Pituitary Tumor-Transforming Gene 1/Delta like Non-Canonical Notch Ligand 1 Signaling in Chronic Liver Diseases
Abstract
1. Introduction
1.1. PTTG1: Gene Structure and Regulation, Protein Synthesis, and Maturation and Tissue Expression
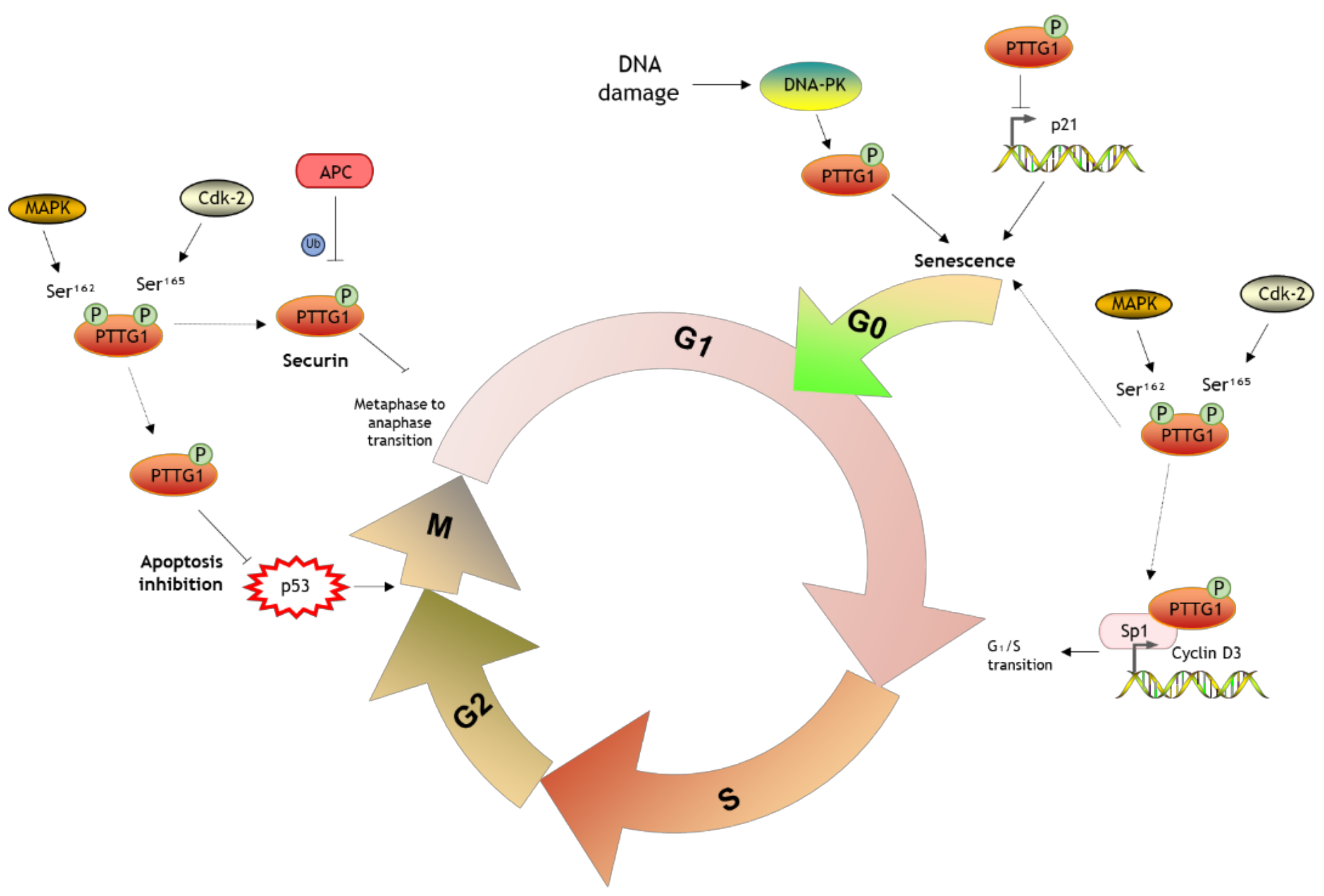
1.2. Biological Functions of PTTG1
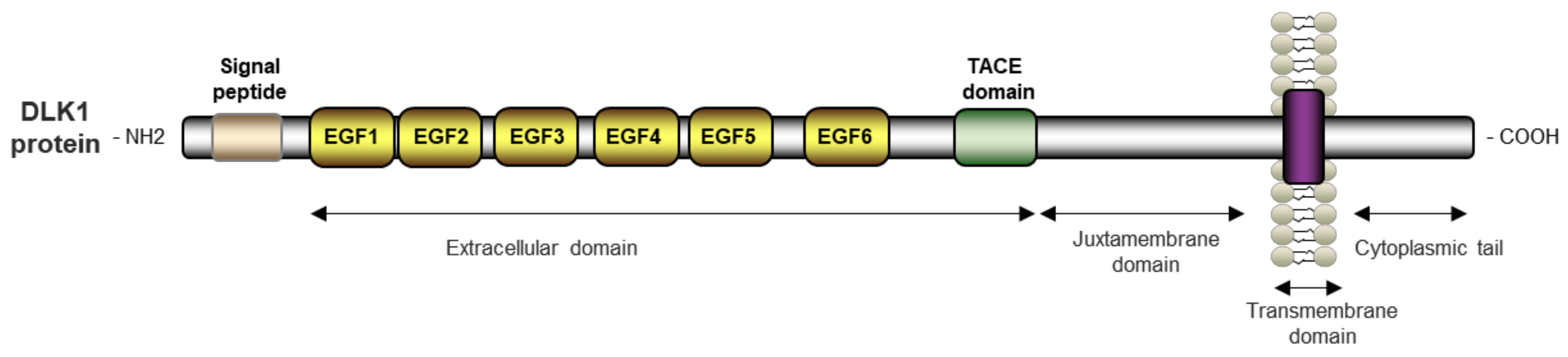
1.3. DLK1: Gene Organization and Regulation, Protein Structure, and Expression in Different Tissues
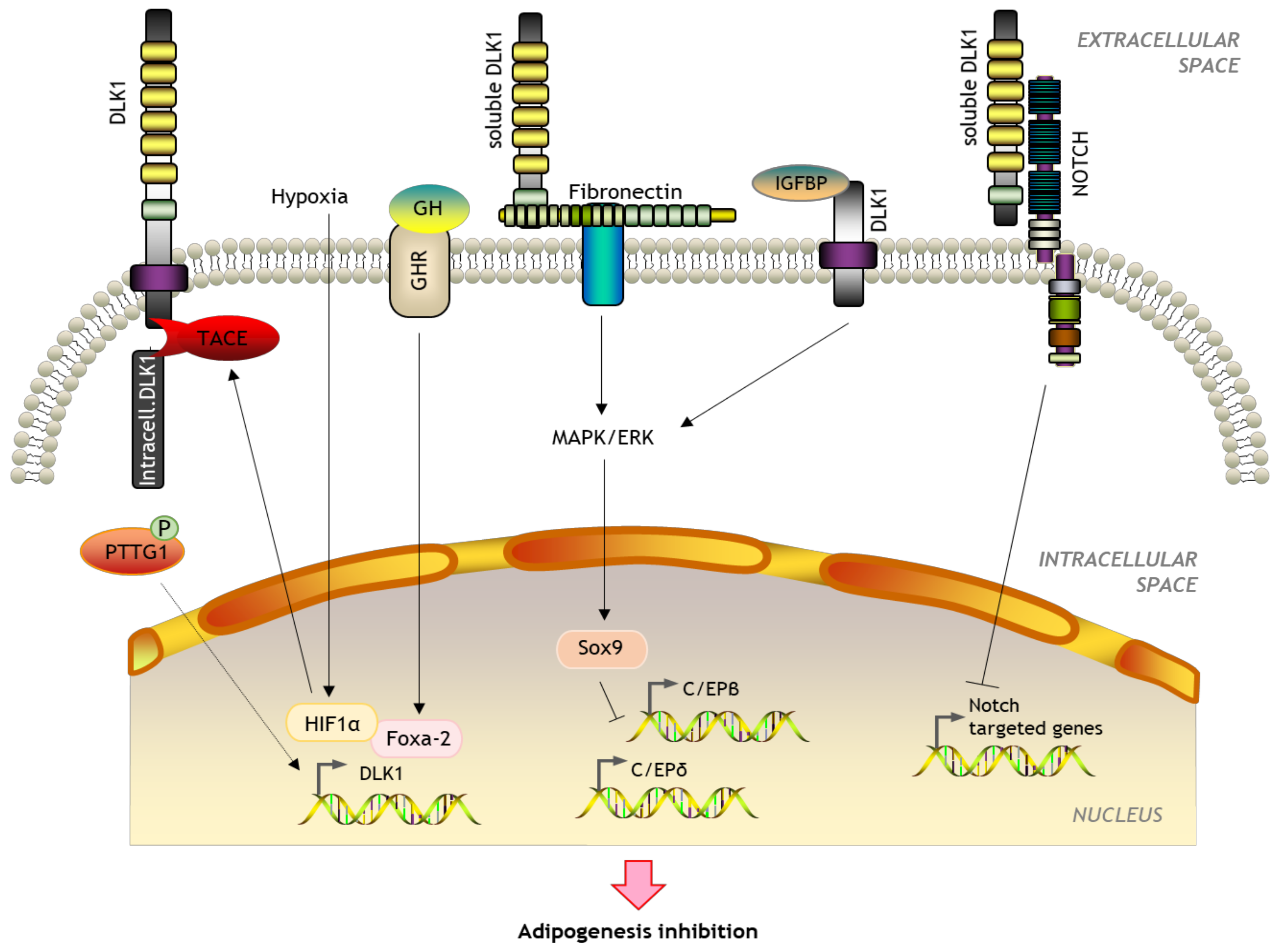
1.4. Biological Functions of DLK1
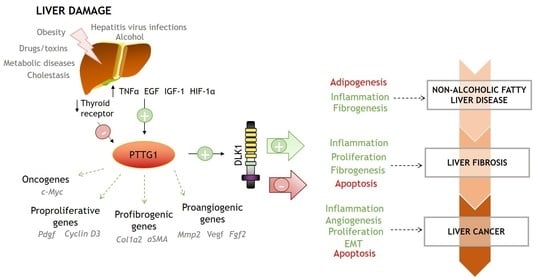
2. Involvement of PTTG1 and DLK1 in the Pathogenesis of Chronic Liver Diseases (CLDs)
2.1. Non-Alcoholic Fatty Liver Disease (NAFLD)
2.2. Fibrosis
2.3. Liver Cancer
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Pimpin, L.; Cortez-Pinto, H.; Negro, F.; Corbould, E.; Lazarus, J.V.; Webber, L.; Sheron, N.; EASL HEPAHEALTH Steering Committee. Burden of liver disease in Europe: Epidemiology and analysis of risk factors to identify prevention policies. J. Hepatol. 2018, 69, 718–735. [Google Scholar] [CrossRef] [PubMed]
- Schuppan, D.; Ashfaq-Khan, M.; Yang, A.T.; Kim, Y.O. Liver fibrosis: Direct antifibrotic agents and targeted therapies. Matrix Biol. 2018, 68, 435–451. [Google Scholar] [CrossRef] [PubMed]
- Pei, L.; Melmed, S. Isolation and characterization of a pituitary tumor-transforming gene (PTTG). Mol. Endocrinol. 1997, 11, 433–441. [Google Scholar] [CrossRef] [PubMed]
- Clem, A.L.; Hamid, T.; Kakar, S.S. Characterization of the role of Sp1 and NF-Y in differential regulation of PTTG/securin expression in tumor cells. Gene 2003, 322, 113–121. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Horwitz, G.A.; Prezant, T.R.; Valentini, A.; Nakashima, M.; Bronstein, M.D.; Melmed, S. Structure, expression, and function of human pituitary tumor-transforming gene (PTTG). Mol. Endocrinol. 1999, 13, 156–166. [Google Scholar] [CrossRef]
- Chen, L.; Puri, R.; Lefkowitz, E.J.; Kakar, S.S. Identification of the human pituitary tumor transforming gene (hPTTG) family: Molecular structure, expression, and chromosomal localization. Gene 2000, 248, 41–50. [Google Scholar] [CrossRef]
- Kakar, S.S. Molecular cloning, genomic organization, and identification of the promoter for the human pituitary tumor transforming gene (PTTG). Gene 1999, 240, 317–324. [Google Scholar] [CrossRef]
- Lee, U.E.; Ghiassi-Nejad, Z.; Paris, A.J.; Yea, S.; Narla, G.; Walsh, M.; Friedman, S.L. Tumor suppressor activity of KLF6 mediated by downregulation of the PTTG1 oncogene. FEBS Lett. 2010, 584, 1006–1010. [Google Scholar] [CrossRef]
- Chen, P.-Y.; Yen, J.-H.; Kao, R.-H.; Chen, J.-H. Down-regulation of the oncogene PTTG1 via the KLF6 tumor suppressor during induction of myeloid differentiation. PLoS ONE 2013, 8, e71282. [Google Scholar] [CrossRef]
- Vlotides, G.; Eigler, T.; Melmed, S. Pituitary tumor-transforming gene: Physiology and implications for tumorigenesis. Endocr. Rev. 2007, 28, 165–186. [Google Scholar] [CrossRef]
- Pei, L. Activation of mitogen-activated protein kinase cascade regulates pituitary tumor-transforming gene transactivation function. J. Biol. Chem. 2000, 275, 31191–31198. [Google Scholar] [CrossRef] [PubMed]
- Ramos-Morales, F.; Domínguez, A.; Romero, F.; Luna, R.; Multon, M.-C.; A Pintor-Toro, J.; Tortolero, M. Cell cycle regulated expression and phosphorylation of hpttg proto-oncogene product. Oncogene 2000, 19, 403–409. [Google Scholar] [CrossRef] [PubMed]
- Romero, F.; Multon, M.C.; Ramos-Morales, F.; Domínguez, Á.; Bernal, J.A.; Pintor-Toro, J.A.; Tortolero, M. Human securin, hPTTG, is associated with Ku heterodimer, the regulatory subunit of the DNA-dependent protein kinase. Nucleic Acids Res. 2001, 15, 1300–1307. [Google Scholar] [CrossRef] [PubMed]
- Espina, A.G.; Méndez-Vidal, C.; Moreno-Mateos, M.A.; Sáez, C.; Romero-Franco, A.; Japón, M.A.; Pintor-Toro, J.A. Induction of Dlk1 by PTTG1 inhibits adipocyte differentiation and correlates with malignant transformation. Mol. Biol. Cell 2009, 20, 3353–3362. [Google Scholar] [CrossRef]
- Panguluri, S.K.; Yeakel, C.; Kakar, S.S. PTTG: An important target gene for ovarian cancer therapy. J. Ovarian Res. 2008, 1, 6. [Google Scholar] [CrossRef]
- Wondergem, B.; Zhang, Z.; Huang, D.; Ong, C.K.; Koeman, J.; Hof, D.V.; Petillo, D.; Ooi, A.; Anema, J.; Lane, B.; et al. Expression of the PTTG1 oncogene is associated with aggressive clear cell renal cell carcinoma. Cancer Res. 2012, 72, 4361–4371. [Google Scholar] [CrossRef]
- Meng, C.; Zou, Y.; Hong, W.; Bao, C.; Jia, X. Estrogen-regulated PTTG1 promotes breast cancer progression by regulating cyclin kinase expression. Mol. Med. 2020, 26, 33. [Google Scholar] [CrossRef]
- Parte, S.; Virant-Klun, I.; Patankar, M.; Batra, S.K.; Straughn, A.; Kakar, S.S. PTTG1: A Unique Regulator of Stem/Cancer Stem Cells in the Ovary and Ovarian Cancer. Stem. Cell Rev. Rep. 2019, 15, 866–879. [Google Scholar] [CrossRef]
- Molina-Jiménez, F.; Benedicto, I.; Murata, M.; Martín-Vílchez, S.; Seki, T.; Antonio Pintor-Toro, J.; Tortolero, M.; Moreno-Otero, R.; Okazaki, K.; Koike, K.; et al. Expression of pituitary tumor-transforming gene 1 (PTTG1)/securin in hepatitis B virus (HBV)-associated liver diseases: Evidence for an HBV X protein-mediated inhibition of PTTG1 ubiquitination and degradation. Hepatology 2010, 51, 777–787. [Google Scholar] [CrossRef]
- Ren, Q.; Jin, B. The clinical value and biological function of PTTG1 in colorectal cancer. Biomed. Pharmacother. 2017, 89, 108–115. [Google Scholar] [CrossRef]
- Wang, Z.; Yu, R.; Melmed, S. Mice lacking pituitary tumor transforming gene show testicular and splenic hypoplasia, thymic hyperplasia, thrombocytopenia, aberrant cell cycle progression, and premature centromere division. Mol. Endocrinol. 2001, 15, 1870–1879. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Moro, E.; Kovacs, K.; Yu, R.; Melmed, S. Pituitary tumor transforming gene-null male mice exhibit impaired pancreatic beta cell proliferation and diabetes. Proc. Natl. Acad. Sci. USA 2003, 100, 3428–3432. [Google Scholar] [CrossRef] [PubMed]
- Yu, R.; Cruz-Soto, M.; Calzi, S.L.; Hui, H.; Melmed, S. Murine pituitary tumor-transforming gene functions as a securin protein in insulin-secreting cells. J. Endocrinol. 2006, 191, 45–53. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhao, Z.; Zhang, M. Pituitary tumor-transforming gene 1: Perspective on an apoptosis pathway. Cancer Biol. Ther. 2011, 11, 346–348. [Google Scholar] [CrossRef][Green Version]
- Vlotides, G.; Cruz-Soto, M.; Rubinek, T.; Eigler, T.; Auernhammer, C.J.; Melmed, S. Mechanisms for growth factor-induced pituitary tumor transforming gene-1 expression in pituitary folliculostellate TtT/GF cells. Mol. Endocrinol. 2006, 20, 3321–3335. [Google Scholar] [CrossRef]
- Chien, W.; Pei, L. A novel binding factor facilitates nuclear translocation and transcriptional activation function of the pituitary tumor-transforming gene product. J. Biol. Chem. 2000, 275, 19422–19427. [Google Scholar] [CrossRef]
- Thompson, A.D., 3rd; Kakar, S.S. Insulin and IGF-1 regulate the expression of the pituitary tumor transforming gene (PTTG) in breast tumor cells. FEBS Lett. 2005, 579, 3195–3200. [Google Scholar] [CrossRef][Green Version]
- Tong, Y.; Tan, Y.; Zhou, C.; Melmed, S. Pituitary tumor transforming gene interacts with Sp1 to modulate G1/S cell phase transition. Oncogene 2007, 26, 5596–5605. [Google Scholar] [CrossRef]
- Pei, L. Identification of c-myc as a down-stream target for pituitary tumor-transforming gene. J. Biol. Chem. 2001, 276, 8484–8491. [Google Scholar] [CrossRef]
- Duffy, M.J.; O’Grady, S.; Tang, M.; Crown, J. MYC as a target for cancer treatment. Cancer Treat. Rev. 2021, 1, 94. [Google Scholar] [CrossRef]
- Malik, M.T.; Kakar, S.S. Regulation of angiogenesis and invasion by human Pituitary tumor transforming gene (PTTG) through increased expression and secretion of matrix metalloproteinase-2 (MMP-2). Mol. Cancer 2006, 5, 61. [Google Scholar] [CrossRef] [PubMed]
- Ying, H.; Furuya, F.; Zhao, L.; Araki, O.; West, B.L.; Hanover, J.A.; Willingham, M.C.; Cheng, S.Y. Aberrant accumulation of PTTG1 induced by a mutated thyroid hormone beta receptor inhibits mitotic progression. J. Clin. Investig. 2006, 116, 2972–2984. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.-N.; Huang, Y.-H.; Yeh, C.-T.; Liao, C.-H.; Lin, K.-H. Thyroid hormone receptors suppress pituitary tumor transforming gene 1 activity in hepatoma. Cancer Res. 2008, 68, 1697–1706. [Google Scholar] [CrossRef] [PubMed]
- Bernal, J.; Luna, R.; Espina, G.; Lázaro, I.; Ramos-Morales, F.; Romero, F.; Arias, C.; Silva, A.; Tortolero, M.; Pintor-Toro, J. Human securin interacts with p53 and modulates p53-mediated transcriptional activity and apoptosis. Nat. Genet. 2002, 32, 306–311. [Google Scholar] [CrossRef] [PubMed]
- Horwitz, G.A.; Miklovsky, I.; Heaney, A.P.; Ren, S.-G.; Melmed, S. Human pituitary tumor-transforming gene (PTTG1) motif suppresses prolactin expression. Mol. Endocrinol. 2003, 17, 600–609. [Google Scholar] [CrossRef][Green Version]
- Zhou, C.; Tong, Y.; Wawrowsky, K.; Melmed, S. PTTG acts as a STAT3 target gene for colorectal cancer cell growth and motility. Oncogene 2014, 33, 851–861. [Google Scholar] [CrossRef]
- Falix, F.A.; Aronson, D.C.; Lamers, W.H.; Gaemers, I.C. Possible roles of DLK1 in the Notch pathway during development and disease. Biochim. Biophys. Acta Mol. Basis. Dis. 2012, 1822, 988–995. [Google Scholar] [CrossRef]
- Smas, C.M.; Sul, H.S. Pref-1, a protein containing EGF-like repeats, inhibits adipocyte differentiation. Cell 1993, 73, 725–734. [Google Scholar] [CrossRef]
- Laborda, J.; Sausville, E.A.; Hoffman, T.; Notario, V. Dlk, a putative mammalian homeotic gene differentially expressed in small cell lung carcinoma and neuroendocrine tumor cell line. J. Biol. Chem. 1993, 268, 3817–3820. [Google Scholar] [CrossRef]
- Smas, C.M.; Green, D.; Sul, H.S. Structural characterization and alternate splicing of the gene encoding the preadipocyte EGF-like protein Pref-1. Biochemistry 1994, 33, 9257–9265. [Google Scholar] [CrossRef]
- Takada, S.; Tevendale, M.; Baker, J.; Georgiades, P.; Campbell, E.; Freeman, T.; Johnson, M.; Paulsen, M.; Ferguson-Smith, A. Delta-like and gtl2 are reciprocally expressed, differentially methylated linked imprinted genes on mouse chromosome 12. Curr. Biol. 2000, 10, 1135–1138. [Google Scholar] [CrossRef]
- Charlier, C.; Segers, K.; Wagenaar, D.; Karim, L.; Berghmans, S.; Jaillon, O.; Shay, T.; Weissenbach, J.; Cockett, N.; Gyapay, G. Human-ovine comparative sequencing of a 250-kb imprinted domain encompassing the callipyge (clpg) locus and identification of six imprinted transcripts: DLK1, DAT, GTL2, PEG11, antiPEG11, and MEG8. Genome Res. 2001, 5, 850–862. [Google Scholar] [CrossRef] [PubMed]
- Buccarelli, M.; Lulli, V.; Giuliani, A.; Signore, M.; Martini, M.; D’Alessandris, Q.G.; Giannetti, S.; Novelli, A.; Ilari, R.; Giurato, G.; et al. Deregulated expression of the imprinted DLK1-DIO3 region in glioblastoma stemlike cells: Tumor suppressor role of lncRNA MEG3. Neuro-Oncol. 2020, 22, 1771–1784. [Google Scholar] [CrossRef] [PubMed]
- Kameswaran, V.; Bramswig, N.C.; McKenna, L.B.; Penn, M.; Schug, J.; Hand, N.; Chen, Y.; Choi, I.; Vourekas, A.; Won, K.J.; et al. Epigenetic Regulation of the DLK1-MEG3 MicroRNA Cluster in Human Type 2 Diabetic Islets. Cell Metab. 2014, 19, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Court, F.; Tayama, C.; Romanelli, V.; Martin-Trujillo, A.; Iglesias-Platas, I.; Okamura, K.; Sugahara, N.; Simón, C.; Moore, H.; Harness, J.V.; et al. Genome-wide parent-of-origin DNA methylation analysis reveals the intricacies of human imprinting and suggests a germline methylation-independent mechanism of establishment. Genome Res. 2014, 24, 554–569. [Google Scholar] [CrossRef]
- Lin, S.-P.; Youngson, N.; Takada, S.; Seitz, H.; Reik, W.; Paulsen, M.; Cavaille, J.; Ferguson-Smith, A. Asymmetric regulation of imprinting on the maternal and paternal chromosomes at the Dlk1-Gtl2 imprinted cluster on mouse chromosome 12. Nat. Genet. 2003, 35, 97–102. [Google Scholar] [CrossRef]
- Li, D.; Yea, S.; Li, S.; Chen, Z.; Narla, G.; Banck, M.; Laborda, J.; Tan, S.; Friedman, J.M.; Friedman, S.L.; et al. Krüppel-like factor-6 promotes preadipocyte differentiation through histone deacetylase 3-dependent repression of DLK1. J. Biol. Chem. 2005, 280, 26941–26952. [Google Scholar] [CrossRef]
- Lu, H.-P.; Lin, C.-J.; Chen, W.-C.; Chang, Y.-J.; Lin, S.-W.; Wang, H.-H.; Chang, C.-J. TRIM28 Regulates Dlk1 Expression in Adipogenesis. Int. J. Mol. Sci. 2020, 21, 7245. [Google Scholar] [CrossRef]
- Kim, Y.; Lin, Q.; Zelterman, D.; Yun, Z. Hypoxia-regulated delta-like 1 homologue enhances cancer cell stemness and tumorigenicity. Cancer Res. 2009, 69, 9271–9280. [Google Scholar] [CrossRef]
- Pittaway, J.F.H.; Lipsos, C.; Mariniello, K.; Guasti, L. The role of delta-like non-canonical Notch ligand 1 (DLK1) in cancer. Endocr. Relat. Cancer 2021, 28, R271–R287. [Google Scholar] [CrossRef]
- Wang, Y.; Kim, K.-A.; Kim, J.-H.; Sul, H.S. Pref-1, a preadipocyte secreted factor that inhibits adipogenesis. J. Nutr. 2006, 136, 2953–2956. [Google Scholar] [CrossRef] [PubMed]
- Traustadóttir, G.Á.; Lagoni, L.V.; Ankerstjerne, L.B.S.; Bisgaard, H.C.; Jensen, C.H.; Andersen, D.C. The imprinted gene Delta like non-canonical Notch ligand 1 (Dlk1) is conserved in mammals, and serves a growth modulatory role during tissue development and regeneration through Notch dependent and independent mechanisms. Cytokine Growth Factor Rev. 2019, 46, 17–27. [Google Scholar] [CrossRef]
- Wang, Y.; Sul, H.S. Ectodomain shedding of preadipocyte factor 1 (Pref-1) by tumor necrosis factor alpha converting enzyme (TACE) and inhibition of adipocyte differentiation. Mol. Cell Biol. 2006, 26, 5421–5435. [Google Scholar] [CrossRef] [PubMed]
- Smas, C.M.; Chen, L.; Sul, H.S. Cleavage of membrane-associated pref-1 generates a soluble inhibitor of adipocyte differentiation. Mol. Cell Biol. 1997, 17, 977–988. [Google Scholar] [CrossRef] [PubMed]
- Mei, B.; Zhao, L.; Chen, L.; Sul, H.S. Only the large soluble form of preadipocyte factor-1 (Pref-1), but not the small soluble and membrane forms, inhibits adipocyte differentiation: Role of alternative splicing. Biochem. J. 2002, 364, 137–144. [Google Scholar] [CrossRef]
- Floridon, C.; Jensen, C.H.; Thorsen, P.; Nielsen, O.; Sunde, L.; Westergaard, J.G.; Thomsen, S.G.; Teisner, B. Does fetal antigen 1 (FA1) identify cells with regenerative, endocrine and neuroendocrine potentials? A study of FA1 in embryonic, fetal, and placental tissue and in maternal circulation. Differentiation 2000, 66, 49–59. [Google Scholar] [CrossRef]
- Cleaton, M.A.; Dent, C.L.; Howard, M.; Corish, J.A.; Gutteridge, I.; Sovio, U.; Gaccioli, F.; Takahashi, N.; Bauer, S.R.; Charnock-Jones, D.S.; et al. Fetus-derived DLK1 is required for maternal metabolic adaptations to pregnancy and is associated with fetal growth restriction. Nat. Genet. 2016, 48, 1473–1480. [Google Scholar] [CrossRef]
- Zhu, N.-L.; Asahina, K.; Wang, J.; Ueno, A.; Lazaro, R.; Miyaoka, Y.; Miyajima, A.; Tsukamoto, H. Hepatic stellate cell-derived delta-like homolog 1 (DLK1) protein in liver regeneration. J. Biol. Chem. 2012, 287, 10355–10367. [Google Scholar] [CrossRef]
- Waddell, J.N.; Zhang, P.; Wen, Y.; Gupta, S.K.; Yevtodiyenko, A.; Schmidt, J.V.; Bidwell, C.A.; Kumar, A.; Kuang, S. Dlk1 is necessary for proper skeletal muscle development and regeneration. PLoS ONE 2010, 5, e15055. [Google Scholar] [CrossRef]
- Moon, Y.S.; Smas, C.M.; Lee, K.; Villena, J.A.; Kim, K.H.; Yun, E.J.; Sul, H.S. Mice lacking paternally expressed Pref-1/Dlk1 display growth retardation and accelerated adiposity. Mol. Cell Biol. 2002, 22, 5585–5592. [Google Scholar] [CrossRef]
- Ansell, P.J.; Zhou, Y.; Schjeide, B.-M.; Kerner, A.; Zhao, J.; Zhang, X.; Klibanski, A. Regulation of growth hormone expression by Delta-like protein 1 (Dlk1). Mol. Cell Endocrinol. 2007, 271, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Raghunandan, R.; Ruiz-Hidalgo, M.; Jia, Y.; Ettinger, R.; Rudikoff, E.; Riggins, P.; Farnsworth, R.; Tesfaye, A.; Laborda, J.; Bauer, S.R. Dlk1 influences differentiation and function of B lymphocytes. Stem. Cells Dev. 2008, 17, 495–507. [Google Scholar] [CrossRef] [PubMed]
- Puertas-Avendaño, R.A.; Gonzalez-Gomez, M.J.; Ruvira, M.D.; Hidalgo, M.J.R.; Morales, N.; Laborda, J.; Díaz, C.; Bello, A.R. Role of the non-canonical notch ligand delta-like protein 1 in hormone-producing cells of the adult male mouse pituitary. J. Neuroendocr. 2011, 23, 849–859. [Google Scholar] [CrossRef] [PubMed]
- Abdallah, B.M.; Ditzel, N.; Mahmood, A.; Isa, A.; A Traustadottir, G.; Schilling, A.F.; Ruiz-Hidalgo, M.-J.; Laborda, J.; Amling, M.; Kassem, M. DLK1 is a novel regulator of bone mass that mediates estrogen deficiency-induced bone loss in mice. J. Bone Miner. Res. Off. J. Am. Soc. Bone Miner. Res. 2011, 26, 1457–1471. [Google Scholar] [CrossRef]
- Miyaoka, Y.; Tanaka, M.; Imamura, T.; Takada, S.; Miyajima, A. A novel regulatory mechanism for Fgf18 signaling involving cysteine-rich FGF receptor (Cfr) and delta-like protein (Dlk). Development 2010, 137, 159–167. [Google Scholar] [CrossRef]
- Surmacz, B.; Noisa, P.; Risner-Janiczek, J.R.; Hui, K.; Ungless, M.; Cui, W.; Li, M. DLK1 promotes neurogenesis of human and mouse pluripotent stem cell-derived neural progenitors via modulating Notch and BMP signalling. Stem Cell Rev. Rep. 2012, 8, 459–471. [Google Scholar] [CrossRef]
- Ohno, N.; Izawa, A.; Hattori, M.; Kageyama, R.; Sudo, T. Dlk inhibits stem cell factor-induced colony formation of murine hematopoietic progenitors: Hes-1-independent effect. Stem Cells. 2001, 19, 71–79. [Google Scholar] [CrossRef]
- Tanimizu, N.; Nishikawa, M.; Saito, H.; Tsujimura, T.; Miyajima, A. Isolation of hepatoblasts based on the expression of Dlk/Pref-1. J. Cell Sci. 2003, 116, 1775–1786. [Google Scholar] [CrossRef]
- Lee, K.; Villena, J.A.; Moon, Y.S.; Kim, K.-H.; Lee, S.; Kang, C.; Sul, H.S. Inhibition of adipogenesis and development of glucose intolerance by soluble preadipocyte factor-1 (Pref-1). J. Clin. Investig. 2003, 111, 453–461. [Google Scholar] [CrossRef]
- Wang, Y.; Sul, H.S. Pref-1 regulates mesenchymal cell commitment and differentiation through Sox9. Cell Metab. 2009, 9, 287–302. [Google Scholar] [CrossRef]
- Wang, Y.; Zhao, L.; Smas, C.; Sul, H.S. Pref-1 interacts with fibronectin to inhibit adipocyte differentiation. Mol. Cell Biol. 2010, 30, 3480–3492. [Google Scholar] [CrossRef] [PubMed]
- Nueda, M.L.; González-Gómez, M.J.; Rodríguez-Cano, M.M.; Monsalve, E.M.; Díaz-Guerra, M.J.; Sánchez-Solana, B.; Laborda, J.; Baladrón, V. DLK proteins modulate NOTCH signaling to influence a brown or white 3T3-L1 adipocyte fate. Sci. Rep. 2018, 8, 16923. [Google Scholar] [CrossRef] [PubMed]
- Hudak, C.; Sul, H.S. Pref-1, a Gatekeeper of Adipogenesis. Front. Endocrinol. 2013, 4, 79. [Google Scholar] [CrossRef] [PubMed]
- Mortensen, S.B.; Jensen, C.H.; Schneider, M.; Thomassen, M.; Kruse, T.A.; Laborda, J.; Sheikh, S.P.; Andersen, D.C. Membrane-tethered delta-like 1 homolog (DLK1) restricts adipose tissue size by inhibiting preadipocyte proliferation. Diabetes 2012, 61, 2814–2822. [Google Scholar] [CrossRef] [PubMed]
- Nueda, M.-L.; García-Ramírez, J.J.; Laborda, J.; Baladrón, V. Dlk1 Specifically Interacts with Insulin-Like Growth Factor Binding Protein 1 to Modulate Adipogenesis of 3T3-L1 Cells. J. Mol. Biol. 2008, 379, 428–442. [Google Scholar] [CrossRef]
- Wolfrum, C.; Shih, D.Q.; Kuwajima, S.; Norris, A.W.; Kahn, C.R.; Stoffel, M. Role of Foxa-2 in adipocyte metabolism and differentiation. J. Clin. Investig. 2003, 112, 345–356. [Google Scholar] [CrossRef]
- Smas, C.M.; Chen, L.; Zhao, L.; Latasa, M.J.; Sul, H.S. Transcriptional repression of pref-1 by glucocorticoids promotes 3T3-L1 adipocyte differentiation. J. Biol. Chem. 1999, 274, 12632–12641. [Google Scholar] [CrossRef]
- Grassi, E.S.; Pantazopoulou, V.; Pietras, A. Hypoxia-induced release, nuclear translocation, and signaling activity of a DLK1 intracellular fragment in glioma. Oncogene 2020, 39, 4028–4044. [Google Scholar] [CrossRef]
- She, H.; Xiong, S.; Hazra, S.; Tsukamoto, H. Adipogenic Transcriptional Regulation of Hepatic Stellate Cells. J. Biol. Chem. 2005, 280, 4959–4967. [Google Scholar] [CrossRef]
- Tsukamoto, H.; She, H.; Hazra, S.; Cheng, J.; Miyahara, T. Anti-adipogenic regulation underlies hepatic stellate cell transdifferentiation. J. Gastroenterol. Hepatol. 2006, 21, S102–S105. [Google Scholar] [CrossRef]
- Levene, A.P.; Goldin, R.D. The epidemiology, pathogenesis and histopathology of fatty liver disease. Histopathology 2012, 61, 141–152. [Google Scholar] [CrossRef] [PubMed]
- Vernon, G.; Baranova, A.; Younossi, Z.M. Systematic review: The epidemiology and natural history of non-alcoholic fatty liver disease and non-alcoholic steatohepatitis in adults. Aliment. Pharmacol. Ther. 2011, 34, 274–285. [Google Scholar] [CrossRef] [PubMed]
- Sanyal, A.J.; Brunt, E.M.; Kleiner, D.E.; Kowdley, K.V.; Chalasani, N.; Lavine, J.E.; Ratziu, V.; McCullough, A. Endpoints and clinical trial design for nonalcoholic steatohepatitis. Hepatology 2011, 54, 344–353. [Google Scholar] [CrossRef] [PubMed]
- Brunt, E.M.; Neuschwander-Tetri, B.A.; Oliver, D.; Wehmeier, K.R.; Bacon, B.R. Nonalcoholic steatohepatitis: Histologic features and clinical correlations with 30 blinded biopsy specimens. Hum. Pathol. 2004, 35, 1070–1082. [Google Scholar] [CrossRef] [PubMed]
- Adams, L.A.; Lymp, J.F.; Sauver, J.S.; Sanderson, S.O.; Lindor, K.D.; Feldstein, A.; Angulo, P. The natural history of nonalcoholic fatty liver disease: A population-based cohort study. Gastroenterology 2005, 129, 113–121. [Google Scholar] [CrossRef]
- Matteoni, C.A.; Younossi, Z.M.; Gramlich, T.; Boparai, N.; Liu, Y.C.; McCullough, A.J. Nonalcoholic fatty liver disease: A spectrum of clinical and pathological severity. Gastroenterology 1999, 116, 1413–1419. [Google Scholar] [CrossRef]
- Ertle, J.; Dechêne, A.; Sowa, J.P.; Penndorf, V.; Herzer, K.; Kaiser, G.; Schlaak, J.F.; Gerken, G.; Syn, W.K.; Canbay, A. Non-alcoholic fatty liver disease progresses to hepatocellular carcinoma in the absence of apparent cirrhosis. Int. J. Cancer 2011, 128, 2436–2443. [Google Scholar] [CrossRef]
- Bugianesi, E.; McCullough, A.J.; Marchesini, G. Insulin resistance: A metabolic pathway to chronic liver disease. Hepatology 2005, 42, 987–1000. [Google Scholar] [CrossRef]
- Greco, D.; Kotronen, A.; Westerbacka, J.; Puig, O.; Arkkila, P.; Kiviluoto, T.; Laitinen, S.; Kolak, M.; Fisher, R.M.; Hamsten, A.; et al. Gene expression in human NAFLD. Am. J. Physiol. Gastrointest. Liver Physiol. 2008, 294, G1281–G1287. [Google Scholar] [CrossRef]
- Fabbrini, E.; Magkos, F.; Mohammed, B.S.; Pietka, T.; Abumrad, N.A.; Patterson, B.W.; Okunade, A.; Klein, S. Intrahepatic fat, not visceral fat, is linked with metabolic complications of obesity. Proc. Natl. Acad. Sci. USA 2009, 106, 15430–15435. [Google Scholar] [CrossRef]
- Dowman, J.K.; Tomlinson, J.W.; Newsome, P.N. Systematic review: The diagnosis and staging of non-alcoholic fatty liver disease and non-alcoholic steatohepatitis. Aliment. Pharmacol. Ther. 2011, 33, 525–540. [Google Scholar] [CrossRef] [PubMed]
- Shulman, G.I. Cellular mechanisms of insulin resistance. J. Clin. Investig. 2000, 106, 171–176. [Google Scholar] [CrossRef] [PubMed]
- Tarantino, G.; Finelli, C. Pathogenesis of hepatic steatosis: The link between hypercortisolism and non-alcoholic fatty liver disease. World J. Gastroenterol. 2013, 19, 6735–6743. [Google Scholar] [CrossRef] [PubMed]
- Bugianesi, E.; Moscatiello, S.; Ciaravella, M.F.; Marchesini, G. Insulin resistance in nonalcoholic fatty liver disease. Curr. Pharm. Des. 2010, 16, 1941–1951. [Google Scholar] [CrossRef] [PubMed]
- Guilherme, A.; Virbasius, J.V.; Puri, V.; Czech, M.P. Adipocyte dysfunctions linking obesity to insulin resistance and type 2 diabetes. Nat. Rev. Mol. Cell Biol. 2008, 9, 367–377. [Google Scholar] [CrossRef] [PubMed]
- Cusi, K. Role of insulin resistance and lipotoxicity in non-alcoholic steatohepatitis. Clin. Liver Dis. 2009, 13, 545–563. [Google Scholar] [CrossRef]
- Villena, J.A.; Choi, C.S.; Wang, Y.; Kim, S.; Hwang, Y.-J.; Kim, Y.-B.; Cline, G.; Shulman, G.I.; Sul, H.S. Resistance to High-Fat Diet–Induced Obesity but Exacerbated Insulin Resistance in Mice Overexpressing Preadipocyte Factor-1 (Pref-1). Diabetes 2008, 57, 3258–3266. [Google Scholar] [CrossRef]
- Kavalkova, P.; Touskova, V.; Roubicek, T.; Trachta, P.; Urbanova, M.; Drapalova, J.; Haluzikova, D.; Mraz, M.; Novak, D.; Matoulek, M.; et al. Serum preadipocyte factor-1 concentrations in females with obesity and type 2 diabetes mellitus: The influence of very low calorie diet, acute hyperinsulinemia, and fenofibrate treatment. Horm. Metab. Res. 2013, 45, 820–826. [Google Scholar] [CrossRef]
- Charalambous, M.; Da Rocha, S.T.; Radford, E.J.; Medina-Gomez, G.; Curran, S.; Pinnock, S.B.; Ferrón, S.R.; Vidal-Puig, A.; Ferguson-Smith, A.C. DLK1/PREF1 regulates nutrient metabolism and protects from steatosis. Proc. Natl. Acad. Sci. USA 2014, 111, 16088–16093. [Google Scholar] [CrossRef]
- Jensen, C.H.; Kosmina, R.; Rydén, M.; Baun, C.; Hvidsten, S.; Andersen, M.; Christensen, L.L.; Gastaldelli, A.; Marraccini, P.; Arner, P.; et al. The imprinted gene Delta like non-canonical notch ligand 1 (Dlk1) associates with obesity and triggers insulin resistance through inhibition of skeletal muscle glucose uptake. EBioMedicine 2019, 46, 368–380. [Google Scholar] [CrossRef]
- Xu, H.; Wang, L. The Role of Notch Signaling Pathway in Non-Alcoholic Fatty Liver Disease. Front. Mol. Biosci. 2021, 8, 792667. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.H.; Yun, M.R.; Kim, H.M.; Jeon, B.H.; Park, B.C.; Lee, B.W.; Kang, E.S.; Lee, H.C.; Park, Y.W.; Cha, B.S. Exogenous administration of DLK1 ameliorates hepatic steatosis and regulates gluconeogenesis via activation of AMPK. Int. J. Obes. 2016, 40, 356–365. [Google Scholar] [CrossRef] [PubMed]
- Cao, C.; Duan, P.; Li, W.; Guo, Y.; Zhang, J.; Gui, Y.; Yuan, S. Lack of miR-379/miR-544 Cluster Resists High-Fat Diet-Induced Obesity and Prevents Hepatic Triglyceride Accumulation in Mice. Front. Cell Dev. Biol. 2021, 9, 720900. [Google Scholar] [CrossRef] [PubMed]
- Friedman, S.L. Liver fibrosis—From bench to bedside. J. Hepatol. 2003, 38, S38–S53. [Google Scholar] [CrossRef]
- Bataller, R.; Brenner, D.A. Liver fibrosis. J. Clin. Investig. 2005, 115, 209–218. [Google Scholar] [CrossRef]
- Arthur, M.J. Fibrogenesis II. Metalloproteinases and their inhibitors in liver fibrosis. Am. J. Physiol. Gastrointest. Liver Physiol. 2000, 279, G245–G249. [Google Scholar] [CrossRef]
- Gäbele, E.; Brenner, D.A.; Rippe, R.A. Liver fibrosis: Signals leading to the amplification of the fibrogenic hepatic stellate cell. Front. Biosci. 2003, 8, d69–d77. [Google Scholar]
- Friedman, S.L.; Rockey, D.C.; McGuire, R.F.; Maher, J.J.; Boyles, J.K.; Yamasaki, G. Isolated hepatic lipocytes and Kupffer cells from normal human liver: Morphological and functional characteristics in primary culture. Hepatology 1992, 15, 234–243. [Google Scholar] [CrossRef]
- Lindquist, J.N.; Stefanovic, B.; Brenner, D.A. Regulation of collagen alpha1(I) expression in hepatic stellate cells. J. Gastroenterol. 2000, 35, 80–83. [Google Scholar]
- Magness, S.T.; Bataller, R.; Yang, L.; Brenner, D.A. A dual reporter gene transgenic mouse demonstrates heterogeneity in hepatic fibrogenic cell populations. Hepatology 2004, 40, 1151–1159. [Google Scholar] [CrossRef]
- Schattenberg, J.M.; Nagel, M.; Kim, Y.O.; Kohl, T.; Wörns, M.A.; Zimmermann, T.; Schad, A.; Longerich, T.; Schuppan, D.; He, Y.W.; et al. Increased hepatic fibrosis and JNK2-dependent liver injury in mice exhibiting hepatocyte-specific deletion of cFLIP. Am. J. Physiol. Gastrointest. Liver Physiol. 2012, 303, G498–G506. [Google Scholar] [CrossRef] [PubMed]
- Zeisberg, E.M.; Potenta, S.; Xie, L.; Zeisberg, M.; Kalluri, R. Discovery of endothelial to mesenchymal transition as a source for carcinoma-associated fibroblasts. Cancer Res. 2007, 67, 10123–10128. [Google Scholar] [CrossRef] [PubMed]
- Jeong, W.; Do, S.H.; Yun, H.S.; Song, B.J.; Kim, S.J.; Kwak, W.J.; Yoo, S.E.; Park, H.Y.; Jeong, K.S. Hypoxia potentiates transforming growth factor-beta expression of hepatocyte during the cirrhotic condition in rat liver. Liver Int. 2004, 24, 658–668. [Google Scholar] [CrossRef] [PubMed]
- Iredale, J.P.; Thompson, A.; Henderson, N.C. Extracellular matrix degradation in liver fibrosis: Biochemistry and regulation. Biochim. Biophys. Acta 2013, 1832, 876–883. [Google Scholar] [CrossRef] [PubMed]
- Cawston, T.E.; Mercer, E. Preferential binding of collagenase to alpha 2-macroglobulin in the presence of the tissue inhibitor of metalloproteinases. FEBS Lett. 1986, 209, 9–12. [Google Scholar] [CrossRef]
- Visse, R.; Nagase, H. Matrix metalloproteinases and tissue inhibitors of metalloproteinases: Structure, function, and biochemistry. Circ. Res. 2003, 92, 827–839. [Google Scholar] [CrossRef]
- Mastroianni, C.M.; Liuzzi, G.M.; D’Ettorre, G.; Lichtner, M.; Forcina, G.; Di Campli, N.F.; Riccio, P.; Vullo, V. Matrix metalloproteinase-9 and tissue inhibitors of matrix metalloproteinase-1 in plasma of patients co-infected with HCV and HIV. HIV Clin. Trials 2002, 3, 310–315. [Google Scholar]
- Huang, C.-C.; Huang, L.L.; Chou, M.-H.; Wu, C.-L.; Chen, C.-M.; Hsieh, C.-S.; Lee, S.-Y.; Chen, C.-L. The human Delta-like 1 homologue is implicated in the progression of liver fibrosis in biliary atresia. J. Pathol. 2004, 202, 172–179. [Google Scholar] [CrossRef]
- Pan, R.L.; Wang, P.; Xiang, L.X.; Shao, J.Z. Delta-like 1 serves as a new target and contributor to liver fibrosis down-regulated by mesenchymal stem cell transplantation. J. Biol. Chem. 2011, 286, 12340–12348. [Google Scholar] [CrossRef]
- Pan, R.-L.; Xiang, L.-X.; Wang, P.; Liu, X.-Y.; Nie, L.; Huang, W.; Shao, J.-Z. Low-molecular-weight fibroblast growth factor 2 attenuates hepatic fibrosis by epigenetic down-regulation of Delta-like1. Hepatology 2015, 61, 1708–1720. [Google Scholar] [CrossRef]
- Buko, V.; Belonovskaya, E.; Naruta, E.; Lukivskaya, O.; Kanyuka, O.; Zhuk, O.; Kranc, R.; Stoika, R.; Sybirna, N. Pituitary tumor transforming gene as a novel regulatory factor of liver fibrosis. Life Sci. 2015, 132, 34–40. [Google Scholar] [CrossRef] [PubMed]
- Perramón, M.; Carvajal, S.; Reichenbach, V.; Fernández-Varo, G.; Boix, L.; Macias-Muñoz, L.; Melgar-Lesmes, P.; Bruix, J.; Melmed, S.; Lamas, S.; et al. The pituitary tumour-transforming gene 1/delta-like homologue 1 pathway plays a key role in liver fibrogenesis. Liver Int. 2022, 42, 651–662. [Google Scholar] [CrossRef] [PubMed]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Yu, L.X.; Ling, Y.; Wang, H.Y. Role of nonresolving inflammation in hepatocellular carcinoma development and progression. Npj Precis. Oncol. 2018, 2, 6. [Google Scholar] [CrossRef]
- Villanueva, A. Hepatocellular Carcinoma. N. Engl. J. Med. 2019, 380, 1450–1462. [Google Scholar] [CrossRef] [PubMed]
- Roberts, S.M.; Kehrer, J.P.; Klotz, L.O. Studies on Experimental Toxicology and Pharmacology, 1st ed.; Roberts, S.M., Kehrer, J.P., Klotz, L.-O., Eds.; Humana Press: Totowa, NJ, USA, 2015; 498p. [Google Scholar]
- Nault, J.C.; Zucman-Rossi, J. TERT promoter mutations in primary liver tumors. Clin. Res. Hepatol. Gastroenterol. 2016, 40, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Xiao, Z.; Yang, L.; Gao, Y.; Zhu, Q.; Hu, L.; Huang, D.; Xu, Q. Hypoxia-inducible factors in hepatocellular carcinoma (Review). Oncol. Rep. 2020, 43, 3–15. [Google Scholar] [CrossRef] [PubMed]
- Capece, D.; Fischietti, M.; Verzella, D.; Gaggiano, A.; Cicciarelli, G.; Tessitore, A.; Zazzeroni, F.; Alesse, E. The Inflammatory Microenvironment in Hepatocellular Carcinoma: A Pivotal Role for Tumor-Associated Macrophages. Biomed. Res. Int. 2013, 2013, 187204. [Google Scholar] [CrossRef] [PubMed]
- Waris, G.; Ahsan, H. Reactive oxygen species: Role in the development of cancer and various chronic conditions. J. Carcinog. 2006, 5, 14. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.; Luan, J.; Huang, C.; Li, J. Tumor-Associated Macrophages in Hepatocellular Carcinoma: Friend or Foe? Gut Liver 2021, 15, 500–516. [Google Scholar] [CrossRef]
- Nault, J.C.; Ningarhari, M.; Rebouissou, S.; Zucman-Rossi, J. The role of telomeres and telomerase in cirrhosis and liver cancer. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 544–558. [Google Scholar] [CrossRef] [PubMed]
- Ogunwobi, O.O.; Harricharran, T.; Huaman, J.; Galuza, A.; Odumuwagun, O.; Tan, Y.; Ma, G.X.; Nguyen, M.T. Mechanisms of hepatocellular carcinoma progression. World J. Gastroenterol. 2019, 25, 2279–2293. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Xiog, Y.; Wang, G.; Li, W.; Tang, T.; Sun, J.; Li, J. miR-374c-5p regulates PTTG1 and inhibits cell growth and metastasis in hepatocellular carcinoma by regulating epithelial-mesenchymal transition. Mol. Med. Rep. 2022, 25, 148. [Google Scholar] [CrossRef]
- Jung, C.R.; Yoo, J.; Jang, Y.J.; Kim, S.; Chu, I.S.; Yeom, Y.I.; Choi, J.Y.; Im, D.S. Adenovirus-mediated transfer of siRNA against PTTG1 inhibits liver cancer cell growth in vitro and in vivo. Hepatology 2006, 43, 1042–1052. [Google Scholar] [CrossRef]
- Liang, M.; Chen, X.; Liu, W.; Li, S.; Li, C.; Jiang, L.; Lv, S. Role of the pituitary tumor transforming gene 1 in the progression of hepatocellular carcinoma. Cancer Biol. Ther. 2011, 11, 337–345. [Google Scholar] [CrossRef][Green Version]
- Xie, Y. Hepatitis B Virus-Associated Hepatocellular Carcinoma. Adv. Exp. Biol. 2017, 1018, 11–21. [Google Scholar]
- Li, C.; Wang, Y.; Wang, S.; Wu, B.; Hao, J.; Fan, H.; Ju, Y.; Ding, Y.; Chen, L.; Chu, X.; et al. Hepatitis B virus mRNA-mediated miR-122 inhibition upregulates PTTG1-binding protein, which promotes hepatocellular carcinoma tumor growth and cell invasion. J. Virol. 2013, 87, 2193–2205. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Yang, Y.; Guo, Y.; Liu, H.; Jiang, J.; Zheng, F.; Wu, B. PTTG1 is involved in TNF-α-related hepatocellular carcinoma via the induction of c-myc. Cancer Med. 2019, 8, 5702–5715. [Google Scholar] [CrossRef]
- Fujii, T.; Nomoto, S.; Koshikawa, K.; Yatabe, Y.; Teshigawara, O.; Mori, T.; Inoue, S.; Takeda, S.; Nakao, A. Overexpression of pituitary tumor transforming gene 1 in HCC is associated with angiogenesis and poor prognosis. Hepatology 2006, 43, 1267–1275. [Google Scholar] [CrossRef]
- El-Masry, S.A.M.; Elsherbini, S.M.; Abdelmoneam, A.A.; Radwan, A. Role of Human Pituitary Tumor-Transforming Gene1 (HPTTG1) as a prognostic biomarker for metastasis in Egyptian patients breast, colon and Liver cancer. J. Biosci. Appl. Res. 2020, 6, 83–97. [Google Scholar] [CrossRef]
- Wang, J.; Peng, R.; Zhang, Z.; Zhang, Y.; Dai, Y.; Sun, Y. Identification and Validation of Key Genes in Hepatocellular Carcinoma by Bioinformatics Analysis. Biomed. Res. Int. 2021, 2021, 6662114. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Guo, J.; Zhou, J.; Lu, J.; Chen, Q.; Zhang, C.; Qing, C.; Koeffler, H.P.; Tong, Y. FoxM1 transactivates PTTG1 and promotes colorectal cancer cell migration and invasion. BMC Med. Genom. 2015, 12, 49. [Google Scholar] [CrossRef]
- Huang, J.-L.; Cao, S.-W.; Ou, Q.-S.; Yang, B.; Zheng, S.-H.; Tang, J.; Chen, J.; Hu, Y.-W.; Zheng, L.; Wang, Q. The long non-coding RNA PTTG3P promotes cell growth and metastasis via up-regulating PTTG1 and activating PI3K/AKT signaling in hepatocellular carcinoma. Mol. Cancer 2018, 17, 93. [Google Scholar] [CrossRef] [PubMed]
- Grassi, E.S.; Pietras, A. Emerging Roles of DLK1 in the Stem Cell Niche and Cancer Stemness. J. Histochem. Cytochem. Off. J. Histochem. Soc. 2022, 70, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhao, X.; Wang, J.; Cheng, Y.; Wu, Q.; Wang, B.; Zhao, F.; Meng, L.; Zhang, Y.; Jin, M.; et al. Distinct roles of Dlk1 isoforms in bi-potential differentiation of hepatic stem cells. Stem Cell Res. Ther. 2019, 10, 31. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.H.; Ren, B.; Keryanov, S.; Tseng, G.C.; Rao, U.N.; Monga, S.P.; Strom, S.; Demetris, A.J.; Nalesnik, M.; Yu, Y.P.; et al. Transcriptomic and genomic analysis of human hepatocellular carcinomas and hepatoblastomas. Hepatology 2006, 44, 1012–1024. [Google Scholar] [CrossRef] [PubMed]
- Dezső, K.; Halász, J.; Bisgaard, H.C.; Paku, S.; Turányi, E.; Schaff, Z.; Nagy, P. Delta-like protein (DLK) is a novel immunohistochemical marker for human hepatoblastomas. Virchows Arch. 2008, 452, 443–448. [Google Scholar] [CrossRef]
- Cairo, S.; Armengol, C.; De Reyniès, A.; Wei, Y.; Thomas, E.; Renard, C.-A.; Goga, A.; Balakrishnan, A.; Semeraro, M.; Gresh, L.; et al. Hepatic stem-like phenotype and interplay of Wnt/beta-catenin and Myc signaling in aggressive childhood liver cancer. Cancer Cell 2008, 14, 471–484. [Google Scholar] [CrossRef]
- López-Terrada, D.; Gunaratne, P.H.; Adesina, A.M.; Pulliam, J.; Hoang, D.M.; Nguyen, Y.; Mistretta, T.-A.; Margolin, J.; Finegold, M.J. Histologic subtypes of hepatoblastoma are characterized by differential canonical Wnt and Notch pathway activation in DLK+ precursors. Hum. Pathol. 2009, 40, 783–794. [Google Scholar] [CrossRef]
- Litten, J.B.; Chen, T.T.; Schultz, R.; Herman, K.; Comstock, J.; Schiffman, J.; Tomlinson, G.E.; Rakheja, D. Activated NOTCH2 is overexpressed in hepatoblastomas: An immunohistochemical study. Pediatr. Dev. Pathol. 2011, 14, 378–383. [Google Scholar] [CrossRef]
- Carrillo-Reixach, J.; Torrens, L.; Simon-Coma, M.; Royo, L.; Domingo-Sàbat, M.; Abril-Fornaguera, J.; Akers, N.; Sala, M.; Ragull, S.; Arnal, M.; et al. Epigenetic footprint enables molecular risk stratification of hepatoblastoma with clinical implications. J. Hepatol. 2020, 73, 328–341. [Google Scholar] [CrossRef] [PubMed]
- Honda, S.; Chatterjee, A.; Leichter, A.L.; Miyagi, H.; Minato, M.; Fujiyoshi, S.; Ara, M.; Kitagawa, N.; Tanaka, M.; Tanaka, Y.; et al. A MicroRNA Cluster in the DLK1-DIO3 Imprinted Region on Chromosome 14q32.2 Is Dysregulated in Metastatic Hepatoblastomas. Front. Oncol. 2020, 10, 513601. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhang, X.; Zhang, M.; Zhu, J.-D.; Zhang, Y.-L.; Lin, Y.; Wang, K.-S.; Qi, X.-F.; Zhang, Q.; Liu, G.-Z.; et al. Up-regulation of DLK1 as an imprinted gene could contribute to human hepatocellular carcinoma. Carcinogenesis 2007, 28, 1094–1103. [Google Scholar] [CrossRef] [PubMed]
- Jin, Z.; Yang, R.; Dong, B.; Xing, B. Progenitor gene DLK1 might be an independent prognostic factor of liver cancer. Expert Opin. Biol. Ther. 2008, 8, 371–377. [Google Scholar] [CrossRef]
- Yanai, H.; Nakamura, K.; Hijioka, S.; Kamei, A.; Ikari, T.; Ishikawa, Y.; Shinozaki, E.; Mizunuma, N.; Hatake, K.; Miyajima, A. Dlk-1, a cell surface antigen on foetal hepatic stem/progenitor cells, is expressed in hepatocellular, colon, pancreas and breast carcinomas at a high frequency. J. Biochem. 2010, 148, 85–92. [Google Scholar] [CrossRef]
- Ko, S.; Russell, J.O.; Shin, D.P.; Monga, S.P. Notch–Sox9 Axis Mediates Hepatocyte Dedifferentiation in KrasG12V-Induced Zebrafish Hepatocellular Carcinoma. Int. J. Mol. Sci. 2022, 23, 4705. [Google Scholar]
- Luk, J.M.; Burchard, J.; Zhang, C.; Liu, A.M.; Wong, K.F.; Shek, F.H.; Lee, N.P.; Fan, S.T.; Poon, R.T.; Ivanovska, I.; et al. DLK1-DIO3 genomic imprinted microRNA cluster at 14q32.2 defines a stemlike subtype of hepatocellular carcinoma associated with poor survival. J. Biol. Chem. 2011, 286, 30706–30713. [Google Scholar] [CrossRef]
- Lempiäinen, H.; Couttet, P.; Bolognani, F.; Müller, A.; Dubost, V.; Luisier, R.; Espinola, A.D.R.; Vitry, V.; Unterberger, E.B.; Thomson, J.P.; et al. Identification of Dlk1-Dio3 imprinted gene cluster noncoding RNAs as novel candidate biomarkers for liver tumor promotion. Toxicol. Sci. 2013, 131, 375–386. [Google Scholar] [CrossRef]
- Xu, X.; Liu, R.-F.; Zhang, X.; Huang, L.-Y.; Chen, F.; Fei, Q.-L.; Han, Z.-G. DLK1 as a potential target against cancer stem/progenitor cells of hepatocellular carcinoma. Mol. Cancer Ther. 2012, 11, 629–638. [Google Scholar] [CrossRef]
- Li, H.; Cui, M.-L.; Chen, T.-Y.; Xie, H.-Y.; Cui, Y.; Tu, H.; Chen, F.-H.; Ge, C.; Li, J.-J. Serum DLK1 is a potential prognostic biomarker in patients with hepatocellular carcinoma. Tumor Biol. J. Int. Soc. Oncodev. Biol. Med. 2015, 36, 8399–8404. [Google Scholar] [CrossRef]
- Cai, C.-M.; Xiao, X.; Wu, B.-H.; Wei, B.-F.; Han, Z.-G. Targeting endogenous DLK1 exerts antitumor effect on hepatocellular carcinoma through initiating cell differentiation. Oncotarget 2016, 7, 71466–71476. [Google Scholar] [CrossRef] [PubMed]
- Liem, I.K.; Adnindya, M.R.; Nasution, A.A. Roles of DLK1 in Liver Development and Oncogenesis. Online J. Biol. Sci. 2017, 17, 309–315. [Google Scholar]
- Khalaf, A.M.; Fuentes, D.; Morshid, A.I.; Burke, M.R.; Kaseb, A.O.; Hassan, M.; Hazle, J.D.; Elsayes, K.M. Role of Wnt/β-catenin signaling in hepatocellular carcinoma, pathogenesis, and clinical significance. J. Hepatocell. Carcinoma 2018, 5, 61–73. [Google Scholar] [CrossRef]
- Seino, S.; Tsuchiya, A.; Watanabe, Y.; Kawata, Y.; Kojima, Y.; Ikarashi, S.; Yanai, H.; Nakamura, K.; Kumaki, D.; Hirano, M.; et al. Clinical outcome of hepatocellular carcinoma can be predicted by the expression of hepatic progenitor cell markers and serum tumour markers. Oncotarget 2018, 9, 21844–21860. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Perramón, M.; Jiménez, W. Pituitary Tumor-Transforming Gene 1/Delta like Non-Canonical Notch Ligand 1 Signaling in Chronic Liver Diseases. Int. J. Mol. Sci. 2022, 23, 6897. https://doi.org/10.3390/ijms23136897
Perramón M, Jiménez W. Pituitary Tumor-Transforming Gene 1/Delta like Non-Canonical Notch Ligand 1 Signaling in Chronic Liver Diseases. International Journal of Molecular Sciences. 2022; 23(13):6897. https://doi.org/10.3390/ijms23136897
Chicago/Turabian StylePerramón, Meritxell, and Wladimiro Jiménez. 2022. "Pituitary Tumor-Transforming Gene 1/Delta like Non-Canonical Notch Ligand 1 Signaling in Chronic Liver Diseases" International Journal of Molecular Sciences 23, no. 13: 6897. https://doi.org/10.3390/ijms23136897
APA StylePerramón, M., & Jiménez, W. (2022). Pituitary Tumor-Transforming Gene 1/Delta like Non-Canonical Notch Ligand 1 Signaling in Chronic Liver Diseases. International Journal of Molecular Sciences, 23(13), 6897. https://doi.org/10.3390/ijms23136897