Conjunctive Analyses of BSA-Seq and BSR-Seq Unveil the Msβ-GAL and MsJMT as Key Candidate Genes for Cytoplasmic Male Sterility in Alfalfa (Medicago sativa L.)
Abstract
:1. Introduction
2. Results
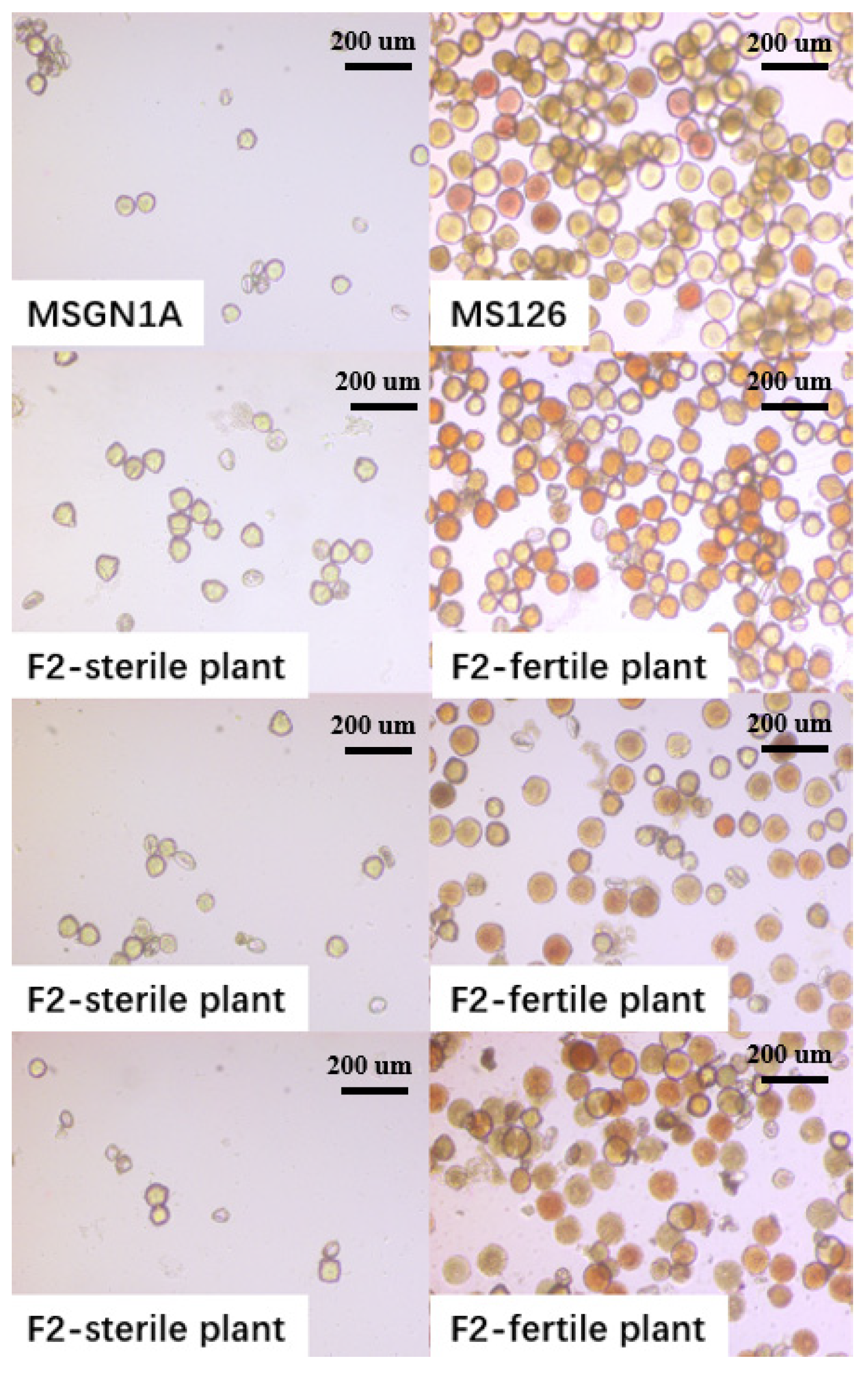
2.1. Sterility Survey of Parental Line and F2 Segregation Population
2.2. Bulked-Segregant Analysis
2.2.1. Sequencing Data Analysis of Sterile and Fertile Bulks
2.2.2. Association Analysis of BSA-Seq Data
2.3. Bulked Segregant RNA-Seq Analysis
2.3.1. Sequencing Data Analysis of Four Bulks
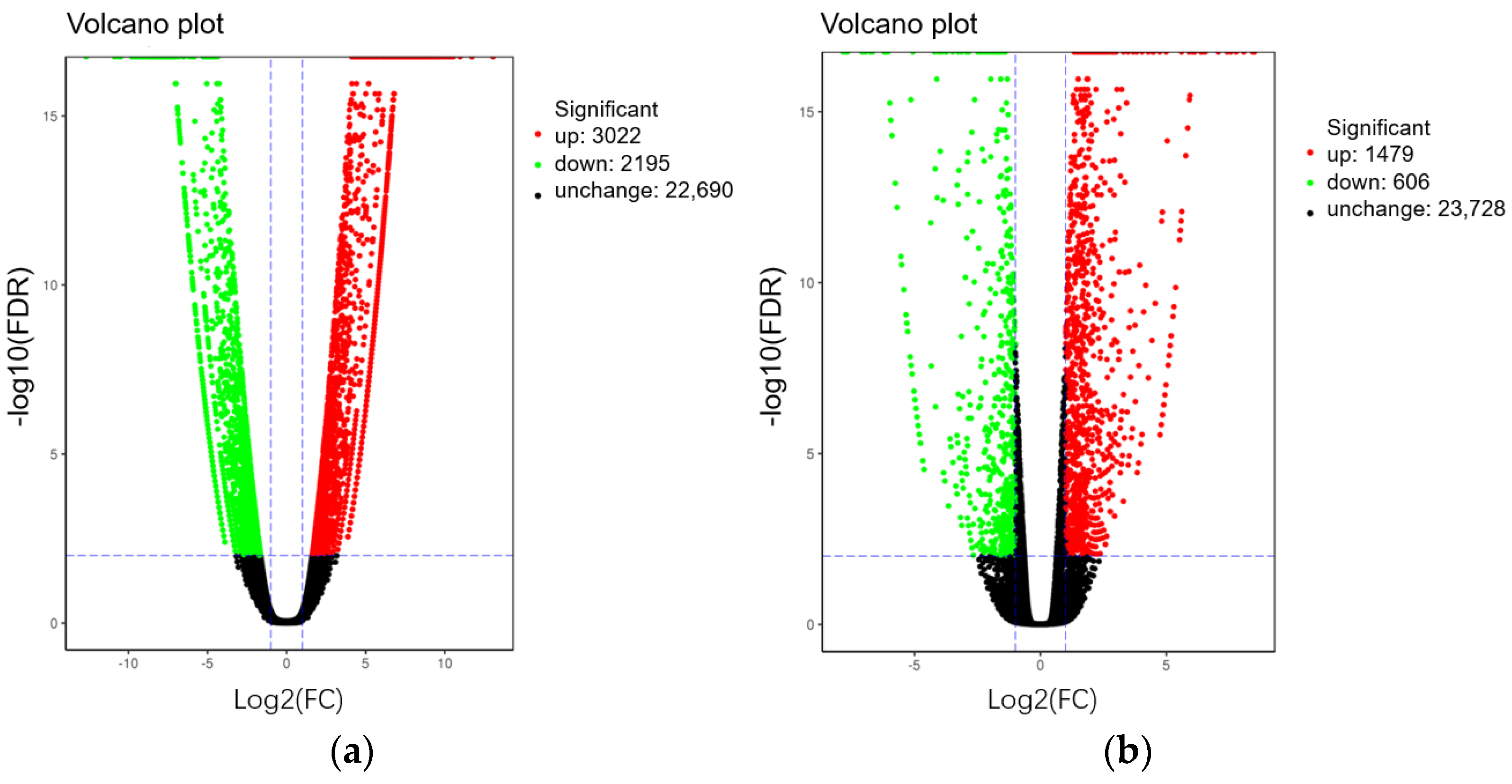
2.3.2. Comparison of Gene Expression Pattern between Two Set Pairs of Sequencing Bulks
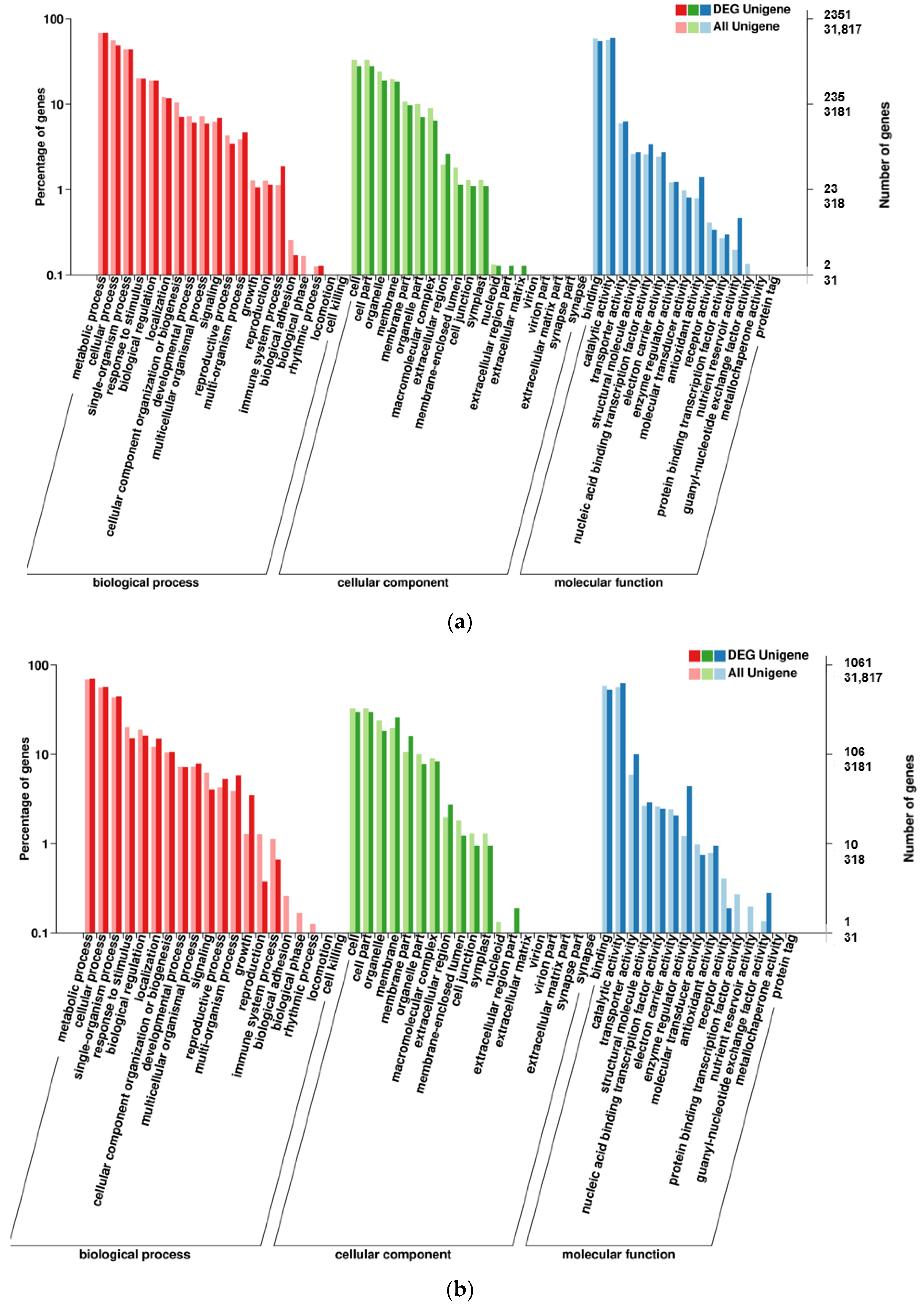
2.3.3. Association Analysis of BSR-Seq Data
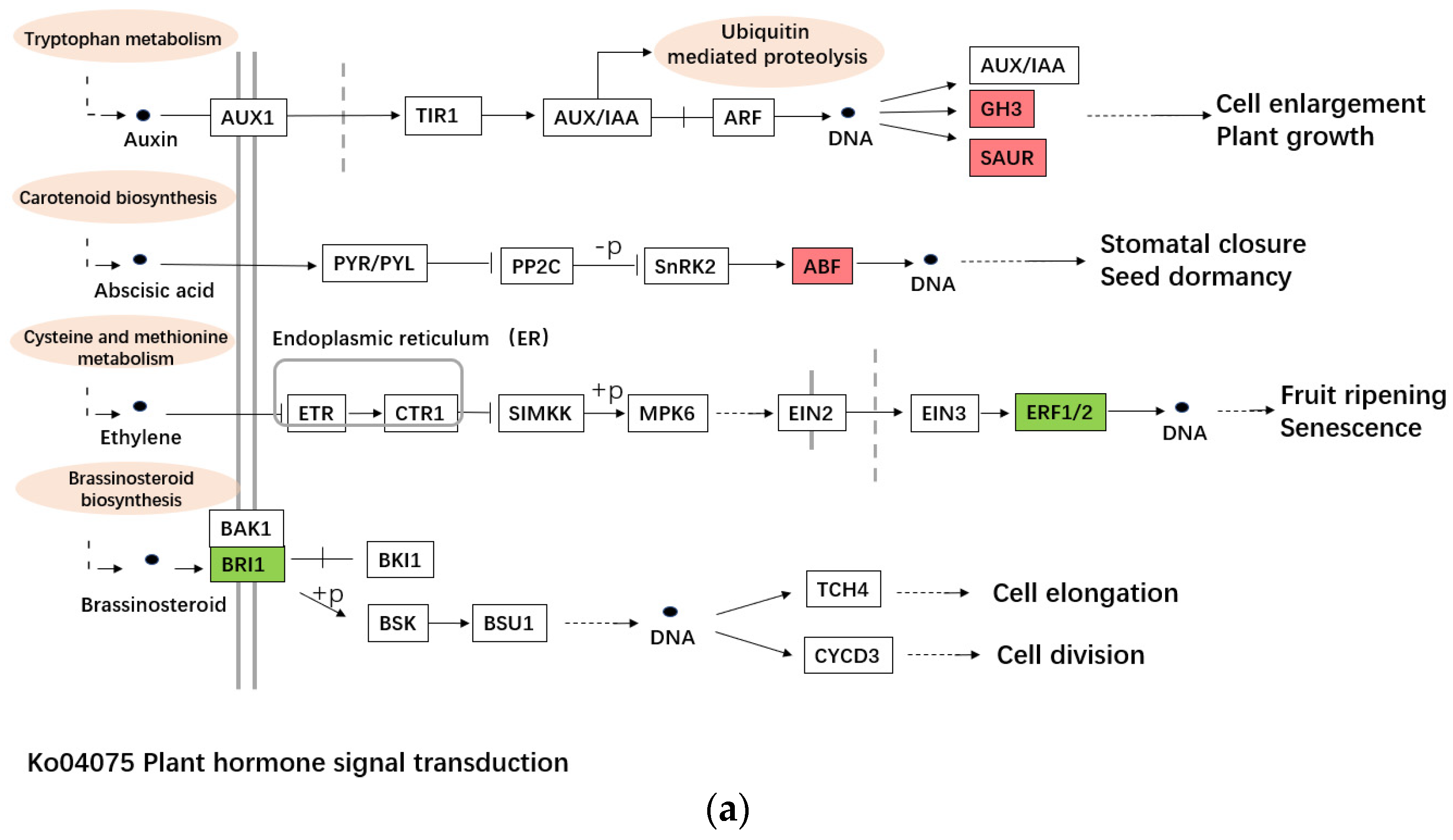
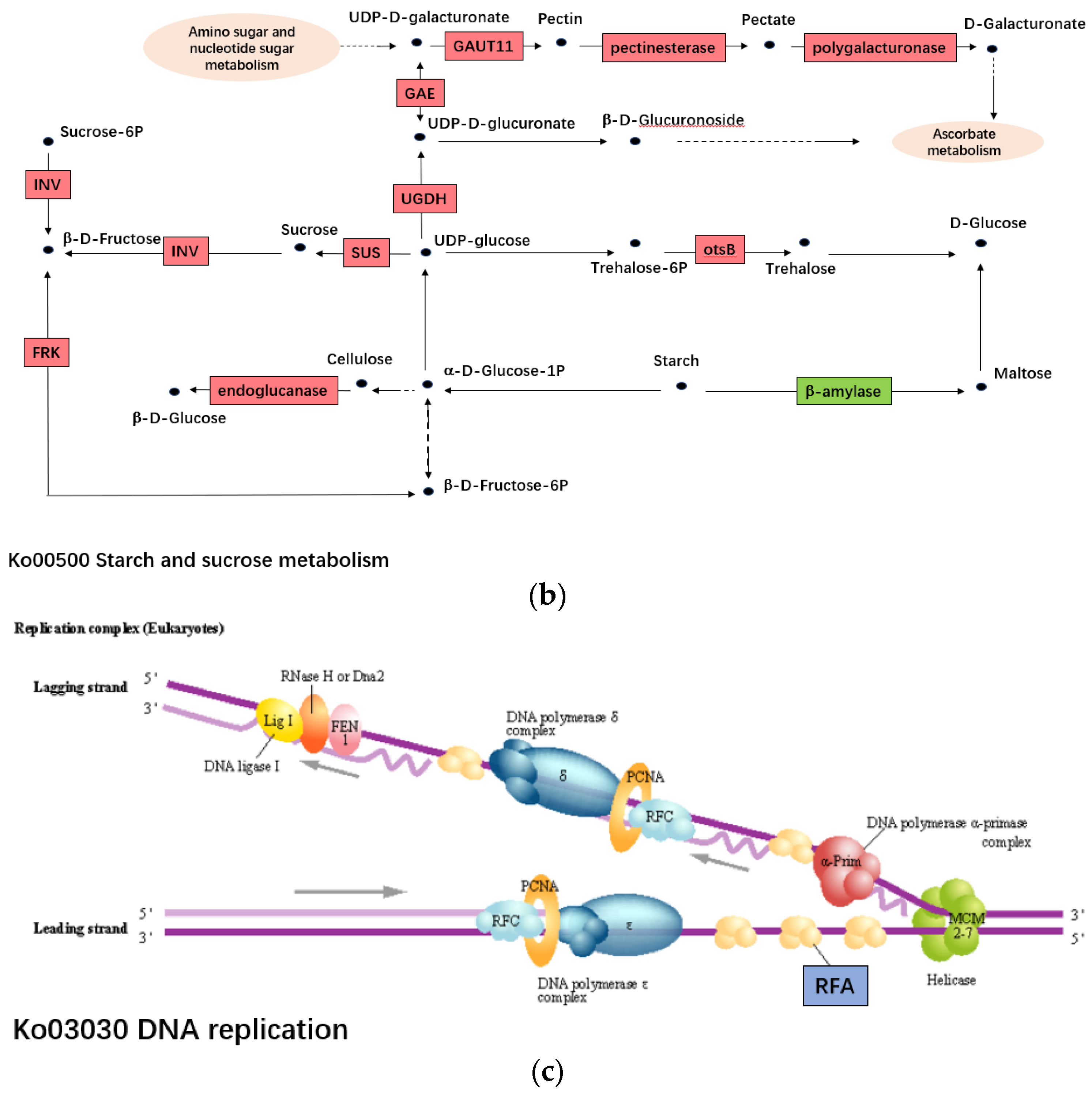
2.4. The DEGs Involved in Cytoplasmic Male Sterility
2.5. The Candidate Genes Involved in Cytoplasmic Male Sterility
2.6. Validation of Quantitative Real Time PCR for Key Genes Associated with Male Sterility in Alfalfa
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Fertility Identification of Alfalfa
4.3. Construction of Illumina Library of BSA-Seq
4.4. Analysis of BSA-Seq Data
4.5. cDNA Library Preparation of BSR-Seq
4.6. Analysis of BSR-Seq Data
4.7. qRT-PCR Validation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Qiu, Y.; Liao, L.; Jin, X.; Mao, D.; Liu, R. Analysis of the meiotic transcriptome reveals the genes related to the regulation of pollen abortion in cytoplasmic male-sterile pepper (Capsicum annuum L.). Gene 2018, 641, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Schnable, P.S.; Wise, R.P. The molecular basis of cytoplasmic male sterility and fertility restoration. Trends Plant Sci. 1998, 3, 175–180. [Google Scholar] [CrossRef]
- Wang, S.; Wang, C.; Zhang, X.-X.; Chen, X.; Liu, J.-J.; Jia, X.-F.; Jia, S.-Q. Transcriptome de novo assembly and analysis of differentially expressed genes related to cytoplasmic male sterility in cabbage. Plant Physiol. Biochem. 2016, 105, 224–232. [Google Scholar] [CrossRef] [PubMed]
- Toriyama, K. Molecular basis of cytoplasmic male sterility and fertility restoration in rice. Plant Biotechnol. 2021, 38, 285–295. [Google Scholar] [CrossRef]
- Zefu, L.; Jiafa, X.; Guangyong, T. Types and genetic mechanisms of plant male sterility. J. Anhui Agric. Sci. 2000, 28, 742–746. [Google Scholar]
- Liu, L.; Guo, W.; Zhu, X.; Zhang, T. Inheritance and fine mapping of fertility restoration for cytoplasmic male sterility in Gossypium hirsutum L. Theor. Appl. Genet. 2003, 106, 461–469. [Google Scholar] [CrossRef]
- Itabashi, E.; Iwata, N.; Fujii, S.; Kazama, T.; Toriyama, K. The fertility restorer gene, Rf2, for Lead Rice-type cytoplasmic male sterility of rice encodes a mitochondrial glycine-rich protein. Plant J. 2010, 65, 359–367. [Google Scholar] [CrossRef]
- Bach, I.C.; Olesen, A.; Simon, P. PCR-based markers to differentiate the mitochondrial genomes of petaloid and male fertile carrot (Daucus carota L.). Euphytica 2002, 127, 353–365. [Google Scholar] [CrossRef]
- Wan, X.; Wu, S.; Li, Z.-W.; Dong, Z.; An, X.; Ma, B.; Tian, Y.; Li, J. Maize Genic Male-Sterility Genes and Their Applications in Hybrid Breeding: Progress and Perspectives. Mol. Plant 2019, 12, 321–342. [Google Scholar] [CrossRef] [Green Version]
- Yuan, Q.; Song, C.; Gao, L.; Zhang, H.; Yang, C.; Sheng, J.; Ren, J.; Chen, D.; Wang, Y. Transcriptome de novo assembly and analysis of differentially expressed genes related to cytoplasmic male sterility in onion. Plant Physiol. Biochem. 2018, 125, 35–44. [Google Scholar] [CrossRef]
- Zhang, X.; Chen, H.; Zhang, Q.; Zhang, Y.; Xiao, Z.; Guo, Y.; Yu, F.; Hu, S. Cytological and genetic characterisation of dominant GMS line Shaan-GMS in Brassica napus L. J. Appl. Genet. 2020, 61, 477–488. [Google Scholar] [CrossRef] [PubMed]
- Pallotta, M.A.; Warner, P.; Kouidri, A.; Tucker, E.J.; Baes, M.; Suchecki, R.; Watson-Haigh, N.; Okada, T.; Garcia, M.; Sandhu, A.; et al. Wheat ms5 male-sterility is induced by recessive homoeologous A and D genome non-specific lipid transfer proteins. Plant J. 2019, 99, 673–685. [Google Scholar] [CrossRef] [PubMed]
- Rieseberg, L.H.; Van Fossen, C.; Arias, D.; Carter, R.L. Cytoplasmic male sterility in sunflower: Origin, inheritance, and frequency in natural populations. J. Hered. 1994, 85, 233–238. [Google Scholar] [CrossRef]
- Bauchan, G. Alfalfa (Medicago sativa ssp. sativa (L.). Genet. Resour. Chromosome Eng. Crop Improv. 2009, 5, 11–39. [Google Scholar]
- Luo, Z. Analysis of the effect of reasonable close planting on respiration characteristics of alfalfa (Medicago sativa L.) artificial grassland. Turk. J. Agric. For. 2021, 45, 533–540. [Google Scholar] [CrossRef]
- Childers, W.R.; McLennan, H.A. Mclennan, Inheritance studies of a completely male sterile character in Medicago sativa L. Genome 1960, 2, 57–66. [Google Scholar]
- Novák, F.; Betlach, J. Development and karyology of the tapetal layer of anthers in sweet pepper (Capsicum annuum L.). Biol. Plant. 1970, 12, 275–280. [Google Scholar] [CrossRef]
- Ruan, M.; Wan, H.; Yang, Y.; Zhou, G.; Wang, R.; Ye, Q.; Yao, Z.; Yang, Y.; Yuna, C.; Li, Z. Invertase activity and related genes expression analusis in cytoplasmic male sterile and fertilr pepper (Capsicum annuum) Lines. J. Agric. Biotechnol. 2018, 26, 2036–2046. [Google Scholar]
- Li, C. Cloning and Functional Study of a Gene Related to S-Type Cytoplasmic Male Sterility in Onin (Allium cepa L.). Master’s Thesis, Shandong Agricultural University, Tai’an, China, 2018. [Google Scholar]
- Heng, S.; Gao, J.; Wei, C.; Chen, F.; Li, X.; Wen, J.; Yi, B.; Ma, C.; Tu, J.; Fu, T.; et al. Transcript levels of orf288 are associated with the hau cytoplasmic male sterility system and altered nuclear gene expression in Brassica juncea. J. Exp. Bot. 2018, 69, 455–466. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Tang, F.; Gao, C.; Gao, X.; Xu, B.; Shi, F. Comparative transcriptome between male fertile and male sterile alfalfa (Medicago varia). Physiol. Mol. Biol. Plants 2021, 27, 1487–1498. [Google Scholar] [CrossRef]
- Guo, J.-X.; Liu, Y.-G. Molecular Control of Male Reproductive Development and Pollen Fertility in Rice. J. Integr. Plant Biol. 2012, 54, 967–978. [Google Scholar] [CrossRef] [PubMed]
- Saumitou-Laprade, P.; Cuguen, J.; Vernet, P. Cytoplasmic male sterility in plants: Molecular evidence and the nucleocytoplasmic conflict. Trends Ecol. Evol. 1994, 9, 431–435. [Google Scholar] [CrossRef]
- Chen, Z.; Zhao, N.; Li, S.; Grover, C.E.; Nie, H.; Wendel, J.F.; Hua, J. Plant Mitochondrial Genome Evolution and Cytoplasmic Male Sterility. Crit. Rev. Plant Sci. 2017, 36, 55–69. [Google Scholar] [CrossRef]
- Ludwig-Müller, J.; Jülke, S.; Bierfreund, N.M.; Decker, E.L.; Reski, R. Moss (Physcomitrella patens) GH3 proteins act in auxin homeostasis. New Phytol. 2009, 181, 323–338. [Google Scholar] [CrossRef]
- Gee, M.A.; Hagen, G.; Guilfoyle, T.J. Tissue-Specific and Organ-Specific Expression of Soybean Auxin-Responsive Transcripts GH3 and SAURs. Plant Cell 1991, 3, 419. [Google Scholar]
- Jakoby, M.; Weisshaar, B.; Dröge-Laser, W.; Vicente-Carbajosa, J.; Tiedemann, J.; Kroj, T.; Parcy, F. bZIP transcription factors in Arabidopsis. Trends Plant Sci. 2002, 7, 106–111. [Google Scholar] [CrossRef]
- Clouse, S.D.; Langford, M.; McMorris, T.C. A Brassinosteroid-Insensitive Mutant in Arabidopsis thaliana Exhibits Multiple Defects in Growth and Development. Plant Physiol. 1996, 111, 671–678. [Google Scholar] [CrossRef] [Green Version]
- Holton, N.; Caño-Delgado, A.; Harrison, K.; Montoya, T.; Chory, J.; Bishop, G.J. Tomato BRASSINOSTEROID INSENSITIVE1 is required for systemin-induced root elongation in Solanum pimpinellifolium but is not essential for wound signaling. Plant Cell 2007, 19, 1709–1717. [Google Scholar] [CrossRef] [Green Version]
- Sharma, M.K.; Kumar, R.; Solanke, A.U.; Sharma, R.; Tyagi, A.K.; Sharma, A.K. Identification, phylogeny, and transcript profiling of ERF family genes during development and abiotic stress treatments in tomato. Mol. Genet. Genom. 2010, 284, 455–475. [Google Scholar] [CrossRef]
- Fujimoto, S.Y.; Ohta, M.; Usui, A.; Shinshi, H.; Ohme-Takagi, M. Arabidopsis ethylene-responsive element binding factors act as transcriptional activators or repressors of GCC box-mediated gene expression. Plant Cell 2000, 12, 393–404. [Google Scholar]
- Caffall, K.H.; Pattathil, S.; Phillips, S.E.; Hahn, M.G.; Mohnen, D. Arabidopsis thaliana T-DNA Mutants Implicate GAUT Genes in the Biosynthesis of Pectin and Xylan in Cell Walls and Seed Testa. Mol. Plant 2009, 2, 1000–1014. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arsovski, A.A.; Haughn, G.W.; Western, T.L. Seed coat mucilage cells of Arabidopsis thaliana as a model for plant cell wall research. Plant Signal. Behav. 2010, 5, 796–801. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; Wang, W.; Wang, Y.-Q.; Liu, Y.-Y.; Wang, J.-X.; Zhang, X.-Q.; Ye, D.; Chen, L.-Q. Arabidopsis Galacturonosyltransferase (GAUT) 13 and GAUT14 Have Redundant Functions in Pollen Tube Growth. Mol. Plant 2013, 6, 1131–1148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vu, D.P.; Rodrigues, C.M.; Jung, B.; Meissner, G.; Klemens, P.A.W.; Holtgräwe, D.; Fürtauer, L.; Nägele, T.; Nieberl, P.; Pommerrenig, B.; et al. Vacuolar sucrose homeostasis is critical for plant development, seed properties, and night-time survival in Arabidopsis. J. Exp. Bot. 2020, 71, 4930–4943. [Google Scholar] [CrossRef]
- Broderick, S.; Rehmet, K.; Concannon, C.; Nasheuer, H.-P. Eukaryotic Single-Stranded DNA Binding Proteins: Central Factors in Genome Stability. Subcell. Biochem. 2010, 50, 143–163. [Google Scholar]
- Bhadula, S.K.; Elthon, T.E.; Habben, J.E.; Helentjaris, T.G.; Jiao, S.; Ristic, Z. Heat-stress induced synthesis of chloroplast protein synthesis elongation factor (EF-Tu) in a heat-tolerant maize line. Planta 2001, 212, 359–366. [Google Scholar] [CrossRef]
- Furukawa, T.; Inagaki, H.; Takai, R.; Hirai, H.; Che, F.-S. Two Distinct EF-Tu Epitopes Induce Immune Responses in Rice and Arabidopsis. Mol. Plant-Microbe Interact. 2014, 27, 113–124. [Google Scholar] [CrossRef] [Green Version]
- Min, S.S.; Dong, G.K.; Sun, H.L. Isolation and characterization of a jasmonic acid carboxyl methyltransferase gene from hot pepper (Capsicum annuum L.). J. Plant Biol. 2005, 48, 292–297. [Google Scholar]
- Suzuki, H.; Rodriguez-Uribe, L.; Xu, J.; Zhang, J. Transcriptome analysis of cytoplasmic male sterility and restoration in CMS-D8 cotton. Plant Cell Rep. 2013, 32, 1531–1542. [Google Scholar] [CrossRef]
- Qu, C.; Fu, F.; Liu, M.; Zhao, H.; Liu, C.; Li, J.; Tang, Z.; Xu, X.; Qiu, X.; Wang, R.; et al. Comparative Transcriptome Analysis of Recessive Male Sterility (RGMS) in Sterile and Fertile Brassica napus Lines. PLoS ONE 2015, 10, e0144118. [Google Scholar] [CrossRef]
- Chantret, N.; Sourdille, P.; Röder, M.; Tavaud, M.; Bernard, M.; Doussinault, G. Location and mapping of the powdery mildew resistance gene MIRE and detection of a resistance QTL by bulked segregant analysis (BSA) with microsatellites in wheat. Theor. Appl. Genet. 2000, 100, 1217–1224. [Google Scholar] [CrossRef]
- Voicescu, M.; Ionescu, S.; Calderon-Moreno, J.M.; Nistor, C.L. Physicochemical characterization of 3,6-diHydroxyflavone binding BSA immobilized on PEG-coated silver nanoparticles. J. Nanopart. Res. 2017, 19, 38. [Google Scholar] [CrossRef]
- Livaja, M.; Wang, Y.; Wieckhorst, S.; Haseneyer, G.; Seidel, M.; Hahn, V.; Knapp, S.J.; Taudien, S.; Schön, C.-C.; Bauer, E. BSTA: A targeted approach combines bulked segregant analysis with next- generation sequencing and de novo transcriptome assembly for SNP discovery in sunflower. BMC Genom. 2013, 14, 628. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.; Yeh, C.-T.; Tang, H.M.; Nettleton, D.; Schnable, P.S. Gene Mapping via Bulked Segregant RNA-Seq (BSR-Seq). PLoS ONE 2012, 7, e36406. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xue, Z.; Li, Y.; Zhang, L.; Shi, W.; Zhang, C.; Feng, M.; Zhang, F.; Tang, D.; Yu, H.; Gu, M.; et al. OsMTOPVIB Promotes Meiotic DNA Double-Strand Break Formation in Rice. Mol. Plant 2016, 9, 1535–1538. [Google Scholar] [CrossRef] [PubMed]
- Jachimowicz, R.D.; Goergens, J.; Reinhardt, H.C. DNA double-strand break repair pathway choice—From basic biology to clinical exploitation. Cell Cycle 2019, 18, 1423–1434. [Google Scholar] [CrossRef]
- Chen, W.; Gong, P.; Guo, J.; Li, H.; Li, R.; Xing, W.; Yang, Z.; Guan, Y. Glycolysis regulates pollen tube polarity via Rho GTPase signaling. PLoS Genet. 2018, 14, e1007373. [Google Scholar] [CrossRef] [Green Version]
- Xu, F.-Q.; Ruan, Y.-L. Preliminary function analysis of OsHXK family in rice and Inhibition of OsHXK10 expression by RNAi technology lead to abnormal pollination and seed abortion in rice. In Proceedings of the 2007 National Conference of Chinese Plant Physiology Society, Shijiazhuang, China,, 1 August 2007. [Google Scholar]
- Li, J.; Baroja-Fernández, E.; Bahaji, A.; Muñoz, F.J.; Ovecka, M.; Montero, M.; Sesma, M.T.; Alonso-Casajús, N.; Almagro, G.; Sánchez-López, A.M.; et al. Enhancing Sucrose Synthase Activity Results in Increased Levels of Starch and ADP-Glucose in Maize (Zea mays L.) Seed Endosperms. Plant Cell Physiol. 2013, 54, 282–294. [Google Scholar] [CrossRef] [Green Version]
- Angeles-Núñez, J.G.; Tiessen, A. Arabidopsis sucrose synthase 2 and 3 modulate metabolic homeostasis and direct carbon towards starch synthesis in developing seeds. Planta 2010, 232, 701–718. [Google Scholar] [CrossRef]
- Chen, J.-J. Study on the Molecular Mechanism of Cytoplasmic Male Sterility in Alfalfa. Master’s Thesis, Jilin Agricultural University, Changchun, China, 2019. [Google Scholar]
- Pego, J.V.; Smeekens, S.C. Plant fructokinases: A sweet family get-together. Trends Plant Sci. 2000, 5, 531–536. [Google Scholar] [CrossRef]
- Wasternack, C.; Song, S. Jasmonates: Biosynthesis, metabolism, and signaling by proteins activating and repressing transcription. J. Exp. Bot. 2017, 68, 1303–1321. [Google Scholar] [CrossRef] [PubMed]
- Wasternack, C.; Hause, B. Jasmonates: Biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann. Bot. 2013, 111, 1021–1058. [Google Scholar] [CrossRef] [PubMed]
- McConn, M.; Browse, J. The Critical Requirement for Linolenic Acid Is Pollen Development, Not Photosynthesis, in an Arabidopsis Mutant. Plant Cell 1996, 8, 403–416. [Google Scholar] [CrossRef] [PubMed]
- Seo, H.S.; Song, J.T.; Cheong, J.-J.; Lee, Y.-H.; Lee, Y.-W.; Hwang, I.; Lee, J.S.; Choi, Y.D. Jasmonic acid carboxyl methyltransferase: A key enzyme for jasmonate-regulated plant responses. Proc. Natl. Acad. Sci. USA 2001, 98, 4788–4793. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sohn, H.B.; Lee, H.Y.; Seo, J.S.; Jung, C.; Jeon, J.H.; Kim, J.-H.; Lee, Y.W.; Lee, J.S.; Cheong, J.-J.; Choi, Y.D. Overexpression of jasmonic acid carboxyl methyltransferase increases tuber yield and size in transgenic potato. Plant Biotechnol. Rep. 2011, 5, 27–34. [Google Scholar] [CrossRef]
- Kim, E.H.; Kim, Y.S.; Park, S.-H.; Koo, Y.J.; Choi, Y.D.; Chung, Y.-Y.; Lee, I.-J.; Kim, J.-K. Methyl Jasmonate Reduces Grain Yield by Mediating Stress Signals to Alter Spikelet Development in Rice. Plant Physiol. 2009, 149, 1751–1760. [Google Scholar] [CrossRef] [Green Version]
- Qi, J.; Li, J.; Han, X.; Li, R.; Wu, J.; Yu, H.; Hu, L.; Xiao, Y.; Lu, J.; Lou, Y. Jasmonic acid carboxyl methyltransferase regulates development and herbivory-induced defense response in rice. J. Integr. Plant Biol. 2016, 58, 564–576. [Google Scholar] [CrossRef]
- Stitz, M.; Hartl, M.; Baldwin, I.T.; Gaquerel, E. Jasmonoyl-l-Isoleucine Coordinates Metabolic Networks Required for Anthesis and Floral Attractant Emission in Wild Tobacco (Nicotiana attenuata). Plant Cell 2014, 26, 3964–3983. [Google Scholar] [CrossRef] [Green Version]
- Zieliński, K.; Dubas, E.; Gerši, Z.; Krzewska, M.; Janas, A.; Nowicka, A.; Matušíková, I.; Żur, I.; Sakuda, S.; Moravčíková, J. β-1,3-Glucanases and chitinases participate in the stress-related defence mechanisms that are possibly connected with modulation of arabinogalactan proteins (AGP) required for the androgenesis initiation in rye (Secale cereale L.). Plant Sci. 2021, 302, 110700. [Google Scholar] [CrossRef]
- Smith, D.L.; Starrett, D.A.; Gross, K.C. A gene coding for tomato fruit beta-galactosidase II is expressed during fruit ripening. Cloning, characterization, and expression pattern. Plant Physiol. 1998, 117, 417–423. [Google Scholar] [CrossRef] [Green Version]
- Zhang, B.; Zhou, Y. Plant cell wall formation and regulation. Sci. Sin. Vitae 2015, 45, 544–556. [Google Scholar] [CrossRef]
- Moneo-Sánchez, M.; Izquierdo, L.; Martín, I.; Labrador, E.; Dopico, B. Subcellular location of Arabidopsis thaliana subfamily a1 β-galactosidases and developmental regulation of transcript levels of their coding genes. Plant Physiol. Biochem. 2016, 109, 137–145. [Google Scholar] [CrossRef] [PubMed]
- Wei, B.; Wang, L.; Bosland, P.W.; Zhang, G.; Zhang, R. Comparative transcriptional analysis of Capsicum flower buds between a sterile flower pool and a restorer flower pool provides insight into the regulation of fertility restoration. BMC Genom. 2019, 20, 837. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, W.; Wang, L.; Chen, C.; Xiong, G.; Tan, X.-Y.; Yang, K.-Z.; Wang, Z.-C.; Zhou, Y.; Ye, D.; Chen, L.-Q. Arabidopsis CSLD1 and CSLD4 are required for cellulose deposition and normal growth of pollen tubes. J. Exp. Bot. 2011, 62, 5161–5177. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, D.; Yuan, S.; Yin, L.; Zhao, J.; Guo, B.; Lan, J.; Li, X. A missense mutation in the transmembrane domain of CESA9 affects cell wall biosynthesis and plant growth in rice. Plant Sci. 2012, 196, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Benstein, R.M.; Ludewig, K.; Wulfert, S.; Wittek, S.; Gigolashvili, T.; Frerigmann, H.; Gierth, M.; Flügge, U.-I.; Krueger, S. Arabidopsis Phosphoglycerate Dehydrogenase1 of the Phosphoserine Pathway Is Essential for Development and Required for Ammonium Assimilation and Tryptophan Biosynthesis. Plant Cell 2013, 25, 5011–5029. [Google Scholar] [CrossRef] [Green Version]
- Toujani, W.; Muñoz-Bertomeu, J.; Flores-Tornero, M.; Rosa-Téllez, S.; Anoman, A.D.; Alseekh, S.; Fernie, A.R.; Ros, R. Functional Characterization of the Plastidial 3-Phosphoglycerate Dehydrogenase Family in Arabidopsis. Plant Physiol. 2013, 163, 1164–1178. [Google Scholar] [CrossRef] [Green Version]
- Shao, Q.-S.; Wang, Y.; Hu, R.-H.; Zhang, W.-S.; Li, M.-Y. Pollen viability, stigma receptivity and fruiting characteristics of botanical origin of Jinxianlian. Zhongguo Zhong Yao Za Zhi 2015, 40, 1061–1065. [Google Scholar]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows—Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kai, W.; Mingyao, L.; Hakon, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, 36, e164. [Google Scholar]
- Takagi, H.; Abe, A.; Yoshida, K.; Kosugi, S.; Natsume, S.; Mitsuoka, C.; Uemura, A.; Utsushi, H.; Tamiru, M.; Takuno, S.; et al. QTL-seq: Rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J. 2013, 74, 174–183. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.D.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]



| Bulk | Clean Reads | Data Generated | Q30 (%) | Genome Coverage (%) | Average Depth (×) | Alignment Efficiency (%) |
|---|---|---|---|---|---|---|
| R03 (sterile) | 238,377,744 | 67,766,104,095 | 94.29 | 83.69 | 45.92 | 90.01 |
| R04 (fertile) | 271,300,098 | 77,009,673,959 | 93.82 | 84.53 | 52.12 | 91.03 |
| Method. | Chromosome | Start (Mb) | End (Mb) | Length (Mb) | Genes Number |
|---|---|---|---|---|---|
| SNP-index | Chromosome 5 | 8.54 | 17.02 | 8.47 | 626 |
| Bulk | Clean Reads | Data Generated (Gb) | Q30 (%) | Mapped Reads | Mapped Efficiency |
|---|---|---|---|---|---|
| T01 (male parent (♂)) | 24,944,663 | 7.43 | 94.14% | 17,230,648 | 85.05% |
| T02 (female parent (♀)) | 27,026,088 | 8.06 | 94.01% | 19,151,540 | 84.11% |
| T03 (sterile) | 39,503,495 | 11.76 | 93.76% | 27,499,773 | 85.60% |
| T04 (fertile) | 38,797,934 | 11.54 | 94.03% | 27,389,445 | 85.18% |
| Pathways | Transcript_ID | FPKM (T03 vs. T04) | Log2FC (T04/T03) | NR_Annotation | Gene Name | |
|---|---|---|---|---|---|---|
| T03 | T04 | |||||
| ko 04075 (Plant hormone signal transduction) | c101056.graph_c0 | 22.93 | 77.45 | 1.90 | SAUR-like auxin-responsive family protein (Medicago truncatula) | SAUR |
| c102778.graph_c0 | 0.05 | 1.21 | 3.84 | Indole-3-acetic acid-amido synthetase (Medicago truncatula) | GH3 | |
| c116678.graph_c0 | 16.29 | 39.07 | 1.35 | indole-3-acetic acid-amido synthetase (Medicago truncatula) | GH3 | |
| c90443.graph_c0 | 0.21 | 1.25 | 2.47 | BZIP transcription factor (Medicago truncatula) | ABF | |
| c108656.graph_c0 | 10.01 | 3.89 | −1.18 | brassinosteroid receptor(Pisum sativum) | BRI1 | |
| c102842.graph_c0 | 3.9 | 0.74 | −2.15 | ethylene response factor 5 (Medicago sativa) | ERF1/2 | |
| ko 00500 (Starch and sucrose metabolism) | c89515.graph_c0 | 87.9 | 326.14 | 2.03 | Pectinesterase (Medicago truncatula) | Pectinesterase |
| c116661.graph_c0 | 5.48 | 16.73 | 1.74 | polygalacturonase (Medicago truncatula) | polygalacturonase | |
| c118955.graph_c0 | 50.45 | 100.64 | 1.17 | UDP-glucuronate 4-epimerase (Medicago truncatula) | GAE | |
| c101072.graph_c0 | 0.2 | 1.03 | 2.25 | CAZy family GT8 glycosyltransferase (Medicago truncatula) | GAUT11 | |
| c100736.graph_c2 | 8.76 | 16.46 | 1.06 | UDP-glucose dehydrogenase (Medicago truncatula) | UGDH | |
| c102786.graph_c0 | 0.68 | 2.28 | 1.84 | Sucrose synthase (Medicago truncatula) | SUS | |
| c106305.graph_c0 | 18.08 | 57.22 | 1.80 | glycosyl hydrolase family 9 protein (Medicago truncatula) | Endoglucanase | |
| c107054.graph_c0 | 54.71 | 119.48 | 1.20 | beta-fructofuranosidase, insoluble protein (Medicago truncatula) | INV | |
| c111356.graph_c0 | 5.08 | 13.78 | 1.63 | Fructokinase (Medicago truncatula) | FRK | |
| c99556.graph_c0 | 3.14 | 12.18 | 2.09 | Trehalose-phosphate phosphatase (Medicago truncatula) | otsB | |
| c114245.graph_c0 | 29.07 | 10.13 | −1.38 | Beta-amylase (Medicago truncatula) | b-amylase | |
| ko 03030 (DNA replication) | c104110.graph_c0 | 0.84 | 2.3 | 1.79 | replication factor-A carboxy-terminal domain protein (Medicago truncatula) | RFA |
| c120364.graph_c1 | 2.83 | 5.28 | 1.14 | ATP-dependent DNA helicase PIF1 (Medicago truncatula) | RFA | |
| c84972.graph_c0 | 0.30 | 1.8 | 5.15 | Replication factor A protein (Medicago truncatula) | RFA | |
| c88743.graph_c0 | 0.20 | 2.41 | 6.54 | Replication factor A protein (Medicago truncatula) | RFA | |
| c96653.graph_c0 | 10.71 | 2.52 | −1.99 | Replication factor A protein (Medicago truncatula) | RFA | |
| c111708.graph_c1 | 1.91 | 0.55 | −1.61 | Replication factor A protein (Medicago truncatula) | RFA | |
| Transcript_ID | Regulation (T04/T03) | Gene Name | NR_Annotation |
|---|---|---|---|
| c123275.graph_c0 | down | Unknow | Unknow |
| c111485.graph_c0 | down | Unknow | Unknow |
| c102760.graph_c0 | up | EF-Tu | elongation factor Tu family protein (Medicago truncatula) |
| c115887.graph_c0 | down | JMT | Jasmonate O-methyltransferase (Medicago truncatula) |
| c102682.graph_c0 | up | β-GAL | Beta-galactosidase (Medicago truncatula) |
| c113540.graph_c0 | down | CESA | Cellulose synthase (Medicago truncatula) |
| c111062.graph_c0 | up | PHGDH | Phosphoglycerate dehydrogenase (Medicago truncatula) |
| Gene_Id | Chr | Start (Kb) | End (Kb) | NR_Annotation |
|---|---|---|---|---|
| MsG0580024998.01 (Transcript_id: c115887.graph_c0) | Chr5 | 12668765 | 12689067 | Jasmonate O-methyltransferase (Medicago truncatula) |
| MsG0580025125.01 | Chr5 | 14126518 | 14128116 | Beta-galactosidase (Medicago truncatula) |
| MsG0580025201.01 | Chr5 | 15181309 | 15201352 | Beta-galactosidase (Medicago truncatula) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, L.; Wang, Y.; Xu, X.; Yan, D.; Yu, W.; Miao, Y.; Xu, B. Conjunctive Analyses of BSA-Seq and BSR-Seq Unveil the Msβ-GAL and MsJMT as Key Candidate Genes for Cytoplasmic Male Sterility in Alfalfa (Medicago sativa L.). Int. J. Mol. Sci. 2022, 23, 7172. https://doi.org/10.3390/ijms23137172
Zhou L, Wang Y, Xu X, Yan D, Yu W, Miao Y, Xu B. Conjunctive Analyses of BSA-Seq and BSR-Seq Unveil the Msβ-GAL and MsJMT as Key Candidate Genes for Cytoplasmic Male Sterility in Alfalfa (Medicago sativa L.). International Journal of Molecular Sciences. 2022; 23(13):7172. https://doi.org/10.3390/ijms23137172
Chicago/Turabian StyleZhou, Le, Yingzhe Wang, Xiaobo Xu, Dong Yan, Weijie Yu, Yifan Miao, and Bo Xu. 2022. "Conjunctive Analyses of BSA-Seq and BSR-Seq Unveil the Msβ-GAL and MsJMT as Key Candidate Genes for Cytoplasmic Male Sterility in Alfalfa (Medicago sativa L.)" International Journal of Molecular Sciences 23, no. 13: 7172. https://doi.org/10.3390/ijms23137172
APA StyleZhou, L., Wang, Y., Xu, X., Yan, D., Yu, W., Miao, Y., & Xu, B. (2022). Conjunctive Analyses of BSA-Seq and BSR-Seq Unveil the Msβ-GAL and MsJMT as Key Candidate Genes for Cytoplasmic Male Sterility in Alfalfa (Medicago sativa L.). International Journal of Molecular Sciences, 23(13), 7172. https://doi.org/10.3390/ijms23137172






