characterization of the cold- and heat-induced Arabidopsis PXL1 gene and its potential role in transduction pathways under temperature fluctuations. J. Plant Physiol. 2015, 176, 138–146. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Xu, L.; Shang, J.; Hu, X.; Yu, H.; Wu, H.; Lv, W.; Zhao, Y. Genome-wide analysis of the maize superoxide dismutase (SOD) gene family reveals important roles in drought and salt responses. Genet. Mol. Biol. 2021, 44, e20210035. [Google Scholar] [CrossRef] [PubMed]
- Si, W.; Hang, T.; Guo, M.; Chen, Z.; Liang, Q.; Gu, L.; Ding, T. Whole-Genome and Transposed Duplication Contributes to the Expansion and Diversification of TLC Genes in Maize. Int. J. Mol. Sci. 2019, 20, 5484. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Tian, F.; Yang, D.C.; Meng, Y.Q.; Jin, J.; Gao, G. PlantRegMap: Charting functional regulatory maps in plants. Nucleic Acids Res. 2020, 48, D1104–D1113. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. clusterProfiler: An R package for comparing biological themes among gene clusters. Omics A J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, Y.; Xu, L.; Huang, Y.; Wu, H.; Zhang, X.; Hu, X.; Ma, Q. Identification and Characterization of the Core Region of ZmDi19-5 Promoter Activity and Its Upstream Regulatory Proteins. Int. J. Mol. Sci. 2022, 23, 7390. https://doi.org/10.3390/ijms23137390
Zhao Y, Xu L, Huang Y, Wu H, Zhang X, Hu X, Ma Q. Identification and Characterization of the Core Region of ZmDi19-5 Promoter Activity and Its Upstream Regulatory Proteins. International Journal of Molecular Sciences. 2022; 23(13):7390. https://doi.org/10.3390/ijms23137390
Chicago/Turabian StyleZhao, Yang, Lijuan Xu, Yuanxiang Huang, Hongying Wu, Xingen Zhang, Xiaolin Hu, and Qing Ma. 2022. "Identification and Characterization of the Core Region of ZmDi19-5 Promoter Activity and Its Upstream Regulatory Proteins" International Journal of Molecular Sciences 23, no. 13: 7390. https://doi.org/10.3390/ijms23137390
APA StyleZhao, Y., Xu, L., Huang, Y., Wu, H., Zhang, X., Hu, X., & Ma, Q. (2022). Identification and Characterization of the Core Region of ZmDi19-5 Promoter Activity and Its Upstream Regulatory Proteins. International Journal of Molecular Sciences, 23(13), 7390. https://doi.org/10.3390/ijms23137390





