Newly Emerged Antiviral Strategies for SARS-CoV-2: From Deciphering Viral Protein Structural Function to the Development of Vaccines, Antibodies, and Small Molecules
Abstract
:1. Introduction of COVID-19
1.1. COVID-19 Pandemic
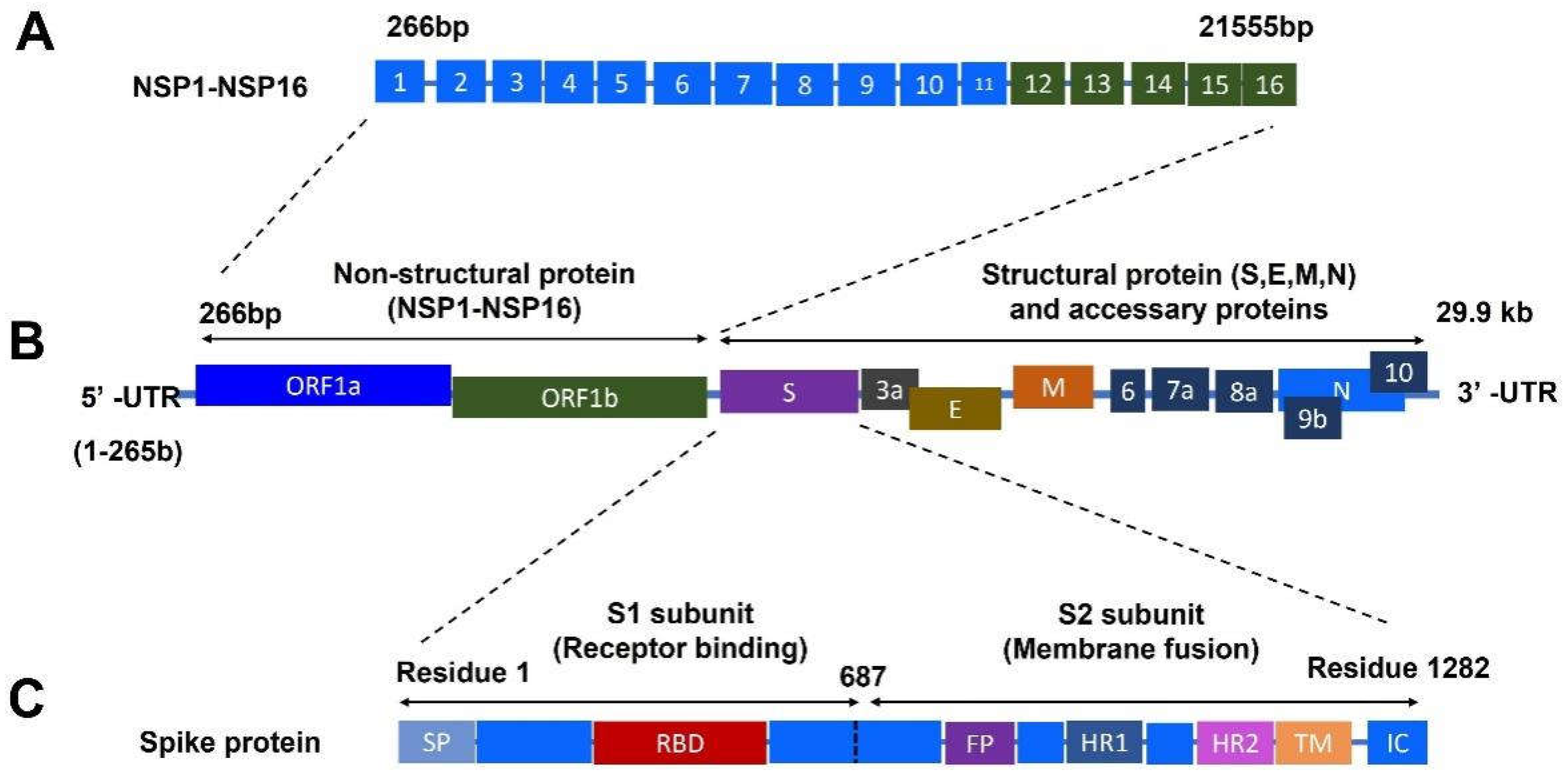
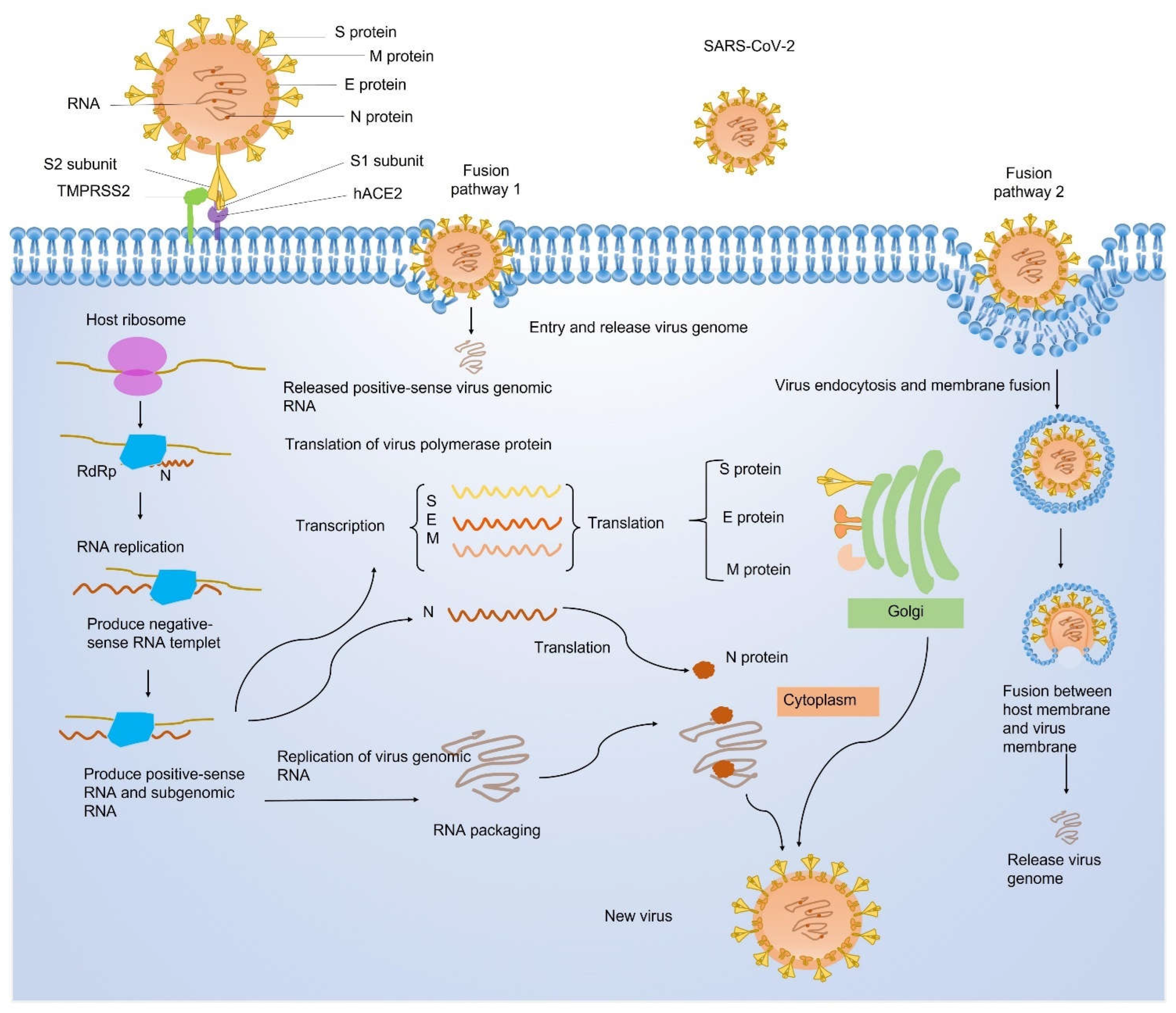
1.2. Structural Proteins of SARS-CoV-2
1.3. Nonstructural Proteins of SARS-CoV-2
1.4. Accessory Proteins
2. Vaccine Approach to Prevent Infection
2.1. SARS-CoV-2 Inactivated Vaccines
2.2. Protein-Based Vaccines
2.3. Nucleic Acid-Based Vaccines
2.3.1. DNA Vaccine
2.3.2. mRNA Vaccine
2.3.3. Circular RNA (circRNA) Vaccine
3. Antiviral Antibodies
3.1. Monoclonal Antibody
3.1.1. Monoclonal Antibody from Human Patients
3.1.2. Nanoparticle Antibody
3.1.3. Cross-Neutralizer Antibody (Across SARS-CoV-2 and SARS-CoV-1)
3.1.4. Constructed Cobinding Antibody Using Paired or Multiple IgG Sequences
3.1.5. Dual Functional Monoclonal Antibody
3.2. Antibody Cocktail
3.3. Engineered Bispecific Monoclonal Antibody
4. Peptide-Based Inhibitor
5. Chemical Compounds
6. CRISPR Cas Technology
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviation
References
- The species Severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [CrossRef] [PubMed] [Green Version]
- Dong, E.; Du, H.; Gardner, L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect. Dis. 2020, 20, 533–534. [Google Scholar] [CrossRef]
- Ejaz, H.; Alsrhani, A.; Zafar, A.; Javed, H.; Junaid, K.; Abdalla, A.E.; Abosalif, K.O.A.; Ahmed, Z.; Younas, S. COVID-19 and comorbidities: Deleterious impact on infected patients. J. Infect. Public Health 2020, 13, 1833–1839. [Google Scholar] [CrossRef] [PubMed]
- Machhi, J.; Herskovitz, J.; Senan, A.M.; Dutta, D.; Nath, B.; Oleynikov, M.D.; Blomberg, W.R.; Meigs, D.D.; Hasan, M.; Patel, M.; et al. The Natural History, Pathobiology, and Clinical Manifestations of SARS-CoV-2 Infections. J. Neuroimmune Pharmacol. 2020, 15, 359–386. [Google Scholar] [CrossRef]
- Yadav, R.; Chaudhary, J.K.; Jain, N.; Chaudhary, P.K.; Khanra, S.; Dhamija, P.; Sharma, A.; Kumar, A.; Handu, S. Role of Structural and Non-Structural Proteins and Therapeutic Targets of SARS-CoV-2 for COVID-19. Cells 2021, 10, 821. [Google Scholar] [CrossRef]
- Malik, Y.A. Properties of Coronavirus and SARS-CoV-2. Malays. J. Pathol. 2020, 42, 3–11. [Google Scholar]
- Wu, C.; Liu, Y.; Yang, Y.; Zhang, P.; Zhong, W.; Wang, Y.; Wang, Q.; Xu, Y.; Li, M.; Li, X.; et al. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm. Sin. B 2020, 10, 766–788. [Google Scholar] [CrossRef]
- Li, F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [Green Version]
- Xia, X. Domains and Functions of Spike Protein in SARS-Cov-2 in the Context of Vaccine Design. Viruses 2021, 13, 109. [Google Scholar] [CrossRef]
- Dakal, T.C. Antigenic sites in SARS-CoV-2 spike RBD show molecular similarity with pathogenic antigenic determinants and harbors peptides for vaccine development. Immunobiology 2021, 226, 152091. [Google Scholar] [CrossRef]
- Koehler, M.; Ray, A.; Moreira, R.A.; Juniku, B.; Poma, A.B.; Alsteens, D. Molecular insights into receptor binding energetics and neutralization of SARS-CoV-2 variants. Nat. Commun. 2021, 12, 6977. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Thambiraja, T.S.; Karuppanan, K.; Subramaniam, G. Omicron and Delta variant of SARS-CoV-2: A comparative computational study of spike protein. J. Med. Virol. 2022, 94, 1641–1649. [Google Scholar] [CrossRef] [PubMed]
- Moreira, R.A.; Chwastyk, M.; Baker, J.L.; Guzman, H.V.; Poma, A.B. Quantitative determination of mechanical stability in the novel coronavirus spike protein. Nanoscale 2020, 12, 16409–16413. [Google Scholar] [CrossRef] [PubMed]
- Casalino, L.; Gaieb, Z.; Goldsmith, J.A.; Hjorth, C.K.; Dommer, A.C.; Harbison, A.M.; Fogarty, C.A.; Barros, E.P.; Taylor, B.C.; McLellan, J.S.; et al. Beyond Shielding: The Roles of Glycans in SARS-CoV-2 Spike Protein. ACS Cent. Sci. 2020, 6, 1722–1734. [Google Scholar] [CrossRef] [PubMed]
- Henderson, R.; Edwards, R.J.; Mansouri, K.; Janowska, K.; Stalls, V.; Gobeil, S.M.C.; Kopp, M.; Li, D.; Parks, R.; Hsu, A.L.; et al. Controlling the SARS-CoV-2 spike glycoprotein conformation. Nat. Struct. Mol. Biol. 2020, 27, 925–933. [Google Scholar] [CrossRef]
- Barreto-Vieira, D.F.; da Silva, M.A.N.; de Almeida, A.L.T.; Rasinhas, A.D.C.; Monteiro, M.E.; Miranda, M.D.; Motta, F.C.; Siqueira, M.M.; Girard-Dias, W.; Archanjo, B.S.; et al. SARS-CoV-2: Ultrastructural Characterization of Morphogenesis in an In Vitro System. Viruses 2022, 14, 201. [Google Scholar] [CrossRef]
- Gorkhali, R.; Koirala, P.; Rijal, S.; Mainali, A.; Baral, A.; Bhattarai, H.K. Structure and Function of Major SARS-CoV-2 and SARS-CoV Proteins. Bioinform. Biol. Insights 2021, 15, 11779322211025876. [Google Scholar] [CrossRef]
- Bianchi, M.; Benvenuto, D.; Giovanetti, M.; Angeletti, S.; Ciccozzi, M.; Pascarella, S. Sars-CoV-2 Envelope and Membrane Proteins: Structural Differences Linked to Virus Characteristics? Biomed. Res. Int. 2020, 2020, 4389089. [Google Scholar] [CrossRef]
- Wong, N.A.; Saier, M.H., Jr. The SARS-Coronavirus Infection Cycle: A Survey of Viral Membrane Proteins, Their Functional Interactions and Pathogenesis. Int. J. Mol. Sci. 2021, 22, 1308. [Google Scholar] [CrossRef]
- Alharbi, S.N.; Alrefaei, A.F. Comparison of the SARS-CoV-2 (2019-nCoV) M protein with its counterparts of SARS-CoV and MERS-CoV species. J. King Saud. Univ. Sci. 2021, 33, 101335. [Google Scholar] [CrossRef]
- Thomas, S. The Structure of the Membrane Protein of SARS-CoV-2 Resembles the Sugar Transporter SemiSWEET. Pathog. Immun. 2020, 5, 342–363. [Google Scholar] [CrossRef] [PubMed]
- Schoeman, D.; Fielding, B.C. Is There a Link Between the Pathogenic Human Coronavirus Envelope Protein and Immunopathology? A Review of the Literature. Front. Microbiol. 2020, 11, 2086. [Google Scholar] [CrossRef] [PubMed]
- Chai, J.; Cai, Y.; Pang, C.; Wang, L.; McSweeney, S.; Shanklin, J.; Liu, Q. Structural basis for SARS-CoV-2 envelope protein recognition of human cell junction protein PALS1. Nat. Commun. 2021, 12, 3433. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.S.; Hoque, M.N.; Islam, M.R.; Islam, I.; Mishu, I.D.; Rahaman, M.M.; Sultana, M.; Hossain, M.A. Mutational insights into the envelope protein of SARS-CoV-2. Gene Rep. 2021, 22, 100997. [Google Scholar] [CrossRef]
- Yalcinkaya, M.; Liu, W.; Islam, M.N.; Kotini, A.G.; Gusarova, G.A.; Fidler, T.P.; Papapetrou, E.P.; Bhattacharya, J.; Wang, N.; Tall, A.R. Modulation of the NLRP3 inflammasome by Sars-CoV-2 Envelope protein. Sci. Rep. 2021, 11, 24432. [Google Scholar] [CrossRef]
- Li, J.; Gong, X.; Wang, Z.; Chen, R.; Li, T.; Zeng, D.; Li, M. Clinical features of familial clustering in patients infected with 2019 novel coronavirus in Wuhan, China. Virus Res. 2020, 286, 198043. [Google Scholar] [CrossRef]
- Gao, T.; Gao, Y.; Liu, X.; Nie, Z.; Sun, H.; Lin, K.; Peng, H.; Wang, S. Identification and functional analysis of the SARS-COV-2 nucleocapsid protein. BMC Microbiol. 2021, 21, 58. [Google Scholar] [CrossRef]
- Chen, H.; Cui, Y.; Han, X.; Hu, W.; Sun, M.; Zhang, Y.; Wang, P.H.; Song, G.; Chen, W.; Lou, J. Liquid-liquid phase separation by SARS-CoV-2 nucleocapsid protein and RNA. Cell Res. 2020, 30, 1143–1145. [Google Scholar] [CrossRef]
- Savastano, A.; Ibáñez de Opakua, A.; Rankovic, M.; Zweckstetter, M. Nucleocapsid protein of SARS-CoV-2 phase separates into RNA-rich polymerase-containing condensates. Nat. Commun. 2020, 11, 6041. [Google Scholar] [CrossRef]
- Iserman, C.; Roden, C.A.; Boerneke, M.A.; Sealfon, R.S.G.; McLaughlin, G.A.; Jungreis, I.; Fritch, E.J.; Hou, Y.J.; Ekena, J.; Weidmann, C.A.; et al. Genomic RNA Elements Drive Phase Separation of the SARS-CoV-2 Nucleocapsid. Mol. Cell 2020, 80, 1078–1091. [Google Scholar] [CrossRef]
- Cubuk, J.; Alston, J.J.; Incicco, J.J.; Singh, S.; Stuchell-Brereton, M.D.; Ward, M.D.; Zimmerman, M.I.; Vithani, N.; Griffith, D.; Wagoner, J.A.; et al. The SARS-CoV-2 nucleocapsid protein is dynamic, disordered, and phase separates with RNA. Nat. Commun. 2021, 12, 1936. [Google Scholar] [CrossRef] [PubMed]
- Mu, J.; Fang, Y.; Yang, Q.; Shu, T.; Wang, A.; Huang, M.; Jin, L.; Deng, F.; Qiu, Y.; Zhou, X. SARS-CoV-2 N protein antagonizes type I interferon signaling by suppressing phosphorylation and nuclear translocation of STAT1 and STAT2. Cell Discov. 2020, 6, 65. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Tang, W.; Wang, J.; Lai, D.; Xu, Q.; Huang, R.; Hu, Y.; Gong, X.; Fan, J.; Shu, Q.; et al. SARS-CoV-2 N Protein Induces Acute Lung Injury in Mice via NF-ĸB Activation. Front. Immunol. 2021, 12, 791753. [Google Scholar] [CrossRef] [PubMed]
- Bai, Z.; Cao, Y.; Liu, W.; Li, J. The SARS-CoV-2 Nucleocapsid Protein and Its Role in Viral Structure, Biological Functions, and a Potential Target for Drug or Vaccine Mitigation. Viruses 2021, 13, 1115. [Google Scholar] [CrossRef]
- Shi, M.; Wang, L.; Fontana, P.; Vora, S.; Zhang, Y.; Fu, T.M.; Lieberman, J.; Wu, H. SARS-CoV-2 Nsp1 suppresses host but not viral translation through a bipartite mechanism. bioRxiv 2020. [Google Scholar] [CrossRef]
- Davies, J.P.; Almasy, K.M.; McDonald, E.F.; Plate, L. Comparative Multiplexed Interactomics of SARS-CoV-2 and Homologous Coronavirus Nonstructural Proteins Identifies Unique and Shared Host-Cell Dependencies. ACS Infect. Dis. 2020, 6, 3174–3189. [Google Scholar] [CrossRef]
- Gupta, M.; Azumaya, C.M.; Moritz, M.; Pourmal, S.; Diallo, A.; Merz, G.E.; Jang, G.; Bouhaddou, M.; Fossati, A.; Brilot, A.F.; et al. CryoEM and AI reveal a structure of SARS-CoV-2 Nsp2, a multifunctional protein involved in key host processes. bioRxiv 2021. [Google Scholar] [CrossRef]
- Moustaqil, M.; Ollivier, E.; Chiu, H.P.; Van Tol, S.; Rudolffi-Soto, P.; Stevens, C.; Bhumkar, A.; Hunter, D.J.B.; Freiberg, A.N.; Jacques, D.; et al. SARS-CoV-2 proteases PLpro and 3CLpro cleave IRF3 and critical modulators of inflammatory pathways (NLRP12 and TAB1): Implications for disease presentation across species. Emerg. Microbes Infect. 2021, 10, 178–195. [Google Scholar] [CrossRef]
- Armstrong, L.A.; Lange, S.M.; Dee Cesare, V.; Matthews, S.P.; Nirujogi, R.S.; Cole, I.; Hope, A.; Cunningham, F.; Toth, R.; Mukherjee, R.; et al. Biochemical characterization of protease activity of Nsp3 from SARS-CoV-2 and its inhibition by nanobodies. PLoS ONE 2021, 16, e0253364. [Google Scholar] [CrossRef]
- Khan, M.T.; Zeb, M.T.; Ahsan, H.; Ahmed, A.; Ali, A.; Akhtar, K.; Malik, S.I.; Cui, Z.; Ali, S.; Khan, A.S.; et al. SARS-CoV-2 nucleocapsid and Nsp3 binding: An in silico study. Arch. Microbiol. 2021, 203, 59–66. [Google Scholar] [CrossRef]
- Li, S.; Kong, L.; Yu, X. The expanding roles of endoplasmic reticulum stress in virus replication and pathogenesis. Crit. Rev. Microbiol. 2015, 41, 150–164. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Qin, C.; Rao, Y.; Ngo, C.; Feng, J.J.; Zhao, J.; Zhang, S.; Wang, T.Y.; Carriere, J.; Savas, A.C.; et al. SARS-CoV-2 Nsp5 Demonstrates Two Distinct Mechanisms Targeting RIG-I and MAVS To Evade the Innate Immune Response. mBio 2021, 12, e0233521. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Qiao, J.; You, Q.; Zong, S.; Peng, Q.; Liu, Y.; Hu, S.; Liu, W.; Li, S.; Shu, X.; et al. SARS-CoV-2 Nsp5 Activates NF-κB Pathway by Upregulating SUMOylation of MAVS. Front. Immunol. 2021, 12, 750969. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Liu, Y.; Huang, Z.; Xu, W.; Hu, W.; Yi, L.; Liu, Z.; Chan, H.; Zeng, J.; Liu, X.; et al. SARS-CoV-2 non-structural protein 6 triggers NLRP3-dependent pyroptosis by targeting ATP6AP1. Cell Death Differ. 2022, 1–15. [Google Scholar] [CrossRef]
- Wang, R.; Chen, J.; Hozumi, Y.; Yin, C.; Wei, G.-W. Decoding Asymptomatic COVID-19 Infection and Transmission. J. Phys. Chem. Lett. 2020, 11, 10007–10015. [Google Scholar] [CrossRef]
- Peng, Q.; Peng, R.; Yuan, B.; Zhao, J.; Wang, M.; Wang, X.; Wang, Q.; Sun, Y.; Fan, Z.; Qi, J.; et al. Structural and Biochemical Characterization of the nsp12-nsp7-nsp8 Core Polymerase Complex from SARS-CoV-2. Cell Rep. 2020, 31, 107774. [Google Scholar] [CrossRef]
- Slanina, H.; Madhugiri, R.; Bylapudi, G.; Schultheiß, K.; Karl, N.; Gulyaeva, A.; Gorbalenya Alexander, E.; Linne, U.; Ziebuhr, J. Coronavirus replication–transcription complex: Vital and selective NMPylation of a conserved site in nsp9 by the NiRAN-RdRp subunit. Proc. Natl. Acad. Sci. USA 2021, 118, e2022310118. [Google Scholar] [CrossRef]
- Baddock, H.T.; Brolih, S.; Yosaatmadja, Y.; Ratnaweera, M.; Bielinski, M.; Swift, L.P.; Cruz-Migoni, A.; Fan, H.; Keown, J.R.; Walker, A.P.; et al. Characterization of the SARS-CoV-2 ExoN (nsp14ExoN-nsp10) complex: Implications for its role in viral genome stability and inhibitor identification. Nucleic Acids Res. 2022, 50, 1484–1500. [Google Scholar] [CrossRef]
- Yuen, C.K.; Lam, J.Y.; Wong, W.M.; Mak, L.F.; Wang, X.; Chu, H.; Cai, J.P.; Jin, D.Y.; To, K.K.; Chan, J.F.; et al. SARS-CoV-2 nsp13, nsp14, nsp15 and orf6 function as potent interferon antagonists. Emerg. Microbes Infect. 2020, 9, 1418–1428. [Google Scholar] [CrossRef]
- Lin, S.; Chen, H.; Ye, F.; Chen, Z.; Yang, F.; Zheng, Y.; Cao, Y.; Qiao, J.; Yang, S.; Lu, G. Crystal structure of SARS-CoV-2 nsp10/nsp16 2′-O-methylase and its implication on antiviral drug design. Signal Transduct. Target. Ther. 2020, 5, 131. [Google Scholar] [CrossRef]
- Rosas-Lemus, M.; Minasov, G.; Shuvalova, L.; Inniss, N.L.; Kiryukhina, O.; Brunzelle, J.; Satchell, K.J.F. High-resolution structures of the SARS-CoV-2 2’-O-methyltransferase reveal strategies for structure-based inhibitor design. Sci. Signal. 2020, 13, eabe1202. [Google Scholar] [CrossRef] [PubMed]
- Gadhave, K.; Kumar, P.; Kumar, A.; Bhardwaj, T.; Garg, N.; Giri, R. NSP 11 of SARS-CoV-2 is an Intrinsically Disordered Protein. bioRxiv 2020. [Google Scholar] [CrossRef]
- Xu, G.; Li, Y.; Zhang, S.; Peng, H.; Wang, Y.; Li, D.; Jin, T.; He, Z.; Tong, Y.; Qi, C.; et al. SARS-CoV-2 promotes RIPK1 activation to facilitate viral propagation. Cell Res. 2021, 31, 1230–1243. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Malone, B.; Llewellyn, E.; Grasso, M.; Shelton, P.M.M.; Olinares, P.D.B.; Maruthi, K.; Eng, E.T.; Vatandaslar, H.; Chait, B.T.; et al. Structural Basis for Helicase-Polymerase Coupling in the SARS-CoV-2 Replication-Transcription Complex. Cell 2020, 182, 1560–1573.e1513. [Google Scholar] [CrossRef] [PubMed]
- Spratt, A.N.; Gallazzi, F.; Quinn, T.P.; Lorson, C.L.; Sönnerborg, A.; Singh, K. Coronavirus helicases: Attractive and unique targets of antiviral drug-development and therapeutic patents. Expert. Opin. Ther. Pat. 2021, 31, 339–350. [Google Scholar] [CrossRef]
- Deng, X.; Baker, S.C. An “Old” protein with a new story: Coronavirus endoribonuclease is important for evading host antiviral defenses. Virology 2018, 517, 157–163. [Google Scholar] [CrossRef]
- Hackbart, M.; Deng, X.; Baker, S.C. Coronavirus endoribonuclease targets viral polyuridine sequences to evade activating host sensors. Proc. Natl. Acad. Sci. USA 2020, 117, 8094–8103. [Google Scholar] [CrossRef] [Green Version]
- Silvas, J.A.; Vasquez, D.M.; Park, J.G.; Chiem, K.; Allué-Guardia, A.; Garcia-Vilanova, A.; Platt, R.N.; Miorin, L.; Kehrer, T.; Cupic, A.; et al. Contribution of SARS-CoV-2 Accessory Proteins to Viral Pathogenicity in K18 Human ACE2 Transgenic Mice. J. Virol. 2021, 95, e0040221. [Google Scholar] [CrossRef]
- Su, C.-M.; Wang, L.; Yoo, D. Activation of NF-κB and induction of proinflammatory cytokine expressions mediated by ORF7a protein of SARS-CoV-2. Sci. Rep. 2021, 11, 13464. [Google Scholar] [CrossRef]
- Flower, T.G.; Buffalo, C.Z.; Hooy, R.M.; Allaire, M.; Ren, X.; Hurley, J.H. Structure of SARS-CoV-2 ORF8, a rapidly evolving immune evasion protein. Proc. Natl. Acad. Sci. USA 2021, 118, e2021785118. [Google Scholar] [CrossRef]
- Murdin, A.D.; Barreto, L.; Plotkin, S. Inactivated poliovirus vaccine: Past and present experience. Vaccine 1996, 14, 735–746. [Google Scholar] [CrossRef]
- Vetter, V.; Denizer, G.; Friedland, L.R.; Krishnan, J.; Shapiro, M. Understanding modern-day vaccines: What you need to know. Ann. Med. 2018, 50, 110–120. [Google Scholar] [CrossRef] [PubMed]
- Pollard, A.J.; Bijker, E.M. A guide to vaccinology: From basic principles to new developments. Nat. Rev. Immunol. 2021, 21, 83–100. [Google Scholar] [CrossRef] [PubMed]
- Xia, S.; Zhang, Y.; Wang, Y.; Wang, H.; Yang, Y.; Gao, G.F.; Tan, W.; Wu, G.; Xu, M.; Lou, Z.; et al. Safety and immunogenicity of an inactivated SARS-CoV-2 vaccine, BBIBP-CorV: A randomised, double-blind, placebo-controlled, phase 1/2 trial. Lancet Infect. Dis. 2021, 21, 39–51. [Google Scholar] [CrossRef]
- Hotez, P.J.; Bottazzi, M.E. Whole Inactivated Virus and Protein-Based COVID-19 Vaccines. Annu. Rev. Med. 2022, 73, 55–64. [Google Scholar] [CrossRef]
- Li, X.N.; Huang, Y.; Wang, W.; Jing, Q.L.; Zhang, C.H.; Qin, P.Z.; Guan, W.J.; Gan, L.; Li, Y.L.; Liu, W.H.; et al. Effectiveness of inactivated SARS-CoV-2 vaccines against the Delta variant infection in Guangzhou: A test-negative case-control real-world study. Emerg. Microbes Infect. 2021, 10, 1751–1759. [Google Scholar] [CrossRef]
- Sapkal, G.N.; Yadav, P.D.; Ella, R.; Deshpande, G.R.; Sahay, R.R.; Gupta, N.; Vadrevu, K.M.; Abraham, P.; Panda, S.; Bhargava, B. Inactivated COVID-19 vaccine BBV152/COVAXIN effectively neutralizes recently emerged B.1.1.7 variant of SARS-CoV-2. J. Travel Med. 2021, 28, taab051. [Google Scholar] [CrossRef]
- Desai, D.; Khan, A.R.; Soneja, M.; Mittal, A.; Naik, S.; Kodan, P.; Mandal, A.; Maher, G.T.; Kumar, R.; Agarwal, A.; et al. Effectiveness of an inactivated virus-based SARS-CoV-2 vaccine, BBV152, in India: A test-negative, case-control study. Lancet Infect. Dis. 2022, 22, 349–356. [Google Scholar] [CrossRef]
- WHO. Status of COVID-19 Vaccines within WHO EUL/PQ Evaluation Process; WHO: Geneva, Switzerland, 2021. [Google Scholar]
- Iglesias, E. Inactivated vaccines: A promising old tool against COVID-19. Res. Rev. Insights 2020, 4, 1–4. [Google Scholar] [CrossRef]
- Iversen, P.L.; Bavari, S. Inactivated COVID-19 vaccines to make a global impact. Lancet Infect. Dis. 2021, 21, 746–748. [Google Scholar] [CrossRef]
- Karch, C.P.; Burkhard, P. Vaccine technologies: From whole organisms to rationally designed protein assemblies. Biochem. Pharmacol. 2016, 120, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Tarke, A.; Coelho, C.H.; Zhang, Z.; Dan, J.M.; Yu, E.D.; Methot, N.; Bloom, N.I.; Goodwin, B.; Phillips, E.; Mallal, S.; et al. SARS-CoV-2 vaccination induces immunological T cell memory able to cross-recognize variants from Alpha to Omicron. Cell 2022, 185, 847–859.e811. [Google Scholar] [CrossRef] [PubMed]
- Heath, P.T.; Galiza, E.P.; Baxter, D.N.; Boffito, M.; Browne, D.; Burns, F.; Chadwick, D.R.; Clark, R.; Cosgrove, C.; Galloway, J.; et al. Safety and Efficacy of NVX-CoV2373 COVID-19 Vaccine. N. Engl. J. Med. 2021, 385, 1172–1183. [Google Scholar] [CrossRef] [PubMed]
- Chavda, V.P.; Hossain, M.K.; Beladiya, J.; Apostolopoulos, V. Nucleic Acid Vaccines for COVID-19: A Paradigm Shift in the Vaccine Development Arena. Biologics 2021, 1, 337–356. [Google Scholar] [CrossRef]
- Voysey, M.; Clemens, S.A.C.; Madhi, S.A.; Weckx, L.Y.; Folegatti, P.M.; Aley, P.K.; Angus, B.; Baillie, V.L.; Barnabas, S.L.; Bhorat, Q.E.; et al. Safety and efficacy of the ChAdOx1 nCoV-19 vaccine (AZD1222) against SARS-CoV-2: An interim analysis of four randomised controlled trials in Brazil, South Africa, and the UK. Lancet 2021, 397, 99–111. [Google Scholar] [CrossRef]
- Sadoff, J.; Gray, G.; Vandebosch, A.; Cárdenas, V.; Shukarev, G.; Grinsztejn, B.; Goepfert, P.A.; Truyers, C.; Fennema, H.; Spiessens, B.; et al. Safety and Efficacy of Single-Dose Ad26.COV2.S Vaccine against COVID-19. N. Engl. J. Med. 2021, 384, 2187–2201. [Google Scholar] [CrossRef]
- Jones, I.; Roy, P. Sputnik V COVID-19 vaccine candidate appears safe and effective. Lancet 2021, 397, 642–643. [Google Scholar] [CrossRef]
- Karikó, K.; Buckstein, M.; Ni, H.; Weissman, D. Suppression of RNA recognition by Toll-like receptors: The impact of nucleoside modification and the evolutionary origin of RNA. Immunity 2005, 23, 165–175. [Google Scholar] [CrossRef] [Green Version]
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Pérez Marc, G.; Moreira, E.D.; Zerbini, C.; et al. Safety and Efficacy of the BNT162b2 mRNA COVID-19 Vaccine. N. Engl. J. Med. 2020, 383, 2603–2615. [Google Scholar] [CrossRef]
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N. Engl. J. Med. 2021, 384, 403–416. [Google Scholar] [CrossRef]
- Wesselhoeft, R.A.; Kowalski, P.S.; Anderson, D.G. Engineering circular RNA for potent and stable translation in eukaryotic cells. Nat. Commun. 2018, 9, 2629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, X.; Lu, Y. Circular RNA: Biosynthesis in vitro. Front. Bioeng. Biotechnol. 2021, 9, 787881. [Google Scholar] [CrossRef] [PubMed]
- Wesselhoeft, R.A.; Kowalski, P.S.; Parker-Hale, F.C.; Huang, Y.; Bisaria, N.; Anderson, D.G. RNA Circularization Diminishes Immunogenicity and Can Extend Translation Duration In Vivo. Mol. Cell 2019, 74, 508–520. [Google Scholar] [CrossRef] [PubMed]
- Qu, L.; Yi, Z.; Shen, Y.; Lin, L.; Chen, F.; Xu, Y.; Wu, Z.; Tang, H.; Zhang, X.; Tian, F.; et al. Circular RNA vaccines against SARS-CoV-2 and emerging variants. Cell 2022, 185, 1728–1744.e16. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhan, W.; Yang, Z.; Tu, C.; Hu, G.; Zhang, X.; Song, W.; Du, S.; Zhu, Y.; Huang, K.; et al. Broad neutralization of SARS-CoV-2 variants by an inhalable bispecific single-domain antibody. Cell 2022, 185, 1389–1401.e1318. [Google Scholar] [CrossRef]
- Chavda, V.P.; Vora, L.K.; Pandya, A.K.; Patravale, V.B. Intranasal vaccines for SARS-CoV-2: From challenges to potential in COVID-19 management. Drug Discov. Today 2021, 26, 2619–2636. [Google Scholar] [CrossRef]
- Dhama, K.; Dhawan, M.; Tiwari, R.; Emran, T.B.; Mitra, S.; Rabaan, A.A.; Alhumaid, S.; Alawi, Z.A.; Al Mutair, A. COVID-19 intranasal vaccines: Current progress, advantages, prospects, and challenges. Hum. Vaccin Immunother. 2022, 1–11. [Google Scholar] [CrossRef]
- Hoseini-Tavassol, Z.; Ejtahed, H.S.; Soroush, A.R.; Sajjadpour, Z.; Hasani-Ranjbar, S.; Larijani, B. Natural Derived Nasal Spray; A Proposed Approach for COVID-19 Disease Control. Infect. Disord. Drug Targets 2021, 21, e160921191568. [Google Scholar] [CrossRef]
- Hassan, A.O.; Kafai, N.M.; Dmitriev, I.P.; Fox, J.M.; Smith, B.K.; Harvey, I.B.; Chen, R.E.; Winkler, E.S.; Wessel, A.W.; Case, J.B.; et al. A Single-Dose Intranasal ChAd Vaccine Protects Upper and Lower Respiratory Tracts against SARS-CoV-2. Cell 2020, 183, 169–184.e113. [Google Scholar] [CrossRef]
- He, J.; Huang, J.-R.; Zhang, Y.-L.; Zhang, J. SARS-CoV-2 nucleocapsid protein intranasal inoculation induces local and systemic T cell responses in mice. J. Med. Virol. 2021, 93, 1923–1925. [Google Scholar] [CrossRef]
- Westendorf, K.; Žentelis, S.; Wang, L.; Foster, D.; Vaillancourt, P.; Wiggin, M.; Lovett, E.; van der Lee, R.; Hendle, J.; Pustilnik, A.; et al. LY-CoV1404 (bebtelovimab) potently neutralizes SARS-CoV-2 variants. bioRxiv 2022. [Google Scholar] [CrossRef] [PubMed]
- Dougan, M.; Azizad, M.; Chen, P.; Feldman, B.; Frieman, M.; Igbinadolor, A.; Kumar, P.; Morris, J.; Potts, J.; Baracco, L.; et al. Bebtelovimab, alone or together with bamlanivimab and etesevimab, as a broadly neutralizing monoclonal antibody treatment for mild to moderate, ambulatory COVID-19. medRxiv 2022. [Google Scholar] [CrossRef]
- Wu, X.; Cheng, L.; Fu, M.; Huang, B.; Zhu, L.; Xu, S.; Shi, H.; Zhang, D.; Yuan, H.; Nawaz, W.; et al. A potent bispecific nanobody protects hACE2 mice against SARS-CoV-2 infection via intranasal administration. Cell Rep. 2021, 37, 109869. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Li, W.; Drabek, D.; Okba, N.M.A.; van Haperen, R.; Osterhaus, A.; van Kuppeveld, F.J.M.; Haagmans, B.L.; Grosveld, F.; Bosch, B.J. A human monoclonal antibody blocking SARS-CoV-2 infection. Nat. Commun. 2020, 11, 2251. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Li, R.; Pan, Z.; Qian, C.; Yang, Y.; You, R.; Zhao, J.; Liu, P.; Gao, L.; Li, Z.; et al. Human monoclonal antibodies block the binding of SARS-CoV-2 spike protein to angiotensin converting enzyme 2 receptor. Cell. Mol. Immunol. 2020, 17, 647–649. [Google Scholar] [CrossRef]
- Gavor, E.; Choong, Y.K.; Er, S.Y.; Sivaraman, H.; Sivaraman, J. Structural Basis of SARS-CoV-2 and SARS-CoV Antibody Interactions. Trends Immunol. 2020, 41, 1006–1022. [Google Scholar] [CrossRef]
- Cathcart, A.L.; Havenar-Daughton, C.; Lempp, F.A.; Ma, D.; Schmid, M.A.; Agostini, M.L.; Guarino, B.; Di iulio, J.; Rosen, L.E.; Tucker, H.; et al. The dual function monoclonal antibodies VIR-7831 and VIR-7832 demonstrate potent in vitro and in vivo activity against SARS-CoV-2. bioRxiv 2022. [Google Scholar] [CrossRef]
- Levin, M.J.; Ustianowski, A.; De Wit, S.; Launay, O.; Avila, M.; Templeton, A.; Yuan, Y.; Seegobin, S.; Ellery, A.; Levinson, D.J. Intramuscular AZD7442 (Tixagevimab–Cilgavimab) for Prevention of COVID-19. N. Engl. J. Med. 2022, 386. [Google Scholar] [CrossRef]
- Levin, M.J.; Ustianowski, A.; De Wit, S.; Launay, O.; Avila, M.; Seegobin, S.; Templeton, A.; Yuan, Y.; Ambery, P.; Arends, R.H. LB5. PROVENT: Phase 3 Study of Efficacy and Safety of AZD7442 (Tixagevimab/Cilgavimab) for Pre-exposure Prophylaxis of COVID-19 in Adults. Open Forum Infect. Dis. 2021, 8, S810. [Google Scholar] [CrossRef]
- Fenwick, C.; Turelli, P.; Ni, D.; Perez, L.; Lau, K.; Lana, E.; Pellaton, C.; Raclot, C.; Esteves-Leuenberger, L.; Campos, J. SARS-CoV-2 Omicron potently neutralized by a novel antibody with unique Spike binding properties. bioRxiv 2022. [Google Scholar] [CrossRef]
- Su, S.-C.; Yang, T.-J.; Yu, P.-Y.; Liang, K.-H.; Chen, W.-Y.; Yang, C.-W.; Lin, H.-T.; Wang, M.-J.; Lu, R.-M.; Tso, H.-C.; et al. Structure-guided antibody cocktail for prevention and treatment of COVID-19. PLoS Pathogens 2021, 17, e1009704. [Google Scholar] [CrossRef] [PubMed]
- Liang, K.-H.; Chiang, P.-Y.; Ko, S.-H.; Chou, Y.-C.; Lu, R.-M.; Lin, H.-T.; Chen, W.-Y.; Lin, Y.-L.; Tao, M.-H.; Jan, J.-T.; et al. Antibody cocktail effective against variants of SARS-CoV-2. J. Biomed. Sci. 2021, 28, 80. [Google Scholar] [CrossRef] [PubMed]
- Ku, Z.; Xie, X.; Lin, J.; Gao, P.; El Sahili, A.; Su, H.; Liu, Y.; Ye, X.; Li, X.; Fan, X.; et al. Engineering SARS-CoV-2 cocktail antibodies into a bispecific format improves neutralizing potency and breadth. bioRxiv 2022. [Google Scholar] [CrossRef]
- Zhang, C.; Yang, M. Antimicrobial Peptides: From Design to Clinical Application. Antibiotics 2022, 11, 349. [Google Scholar] [CrossRef]
- Basit, A.; Karim, A.M.; Asif, M.; Ali, T.; Lee, J.H.; Jeon, J.H.; Rehman, S.U.; Lee, S.H. Designing Short Peptides to Block the Interaction of SARS-CoV-2 and Human ACE2 for COVID-19 Therapeutics. Front. Pharmacol. 2021, 12, 731828. [Google Scholar] [CrossRef]
- Karoyan, P.; Vieillard, V.; Gómez-Morales, L.; Odile, E.; Guihot, A.; Luyt, C.E.; Denis, A.; Grondin, P.; Lequin, O. Human ACE2 peptide-mimics block SARS-CoV-2 pulmonary cells infection. Commun. Biol. 2021, 4, 197. [Google Scholar] [CrossRef]
- Cao, L.; Goreshnik, I.; Coventry, B.; Case, J.B.; Miller, L.; Kozodoy, L.; Chen, R.E.; Carter, L.; Walls, A.C.; Park, Y.J.; et al. De novo design of picomolar SARS-CoV-2 miniprotein inhibitors. Science 2020, 370, 426–431. [Google Scholar] [CrossRef]
- Riva, L.; Yuan, S.; Yin, X.; Martin-Sancho, L.; Matsunaga, N.; Pache, L.; Burgstaller-Muehlbacher, S.; De Jesus, P.D.; Teriete, P.; Hull, M.V.; et al. Discovery of SARS-CoV-2 antiviral drugs through large-scale compound repurposing. Nature 2020, 586, 113–119. [Google Scholar] [CrossRef]
- Bung, N.; Krishnan, S.R.; Bulusu, G.; Roy, A. De novo design of new chemical entities for SARS-CoV-2 using artificial intelligence. Future Med. Chem. 2021, 13, 575–585. [Google Scholar] [CrossRef]
- Pushpakom, S.; Iorio, F.; Eyers, P.A.; Escott, K.J.; Hopper, S.; Wells, A.; Doig, A.; Guilliams, T.; Latimer, J.; McNamee, C.; et al. Drug repurposing: Progress, challenges and recommendations. Nat. Rev. Drug Discov. 2019, 18, 41–58. [Google Scholar] [CrossRef]
- Farooq, S.; Ngaini, Z. Natural and Synthetic Drugs as Potential Treatment for Coronavirus Disease 2019 (COVID-2019). Chem. Afr. 2020, 1–13. [Google Scholar] [CrossRef]
- Bojadzic, D.; Alcazar, O.; Chen, J.; Chuang, S.T.; Condor Capcha, J.M.; Shehadeh, L.A.; Buchwald, P. Small-Molecule Inhibitors of the Coronavirus Spike: ACE2 Protein-Protein Interaction as Blockers of Viral Attachment and Entry for SARS-CoV-2. ACS Infect. Dis. 2021, 7, 1519–1534. [Google Scholar] [CrossRef]
- Self, W.H.; Semler, M.W.; Leither, L.M.; Casey, J.D.; Angus, D.C.; Brower, R.G.; Chang, S.Y.; Collins, S.P.; Eppensteiner, J.C.; Filbin, M.R.; et al. Effect of Hydroxychloroquine on Clinical Status at 14 Days in Hospitalized Patients With COVID-19: A Randomized Clinical Trial. JAMA 2020, 324, 2165–2176. [Google Scholar] [CrossRef] [PubMed]
- Tian, L.; Qiang, T.; Liang, C.; Ren, X.; Jia, M.; Zhang, J.; Li, J.; Wan, M.; YuWen, X.; Li, H.; et al. RNA-dependent RNA polymerase (RdRp) inhibitors: The current landscape and repurposing for the COVID-19 pandemic. Eur. J. Med. Chem. 2021, 213, 113201. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.Z.; Li, K.; Maskey, A.R.; Huang, W.; Toutov, A.A.; Yang, N.; Srivastava, K.; Geliebter, J.; Tiwari, R.; Miao, M.; et al. A small molecule compound berberine as an orally active therapeutic candidate against COVID-19 and SARS: A computational and mechanistic study. FASEB J. 2021, 35, e21360. [Google Scholar] [CrossRef]
- Rothlin, R.P.; Vetulli, H.M.; Duarte, M.; Pelorosso, F.G. Telmisartan as tentative angiotensin receptor blocker therapeutic for COVID-19. Drug Dev. Res. 2020, 81, 768–770. [Google Scholar] [CrossRef]
- Mahoney, M.; Damalanka, V.C.; Tartell, M.A.; Chung, D.H.; Lourenco, A.L.; Pwee, D.; Mayer Bridwell, A.E.; Hoffmann, M.; Voss, J.; Karmakar, P.; et al. A novel class of TMPRSS2 inhibitors potently block SARS-CoV-2 and MERS-CoV viral entry and protect human epithelial lung cells. bioRxiv 2021. [Google Scholar] [CrossRef]
- Panoutsopoulos, A.A. Known drugs and small molecules in the battle for COVID-19 treatment. Genes Dis. 2020, 7, 528–534. [Google Scholar] [CrossRef]
- Nguyen, H.L.; Thai, N.Q.; Truong, D.T.; Li, M.S. Remdesivir Strongly Binds to Both RNA-Dependent RNA Polymerase and Main Protease of SARS-CoV-2: Evidence from Molecular Simulations. J. Phys. Chem. B 2020, 124, 11337–11348. [Google Scholar] [CrossRef]
- Patel, T.K.; Patel, P.B.; Barvaliya, M.; Saurabh, M.K.; Bhalla, H.L.; Khosla, P.P. Efficacy and safety of lopinavir-ritonavir in COVID-19: A systematic review of randomized controlled trials. J. Infect. Public Health 2021, 14, 740–748. [Google Scholar] [CrossRef]
- Udwadia, Z.F.; Singh, P.; Barkate, H.; Patil, S.; Rangwala, S.; Pendse, A.; Kadam, J.; Wu, W.; Caracta, C.F.; Tandon, M. Efficacy and safety of favipiravir, an oral RNA-dependent RNA polymerase inhibitor, in mild-to-moderate COVID-19: A randomized, comparative, open-label, multicenter, phase 3 clinical trial. Int. J. Infect. Dis. 2021, 103, 62–71. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wu, Y.; Yao, S.; Ge, H.; Zhu, Y.; Chen, K.; Chen, W.-Z.; Zhang, Y.; Zhu, W.; Wang, H.-Y.; et al. Discovery of potential small molecular SARS-CoV-2 entry blockers targeting the spike protein. Acta Pharmacol. Sin. 2022, 43, 788–796. [Google Scholar] [CrossRef] [PubMed]
- Vangeel, L.; Chiu, W.; De Jonghe, S.; Maes, P.; Slechten, B.; Raymenants, J.; André, E.; Leyssen, P.; Neyts, J.; Jochmans, D. Remdesivir, Molnupiravir and Nirmatrelvir remain active against SARS-CoV-2 Omicron and other variants of concern. Antivir. Res. 2022, 198, 105252. [Google Scholar] [CrossRef] [PubMed]
- Haggag, Y.A.; El-Ashmawy, N.E.; Okasha, K.M. Is hesperidin essential for prophylaxis and treatment of COVID-19 Infection? Med. Hypotheses 2020, 144, 109957. [Google Scholar] [CrossRef] [PubMed]
- Umemura, Y.; Mitsuyama, Y.; Minami, K.; Nishida, T.; Watanabe, A.; Okada, N.; Yamakawa, K.; Nochioka, K.; Fujimi, S. Efficacy and safety of nintedanib for pulmonary fibrosis in severe pneumonia induced by COVID-19: An interventional study. Int. J. Infect. Dis. 2021, 108, 454–460. [Google Scholar] [CrossRef]
- Jayk Bernal, A.; Gomes da Silva, M.M.; Musungaie, D.B.; Kovalchuk, E.; Gonzalez, A.; Delos Reyes, V.; Martín-Quirós, A.; Caraco, Y.; Williams-Diaz, A.; Brown, M.L.; et al. Molnupiravir for Oral Treatment of COVID-19 in Nonhospitalized Patients. N. Engl. J. Med. 2022, 386, 509–520. [Google Scholar] [CrossRef]
- Bahadoram, M.; Keikhaei, B.; Saeedi-Boroujeni, A.; Mahmoudian-Sani, M.R. Chloroquine/hydroxychloroquine: An inflammasome inhibitor in severe COVID-19? Naunyn Schmiedebergs Arch. Pharmacol. 2021, 394, 997–1001. [Google Scholar] [CrossRef]
- Andreani, J.; Le Bideau, M.; Duflot, I.; Jardot, P.; Rolland, C.; Boxberger, M.; Wurtz, N.; Rolain, J.M.; Colson, P.; La Scola, B.; et al. In vitro testing of combined hydroxychloroquine and azithromycin on SARS-CoV-2 shows synergistic effect. Microb. Pathog. 2020, 145, 104228. [Google Scholar] [CrossRef]
- Omrani, A.S.; Pathan, S.A.; Thomas, S.A.; Harris, T.R.E.; Coyle, P.V.; Thomas, C.E.; Qureshi, I.; Bhutta, Z.A.; Mawlawi, N.A.; Kahlout, R.A.; et al. Randomized double-blinded placebo-controlled trial of hydroxychloroquine with or without azithromycin for virologic cure of non-severe COVID-19. EClinicalMedicine 2020, 29, 100645. [Google Scholar] [CrossRef]
- Johnston, C.; Brown, E.R.; Stewart, J.; Karita, H.C.S.; Kissinger, P.J.; Dwyer, J.; Hosek, S.; Oyedele, T.; Paasche-Orlow, M.K.; Paolino, K.; et al. Hydroxychloroquine with or without azithromycin for treatment of early SARS-CoV-2 infection among high-risk outpatient adults: A randomized clinical trial. EClinicalMedicine 2021, 33, 100773. [Google Scholar] [CrossRef]
- Ding, R.; Long, J.; Yuan, M.; Jin, Y.; Yang, H.; Chen, M.; Chen, S.; Duan, G. CRISPR/Cas System: A Potential Technology for the Prevention and Control of COVID-19 and Emerging Infectious Diseases. Front. Cell. Infect. Microbiol. 2021, 11, 639108. [Google Scholar] [CrossRef] [PubMed]
- Safari, F.; Afarid, M.; Rastegari, B.; Borhani-Haghighi, A.; Barekati-Mowahed, M.; Behzad-Behbahani, A. CRISPR systems: Novel approaches for detection and combating COVID-19. Virus Res. 2021, 294, 198282. [Google Scholar] [CrossRef] [PubMed]
- Bayoumi, M.; Munir, M. Potential Use of CRISPR/Cas13 Machinery in Understanding Virus–Host Interaction. Front. Microbiol. 2021, 12, 743580. [Google Scholar] [CrossRef]
- Ullah, M.F.; Ali, Y.; Khan, M.R.; Khan, I.U.; Yan, B.; Ijaz Khan, M.; Malik, M.Y. A review of COVID-19: Treatment strategies and CRISPR/Cas9 gene editing technology approaches to the coronavirus disease. Saudi. J. Biol. Sci. 2022, 29, 860–871. [Google Scholar] [CrossRef]
- Kannan, S.; Altae-Tran, H.; Jin, X.; Madigan, V.J.; Oshiro, R.; Makarova, K.S.; Koonin, E.V.; Zhang, F. Compact RNA editors with small Cas13 proteins. Nat. Biotechnol. 2022, 40, 194–197. [Google Scholar] [CrossRef] [PubMed]
- Kellner, M.J.; Koob, J.G.; Gootenberg, J.S.; Abudayyeh, O.O.; Zhang, F. SHERLOCK: Nucleic acid detection with CRISPR nucleases. Nat. Protoc. 2019, 14, 2986–3012. [Google Scholar] [CrossRef] [PubMed]
- Joung, J.; Ladha, A.; Saito, M.; Segel, M.; Bruneau, R.; Huang, M.W.; Kim, N.G.; Yu, X.; Li, J.; Walker, B.D.; et al. Point-of-care testing for COVID-19 using SHERLOCK diagnostics. medRxiv 2020. [Google Scholar] [CrossRef]
- Kaminski, M.M.; Abudayyeh, O.O.; Gootenberg, J.S.; Zhang, F.; Collins, J.J. CRISPR-based diagnostics. Nat. Biomed. Eng. 2021, 5, 643–656. [Google Scholar] [CrossRef]
- Li, P.Y.; Li, S.Q.; Gao, S.G.; Dong, D.Y. A one-step platform for screening high-efficient and minimal off-target CRISPR/Cas13 crRNAs to eradicate SARS-CoV-2 virus for treatment of COVID-19 patients. Med. Hypotheses 2022, 159, 110754. [Google Scholar] [CrossRef]
- Dixon, B.E.; Wools-Kaloustian, K.; Fadel, W.F.; Duszynski, T.J.; Yiannoutsos, C.; Halverson, P.K.; Menachemi, N. Symptoms and symptom clusters associated with SARS-CoV-2 infection in community-based populations: Results from a statewide epidemiological study. medRxiv 2020. [Google Scholar] [CrossRef]
- Hu, B.; Huang, S.; Yin, L. The cytokine storm and COVID-19. J. Med. Virol. 2021, 93, 250–256. [Google Scholar] [CrossRef] [PubMed]
- Pelaia, C.; Calabrese, C.; Garofalo, E.; Bruni, A.; Vatrella, A.; Pelaia, G. Therapeutic Role of Tocilizumab in SARS-CoV-2-Induced Cytokine Storm: Rationale and Current Evidence. Int. J. Mol. Sci. 2021, 22, 3059. [Google Scholar] [CrossRef] [PubMed]
- Kontoghiorghes, G.J.; Kolnagou, A.; Fetta, S.; Kontoghiorghe, C.N. Conventional and Unconventional Approaches for Innovative Drug Treatments in COVID-19: Looking Outside of Plato’s Cave. Int. J. Mol. Sci. 2021, 22, 7208. [Google Scholar] [CrossRef]
- Asrani, P.; Hassan, M.I. SARS-CoV-2 mediated lung inflammatory responses in host: Targeting the cytokine storm for therapeutic interventions. Mol. Cell. Biochem. 2021, 476, 675–687. [Google Scholar] [CrossRef] [PubMed]
- Feng, L.; Qi, Q.; Wang, P.; Chen, H.; Chen, Z.; Meng, Z.; Liu, L. Serum levels of IL-6, IL-8, and IL-10 are indicators of prognosis in pancreatic cancer. J. Int. Med. Res. 2018, 46, 5228–5236. [Google Scholar] [CrossRef] [Green Version]
- Gritti, G.; Raimondi, F.; Ripamonti, D.; Riva, I.; Landi, F.; Alborghetti, L.; Frigeni, M.; Damiani, M.; Micò, C.; Fagiuoli, S.; et al. IL-6 signalling pathway inactivation with siltuximab in patients with COVID-19 respiratory failure: An observational cohort study. medRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Investigators, T.R.-C.; Derde, L.P.G. Effectiveness of Tocilizumab, Sarilumab, and Anakinra for critically ill patients with COVID-19 The REMAP-CAP COVID-19 Immune Modulation Therapy Domain Randomized Clinical Trial. medRxiv 2021. [Google Scholar] [CrossRef]
- Atal, S.; Fatima, Z. IL-6 Inhibitors in the Treatment of Serious COVID-19: A Promising Therapy? Pharm. Med. 2020, 34, 223–231. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, Y.; Vilekar, P.; Yang, S.P.; Gupta, M.; Oh, M.I.; Meek, A.; Doyle, L.; Villar, L.; Brennecke, A.; et al. Small molecule therapeutics for COVID-19: Repurposing of inhaled furosemide. PeerJ 2020, 8, e9533. [Google Scholar] [CrossRef]
- Brennecke, A.; Villar, L.; Wang, Z.; Doyle, L.M.; Meek, A.; Reed, M.; Barden, C.; Weaver, D.F. Is Inhaled Furosemide a Potential Therapeutic for COVID-19? Am. J. Med. Sci. 2020, 360, 216–221. [Google Scholar] [CrossRef]

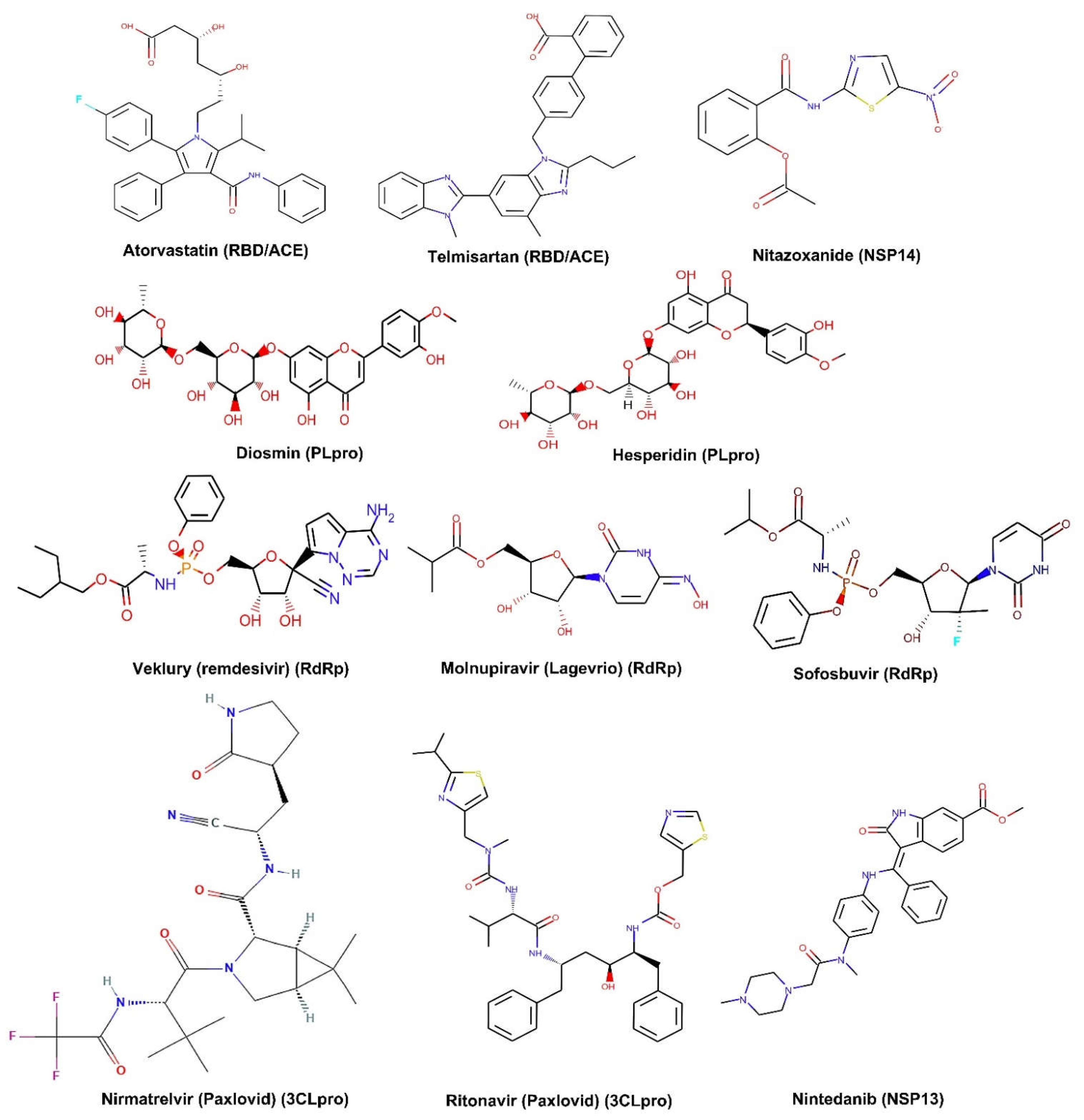
| Protein | Durg Target | Chemical Compound | Clinical Number | Status |
|---|---|---|---|---|
| Spike | RBD/ACE2 blocker | Atorvastatin | NCT04380402 | Phase 2 |
| Telmisartan | NCT04355936 | Completed | ||
| Chloroquine | NCT04303507 | N/A | ||
| NSP3 | Papain-like protease | Diosmin/Hesperidin (ACE2) | NCT04452799 | Early Phase 1 |
| NSP5 | 3CLpro | Nirmatrelvir (Paxlovid) | NCT04960202 NCT05047601 NCT05011513 | FDA approved |
| Ritonavir (Paxlovid) | ||||
| NSP12 | RdRp | Veklury (remdesivir) | NCT04292730 | FDA approved |
| Molnupiravir (Lagevrio) | NCT04575597 | FDA approved for EUA | ||
| Sofosbuvir | NCT04460443 | Phase 2 | ||
| NSP13 | Helicase | Nintedanib | NCT04541680 | Phase 3 |
| NSP14 | Endoribonuclease (3′-5′) | Nitazoxanide | NCT04561219 | Phase 2 |
| NSP15 | Uridine-specific endoribonuclease | Paritaprevir | NCT02504099 | N/A |
| NSP16 | Methyltransferase | Bemcentinib | NCT04890509 | Phase 2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, C.; Yang, M. Newly Emerged Antiviral Strategies for SARS-CoV-2: From Deciphering Viral Protein Structural Function to the Development of Vaccines, Antibodies, and Small Molecules. Int. J. Mol. Sci. 2022, 23, 6083. https://doi.org/10.3390/ijms23116083
Zhang C, Yang M. Newly Emerged Antiviral Strategies for SARS-CoV-2: From Deciphering Viral Protein Structural Function to the Development of Vaccines, Antibodies, and Small Molecules. International Journal of Molecular Sciences. 2022; 23(11):6083. https://doi.org/10.3390/ijms23116083
Chicago/Turabian StyleZhang, Chunye, and Ming Yang. 2022. "Newly Emerged Antiviral Strategies for SARS-CoV-2: From Deciphering Viral Protein Structural Function to the Development of Vaccines, Antibodies, and Small Molecules" International Journal of Molecular Sciences 23, no. 11: 6083. https://doi.org/10.3390/ijms23116083
APA StyleZhang, C., & Yang, M. (2022). Newly Emerged Antiviral Strategies for SARS-CoV-2: From Deciphering Viral Protein Structural Function to the Development of Vaccines, Antibodies, and Small Molecules. International Journal of Molecular Sciences, 23(11), 6083. https://doi.org/10.3390/ijms23116083






