Structural Basis for the Regiospecificity of a Lipase from Streptomyces sp. W007
Abstract
:1. Introduction
2. Results
2.1. Catalytic Pocket of MAS1
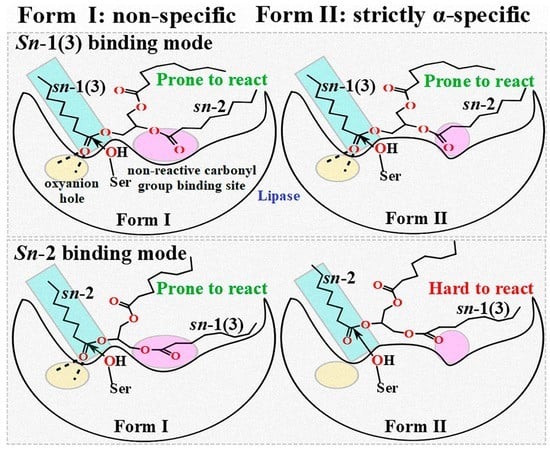
2.2. Comparison of Different Reaction Conformations of TC
2.3. Binding Geometry of TC in the sn-2 Reaction Mode
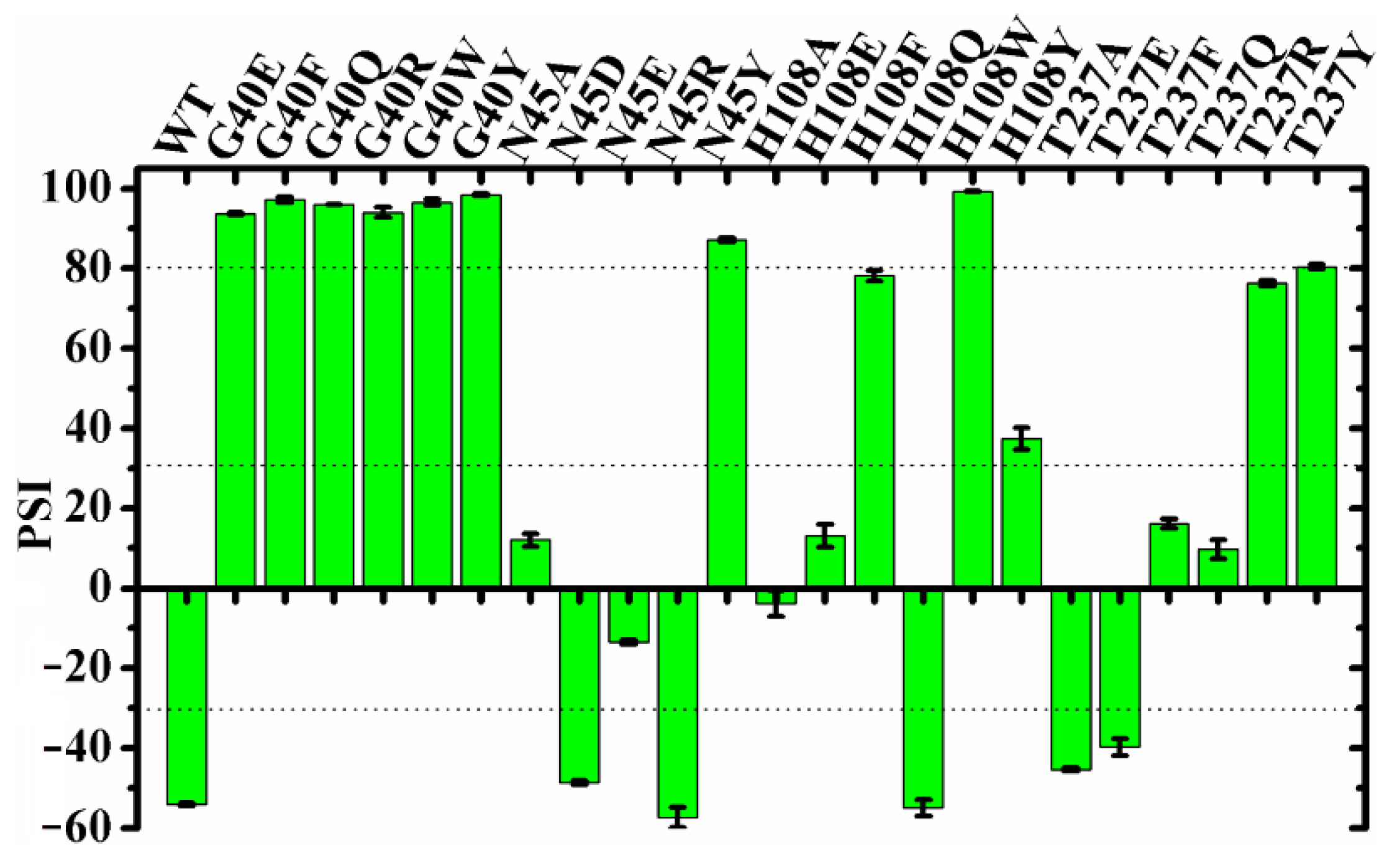
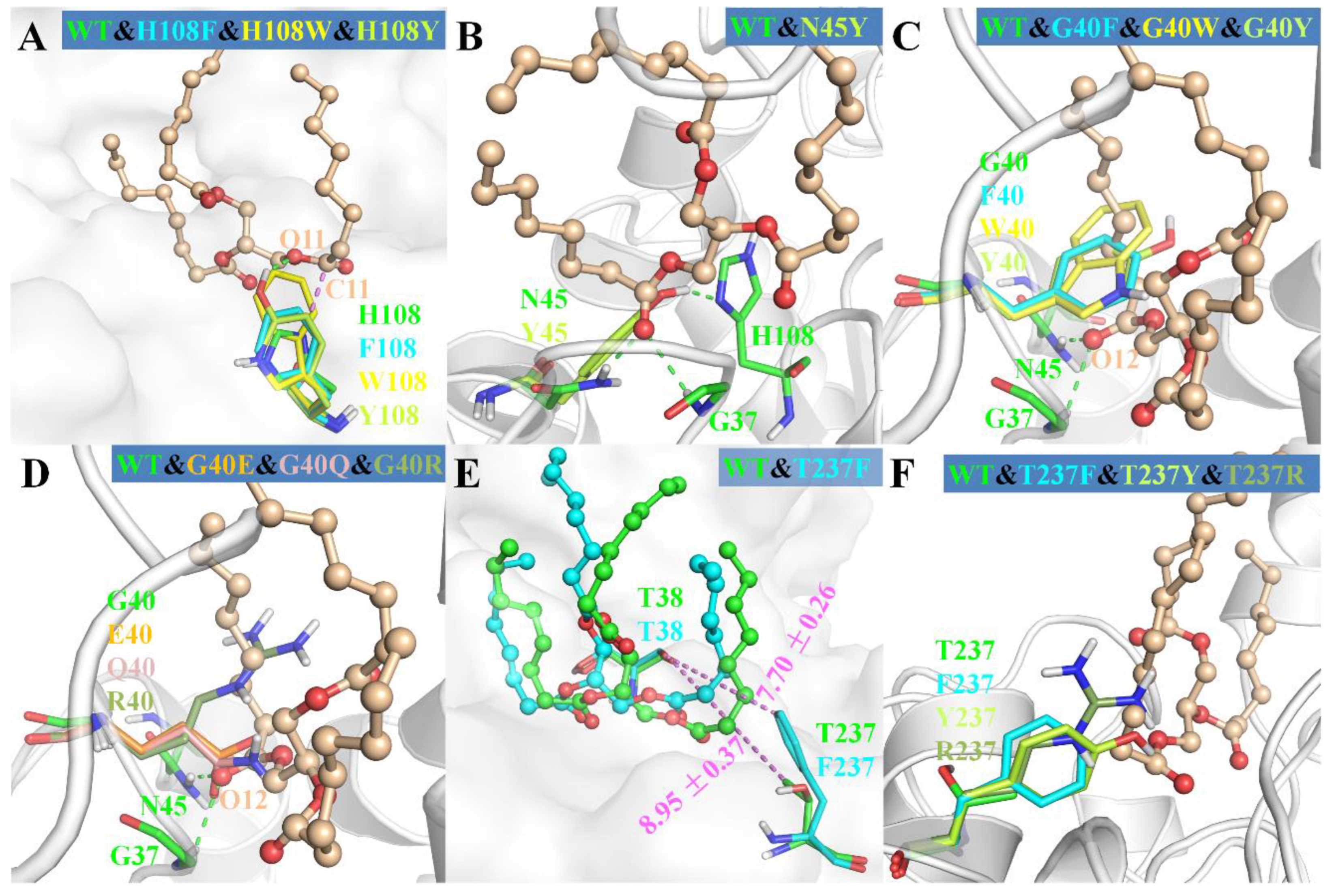
2.4. Regiospecificity of Mutants
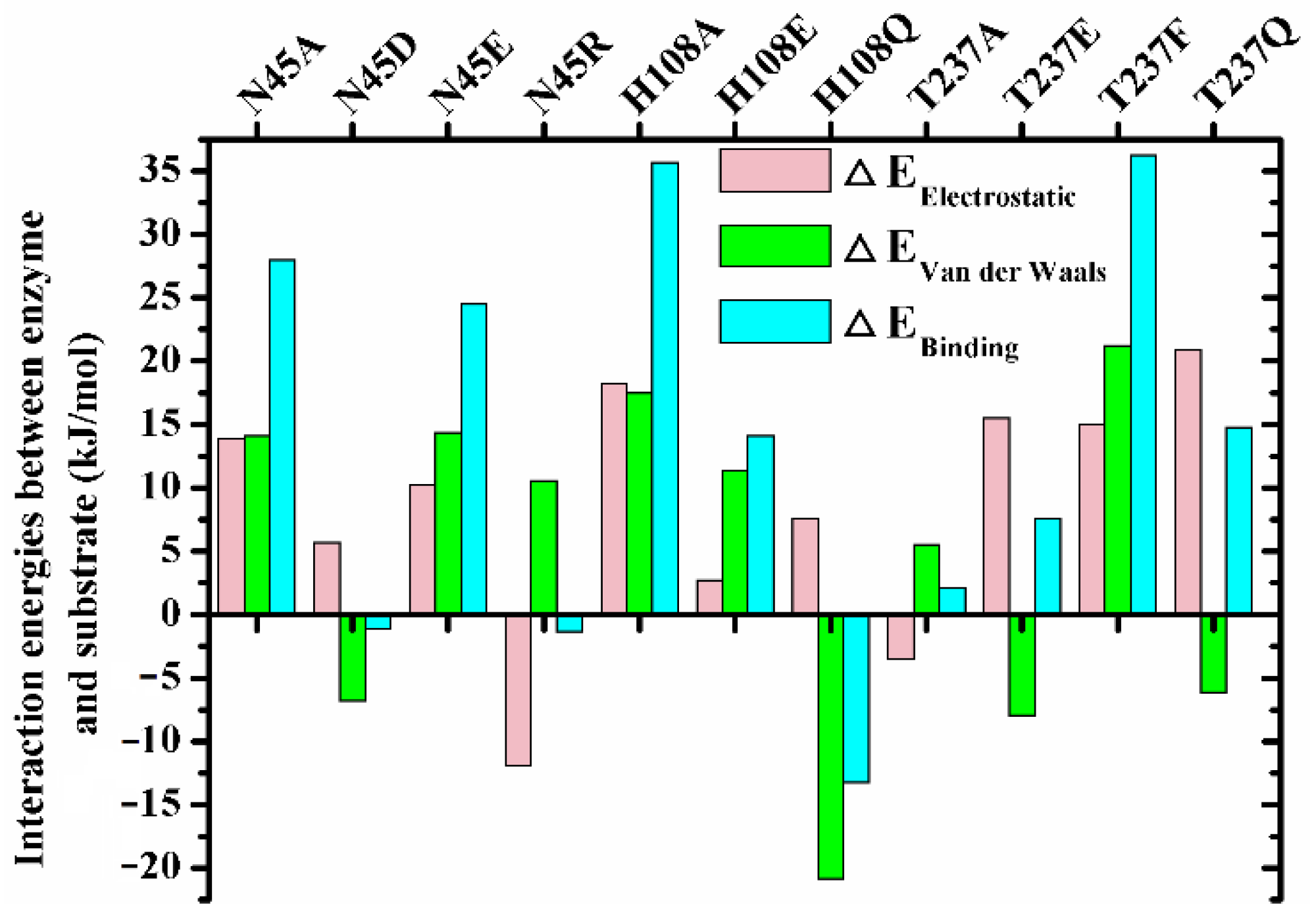
2.5. Substrate Binding Stability
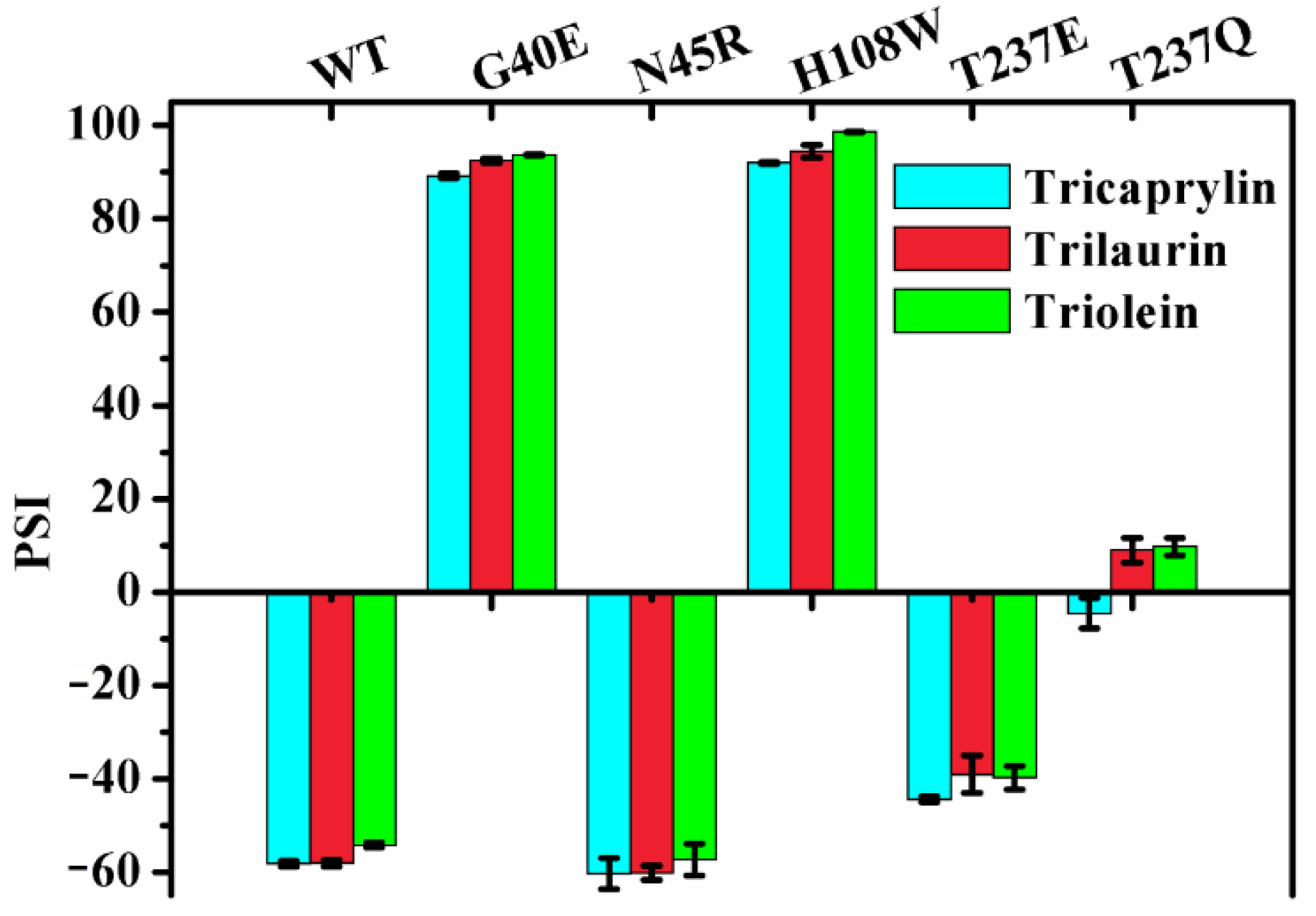
2.6. Effect of Chain Length on the Regiospecificity
3. Discussion
3.1. Role of H108
3.2. Role of N45
3.3. Effect of Substitution of G40 and T237
4. Materials and Methods
4.1. Strains, Plasmids and Materials
4.2. Construction of E. coli Expression Host and Protein Purification
4.3. Mutants Constructing
4.4. Hydrolytic Activity Assay
4.5. Regiospecificity Analysis
4.6. Model Construction of Enzyme-Substrate Complexes
4.7. Preparation of Simulation Systems
4.8. Molecular Dynamic Simulations
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Xia, Q.; Akanbi, T.O.; Li, R.; Wang, B.; Yang, W.; Barrow, C.J. Lipase-catalysed synthesis of palm oil-omega-3 structured lipids. Food Funct. 2019, 10, 3142–3149. [Google Scholar] [CrossRef] [PubMed]
- Cui, R.; Xu, L.; Lan, D.; Yang, B.; Wang, Y. A novel sn-1,3 specific lipase from Janibacter sp. As catalysts for the high-yield synthesis of long-medium-long type structured triacylglycerols. Food Chem. 2022, 366, 130523. [Google Scholar] [CrossRef] [PubMed]
- Mota, D.A.; Rajan, D.; Heinzl, G.C.; Osório, N.M.; Gominho, J.; Krause, L.C.; Soares, C.M.F.; Nampoothiri, K.M.; Sukumaran, R.K.; Ferreira-Dias, S. Production of low-calorie structured lipids from spent coffee grounds or olive pomace crude oils catalyzed by immobilized lipase in magnetic nanoparticles. Bioresour. Technol. 2020, 307, 123223. [Google Scholar] [CrossRef]
- Guo, Y.; Cai, Z.; Xie, Y.; Ma, A.; Zhang, H.; Rao, P.; Wang, Q. Synthesis, physicochemical properties, and health aspects of structured lipids: A review. Compr. Rev. Food Sci. F 2020, 19, 759–800. [Google Scholar] [CrossRef]
- Matori, M.; Asahara, T.; Ota, Y. Positional specificity of microbial lipases. J. Ferment. Bioeng. 1991, 72, 397–398. [Google Scholar] [CrossRef]
- Rogalska, E.; Cudrey, C.; Ferrato, F.; Verger, R. Stereoselective hydrolysis of triglycerides by animal and microbial lipases. Chirality 1993, 5, 24–30. [Google Scholar] [CrossRef]
- Kovac, A.; Scheib, H.; Pleiss, J.; Schmid, R.D.; Paltauf, F. Molecular basis of lipase stereoselectivity. Eur. J. Lipid Sci. Technol. 2000, 102, 61–77. [Google Scholar] [CrossRef]
- Chen, H.; Meng, X.; Xu, X.; Liu, W.; Li, S. The molecular basis for lipase stereoselectivity. Appl. Microbiol. Biotechnol. 2018, 102, 3487–3495. [Google Scholar] [CrossRef]
- Wang, X.; Qin, X.; Li, D.; Yang, B.; Wang, Y. One-step synthesis of high-yield biodiesel from waste cooking oils by a novel and highly methanol-tolerant immobilized lipase. Bioresour. Technol. 2017, 235, 18–24. [Google Scholar] [CrossRef]
- Wang, X.; Li, D.; Qu, M.; Durrani, R.; Yang, B.; Wang, Y. Immobilized MAS1 lipase showed high esterification activity in the production of triacylglycerols with n-3 polyunsaturated fatty acids. Food Chem. 2017, 216, 260–267. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, D.; Wang, W.; Yang, B.; Wang, Y. A highly efficient immobilized MAS1 lipase for the glycerolysis reaction of n-3 PUFA-rich ethyl esters. J. Mol. Catal. B-Enzym. 2016, 134, 25–31. [Google Scholar] [CrossRef]
- Zhao, Z.; Hou, S.; Lan, D.; Wang, X.; Liu, J.; Khan, F.I.; Wang, Y. Crystal structure of a lipase from Streptomyces sp. strain W007—Implications for thermostability and regiospecificity. FEBS J. 2017, 284, 3506–3519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lang, D.A.; Mannesse, M.L.M.; De Haas, G.H.; Verheij, H.M.; Dijkstra, B.W. Structural basis of the chiral selectivity of Pseudomonas cepacia lipase. Eur. J. Biochem. 1998, 254, 333–340. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nardini, M.; Lang, D.A.; Liebeton, K.; Jaeger, K.E.; Dijkstra, B.W. Crystal structure of Pseudomonas aeruginosa lipase in the open conformation. J. Biol. Chem. 2000, 275, 31219–31225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matori, M.; Asahara, T.; Ota, Y. Reaction conditions influencing positional specificity index (PSI) of microbial lipases. J. Ferment. Bioeng. 1991, 72, 413–415. [Google Scholar] [CrossRef]
- Lee, W.J.; Zhang, Z.; Lai, O.M.; Tan, C.P.; Wang, Y. Diacylglycerol in food industry: Synthesis methods, functionalities, health benefits, potential risks and drawbacks. Trends Food Sci. Techol. 2020, 97, 114–125. [Google Scholar] [CrossRef]
- Ranganathan, S.V.; Narasimhan, S.L.; Muthukumar, K. An overview of enzymatic production of biodiesel. Bioresour. Technol. 2008, 99, 3975–3981. [Google Scholar] [CrossRef]
- Tang, Q.; Popowicz, G.M.; Wang, X.; Liu, J.; Pavlidis, I.V.; Wang, Y. Lipase-driven epoxidation is a two-stage synergistic process. ChemistrySelect 2016, 1, 836–839. [Google Scholar] [CrossRef]
- Wang, F.; Liu, S.; Mao, X.; Cui, R.; Yang, B.; Wang, Y. Crystal structure of a phospholipase d from the plant-associated bacteria Serratia plymuthica strain AS9 reveals a unique arrangement of catalytic pocket. Int. J. Mol. Sci. 2021, 22, 3219. [Google Scholar] [CrossRef]
- Yuan, D.; Lan, D.; Xin, R.; Yang, B.; Wang, Y. Screening and characterization of a thermostable lipase from marine Streptomyces sp. strain W007. Biotechnol. Appl. Biochem. 2016, 63, 41–50. [Google Scholar] [CrossRef]
- Yang, B.; Wang, Y.H.; Yang, J.G. Optimization of enzymatic degumming process for rapeseed oil. J. Am. Oil Chem. Soc. 2006, 83, 653–658. [Google Scholar] [CrossRef]
- Uppenberg, J.; Oehrner, N.; Norin, M.; Hult, K.; Kleywegt, G.J.; Patkar, S.; Waagen, V.; Anthonsen, T.; Jones, T.A. Crystallographic and molecular-modeling studies of lipase B from Candida antarctica reveal a stereospecificity pocket for secondary alcohols. Biochemistry 1995, 34, 16838–16851. [Google Scholar] [CrossRef] [PubMed]
- Eswar, N.; Webb, B.; Marti-Renom, M.A.; Madhusudhan, M.S.; Eramian, D.; Shen, M.-y.; Pieper, U.; Sali, A. Comparative protein structure modeling using modeller. Curr. Protoc. Bioinf. 2006, 15, 5.6.1–5.6.30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. Gromacs: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 2, 19–25. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.M.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef] [PubMed]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Petersson, G.A.; Nakatsuji, H.; et al. Gaussian, Inc., Wallingford CT. Available online: https://gaussian.com/g09citation (accessed on 1 November 2016).
- Yang, B.; Wang, H.; Song, W.; Chen, X.; Liu, J.; Luo, Q.; Liu, L. Engineering of the conformational dynamics of lipase to increase enantioselectivity. ACS Catal. 2017, 7, 7593–7599. [Google Scholar] [CrossRef]
- Zhao, Z.; Lan, D.; Tan, X.; Hollmann, F.; Bornscheuer, U.T.; Yang, B.; Wang, Y. How to break the Janus effect of H2O2 in biocatalysis? Understanding inactivation mechanisms to generate more robust enzymes. ACS Catal. 2019, 9, 2916–2921. [Google Scholar] [CrossRef] [Green Version]
- Olsson, M.H.M.; Søndergaard, C.R.; Rostkowski, M.; Jensen, J.H. Propka3: Consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 2011, 7, 525–537. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Cornell, W.D.; Cieplak, P.; Bayly, C.I.; Gould, I.R.; Merz, K.M.; Ferguson, D.M.; Spellmeyer, D.C.; Fox, T.; Caldwell, J.W.; Kollman, P.A. A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 1995, 117, 5179–5197. [Google Scholar] [CrossRef] [Green Version]
- Hornak, V.; Abel, R.; Okur, A.; Strockbine, B.; Roitberg, A.; Simmerling, C. Comparison of multiple amber force fields and development of improved protein backbone parameters. Proteins Struct. Funct. Bioinf. 2006, 65, 712–725. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.M.; Cieplak, P.; Kollman, P.A. Geometrical and electronic structure variability of the sugar-phosphate backbone in nucleic acids. J. Comput. Chem. 2000, 21, 1049–1074. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, Z.; Chen, S.; Xu, L.; Cai, J.; Wang, J.; Wang, Y. Structural Basis for the Regiospecificity of a Lipase from Streptomyces sp. W007. Int. J. Mol. Sci. 2022, 23, 5822. https://doi.org/10.3390/ijms23105822
Zhao Z, Chen S, Xu L, Cai J, Wang J, Wang Y. Structural Basis for the Regiospecificity of a Lipase from Streptomyces sp. W007. International Journal of Molecular Sciences. 2022; 23(10):5822. https://doi.org/10.3390/ijms23105822
Chicago/Turabian StyleZhao, Zexin, Siyue Chen, Long Xu, Jun Cai, Jia Wang, and Yonghua Wang. 2022. "Structural Basis for the Regiospecificity of a Lipase from Streptomyces sp. W007" International Journal of Molecular Sciences 23, no. 10: 5822. https://doi.org/10.3390/ijms23105822
APA StyleZhao, Z., Chen, S., Xu, L., Cai, J., Wang, J., & Wang, Y. (2022). Structural Basis for the Regiospecificity of a Lipase from Streptomyces sp. W007. International Journal of Molecular Sciences, 23(10), 5822. https://doi.org/10.3390/ijms23105822